Abstract
Defective DNA mismatch repair creates a strong mutator phenotype, recognized as microsatellite instability (MSI). Various next-generation sequencing-based methods for evaluating cancer MSI status have been established, and NGS-based studies have thoroughly described MSI-driven tumorigenesis. Accordingly, high-frequency MSI (MSI-H) has been detected in 81 tumor types, including those in which MSI was previously underrated. The findings have increased the use of immunotherapy, which is assumed to be efficient in tumors having a high mutation burden and/or neoantigen load. In MSI tumorigenesis, positively and negatively selected driver gene mutations have been characterized in colorectal cancers. Recent advancements in genome-wide studies of MSI-H cancers have developed novel diagnostic and therapeutic approaches, including CXCR2 inhibitor, a synthetic lethal therapy targeting the Werner gene and inhibition of nonsense-mediated mRNA decay. MSI is a predictive marker for chemotherapy as well as immunotherapy. Thus, analyses of MSI status and MSI-related alterations in cancers are clinically relevant. We present an update on MSI-driven tumorigenesis, focusing on a novel landscape of diagnostic and therapeutic approaches.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microsatellite instability (MSI) is characterized by alterations in the genome-wide microsatellite repeats. MSI is observed in most cancers developed in patients with Lynch syndrome (Aaltonen et al. 1993; Ionov et al. 1993; Thibodeau et al. 1993), and in a subclass of sporadic cancers (Eso et al. 2020; Imai and Yamamoto 2008; Gelsomino et al. 2016; Yamamoto et al. 2012, 2014; Yamamoto and Imai 2015, 2019), in which it is primarily triggered by MLH1 methylation. DNA mismatch repair (MMR) genes are inactivated genetically or epigenetically, leading to increased frameshift mutations in various cancer-related genes, and resulting in tumor development. High frequency MSI (MSI-H) cancers have specific molecular, pathological, and clinical characteristics different from low-frequency MSI (MSI-L) or microsatellite stable (MSS) cancers, irrespective of tumor tissue origin.
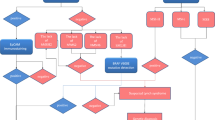
The use of next-generation sequencing (NGS) to find genetic changes in many tumor types is a practical method for investigating MSI-H tumorigenesis, and for discovering novel diagnostic biomarkers and therapeutic targets (Yamamoto and Imai 2019). We present an update on MSI-driven tumorigenesis, focusing on a novel landscape for diagnostic and therapeutic approaches (Fig. 1).
MSI analyses in various tumor types by NGS
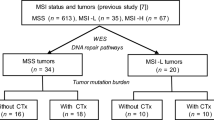
Evaluation of MSI status by polymerase chain reaction (PCR)-based assays and/or immunohistochemistry, has become a regular clinical practice for various types of advanced cancers (Ruiz-Bañobre and Goel 2019). From NGS-based techniques, alternative computational methods using targeted or whole exome sequencing data have been developed to evaluate MSI (Foltz et al. 2017; Hirotsu et al. 2020; Ruiz-Bañobre and Goel 2019; Wang and Liang 2018) (Fig. 2). These methods are shown to precisely evaluate MSI status (Ganesh et al. 2019; Ruiz-Bañobre and Goel 2019). Recently, Fujimoto et al. characterized the whole genome mutational landscape of microsatellites in 21 cancer types (Fujimoto et al. 2020). Moreover, deep learning could reportedly predict MSI directly from tissues sections stained with hematoxylin and eosin, facilitating common MSI screening in the near future (Kather et al. 2019).
Noninvasive analyses of MSI and tumor mutation burden (TMB) using cell-free DNA (cfDNA)
Using a pan-cancer gene panel that included selected microsatellite sequences, Georgiadis et al. (2019) established a cfDNA-based analysis of MSI and TMB. The sensitivities for MSI-H (n = 23) and TMB-High (n = 15) were 78% and 67%, respectively, and specificity was > 99% (n = 163). The findings showed the possibility of noninvasive pan-cancer detection and follow-up for cancer patients with MSI-H and/or TMB-High, presumed to have high sensitivity to immune checkpoint inhibitors (ICIs).
