Abstract
Recently, there has been growing interest in the characterization of native yeasts for their use in production of wines with regional characteristics. This study aimed to investigate Saccharomyces and non-Saccharomyces yeasts present in the spontaneous fermentation of Tannat and Marselan grape musts collected from Concordia (Entre Ríos, Argentina) over 2019, 2020, and 2021 vintages. The evolution of these fermentative processes was carried out by measuring total soluble solids, total acidity, volatile acidity, pH, ethanol concentration, and total carbon content. Isolated Saccharomyces and non-Saccharomyces yeasts were identified based on colony morphology in WL medium, 5.8S-ITS-RFLP analysis, and 26S rDNA D1/D2 gene sequencing. Two hundred and ten yeast colonies were isolated and identified as Pichia kudriavzevii, Saccharomyces cerevisiae, Hanseniaspora uvarum, Metschnikowia pulcherrima, Candida albicans, Candida parapsilosis, Pichia occidentalis, Pichia bruneiensis, Hanseniaspora opuntiae, Issatchenkia terricola, and Hanseniaspora vineae. P. kudriavzevii isolated from all vintages was associated with the spontaneous fermentation of grape musts from the Concordia region.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alcoholic fermentation is a complex process with many biochemical changes due to the activities of fermenting microorganisms, such as several yeast species, as well as external physical factors. Although ethanol and carbon dioxide are the main fermentation products, other compounds that influence beverage flavor and color are also produced. These products vary according to the sugar content of the raw material and yeast activity, though beverage composition may differ depending on their origin (Del Fresno et al. 2017).
During the spontaneous fermentation of grape musts, non-Saccharomyces yeasts develop first. They are naturally present in grapes, in greater numbers than Saccharomyces cerevisiae, and are adapted to the environment (Cray et al. 2013). Hanseniaspora, Pichia, Debaryomyces, Issatchenkia, Candida, and Metschnikowia stand out among the non-Saccharomyces genera (Jolly et al. 2014; Grangeteau et al. 2016; Padilla et al. 2016). Since S. cerevisiae is more tolerant to ethanol and is competitive to grow under such environmental conditions, it shows better adaptation and subsequent develops (Jolly et al. 2014; Vaudano et al. 2019). However, some non-Saccharomyces yeasts can survive until the end of fermentation because of their high ethanol resistance (Combina et al. 2005).
Until a few years ago, non-Saccharomyces yeasts were considered responsible for microbiological problems and wine defects because they were isolated from altered wines (Padilla et al. 2016). However, current researches recognize their fundamental role in winemaking processes because they provide distinctive characteristics in wines (Maturano et al. 2016; Barkhuizen et al. 2021; Cioch-Skoneczny et al. 2021; Drumonde-Neves et al. 2021). Although they are known to be poor fermenters because of their low tolerance to ethanol, many are being investigated for winemaking purposes (Varela and Borneman 2017; Martin et al. 2018; Drumonde-Neves et al. 2021). Some non-Saccharomyces yeasts can be used to reduce ethanol levels in wines (Mestre Furlani et al. 2017; Maturano et al. 2019), whereas others can degrade malic acid during malolactic fermentation (del Mónaco et al. 2014). Moreover, the use of non-Saccharomyces yeasts in single or mixed/sequential fermentations is a powerful tool for improving the fruity aromatic quality and complexity of wines, and thus, to achieve a better definition of the regional flavor style (Padilla et al. 2016; Del Fresno et al. 2017; Shi et al. 2019; Lai et al. 2022). There is a modern approach, supported by rigorous scientific research, to apply ‘multispecies’ wine ferments, specifically native S. cerevisiae and non-Saccharomyces species (Jolly et al. 2014; Martin et al. 2018; Shi et al. 2019). Consequently, the use of autochthonous yeast species requires isolation and characterization procedures as well as molecular techniques for their identification.
The study of the effects of non-Saccharomyces yeasts on vinification is a trending topic among researchers in different countries (Del Fresno et al. 2017; Cimini and Moresi 2022). In recent years, several studies have focused on the characterization of native yeasts involved in spontaneous fermentation, mainly to understand the ecology, physiology, biochemistry, and molecular biology of Saccharomyces and non-Saccharomyces species (Maturano et al. 2016; Raymond Eder et al. 2017; Mendoza et al. 2019; Raymond Eder and Rosa 2019; Shi et al. 2019; García-Béjar et al. 2021; Zhang et al. 2021).
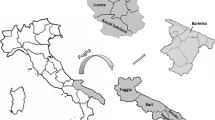
In Argentina, several researchers have studied the yeasts isolated from different grape varieties during spontaneous fermentation. Malbec varieties were analyzed in Cuyo (western Argentina) (Combina et al. 2005) and Patagonia regions (southern Argentina) (del Mónaco et al. 2014, 2016). In Córdoba (central Argentina), Isabella and Malbec varieties have been investigated (Raymond Eder et al. 2017; Raymond Eder and Rosa 2019), while studies in Malbec, Merlot, Syrah, and Torrontes from northern Argentina (Mendoza et al. 2019) have also been reported. However, no studies have been found on native yeasts from the east of the country (Entre Ríos, Argentina).
The province of Entre Ríos, located in eastern Argentina, is positioned as a new producer of vines and wines. Tannat and Marselan varieties are among the Vitis vinifera mainly cultivated in this region. The microbial communities of these grapes, particularly yeasts, have not yet been studied. Since no investigations have been reported, a special interest in their study has increased.
Many factors affect the diversity of microorganisms in grapes, such as climate, location, and grape physicochemical parameters (Combina et al. 2005; Vaudano et al. 2019; Sumby et al. 2021). This observation reinforces the interest in searching for wine yeast diversity in ecological niches alternative to traditional environments.
