Abstract
Salmonella Typhimurium (STM) is one of the most important food-borne bacteria that seriously harms livestock and human beings, which is capable of regulating the expression of its own genes in a variety of ways to adapt to a wide variety of adverse environmental stresses. To understand the regulatory roles of sRNA STnc1480 on the capability of STM, the STnc1480 gene-deficient strain △STnc1480 and its complement strain △STnc1480/STnc1480 were generated, and the impacts of STnc1480 gene deficiency on the capability of responding to different environmental stresses, biofilm(BF)formation and pathogenicity were analyzed, respectively. Then the target genes that were regulated by STnc1480 were also analyzed and explored. Compared with parent and complement strains, the deficiency of the STnc1480 gene significantly reduced the BF formation. Moreover, the capacities of adhesion and invasiveness of the △STnc1480 strain to macrophages were also significantly reduced, while the LD50 in mice was significantly increased. The bacterial loads in liver and spleen were significantly reduced, and the pathological damage was alleviated. It was confirmed that the STnc1480 could be complementary to the 5’-UTR (-52 to -71 bases) region of lpfA mRNA. The bacterial dual-plasmid reporting system confirmed that STnc1480 was capable of interacting with the mRNA of the lpfA gene, suggesting that STnc1480 can regulate the 5’-UTR of the lpfA mRNA at post-transcription level to reduce the expression of the bacterial fimbria, thus reducing the BF formation and pathogenicity of STM.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salmonella Typhimurium (STM) is a Gram-negative intracellular bacterium with a wide range of host, which is widely distributed in the digestive tracts of livestock, birds and human (Fàbrega and Vila 2013; Majowicz et al. 2010). As one of important food-borne pathogens, STM can infect human through the contaminated meat, milk, eggs, and other animal-derived foods, resulting in human acute gastroenteritis, which has posed a serious threat to global food hygiene and safety. In the recent years, food poisoning caused by STM has occurred frequently in many countries, especially in the developing countries, causing serious harm to human health (Senior 2009).
Small RNA (sRNA) is non-coding RNA with about 50–400 nucleotides (nt) in length (Fröhlich and Vogel 2009). They are widely found in the genomes of many bacterial species (Harfouche et al. 2015). The currently available studies have found that sRNA is involved in regulation of the expression of certain genes of bacteria in adverse stress environments, and play an important role in responding to the environmental stresses, community effect, and the pathogenicity of bacteria(Fröhlich and Vogel 2009; Wagner and Romby 2015). A number of studies have confirmed that bacterial sRNA can perform a variety of biological functions by binding to mRNAs of their target genes to adapt to the environmental changes by regulating them at the post-transcriptional level (Papenfort and Vogel 2009).
Recently, more than 100 non-coding-sRNAs have been found and identified in Salmonella (Kröger et al. 2012). However, the biological functions of most of these sRNAs are not yet known. A few studies have revealed that the expression level of sRNA STnc1480 with 395 nucleotides in length was significantly increased during the infection process of STM into macrophages (Kröger et al. 2012; Srikumar et al. 2015). However, the regulatory roles of STnc1480 on the response of STM to environmental stress, biofilm (BF) formation, and pathogenicity of STM remain unclear. Therefore, the main purpose of this study was to understand the regulatory roles of STnc1480 on environmental stress, BF formation, and pathogenicity in STM. Here, we constructed and analyzed the response to environmental stresses, BF formation, and pathogenicity of the STnc1480 gene-deficient strain and complement strain, and identified the target genes regulated by sRNA STnc1480, aiming to provide new insights into the molecular mechanism of sRNA regulation in STM.
Materials and methods
Design of the primers
According to the STM SL1344 (accession number: FQ312003.1) sequence and pKD3 (AY048742.1) plasmid sequence deposited in the GeneBank, we designed the specific primers using Primer 5.0 software (Premier Inc., Canada), and synthesized by Beijing Genomics Institute (China). All the primers used in this study are listed in Table 1.
Generation of STnc1480-deficient mutant strain and complement strains
The strains and plasmid used are listed in Table 2. STM Wild Type (WT) SL1344 strains were cultured with brain–heart immersion (BHI) broth (Hopebio, China) at 37 °C for 12–16 h. The gene sequence of STM STnc1480 was amplified with F1/R1 primers. The STnc1480-deficient mutant strain was constructed using λ-Red recombination technology (Datsenko and Wanner 2000). Briefly, chloramphenicillin-containing gene products were first amplified by PCR with F2/R2 primer pair, which were used to construct recombinant strain △: STnc1480:: cat. Then the STnc1480-deficient mutant strain was identified by PCR amplification with F1/R1 primer pair in combination with sequencing technology. The complement strain △STnc1480/STnc1480 was constructed using F3/R3 primer pair.
