Abstract
A novel actinomycete, designated TRM68295T, was isolated from the sediments of the Tailan River, Xinjiang, northwest China. The study of the polyphasic approach showed that the characteristics of the strain were consistent with the genus Streptomyces. Phylogenetic analysis indicated that the strain had a high sequence similarity with Streptomyces phaeochromogenes ATCC 3338T (97.9%). The diagnostic diamino acid of cell walls was identified as ll-diaminopimelic acid. The whole cell sugars were identified as ribose, xylose, glucose and galactose. The major fatty acids were iso-C14:0, iso-C15:0, anteiso-C15:0, iso-C16:0, C16:1ω9c, C16:0 and anteiso-C17:0. The major menaquinones were MK-9 (H10), MK-9 (H6) and MK-9 (H2). The polar lipids were composed of diphosphatidylglycerol (DPG), phosphatidylethanolamine (PE), phosphatidylglycerol (PG), phosphatidylcholine (PC), phosphatidylinositol (PI) and phosphatidylinositol mannoside (PIM). The G+C content in the draft genome sequence of the strain was identified as 70.0 mol%. The average nucleotide identity (ANI) between strains TRM68295T and S. phaeochromogenes ATCC 3338T was 88.1%. Digital DNA-DNA hybridization (dDDH) value between strain TRM68295T and S. phaeochromogenes ATCC 3338T was 40.7%. The multilocus sequence analysis of five house-keeping genes (atpD, gyrB, recA, rpoB, and trpB) indicated that the MLSA distances between the strain and similar species were greater than the threshold of 0.007. Based on the above studies, strain TRM68295T is considered as a novel species of the genus Streptomyces, for which the name Streptomyces albicerus sp. nov. is proposed. The type strain is TRM68295T (= CCTCC AA 2018085T = KCTC 49272T).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Waksman and Henrici first proposed Streptomyces in 1943. As of this writing, more than 850 species with valid names of Streptomyces have been published (https://www.bacterio.net/streptomyces.html) (Zhang et al. 2017). Streptomyces are aerobic Gram-positive bacteria with well-developed mycelia, which can produce a large number of conidia for reproduction. Streptomyces can produce a variety of secondary metabolites, such as antimicrobial drugs, antineoplastic drugs, enzyme inhibitory activities, herbicides, insecticides, antiparasitic agents, hypoglycemic agents (Deshpande et al. 1988). To find microorganisms that can produce new antibiotics, more attention focuses on the unexplored natural environments, such as deep sea (Tian et al. 2013), desert (Calasanz et al. 2013), polar (Encheva et al. 2013) and salt lake (Jose and Jebakumar 2012).
The Tailan River Basin is located in Wensu, Aksu District, Xinjiang. It originates from the Tuomuer Peak in the southwestern Tianshan Mountains. The upper reaches are the Big Tailan River and the Small Tailan River. The two rivers are called Tailan River after merging in the area of 8 km before the mountain. The climate of the basin belongs to the continental temperate arid climate with properties of dry climate, abundant sunshine, windy sand, rare precipitation, large evaporation and large temperature difference between day and night. We sampled sediments of the Tailan River to isolate actinomycetes and analyze the diversity of actinomycetes. During the process, a novel species TRM68295T was isolated. This study determined the taxonomic status of the strain based on genotype, physiological and biochemical characteristics, and phylogeny.
Materials and methods
Strain isolation and culturing
Strain TRM68295T was isolated from the sediments of the Tailan River at Aksu, Xinjiang, northwest China (latitude 40° 71′; longitude 81° 16′). The strain was isolated by the spread plate method on glycerol–arginine medium which contained (g/l): arginine 2 g, glycerol 12 g, MgSO4 0.5 g, K2HPO4 1 g, agar 17 g, pH 7.5, supplemented with 100 µl of 50 mg K2Cr2O7/ml in a 100 ml medium to reduce fungal contamination. The strain was isolated after culturing at 28 °C for 10 days. The strain was purified on Gauze's No. 1 medium at 28 °C. The strain was stored in 20% glycerol for a short term, and lyophilized in 20% skim milk powder for long-term storage. Streptomyces phaeochromogenes ATCC 3338T was purchased from American Type Culture Collection (ATCC). Streptomyces ederensis JCM 4958T was purchased from the Japan Collection of Microorganisms (JCM).