The origin of plasma circulating cfDNA variants determined by high-intensity sequencing
Using high-intensity sequencing, Razavi et al. (2019) determined the origin of cfDNA variants by analyzing cfDNA and corresponding leukocytes and tissue DNA in 124 metastatic cancer patients and 47 controls. The analyses showed high sensitivity and specificity, enabling detection of cancer-derived mutations and evaluation of TMB, MSI, mutation signatures and origins of somatic mutations detected in cfDNA. A large portion of cfDNA mutations (53% of cancer patients and 82% of controls) showed characteristics corresponding to clonal hematopoiesis. These findings suggest the importance of clonal hematopoiesis, and the need to further analyze corresponding leukocytes cfDNA sequencing for precise interpretation of variants.
Positively and negatively selected driver gene mutations in MSI-H CRCs
In MSI tumorigenesis, excessive genome instability causes not only positively selected mutations that contribute to tumor development, but also negatively selected mutations which have detrimental effects on cancer cells (Jonchere et al. 2018) (Fig. 3). The positively and negatively selected driver gene mutations identified, indicate that genome instability has a dual role in MSI-H CRC development.
Except for a small number of detrimental mutations detected in coding regions, negatively selected mutations were found mainly in long noncoding repeat sequences of the 5′ or 3′ UTR. Although around 10% of mutations were shown to change RNA expression, their functional effects need further analysis. Intriguingly, several negatively selected mutations were shown to upregulate tumor suppressive roles in MSI tumorigenesis, while others were shown to downregulate oncogenic roles. These results are in keeping with the contradictory activation or inactivation, respectively, in tumorigenesis. Results also emphasize that MSI in noncoding UTRs have an essential tumor suppressive effect in MSI tumorigenesis.
Ultra-hypermutated phenotype caused by DNA polymerase epsilon inactivation
For DNA replication fidelity, replicative DNA polymerases, exonucleolytic proofreading, and DNA MMR are necessary. Inactivating mutations of the polymerase epsilon (POLE) or delta (POLD1) result in an ultra-hypermutated phenotype that is represented by an excessive number of mutations (Gargiulo et al. 2016; Mittica et al. 2017; Nebot-Bral et al. 2017) (Figs. 1, 2). Owing to the high immunogenicity, patients with POLE-mutant tumors may also be sensitive to immunotherapy (Domingo et al. 2016; Ganesh et al. 2019) (Fig. 2).
Immune checkpoint immunotherapy
Frameshift mutation-enriched MSI-H tumors have been characterized by high Immunoscores, marked infiltration of immune cells (particularly mutation-specific CD8 + and CD4 + tumor-infiltrating lymphocytes (TILs) and macrophages), inhibitory PD-1/PD-L1 cells, and type I interferons-enriched microenvironment (Ganesh et al. 2019; Mlecnik et al. 2016). Accordingly, the outcomes of clinical trials that evaluated ICIs in patients with MSI-H tumors are encouraging (Ganesh et al. 2019; Le et al. 2015, 2017; O’Neil et al. 2017; Overman et al. 2017, 2018). The efficacy of the anti-PD-1 antibody was shown in 12 types of MSI-H cancers (Le et al. 2017). These results further suggest that a substantial portion of mutant-derived neoantigens render MSI-H tumors sensitive to ICIs, irrespective of tissue origin. Many clinical trials are presently underway to evaluate the efficiency of various ICIs in MSI-H metastatic CRC (mCRC) and other tumors (Ganesh et al. 2019).