Environmental stresses assay
Briefly, the strains of SL1344, △STnc1480, and △STnc1480/STnc1480 were inoculated into 50 mL of BHI liquid medium, and cultured at 37 °C overnight; This was followed by inoculation of the bacteria on an LB agar plate overnight and observation of the individual colony sizes by continuous zoom microscopy. The bacterial optical density at 600 nm (OD600nm) value was adjusted to 0.5, the bacteria solution was inoculated at 1:100 ratio in the BHI liquid medium and cultured continuously at 37 °C, their OD600 values were measured every hour and their growth curves were plotted. At the same time and 37℃, the bacterial solutions were inoculated at a ratio of 1:100, respectively, to HCl-adjusted pH 4.8 BHI liquid medium, NaOH-adjusted pH 10 BHI liquid medium, and BHI liquid medium containing 2 M NaCl and 2 mM 30% H2O2. The OD600 values were measured every hour and their growth curves were plotted. The effects of STnc1480 gene deficiency on the responses of STM to different environmental stress conditions were examined and analyzed.
Biofilm formation assay
The overnight-cultured active bacterial solution was inoculated in 1:100 ratio into the BHI liquid medium and cultured up to its OD600nm of 0.2. Then 100 μL of bacteria solution was aspired and transferred to 96-well microplates. Each sample was divided into two groups and eight parallel repetitions were set for each group. BF of STM was measured according to methods reported in the literature (Kint et al. 2010). After being cultured for 22, 23, and 24 h, the differences in BF formation capability between SL1344, △STnc1480, and △STnc1480/STnc1480 strains were compared and the effects of STnc1480 gene deficiency on STM BF formation capacity were analyzed.
Cell adhesion, invasion, and intracellular survival assay
Briefly, mouse macrophages RAW264.7 were placed in culture incubator at 37 °C and 5% carbon dioxide (CO2) and cultured overnight to grow into a single layer with about 2 × 105 cells per well. The bacterial adhesion and invasion tests were performed according to the methods described in the literature with a slight modification (Ryan et al. 2018). After infection of 1 h, for the cell invasion test, the bacterial cells that did not enter the macrophages were killed with Dulbecco’s modified Eagle medium (DMEM) (Gibco, USA) containing gentamicin [(100 mg/mL) (BIOTOPPED, China)] at 37 °C. The remaining steps were the same as those described above. The adhesion rates and invasion rates were calculated. Intracellular survival tests were conducted with reference to the literature (Hölzer et al. 2009). At 2, 4, 6, 8, 10, and 12 h after culture, cultured solution was aspired, cleaned, and lysed. The bacterial colonies were counted. The test was repeated three times and three parallels were set each time.
Animal infections
One hundred and fourteen 6-week-old BALB/c mice were randomly divided into one control group and six different dilution of bacterial infection groups 2 × 102 cfu/mL, 2 × 103 cfu/mL, 2 × 104 cfu/mL, 2 × 105 cfu/mL, 2 × 106 cfu/mL, and 2 × 107 cfu/mL, respectively. STM SL1344, △STnc1480, and △STnc1480/STnc1480 strains were cultured in BHI liquid medium, respectively, and used to infect mice when the logarithmic phase OD600 was 0.58. The mice in each subgroup were intraperitoneally injected with 0.5 mL of serially diluted cultures and the control group was treated with the same volume of phosphate buffered saline (PBS) and observed continuously for 10 days, and the LD50 of the mice was calculated by the modified Kurt method (Angalabiri-Owei and Isirima 2014). Six-week-old mice were infected with 2 × 104 cfu/ mL SL1344, △STnc1480, and △STnc1480/STnc1480 strains, respectively. Mice were sacrificed on days 1, 3, 5, 7, and 9 after infection, and the amounts of bacteria in these organs were simultaneously measured (Kumawat et al. 2016). Meanwhile, the liver and spleen were collected and pathological tissue sections were prepared 5 days after injection and the histopathological changes were observed by optical microscopy (Olympus, Japan).
