Abstract
This study aimed to compare the fungal rhizosphere communities of Rhazya stricta, Enneapogon desvauxii, Citrullus colocynthis, Senna italica, and Zygophyllum simplex, and the gut mycobiota of Poekilocerus bufonius (Orthoptera, Pyrgomorphidae, “Usherhopper”). A total of 164,485 fungal reads were observed from the five plant rhizospheres and Usherhopper gut. The highest reads were in S. italica rhizosphere (29,883 reads). Species richness in the P. bufonius gut was the highest among the six samples. Ascomycota was dominant in all samples, with the highest reads in E. desvauxii (26,734 reads) rhizosphere. Sordariomycetes and Dothideomycetes were the dominant classes detected with the highest abundance in C. colocynthis and E. desvauxii rhizospheres. Aspergillus and Ceratobasidium were the most abundant genera in the R. stricta rhizosphere, Fusarium and Penicillium in the E. desvauxii rhizosphere and P. bufonius gut, Ceratobasidium and Myrothecium in the C. colocynthis rhizosphere, Aspergillus and Fusarium in the S. italica rhizosphere, and Cochliobolus in the Z. simplex rhizosphere. Aspergillus terreus was the most abundant species in the R. stricta and S. italica rhizospheres, Fusarium sp. in E. desvauxii rhizosphere, Ceratobasidium sp. in C. colocynthis rhizosphere, Cochliobolus sp. in Z. simplex rhizosphere, and Penicillium sp. in P. bufonius gut. The phylogenetic results revealed the unclassified species were related closely to Ascomycota and the species in E. desvauxii, S. italica and Z. simplex rhizospheres were closely related, where the species in the P. bufonius gut, were closely related to the species in the R. stricta, and C. colocynthis rhizospheres.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The climate in Saudi Arabia is hot and dry throughout most of the country. Due to its arid climate, the flora of Saudi Arabia has been neglected like other Gulf countries in the Arab peninsula. Folk medicine, which is widely practiced, maintains an important part of Saudi Arabia’s health legacy (Al-Essa et al. 1998). Public interest in traditional medicine, especially in towns, has decreased with the integration of western modern medicine in Saudi Arabia.
Rhazya stricta Decne (Gentianales, Apocynaceae) is a widely distributed dogbane plant that is a dominant species in the rangeland of Saudi Arabia due to its allelopathic effects (Assaeed 1996; Allred 1968). To characterize the fungal diversity among the flowering plants, Aristolochia bracteolate (Piperales, Aristolochieae), Haplophyllum tuberculatum (Sapindales, Rutaceae), Nigella sativa (Ranunculales, Ranunculaceae), Rhazya stricta, Teucrium muscatense (Lamiales, Lamiaceae), Trigonella foenum-graecum (Fabales, Fabaceae), and Zataria multiflora (Lamiales, Lamiaceae) have been monitored (Elshafie et al. 2003). Twenty-four fungal species were isolated from the different parts of plants and there were no differences in significance among the herbal plant species mycoflora. The common fungal species isolated were Aspergillus niger and Penicillium sp. (Eurotiales, Trichocomaceae), followed by Rhizopus spp., and Aspergillus flavus (Elshafie et al. 2003).
The study of soil fungal flora associated with R. stricta showed that there were 14 species belonging to five genera, representing three orders: Penicillium (Eurotiales, Trichocomaceae), Aspergillus (Eurotiales, Trichocomaceae), Mucor and Rhizopus (Mucorales, Mucoraceae), and Fusarium (Hypocreales, Nectriaceae). Aspergilli were most frequently A. terreus, the most dominant among Aspergillus spp. These results suggest that soil fungal flora of the R. stricta plant is complex and diverse (Baeshen et al. 2014). In another study, fungal communities at Mecca old road showed that three phyla, Ascomycota, Basidiomycota and Chytridiomycota were recorded (Moussa et al. 2017). Metagenomic techniques have been utilized to study the fungal diversity of soil and rapidly characterize the biodiversity of many soils all over the world (Schmit and Mueller 2007; DeSalle et al. 2008; Hirsch et al. 2010; Blackwell 2011). Few metagenomic studies have been used to characterize microflora of soils in developing countries, or in Saudi Arabia (Yasir et al. 2015).
