Abstract
Introduction & hypothesis
Previous studies have suggested that women with urinary incontinence have an altered urinary microbiome. We hypothesized that the microbiome in women with mixed urinary incontinence (MUI) differed from controls and tested this hypothesis using bacterial gene sequencing techniques.
Methods
This multicenter study compared the urinary microbiome in women with MUI and similarly aged controls. Catheterized urine samples were obtained; v4–6 regions of the 16S rRNA gene were sequenced to identify bacteria. Bacterial predominance (> 50% of an individual’s genera) was compared between MUI and controls. Bacterial sequences were categorized into “community types” using Dirichlet multinomial mixture (DMM) methods. Generalized linear mixed models predicted MUI/control status based on clinical characteristics and community type. Post-hoc analyses were performed in women < 51 and ≥ 51 years. Sample size estimates required 200 samples to detect a 20% difference in Lactobacillus predominance with P < 0.05.
Results
Of 212 samples, 97.6% were analyzed (123 MUI/84 controls, mean age 53 ± 11 years). Overall Lactobacillus predominance did not differ between MUI and controls (45/123 = 36.6% vs. 36/84 = 42.9%, P = 0.36). DMM analyses revealed six community types; communities differed by age (P = 0.001). A High-Lactobacillus (89.2% Lactobacillus) community had a greater proportion of controls (19/84 = 22.6%, MUI 11/123 = 8.9%). Overall, bacterial community types did not differ in MUI and controls. However, post-hoc analysis of women < 51 years found that bacterial community types distinguished MUI from controls (P = 0.041); Moderate-Lactobacillus (aOR 7.78, CI 1.85–32.62) and Mixed (aOR 7.10, CI 1.32–38.10) community types were associated with MUI. Community types did not differentiate MUI and controls in women ≥ 51 years (P = 0.94).
Conclusions
Women with MUI and controls did not differ in overall Lactobacillus predominance. In younger women, urinary bacterial community types differentiated MUI from controls.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mixed urinary incontinence (MUI) is the involuntary leakage of urine associated with both urgency and stress provocation. Of all incontinence types, MUI is the most refractory to treatment, its pathophysiology the least understood and therapies the least standardized, posing significant treatment challenges [1]. Women with MUI face a therapeutic dilemma because of the dual nature of their condition; treatment of stress urinary incontinence (SUI) symptoms may exacerbate urgency while treatment of urgency urinary incontinence (UUI) symptoms may worsen SUI. The study, Effects of Surgical Treatment Enhanced with Exercise for Mixed Urinary Incontinence (ESTEEM), was designed to compare stress, urgency and MUI symptom outcomes in an MUI population treated with surgery alone versus surgery and behavioral/pelvic floor therapy [2]. The ESTEEM study’s population provided a singular opportunity to evaluate the urinary microbiome as a biologically plausible contributor to MUI.
Prior work has established that the female urinary microbiome is associated with UUI [3,4,5]. With the advent of precise bacterial DNA testing, it is clear that a female urinary microbiome exists and that it may have a role in UUI treatment response [3, 5]. As the urinary microbiome may have a role in UUI, the question arises whether the urinary microbiome may also play a role in MUI, an even more severe form of incontinence that includes UUI. The current study builds on the existing data regarding the relationship between the urinary microbiome and UUI and expands its scope to include those participants with MUI. This is clinically relevant, as MUI is enigmatic and difficult to treat. For example, improvement of the UUI component of MUI following stress incontinence surgery has never been clearly understood. If change in the MUI microbiome occurs, particularly in the microbiota associated with UUI, clinicians would then have a potential explanation for improvement of the UUI component of MUI following stress incontinence surgery. This MUI microbiome study’s over-arching goal is to provide insight into MUI’s underlying pathophysiology and to gain insight into MUI’s varying clinical responses following treatment.
