Abstract
Pectinase is widely used in numerous industrial fields, including the food, wine, and paper industries. In this work, seven bacteria were isolated from orange peel and their pectinase production activity was assayed. One bacterium (OR-B2) identified as a Bacillus sp. showed the highest enzyme activity towards others. A gene encoding a pectate lyase designed as PelB-B2 in this work was amplified and heterogeneous expressed in E.coli. PelB-B2 was defined as a member of the PelB pectate lyase family after phylogenic tree analysis. 3D model of PelB-B2 was constructed by SWISS-MODEL and PelB-B2 showed conserved para-β structure. After inducing culture and purified by Ni-affinity chromatography, the properties of the purified PelB-B2 were assayed. Optimal pH and temperature for PelB-B2 was pH 8.0 and 50 °C, respectively. PelB-B2 showed excellent pH stability and thermostability. It was stable within pH range 3.0–11.0 and retained more than 51% activity after incubation at 40 °C, 50 °C, or 60 °C for 1 h. Furthermore, we determined that PelB-B2 was a Ca2+-dependent pectinase and the pectin extracted from citrus was the benefit substrate for PelB-B2. The Km and Vmax of PelB-B2 were 1.64 g/L and 232.56 mol/(L min), respectively. The OR-B2 can be a new resource for pectinase production and the PelB-B2 has potential for industrial application.
Graphic abstract
7 bacteria were isolated from orange peel, namely OR-B1 to OR-B7 and their pectinase activities were assayed. One pectate lyase belongs to PelB family was cloned from OR-B2 and heterogeneous expressed in E. coli. Purified PelB-B2 was further studied with its properties. Effects of pH, temperature, chemicals, substrate on the enzyme activity were assayed and the enzyme kinetic was also measured.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pectic substances and pectinases
Pectic substance is a generic name for compounds occurring as major components of the middle lamella between plant cells, having high molecular weight and negative charge, and which are acidic, complex glycosidic macromolecules (Jayani et al. 2005). They account for 0.5–4.0% of the fresh weight of plant material (Kashyap et al. 2001; Sakai et al. 1993). In the case of dried fruits, the percentage of pectin may be greater than 30% (Tapre and Jain, 2014). Chemically, pectic substances are composed of polymers of galacturonic acid residues that are α-d-(1,4)-linked at the O1 and O4 positions and sometimes methylated at the C6 and or acetylated at the O2 and or O3 positions (Lerouxel et al. 2006; Soumpourou et al. 2007). Pectic substances are present in various forms and classified by the American Chemical Society into four main types based on their structures (Agarwal et al. 1999; Jayani et al. 2005): (1) protopectins, which are water insoluble and yield pectin or pectic acids; (2) pectic acids, which contain negligible amounts of methoxyl groups; (3) pectinic acids, a term referring to polygalacturonan chains containing > 0 but < 75% methylated galacturonate units; and (4) pectin (polymethylgalacturonate), which is a polymeric material wherein at least 75% of the carboxyl groups of the galacturonate units are esterified with methanol.
Pectinases, also known as pectolytic or pectic enzymes, belong to the family of polysaccharides that contribute to the breakdown of pectins (Henrissat and Bairoch 1993; Kertesz 1955). Pectinases catalyze degradation of pectic substances through depolymerization (hydrolases and lyases) or by deesterification (esterases) reactions. Resulting from the various forms of pectic substances, pectolytic enzymes appeared in various forms and have been classified under three headings according to their preferred substrate and method of action. The three major types of pectinases are as follow: (1) protopectinase, which enzyme solubilizes protopectin forming highly polymerized soluble pectin; (2) pectinesterases, which catalyze deesterification of the methoxyl group of pectin forming pectic acid; and (3) depolymerizing enzymes, which degrade substrates by hydrolyzing or cleaving the glycosidic linkages (Alimardani-Theuil et al. 2011; Kashyap et al. 2001). The major types and action modes of pectinases are classified in Table S1.