Rhazya stricta, Enneapogon desvauxii, Citrullus colocynthis, Senna italica, and Zygophyllum simplex were chosen because they are native plants that Bedouin use in folk medicine to treat many diseases. This study aimed to compare rhizosphere fungal communities of the five selected plants and the gut mycobiota of Poekilocerus bufonius, which is a type of grasshopper, known as Usherhopper that feeds on milkweed plants.
Methods
Area of study
The chosen study site was a Rhazya stricta community and public site (Hadda, Mecca-Jeddah road) at N21° 45′ 04.03", E39° 53′ 88.92". Field samples were taken from the rhizosphere of five plants (1A, Rhazya stricta; 2A, Enneapogon desvauxii; 3A, Citrullus colocynthis; 4A, Senna italica; soil5, Zygophyllum simplex) in the morning in June 2014 at 40 ºC. Soil cores were taken and pooled. Samples were stored in Falcon tubes (50 ml) at − 20 ºC until analysis. In addition, the gut of P. bufonius (Usherhopper DNA) was collected and stored.
DNA Extraction, library construction, and PCR
Next-generation sequencing followed Nilsson et al. (2008). Genomic DNA was extracted using MoBio Power kit from 5 g soil. A quality test was done, once the DNA samples were obtained, and then the whole qualified DNA was used to construct libraries. PCR products were converted into blunt ends. To add adapters, an ‘A’ nucleotide was added to each 3′ end. The fusion primer with dual adapters and index was used for PCR, and fragments that were too short were removed. Only the qualified library was used for sequencing. Illumina HiSeq/MiSeq encoded amplicons were used for monitoring the rhizosphere fungal diversity in the five plants and usherhopper gut. The primer pair ITS1 and ITS4 were used to amplify the qualified DNA (Moussa et al., 2017).
Bioinformatics analysis
Bioinformatics analysis for sequencing data was carried out. Filtration of the raw data was done to obtain clean reads, and tags were clustered at 97% sequence identity to an operational taxonomic unit (OTU), which assigned its taxonomy.
Processing of data and statistical analysis
Data statistics
The raw data were processed by an in-house procedure to obtain more reliable and accurate results (Douglas et al. 2014). For pooling of libraries, clean reads were assigned to samples. The results of data processing are listed in Table S1.
Paired-end reads
Both two paired-end reads overlapping regions were merged using FLASH v1.2.11 (Table S2) (Magoc and Salzberg 2011).
Analysis of community patterns
The sequences of the Internal Transcript Spacer [ITS1 and 4] were screened using the gold database (v20110519) for chimeras and UNITE (v20140703) separately for 18S rDNA sequences. The de novo chimera detection was done, and all tags were mapped to each representative OTU sequence. Sequences representing OTUs were taxonomically classified using 0.8 confidence values as a cutoff. Databases used for species annotation: 18S rDNA and ITS for the fungal community were used Silva V119 (Quast et al. 2013) and UNITE Version 6 (Abarenkov et al. 2010). OTUs that were not assigned to certain species were removed. A Venn diagram was drawn to the illustrated overlap of OTUs for each group. Different colors represent different samples or groups.
Results
OTU cluster and abundance
OTU statistics
Filtered tags were clustered into OTU at 97% similarity (Table 1). The results revealed that the P. bufonius gut sample was the richest and most diversity (162 OTUs), while the least diversity was obtained in the S. italica rhizosphere sample (66 OTUs), where the degree of sample diversity represents primarily the number per sample. In addition to the OTU count, the C. colocynthis rhizosphere sample had the highest tag sequence (29,883 bp), while the least tag sequences number was obtained in the P. bufonius gut sample (18,816 bp) (Table 1).
