Abstract
Key message
We found that the flowering time order of accessions in a genetic population considerably varied across environments, and homolog copies of essential flowering time genes played different roles in different locations.
Abstract
Flowering time plays a critical role in determining the life cycle length, yield, and quality of a crop. However, the allelic polymorphism of flowering time-related genes (FTRGs) in Brassica napus, an important oil crop, remains unclear. Here, we provide high-resolution graphics of FTRGs in B. napus on a pangenome-wide scale based on single nucleotide polymorphism (SNP) and structural variation (SV) analyses. A total of 1337 FTRGs in B. napus were identified by aligning their coding sequences with Arabidopsis orthologs. Overall, 46.07% of FTRGs were core genes and 53.93% were variable genes. Moreover, 1.94%, 0.74%, and 4.49% FTRGs had significant presence-frequency differences (PFDs) between the spring and semi-winter, spring and winter, and winter and semi-winter ecotypes, respectively. SNPs and SVs across 1626 accessions of 39 FTRGs underlying numerous published qualitative trait loci were analyzed. Additionally, to identify FTRGs specific to an eco-condition, genome-wide association studies (GWASs) based on SNP, presence/absence variation (PAV), and SV were performed after growing and observing the flowering time order (FTO) of plants in a collection of 292 accessions at three locations in two successive years. It was discovered that the FTO of plants in a genetic population changed a lot across various environments, and homolog copies of some key FTRGs played different roles in different locations. This study revealed the molecular basis of the genotype-by-environment (G × E) effect on flowering and recommended a pool of candidate genes specific to locations for breeding selection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rapeseed (Brassica napus L.) (AACC, n = 19), which originated from interspecific hybridization between Brassica rapa (AA, n = 10) and Brassica oleracea (CC, n = 9) approximately 7500 years ago, is a major source of edible oil and livestock feed rich in protein (Chalhoub et al. 2014; Nagaharu 1935). Allotetraploid rapeseed has been used in some major production regions, such as China and Canada, for less than a century. It has widely adapted to adverse climate zones across Eurasia, North America, and Australia by developing a broad variation in flowering time (FT) from early spring to late winter types. Therefore, variations in FT of rapeseed are crucial for its adaptation to new environments. Moreover, FT plays a critical role in determining the length of the life cycle period, yield, and quality of seeds and is one of the major targets for rapeseed breeding programs (Guo et al. 2014).
Flowering is regulated by a variety of endogenous and environmental cues such as changes in hormone levels (e.g., gibberellin, GA), age, carbohydrate status, ambient temperature, and photoperiod. Flowering is promoted by florigen, a mobile signal substance encoded by the floral integrator gene Flowering LOCUS T (FT). Florigens activate floral identity genes in the shoot apical meristem (SAM) and induce floral organ development (Freytes et al. 2021; Kinoshita and Richter 2020). In Arabidopsis, the CONSTANS (CO)-FT regulon genes are involved in various pathways that regulate flowering. Over the past decades, key regulators of the main flowering pathways have been identified, such as microRNA156 which targets SQUAMOSA PROMOTER BINDING-LIKE (SPL), a flowering promoter, in age-related pathways (Hyun et al. 2017); FLOWERING CONTROL LOCUS A (FCA) and LUMINIDEPENDENS (LD) in the autonomous pathways (Chakrabortee et al. 2016; Macknight et al. 1997); GIBBERELLIC ACID INSENSITIVE (GAI), REPRESSOR OF GA (RGA), and SPINDLY (SPY) in the gibberellic acid (GA) pathway (Jacobsen and Olszewski 1993; Peng and Harberd 1993; Tyler et al. 2004); CO, FT, DE-ETIOLATED 1 (DET1), and PHYTOCHROME A (PHYA) in the photoperiod pathway (Li et al. 2015; Putterill et al. 1995; Whitelam et al. 1993; Yoo et al. 2005); and FLOWERING LOCUS C (FLC), VERNALIZATION 1 (VRN1), and VRN2 in the vernalization pathway (Gendall et al. 2001; King et al. 2013; Sharma et al. 2020). A large number of these regulators are transcriptional factors (TFs), cofactors for TFs that have numerous downstream targeting genes, and chromatin remodelers. Additionally, hundreds of genes influence FT in Arabidopsis. Due to its much larger genome, B. napus, a relative of Arabidopsis in the family Brassicaceae, possesses 3–6 times more flowering time-related genes (FTRGs) in its allotetraploid genome than Arabidopsis.