Morphological, cultural, physiological, and biochemical characteristics
To determine the culture characteristics, strain TRM68295T was cultured on ISP 1-7 medium (Shirling and Gottlieb 1966), Czapek’s agar, Nutrient agar (Waksman 1967) and Gauze’s No. 1 medium for 14 days at 28 °C. The colors of the substrate mycelium, aerial mycelium and soluble pigment were determined by the ISCC NBS color chart (Kelly 1964). Morphology properties were observed by light microscopy (Leica DM1000) and scanning electron microscopy (Thermo Apreo) using cultures grown on Gauze's No. 1 medium at 28 °C for 20 days. Different temperatures (4, 16, 20, 25, 28, 32, 37, 40, 45 and 50 °C), pH (pH 4.0–12.0, interval 1.0 pH unit) (Xu et al. 2005) and concentrations of NaCl (0–10%, interval 1%, w/v) for strain growth were examined on Gauze’s No. 1 medium and were observed after at 28 °C for 14 days. The productions of peroxidase, urease, esterase and catalase were tested using the method described by Gerhardt et al (1994). The uses of sole carbon source (0.5%, w/v), cellulose decomposition, starch hydrolysis, liquefaction of gelatin, milk peptonization and solidification, nitrate reduction and production of H2S (Gordon 1974; Yokota et al. 1993) were studied.
Chemotaxonomy
Biomass used for studies was obtained by culture in liquid Gauze's No. 1 medium for 7 days in shake flasks at 28 °C. The cells collected by centrifugation were washed with distilled water, and then freeze-dried. Cell wall amino acids were determined by the method proposed by Staneck and Roberts (1974). The whole cell sugars were determined by the method proposed by Tang et al. (2009). Polar lipids were detected by two-dimensional TLC and identified by the method proposed by Minnikin et al. (1984). Menaquinones were extracted from freeze-dried biomass according to the method proposed by Collins et al. (1977), and analyzed by HPLC (Wang et al. 2011). Cellular fatty acids were extracted from the fresh cells according to the method proposed by Sasser (1990), and analyzed by GS chromatography using the Microbial Identification System (Sherlock version 6.1; MIDI database: RTSBA6).
Genome sequencing and phylogenetic analysis
The extraction of genomic DNA and PCR amplification of 16S rRNA sequence was performed by the method of Chun and Goodfellow (1995). The purified PCR product was cloned into the vector pMD18-T (Takara) and sent to Sangon for gene sequencing. The two sequences obtained by gene sequencing were spliced using SeqMan software. After using vecscreen (https://www.ncbi.nlm.nih.gov/tools/vecscreen/) to remove the vector sequences, an almost complete 16S rRNA gene sequence (1494 bp) was obtained. EzBioCloud (https://www.ezbiocloud.net/identify, Yoon et al. 2017) was used to calculate the 16S rRNA gene sequence similarity with other strains, and then the sequences of closely related strains were selected to construct the phylogenetic tree. The initial alignment was performed using clustal_x software (Thompson et al. 1997). The phylogenetic tree was constructed using the Neighbor-Joining (Saitou and Nei 1987), Maximum-Parsimony (Fitch 1971) and Maximum-Likelihood (Felsenstein 1981). The corresponding algorithms were realized using MEGA 7.0 (Kumar et al. 2016). The stability of the topology of the phylogenetic trees was evaluated by performing 1000 resamplings (Felsenstein 1985).