Venn chart of OTUs
A Venn diagram was used to visualize the number of common and unique OTUs in multiple samples. The microbiome core from different environments could be estimated from the OTU representative species. The overlapping area and/or intersections represents the set of OTUs shared with the counterpart. There were 15 shared OTUs among all five samples (Fig. 1). The rhizospheres of S. italica, C. colocynthis, E. desvauxii, R. stricta, and the gut of P. bufonius were composed of 10, 19, 26, 42, and 117 unique OTUs, respectively.
Principal component analysis (PCA)
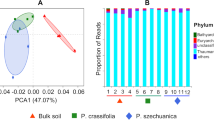
The fungal metagenomic differences among the five samples were reflected in a Principal Component Analysis (PCA) plot. The PCA plot compares OTU distribution among the different rhizosphere soil samples. The 1st ordination axis (35.39% of the total variance) separated the fungal community in the rhizosphere soil of R. stricta from those of the other samples, whereas the 2nd ordination axis (29.55% of the total variance) distinguished the communities of fungi in C. colocynthis rhizosphere soil and Z. simplex from the other samples (Fig. 2).
Sampling depth and species richness
The horizontal asymptote was reached in only the curves representing the rhizosphere samples of C. colocynthis and S. italica, suggesting sufficient sampling depth from these samples. The curves of R. stricta and E. desvauxii rhizosphere samples appeared to have reached the plateau (Figs. 3 and S1–S4).
Species composition and abundance
The total number of fungal reads was 164,485 from all six metagenomes. The number and ranking of fungal reads are shown in Table 2. The highest number was in the S. italica rhizosphere (29,883), and the lowest was in the gut of P. bufonius (18,816 reads).
Species annotation
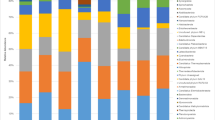
At phylum level, the rhizospheres of the five plants and the gut of P. bufonius were composed of only two fungal phyla (Ascomycota and Basidiomycota), with Ascomycota as the dominant phylum (Fig. 4). At the class level, the two classes Sordariomycetes and Dothideomycetes, which belong to Ascomycota, were dominant over all the classes of Ascomycota and Basidiomycota (Fig. 5). At the order level, the most abundant order was Eurotiales (Eurotiomycetes), followed by Hypocreales (Sordariomycetes) and Pleosporales (Dothideomycetes) (Table S4 and Fig. 6). At the family level, the dominant families detected were Trichocomaceae and Didmellaceae (Ascomycota), also family Ceratobasidiaceae, (Basidiomycota) (Table S5 and Fig. 7).
The most abundant genera and species were Aspergillus (Aspergillus terreus), and Ceratobasidium (Ceratobasidium sp.) in the R. stricta rhizosphere, while Fusarium (Fusarium sp.) and Penicillium (Penicillium sp.) in the rhizosphere of E. desvauxii and P. bufonius gut. Ceratobasidium (Ceratobasidium sp.), and Myrothecium (Myrothecium roridum) in the C. colocynthis rhizosphere, Aspergillus (Aspergillus terreus) and Fusarium (Fusarium sp.) in the S. italica rhizosphere, and Cochliobolus (Cochliobolus sp.) in the Z. simplex rhizosphere (Tables S6 & S7, and Figs. 8 & 9).
Phylogenetic analysis
From Figures S5A-F, the unclassified species were closely related to Ascomycota and the species in the rhizosphere of E. desvauxii, S. italica, and Z. simplex were closely related to each other, while that of P. bufonius gut (Usherhopper) and the rhizospheres of R. stricta and C. colocynthis were closely related to each other. From the phylogenetic tree at the genus level (Fig. 10), the genera Cladosporium, Arthrobotrys, Phoma, Kabatiella, Aspergillus, Myrothecium, and Candida were classified under Ascomycota, while the genus Jaminaea was classified under Basidiomycota.