Owing to advances in sequencing technology such as genomics, genome assembly and annotation have considerably improved (Bayer et al. 2021). The first reference genome of B. napus, the Darmor-bzh assembly, was published in 2014 (Chalhoub et al. 2014). The genomes of two winter ecotype accessions, Tapidor and Express 617 (Bayer et al. 2017; Lee et al. 2020), and those of two semi-winter ecotype accessions, ZS11 and Ningyou-7 (Chen et al. 2021; Zou et al. 2019), were de novo assembled and annotated in succession. However, none of these single reference genomes can represent the entire gene content of B. napus, since structural variations (SVs), such as gene presence and absence variations (PAVs) and copy number variations (CNVs), exist among these plants. To overcome this limitation, a pangenome for rapeseed was constructed and analyzed for SVs that were associated with agronomic traits (Song et al. 2020), disease resistance (Dolatabadian et al. 2020), and gene-loss propensity by interspecific hybridization (Bayer et al. 2021).
In this study, we aimed to improve the resolution of the FTRG panorama of B. napus on a pangenome-wide scale, detect single nucleotide polymorphisms (SNPs) associated with FTRGs specific to ecotypes, and identify SVs across a collection of germplasms belonging to different ecotypes. Finally, we performed multiple-year and multiple-location experiments to determine FTRGs associated with flowering time order variations (FTOV) specific to geographic locations and annual climate conditions by observing the changes in flowering time rank (FTR) of plants and performing genome-wide association studies (GWAS) based on SNPs, PAVs, and SVs in a genetic population consisting of 292 accessions. The results of this study provide insights into the FTRG network of B. napus and the molecular nature of a genotype-by-environment (G × E) effect and provide a reference to select and/or manipulate candidate FTRGs specific to geographic locations for introduction and domestication.
Materials and methods
Number of accessions investigated in each analysis
We investigated 1626, 991, and 292 accessions in PAV analysis, SV analysis, and multiple-environment GWAS, respectively (Table 1). The ID of these accessions and the proportion of ecotypes in each analysis are indicated or noted in the table.
Pangenome
The pangenome of B. napus constructed by Song et al. (2021) was used in this study. ZS11 was used as the reference genome, and 1688 resequenced genomes were combined to construct the pangenome (Lu et al. 2019; Parkin et al. 2005; Song et al. 2020; Wang et al. 2018; Wu et al. 2019). A total of 62 accessions were removed due to the deficiency of ecotype information. The ID of the accessions and the ecotypes to which they belong are provided in Table S23.
Identification of candidate FTRGs
Sequences of Arabidopsis FTGRs were downloaded from the flowering interaction database (http://www.phytosystems.ulg.ac.be/florid) (Bouché et al. 2016). The coding sequences (CDSs) of the Brassica napus pangenome were downloaded from the BnPIR database (http://cbi.hzau.edu.cn/bnapus/) (Song et al. 2021). The CDSs were compared with Arabidopsis FTRGs using blastn with an e-value of < 1e-20 and identity of > 80%.
SNP discovery
The SNP information for B. napus was downloaded from BnPIR, and the variations were annotated using VEP v99 (McLaren et al. 2016).
SV identification and selection analysis
Fastp (Chen et al. 2018) was used to filter out low-quality sequences from the raw data created by Wu et al. (2019) The ID of the 991 accessions and the ecotypes to which they belong are listed in Table S23. Clean data were mapped to the ZS11 reference genome using bowtie2 (Langmead and Salzberg 2012). BreakDancer and Delly were used to identify SVs (Chen et al. 2009).The results of BreakDancer were converted into VCF format. Then, SURVIVOR (version 1.0.6) (Jeffares et al. 2017) was used to identify SVs detected by both tools using the following parameters: “SURVIVOR merge Name 1000 2 1 1 0 50.” SURVIVOR defines and merges SVs based on breakpoints, SV types, and distances between SV chains. The SVs of all samples were merged using bcftools (Danecek et al. 2021). When merging, the depth of the reads at the breakpoints of SVs must be ≥ 4x, the distance between the start and end points of two adjacent SVs should not exceed 1000 bases, and the length of the overlap between the two SVs should > 50% of the total length.
To analyze SV under selection, the occurrence frequencies of each SV were calculated in the spring, winter, and semi-winter groups. Similar to the gene PAV analysis, the significance of the differences in the frequencies for each SV between groups was determined using Fisher’s exact test. p-values were then corrected using the Benjamini and Hochberg method. SVs under selection were identified with a false discovery rate (FDR) < 0.001 and a fold-change > 2. We set a 2-kb interval upstream of the genes as the promoter regions. Based on genome annotation and SV locations, we evaluated the effect of putative SV on the gene, CDS, and promoter regions. The SVs under selection were then analyzed to identify the associated genes.