The aim of this study was to analyze the population dynamics of native yeasts (Saccharomyces and non-Saccharomyces) during spontaneous fermentation of Tannat and Marselan grape musts and their identification using both culture-based and molecular identification approaches.
Materials and methods
Grape sampling
Tannat and Marselan grape varieties were collected during 2019, 2020, and 2021 vintages, from a vineyard located in La Criolla (Concordia department, Entre Ríos province, latitude − 31°14′39″, longitude − 58°07′17″). Samples consisted of healthy grape bunches, not damaged, and randomly harvested at their optimal ripeness, across three vineyard lines. Bunches were placed in sterile bags, transported to the laboratory under cold storage, and maintained at 5 ± 2 °C until assay development.
Yeast isolation and cell count. Macroscopic and microscopic characteristics
Native yeast counts from Tannat and Marselan grapes were determined according to the methodology described below. 200 g of each variety were destemmed and aseptically crushed in a stomacher (IUL Instruments, Spain) for 20 s. Musts were supplemented with 85 mg/L sodium metabisulfite (Cicarelli, Argentina) and incubated at 25 ± 2 °C for 12 days to allow spontaneous fermentation. Sample aliquots were taken regularly, and suitable dilutions were plated in duplicate in order to yeast count. YPD agar (1% yeast extract (Britania, Argentina), 2% peptone (Britania), 2% dextrose (Biopack, Argentina), 1.5% agar (Britania) with 30 µg/mL chloramphenicol (Merck, Germany) (YPDC) was employed for total yeast counts while YPDC agar with 0.4 µg/mL cycloheximide (Merck) (YPDCL) to inhibit Saccharomyces, was used for non-Saccharomyces. Plates were incubated at 25 ± 2 °C for 72 h.
Yeast isolation was carried out on Wallerstein Laboratories (WL) differential nutrient agar (Oxoid, England) supplemented with 30 µg/mL chloramphenicol. Volumes of 0.1 mL from serially diluted samples were plated in duplicate. After 72 h of incubation at 25 ± 2 °C, yeast colonies showing different phenotypes (morphology and/or color) were isolated and cultured on WL agar to obtain a pure culture. From each grape variety and vintage, 27–41 representative colonies of all morphologies were selected. The microscopic characteristics (morphology, budding, etc.) were observed using an optical microscope (Leica, USA) at 100 × magnification. For long-term storage, yeast cells were inoculated into YPD broth, incubated for 48 h at 25 ± 2 °C, and then frozen at -80 °C using sterile glycerol (15% v/v) (Biopack) as a cryoprotective agent.
Monitoring of spontaneous alcoholic fermentation
The evolution of spontaneous fermentative processes in Tannat and Marselan grapes was carried out simultaneously with yeast counts. The fermenting musts were previously described, and the following physicochemical parameters were periodically determined over a 12-days period:
Total soluble solids
Refractometric method with a Hanna HI 96801 refractometer (Romania). Results were expressed as °Brix.
Total acidity
Potentiometric titration with sodium hydroxide (Cicarelli), according to MA-E-AS313-01: R2015 OIV technique (2020). Results were expressed as g tartaric acid/L.
Volatile acidity
Steam distillation (Jaulmes method), according to MA-E-AS313-02: R2015 OIV technique (2020). Results were expressed as g acetic acid/L.
pH
Potentiometric method with a BOECO BT-500 pHmeter (Germany), according to MA-E-AS313-15: R2011 OIV technique (2020).
Ethanol concentration
Enzymatic method (Boehringer Mannheim/R-Biopharm, Cat. N° 10,176,290,035, Germany). Results were expressed as % (v/v).
Total carbon concentration
Dumas method, dry digestion, and quantification with LECO CHN 628 according to OMA (2019). Results were expressed as g/100 g dry matter.
Molecular identification
Standard strain: S. cerevisiae ATCC 9763 provided by Administración Nacional de Laboratorios e Institutos de Salud (ANLIS) “Dr. Carlos G. Malbrán” (Argentina) was used as a standard strain for molecular assays. Species identification was carried out using the Yeast-ID.org database (https://www.yeast-id.org/) based on restriction analysis of the region including the gene codifying 5,8S rRNA and the transcribed intergenic regions ITS (5,8S-ITS).
DNA extraction
Each strain was cultured in test tubes containing 10 mL of YPD broth at 30 ± 1 ºC for 48 h. A volume of 1 mL was centrifuged at 2400 g for 10 min and DNA was extracted according to the CTAB method (Wilson 2001). DNA was visualized by electrophoresis on 1% (w/v) agarose gel (Genbiotech, Argentina) in 1 × TBE buffer at 100 V for 60 min. Gels were stained with 0.5 μg/mL ethidium bromide (Genbiotech) and visualized under UV light (Labnet International, Inc. USA). A 1 kb molecular weight marker was used (Genbiotech, Argentina). DNA was stored at − 20 ± 1 °C until use.
PCR amplification and analysis
All isolated strains were identified by PCR amplification of the 5.8-ITS rDNA region using ITS1 and ITS4 primers (White et al. 1990). DNA amplifications were carried out in 40 µL final volume containing 0.3 U GoTaq G2 DNA polymerase (Promega®, USA), 1 X PCR reaction buffer, 0.4 mM dNTP, 0.6 μM from each primer, 2 μL DNA (50–100 ng/µL). The PCR was performed on a Longgene MG96G (China) thermal cycler, under the following conditions: initial denaturation at 95 °C for 5 min, 35 cycles of denaturation at 95 °C for 1 min, annealing at 55 °C for 2 min, extension at 72 °C for 2 min, and a final extension step of 10 min at 72 °C. PCR amplification products were analyzed by electrophoresis on a 1.5% (w/v) agarose gel (Genbiotech) in 1 × TBE buffer, separated at 100 V for 100 min. Gels were stained with 0.5 μg/mL ethidium bromide (Genbiotech) and visualized under UV light. A 100 kb molecular weight marker was used (Genbiotech).