Data availability
All sequence data were submitted to NCBI and have accession numbers: SRR6447717, SRR6447718, SRR6447719, SRR6447720, SRR6447721, and SRR6447722, with direct link http://www.ncbi.nlm.nih.gov/sra?term=SRP128163.
Discussion
The rapid evaluation of the genetic structure of complex communities in diverse environments was carried out using DNA fingerprinting (Muyzer and Smalla 1998). This permits the correlation of environmental disturbances and the extent of changes (Massol-Deya et al. 1997; Smit et al. 1997; Engelen et al. 1998; Schäfer et al. 2001). Fungal diversity, especially the mechanisms leading to their large-scale ecological and geographic ranges, have not been previously understood. Fungi associated with roots play a crucial role in the nutrition and health of plants (Verbruggen et al. 2012).
Microbiomes of the rhizosphere differ among plant species and in bulk soil. Pea plants had a stronger effect than wheat and oat on the rhizosphere, resulting in a great difference in the rhizosphere community. The eukaryotes relative abundance in the rhizospheres of pea and oat was higher than five-fold the abundance in bulk soil or the wheat rhizosphere (Turner et al. 2013). Our findings were in agreement, as the fungal community of R. stricta had 42 unique OTUs, E. dessvauxii had 26, C. colocynthis had 19 and S. italica had 10 unique OTUs.
Over the last decade, rapid development in microbiome research has shown that the microbial communities make significant contributions to immunity, reproduction, digestion, and other functions in insect hosts (Warnecke et al. 2007; Werren et al. 2008; Lehman et al. 2009; Fraune and Bosch 2010; Lee and Mazmanian 2010; Sharon et al. 2010). Our findings showed that the gut of P. bufonius was rich in saprophytic fungi like Penicillium spp., which may supply the insect with enzymes that aid in digestion, and antimicrobial peptides that boost insect immunity.
The root-associated fungal diversity was studied using 454 pyrosequencing and revealed 164 non-singleton OTUs (Yu et al. 2013). In this connection, our findings concluded that there are 15 non-singleton OTUs assigned to all five plants.
The richness of plant species diversity increases soil microbe diversity, particularly fungal communities (Hollister et al. 2010; Kernaghan 2005; Peay et al. 2013; Wu et al. 2013; Zak et al. 2003). Our study found that areas rich in R. stricta showed the highest number of fungi in its rhizosphere (42 OTUs). Fungal diversity is directly proportional to plant diversity. However, precipitation levels may have a greater effect (McGuire et al., 2012). Despite essential fluctuations due to different conditions in the host (Hongoh et al. 2005, 2006; Moran et al. 2008; Andert et al. 2010; Huang et al. 2013), a specific set of microbes is often associated with a specific host (Turnbaugh et al. 2007; Hamady and Knight 2009; Huse et al. 2012). The set of microbes can be considered as a host family-, genus-, or species-specific that may play important co-evolutionary roles (Hongoh 2010; Hongoh et al. 2005; Andert et al. 2010; Brucker and Bordenstein 2012). Our findings concluded that Aspergillus terreus and Ceratobasidium spp. were abundant in the R. stricta rhizosphere while Fusarium spp. and Penicillium spp. were abundant in the E. desvauxi rhizosphere and P. bufonius gut.
Baldrian et al. (2012) concluded that fungal sequences closely related to Basidiomycota represented 53.5% of the OTUs in forest soil, and sequences close to Ascomycota represented 41.1%. Geml et al. (2014) reported that Ascomycota was the dominant genus, representing 33.18% of the OTUs, followed by Basidiomycota with 22.73% of the OTUs, Glomeromycota with 5.29%, Mucoromycotina, incertae sedis with 1.94%, and Chytridiomycota with 0.53% of the OTUs, while 36.33% of the fungal OTUs were unidentified. In spruce plots, 28% of the soil fungal OTUs belonged to Basidiomycota and in spruce plots, Basidiomycota represented 65% of OTUs (Buée et al. 2009). The heterogeneity of the spatial distribution of populations is determined by the soil organic matter composition and host tree. Ascomycota was the most dominant of fungal community across Italy and France with OTU percentages ranging from 36.7% to 93% for samples from different ecosystems (Orgiazzi et al. 2013). As a result of our work, we noticed that OTUs were 51.7%, 90.1%, 81.4%, 60.2 and 85.5% belonging to Ascomycota in the rhizosphere of R. stricta, E. desvauxi, C. colocynthis, S. italica and Z. simplex, respectively, while the unidentified fungal OTUs were 47.5%, 9.9%, 17.3%, 39.8% and 14.4% in the rhizosphere of R. stricta, E. desvauxi, C. colocynthis, S. italica and Z. simplex, respectively.