The draft genome sequence of strain TRM68295T was sequenced using the Illumina platform. Velvet (Bankevich et al. 2012) software was used to assemble the sequencing data of the strain. The completeness and contamination of the genome were evaluated by the software CheckM (Parks et al. 2015). The average depth of genome coverage was 137-fold. The assembled genome was predicted by prodigal (Hyatt et al. 2010). The software tRNAscan-SE (Lowe and Eddy 1997) was used to predict the tRNA in the genome, and the software Infernal1.1 (Nawrocki and Eddy 2013) was used to predict the rRNA in the genome based on the Rfam (Nawrocki et al. 2015) database. The putative biosynthetic gene clusters for secondary metabolites were identified using the AntiSMASH program, version 4.0 (Weber et al. 2015). The genome sequences of strain TRM68295T (accession no. VWMY00000000) and strain S. phaeochromogenes (accession no. LIQZ00000000) have been submitted to GenBank. The G+C content of strain TRM68295T was determined from the draft genome sequence by Average Nucleotide Identity (ANI) calculator (Yoon et al. 2017). The average nucleotide identity (ANI) was determined using the web-service (https://www.ezbiocloud.net/tools/ani) (Lee et al. 2015). The digital DNA–DNA hybridization (dDDH) values were calculated on the GGDC website using formula 2, originally described by Auch et al. (2010) and updated by Meier-Kolthoff et al (2013). Five housekeeping gene sequences of strain TRM68295T were obtained by PCR amplification. The housekeeping genes of closely related strains were downloaded from the website (https://199.133.98.43/cgi-bin/bigsdb/bigsdb.pl?db=public_ars_streptomyces_isolates&page=query). The phylogenetic tree was constructed using the Neighbor-Joining algorithm in MEGA 7.0 (Kumar et al. 2016). The MLSA distances were calculated using MEGA 7.0.
Results and discussion
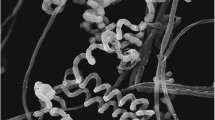
After 14-day culture at 28 °C, the growth of strain TRM68295T on ten standard media, the colors of aerial mycelia and substrate mycelia were recorded (Table 1). The strain grew well on ISP 4, ISP 5, ISP 7, Gauze’s No. 1 agar and Czapek’s agar, and grew moderately on ISP 1, ISP 2, ISP 3, ISP 6 and nutrient agar medium. A soluble pigment (black) was produced on ISP 7 medium. The aerial mycelium was dense, cylindrical, with a smooth surface and no branches. The spore chain was helical. The spores were cylindrical, of which the surface was spiny (Supplementary Fig. 1). The strain could grow at 16–45 °C, pH 6.0–10.0 and 0–6% NaCl, and grow best at 2% (w/v) NaCl at 28 °C and pH 7.0. The strain was found to be positive for catalase, urease, gelatin liquefaction, milk peptonization and solidification, starch hydrolysis, nitrate reduction, cellulose decomposition, but negative for oxidase, melanin production, H2S production. The strain could degrade Tweens 20, 40, 60 and 80.
The diagnostic diamino acid of cell wall was identified as ll-diaminopimelic acid. The whole cell sugars were found with ribose, xylose, glucose and galactose as main sugars. The polar lipids were composed of diphosphatidylglycerol (DPG), phosphatidylethanolamine (PE), phosphatidylglycerol (PG), phosphatidylcholine (PC), phosphatidylinositol (PI) and phosphatidylinositol mannoside (PIM) (Supplementary Fig. 2). The menaquinones were identified as MK-9 (H10), MK-9 (H6) and MK-9 (H2). The major fatty acids were identified (> 5%) as iso-C14:0 (5.5%), iso-C15:0 (11.2%), anteiso-C15:0 (19.4%), iso-C16:0 (21.0%), C16:1ω9c (5.3%), C16:0 (9.1%) and anteiso-C17:0 (9.0%).