Gene PAV selection analysis
The PAV of the pangenome was downloaded from the BnPIR database (http://cbi.hzau.edu.cn/bnapus/) (Song et al. 2021). Then, the frequencies of each gene in different B. napus ecotypes (spring, winter, and semi-winter) were calculated. The significance of the differences in gene frequencies between the groups was determined using Fisher's exact test. p-values were corrected using the Benjamini and Hochberg method to obtain the adjusted p-value. Genes with a fold-change in frequency > 2 between the two groups and a false discovery rate value < 0.001 were defined as genes under gene PAV selection.
The protein sequences of all genes in the B. napus pangenome were compared with those in the UniPort and Kyoto Encyclopedia of Genes and Genomes (KEGG) databases using BLASTP with a threshold of 1E-5. Using the Retrieve/ID mapping tool (https://www.uniprot.org/uploadlists/), UniPort IDs were converted to GO IDs to perform Gene Ontology (GO) annotation for all genes in the B. napus pangenome. Based on the GO and KEGG annotations, a hypergeometric test was used to perform functional enrichment analysis of the genes under gene PAV selection. The dhyper function in R was used to calculate p-values in the enrichment analysis. Additionally, the multiple-test corrected p-value (q-value) was calculated using the Benjamini–Hochberg function implemented in the R. q-value 0.05 was considered statically significant.
Linking known QTL and FTRGs
Chromosomal coordinates of QTL intervals for FT were obtained from previous studies (Jian et al. 2019; Liu et al. 2022; Scheben et al. 2020; Xu et al. 2021). The coordinates of the QTL intervals were converted to coordinates of the ZS11 genome.
Plots and graphs
Waterfall plots were drawn using GenVisR v1.11.3 (Skidmore et al. 2016). PAV information of FTRGs was used to generate a heatmap using R (version 4.0.3) with the ComplexHeatmap package (version 2.6.2, https://bioconductor.org/packages/release/bioc/html/ComplexHeatmap.html) (Gu et al. 2016). Scatter and line plots were drawn using the R package ggplot2 (Villanueva and Chen 2019). Sankey plots were drawn using the R package networkD3 (Allaire et al. 2017).
Construction of core germplasm
The 292 accessions for GWAS were selected based on the phylogenetic tree and principal component analyses of the genetic diversity of the 991 accessions. This GWAS population represented around 97.0% SNPs and 97.0% InDel polymorphisms of the large population consisting of 991 global accessions, which were sequenced on a genome-wide scale in our previous study (Wu et al. 2019).
Plant growth conditions
The 292 core accessions were grown at three locations, namely HZ (30.89°N and 119.63°E), JX (30.86°N and 120.70°E), and XY (34.80°N and 108.10°E), in two successive years during rapeseed growing seasons in 2019–2021. Sixteen plants from each accession were planted in a block (120 × 100 cm) with three replicates. Field management measures were carried out in accordance with the local rapeseed planting practices.
Observation of FT
The FT date for each accession was recorded when 50% of the plants had visible open flowers on their main inflorescence. The DSF was then calculated based on the number of days between sowing dates and FT dates.
GWAS-SNP, GWAS-PAV, and GWAS-SV
High-quality SNPs with minor allele frequency (MAF) > 0.05 discovered in the core accessions were used for GWAS-SNP. GWAS-SNP was performed using the BnaGVD website (http://rapeseed.biocloud.net/home) (Yan et al. 2021) with the default parameters of the EMMAX model. Associated genes were searched for in 75-kb sequence regions adjacent to SNPs. The SV and gene PAV information was used as the genotype instead of the SNP to perform GWAS-SV and GWAS-PAV using FarmCPU (default parameter) (Liu et al. 2016) in rMVP, with the threshold of significance set at 0.05/SV or 0.05/PAV number. SNPs were used for PCA using GCTA (Yang et al. 2011), and SNP-based principal components were used as covariates.
Results
PAV of B. napus FTRGs
In this study, we used the orthologous genes of Arabidopsis thaliana as a reference and identified 1337 FTRGs in the B. napus pangenome. Of these, 616 (46.07%) were core genes and 721 (53.93%) were variable genes; 482 (36.05%) genes (207 core and 275 variable genes) and 537 (40.16%) genes (310 core and 227 variable genes) were distributed in the A and C subgenomes, respectively. A total of 297 (22.21%) genes (all variables) were assigned to the contigs of the pangenome in addition to the two subgenomes. A group of 21 (1.57%) (two core and 19 variable) genes were identified in unplaced contigs (Table 2).