The most abundant phylum at all four sites in the western coastal region of Saudi Arabia was Ascomycota followed by Basidiomycota (Moussa et al. 2017). Our results are consistent with this, in that Ascomycota was the most abundant phylum in all rhizosphere of the five plants studied.
Our previous work in the traditional isolation of fungi from the soil rhizosphere of R. stricta showed that there were 14 fungal species belonging to five genera: Aspergillus, Fusarium, Mucor, Penicillium, and Rhizopus, with Aspergillus spp. most frequent. (Baeshen et al. 2014). This study confirmed the previous findings using next-generation sequencing (NGS), where Aspergillus spp. were the most abundant in R. stricta rhizosphere.
In the western coastal region of Saudi Arabia, Sordariomycetes was observed predominantly at Asfan road, Thuwal and Khulais, while at Mecca old road, Pezizomycetes was dominant, but absent at Asfan road. At Mecca old road, Agaricomycetes was present only; while Tremellomycetes, Microbotryomycetes and Malasseizomycetes were found at Asfan road only (Moussa et al. 2017). Due to the dominance of Ascomycota, the dominant classes were Dothideomycetes in the rhizosphere of both E. desvauxi and S. italica, Eurotiomycetes in the rhizosphere of R. stricta, and Sordariomycetes in the rhizosphere of both C. colocynthis and Z. simplex.
Nine fungal species of genera Cladosporium, Alternaria, Exserohilum, Pyrenochaeta, Phoma, Neosartorya, Aspergillus, Penicillium, and Fusarium have been isolated from the plants roots at Ulleungdo Island (South Korea), which belonged to classes Eurotiomycetes, Dothideomycetes, and Sordariomycetes (Kim et al. 2013). Fourteen fungal species (Eurotium, Aspergillus, Penicillium, Cladosporium, Talaromyces, Exserohilum, Fusarium, Acremonium, Gibberella, Microsphaeropsis, Pestalotiopsis, Mucor, Zygorhynchus and Umbelopsis have been isolated from the plants roots at Dokdo Island (S. Korea/Japan); eleven of which belong to four classes (Dothideomycetes, Ascomycetes, Sordariomycetes, and Eurotiomycetes) and three species belonging to Mucoromycotina (You et al. 2011a; b, 2013). In our study, we found four fungal genera (Aspergillus, Ceratobasidium, Fusarium and Myrothecium) in the R. stricta rhizosphere, two genera (Fusarium and Penicillium) in the rhizosphere of E. desvauxii, three genera (Aspergillus, Ceratobasidium and Myrothecium) in the C. colocynthis rhizosphere, two genera (Aspergillus and Fusarium) in the S. italica rhizosphere, and three genera (Cochliobolus, Fusarium and Myrothecium) in the Z. simplex rhizosphere.
Ranjard et al. (2001) investigated 18 genera belonging to Basidiomycota, 78 genera belonging to Ascomycota, 3 genera belonging to Zygomycota, one genus belonging to Chytridiomycota, 3 genera belonging to Plasmodiophoromycota and one genus belonging to Oomycota. In our study 226 genera belonging to Ascomycota and 17 genera belonging to Basidimycota were investigated.