KEYNOTE-164 (NCT02460198) study assessed the efficacy of pembrolizumab in previously treated MSI-H/deficient MMR (dMMR) mCRC. Registered patients with MSI-H/dMMR mCRC were 124 (61 for cohort A and 63 for cohort B). Overall response rate (ORR) was 33% for both cohorts. Thus, pembrolizumab was effective in patients with MSI-H/dMMR mCRC, and had manageable side effects (Le et al. 2020).
KEYNOTE-158 (NCT02628067) phase II study of pembrolizumab, was conducted in patients with formerly treated, unresectable or metastatic MSI-H/dMMR non-colorectal cancer. In total, 233 patients with 27 tumor types were registered. High-ranking tumor types were endometrial (21.0%), gastric (10.3%), cholangiocarcinoma (9.4%), and pancreatic (9.4%) cancers. Median follow-up period was 13.4 months and ORR was 34%. These results suggest that anti-PD-1 therapy using pembrolizumab is effective in patients with formerly treated advanced MSI-H/dMMR non-colorectal cancer (Marabelle et al. 2020).
In addition to PD-1 blocking, monospecific and bispecific antibodies, cellular therapies, cytokines, and vaccines that target various immune checkpoint components, macrophages and other molecules involved in innate immunity are actively investigated (Fig. 4).
Biomarkers for immune checkpoint immunotherapy
Various biomarkers predicting response to ICIs are presently being investigated. MSI-H accumulates frameshift mutations in cancer cells, leading to specific molecular traits, including higher tumor mutational burden/load (TMB/TML) (Innocenti et al. 2019), increased tumor neoantigen load (TNL), and marked TILs (Fig. 5). These alterations are correlated with increased sensitivity to ICIs (Chang et al. 2018), and hence high TMB and/or TNL is associated with a better response to ICIs (Panda et al. 2018). Moreover, compared with subclonal neoantigens, clonal neoantigens reportedly play more important roles in producing efficient anti-tumor immunity (McGranahan et al. 2016). The immunoscore is determined as the lymphocyte distribution in invasion margins and the center of tumors (Galon et al. 2013). Estimation of immune status by means of Immunoscore is reportedly helpful in predicting response to ICIs and tumor recurrence beyond MSI (Mlecnik et al. 2016).
Immunopathologic stratification of MSI-H CRC for ICIs
To classify MSI-H CRC patients, Llosa et al. integrated PD-L1 expression at the invasive front and histopathologic characteristics (percentage of extracellular mucin) to create a combined PD-L1 and mucin (CPM) score. This score distinguished which patients respond to ICIs. If confirmed in larger studies, MSI testing with CPM score may optimize immunotherapeutic interventions for MSI-H CRC patients (Llosa et al. 2019).
TMB is a predictive marker for response to ICIs in MSI-H mCRC
MSI is a marker for good response to ICIs. Patients with MSI-H mCRC treated with PD-1 had good disease control as well as promising progression-free survival (PFS). Nevertheless, ORR to pembrolizumab and nivolumab is variable and frequently less than half, indicating that further predictive markers are necessary. All 13 TMB-high patients and 3 of 9 TMB-low patients reportedly responded to ICIs, suggesting that TMB is a significant independent marker to predict response to ICIs in MSI-H mCRC patients. If confirmed in prospective studies, TMB may guide the order and/or combinations of ICIs in MSI-H mCRC (Schrock et al. 2019).
Genetic diversity of MSI-H tumors influences anti-PD-1 immunotherapy response
Despite tumor immunogenicity, patients with MSI-H tumors show variable responses to ICIs, and about half are resistant to treatment. It is reported that the degree of MSI and subsequent TMB partially affects variable responses to PD-1 blocking in MSI-H cancers. The accumulation of insertion–deletion (indel) mutational load was especially correlated with response. The genome-wide evaluation of MSI degree and TMB may be necessary to predict responses to anti-PD-1 blocking across MSI-H tumors (Mandal et al. 2019).