Occurrence and application of pectinase
Pectinases may have been among the first enzymes used in the home and they hold a leading position among the commercially produced industrial enzymes (Kashyap et al. 2001, Rashmi 2008). They are reported to be produced efficiently by microbes and plants (Saranraj and Naidu 2014). Fifty percent of industrial enzymes originate from fungi and yeast, while 35% are from bacteria and the remaining 15% are of plant or animal origin (Rubio et al. 2005). In the field of microbial pectinases, the search for microbes capable to produce pectinases is never-ending. In recent years, numerous microbes producing pectic enzymes have been isolated and the properties of those enzymes characterized. For example, fungi of Aspergillusniger and Rhizomucorpusillus were used for pectinase production (Anand et al. 2017; Reginatto et al. 2017; Siddiqui et al. 2012). In addition, bacteria such as Bacillusclausii isolated from alkaline soil and Bacillusfirmus isolated from a cow also can produce pectinase efficiently (Li et al. 2012; Roosdiana et al. 2013).
The application of pectinases can be summarized according to their acid–base properties. Acidic pectinases have found widespread use within the beverage industry for extraction and clarification of juice and wine (Alkorta et al. 1998; Blanco et al. 1998; Gainvors et al. 1994; Hugouvieux-Cotte-Pattat 1996, Kashyap et al. 2001; Pretel et al. 1997). Alkaline pectinases are mainly used in the degumming and retting of fiber crops and pretreatment of pectic wastewater from fruit juice industries including in the areas of papermaking, oil extraction, and coffee and tea fermentation (Kashyap et al. 2001). In addition, bacterial pectinases also have shown biological control potential in the biopesticide industry (Ju 2011).
In this study, we sought to isolate from citrus peel several bacterial capable to produce pectinases. We then successfully expressed a pectate lyase produced by OR-B2, a Bacillus sp. in Escherichiacoli, and characterized this enzyme according to their properties. The research findings presented here will enrich the pectinase-producing microbial resource, enhance our current knowledge of such microbes, and promote further studies and applications of microbial produced pectinases.
Materials and methods
Materials and media for microculture and production of enzyme
Fresh oranges were acquired from a local market in Fuzhou, Fujian province of China. The peels were pared and moisturizing-cultured in petri dish at room temperature for 1 week to promote microorganisms’ growth. Bacteria were isolated on nutrient agar (NA) (3% beer-extract, 10% peptone, 5% NaCl and 2% agar, pH 7.5). Single colony of these bacteria was obtained by streak plate method and preserve for further study. For enzyme production, microbes were first cultured in seed medium: bacteria in nutrient broth (NA without agar) followed by overnight culture at 37 °C for 2 days. The seed cultures were transferred to fermentation culture (3% orange peel powder, 0.3% (NH4)2SO4, 0.05% MgSO4, 0.1% K2HPO4, 0.02% CaCl2, 0.001 FeSO4, and 0.1 NaCl) for fermentation.
Enzyme assay
The activity of pectinases produced by microbes isolated in this study was determined by measuring the release of reducing sugar from substrates using the 3,5-dinitrosalicylic acid (DNS) method as described previously (Guan et al. 2018a, b). d-( +)-Galacturonic acid monohydrate (Sigma) was used to generate the standard curve. The initial concentration of d-( +)-Galacturonic acid monohydrate was 0.4 mg/mL and the reaction system was as shown in Table S2. Fermented products were centrifuged at 4 °C for 5 min and the supernatant was used for enzyme assay. The reaction system was 0.1 mL diluted enzyme and 0.9 mL 1% (w/v) pectin (Sigma) dissolved by citric acid-Na2PO4 buffer (pH 7.0). After incubating at 50 °C for 30 min, the reaction was stopped by adding 1.5 mL DNS and boiling 10 min for color development. The OD540 was read and the production of reducing sugar was calculated according to the standard curve. One unit (U) of pectinase was defined as the amount of enzyme required to release 1 μmol of d-( +)-Galacturonic acid per minute. Triplicate experiments were carried out for every sample.
Identification of microbes
16S rDNA of OR-B2 was amplified with specific primers (Table S3) and sequenced for verification at Invitrogen. The feedback sequences were searched using BLAST at the National Center for Biotechnology Information (NCBI) and BLAST search of nucleotides was conducted in relation to other microbes for purposes of DNA similarity assay. Phylogenetic analysis was then performed using a neighbor-joining method in MEGA5 software.