Restriction analysis
PCR products were digested with the restriction endonucleases CfoI, HaeIII, and HinfI (Pham et al. 2011) according to the manufacturer’s instructions (Promega®). Restriction fragments were separated on a 2% (w/v) agarose gel (Genbiotech) at a constant voltage of 80 V for 150 min and stained with ethidium bromide. A 25 pb molecular weight marker (Inbio Highway, Argentina) was used and species assignations were performed by comparison with profiles recorded in the Yeast-ID database.
26S rDNA D1/D2 gene sequencing and sequence analyses
In order to confirm the found species, some isolates, representative of each identified profile, were selected and the D1/D2 domain was sequenced from the 26S rDNA and amplified by NL1 (5′-GCATATCAATAAGCGGAGGAAAAG-3′) and NL4 (5′-GGTCCG TGTTTCAAGACGG-3′) primers (O´Donnel et al. 1993), using the PCR conditions described by Wang and Liu (2013). PCR products (600 bp) were sent for purification and subsequent sequencing (Macrogen Inc., Seoul, Korea) and the results were compared with those available in the NCBI GenBank nucleotide sequence database (http://www.ncbi.nlm.nih.gov/genbank/). Sequences from the representative strains were then deposited in the database with accession numbers.
Yeast diversity
The percentage distribution of yeast species isolated from Tannat and Marselan grapes was calculated by comparing the number of species detected with the total isolated yeasts per vintage and grape variety. It was calculated as follows: % = NS/NT × 100, where NS is the total strain per species and NT is the total isolated yeast.
Results and discussion
Dynamic of yeast populations in spontaneous fermentation of Tannat and Marselan grape musts
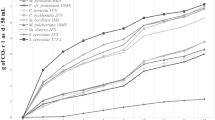
The population dynamics of S. cerevisiae and non-Saccharomyces yeasts for Tannat and Marselan grapes, during the 2019, 2020, and 2021 vintages, are shown in Fig. 1. The difference between the total yeast counts in YPDC agar and YPDCL agar indicates the relative contribution of S. cerevisiae. Initially, the total yeast count reached a population of 103 CFU/mL, similar to non-Saccharomyces yeast. These values are comparable to those reported by other authors (Combina et al. 2005; Maturano et al. 2016; Zabukovec et al. 2020). However, unexpectedly, for both varieties in the 2020 vintage, S. cerevisiae was observed at the beginning of fermentation. This was in accordance with Maturano et al. (2016), who found S. cerevisiae in large quantities (39%) in grape must. As fermentation time advanced, total yeast counts increased up to 106 CFU/mL, and simultaneously, S. cerevisiae showed a proliferation in their population to the detriment of non-Saccharomyces yeasts. Although non-Saccharomyces yeasts decreased in number during fermentation, they remained viable until the end of fermentation.
Spontaneous fermentation of grape musts. Population dynamics of Tannat yeasts from the 2019 vintage (a), 2020 vintage (c), and 2021 vintage (e): total yeasts (T–T), S. cerevisiae (T–S), and non-Saccharomyces yeasts (T-nS). Population dynamics of Marselan yeasts from the 2019 vintage (b), 2020 vintage (d), and 2021 vintage (f): total yeasts (M–T), S. cerevisiae (M–S), and non-Saccharomyces yeasts (M-nS)
Although only a few researchers have carried out S. cerevisiae and non-Saccharomyces yeast counts on YPDCL and YPDC agar during musts spontaneous fermentation, they agreed that only S. cerevisiae was identified at the final stages (Combina et al. 2005; Raymond Eder et al. 2017; Zabukovec et al. 2020). It is important to note that Pichia kudriavzevii (a non-Saccharomyces species) was also identified in this research work at the end of fermentation assays and in large counts, probably due to its ethanol resistance (experimentally demonstrated but not shown). Nieto-Sarabia et al. (2022) reported similar results.
Identification of yeast species
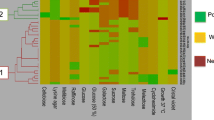
Yeast species from spontaneously fermenting musts of Tannat and Marselan grapes harvested in 2019, 2020, and 2021 vintages were isolated and identified. Two hundred and ten (210) colonies of native yeasts were isolated: 34, 35, and 41 from Tannat variety and 27, 38, and 35 from Marselan, respectively. Initially, yeast colonies were analyzed according to their morphology and color on WL nutrient agar in addition to microscopic observations (Table 1). Cavazza et al. (1992), Pallmann et al. (2001), Polizotto et al. (2016) and Li et al. (2018) reported that most yeast species typically found in grape musts fermentation could be differentiated according to their morphology and/or colony color on WL medium. However, it was observed that both characteristics in this medium were modified over time. As can be seen in Fig. 2 (VII a, b), C. parapsilosis initially formed pale green colonies with a white rim, glossy, and after 7 days, the color turned to emerald green.