Immune escape in MSI-H cancer
Immune evasion mechanisms help MSI-H tumor cells evade the immune response despite high TMB/TNL and prominent TILs. This mechanism explains, at least in part, why not all MSI-H tumors show good response to ICIs. Since 30–50% of MSI-H tumors do not respond to ICIs, a predictive biomarker is clinically needed.
Genetic and/or epigenetic inactivation of the human leukocyte antigen (HLA) class I, β2-microglobulin (B2M), and antigen-processing genes are often found in MSI-H tumors (Fig. 4). Frameshift mutations are frequently detected in target coding microsatellites in these genes (Hirata et al. 2007). Markers predicting resistance to ICI therapy, including acquired mutations in the JAK1, JAK2 and B2M genes, were found in melanoma patients, but the role in other MSI-H tumors is not well defined (Ganesh et al. 2019) (Fig. 5). Ongoing clinical trials focusing on ICIs in MSI-H tumors will investigate immune evasion mechanisms to identify predictive biomarkers for primary and acquired resistance (Ganesh et al. 2019; Havel et al. 2019; Llosa et al. 2015; Ozcan et al. 2018).
Immune suppression and resistance to ICIs by KRAS-IRF2 axis in CRC
Oncogenic KRAS reportedly suppresses the immune system in CRC by inhibiting IRF2 expression, resulting in downregulation of interferon responsive genes, increased expression of CXCL3, enrollment of myeloid-derived suppressor cells, and consequent resistance to ICI therapy (Hänggi and Ruffell 2019; Liao et al. 2019).
Although MSI-H is a potent predictive marker of good response to ICIs, a subset of MSI-H CRC patients does not respond to anti-PD-1 therapy (O’Neil et al. 2017; Overman et al. 2017). IRF2 expression levels were positively correlated with response to anti-PD-1 therapy, and could be a novel predictive biomarker of response to therapy in MSI-H CRC. Accordingly, IRF2 reduction in MC38 cells, an MSI-H cell line that responds to ICIs, leads to resistance to ICIs. KRAS status is reportedly not related to anti-PD-1 therapy efficacy (Overman et al. 2018). Therefore, while KRAS/BRAF status is not a predictive marker of response to ICIs in MSI-H CRC, KRAS mutation may be a predictive marker of primary resistance to ICIs by IRF2 reduction in MSS CRC. Accordingly, the combination of ICIs with CXCR2 inhibitor may be promising in KRAS-mutated CRC patients (Katoh et al. 2013; Steele et al. 2016).
Identification of WRN as a synthetic lethal therapeutic target in MSI-H tumors
Inhibition of poly (ADP-ribose) polymerase (PARP) in patients with deficient homologous recombination DNA repair pathway is the first successful synthetic lethal cancer therapy. Other targets for synthetic lethal therapy have been studied (Chan et al. 2019; Kategaya et al. 2019; Lieb et al. 2019). Based on large-scale functional genomic screening of cancer cell lines, MSI-H cancer cells have been reported to be selectively susceptible to inactivation of WRN (Behan et al. 2019) (Fig. 6). WRN is a RecQ family DNA helicase, which is possibly inhibited using small molecules. Preclinical study of such small molecules in patients with MSI-H tumors is expected in the near future (Kawasaki et al. 2008).
Patient-derived xenografts (PDXs) and matched cell lines identify pharmacogenomic susceptibilities in CRC
Lazzari et al. (Lazzari et al. 2019) developed a 2D cell line (xeno-cell lines, XL) platform, which originated from PDXs of CRC with matched germline genomic DNA. Dependency on the WRN gene was evaluated in MSS, MSI-H, and MSI-like XLs using a functional approach with reverse genetics. MSI-H XLs were dependent on WRN gene expression, while MSS XLs were not. Interestingly, one MSS XL showing transcriptionally MSI-like characters was vulnerable to WRN loss. The XL platform is a useful method for validation of genes and studies to find new drug targets in CRC.