Gene cloning, sequence analysis, and heterologous expression of PelB-B2
The full length of pelB-B2 was amplified from genome of OR-B2 with specific primers (Table S3), inserted into pET22b, forming the recombinant vector of pET22b-pelB-B2, and then sequenced at Invitrogen. The protein sequence similarity was assessed using BLAST. Phylogenetic analysis was then performed for the PelB-B2 homologues in other Bacillus sp. using a neighbor-joining method in MEGA5 software. 3D model of PelB-B2 was constructed by SWISS-MODEL.
The recombinant plasmid of pET22b-pelB-B2 was transferred into E.coli BL21(DE3), and then incubated at 37 °C till the OD600 reached 0.6–0.8, followed by 0.6 mM IPTG added for induction culture at 15 °C. After 200 rpm, 24 h incubation, the culture was collected, resuspended with Tris–HCl (pH 7.0) and lysis by ultrasonication. The supernatant was purified by Ni-affinity chromatography.
Characterization of PelB-B2
The activity of PelB-B2 was assayed using 3,5-dinitrosalicylic acid (DNS) as previously described. The optimal temperature for enzyme was determined as follows: enzyme were co-incubated with substrate (pectin extracted from citrus, pectin extracted from apple or Polygalacturonic acid) dissolved in citric acid-Na2PO4 buffer with pH 7.0, and reaction temperature was ranging from 30 to 70 °C. The enzyme activity also was measured as described above. The thermostability of enzyme was determined by measuring the activity of enzyme remaining after incubating at different temperatures for 5 min, 10 min, 20 min, 30 min, and 60 min, respectively. The initial enzyme activity was set equal to 100%.
The optimal pH for PelB-B2 was determined as follows: enzyme was co-incubated with substrate dissolved in buffer with different pH (citric acid-Na2PO4 buffer for pH 3.0–7.0, Tris–HCl for pH 7.0–9.0, and Gly–NaOH for pH 9.0–11.0) and the enzyme activity was measured at 50 °C. The pH stability of pectinases was determined by incubating the enzyme with buffer without substrate at varying pH at 37 °C for 1 h and the enzyme activity was measured as described above. The initial enzyme activity was set equal to 100%.
The effect of chemicals including metal ions, surfactant, or chelant on the enzyme activity of PelB-B2 was assayed. In detail, NiSO4 (Ni2+), MnSO4 (Mn2+), MgSO4 (Mg2+), KNO3 (K+), NaSO4 (Na+), Li2SO4 (Li+), FeCl3 (Fe3+), ZnSO4 (Zn2+), CaCl2 (Ca2+), CuSO4 (Cu2+), CrCl3 (Cr3+), BaCl2 (Ba2+), HgCl2 (Hg2+), AgCl (Ag+), Co (NO3)2(Co2+), SDS, and EDTA were added in the reaction system with the final concentration with 1 mM and the enzyme activity was measured as described above. The enzyme activity in the absence of any effector was set as the control.
0.5% (v/w) Pectin extracted for citrus or apple and polygalacturonic acid (PGA) purchased from Sigma-Aldrich was used as substrate for PelB-B2 to identify the substrate specificity. The highest enzyme activity was set as 100%. Triplicate experiments were carried out for every sample.
Kinetic studies
PelB-B2 was reacted with pectin extracted from citrus (Sigma-Aldrich) with different concentration of 0.025, 0.05, 0.1, 0.2, 0.4, 0.5, 0.8, and 1.0% (w/v), and the enzyme activity was measured under optimal pH and temperature. The kinetic parameters of PelB-B2 using pectin extracted from citrus as the substrate were evaluated by Lineweaver–Burk plot, and the Km and Vmax were calculated according to the kinetic equation.
Results
Pectinase activity of microorganisms isolated from orange peel
In this study, seven bacterial strains were isolated from orange peel and named as OR-B1–OR-B7, respectively. These microbes showed different pectinase activity. Among the seven strains, OR-B2 showed highest ability to degraded pectin substrate with enzyme activity of 0.45 U/mL (Fig. 1a). The 16S rDNA of the seven strains were amplified from genomes and the PCR products were verified by gel electrophoresis and sequenced at Invitrogen (Table S3). The 16S rDNA of OR-B2 was submitted to Genbank and the accession number was SUB6462140 OR-B2-16S MN606168. The sequence identity of OR-B2 with other microbes was then searched using BLAST and the phylogenetic analysis was performed using a neighbor-joining method in MEGA5. The results showed that OR-B2 showed more than 99% sequence identity to Bacillus sp., Therefore, OR-B2 was identified as a Bacillus sp. (Fig. 1b) In addition, strains besides OR-B2 were also identified by phylogenetic analysis (Fig. S1). B1 and B4 showed highest identity to Erwinia sp., B3 showed highest identity to Pantoeaagglomerans, B5 showed highest identity to Enterobacteraerogenes, B6 showed highest identity to Raoultella sp., and B7 showed highest identity to Klebsiellapneumoniae.