The distribution of the FTRGs in the reference genome is shown in Fig. 1. The largest number of FTRGs (99 genes) was identified on chromosome C03, whereas the smallest number of FTRGs (33 genes) was identified on chromosomes A04, A08, and A10. For each chromosome in the A genome, more variable FTRGs were identified than core FTRGs. In the C genome, except for C02 and C09, fewer variable FTRGs were identified than core FTRGs on most chromosomes. Chromosome C03 had the highest frequency (86.79%) of core FTRGs, whereas chromosome C02 had the lowest frequency (30.19%). Detailed information on variable and core FTRGs in the pangenome is provided in Table S1.
The distribution of dispensable flowering time-related genes (FTRGs) and all FTRGs across the reference genomes. The Y-axis represents gene densities, which were normalized to the genome-wide maximum of each measurement peaking at 1. The X-axis stands for the ID and size of each chromosome of the genome
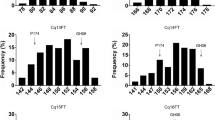
PAV-based pairwise comparisons were performed to analyze the presence-frequency difference (PFD) of FTRGs among the three ecotypes. The comparison of PFDs between the spring and winter ecotypes revealed five significantly more frequent and five significantly less frequent FTRGs in the spring ecotype than in the winter ecotype (Fig. 2a, d). Similarly, three FTRGs were significantly more frequent and 23 FTRGs were significantly less frequent in the spring ecotype than in the semi-winter ecotype (Fig. 2b, e). Moreover, six FTRGs were significantly more frequent in the winter ecotype than in the semi-winter ecotype and 54 FTRGs were significantly less frequent in the winter ecotype than in the semi-winter ecotype (Fig. 2c, f). The PFDs of the FTRGs in the ecotypes are provided in Table S2.
Comparison of the frequencies of flowering time-related genes (FTRGs) between a winter and spring, b semi-winter and spring, and c winter and semi-winter ecotypes. The red and green dots indicate significant differences (p < 0.01) between two comparing pairs, and they are shown as red and green lines in (d–f). The blue dots in (a–c) were not significantly different between the comparing pairs (color figure online)
Collectively, 26 (1.94%), 10 (0.74%), and 60 (4.49%) FTRGs had significant PFDs between the spring and semi-winter, spring and winter, and winter and semi-winter ecotypes, respectively. Excluding duplicates, 75 (5.61%) FTRGs had PFDs among the three ecotypes. The IDs of the PFD genes are listed in Table S3, and the overlapping numbers of the FTRGs between the ecotypes are indicated in a Venn diagram (Fig. S1). To intuitively overview the PAV status of each FTRG across the 1626 individuals (Table S4), we drew a fine-resolution (75 × 1626 pixels) heatmap (Fig. 3). As shown in Fig. 3, most FTRGs present in the semi-winter ecotype were absent in the spring and winter ecotypes, and seven FTRGs, namely BnaSORG0131600UN (PROTEIN ARGININE METHYLTRANSFERASE 4A, PRMT4A), BnaA02G0022900ZS (UBIQUITIN-SPECIFIC PROTEASE 12, UBP12), BnaA06G0299300ZS (CYCLIN-DEPENDENT KINASE C2, CDKC2), BnaA10G0150400ZS (SENSITIVITY TO RED LIGHT REDUCED 1, SRR1), BnaA03G0188600ZS (NUCLEAR FACTOR Y SUBUNIT B1, NF-YB1), BnaA03G0194600ZS (EARLY IN SHORT DAYS 6, ESD6), and BnaA07G0171100ZS (CYCLING DOF FACTOR 2, CDF2), were mostly different across the three ecotypes in terms of PAV.
Waterfall plot of presence/absence variation PAV of the flowering time-related genes (FTRGs) among three ecotypes. The IDs of the six FTRGs with the most significant differences between the ecotypes are indicated on the left site of the plot. The blue and yellow colors of each pixel represent the presence or absence of a particular gene in a genotype, respectively. Accession number = 1626, gene number = 75 (color figure online)
SV analyses of FTRGs
We previously resequenced a global collection of 991 accessions and investigated the SNPs for genomic polymorphisms (Table S23; Wu et al. 2019). Here, we identified 124,287 SVs, including insertions/deletions (indels), inversions, and duplications in the collection for the subsequent GWAS analyses (Table 3). Most SVs were of low frequency (< 0.01) (Fig. S2). Of these, 55,662 SVs were identified in the A genome, 68,565 SVs were identified in the C genome, and 60 SVs were identified in unplaced contigs in the ZS11 reference genome. On average, more SVs per chromosome were identified in the C genome (7618.3) than in the A genome (5566.2). Most SVs were indels (96.95% overall), varying from 95.53% to 98.31% on each chromosome (Table 3).