According to the EzBioCloud analysis, the 16S rRNA gene sequence of strain TRM68295T (GenBank accession no. MK795696) had high similarity with members of the genus Streptomyces, and the closest phylogenetic neighbor was S. phaeochromogenes ATCC 3338T (97.9%). The phylogenetic tree constructed from the 16S rRNA gene sequences by the Neighbor-Joining method showed that strain TRM68295T formed a distinct clade (Fig. 1), which was also recovered in the Maximum-Parsimony and Maximum-Likelihood trees (Supplementary Figs. 3, 4). The draft genome sequence consists of 307 contigs and 200 scaffolds covering a total sequence length of 10,359,585 bp, with an N50 contig size of 82,685 bp and an N50 scaffold size of 138,999 bp. The completeness and contamination of the genome were 99.5% and 0.6%, respectively. The number of rRNAs and tRNAs predicted by genome was 5 and 68, respectively. The draft genome contains 47 putative biosynthetic gene clusters capable of producing various secondary metabolites, including polyketides, non-ribosomal peptides, terpenes, lanthipeptides, siderophores, ectoine, bacteriocins, lassopeptides, melanin and thiopeptide, which suggested that strain TRM68295T had the potential to produce novel antimicrobial compounds or their analogues.
The G+C content in the draft genome sequence of strain TRM68295T was identified as 70.0 mol%. The average nucleotide identity (ANI) value of strains TRM68295T and S. phaeochromogenes ATCC 3338T was 88.1%. The average nucleotide identity value is significantly lower than the widely accepted threshold for describing prokaryote species (95–96%; Kim et al. 2014). The dDDH value of strain TRM68295T and its closest strain S. phaeochromogenes ATCC 3338T was 40.7%, which is significantly lower than 70% of the species defined threshold (Richter et al. 2016; Wayne et al. 1987). The MLSA phylogenetic analysis showed Streptomyces dioscori (MLSA distance = 0.049), Streptomyces cinereoruber subsp. fructofermentans (MLSA distance = 0.051), Streptomyces tauricus (MLSA distance = 0.053) and S. phaeochromogenes (MLSA distance = 0.056) as the nearest neighbors (Fig. 2). The MLSA distance was much larger than the threshold for species classification (> 0.007) (Rong and Huang 2012). The characteristics differentiating between strain TRM68295T and closely related strains are shown in Tables 2 and 3. Based on the above studies, strain TRM68295T is considered a novel species of the genus Streptomyces, for which the name Streptomyces albicerus sp. nov. is proposed. The type strain is TRM68295T (= CCTCC AA 2018085T = KCTC 49272T).
Neighbour-joining tree based on concatenated partial sequences of five housekeeping genes (atpD, gyrB, recA, rpoB and trpB) shows the position of strain TRM68295T amongst its phylogenetic neighbours. Only bootstrap values above 50% (percentages of 1000 replications) are indicated. Bar, 0.01 substitutions per nucleotide position
Description of Streptomyces albicerus sp. nov.
Streptomyces albicerus (al.bi'cer.us L. masc. adj. albicerus, Pale yellow, because of the pale yellow aerial hyphae of the strain).
Gram-positive, aerobic actinomycetes. The aerial mycelium is dense, cylindrical, and the surface is smooth. The spore chain is spiral, and the spore surface is spiny. Grow well on ISP 4, ISP 5, ISP 7, Gauze's No. 1 medium and Czapek's agar; grow moderately on ISP 1, ISP 2, ISP 3, ISP 6 and Nutrient agar medium. Soluble pigment (black) is produced on ISP 7 medium. The strain can grow at 16–45 °C, pH 6.0–10.0 and 0–6% NaCl, and grow best at 2% (w/v) NaCl at 28 °C and pH 7.0. Maltose, d-trehalose, xylose, raffinose, d-galactose, chitosan, l-sorbose, lactose, d-cellobiose, glucose, l-fucose, sucrose, xylan, Inositol, d-ribose, l-arabinose, d-fructose, starch or l-rhamnose are utilized. Melezitose, d-Salicin and xylitol are not utilized. Positive for catalase, urease, gelatin liquefaction, milk peptonization and solidification, starch hydrolysis, nitrate reduction, cellulose decomposition, degradations of Tweens 20,40, 60 and 80, whereas negative for oxidase, melanin production, production of H2S; contains ll-diaminopimelic as diagnostic amino acid of the peptidoglycan and ribose, xylose, glucose and galactose in whole cell hydrolysates. The polar lipids contain diphosphatidylglycerol (DPG), phosphatidylethanolamine (PE), phosphatidylglycerol (PG), phosphatidylcholine (PC), phosphatidylinositol (PI) and phosphatidylinositol mannoside (PIM). The menaquinone system contains MK-9 (H10), MK-9 (H6) and MK-9 (H2). The major fatty acids are iso-C14:0, iso-C15:0, anteiso-C15:0, iso-C16:0, C16:1ω9c, C16:0, anteiso-C17:0. The G+C content in the draft genome sequence of strain TRM68295T is 70.0 mol%.