Enzyme activity of bacteria and the molecular identification of OR-B2. a Enzyme activity of pectinases produced by microbes isolated in this study. Error bars: SD from three replicates. b Unrooted neighbor-joining tree of strain OR-B2 and related genera based on 16S rDNA gene sequences. Note: the candidate highlight with red is the strain OR-B2 isolated in this study. Bootstrap values of 1000 replications are given at nodes
Cloning, expression, and features analysis of PelB-B2
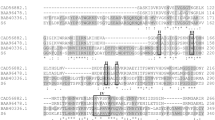
The pelB-B2 was amplified from genome of OR-B2 with specific primers and verified at Invitrogen. The nucleotide of pelB-B2 was submitted to Genbank and the accession number was given by the GenBank Direct Submission Staff with BankIt2276347 PelB MN614144. The full length of pelB-B2 was 1065 bp, encoding a protein of PelB-B2 with 354 amino acids and the pI/Mw was 9.21/38.34 kDa. The deduced PelB-B2 shares 99% identity to the orthologues found in Bacillusvelezensis (Fig. 2a). 3D model of PelB-B2 was constructed by SWISS-MODEL and the structure of PelB-B2 showed a typical para-β structure (Fig. 2b). The identity of PelB-B2 and the template for model building, a pectin lyase B (SMTL ID: 1qcx.1), was 38.71%. All these results supplied that PelB-B2 was a conserved pectate lyase which belongs to PelB superfamily (E.C. 4.2.2.2). Furthermore, PelB-B2 was successfully heterogeneous expressed in E.coli by recombined with the vector of pET22b. After induced cultured with 0.6 mM IPTG and purification by Ni-affinity chromatography, the recombinant PelB-B2 migrated as a single band of approximately 38 kDa in SDS-PAGE (Fig. 2c).
Structure analysis, purification, and 3D model construction of PelB-B2. a Phylogenetic tree for PelB homologues (NCBI code in parentheses) produced by OR-B2 or other microbes. Note: the candidate highlighted in red is the pectate lyase produced by OR-B2. Bootstrap values of 1000 replications are given at nodes. b Construction of 3D model of PelB-B2 by SWISS-MODEL. The template was a pectin lyase B in the database (SMTL ID: 1qcx.1) and the identity of PelB-B2 and the template was 38.71%. c Recombinant PelB-B2 from E.coli BL21 on an SDS-PAGE gel. Lanes: M, molecular standard; line 1: supernatant of crude extract from E.coli BL21 harboring pET22b; line 2: supernatant of crude extract from E.coli BL21 harboring pET-pelB-B2; line 3: purified recombinant PelB-B2 after Ni-affinity chromatography
Effect of pH and temperature on enzyme activity
After heterogeneous expression in E.coli, the activity of PelB-B2 was 198.18 U/mL. The optimal temperature and pH for PelB-B2 was 50 °C and 8.0 (Fig. 3a, b). In this study, we found that PelB-B2 showed good thermostability when stressed with heat shock at 40 °C, 50 °C or 60 °C for 1 h. After incubated for 10 min, the remained activity was more than 90% compared with the control. After incubated for 60 min, the enzyme activity remained more than 51% compared with the control (Fig. 3c). PelB-B2 also showed excellent pH stability between pH 6 and 10, with the remained activity more than 99.25%. The enzyme activity even remained approximately 40% when incubated in buffer with pH 4 or 11 for 1 h (Fig. 3d).