Photographs of yeast colony morphotypes on WL nutrient agar. I: M. pulcherrima; II: I. terricola; III: P. occidentalis; IV: P. bruneiensis; V: P. kudriavzevii; VI: C. albicans; VII: C. parapsilosis, a: 72 h. after inoculation, b: 7 days after inoculation; VIII: H. opuntiae; IX: H. uvarum; X: H. vineae; XI: S. cerevisiae
According to Wang and Liu (2013), some I. terricola strains exhibited pale green colonies with white rims, surfaces with circular dents, and consistency of flour, whereas others were white with a hint of yellow, surface with circular dents, and consistency of flour. In the present study, colonies of these strains were green, black in the center, convex, and had an elevated dome. Nevertheless, WL agar is very useful for the preliminary differentiation of colonies prior to molecular identification.
To the best of our knowledge, macroscopic characteristics of some isolated yeasts such as H. opuntiae, P. occidentalis and P. bruneiensis grown on WL agar, have not been previously described (Table 1, Fig. 2). The first two species have been often found in grape musts and wines (Drumonde-Neves et al. 2021). Some authors have isolated P. bruneiensis from Hibiscus flowers (Sipiczki 2012) and apples (Liu et al. 2022) but nothing has been found in grapes, wines, or vineyards.
Since P. kudriavzevii exhibited white and opaque colonies, very similar to S. cerevisiae, its differentiation became too difficult (Fig. 2). Therefore, during the middle and final stages of yeast isolation, and according to the morphology observed on WL agar, spontaneous fermentation was thought to be dominated by S. cerevisiae. However, molecular identification assays also detected P. kudriavzevii. In addition, the growth of this species in tubes with broths showed a different behavior because it formed white agglomerated particles on the tube's wall, above the liquid surface (up to 2 cm). These characteristics have not been reported previously.
Isolated yeasts were subjected to molecular analysis using the PCR method of the internal transcribed spacer region (ITS), which comprises 5.8S rRNA and two flanking regions (ITS1 and ITS2) (White et al. 1990). The isolates showed different PCR product sizes ranging from 380 to 880 bp (Table 1). Subsequently, the products were digested with CfoI, HaeIII, and HinfI restriction enzymes (Pham et al. 2011). Digestion with each endonuclease yielded eleven different restriction profiles (Table 1). Isolated species were mostly differentiated based on these patterns. However, due to a high level of homology between groups VIII and IX, their differentiation was not possible with the aforementioned restriction enzymes. The cited patterns belong to three Hanseniaspora species: H. guillermondi, H uvarum, and H. opuntiae (Garofalo et al. 2016; Wang et al. 2019). Some authors have reported the possibility of using the DdeI and MboII restriction enzymes (Nisiotou et al. 2007; Wang et al. 2019). However, they were not available in the laboratory, therefore, identification by sequencing the 26S rDNA D1/D2 domain genes was necessary.
Similar diversity of non-Saccharomyces yeasts was observed in both varieties and also in all analysed vintages during the first stages of fermentation (Fig. 3). S. cerevisiae and P. kudriavzevii were the most dominant species. They contributed 40% and 35% of all isolates in both varieties, respectively, followed by H. uvarum (13%). Other non-Saccharomyces species were less frequently identified (Fig. 3). M. pulcherrima was isolated from both varieties, whereas P. occidentalis, P. bruneiensis, C. albicans, and C. parapsilosis were found only in Marselan, and H. opuntiae, I. terricola, and H. vineae, in Tannat grapes. Some of these yeast species (i.e., H. uvarum, H. vineae, C. albicans, P. opuntanie, C. parapsilosis, I. terricola, and P. kudriavzevii) have been widely described in grapes from other regions (Raymond Eder et al. 2017; Guaragnella et al. 2020; Zabukovec et al. 2020; Drumonde-Neves et al. 2021).
It is well known that S. cerevisiae is the dominant species in spontaneous fermentation of grape musts. However, only a few studies have recognized P. kudriavzevii as a fermenting species suitable for winemaking processes (Aponte and Giuseppe 2016; del Mónaco et al. 2016; Shi et al. 2019).
For all three vintages, H. uvarum was the third most abundant species in Tannat and Marselan varieties. Several studies have reported its presence in both grapes and musts (Maturano et al. 2016; Vaudano et al. 2019; Drumonde-Neves et al. 2021).
Sequencing analysis
All isolates identified by PCR–RFLP patterns were consistent with the sequencing results. However, some yeasts (genus Hanseniaspora) could not be differentiated in the ID Yeast database because they produced similar patterns to the assayed enzymes. Therefore, they can only be identified by sequencing.
Sequences obtained were uploaded to the NCBI GenBank nucleotide sequence database, and the following accession numbers were obtained: Group I, OQ553803 (99.79%); Group II, OQ520340 (98.03%); Group III, OQ553931(100%); Group IV, OQ559391 (98.14%); Group V, OQ520881 (99.31%), OQ553797 (99.64%); Group VI, OQ553801 (94.04%); Group VII, OQ521663 (99.82%); Group VIII, OQ521667 (97.21%), OQ520564 (99.65%); Group IX, OQ520337 (97.58%); Group X, OQ550975 (100%); Group XI, OQ520880 (95.76%), OQ553805 (98.76%), OQ559564 (98.57%), OQ521665 (99.48%). Query coverage ranged between 80 and 100%.
Spontaneous fermentation monitoring
Spontaneous fermentation of Tannat and Marselan grape musts was complete after 12 days. The results of the physicochemical analyses of these musts are shown in Tables 2 and 3.
Tannat grapes from the 2019 and 2021 vintages registered the highest initial total acidity (Table 2), whereas the lowest values were determined in the Marselan variety from the 2020 vintage (Table 3). At the end of the fermentation process, total acidity was slightly higher than that reported in other studies (Franco-Bañuelos et al. 2017; Piccardo and Zamora 2021). Despite this increase, these values were equally low. For other varieties, some authors have informed a decrease in this parameter under similar conditions (Raymond Eder et al. 2017; Raymond Eder and Rosa 2019).