Conclusion
The most dominant phyla in all samples were Ascomycota and Basidiomycota. The fungal species found in the rhizosphere of the plants under study included saprophytic and pathogenic fungi only (Aspergillus, Fusarium, Penicillium, Ceratobasidium Myrothecium, Cochliobolus, Phoma, and Pestalotiopsis). Glomeromycota were absent. The P. bufonius gut mycoflora was composed of Fusarium and Penicillium, Candida that aid in the digestion of cellulose and hemicellulose feeds.
References
Abarenkov K, Henrik NR, Larsson KH et al (2010) The UNITE database for molecular identification of fungi–recent updates and future perspectives. New Phytol 186(2):281–285
Al-Essa MA, Al-Mehaidib A, Al-Gain S (1998) Parental awareness of liver disease among children in Saudi Arabia. Ann Saudi Med 18(1):79–81
Allred BW (1968) Range management training handbook for Saudi Arabia. FAO, Rome
Andert J, Marten A, Brandl R, Brune A (2010) Inter- and intraspecific comparison of the bacterial assemblages in the hindgut of humivorous scarab beetle larvae (Pachnoda spp.). FEMS Microbiol Ecol 74(2):439–449
Assaeed AM (1996) Hammada elegans-Rhazya stricta relationships in deteriorated range site in Raudhat Al-Khafs, Saudi Arabia. J Agric Sci Mansoura Univ 21(3):957–964
Baeshen NA, Sabir JS, Zainy MM et al (2014) Biodiversity and DNA barcoding of soil fungal flora associated with Rhazya stricta in Saudi Arabia. Bothalia 44(5):301–314
Baldrian P, Kolařík M, Stursová M et al (2012) Active and total microbial communities in forest soil are largely different and highly stratified during decomposition. ISME J 6:248–258
Blackwell M (2011) The Fungi: 1, 2, 3, 5.1 million species? Am J Bot 98(3):426–438
Bloemberg GV, Lugtenberg BJJ (2001) Molecular basis of plant growth promotion and biocontrol by rhizobacteria. Curr Opin Plant Biol 4:343–350
Brucker RM, Bordenstein SR (2012) Speciation by symbiosis. Trends Ecol Evol 27(8):443–451
Buée M, Reich M, Murat C et al (2009) 454 Pyrosequencing analyses of forest soils reveal an unexpectedly high fungal diversity. New Phytol 184:449–456
DeSalle R, Graham SW, Fazekas AJ et al (2008) Multiple multilocus DNA barcodes from the plastid genome discriminate plant species equally well. PLoS ONE 3:e2802
Douglas WF, Bing M, Pawel G, Naomi S, Sandra O, Rebecca MB (2014) An improved dual-indexing approach for multiplexed 16S rRNA gene sequencing on the Illumina MiSeq platform. Microbiome 2:6
Elshafie AE, Al Siyabi FM, Salih FM, Ba Omar T, Al Bahry SN, Al Kindi S (2003) The mycobiota of herbal drug plants in Oman and possible decontamination by gamma radiation. Phytopathol Mediterr 42(2):149–154
Engelen B, Meinkein K, von Wintzingerode F, Heuer H, Malkomes H-P, Backhaus H (1998) Monitoring impact of a pesticide treatment on bacterial soil communities by metabolic and genetic fingerprinting in addition to conventional testing procedures. Appl Environ Microbiol 64:2814–2821
Fraune S, Bosch TC (2010) Why bacteria matter in animal development and evolution. BioEssays 32:571–580
Geml J, Gravendeel B, van der Gaag KJ et al (2014) The contribution of DNA metabarcoding to fungal conservation: diversity assessment, habitat partitioning and mapping red-listed fungi in protected coastal Salix repens communities in the Netherlands. PLoS ONE 9(6):e99852
Hamady M, Knight R (2009) Microbial community profiling for human microbiome projects: tools, techniques, and challenges. Genome Res 19:1141–1152
Hirsch PR, Mauchline TH, Clark IM (2010) Culture-independent molecular techniques for soil microbial ecology. Soil Biol Biochem 42:878–887