The type strain is TRM68295T (= CCTCC AA 2018085T = KCTC 49272T), isolated from sediment samples of the Tailan River at Aksu, Xinjiang, northwest China. The GenBank accession number of 16S rRNA gene sequence is MK795696. The GenBank accession number for the whole genome sequence is VWMY00000000.
References
Auch AF, von Jan M, Klenk HP, Göker M (2010) Digital DNA–DNA hybridization for microbial species delineation by means of genome-to-genome sequence comparison. Stand Genomic Sci 2:117–134
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA (2012) Spades: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477
Calasanz M, Göker M, Pötter G, Rohde M, Spröer C, Schumann P, Klenk HP, Gorbushina AA (2013) Geodermatophilus telluris sp. nov., an actinomycete isolated from Saharan desert sand. Int J Syst Evol Microbiol 63:2254–2259
Chun JS, Goodfellow M (1995) A phylogenetic analysis of the genus Nocardia with 16S rRNA gene sequences. Int J Syst Bacteriol 45:240–245
Collins MD, Pirouz T, Goodfellow M, Minnikin DE (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230
Deshpande BS, Ambedkar SS, Shewale JG (1988) Biologically active secondary metabolites from Streptomyces. Enzyme Microb Tech 10:455–473
Encheva M, Zaharieva N, Kenarova A, Chipev N, Chipeva V, Hristova P, Ivanova I, Moncheva P (2013) Abundance and activity of soil actinomycetes from Livingston Island, Antarctica. Bulg J Agric Sci 19:68–71
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–789
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Sys Zool 20:406–416
Gerhardt P, Murray RGE, Wood WA, Smibert RM, Krieg NR (1994) Methods for general and molecular bacteriology. American Society for Microbiology, Washington
Gordon RA (1974) Nocardia coeliaca, Nocardia autotrophica, and the Nocardin strain. Int J Syst Bacteriol 24:54–63
Hyatt D, Chen GL, Locascio PF, Land ML, Larimer FW, Hauser LJ (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform 11:119
Jose PA, Jebakumar SRD (2012) Phylogenetic diversity of actinomycetes cultured from coastal multipond solar saltern in Tuticorin. India Aquat Biosyst 8:23
Kelly KL (1964) Inter-Society Color Council—National Bureau of Standards Color Name Charts illustrated with centroid colors. US Government Printing Office, Washington
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lee I, Kim YO, Park SC, Chun J (2015) Orthoani: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964
Meier-Kolthoff PJ, Alexander FA, Klenk HP, Göker M (2013) Genome sequence based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal K, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Nawrocki EP, Eddy SR (2013) Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 29:2933–2935
Nawrocki EP, Burge SW, Bateman A, Daub J, Eberhardt RY, Eddy SR, Floden EW, Gardner PP, Jones TA, Tate J, Finn RD (2015) Rfam 12.0: updates to the RNA families database. Nucleic Acids Res 43:130–137
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW (2015) CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25:1043–1055
Piao C, Zheng W, Li Y, Liu C, Jin L, Song W, Yan K, Wang X, Xiang W (2017) Two new species of the genus Streptomyces: Streptomyces camponoti sp. nov. and Streptomyces cuticulae sp. nov. isolated from the cuticle of Camponotus japonicus Mayr. Arch Microbiol 199:963–970
Richter M, Rosselló-Móra R, Oliver Glöckner F, Peplies J (2016) J SpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 32:929–931
Rong X, Huang Y (2012) Taxonomic evaluation of the Streptomyces hygroscopicus clade using multi-locus sequence analysis and DNA–DNA hybridization, validating the MLSA scheme for the systematics of the whole genus. Syst Appl Microbiol 35:7–18
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Bio Evol 4:406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. Technical note 101. Microbial ID, Newark