The volatile acidity of wines is constituted by 99% acetic acid. During alcoholic fermentation, fermentative yeasts produce variable quantities of volatile acidity, depending on the yeast strain, sugar content, and temperature of fermentation. Amounts from 0.2 to 0.8 g/L are acceptable but should not exceed 1.3 g/L (Cioch-Skoneczny et al. 2021). As shown in Tables 2 and 3, the registered values were within this range.
A constant reduction in total carbon concentration was observed at the end of each assay and was attributed to carbon dioxide loss during alcoholic fermentation. Likewise, ethanol concentration increased during the same period, indicating the advancement of the fermentative process. Final alcohol concentrations resulted similar to the values reported in vinifications carried out with musts in analogous physicochemical (initial total soluble solids) and environmental conditions (Raymond Eder et al. 2017; Raymond Eder and Rosa 2019).
The evolution of spontaneous fermentation of Tannat and Marselan musts is shown in Fig. 4a and b. In general, after 8 days of fermentation, no variation in total soluble solids was observed thus indicating the end of the process.
In the 2019 vintage, total soluble solids at harvest were 19°Bx for both grape varieties. This value results extremely low if the aim is to obtain wines with an alcoholic graduation greater than 10% v/v. According to information provided by Instituto Nacional de Tecnología Agropecuaria (2019), it was verified that during the period among November 2018 and January 2019, the monthly average rainfall was much higher than the historical measure. This hydrological excess increases fruit size; they develop more aqueous, with poor sugar content and richer in acids, which could cause a delay in ripening (Ramos and Martínez De Toda 2022; Veselá et al. 2022). Therefore, it can be assumed that excessive rainfall could be the reason for the lower total soluble solids content.
Yeast diversity during spontaneous fermentation of grape musts
The contribution of yeast species during the different stages (initial, middle, and final) of spontaneous fermentation of Marselan and Tannat grape musts is shown in Fig. 5 and 6. A great variability in species was observed at the beginning of the process, except for the Tannat and Marselan 2021 vintage. As alcoholic fermentation progressed, some species disappeared, and only those that could adapt to the new environmental conditions (higher ethanol content) remained viable (Albergaria and Arneborg 2016).
From the 2019 and 2021 vintages, P. kudriavzevii was the main non-Saccharomyces species coexisting with S. cerevisiae at advanced stages during Tannat fermentation. It was also found in 2020 vintage musts, but in a low number. S. cerevisiae was not isolated during fermentation of Marselan grapes from the 2019 vintage whereas P. kudriavzevii was the dominant species. In contrast, in the 2020 vintage, S. cerevisiae was the only species isolated during the final stages of fermentation. The results for both varieties differ from previous reports that identified Aureobasidium, Hanseniaspora, Metschnikowia, Starmerella, Lachancea, and Candida as the dominant non-Saccharomyces genera in grape musts from different wine regions, while the genus Pichia was less frequently identified (Maturano et al. 2016; Raymond Eder et al. 2017; Vaudano et al. 2019; Mateus et al. 2020).
Ethanol production varied in Tannat and Marselan spontaneous fermentations over the three studied vintages. Considering the 2020 vintage, P. kudriavzevii was not found in Marselan but appeared in low quantities in Tannat. As shown in Table 3, this situation corresponds to a lower ethanol content. On the other hand, the highest ethanol concentration (9% v/v) was determined in Marselan musts from the 2021 vintage. When analyzing the species present at the end of fermentation, S. cerevisiae and P. kudriavzevii were isolated in almost the same proportion. This could indicate their role in ethanol production. Kaur et al. (2019) studied a P. kudriavzevii isolated from fruits and reported their potential to ferment sugars with ethanol production. In addition to high ethanol production, Nieto-Sarabia et al. (2022) showed that P. kudriavzevii had an ethanol tolerance superior to that of commercial S. cerevisiae. Aponte and Giuseppe (2016) reported that P. kudriavzevii isolated from Aglianico grapes produced 11% (v/v) ethanol.
In general, non-Saccharomyces yeasts have been reported to have lower fermentative capacities than S. cerevisiae (Polizotto et al. 2016). However, according to the results of this study (and others not shown), it can be stated that P. kudriavzevii could carry out grape musts fermentation with a good ethanol ratio.
In Argentina, P. kudriavzevii has been associated with spontaneous grape fermentation. One of these was isolated from the Isabella variety in the Córdoba province (Argentina) (Raymond Eder et al. 2017). In addition, del Mónaco et al. (2014) found it in Malbec grapes from Patagonia, Argentina, during the initial stages of spontaneous fermentation. In Zona Alta del Río Mendoza (Cuyo region), spontaneous fermentation of Malbec grapes has been studied, but the presence of this species has not been reported (Combina et al. 2005). On the other hand, Maturano et al. (2016) isolated it from Malbec grape must in a low proportion with respect to others. It is important to note that in all cases, the number of isolates was low, and P. kudriavzevii was not the main isolated species.
It is well known that S. cerevisiae belongs to the native grape microorganisms, it can be isolated from spontaneous fermentations and is responsible for the alcoholic fermentation during winemaking processes. However, the results found in Marselan grapes from the 2019 vintage showed that alcoholic fermentation was mainly carried out by P. kudriavzevii (Fig. 6). It can be seen that in middle and final stages over the three vintages (with the exception of Marselan 2020 because P. kudriavzevii was not isolated), S. cerevisiae and P. kudriavzevii were present, which indicated that spontaneous fermentation was carried out by both species, and sometimes P. kudriavzevii was the dominant one.