In the current study, we evaluated the urinary microbiome in women with MUI at baseline compared with asymptomatic, similarly aged controls. Subsequent work may determine whether MUI microbiome characteristics have a role in MUI treatment response. Based on previous studies that found that women with UUI have a more diverse microbiome and decreased Lactobacillus compared with controls [3, 4], we hypothesized that the microbiome of women with MUI differed from asymptomatic controls in both overall Lactobacillus predominance and microbial community types.
Methods
Study population & design
This multi-site, IRB-approved, observational study evaluated the urinary microbiome in women with and without MUI. The methodology for this study has been previously published [6]. We excluded patients who had used oral or intravenous antibiotics within the last month, who had used vaginal antibiotics within the prior 7 days and who were currently using vaginal probiotics or spermicides. Furthermore, we discouraged participants from engaging in vaginal intercourse/douching/using vaginal sprays or wipes for 48 h prior to study visits, and specimens were collected at least 48 h following cessation of menses. MUI participants consisted of a subset of women enrolled in a randomized trial of MUI treatment comparing the efficacy of mid-urethral sling (MUS) surgery alone to MUS surgery and behavioral therapy (NCT# 01959347) [2]. For this supplementary study, we also recruited controls who were of similar age as the MUI cases, as described in the methods paper [6].
Participant characteristics and baseline symptom questionnaire data were obtained including the Urogenital Distress Inventory (UDI) to measure incontinence symptom severity in the MUI participants [7]. The UDI irritative subscale score reflected UUI, the UDI stress subscale score reflected SUI, and the total UDI score reflected MUI severity. The parent study required all MUI participants to have at least “moderate bother” on the UDI for the UUI and SUI items: “Do you usually experience leakage associated with a feeling of urgency…?” and “Do you usually experience urinary leakage related to coughing, sneezing, or laughing?” Controls had slight or no incontinence (Incontinence Severity Index scores ≤ 2) and no significant overactive bladder symptoms (Overactive Bladder Awareness Scores < 8) [8, 9].
As described previously [6], catheterized urine (5–10 ml) was placed in commercially available tubes containing Assay Assure® DNA protectant after screening negative for infection (absence of dysuria and negative dipstick analysis with nitrites/leukocyte esterase ≤ trace). An additional 4 ml of urine was placed in BDVacutainers® (Sierra Molecular Corp., Incline Village, NV, USA) for routine urine culture. Tubes were shipped on cold pack to the UNM CTSC Laboratory and received within 24 h. A clinical laboratory (Tricore Reference Laboratories, Albuquerque, NM, USA) performed routine urine culture, and study investigators at the UNM CTSC Laboratory performed immediate DNA extraction. Following DNA extraction, samples were stored at −80 °C until completion of specimen collection at all sites. Investigators then performed polymerase chain reaction (PCR) amplification and 16S sequencing.
Laboratory methods
By design, the study masked laboratory investigators to samples’ MUI or control status. DNA isolation procedures, library preparation, sequencing techniques, and analytics have been previously described in detail [6]. As noted in our methods manuscript, variable regions 4–6 of the 16S rRNA gene were amplified by PCR using primers 515F and 1114R with the addition of Illumina® Nextera linker sequences [6, 10]. If a single PCR amplicon of the correct size was not identified during gel electrophoresis (Agilent Bioanalyzer 1000), DNA was re-isolated from remaining urine and the sample was reprocessed. If upon resampling the sample failed again, it was excluded from further processing and analysis. We were unable to recover suitable DNA from five urine samples collected in this study. A second PCR of ten cycles was used to complete the Illumina adapter sequence and to add dual 8-nucleotide index sequences. Completed sequencing libraries were purified, and unused primers were removed using Agencourt AMPure XP beads. After quantification and purity assessment by Qubit fluorometeric and gel electrophoresis bioanalysis (Agilent Bioanalyzer 1000), samples were pooled in equimolar ratios for sequencing.