A comprehensive gastric cancer (GC) PDX collection captures cancer cell-intrinsic transcriptional MSI characters
Corso et al. (2019) developed a comprehensive GC PDX platform that could find and confirm novel drug targets and optimize therapeutic strategies. Moreover, transcriptomic analysis of GC PDXs identified a cancer cell intrinsic MSI signature that recognized a subset of MSS patients with MSI transcriptional characters and better prognosis. Thus, a multilevel GC PDX platform could identify an MSI gastric signature, contributing to precision medicine progress in GC.
Targeting nonsense-mediated mRNA decay (NMD) in MSI-H CRCs
mRNAs with a premature termination codon (PTC) are usually degraded by NMD system. NMD increases deficiencies caused by partially inactivated function of tumor suppressor genes with frameshift mutations (Bhuvanagiri et al. 2010; Popp and Maquat 2018; Usuki et al. 2004). Owing to the large number of mRNAs with PTC in MSI-H cancers, NMD can significantly inhibit the complete expression of tumor suppressor genes (Chan et al. 2009; El-Bchiri et al. 2005, 2008).
Compared with MSS CRC, MSI-H CRC reportedly expresses high levels of SMG1/6/7 and UPF1/2, which are crucial NMD activators (Bokhari et al. 2018). Many mRNAs with PTC are re-expressed by inhibition of the NMD activity. There are some mRNAs encoding mutated proteins with detrimental action against MSI carcinogenesis. Suppression of NMD by its inhibitor amlexanox, reportedly decreased MSI-H cancer growth in vivo, but not in MSS cancers. Amlexanox is useful for patients with recurring aphthous stomatitis (Ballal 2014). Thus, suppression of the oncogenic action of NMD appears to be an efficient approach for tailoring treatment of MSI-H CRC.
From the viewpoint of immunotherapy, NMD inhibitors receive the attention. NMD inhibitors are estimated to increase anti-tumor T cell immunity in MSI-H CRC by the cell surface presentation of abundant MSI-derived neoantigens. Whether NMD inhibitors synergistically function with ICIs in patients with MSI-H tumor needs further clarification.
Adaptive mutability of CRCs in response to targeted therapies
The occurrence of drug resistance restricts the efficacy of targeted therapies in human cancers. It is reported that homologous recombination repair and MMR genes were downregulated, and error-prone polymerases were upregulated by inhibition of epidermal growth factor receptor (EGFR)/BRAF in drug-resistant cells (Russo et al. 2019). Expression of MMR proteins was decreased in PDXs and cancer specimens through therapy. Thus, DNA damage, MSI, and increased mutability were induced by inhibition of EGFR/BRAF, suggesting that cancer cells manipulate adaptive mutability to escape therapeutic pressures.
Concluding remarks
MSI-H is observed in various tumor types, including those for which MSI-H was previously underrated. The recent advancements in genome-wide investigation of MSI-H cancers has led to the discovery of new diagnostic and therapeutic strategies, including CXCR2 inhibitor, a synthetic lethal therapy targeting WRN, and NMD inhibition. Combining genome and phenotype-based information with clinical data will facilitate the development of innovative diagnostic and therapeutic approaches in the era of NGS and precision medicine.