Conclusions
The yeast microbiota isolated and identified in this study, constitutes the first study of Tannat and Marselan varieties in Argentina. The evolution of Saccharomyces and non-Saccharomyces yeasts during must fermentation was investigated.
In addition, the morphologies of H. opuntiae, P. bruneiensis, and P. occidentalis on WL agar, which have not been previously reported, were described. Sequencing analysis confirmed the identification based on PCR–RFLP analysis. A similar diversity of yeast species was observed in both varieties.
In contrast, this work allowed the association of P. kudriavzevii (non-Saccharomyces yeast) with spontaneous fermentation of grape musts from the Concordia (Entre Ríos, Argentina) region. This species coexisted with S. cerevisiae at different stages of alcoholic fermentation. Moreover, P. kudriavzevii was the dominant species in some fermentations and produced good ethanol yield. Therefore, it can be concluded that this yeast species exhibits a high potential for further exploration since it seems a good candidate to formulate a mixed culture with S. cerevisiae. In this sense, further research such as oenological characterization (sulfite tolerance, production of aroma compounds and biogenic amine) is essential to validate its role.
References
Albergaria H, Arneborg N (2016) Dominance of Saccharomyces cerevisiae in alcoholic fermentation processes: role of physiological fitness and microbial interactions. Appl Microbiol Biotechnol 100:2035–2046. https://doi.org/10.1007/s00253-015-7255-0
Aponte M, Giuseppe B (2016) Potential role of yeast strains isolated from grapes in the production of Taurasi DOCG front. Microbiol 1:809. https://doi.org/10.3389/fmicb.2016.00809
Barkhuizen JH, Coetzee G, van Rensburg E, Görgens JF (2021) The effect of growth rate on the production and vitality of non-Saccharomyces wine yeast in aerobic fed-batch culture. Bioprocess Biosyst Eng 44(12):2655–2665. https://doi.org/10.1007/s00449-021-02634-3
Cavazza A, Grando M, Zini C (1992) Rilevazione della flora microbica di mosti e vini. Dossier Biotecnologie 9:1–4
Cimini A, Moresi M (2022) Research trends in the oenological and viticulture sectors. Aust J Grape Wine Res 28(3):475–491. https://doi.org/10.1111/ajgw.12546
Cioch-Skoneczny M, Satora P, Skoneczny S, Klimczak K (2021) Physicochemical characterization of wines produced using indigenous yeasts from cold climate grapes. Eur Food Res Technol 247(1):201–209. https://doi.org/10.1007/s00217-020-03618-5
Combina M, Elía A, Mercado L, Catania C, Ganga A, Martinez C (2005) Dynamics of indigenous yeast populations during spontaneous fermentation of wines from Mendoza. Argentina Int J Food Microbiol 99(3):237–243. https://doi.org/10.1016/j.ijfoodmicro.2004.08.017
Cray JA, Bell ANW, Bhaganna P, Mswaka AY, Timson DJ, Hallsworth JE (2013) The biology of habitat dominance; can microbes behave as weeds? Microb Biotechnol 6(5):453–492. https://doi.org/10.1111/1751-7915.12027
Del Fresno JM, Morata A, Loira I, Bañuelos MA, Escott C, Benito S, González Chamorro C, Suárez-Lepe JA (2017) Use of non-Saccharomyces in single-culture, mixed and sequential fermentation to improve red wine quality. Eur Food Res Technol 243(12):2175–2185. https://doi.org/10.1007/s00217-017-2920-4
del Mónaco SM, Barda NB, Rubio NC, Caballero AC (2014) Selection and characterization of a Patagonian Pichia kudriavzevii for wine deacidification. J Appl Microbiol 117(2):451–464. https://doi.org/10.1111/jam.12547
del Mónaco SM, Rodríguez ME, Lopes CA (2016) Pichia kudriavzevii as a representative yeast of North Patagonian winemaking terroir. Int J Food Microbiol 230:31–39. https://doi.org/10.1016/j.ijfoodmicro.2016.04.017
Drumonde-Neves J, Fernandes T, Lima T, Pais C, Franco-Duarte R (2021) Learning from 80 years of studies: A comprehensive catalogue of non-Saccharomyces yeasts associated with viticulture and winemaking. FEMS Yeast Res 21(3):1–3. https://doi.org/10.1093/femsyr/foab017
Franco-Bañuelos A, Contreras-Martínez CS, Carranza-Téllez J, Carranza-Concha J (2017) Total phenolic content and antioxidant capacity of non-native wine grapes grown in Zacatecas. Mexico Agrociencia 51(6):661–671
García-Béjar B, Árevalo-Villena M, Briones A (2021) Characterization of yeast population from unstudied natural sources in La Mancha region. J Appl Microbiol 130(3):650–664. https://doi.org/10.1111/jam.14795
Garofalo C, Russo P, Beneduce L, Massa S, Spano G, Capozzi V (2016) Non-Saccharomyces biodiversity in wine and the ‘microbial terroir’: a survey on Nero di Troia wine from the Apulian region. Italy Ann Microbiol 66:143–150. https://doi.org/10.1007/s13213-015-1090-5