Sequencing utilized version 3 sequencing chemistry and paired 300-bp sequencing reads on the Illumina MiSeq®. Each batch of 96 samples included 88 unique patient samples, 6 repeated samples to assess sample reproducibility and 2 negative controls, consisting of a no-DNA extraction control and a no-template PCR control that underwent all PCR, library preparation and sequencing steps. With the given depth of sequencing in this study, no-template negative control samples produced a mean of 3077 (range 2410–4242) classified sequencing reads; consequently, to ensure a low likelihood of false-positive results, we used a threshold of > 12,000 classified sequencing reads for samples to be included in the final analysis.
Bioinformatics analysis
Overlapping sequencing reads were combined into a single read and operational taxonomic unit classifications were completed and compared via a high-performance implementation of the Ribosomal Database Project (RDP) Classifier [11] to a curated version of the May 2013 Greengenes database using the Illumina® BaseSpace 16S Metagenomics App (v1.01). Specifically, the Greengenes data were modified to remove 16S sequences with a length was below 1250 bp, entries that had more than 50 wobble bases, and ambiguous epithets and partial classifications. The database includes 16S rRNA sequences representing 33 phyla, 74 classes, 148 orders, 321 families, 1086 genera and 6466 distinct species. The use of the Illumina BaseSpace 16S Metagenomics pipeline was an effort to increase the reproducibility and comparability of our data by using a standardized bioinformatic process that can be easily replicated by other researchers. Urinary bacteria were identified to the genus level in this portion of study analysis.
Statistical analysis
The primary aim of this study was to examine the difference in Lactobacillus predominance between MUI and controls. Predominance was defined as a sample in which a specific genus constituted > 50% of an individual’s taxonomic community. The overall proportions of women with Lactobacillus predominance were compared between MUI and controls using chi-square tests. Sample size analysis, with 80% power and alpha of 0.05, indicated that urine specimens from 200 women (120 MUI, 80 controls) would be required to detect a 20% difference in Lactobacillus predominance [6].
The secondary aim was to compare the bacterial taxa between MUI and controls utilizing Dirichlet Multinomial Mixture (DMM) modeling, which identified bacterial communities across MUI cases and controls [12, 13]. The DMM method clustered samples based on the relative abundance of identified microbiota. DMM methodology effectively manages the large data dimensionality associated with microbiome analyses. This facilitated use of sequencing results in the form of bacterial community types and clinical variables in multivariable analyses. Prior to the quantitative analysis, investigators reviewed community types for clinical relevance. Chi-square testing was used to evaluate whether the DMM community types differentiated participants with MUI from controls.
Participant characteristics were compared across the DMM community types in bivariate analysis using chi-square tests and analysis of variance (ANOVA) models. A multivariable generalized linear mixed model with a logit link was constructed to predict the MUI or control status based on DMM community type, with demographic and medical history variables significantly associated with DMM community type in bivariate analysis (p < 0.05) and variables associated with MUI versus control status and other variables of clinical significance as covariates. The model also included a random effect for clinical site.
The study recruited controls of similar ages to MUI participants because of the known age-dependent differences in the female vaginal microbiome and the potential influence of age upon the human microbiome in general [14, 15]. Since the six DMM community types differed by age, separate post hoc sub-analyses were performed in women < 51 and ≥ 51 years, the median age of menopause [16]. Self-reported menopausal status was not used to discriminate community types, as approximately 20% of women in this study did not know their menopausal status.
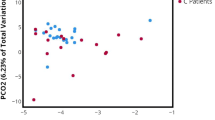
Overall alpha and beta diversities were compared between MUI and controls. Alpha diversity (Shannon index as a measure of bacterial genus diversity within individuals) was evaluated using a similar linear mixed model to those described previously, with the following covariates: DMM community type, MUI versus control, age, BMI, smoking status and ethnicity. Beta diversity (provides a measure of dissimilarity between individuals) was evaluated using ordination analysis, including nonmetric multidimensional scaling distance-based redundancy analysis and multivariate homogeneity of group dispersions [17,18,19]. Additionally, multivariate analysis of variance (MANOVA) association testing evaluated beta diversity using the same covariates as the alpha diversity model [20].