To identify the SVs that contributed to the differentiation of the three ecotypes, we compared the frequency of each SV among the spring, winter, and semi-winter ecotypes. A total of 1867 SVs with significantly different frequencies between ecotypes were identified. The position, SV type, length, allelic frequency, and pairwise ecotype comparisons are shown in detail in Table S5. The comparison results revealed five SVs that had significantly higher frequencies and 31 SVs that had significantly lower frequencies in the spring ecotype than in the semi-winter ecotype (Figs. S3 and S4). Moreover, 524 SVs with significantly higher frequencies and 263 SVs with significantly lower frequencies in the spring ecotype than in the winter ecotype were identified (Figs. S5 and S6). The results of the comparison of the SVs between the winter and semi-winter ecotypes revealed 187 SVs with significantly higher frequencies and 971 SVs with significantly lower frequencies in the winter ecotype than in the semi-winter ecotype (Figs. S7 and S8). Notably, 52 SVs had significantly different frequencies among the three ecotypes, which might have contributed to the differentiation of the ecotypes. The position, SV type, length, allelic frequency, pairwise ecotype comparisons, and IDs of the SV-associated FTRGs are summarized in Table S6. Collectively, we identified 3204 SVs on FTRGs in B. napus. Of these, 61.77% were indels, 38.14% were inversions, and only 0.06% were duplications.
Analysis of known quantitative trait loci (QTL) that regulates FT
Numerous QTL that control FT were mapped in B. napus (Jian et al. 2019; Liu et al. 2022; Scheben et al. 2020; Xu et al. 2021). These QTL contributed considerably to flowering time variations. To investigate the FTRGs underlying the known QTL, we remapped the QTL to the pangenome. SNPs, PAVs, and SVs of the FTRGs underlying 14 QTL were analyzed. Seventy-eight FTRGs underlying those QTL were identified. Of these, 39 FTRGs varied significantly across ecotypes. A waterfall plot was drawn to display the variant types, such as gene-loss, stop-codon-loss, missense, synonymous, intron, splice-region, 3′-UTR variants, and 5′-UTR variants of the FTRGs underlying the QTL (Fig. 4). Among these FTRGs, BnaA06G0332400ZS (FLOWERING-PROMOTING FACTOR 1, FPF1) had the lowest mutation rate, and the mutations were mainly synonymous. BnaA06G0277300ZS (CYCLIC DOF FACTOR 1-LIKE, CDF1) had the highest mutation rate and the mutations were gene-loss and missense variants, indicating considerably high genetic polymorphism and functional diversity. Although BnaA06G0126600ZS (ACTIN-RELATED PROTEIN 4, ARP4) had a relatively low mutation rate, the mutations were mostly gene-loss variants, indicating PAVs across the 1626 individuals. BnaA06G0256500ZS (DICER-LIKE 3, DCL3) was featured with mutations in the stop-codon-loss variants. BnaA06G0270200ZS (UBIQUITIN-SPECIFIC PROTEASE 26, UBP26) had a considerable proportion of synonymous mutations that did not lead to any functional differences. On average, individuals of each ecotype had a similar mutation frequency. However, it was obvious that the differences in mutation frequency between the individuals in the semi-winter type were bigger than those in the winter and spring types. The allelic variation of the 39 FTRGs across 1626 accessions (including the 991 accessions) is provided in detail in Table S7.