Shirling EB, Gottlieb D (1966) Methods for characterization of streptomyces species. Int J Sys Bacteriol 16:313–340
Staneck JL, Roberts GD (1974) Simplified approach to identification of aerobic actinomycetes by thin-layer chromatography. Appl Microbiol 28:226–231
Tang SK, Wang Y, Chen Y, Lou K, Cao LL, Xu LH, Li WJ (2009) Zhihengliuella alba sp. nov., and emended description of the genus Zhihengliuella. Int J Syst Evol Microbiol 59:2025–2031
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tian XP, Long LJ, Li SM, Zhang J, Xu Y, He J, Li J, Wang FZ, Li WJ, Zhang CS, Zhang S (2013) Pseudonocardia antitumoralis sp. nov., a deoxynyboquinone-producing actinomycete isolated from a deep-sea sediment. Int J Syst Evol Microbiol 63:893–899
Waksman SA (1967) The actinomycetes a summary of current knowledge. Ronald Press, New York
Waksman SA, Henrici AT (1943) The nomenclature and classification of the actinomycetes. J Bacteriol 46:337–341
Wang C, Xu XX, Qu Z, Wang HL, Lin HP, Xie QY, Ruan JS, Hong K (2011) Micromonospora rhizosphaerae sp. nov., isolated from mangrove rhizosphere soil. Int J Syst Evol Microbiol 61:320–324
Wang Z, Tian J, Li X, Gan L, He L, Chu Y, Tian Y (2018) Streptomyces dioscori sp. nov., a novel endophytic actinobacterium isolated from bulbil of Dioscorea bulbifera L. Curr Microbiol 75:1384–1390
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE, Stackebrandt E, Starr MP, Trüper HG (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Weber T, Blin K, Duddela S, Krug D, Kim HU, Bruccoleri R, Lee SY, Fischbach MA, Müller R, Wohlleben W, Breitling R, Takano E, Medema MH (2015) antiSMASH 3.0—a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res 43:237–243
Xu P, Li W, Tang S, Zhang Y, Chen G, Chen H, Xu L, Jiang C (2005) Naxibacter alkalitolerans gen. nov., sp. nov., a novel member of the family 'Oxalobacteraceae' isolated from China. Int J Syst Evol Microbiol 55:1149–1153
Yokota A, Tamura T, Hasegawa T, Huang LH (1993) Catenuloplanes japonicus gen. nov., sp. nov., nom. rev., a new genus of the order actinomycetales. Int J Syst Bacteriol 43:805–812
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol 67:1613–2161
Zhang R, Han X, Xia Z, Luo X, Wan C, Zhang L (2017) Streptomyces luozhongensis sp. nov., a novel actinomycete with antifungal activity and antibacterial activity. Antonie Van Leeuwenhoek 110:195–203
Acknowledgements
Thanks to Key Laboratory of Protection and Utilization of Biological Resources in Tarim Basin of Tarim Basin of Xinjiang Production & Construction Corps for providing research facilities.
Funding
This research was supported by the National Natural Science Foundation of China-Xinjiang Joint Fund Key Project (project no. U1703236) and Tarim University graduate research and innovation projects (project no. XJ2019G270).
Author information
Authors and Affiliations
Contributions
BS contributed to performing the experiments and writing the initial draft. LY, ZX and CW contributed to the guidance of experimental operations. LZ contributed to reagents, instrumentation and the financial support for this work.
Corresponding author
Ethics declarations
Conflict of interest
The authors state that there is no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The GenBank/EMBL/DDBJ accession number for the genome and 16S rRNA gene sequence of strain TRM68295T is VWMY00000000 and MK795696.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sun, B., Yuan, L., Xia, Z. et al. Streptomyces albicerus sp. nov., a novel actinomycete isolated from the sediments of the Tailan River in Xinjiang, China. Arch Microbiol 202, 1639–1646 (2020). https://doi.org/10.1007/s00203-020-01871-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-01871-6