Results
Two hundred twelve women (128 MUI, 84 controls) contributed specimens for this study from January 2015 through April 2016. DNA was successfully extracted, sequenced and analyzed from 97.6% (207/212) of the specimens (123/128 MUI, 84/84 controls). Participants’ mean age was 53 years and did not differ between MUI and controls. MUI participants had higher mean BMI, were more commonly Hispanic, more commonly had recurrent UTIs and more commonly used vaginal estrogen compared with controls (Table 1). Bacterial DNA sequencing resulted in a median sequencing read depth per sample of 80,949 reads (range 12,517–450,993). Bioinformatic processing of the 16S rRNA sequences resulted in the classification of 28 phyla, 60 classes, 82 orders, 191 families and 581 genera.
The proportion of women with Lactobacillus-predominant microbiota (> 50% Lactobacillus in their samples) did not differ between MUI and controls [45/123 (36.6%), 36/84 (42.9%), P = 0.364]. This was true for both younger and older women [< 51 years: 27/57 (47.4%) MUI; 20/38 (52.6%) controls, P = 0.615; ≥ 51 years: 18/66 (27.3%) MUI, 16/46 (34.8%) controls, P = 0.395]. Although Lactobacillus predominance did not distinguish between MUI and controls, its predominance differed between younger and older women overall, with higher Lactobacillus predominance in younger women [47/95 (49.5%) in < 51 years; 34/112 (30.4%) in ≥ 51 years, P = 0.005].
DMM analysis identified six DMM community types. Figure 1 qualitatively and quantitatively describes the composition of the six communities. In bivariate analysis, the proportion of women belonging to the DMM community types differed between MUI and controls (P = 0.032) (Table 1). Additional bivariate analyses of clinical characteristics found that DMM communities also differed in age and smoking status (Table 2).
On initial multivariable analysis, higher BMI and Hispanic ethnicity remained significantly associated with MUI (Table 3). Although history of recurrent UTIs and the use of vaginal estrogen were significant on bivariate analysis, there were too few subjects in these groups to reliably include them as separate covariates in this model. There was no significant effect overall in community type composition between MUI and controls (P = 0.196) (Table 3). However, as bivariate analysis indicated that DMM communities varied by age, post hoc multivariable analysis of women < 51 years of age (controlling for DMM community types, smoking status, ethnicity, age and BMI) revealed that DMM community type and BMI remained associated with MUI (Table 4). In this multivariable model, the community with a very high proportion (89.2%) of Lactobacillus (High-Lactobacillus community type 1, Fig.1) served as the comparator community as it had the highest proportion of controls. In these younger women, those with the mixed genera community type (Mixed community type 2, Fig.1) or those with the moderate Lactobacillus community type (Moderate-Lactobacillus community type 6, composed on average of 61.1% Lactobacillus, Fig. 1) had approximately 7- to 8-fold increased odds of having MUI (Table 4). Post-hoc multivariable analysis of women ≥ 51 years revealed that bacterial community types did not differ between MUI and controls (P = 0.938); only body mass index (P = 0.010) remained independently associated with MUI (Appendix Table 1).
Alpha diversity (Shannon and Simpson Indexes) did not differ between MUI and controls, but did differ based on DMM community types (Table 5). The High-Lactobacillus community type 1 was the most homogeneous and least diverse, with all other community types being more diverse (p < 0.001). Beta diversity also did not differ between the MUI and control groups (Appendix Fig. 1).
Discussion
We found no difference in Lactobacillus predominance between women with MUI compared with similarly aged controls. However, analysis of community types resulting from DMM clustering of individuals into groups based on the entire composition of the urinary microbiome revealed that several urinary bacterial communities significantly differed between MUI and controls in women < 51 years of age.