References
Aaltonen LA, Peltomaki P, Leach FS et al (1993) Clues to the pathogenesis of familial colorectal cancer. Science 260:812–816
Ballal VVJ (2014) Oral medicine: Amlexanox. Br Dent J 217:208
Behan FM, Iorio F, Picco G et al (2019) Prioritization of cancer therapeutic targets using CRISPR-Cas9 screens. Nature 568:511–516
Bhuvanagiri M, Schlitter AM, Hentze MW, Kulozik AE (2010) NMD: RNA biology meets human genetic medicine. Biochem J 430:365–377
Bokhari A, Jonchere V, Lagrange A et al (2018) Targeting nonsense-mediated mRNA decay in colorectal cancers with microsatellite instability. Oncogenesis 7:70
Chan WK, Bhalla AD, Le Hir H et al (2009) A UPF3-mediated regulatory switch that maintains RNA surveillance. Nat Struct Mol Biol 16:747–753
Chan EM, Shibue T, McFarland JM et al (2019) WRN helicase is a synthetic lethal target in microsatellite unstable cancers. Nature 568:551–556
Chang L, Chang M, Chang HM, Chang F (2018) Microsatellite instability: a predictive biomarker for cancer immunotherapy. Appl Immunohistochem Mol Morphol 26:e15–e21
Corso S, Isella C, Bellomo SE et al (2019) A comprehensive PDX gastric cancer collection captures cancer cell-intrinsic transcriptional MSI traits. Cancer Res 79:5884–5896
Domingo E, Freeman-Mills L, Rayner E et al (2016) Somatic POLE proofreading domain mutation, immune response, and prognosis in colorectal cancer: a retrospective, pooled biomarker study. Lancet Gastroenterol Hepatol 1:207e16
El-Bchiri J, Buhard O, Penard-Lacronique V et al (2005) Differential nonsense mediated decay of mutated mRNAs in mismatch repair deficient colorectal cancers. Hum Mol Genet 14:2435–2442
El-Bchiri J, Guilloux A, Dartigues P et al (2008) Nonsense-mediated mRNA decay impacts MSI-driven carcinogenesis and anti-tumor immunity in colorectal cancers. PLoS One 3:e2583
Eso Y, Shimizu T, Takeda H, Takai A, Marusawa H (2020) Microsatellite instability and immune checkpoint inhibitors: toward precision medicine against gastrointestinal and hepatobiliary cancers. J Gastroenterol 55:15–26
Foltz SM, Liang WW, Xie M, Ding L (2017) MIRMMR: binary classification of microsatellite instability using methylation and mutations. Bioinformatics 33:3799–3801
Fujimoto A, Fujita M, Hasegawa T, Wong JH, Maejima K, Oku-Sasaki A, Nakano K, Shiraishi Y, Miyano S, Yamamoto G, Akagi K, Imoto S, Nakagawa H (2020) Comprehensive analysis of indels in whole-genome microsatellite regions and microsatellite instability across 21 cancer types. Genome Res 30:334–346
Galon J, Angell HK, Bedognetti D, Marincola FM (2013) The continuum of cancer immunosurveillance: prognostic, predictive, and mechanistic signatures. Immunity 39:11–26
Ganesh K, Stadler ZK, Cercek A et al (2019) Immunotherapy in colorectal cancer: rationale, challenges and potential. Nat Rev Gastroenterol Hepatol 16:361–375
Gargiulo P, Della Pepa C, Berardi S et al (2016) Tumor genotype and immune microenvironment in POLE-ultramutated and MSI-hypermutated endometrial cancers: new candidates for checkpoint blockade immunotherapy? Cancer Treat Rev 48:61–68
Gelsomino F, Barbolini M, Spallanzani A, Pugliese G, Cascinu S (2016) The evolving role of microsatellite instability in colorectal cancer: a review. Cancer Treat Rev 51:19–26
Georgiadis A, Durham JN, Keefer LA et al (2019) Noninvasive detection of microsatellite instability and high tumor mutation burden in cancer patients treated with PD-1 blockade. Clin Cancer Res 25:7024–7034
Hänggi K, Ruffell B (2019) Oncogenic KRAS drives immune suppression in colorectal cancer. Cancer Cell 35:535–537
Havel JJ, Chowell D, Chan TA (2019) The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy. Nat Rev Cancer 19:133–150