Grangeteau C, Gerhards D, von Wallbrunn C, Alexandre H, Rousseaux S, Guilloux-Benatier M (2016) Persistence of two non-Saccharomyces yeasts (Hanseniaspora and Starmerella) in the cellar. Front Microbiol 7:268. https://doi.org/10.3389/fmicb.2016.00268
Guaragnella N, Chiara M, Capece A, Romano P, Pietrafesa R, Siesto G, Manzari C, Pesole G (2020) Genome sequencing and comparative analysis of three Hanseniaspora uvarum indigenous wine strains reveal remarkable biotechnological potential. Front Microbiol 10:1–14. https://doi.org/10.3389/fmicb.2019.03133
Instituto Nacional de Tecnología Agropecuaria (INTA) (2019) Informe sobre el efecto de exceso hídrico en los distintos cultivos en el área influencia de la EEA Concordia. Available from: https://inta.gob.ar/sites/default/files/inta-concordia_informe_exceso_hidrico_situacion_area_de_influencia_eea_inta_concordia_2019.pdf
Jolly NP, Varela C, Pretorius IS (2014) Not your ordinary yeast: non-Saccharomyces yeasts in wine production uncovered. FEMS Yeast Res 14(2):215–237. https://doi.org/10.1111/1567-1364.12111
Kaur S, Oberoi HS, Phutela R (2019) Isolation and characterization of a non-Saccharomyces yeast with improved functional characteristics for ethanol production. Int J Microbiol Res 26(4):42829. https://doi.org/10.9734/mrji/2018/v26i43007
Lai YT, Hsieh CW, Lo YC, Liou BK, Lin HW, Hou CY, Cheng KC (2022) Isolation and identification of aroma-producing non-Saccharomyces yeast strains and the enological characteristic comparison in wine making. LWT 154:112653. https://doi.org/10.1016/j.lwt.2021.112653
Li J, Hu W, Huang X, Xu Y (2018) Investigation of yeast population diversity and dynamics in spontaneous fermentation of Vidal blanc icewine by traditional culture-dependent and high-throughput sequencing methods. Food Res Int 112:66–77. https://doi.org/10.1016/j.foodres.2018.06.011
Liu L, Zhao PT, Hu CY, Tian D, Deng H, Meng YH (2022) Screening low-methanol and high-aroma produced yeasts for cider fermentation by transcriptive characterization. Front Microbiol 13:1042613. https://doi.org/10.3389/fmicb.2022.1042613
Martin V, Valera M, Medina K, Boido E, Carrau F (2018) Oenological impact of the Hanseniaspora/Kloeckera yeast genus on wines—a review. Ferment 4(3):76. https://doi.org/10.3390/fermentation4030076
Mateus D, Sousa S, Coimbra C, Rogerson FS, Simões J (2020) Identification and characterization of non-Saccharomyces species isolated from port wine spontaneous fermentations. Foods 9(2):120. https://doi.org/10.3390/foods9020120
Maturano YP, Mestre MV, Combina M, Toro ME, Vazquez F, Esteve-Zarzoso B (2016) Culture-dependent and independent techniques to monitor yeast species during cold soak carried out at different temperatures in winemaking. Int J Food Microbiol 21(237):142–149. https://doi.org/10.1016/j.ijfoodmicro.2016.08.013
Maturano YP, Mestre MV, Kuchen B, Toro ME, Mercado LA, Vazquez F, Combina M (2019) Optimization of fermentation-relevant factors: a strategy to reduce ethanol in red wine by sequential culture of native yeasts. Int J Food Microbiol 289:40–48. https://doi.org/10.1016/j.ijfoodmicro.2018.08.016
Mendoza LM, Vega-Lopez GA, Fernández de Ullivarri M, Raya RR (2019) Population and oenological characteristics of non-Saccharomyces yeasts associated with grapes of Northwestern Argentina. Arch Microbiol 201(2):235–244. https://doi.org/10.1007/s00203-018-1601-4
Mestre Furlani MV, Maturano YP, Combina M, Mercado LA, Toro ME, Vázquez F (2017) Selection of non-Saccharomyces yeasts to be used in grape musts with high alcoholic potential: a strategy to obtain wines with reduced ethanol content. FEMS Yeast Res 17(2):1–10. https://doi.org/10.1093/femsyr/fox010
Nieto-Sarabia VL, Ballinas-Cesatti CB, Melgar-Lalanne G, Cristiani-Urbina E, Morales-Barrera L (2022) Isolation, identification, and kinetic and thermodynamic characterization of a Pichia kudriavzevii yeast strain capable of fermentation. Food Bioprod Process 131:109–124. https://doi.org/10.1016/j.fbp.2021.10.013
Nisiotou AA, Spiropoulos AE, Nychas GJ (2007) Yeast community structures and dynamics in healthy and Botrytis-affected grape must fermentations. Appl Environ Microbiol 73(21):6705–6713. https://doi.org/10.1128/AEM.01279-07
O’Donnell K (1993) Fusarium and its near relatives. In: Reynolds DR, Taylor JW (eds) The fungal holomorph: mitotic, meiotic and pleomorphic speciation in fungal systematics. CAB International, Wallingford, UK, pp 225–233
Official Methods of Analysis of AOAC INTERNATIONAL (2019) 21st Eds, AOAC INTERNATIONAL, Gaithersburg, MD, USA, Official Method 920.70.