DMM methodology concomitantly uses the presence and abundance of all genera across both cases and controls to identify groups with similar microbial communities [14, 15]. This method accommodates data with rare or sparse genera and varying numbers and types of identified genera across samples [12]. The DMM community types, derived based on commonality of genera, served as independent variables in multivariable analysis along with clinical parameters such as age and BMI. Other clustering methods measure pairwise distances between observations based on a single metric derived from the differences in individual genera (for example, the maximum or minimum difference). Instead of using hierarchical clustering of individuals based on these metrics, we chose to use DMM methods to more fully account for the diversity and particular composition of the microbiome [12, 13].
Six urinary community types were identified by DMM analysis. Communities differed by age, smoking history and presence of MUI. We found that the High-Lactobacillus community (89% Lactobacillus) (community type 1) had the largest proportion of asymptomatic controls, while two other microbial communities were associated with MUI. In younger women, both BMI and specific DMM communities were associated with MUI. Compared with the High-Lactobacillus community type 1, a Mixed (low-Lactobacillus, community type 2) and a Moderate-Lactobacillus community type 6 were associated with 7- to 8-fold higher odds of MUI. These results are similar to those of another female urinary microbiome study, which noted that the UUI microbiome was characterized by decreased Lactobacillus and increased Gardnerella abundance, characteristics comparable to our MUI Mixed DMM community type 2 [4]. Thus, two research groups using different analytic techniques found congruent results in women with UUI and MUI. These findings suggest that Lactobacillus may be associated with continence, whereas Gardnerella may be associated with incontinence, and the proportions or combinations of these bacteria may influence urinary symptoms, particularly in younger women.
In older women, only BMI, not microbiome communities, remained associated with MUI in multivariable analysis. The differential impact of age on urinary microbiome findings underscores the importance of recruitment of similarly aged, asymptomatic controls. The effect of age on microbial communities may contribute to treatment response and disease severity in UI. In the vaginal microbiome, Lactobacillus dominates the microbiota of premenopausal women (83%) compared with dominance in only 54% of post-menopausal women [14]. Similarly, the current study found that in the urinary microbiome, Lactobacillus dominance in women in women < 51 was greater than in women ≥ 51 years. Although studies of the relationship between the urinary and vaginal microbiomes are scarce, the possibility that the two microbiomes may be linked will be investigated in a planned future analysis.
A more recent culture-based study that compared women with overactive bladder and healthy controls also demonstrated differences in bacterial genera. Lactobacillus was found less commonly in women with overactive bladder [21]. Our study also found that a highly predominant Lactobacillus community (High-Lactobacillus, community type 1) was more common in controls, while women with MUI (Mixed and Moderate-Lactobacillus community types 2 and 6) more commonly had communities with lesser proportions of Lactobacillus. In aggregate, these studies support the concept that Lactobacillus may be associated with lack of urinary symptoms. Importantly, Lactobacillus abundance alone did not distinguish between the absence and presence of urinary symptoms. The proportion of Lactobacillus as well as the combination of Lactobacillus, Gardnerella and other genera (such as Prevotella, Serratia, Eschericia, Streptococcus and Tepidomonas) may distinguish those with or without urinary symptoms. For example, despite the Lactobacillus predominance in the Moderate-Lactobacillus community type 6, Gardnerella, in combination with other genera, more commonly represented MUI. The current study suggests that Lactobacillus occurrence or predominance may not be the only predictor of urinary symptoms, but that Lactobacillus, in addition to combinations and proportions of other bacterial taxa, may influence MUI communities and the MUI microbiome.