Waterfall plot of known qualitative trait locus (QTL) that control flowering time in a recombinant-inbred-line population. The IDs of flowering time-related genes (FTRG) according to the ZS11 reference genome are listed on the left side with histograms indicating the rate of mutation of each gene of the 1626 accessions. The graph on the top represents the mutation frequency of the accessions across the 39 FTRGs. The different colors of the 39 × 1626 pixels represent the mutation types as indicated on the right side of the plot (color figure online)
FTOV observation in a genetic population
Based on the genome-wide distribution of FTRGs in B. napus, we attempted to identify the FTRGs associated with flowering time order (FTO) in a given environment. We grew the genetic population consisting of 292 core accessions (Table S23), which were selected based on the resequencing data of 991 global germplasm collections at three locations: Xian-Yang (XY, 34.80°N and 108.10°E), Hu-Zhou (HZ, 30.89°N and 119.63°E), and Jia-Xing (JX, 30.86°N and 120.70°E) in two successive years. The 292 core accessions contains most polymorphisms of 991 populations, and its validity in GWAS analysis was confirmed in previous study. We recorded the days from sowing to flowering (DSF) for each individual in the six environments (location × year). DSF varied from 135 to 194 at HZ, 102 to 202 at JX, and 158 to 213 at XY (Table S8). The number of flowering accessions displayed a bimodal distribution, peaking twice in all six environments, which was particularly clear at XY (Fig. S9). We divided the DSF into five grades, G1 to G5, corresponding to increasing DSF. We determined the FTR of each accession and arranged the FTO of the population in each environment (Table S8). We drew a Sankey diagram to intuitively show the FTOV across the six environments (Fig. 5). The FTRs of 72 (24.66%) accessions were relatively consistent in various environments. However, the FTRs of 193 (66.10%) accessions fluctuated with changes in FTRs between neighboring grades across the environments. Notably, the FTRs of 27 (9.25%) accessions changed drastically between non-adjacent grades across environments (Fig. 5; Table S8). Therefore, the FTO of a population changes with time and location, and the FTR for a specific accession in two different environments may be considerably different.
Comparison of flowering time rank in the population comprising 292 accessions across different locations and years. Time is divided into five periods from early to late as shown with the erect columns. The darker the line that links an accession, the more drastic the change in the rank of flowering time of the accession. The lavender color represents the collinearity between years at the same location, and the yellow color represents the collinearity across locations. HZ, JX, and XY indicate the location of field experiments (Hu-Zhou, 30.89°N and 119.63°E; Jia-Xing 30.86°N and 120.70°E; and Xian-Yang 34.80°N and 108.10°E, respectively). Y1 and Y2 stand for the repetition of field experiments in different years (color figure online)
GWAS on FT specific to a given environment
To identify the FTRGs associated with FT in a specific environment, we performed GWAS-SNP, GWAS-PAV, and GWAS-SV (excluding PAV). A set of Manhattan plots showed the results of the GWAS-SNP in the six environments (Fig. S10). Genes associated with FT specific to the environment were analyzed (Tables S9–S14). Cross-analyses between locations were performed, and the number of overlapped associated genes is shown in a Venn diagram (Fig. 7a). Eighteen genes were significantly associated with FT at the three locations. We identified 22 and 134 significantly associated genes that overlapped between JX and XY, HZ, and JX, respectively. The numbers of genes that were identified only at HZ, JX, and XY were 758, 1853, and 176, respectively (Table S15). Several associated genes belonged to the 1337 FTRGs that were predicted above (Table S16). Interestingly, homologous gene copies play different roles in various environments. For example, both BnaA03G0144400ZS and BnaC02G0039100ZS are FLC homologous genes; however, they were identified as associated genes at different locations, that is, BnaA03G0144400ZS was identified in HZ, whereas BnaC02G0039100ZS was identified at JX. Moreover, both BnaC03G0129200ZS and BnaA02G0118200ZS were VERNALISATION INSENSITIVE 3 (VIN3) homologs; however, BnaC03G0129200ZS was identified as an associated gene at JX, whereas BnaA02G0118200ZS was identified at XY (Table S16).
GWAS-PAV and GWAS-SV (excluding PAV) were also performed to supplement the results of GWAS-SNP (Fig. 6). PAV-associated genes in all six environments are provided in Table S17. GWAS-PAV and GWAS-SV (excluding PAV) did not reveal associated genes common to the three locations. GWAS-PAV revealed 11, 17, and 7 genes associated with HZ, JX, and XY, respectively (Figs. 6, 7b). GWAS-SV (excluding PAV) revealed 140, 24, and 3 genes associated with HZ, JX, and XY, respectively (Figs. 6, 7c; Table S18). GWAS-PAV revealed one overlapping gene, BnaA10G0193800ZS (TatD-related DNase), between HZ and JX, and one overlapping gene, BnaA02G0374400ZS (Topless-related Protein), between JX and XY (Fig. 7b). In contrast, GWAS-SV (excluding PAV) revealed other overlapping genes BnaC08G0316200ZS (Spermatogenesis-associated Protein 7, associated with SV114034) between HZ and JX (Fig. 7c). Taken together, these results indicated that GWAS-SNP, GWAS-PAV, and GWAS-SV (excluding PAV) enriched the number of FTRG candidates and provided the molecular nature underlying the genotype-by-environment (G × E) effect on FT.