Hirata T, Yamamoto H, Taniguchi H et al (2007) Characterization of the immune escape phenotype of human gastric cancers with and without high-frequency microsatellite instability. J Pathol 211:516–523
Hirotsu Y, Nagakubo Y, Amemiya K et al (2020) Microsatellite instability status is determined by targeted sequencing with MSIcall in 25 cancer types. Clin Chim Acta 502:207–213
Imai K, Yamamoto H (2008) Carcinogenesis and microsatellite instability: the interrelationship between genetics and epigenetics. Carcinogenesis 29:673–680
Innocenti F, Ou FS, Qu X et al (2019) Mutational analysis of patients with colorectal cancer in CALGB/SWOG 80405 identifies new roles of microsatellite instability and tumor mutational burden for patient outcome. J Clin Oncol 37:1217–1227
Ionov Y, Peinado MA, Malkhosyan S, Shibata D, Perucho M (1993) Ubiquitous somatic mutations in simple repeated sequences reveal a new mechanism for colonic carcinogenesis. Nature 363:558–561
Jonchere V, Marisa L, Greene M et al (2018) Identification of positively and negatively selected driver gene mutations associated with colorectal cancer with microsatellite instability. Cell Mol Gastroenterol Hepatol 6:277–300
Kategaya L, Perumal SK, Hager JH, Belmont LD (2019) Werner syndrome helicase is required for the survival of cancer cells with microsatellite instability. iScience 13:488–497
Kather JN, Pearson AT, Halama N et al (2019) Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat Med 25:1054–1056
Katoh H, Wang D, Daikoku T et al (2013) CXCR2-expressing myeloid-derived suppressor cells are essential to promote colitis-associated tumorigenesis. Cancer Cell 24:631–644
Kawasaki T, Ohnishi M, Suemoto Y et al (2008) WRN promoter methylation possibly connects mucinous differentiation, microsatellite instability and CpG island methylator phenotype in colorectal cancer. Mod Pathol 21:150–158
Lazzari L, Corti G, Picco G et al (2019) Patient-derived xenografts and matched cell lines identify pharmacogenomic vulnerabilities in colorectal cancer. Clin Cancer Res 25:6243–6259
Le DT, Uram JN, Wang H et al (2015) PD-1 blockade in tumors with mismatch-repair deficiency. N Engl J Med 372:2509–2520
Le DT, Durham JN, Smith KN et al (2017) Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357:409–413
Le DT, Kim TW, Van Cutsem E et al (2020) Phase II open-label study of pembrolizumab in treatment-refractory, microsatellite instability-jigh/mismatch repair-deficient metastatic colorectal cancer: KEYNOTE-164. J Clin Oncol 38:11–19
Liao W, Overman MJ, Boutin AT et al (2019) KRAS-IRF2 axis drives immune suppression and immune therapy resistance in colorectal cancer. Cancer Cell 35:559–572
Lieb S, Blaha-Ostermann S, Kamper E et al (2019) Werner syndrome helicase is a selective vulnerability of microsatellite instability-high tumor cells. Elife. 8:e43333
Llosa NJ, Cruise M, Tam A et al (2015) The vigorous immune microenvironment of microsatellite instable colon cancer is balanced by multiple counter-inhibitory checkpoints. Cancer Discov 5:43–51
Llosa NJ, Luber B, Siegel N et al (2019) Immunopathologic stratification of colorectal cancer for checkpoint blockade immunotherapy. Cancer Immunol Res 7:1574–1579
Mandal R, Samstein RM, Lee KW et al (2019) Genetic diversity of tumors with mismatch repair deficiency influences anti-PD-1 immunotherapy response. Science 364:485–491
Marabelle A, Le DT, Ascierto PA et al (2020) Efficacy of pembrolizumab in patients with noncolorectal high microsatellite instability/mismatch repair-deficient cancer: results from the phase II KEYNOTE-158 study. J Clin Oncol 38:1–10
McGranahan N, Furness AJS, Rosenthal R et al (2016) Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351:1463–1469
Mittica G, Ghisoni E, Giannone G et al (2017) Checkpoint inhibitors in endometrial cancer: preclinical rationale and clinical activity. Oncotarget 8:90532–90544