Prediction and screening of target genes regulated by STnc1480
TargetRNA2 online software was used to predict the target genes of STnc1480 (Kery et al. 2014). The STM SL1344 genome was selected and the STnc1480 RNA sequence was entered. The genes with longer bases being paired with STnc1480 were selected as the target genes.
RNA extraction and real-time quantitative PCR
Briefly, the bacteria were cultured to BF formation state, and then collected by centrifugation. Total RNA was extracted with TRIzol reagent (Invitrogen, USA). Then complementary DNA (cDNA) was synthesized using PrimeScript-RRT kit including gDNA Eraser (TaKaRa Bio, Inc., Japan). Briefly, using 16sF/16sR and lpfAF/lpfAR as primers, the transcription levels of the target genes were determined by real-time quantitative PCR (RT-qPCR) using the ABI7500 instrument (Applied Biosystems, USA). Using the 16sRNA gene as the reference gene, the relative transcription levels of the target genes were calculated by the 2−△△CT method (Livak and Schmittgen 2001). Each test was repeated three times.
The bacterial dual-plasmid reporting system experiment
The interaction between STnc1480 and its target genes was analyzed using the bacterial dual-plasmid reporter system. In brief, F4/R4 and F5/R5 primers were used to amplify the operon DNA sequences of STnc1480 and lpfA. Then the pUT18C-STnc1480 and pMR-LacZ-lpfA plasmids were constructed; the constructed recombinant plasmids were co-transfected into BTH101 E. coli competent cells via electroporation. The positive clones on a Kanr (100 mg/ mL) Ampr (100 mg/ 9 mL) (BIOTOPPED, China) double-resistant plate coated with X-gal (TaKaRa Bio, Inc., Japan) and isopropyl-beta-D-thiogalactopyranoside(IPTG) (Solarbio Science & Technology Co., Ltd., China) were screened out and were placed at 37 °C overnight, and the color change of the bacterial lawn was observed.
Statistical analysis of data
GraphPad Prism 5.0 software (https://www.graphpad.com/, USA) was used for data analysis, and t test was used to conduct data significance analysis. The value of P < 0.05 was considered significantly different, while P < 0.01 was considered extremely significant different.
Results
STnc1480 does not affect the growth of STM in different environments
Through PCR and sequencing confirmation, the deletion strain △STnc1480 and the complement strain △STnc1480/STnc1480 were obtained (Supplementary Fig. S1 and Supplementary Fig. S2), respectively. The growth rates and bacterial single colony of the SL1344, △STnc1480, and △STnc1480/STnc1480 at 37 °C, were not significantly different (P > 0.05) (Fig. 1A) and Supplementary Fig. S4). When being cultured under the condition of pH4.8, pH10, 2 M NaCl, and 2 mM 30% H2O2, the growth of △STnc1480 was not significantly different from those of the complement strain and the parent strain (P > 0.05) (Fig. 1B, C, D and E), suggesting that STnc1480 does not affect the growth of STM in different environments.
STnc1480 affects biofilm formation of STM
Cultured for 22, 23, and 24 h, respectively, the SL1344, △STnc1480, and △STnc1480/STnc1480 were all capable of forming BF (Fig. 2A). However, the BF-forming ability of △STnc1480 strain was significantly weakened compared to the others at the indicated time (P < 0.05) (Fig. 2B), suggesting that STnc1480 had a potential influence on BF formation of STM.
STnc1480 affects adherence, invasion, and intracellular survival of STM to cells
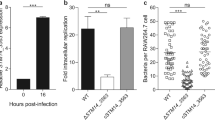
Compared with those of SL1344 and △STnc1480/STnc1480, the adhesion of △STnc1480 to RAW264.7 cells was extremely significantly weakened (P < 0.01) (Fig. 3A), and the invasion ability was also significantly weakened (P < 0.05) (Fig. 3B). At 6, 8, 10, and 12 h after infection, the quantity of △STnc1480 strains was significantly lower than those of SL1344 and △STnc1480/STnc1480 strains (P < 0.01) (Fig. 3C and Supplementary Fig. S5), indicating that STnc1480 affects the intracellular survival of STM.