Venn diagrams comparing flowering time-related genes determined by a GWAS-SNP, b GWAS-PAV, and c GWAS-SV between different locations. The blue, red, and yellow circles represent XY, HZ, and JX, respectively. The overlapped genes were indicated with numbers, and their IDs are provided in Supplemental Tables S15, S17, and S18. HZ, Hu-Zhou, 30.89°N and 119.63°E; JX, Jia-Xing 30.86°N and 120.70°E; XY, Xian-Yang 34.80°N and 108.10°E (color figure online)
Discussion
The transition from vegetative growth to flowering is an important developmental stage of a plant during its life cycle. FT determines not only the life cycle length but also the yield and quality of a crop. A correct time of flowering is essential in multiple-cropping systems, where one or even two additional cropping seasons for rice production closely follow the harvest of winter crops. The late flowering of the preceding crop delays the planting time of the next crop. Moreover, the cultivation of plants with appropriate FT requires knowledge of allelic variations in FTRGs. In this study, we investigated B. napus FTRGs on a pangenome-wide scale and identified SNPs, PAVs, and SVs in a collection of worldwide rapeseed germplasm. Here, we defined SVs as indels, inversions, and duplications, but not PAV of genes (Lu et al. 2019; Song et al. 2020, 2021; Wang et al. 2018; Wu et al. 2019).
The concept of the pangenome was first developed in bacteria, describing an organism’s complete genetic material, including core and variable genomes (Tettelin et al. 2005). Various strategies have been developed to improve the pangenome. First, methods were applied to align reads from multiple individuals to a reference genome of good quality, assemble unaligned reads into novel contigs, and add novel contigs to the original reference sequence to construct a pangenome. Second, different strategies were applied to de novo assemble genomes of multiple accessions and align the whole genomes to identify variable genomic regions. More recently, a new strategy was proposed to construct a pangenome graph by aligning whole genomes and storing information on variable regions through the graph (Bayer et al. 2020). In the present study, we used the pangenome constructed by Song et al. (2021), which was developed by mapping 1688 resequenced accessions to the ZS11 reference genome (Lu et al. 2019; Wang et al. 2018; Wu et al. 2019). Dolatabadian et al. (2020) adopted a similar strategy to build a B. napus pangenome by mapping 50 resequenced accessions to the Darmor-bzh reference genome (v8.1). They identified 1749 resistant gene analogs, of which 996 were core, 753 were variable, and 368 were not present in the reference. The number of resequenced accessions involved in our study was considerably larger than that in the study by Dolatabdian et al. (2020), and consequently, the pangenome was also larger (1789.9 vs. 1040 Mb). We identified 1337 FTRGs in the pangenome, including 616 core and 721 variable genes, indicating high diversity (Fig. 1, Table 2).
In the last two decades, knowledge of FT regulation in model plants has rapidly increased, and analyses of the complicated genetic networks underlying each pathway have revealed multiple connections between the components of these pathways (Berry and Dean 2015). Genetic analysis of FT regulation was greatly facilitated by referring to Arabidopsis thaliana, a close relative of the family Brassicaceae. In our study, the B. napus FTRGs used for the PAV and SV analyses were identified by aligning Arabidopsis FTRGs to the coding sequences of the B. napus pangenome. Arabidopsis FTRGs were downloaded from the Flowering-Interactive Database (FLOR-ID), a hand-curated database containing information about 306 FTRGs and linked to 1595 publications (Bouché et al. 2016). Due to its polyploid nature, the network of B. napus FTRGs comprising approximately 1337 genes was considerably more complicated than that of Arabidopsis. The A subgenome of B. napus had more variable FTRGs (275 FTRGs) than the C subgenome (227 FTRGs), and the frequencies of PAVs and SVs were higher in the A genome than in the C genome (Tables 2 and 3), indicating that the A genome had higher diversity than the C genome in the FT control. This result was consistent with that of our previous study that compared the overall genetic diversity between the two subgenomes by analyzing SNPs (Wu et al. 2019). The A genome had a higher degree of genetic diversity, possibly due to occasional outcrossing between rapeseed (AACC) and its diploid ancestor species B. rapa (AA). In contrast, outcrossing between rapeseed and B. oleracea (CC) is very rare. Therefore, it is likely that the genetic diversity of the C genome was limited to a few donors of