Thomas-White recently evaluated the microbiome of women undergoing SUI surgery and found no association between SUI symptoms and microbiome status [22]. The current study could not attribute the observed differences in the MUI microbiome to UUI or SUI predominance, but findings from Thomas-White suggest that the driving influence on the MUI microbiome may be the UUI component [22]. Those investigators also reported that UUI symptoms were associated with BMI and menopausal status. Our findings, too, found that BMI and older age (as a reflection of menopausal status) were independently associated with MUI. Our study, similar to reports from Pearce [4] and Karstens [23], found that overall microbiome diversity measures did not differ between MUI and controls. We did find, however, that the community with the least alpha diversity (community type 1) was most commonly found in controls, whereas all other communities, including those representative of MUI (community types 2 and 6), were more heterogeneous. This is consistent with microbiome studies from other body regions, which have reported that increasingly heterogeneous communities may be associated with disturbed habitats [12].
Strengths of this study include strategies to improve generalizability and minimize bias: rigorous definitions distinguishing MUI cases and controls, a well-characterized and age-matched asymptomatic control group recruited from multiple geographic sites, the use of state-of-the-art metagenomics and masking of laboratory investigators to case/control status. The analysis was further strengthened by the use of DMM to decrease data dimensionality and facilitate multivariable analysis. Limitations include its analysis to the genus rather than species level; in our future work comparing the urinary and vaginal microbiomes we will analyze to the species level. The use of a group demarcation of 51 years may not precisely reflect the age of menopause. We chose this cutoof point because self-identified menopausal status was missing in approximately 20% of our participants and self-identification of menopause misclassifies 30–40% of women [24, 25]. Finally, given that the analysis characterizing the MUI microbiome was complex and numbers within the final six DMM groups were relatively small, our findings will require confirmation by studies with larger cohorts.
In summary, Lactobacillus predominance did not differ between MUI and controls. We did find that although membership in distinct DMM communities did not distinguish between MUI and controls overall, it did distinguish between MUI and controls in younger women < 51 years. Younger MUI participants more commonly had Moderate-Lactobacillus or Mixed communities rather than a High-Lactobacillus community. Whether the high preponderance of health-associated bacteria (Lactobacillus) or the shift in the balance of MUI-associated bacteria (e.g., Gardnerella, Prevotella) contributes to bladder symptoms warrants further investigation. Furthermore, whether these communities are found to be predictive of treatment success in these participants of the parent trial is yet to be determined [6].
References
Minassian VA, Bazi T, Stewart WF. Clinical epidemiological insights into urinary incontinence. Int Urogynecol J. 2017;28:687–96.2.
Sung VW, Borello-France D, Dunivan G, et al. Methods for a multicenter randomized trial for mixed urinary incontinence: rationale and patient- centeredness of the ESTEEM trial. Int Urogynecol J. 2016;27:1479–90.
Brubaker L, Wolfe AJ. The female urinary microbiota/microbiome: clinical and research implications. Rambam Maimonides Med J 2017;8.
Pearce MM, Hilt EE, Rosenfeld AB, et al. The female urinary microbiome: a comparison of women with and without urgency urinary incontinence. mBio. 2014;5:e01283–14.
Thomas-White KJ, Hilt EE, Fok C, et al. Incontinence medication response relates to the female urinary microbiota. Int Urogynecol J. 2016;27:723–33.
Komesu YM, Richter HE, Dinwiddie DL, et al. Methodology for a vaginal and urinary microbiome study in women with mixed urinary incontinence. Int Urogynecol J. 2017;28:711–20.
Shumaker SA, Wyman JF, Uebersax JS, McClish D, Fantl JA. Health-related quality of life measures for women with urinary incontinence: the incontinence impact questionnaire and the urogenital distress inventory. Continence program in women (CPW) research group. Qual Life Res. 1994;3:291–306.
Coyne KS, Margolis MK, Bavendam T, Roberts R, Elinoff V. Validation of a 3-item OAB awareness tool. Int J Clin Pract. 2011;65:219–24.
Sandvik H, Seim A, Vanvik A. Hunskaar S. A severity index for epidemiological surveys of female urinary incontinence: comparison with 48-hour pad-weighing tests. Neurourol Urodyn. 2000;19:137–45.
Kumar PS, Brooker MR, Dowd SE, Camerlengo T. Target region selection is a critical determinant of community fingerprints generated by 16S pyrosequencing. PLoS One. 2011;6(6):e20956.