Hollister EB, Schadt CW, Palumbo AV, Ansley RJ, Boutton TW (2010) Structural and functional diversity of soil bacterial and fungal communities following woody plant encroachment in the southern Great Plains. Soil Biol Biochem 42:1816–1824
Hongoh Y (2010) Diversity and genomes of uncultured microbial symbionts in the termite gut. Biosci Biotechnol Biochem 74:1145–1151
Hongoh Y, Deevong P, Inoue T et al (2005) Intra- and interspecific comparisons of bacterial diversity and community structure support coevolution of gut microbiota and termite host. Appl Environ Microbiol 71:6590–6599
Hongoh Y, Ekpornprasit L, Inoue T et al (2006) Intracolony variation of bacterial gut microbiota among castes and ages in the fungus-growing termite Macrotermes gilvus. Mol Ecol 15:505–516
Huang XF, Bakker MG, Judd TM, Reardon KF, Vivanco JM (2013) Variations in diversity and richness of gut bacterial communities of termites (Reticulitermes flavipes) fed with grassy and woody plant substrates. Microb Ecol 65:531–536
Huse SM, Ye Y, Zhou Y, Fodor AA (2012) A core human microbiome as viewed through 16S rRNA sequence clusters. PLoS ONE 7:e34242
Kernaghan G (2005) Mycorrhizal diversity: cause and effect? Pedobiologia 49:511–520
Kim M, Yoon H, You Y-H et al (2013) Metagenomic analysis of fungal communities inhabiting the fairy ring zone of Tricholoma matsutake. J Microbiol Biotechnol 23(10):1347–1356
Lee YK, Mazmanian SK (2010) Has the microbiota played a critical role in the evolution of the adaptive immune system? Science 330:1768–1773
Lehman RM, Lundgren JG, Petzke LM (2009) Bacterial communities associated with the digestive tract of the predatory ground beetle, Poecilus chalcites, and their modification by laboratory rearing and antibiotic treatment. Microb Ecol 57:349–358
Magoc T, Salzberg S (2011) FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27(21):2957–2963
Massol-Deya A, Weller R, Rios-Hernandez L, Zhou J-Z, Hickley RF, Tiedje JM (1997) Succession and convergence of biofilm communities in fixed-film reactors treating aromatic hydrocarbons in groundwater. Appl Environ Microbiol 63:270–276
McGuire KL, Fierer N, Bateman C, Treseder KK, Turner BL (2012) Fungal community composition in neotropical rain forests: the influence of tree diversity and precipitation. Microb Ecol 63:804–812
Moran NA, McCutcheon JP, Nakabachi A (2008) Genomics and evolution of heritable bacterial symbionts. Annu Rev Genet 42:165–190
Moussa TAA, Al-Zahrani HS, Almaghrabi OA, Abdelmoneim TS, Fuller MP (2017) Comparative metagenomics approaches to characterize the soil fungal communities of western coastal region. Saudi Arabia PLoS ONE 12(9):e0185096
Muyzer G, Smalla K (1998) Application of denaturing gradient gel electrophoresis (DGGE) and temperature gradient gel electrophoresis (TGGE) in microbial ecology. Antonie Van Leeuwenhoek 73:127–141
Nilsson RH, Kristiansson E, Ryberg M, Hallenberg N, Larsson KH (2008) Intraspecific ITS variability in the kingdom fungi as expressed in the international sequence databases and its implications for molecular species identification. Evol Bioinformatics Online 4:193–201
Orgiazzi A, Bianciotto V, Bonfante P et al (2013) 454 pyrosequencing analysis of fungal assemblages from geographically distant, disparate soils reveals spatial patterning and a core mycobiome. Diversity 5:73–98
Peay KG, Baraloto C, Fine PV (2013) Strong coupling of plant and fungal community structure across western Amazonian rainforests. ISME J 7:1852–1861
Quast C, Pruesse E, Yilmaz P et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucl Acids Res 41(D1):D590–D596