Mlecnik B, Bindea G, Angell HK et al (2016) Integrative analyses of colorectal cancer show immunoscore is a stronger predictor of patient survival than microsatellite instability. Immunity 44:698–711
Nebot-Bral L, Brandao D, Verlingue L et al (2017) Hypermutated tumours in the era of immunotherapy: the paradigm of personalised medicine. Eur J Cancer 84:290–303
O’Neil BH, Wallmark JM, Lorente D et al (2017) Safety and antitumor activity of the anti-PD-1 antibody pembrolizumab in patients with advanced colorectal carcinoma. PLoS One 12:e0189848
Overman MJ, McDermott R, Leach JL et al (2017) Nivolumab in patients with metastatic DNA mismatch repair-deficient or microsatellite instability-high colorectal cancer (CheckMate 142): an open-label, multicentre, phase 2 study. Lancet Oncol 18:1182–1191
Overman MJ, Lonardi S, Wong KYM et al (2018) Durable clinical benefit with nivolumab plus ipilimumab in DNA mismatch repair-deficient/microsatellite instability-high metastatic colorectal cancer. J Clin Oncol 36:773–779
Ozcan M, Janikovits J, von Knebel Doeberitz M, Kloor M (2018) Complex pattern of immune evasion in MSI colorectal cancer. Oncoimmunology 7:e1445453
Panda A, Mehnert JM, Hirshfield KM et al (2018) Immune activation and benefit from Avelumab in EBV-positive gastric cancer. J Natl Cancer Inst 110:316–320
Popp MW, Maquat LE (2018) Nonsense-mediated mRNA decay and cancer. Curr Opin Genet Dev 48:44–50
Razavi P, Li BT, Brown DN et al (2019) High-intensity sequencing reveals the sources of plasma circulating cell-free DNA variants. Nat Med 25:1928–1937
Ruiz-Bañobre J, Goel A (2019) DNA mismatch repair deficiency and immune checkpoint inhibitors in gastrointestinal cancers. Gastroenterology 156:890–903
Russo M, Crisafulli G, Sogari A et al (2019) Adaptive mutability of colorectal cancers in response to targeted therapies. Science 366:1473–1480
Schrock AB, Ouyang C, Sandhu J et al (2019) Tumor mutational burden is predictive of response to immune checkpoint inhibitors in MSI-high metastatic colorectal cancer. Ann Oncol 30:1096–1103
Steele CW, Karim SA, Leach JDG et al (2016) CXCR2 inhibition profoundly suppresses metastases and augments immunotherapy in pancre- atic ductal adenocarcinoma. Cancer Cell 29:832–845
Thibodeau S, Bren G, Schaid D (1993) Microsatellite instability in cancer of the proximal colon. Science 260:816–819
Usuki F, Yamashita A, Higuchi I et al (2004) Inhibition of nonsense-mediated mRNA decay rescues the phenotype in Ullrich’s disease. Ann Neurol 55:740–744
Wang C, Liang C (2018) MSIpred: a python package for tumor microsatellite instability classification from tumor mutation annotation data using a support vector machine. Sci Rep 8:17546
Yamamoto H, Imai K (2015) Microsatellite instability: an update. Arch Toxicol 89:899–921
Yamamoto H, Imai K (2019) An updated review of microsatellite instability in the era of next-generation sequencing and precision medicine. Semin Oncol 46:261–270
Yamamoto H, Adachi Y, Taniguchi H et al (2012) The interrelationship between microsatellite instability and microRNA in gastrointestinal cancer. World J Gastroenterol 18:2745–2755
Yamamoto H, Watanabe Y, Maehata T et al (2014) An updated review of gastric cancer in the next-generation sequencing era: insights from bench to bedside and vice versa. World J Gastroenterol 20:3927–3937
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yamamoto, H., Watanabe, Y., Maehata, T. et al. Microsatellite instability in cancer: a novel landscape for diagnostic and therapeutic approach. Arch Toxicol 94, 3349–3357 (2020). https://doi.org/10.1007/s00204-020-02833-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-020-02833-z