Deleting STnc1480 attenuates virulence of STM in mice
The LD50 of SL1344, △STnc1480, and △STnc1480/STnc1480 were 3.6 × 104 cfu/ mL, 4 × 105 cfu/ mL, and 8.5 × 104 cfu/ mL, respectively (Table 3). The survival curves indicated that the pathogenicity of △STnc1480 group was significantly reduced (Fig. 4C). On the 5th, 7th, and 9th days after infection, as compared with those of SL1344 and △STnc1480/STnc1480 groups, the bacterial loads of △STnc1480 group in the liver and spleen of mice were significantly lower (P < 0.01) (Fig. 4A, B). Compared with those of SL1344 and △STnc1480/STnc1480 groups, the histopathological damages in liver, spleen, and small intestine of the mice infected with the △STnc1480 were alleviated (Fig. 5).
STnc1480 regulates lpfA expression
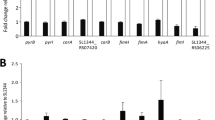
TargetRNA2 prediction of the target genes of STnc1480 led to the finding that the STnc1480 were complementarily paired with the 5'-UTR (− 52 to − 71 bases) of lpfA mRNA (Fig. 6A, B). In the state of BF formation, RT-qPCR analysis results showed that compared with SL1344 and △STnc1480/STnc1480, the transcription level of lpfA gene in △STnc1480 strain was significantly lower (P < 0.01) (Fig. 7), suggesting that the expression of lpfA gene may be regulated by the STnc1480.
Complementary pairing position of sRNA STnc1480 with mRNA of lpfA gene of STM. A The schematic represents the pairing region of STnc1480 sRNA with lpfA gene, B The − 35, − 10 region of lpfA gene promoter were underlined. The complementary sequences for STnc1480 targeting region are shown by black spots
STnc1480 can interact with target gene lpfA
The pMR-LacZ-lpfA and pUT18C-STnc1480 plasmids were constructed (Supplementary Fig. S3). Compared with the transformed pUT18C, pMR-LacZ and the co-transformed pUT18C and pMR-LacZ empty plasmids, the E. coli moss transformed with pUT18C-STnc1480 and pMR-LacZ-lpfA turned blue on the X-gal plate (Fig. 8), indicating that STnc1480 plays a positively regulatory role on the expression of lpfA gene.
Verification of interaction between STnc1480 and target gene lpfA using two plasmids co-expression system. A Bacterial lawn of BTH101 transformed by pUT18C, B bacterial lawn of BTH101 transformed by pMR-LacZ, C bacterial lawn of BTH101 co-transformed by pUT18C and pMR-LacZ, D bacterial lawn of BTH101 co-transformed by pUT18C-STnc1480 and pMR-LacZ-lpfA
Discussion
Many studies have shown that sRNA may regulate the expression of target genes in a variety of ways at the post-transcriptional levels. A large majority of sRNA regulatory factors block the entry of 30S ribosome by combining mRNA’s Shine–Dalgarno (SD) sequences or AUG start codon, thus preventing the start of translation. Alternatively, sRNA can also be paired with the target gene base to protect it from cell RNA degradation, thereby stabilizing the protein expression of the target gene (Bouvier et al. 2008; Opdyke et al. 2004). In addition, sRNA can be paired with the 5'-non-coding (UTR) regions of mRNA to open the secondary structures of the masking SD sequence and promote the binding of the SD sequence to the 30S ribosome, thus activating the expression of the target genes (Bobrovskyy and Vanderpool 2014).
In the recent years, many investigators have carried out research work on sRNAs of STM. To date, approximately 280 sRNAs expressed under various conditions have been detected in the STM LT2 strain (Kröger et al. 2013). Shabarinath et al. (Srikumar et al. 2015) found a significant increase in the expression level of 31 sRNAs through transcriptome studies of STM-infected mouse macrophages, while expression levels of some sRNAs were very low during extracellular growth, but were significantly higher in macrophages, suggesting that these sRNAs may be related to STM's pathogenicity and intracellular survival (Sittka et al. 2008).
It has been found that expression level of sRNA STnc1480 was significantly increased after STM-infected cells (Srikumar et al. 2015). However, the regulatory roles of STnc1480 and its target genes have not yet been studied. Here, we confirmed that the cell invasion ability and pathogenicity of the STnc1480-defficient strain were significantly reduced, suggesting that the STnc1480 was involved in regulation of pathogenicity in STM. Bioinformational predictions revealed that lpfA was one of the target genes regulated by STnc1480. It has been proved that lpfA was the main subunit of lpf fimbria, which helps STM form BF in the infected intestinal epithelia and increases ability of STM's intestinal colonization and pathogenicity to mice (Quan et al. 2019; Ledeboer et al. 2006). RT-qPCR analysis further confirmed that expression level of lpfA in the △STnc1480 was lower than that of SL1344, indicating that STnc1480 contributes to the expression regulation of lpfA gene, thus resulting in the reduced capacities of BF formation, cell invasion, and pathogenicity of STM.