Enzymatic properties of purified PelB-B2. a Effect of temperature on PelB-B2. Activity was assayed in citric acid–Na2PO4 buffer with pH 7.0, while the reaction temperature was ranging from 30 to 50 °C. b Effect of pH on PelB-B2 activity. Activities at various pHs were assayed at 50 °C. c Thermostability of PelB-B2. Residual activity was assayed at 50 °C with pH 8.0 after pre-incubation at 40 °C, 50 °C, and 60 °C for different time. d pH stability of PelB-B2. Residual activity was assayed after incubation at various pHs for 1 h. Error bars SD from three replicates
Effect of chemicals and substrate on enzyme activity and enzyme kinetic assay
The effect of 15 metal ions, SDS, and EDTA on the enzyme activity was assayed. Among the 15 metal ions, Hg2+, Ag+, and Co2+ showed strong inhibition effect on PelB-B2 with approximately 90% enzyme activity reduction. On the other hand, addition of Ca2+ into the reaction system can significantly promote the enzyme activity of PelB-B2. For surfactant or chelant, SDS and EDTA both play a role as enzyme activity inhibitor (Fig. 4a). These results showed that PelB-B2 was a Ca2+-dependant pectate lyase.
Effect of metal ions, chemical reagents, and substrate on the enzyme activity and the enzyme kinetic of PelB-B2 with the substrate of pectin. a Effect of metal ions and chemical reagents on PelB-B2. Enzyme activity was assayed under reaction system with 10 mM metal ions or chemical reagents. Enzyme activity of PelB-B2 assayed with no addition of metal ions or chemical reagents was set as the control. Error bars: SD from three replicates. b Enzyme activity on different substrate. Using pectin extracted from citrus, pectin extracted from apple or PGA (polygalacturonic acid) as substrate, the enzyme activity was assayed at the optimal reaction condition. Error bars: SD from three replicates. c Enzyme kinetic of PelB-B2 with the substrate of pectin. PelB-B2 was reacted with pectin extracted from citrus with different concentration of 0.025, 0.05, 0.1, 0.2, 0.4, 0.5, 0.8, and 1.0% (w/v), and the enzyme activity was measured under optimal pH and temperature. The kinetic parameters of PelB-B2 reaction with pectin extracted from citrus were evaluated by Lineweaver–Burk plot and the Km and Vmax was calculated according to the kinetic equation
PelB-B2 showed different activity when reacted with different substrate. Three substrates including pectin extracted from citrus, pectin extracted from apple, and polygalacturonic acid (PGA) was used to assay the substrate specificity of PelB-B2. Result showed that pectin extracted from citrus was the optimal substrate for PelB-B2 among the three substrates (Fig. 4b).
Pectin extracted from citrus was used as the substrate to calculate the Km and Vmax for PelB-B2 (Fig. 4c). The Michaelis constant (Km) and maximum reaction rate (Vmax) of PelB-B2 were 1.64 g/L and 232.56 mol/(L min), respectively.
Discussion
Pectinases are widely applied in various industrial fields and microorganisms have been exploited by industry for pectinases production. Peel of citrus fruits such as lemon or grapefruit is pectin rich, which make it beneficial for growing pectinolytic microbes. In our study, the peel of fresh orange was used as raw material for isolating microorganisms. This study sought to broaden the source of pectinolytic microbes and provide a theoretical foundation for pectinases production and application.
Bacillus sp. was widely studied for its beneficial products. Studies have indicated that Bacillus sp. harbors potential to produce various enzymes, such as alginate lyase, poly (butylene adipate-co-terephthalate) hydrolase, protease, cellulose, endoglucanase, and xylanase (Chen et al. 2018; Hero et al. 2017; Muroi et al. 2017; Sun et al. 2017; Yildirim et al. 2017). In addition, Bacillus sp. has been studied recently also as a pectin-degrading bacteria. Bacillussubtilis SAV-21 isolated from fruit and vegetable market waste soil in India produce pectin-degrading enzymes with 3315 U/gds (pectinase) and 10.5 U/gds (pectin lyase), and the optimal pH and temperature for reaction was 4.0 and 35 °C for both the two pectinase (Kaur and Gupta 2017). Pectate lyase BsPel-PB1 produced by Bacillussubtilis PB1 can degrade citrus pectin as substrate at optimal temperature of 50 °C and optimal pH 9.5, and the Km and Vmax were 0.312 mg/mL and 1248 U/mL (Zhou et al. 2017b). Bacillussafensis M35, Bacillusaltitudinis R31, and B.altitudinis J208 can produce pectin lyases and the maximum activity was obtained at pH 10.0 and 60 °C (Thite and Nerurkar 2018).