Wang Q, Garrity GM, Tiedje JM, Cole JR. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol. 2007;73(16):5261–7.
Holmes I, Harris K, Quince C. Dirichlet multinomial mixtures: generative models for microbial metagenomics. PLoS On3. 2012;7(2):e30126.
La Rosa PS, Brooks JP, Deych E, Boone EL, Edwards DJ, Wang Q, et al. Hypothesis testing and power calculations for taxonomic-based human Mcrobiome data. PLoS One. 2012;7(12):e52708.
Brotman RM, Shardel MD, Gajer P, Fadrosh D, Change K, Silver MI, et al. Association between the vaginal microbiota, menopause status and signs of vulvovaginal atrophy. Menopause. 2013;21(5):1–9.
Whiteside SA, Razvi H, Dave S, Reid G, Burton JP. The microbiome of the urinary tract—a role beyond infection. Nat. RevUrol. 2015;12:81–90.
Practice Bulletin No ACOG. 141: management of menopausal symptoms. Obstet Gynecol. 2014;123:202–16.
Minchin PR. An evaluation of the relative robustness of techniques for ecological ordination. Vegetatio. 1987;69:89–107.
Distance-based redundancy analysis: testing multispecies responses in multifactorial ecological experiments (vol 69, pg 1, 1999). Ecol Monogr. 1999;69:512–2.
Anderson MJ. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 2006;26:32–46.
Anderson MJ. Permutation tests for univariate or multivariate analysis of variance and regression. Can J Fish Aquat Sci. 2001;58:626–39.
Curtiss N, Balachandran A, Krska L, Peppiatt-Wildman C, Wildman S, Duckett J. A case controlled study examining the bladder microbiome in women with overactive bladder (OAB) and healthy controls. Eur J Obstet Gynecol Reprod Biol. 2017;214:31–5.
Thomas-White KJ, Kliethermes S, Rickey L, et al. Evaluation of the urinary microbiota of women with uncomplicated stress urinary incontinence. Am J Obstet Gynecol 2017;216:55 e1–55 e16.
Karstens L, Asquith M, Davin S, Stauffer P, Fair D, Gregory WT, et al. Does the urinary microbiome play a role in urgency urinary incontinence and its severity? Front Cell Infect Microbiol. 2016;6(78):1–13.
Kroke A, Schulz M, Hoffman K, Bermann MM, Boeing H. Assignment to menopausal status and estimation of age at menopause for women with missing or invalid data—a probabilistic approach with weighting factors in a large-scale epidemiologic study. Maturitas. 2001;40:39–46.
Harlow SD, Crawford SL, Sommer B, Greendale GA. Self-defined menopausal status in a multi-ethnic sample of midlife women. Maturitas. 2000;36:93–122.