To confirm the interaction between STnc1480 and 5’-UTR of lpfA mRNA, the bacterial dual-plasmid reporting system was employed in this study. By co-transformated the plasmid expressing sRNA and the plasmid containing their target mRNA-binding sites and the reporting gene, the effects of STnc1480 and lpfA on LacZ gene expression using X-gal as color agent were analyzed. The results revealed that STnc1480 could bind with the 5’UTR region of lpfA mRNA. Therefore, we speculated that the molecular mechanism of STnc1480 is to regulate the expression level of the lpfA gene by binding to mRNA of lpfA gene, protecting it from RNase degradation, and thus controlling the expression levels of the target genes, which posses similar regulatory mechanisms with the literature (Fröhlich et al. 2013).
In conclusion, this study revealed the regulatory roles of sRNA STnc1480 on BF formation and pathogenicity in STM, which provided new insights into the regulatory mechanism of sRNAs in STM.
References
Angalabiri-Owei BE, Isirima JC (2014) Evaluation of the lethal dose of the methanol extract of rhizophora racemosa leaf using Karbers method. Cell Pathol 2:65–68. https://doi.org/10.5897/AJCPATH14.009
Bobrovskyy M, Vanderpool CK (2014) The small RNA SgrS: roles in metabolism and pathogenesis of enteric bacteria. Front Cell Infect Microbiol 4:61–68. https://doi.org/10.3389/fcimb.2014.00061
Bouvier M, Sharma CM, Mika F, Nierhaus KH, Vogel J (2008) Small RNA binding to 5’ mRNA coding region inhibits translational initiation. Mol Cell 32:827–837. https://doi.org/10.1016/j.molcel.2008.10.027
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97:6640–6645. https://doi.org/10.1073/pnas.120163297
Fàbrega A, Vila J (2013) Salmonella enterica serovar Typhimurium skills to succeed in the host: virulence and regulation. Clin Microbiol Rev 26:308–341. https://doi.org/10.1128/CMR.00066-12
Fröhlich KS, Vogel J (2009) Activation of gene expression by small RNA. Curr Opin Microbiol 12:674–682. https://doi.org/10.1016/j.mib.2009.09.009
Fröhlich KS, Papenfort K, Fekete A, Vogel J (2013) A small RNA activates CFA synthase by isoform-specific mRNA stabilization. EMBO J 32:2963–2979. https://doi.org/10.1038/emboj.2013.222
Harfouche L, Haichar FZ, Achouak W (2015) Small regulatory RNAs and the fine-tuning of plant-bacteria interactions. New Phytol 206:98–106. https://doi.org/10.1111/nph.13195
Hölzer SU, Schlumberger MC, Jäckel D, Hensel M (2009) Effect of the O-antigen length of lipopolysaccharide on the functions of type III secretion systems in Salmonella enterica. Infect Immun 77:5458–5470. https://doi.org/10.1128/IAI.00871-09
Kery MB, Feldman M, Livny J, Tjaden B (2014) TargetRNA2: identifying targets of small regulatory RNAs in bacteria. Nucleic Acids Res 2:124–129. https://doi.org/10.1093/nar/gku317
Kint G, De Coster D, Marchal K, Vanderleyden J, De Keersmaecker SC (2010) The small regulatory RNA molecule MicA is involved in Salmonella enterica serovar Typhimurium biofilm formation. BMC Microbiol 10:276–284. https://doi.org/10.1186/1471-2180-10-276
Kröger C, Dillon SC, Cameron AD, Papenfort K, Sivasankaran SK, Hokamp K, Chao Y, Sittka A, Hébrard M, Händler K, Colgan A, Leekitcharoenphon P, Langridge GC, Lohan AJ, Loftus B, Lucchini S, Ussery DW, Dorman CJ, Thomson NR, Vogel J, Hinton JC (2012) The transcriptional landscape and small RNAs of Salmonella enterica serovar Typhimurium. Proc Natl Acad Sci USA 109:E1277-1286. https://doi.org/10.1073/pnas.1201061109
Kröger C, Colgan A, Srikumar S, Händler K, Sivasankaran SK, Hammarlöf DL, Canals R, Grissom JE, Conway T, Hokamp K, Hinton JC (2013) An infection-relevant transcriptomic compendium for Salmonella enterica Serovar typhimurium. Cell Host Microbe 14:683–695. https://doi.org/10.1016/j.chom.2013.11.010