OIV Compendium of International Methods of Wine and Must Analysis (2020) Volume 1, Paris, France: International Organisation of Vine and Wine
Padilla B, Gil JV, Manzanares P (2016) Past and future of non-Saccharomyces yeasts: from spoilage microorganisms to biotechnological tools for improving wine aroma complexity. Front Microbiol 7(411):1–20. https://doi.org/10.3389/fmicb.2016.00411
Pallmann CL, Brown JA, Olineka TL, Cocolin L, Mills DA, Bisson LF (2001) Use of WL medium to profile native flora fermentations. Am J Enol Vitic 52(3):198–203
Pham T, Wimalasena T, Box WG, Koivuranta K, Storgårds E, Smart KA, Gibson BR (2011) Evaluation of ITS PCR and RFLP for differentiation and identification of brewing yeast and brewery “wild” yeast contaminants. J Inst Brew 117(4):556–568. https://doi.org/10.1002/j.2050-0416.2011.tb00504.x
Piccardo D, Zamora F (2021) González-Neves G (2021) Desarrollo y evaluación de alternativas tecnológicas para reducir el contenido de alcohol y el pH de vinos tintos. Memoria Investig Ing 20:24–33. https://doi.org/10.36561/ing.20.4
Polizzotto G, Barone E, Ponticello G, Fasciana T, Barbera D, Corona O, Amore G, Giammanco A, Oliva D (2016) Isolation, identification and oenological characterization of non-Saccharomyces yeasts in a Mediterranean island. Lett Appl Microbiol 63(2):131–138. https://doi.org/10.1111/lam.12599
Ramos MC, Martínez De Toda F (2022) Influence of weather conditions and projected climate change scenarios on the suitability of Vitis vinifera cv. Carignan in Rioja DOCa. Spain Int J Biometeorol 66:1067–1078. https://doi.org/10.1007/s00484-022-02258-6
Raymond Eder ML, Rosa AL (2019) Yeast diversity in Vitis non-vinifera ecosystems. Rev Argent Microbiol 51(3):278–283. https://doi.org/10.1016/j.ram.2018.09.004
Raymond Eder ML, Reynoso C, Lauret SC, Rosa AL (2017) Isolation and Identification of the indigenous yeast population during spontaneous fermentation of Isabella (Vitis labrusca L.) grape must. Front Microbiol. https://doi.org/10.3389/fmicb.2017.00532
Shi WK, Wang J, Chen FS, Zhang XY (2019) Effect of Issatchenkia terricola and Pichia kudriavzevii on wine flavor and quality through simultaneous and sequential co-fermentation with Saccharomyces cerevisiae. LWT. https://doi.org/10.1016/j.lwt.2019.108477
Sipiczki M (2012) Pichia bruneiensis sp. nov., a biofilm-producing dimorphic yeast species isolated from flowers in Borneo. Int J Syst Evol Microbiol 62:3099–3104. https://doi.org/10.1099/ijs.0.044974-0
Sumby KM, Caliani NS, Jiranek V (2021) Yeast diversity in the vineyard: how it is defined, measured and influenced by fungicides. Aust J Grape Wine Res 27(2):169–193. https://doi.org/10.1111/ajgw.12479
Varela C, Borneman AR (2017) Yeasts found in vineyards and wineries. Yeast 34(3):111–128. https://doi.org/10.1002/yea.3219
Vaudano E, Quinterno G, Costantini A, Pulcini L, Pessione E, Garcia-Moruno E (2019) Yeast distribution in Grignolino grapes growing in a new vineyard in Piedmont and the technological characterization of indigenous Saccharomyces spp. strains. Int J Food Microbiol 289:154–161. https://doi.org/10.1016/j.ijfoodmicro.2018.09.016
Veselá K, Severová L, Svoboda R (2022) The impact of temperature and precipitation change on the production of grapes in the Czech Republic. Sustainability. https://doi.org/10.3390/su14063202
Wang C, Liu Y (2013) Dynamic study of yeast species and Saccharomyces cerevisiae strains during the spontaneous fermentations of Muscat blanc in Jingyang. China Food Microbiol 33(2):172–177. https://doi.org/10.1016/j.fm.2012.09.014
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, New York, NY, USA, pp 315–22. https://doi.org/10.1016/B978-0-12-372180-8.50042-1
Wilson K (2001) Preparation of genomic DNA from bacteria. Curr Protoc Mol Biol 56:241–245. https://doi.org/10.1002/0471142727.mb0204s56
Zabukovec P, Čadež N, Čuš F (2020) Isolation and identification of indigenous wine yeasts and their use in alcoholic fermentation. Food Technol Biotechnol 58(3):337–347. https://doi.org/10.17113/ftb.58.03.20.6677
Zhang J, Plowman JE, Tian B, Clerens S, On SLW (2021) Application of MALDI-TOF analysis to reveal diversity and dynamics of winemaking yeast species in wild-fermented, organically produced, New Zealand Pinot Noir wine. Food Microbiol. 99:103824. https://doi.org/10.1016/j.fm.2021.103824
Acknowledgements
The authors would like to thank Administración Nacional de Laboratorios e Institutos de Salud (ANLIS) “Dr. Carlos G. Malbrán” (Argentina) for providing S. cerevisiae ATCC 9763. In addition, special thanks to Lic. Carolina Chacón for language assistance.
Funding
This work was funded by Secretaría de Ciencia y Técnica, Universidad Nacional de Entre Ríos (Argentina), PID UNER N° 8109, Res. C.S. N° 186/19.
Author information
Authors and Affiliations
Contributions
Liliana Gerard made the design of the work, analysed, interpreted the obtained results and contributed to the manuscript writing and its final revision. She also developed the molecular identification of the isolated yeasts. María Belén Corrado and Carina Soldá performed the tracking of the grapes spontaneous fermentation, isolated and characterised yeasts species.They also managed activities to annotate (produce metadata), scrubbed data and maintained research data for initial and subsequent use. Cristina Davies managed and coordinated the responsibility for the activity planning and execution. She also made the manuscript writing, final critical revision and approval of the version to be published. María Gabriela Dalzotto contributed to the manuscript writing, editing and final revision. Sofía Esteche contributed to developing the molecular identification of the isolated yeasts. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by Yusuf Akhter.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gerard, L.M., Corrado, M.B., Davies, C.V. et al. Isolation and identification of native yeasts from the spontaneous fermentation of grape musts. Arch Microbiol 205, 302 (2023). https://doi.org/10.1007/s00203-023-03646-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-023-03646-1