In this study, seven bacteria were isolated from orange peels. The activity of pectinase was determined by the DNS method while using pectin as substrate. Results showed that the isolates can produce pectinase. One bacterium (Bacillus sp., OR-B2) produced pectinase with the highest activity among the isolates with 0.45 U/mL. A gene designed as pelB-B2 which encoding a pectate lyase in OR-B2, was obtained and heterologous expressed in E.coli successfully. PelB-B2 was considered as a member of pectate lyase family belongs to PelB superfamily after bioinformatics analysis. The maximum activity of recombined pectate lyase was 198.18 U/mL and the optimal temperature for PelB-B2 was 8.0 and 50 °C. The optimal reaction condition for PelB-B2 is close to the pectate lyase BsPel-PB1, with the optimal pH and temperature with 9.5 and 50 °C (Zhou et al. 2017b). Importantly, the PelB-B2 has excellent pH stability and thermostability. After incubated with buffer with pH ranging from 6 to 10 for 1 h, the enzyme activity still remained more than 99.25%. After incubated at 40 °C, 50 °C or 60 °C for 10 min, the remained activity was more than 90% compared with the control. After incubated for 60 min, the enzyme activity still remained more than 51%. All of these properties of PelB-B2 reveal that this pectate lyase has good potential for industrial application.
Chemicals affected enzyme activity. As the results showed in this study, Ca2+ accelerates the enzyme activity, while Ni+, Mn2+, Zn2+, Hg2+, Ag+, and Co2+ inhibited the enzyme activity. This is in line with forecast, since Ca2+ was proven to be the required for pectinase reaction in previous study (Peng et al. 2011; Zhou et al. 2017a). Substrate specificity is important for the application of pectinase. PelB-B2 has maximum activity when reaction with citrus pectin, and then followed by apple pectin and PGA. This may contribute to the different structure of these substrates. This result is different with BacPelA, which showed 114.7% activity on apple pectin relative to PGA (taken as 100.0%), whereas activity on citrus pectin was only 60.7% relative to that on PGA (Zhou et al. 2017a). Various pectinase were heterogeneous expressed in E.coli recently years. A metagenome-derived alkaline pectate lyase designed as PelB was purified and the enzyme kinetic was assayed. The Km and Vmax of PelB were 1.65 g/L and 63.34 mol/(L min) (Wang et al. 2014). Compared to the PelB, PelB-B2 showed that higher substrate affinity with Km and Vmax of PelB-B2 was 1.64 g/L and 232.56 mol/(L min).
Overall, pectinase is widely required in various industrial fields, and efforts directed to isolating microorganisms which can produce pectinase efficiently are important. The bacteria isolated in this study showed great potential for commercial pectinase production.
Change history
15 May 2020
In the original article,
References
Agarwal RP, Thanvi I, Vachhani G, Kochar DK, Rastogi A (1999) Exercise induced proteinuria as an early indicator of diabetic nephropathy. J Assoc Physicians India 46:1127–1128
Alimardani-Theuil P, Gainvors-Claisse A, Duchiron F (2011) Yeasts: an attractive source of pectinases-from gene expression to potential applications: a review. Process Biochem 46:1525–1537. https://doi.org/10.1016/j.procbio.2011.05.010
Alkorta I, GarbisuC LMJ, Serra JL (1998) Industrial applications of pectic enzymes: a review. Process Biochem 33:21–28
Anand G, Yadav S, Yadav D (2017) Production, purification and biochemical characterization of an exo-polygalacturonase from Aspergillusniger MTCC 478 suitable for clarification of orange juice. Biotech 7:122. https://doi.org/10.1007/s13205-017-0760-3
Blanco P, Sieiro C, Reboredo NM, Villa TG (1998) Cloning, molecular characterization, and expression of an endo-polygalacturonase-encoding gene from Saccharomycescerevisiae IM1-8b. Fems Microbiol Lett 164:249–255. https://doi.org/10.1111/j1574-6968.1998.tb13094.x
Chen P, Zhu Y, Men Y, Zeng Y, Sun Y (2018) Purification and characterization of a novel alginate lyase from the marine Bacterium bacillus sp. Alg07. Mar Drugs 16:86. https://doi.org/10.3390/md16030086