Acknowledgements
The authors thank the research coordinators and all those who made this work possible, including University of Alabama at Birmingham: R. Edward Varner, MD, Robert L. Holley, MD, David R. Ellington, MD, Isuzu Meyer, MD, Alison Parden, MD, Alicia Ballard, MD, Velria Willis, BSN, Nancy Saxon, BSN, and Kathy Carter, BSN. Brown University: Deborah Myers, MD, Charles Rardin, MD, Brittany Star Hampton, MD, Cassandra Carberry, MD, Kyle Wohlrab, MD, Ann Meers, BSN, and Katheryn Rhodes, BA. Cleveland Clinic Foundation: Matthew Barber, MD, Marie Paraiso, MD, Eric Jelovsek, MD, Cecile Unger, MD, Audra Hill, MD, Ly Pung, RN, Kathleen Dastoli, RN, Annette Graham, RN, and Maryori Edington. Duke University: Cindy Amundsen, MD, Anthony Visco, MD, Alison Weidner, MD, Amie Kawasaki, MD, Shantae McLean, Nicole Longoria and Akira Hayes. University of California San Diego: Marianna Alperin, MD, Michael Albo, MD, Laura Aughinbaugh, RNP, Joann Columbo, Cindy Furey, PT, Sherella Johnson, Charles Nager, MD, and Patsy Riley, RN. Kaiser San Diego: Shawn Menefee, MD, Jasmine Tan-Kim, MD, Keisha Dyer, MD, Gouri Diwadkar, MD, Karl Luber, MD, Lynn Hall, RNP, Gisselle Zazueta-Damian and Linda MacKinnon. Kaiser Bellflower: John N. Nguyen, MD, Sharon Jakus-Walkman, MD, Azadeh Rezvan, MD, Christina Liao, MD, Arty Patel, PT, Mary Simmons, PT, Mercedes Cardona and Nancy Flores. University of New Mexico: Gena Dunivan, MD, Peter Jeppson, MD, Sara Cichowski, MD, Karen Taylor, BA, Cassandra Castaneda, BA, Julia Middendorf, BSN, Susan Tigert, BA/BS, Kurt Schwalm, BS, and Amy Overby, BS, CG/MB/PA(ASCP)CM. University of Pennsylvania: Heidi Harvie, MD, Uduak Andy, MD, Lorraine Flick and Michelle Kinglee. University of Pittsburgh: Pamela Moalli, PhD, MD, Michael Bonidie, Gary Sutkin, MD, Jonathan Shepherd, MD, Christopher Chermansky, MD, Judy Gruss, Karen Mislanovich and Lori Geraci. RTI International: Dennis Wallace, PhD, Carolyn Huitema, MS, Michael Ham, BS, and Joshua Levy, MS.
Funding source and sponsor’s role
The Eunice Kennedy Shriver National Institute of Child Health and Human Development sponsored this Pelvic Floor Disorders Network (PFDN) study (1-U10-HD069025–01, 2-U10-HD041261–11, 2-U10-HD041267–12, 1-U10-HD069013–01, 2-U10-HD054214–06, 2-U10-HD054215–06, 1-U10-HD069010–01, 1-U10-HD069006–01, 1-U01HD069031–01). ClinicalTrials.gov Number NCT01959347.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Presentation of information in this manuscript
Oral poster presented at the Pelvic Floor Disorders Week Conference (sponsored by the American Urogynecologic Society), October 5, 2017, Providence, Rhode Island; e-oral poster presented at the International Continence Society annual meeting, Florence, Italy, September 13, 2017.
Other support
National Center for Research Resources and the National Center for Advancing Translational Sciences of the National Institutes of Health (grant no. ULTR001449, the University of New Mexico Clinical and Translational Science Center) (for performance of DNA extraction, 16S sequencing).
Author’s potential conflicts of interest/disclosures
Yuko Komesu, MD: Co-PI Grant 1R01AT007171, National Center Complementary and Integrative Health, NIH and “Funding Source” above. Holly Richter, PhD, MD: UpToDate, Renovia. Benjamin Carper: None.Darrell L. Dinwiddie, PhD: None. Emily S. Lukacz, MD: AMS/Astora, Axonics, Boston Scientific, Uroplasty/Coloplast, UptoDate. Nazema Y. Siddiqui, MD: Medtronic. Vivian W. Sung, MD: None. Halina M. Zyczynski, MD: AUGS Board. Beri Ridgeway, MD: Coloplast, Ethicon. Rebecca G. Rogers, MD: UptoDate, ABOG Board member, Transform Trial, International Urogynecology Journal. Lily A. Arya, MD: None. Donna Mazloomdoost, MD: None. Marie G. Gantz, PhD: None.
Rights and permissions
About this article
Cite this article
Komesu, Y.M., Richter, H.E., Carper, B. et al. The urinary microbiome in women with mixed urinary incontinence compared to similarly aged controls. Int Urogynecol J 29, 1785–1795 (2018). https://doi.org/10.1007/s00192-018-3683-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00192-018-3683-6