Kumawat M, Pesingi PK, Agarwal RK, Goswami TK, Mahawar M (2016) Contribution of protein isoaspartate methyl transferase (PIMT) in the survival of Salmonella typhimurium under oxidative stress and virulence. Int J Med Microbiol 306:222–230. https://doi.org/10.1016/j.ijmm.2016.04.005
Ledeboer NA, Frye JG, McClelland M, Jones BD (2006) Salmonella enterica serovar Typhimurium requires the Lpf, Pef, and Tafi fimbriae for biofilm formation on HEp-2 tissue culture cells and chicken intestinal epithelium. Infect Immun 74:3156–3169. https://doi.org/10.1128/IAI.01428-05
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Majowicz SE, Musto J, Scallan E, Angulo FJ, Kirk M, O’Brien SJ, Jones TF, Fazil A, Hoekstra RM (2010) The global burden of nontyphoidal Salmonella gastroenteritis. Clin Infect Dis 50:882–889. https://doi.org/10.1086/650733
Opdyke JA, Kang JG, Storz G (2004) GadY, a small-RNA regulator of acid response genes in Escherichia coli. J Bacteriol 186:6698–6705. https://doi.org/10.1128/JB.186.20.6698-6705.2004
Papenfort K, Vogel J (2009) Multiple target regulation by small noncoding RNAs rewires gene expression at the post-transcriptional level. Res Microbiol 160:278–287. https://doi.org/10.1016/j.resmic.2009.03.004
Quan G, Xia P, Zhao J, Zhu C, Meng X, Yang Y, Wang Y, Tian Y, Ding X, Zhu G (2019) Fimbriae and related receptors for Salmonella enteritidis. Microb Pathog 126:357–362. https://doi.org/10.1016/j.micpath.2018.10.025
Ryan D, Mukherjee M, Nayak R, Dutta R, Suar M (2018) Biological and regulatory roles of acid-induced small RNA RyeC in Salmonella typhimurium. Biochimie 150:48–56. https://doi.org/10.1016/j.biochi.2018.05.001
Senior K (2009) Estimating the global burden of foodborne disease. Lancet Infect Dis 9:80–81. https://doi.org/10.1016/s1473-3099(09)70008-8
Sittka A, Lucchini S, Papenfort K, Sharma CM, Rolle K, Binnewies TT, Hinton JC, Vogel J (2008) Deep sequencing analysis of small noncoding RNA and mRNA targets of the global post-transcriptional regulator. Hfq Plos Genet 4:e1000163. https://doi.org/10.1371/journal.pgen.1000163
Srikumar S, Kröger C, Hébrard M, Colgan A, Owen SV, Sivasankaran SK, Cameron AD, Hokamp K, Hinton JC (2015) RNA-seq brings new insights to the intra-macrophage transcriptome of Salmonella typhimurium. PLoS Pathog 11:e1005262. https://doi.org/10.1371/journal.ppat.1005262
Wagner EGH, Romby P (2015) Small RNAs in bacteria and archaea: who they are, what they do, and how they do it. Adv Genet 90:133–208. https://doi.org/10.1016/bs.adgen.2015.05.001
Acknowledgements
The authors thank the field staff for providing the materials for this study.
Funding
This work was supported by grant from the national key research and development program (No. 2016YFD0500900), the Grant from Youth Science and Technology Innovation Leader of Xinjiang Production and Construction Corps (No. 2016BC001), and the Key Scientific and Technological Project in Agriculture of Xinjiang Production and Construction Corps (No. 2019GG026).
Author information
Authors and Affiliations
Contributions
LJ and QJ wrote the main manuscript text. LJ, Ning CC, LN, GY, JC, ZX, and ZX prepared figures. MQ, XX, CX, and QJ reviewed and edited. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interests
This manuscript has not been simultaneously submitted for publication in another journal and been approved by all co-authors. The authors declare that they do not have any conflict of interest.
Ethical approval
The experiments were carried out in accordance with the guidelines issued by the Ethical Committee of Shihezi University.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, J., Ning, C., Li, N. et al. The small RNA STnc1480 contributes to the regulation of biofilm formation and pathogenicity in Salmonella typhimurium. Arch Microbiol 204, 716 (2022). https://doi.org/10.1007/s00203-022-03331-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-022-03331-9