Gainvors A, Karam N, Lequart C, Belarbi A (1994) Use of Saccharomycescerevisiae for the clarification of fruit juices. Biotechnol Lett 16:1329–1334. https://doi.org/10.1007/BF00149641
Guan Y, Yin D, Du X, Ye X (2018a) Functional metabolomics approach reveals the reduced biosynthesis of fatty acids and TCA cycle is required for pectinase activity in Bacilluslicheniformis. J Ind Microbiol Biotechnol 45:951–960. https://doi.org/10.1007/s10295-018-2071-z
Guan Y, Yin D, Du X, Ye X (2018b) Metabolomics approach used for understanding temperature-related pectinase activity in Bacillus licheniformis DY2. Fems Microbiol Lett 365:23. https://doi.org/10.1093/femsle/fny255
Henrissat B, Bairoch A (1993) New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 293:781–788. https://doi.org/10.1042/bj2930781
Hero JS, Pisa JH, Perotti NI, Romero CM, Martínez MA (2017) Endoglucanase and xylanase production by Bacillus sp. AR03 in Co-Culture. Prep Biochem Biotechnol 47:589–596. https://doi.org/10.1080/10826068.2017.1280826
Hugouvieux-Cotte-Pattat NCG, Nasser W, Reverchon S (1996) Regulation of pectinolysisin Erwiniachrysanthemi. Annu Rev Microbiol 50:213–257. https://doi.org/10.1146/annurev.micro.50.1.213
Jayani RS, Saxena S, Gupta R (2005) Microbial pectinolytic enzymes: a review. Process Biochem 40:2931–2944. https://doi.org/10.1016/j.procbio.2005.03.026
Ju Z (2011) Research progress on pectinase from bacteria. Biotechnol Bull 2:56–60
Kashyap DR, Vohra PK, Chopra S, Tewari R (2001) Applications of pectinases in the commercial sector: a review. Bioresour Technol 77:215–227
Kaur SJ, Gupta VK (2017) Production of pectinolytic enzymes pectinase and pectin lyase by Bacillussubtilis SAV-21 in solid state fermentation. Ann Microbiol 67:333–342. https://doi.org/10.1007/s13213-017-1264-4
Kertesz ZI (1955) Pectic enzymes. Methods Enzymol 1:158–166
Lerouxel O, Cavalier DM, Liepman AH, Keegstra K (2006) Biosynthesis of plant cell wall polysaccharides—a complex process. Curr Opin Plant Biol 9:621–630
Li ZM, Bai ZH, Zhang BG, Li BJ, Jin B, Zhang M, Lin F, Zhang HX (2012) Purification and characterization of alkaline pectin lyase from a newly isolated Bacillusclausii and its application in elicitation of plant disease resistance. Appl Biochem Biotechnol 167:2241–2256. https://doi.org/10.1007/s12010-012-9758-9
Muroi F, Tachibana Y, Soulenthone P, Yamamoto K, Mizuno T, Sakurai T, Kobayashi Y, Kasuya KI (2017) Characterization of a poly(butylene adipate-co-terephthalate) hydrolase from the aerobic mesophilic bacterium Bacilluspumilus. Polym Degrad Stab 137:11–22. https://doi.org/10.1016/j.polymdegradstab.2017.01.006
Peng Y, Meng K, Luo H, Shi P, Huang H, Bai Y, Yang P, Yao B (2011) A novel low-temperature active alkaline pectate lyase from Klebsiella sp. Y1 with potential in textile industry. Process Biochem 46:1921–1926. https://doi.org/10.1016/j.procbio.2011.06.023
Pretel MT, Lozano P, Riquelme F, Romojaro F (1997) Pectic enzymes in fresh fruit processing: optimization of enzymic peeling of oranges. Process Biochem 32:43–49
Rashmi RMK, Sneha G, Shabana S, Syama A, Radhika VS (2008) Partial purification and biochemical characterization of extracellular pectinase from Aspergillusniger isolated from groundnut seeds. J Appl Biosci 9:378–384
Reginatto C, Rossi C, Miglioranza BG, Santos MD, Meneghel L, Silveira MMD, Malvessi E (2017) Pectinase production by Aspergillusniger LB-02-SF is influenced by the culture medium composition and the addition of the enzyme inducer after biomass growth. Process Biochem 58:1–8. https://doi.org/10.1016/j.procbio.2017.04.018
Roosdiana A, Prasetyawan S, Mahdi C, Sutrisno X (2013) Production and characterization of Bacillus firmus pectinase. J Pure Appl Chem Res 2:35–41. https://doi.org/10.21776/ub.jpacr.2013.002.01.111
Rubio MJ, Roberto C, Rubio FA, Fernando RJ (2005) Continuous enzymatic precooking for the production of an instant corn flour for snack and tortilla. EP 1:242–545.