Ranjard L, Poly F, Lata J-C, Mougel C, Thioulouse J, Nazaret S (2001) Characterization of bacterial and fungal soil communities by automated ribosomal intergenic spacer analysis fingerprints: biological and methodological variability. Appl Environ Microbiol 67(10):4479–4487
Schäfer H, Bernard L, Courties C et al (2001) Microbial community dynamics in Mediterranean nutrient-enriched seawater mesocosms: changes in the genetic diversity of bacterial populations. FEMS Microbiol Ecol 34:243–253
Schmit JP, Mueller GM (2007) An estimate of the lower limit of global fungal diversity. Biodivers Conserv 16:99–111
Sharon G, Segal D, Ringo JM, Hefetz A, Zilber-Rosenberg I, Rosenberg E (2010) Commensal bacteria play a role in mating preference of Drosophila melanogaster. PNAS USA 107:20051–20056
Smit E, Leeflang P, Wernars K (1997) Detection of shifts in microbial community structure and diversity in soil caused by copper contamination using amplified ribosomal DNA restriction analysis. FEMS Microbiol Ecol 23:249–261
Turnbaugh PJ, Ley RE, Hamady M, Fraser-Liggett CM, Knight R, Gordon JI (2007) The human microbiome project. Nature 449:804–810
Turner TR, Ramakrishnan K, Walshaw J et al (2013) Comparative metatranscriptomics reveals kingdom level changes in the rhizosphere microbiome of plants. ISME J 7:2248–2258
Verbruggen E, Kiers ET, Bakelaar PNC, Röling WFM, van der Heijden MGA (2012) Provision of contrasting ecosystem services by soil communities from different agricultural fields. Plant Soil 350:43–55
Warnecke F, Luginbuhl P, Ivanova N et al (2007) Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature 450:560–565
Werren JH, Baldo L, Clark ME (2008) Wolbachia: master manipulators of invertebrate biology. Nat Rev Microbiol 6:741–751
Wu YT, Wubet T, Trogisch S, Both S, Scholten T, Bruelheide H, Buscot F (2013) Forest age and plant species composition determine the soil fungal community composition in a Chinese subtropical forest. PLoS ONE 8:e66829
Yasir M, Azhar EI, Khan I et al (2015) Composition of soil microbiome along elevation gradients in southwestern highlands of Saudi Arabia. BMC Microbiol 15:65
You YH, Yoon H, Lee GS et al (2011) Diversity and plant growth-promotion of endophytic fungi isolated from the roots of plants in Dokdo islands. J Life Sci 21:992–996
You YH, Yoon H, Woo JR et al (2011) Plant growth-promoting activity of endophytic fungi isolated from the roots of native plants in Dokdo islands. J Life Sci 21:1619–1624
You YH, Yoon H, Kim H et al (2013) Plant growth-promoting activity and genetic diversity of endophytic fungi isolated from native plants in Dokdo Islands for restoration of a coastal ecosystem. J Life Sci 23:95–101
Yu L, Nicolaisen M, Larsen J, Ravnskov S (2013) Organic fertilization alters the community composition of root-associated fungi in Pisum sativum. Soil Biol Biochem 58:36–41
Zak DR, Holmes WE, White DC, Peacock AD, Tilman D (2003) Plant diversity, soil microbial communities, and ecosystem function: are there any links? Ecology 84:2042–2050
Acknowledgment
This project was funded by the Deanship of Scientific Research (DSR) at King Abdulaziz University, Jeddah, under grant no. (HiCi 1-363-1434H). The authors, therefore, acknowledge with thanks DSR for technical and financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Noor, S.O., Al-Zahrani, D.A., Hussein, R.M. et al. Assessment of fungal diversity in soil rhizosphere associated with Rhazya stricta and some desert plants using metagenomics. Arch Microbiol 203, 1211–1219 (2021). https://doi.org/10.1007/s00203-020-02119-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-02119-z