Sakai T, Sakamoto T, Hallaert J, Vandamme EJ (1993) Pectin, pectinase, and protopectinase—production, properties, and applications. Adv Appl Microbiol 39:213–294
Saranraj P, Naidu MA (2014) Microbial pectinases: a review. Glob J Trad Med Syst 3:1–9
Siddiqui MA, Pande V, Arif M (2012) Production, purification, and characterization of polygalacturonase from Rhizomucorpusillus isolated from decomposting orange peels. Enzyme Res 9:138634. https://doi.org/10.1155/2012/138634
Soumpourou E, Lakovidis M, Chartrain L, Lyall V, Thomas CM (2007) The Solanumpimpinellifolium Cf-ECP1 and Cf-ECP4 genes for resistance to Cladosporiumfulvam are located at the Milky Way locus on the short arm of chromosome 1. Theor Appl Genet 115:1127–1136. https://doi.org/10.1007/s00122-007-0638-6
Sun L, Cao J, Liu Y, Wang J, Guo P, Wang Z (2017) Gene cloning and expression of cellulase of Bacillusamyloliquefaciens isolated from the cecum of goose. Anim Biotechnol 28:74–82. https://doi.org/10.1080/10495398.2016.1205594
Tapre AR, Jain RK (2014) Pectinases: enzymes for fruit processing industry. Int Food Res J 21:447–453
Thite VS, Nerurkar AS (2018) Physicochemical characterization of pectinase activity from Bacillus spp. and their accessory role in synergism with crude xylanase and commercial cellulase in enzyme cocktail mediated saccharification of agrowaste biomass. J Appl Microbiol 124:1147–1163. https://doi.org/10.1111/jam.13718
Wang H, Li X, Ma Y, Song J (2014) Characterization and high-level expression of a metagenome-derived alkaline pectate lyase in recombinant Escherichiacoli. Process Biochem 49:69–76
Yildirim V, Baltaci MO, Ozgencli I, Şişecioğlu M, Adiguzel A, Adiguzel G (2017) Purification and biochemical characterization of a novel thermostable serine alkaline protease from Aeribacilluspallidus C10: a potential additive for detergents. J Enzyme Inhib Med Chem 32:468–477. https://doi.org/10.1080/14756366.2016.1261131
Zhou C, Xue Y, Ma Y (2017a) Cloning, evaluation, and high-level expression of a thermo-alkaline pectate lyase from alkaliphilic Bacillusclausii with potential in ramie degumming. Appl Microbiol Biotechnol 101:3663–3676. https://doi.org/10.1007/s00253-017-8110-2
Zhou M, Wu J, Wang T, Gao L, Yin H, Xin L (2017b) The purification and characterization of a novel alkali-stable pectate lyase produced by Bacillussubtilis PB1. World J Microbiol Biotechnol 33:190. https://doi.org/10.1007/s11274-017-2357-8
Acknowledgements
The authors would like to thank Gale A. Kirking at English Editorial Services, s.r.o. for language correction and additional assistance in improving the text.
Funding
This work was supported by National Natural Science Foundation of China (31701844), Natural Science Foundation of Fujian province (2018J05057), the Foundation of Fujian Educational Bureau (JAT170078), Fuzhou University Testing Fund of precious apparatus (2019T018), and with institutional support RVO: 60077344 of the Biology Centre CAS, Institute of Entomology, Czech Republic.
Author information
Authors and Affiliations
Contributions
YG, DW, CL, YZ, IG, and XY contributed equally to the study design, collection of data, development of the sampling, analyses, interpretation of results, and preparation of the paper. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Ethical statement
No special permit was required for this work.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Guan, Y., Wang, D., Lv, C. et al. Archives of microbiology: screening of pectinase-producing bacteria from citrus peel and characterization of a recombinant pectate lyase with applied potential. Arch Microbiol 202, 1005–1013 (2020). https://doi.org/10.1007/s00203-020-01807-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-01807-0