the original hybridization events that had created the species. In general, rapeseed breeding, especially that of the canola type, is subject to a narrow genetic diversity, and the PAVs and SVs of FTRGs could serve as efficient markers for breeding the ideal FT. Schiessl et al. (2017a; 2017b) identified 184 FTRGs orthologous to 35 Arabidopsis FTRGs. The 1337 FTRGs included 145 (78.8%) genes that they identified. The discrepancy might arise from the different thresholds applied to determine orthologous genes, and different reference genomes applied. For a part of the FTRGs such as FLC, FT, TEMPRANILLO 1 (TEM1), and EARLY FLOWERING 7 (ELF7), we found the same number of copies as they did. However, the positions of the copies might be different. For example, they localized one of the two ELF7 copies on A10 that was the same as we did, but the other ELF7 copy, which we recognized as BnaC09G0616900ZS, on unknown chromosomes. For some other Arabidopsis FTRGs such as AGAMOUS-LIKE 24 (AGL24), APETALA 1 (AP1), CIRCADIAN CLOCK ASSISTED 1 (CCA1), CDF1, EARLY FLOWERING IN SHORT DAYS (EFS), FLOWERING LOCUS D (FD), FRUITFUL (FUL), GLYCIN-RICH PROTEIN 7 (GRP7), PHYA, SRR1, VIN3, we identified more copies than they did. For a few genes such as CO, they found more copies.
Based on PAV and SV analyses of FTRGs in B. napus, it seems that the spring ecotype and semi-winter ecotype had higher similarity between each other compared with the winter ecotype (Figs. 2, 3). According to Fussell (1955) and our previous study (Wu et al. 2019), Europe was the center of ancient rapeseed cultivars. B. napus could have been cultivated in the Mediterranean areas as early as Classical Times (between the eighth century BC and the sixth century AD). It was spread to other areas by soldiers, merchants and/or birds, and animals, adapting to diverse climate zones and latitudes and forming ecotypes different from the winter type. Winter rapeseed distinguishes from semi-winter and spring types with a longer life cycle and more stringent requirements for vernalization. It is the ancestor type, whereas the semi-winter and spring types were formed in the process of adaptation to eco-environments where seedlings grow in warmer weather conditions.
QTL is a section of DNA that correlates with trait variations such as FT. Mapping of QTL is often an early step in finding genes that are responsible for trait variations. Even with a fine-mapping approach, a QTL interval may cover several candidate genes. In this study, we remapped some known QTL that control FT to the pangenome and analyzed the genetic polymorphisms of the FTRGs underlying 14 known QTL. Mutational variants such as gene loss, stop codon, missense, synonymous, intron, splice regions, and 3′-UTR of the FTRGs underlying the QTL were intuitively displayed (Fig. 4). Mutational variants, such as synonymous variants, can be excluded and PAVs can be prioritized as candidates in the analysis. Moreover, the graph can help in understanding the diversity of a specific FTRG and narrowing the candidate genes that contribute to a QTL effect. Overall, the winter ecotype did not have a higher mutation frequency than the semi-winter and spring ecotypes. However, the deviation of mutation frequency between the individuals within the semi-winter type was obviously bigger than that within the winter and spring types (Fig. 4). This might arise from the choice of the reference genome. In our research, we used ZS11 as the reference, which is a typical native Chinese cultivar. The semi-winter group contain both Chinese cultivars/breeding lines that are genetically close to ZS11, and foreign cultivars/breeding lines from, e.g., Pakistan and/or Australia that may have large genetic distances from ZS11.
Furthermore, to gain more knowledge about the FTRGs and expand the number of FTRG candidates, we performed GWAS-SNP, GWAS-PAV, and GWAS-SV to identify FTRGs after growing a genetic population in three locations in two successive years. XY is located in the northwest region of China (34.80°N and 108.10°E) with a relatively higher latitude where winter is rather cold and dry, and the daily photoperiod in spring is relatively long. HZ (30.89°N and 119.63°E) and JX (30.86°N and 120.70°E) are located at a ~ 100-km distance within the same province Southeast of China, where winter was milder and rainfall was more plentiful than at XY (Fig. S11, Table S19). We observed that the FTO varied across different locations and years. In particular, 24.66% of accessions had a relatively consistent FTR, and 66.10% of accessions had moderate FTR changes. In contrast, 9.25% of accessions had rather drastic FTOV across non-adjacent DSF grades (Fig. 5). To understand whether the FTR change across the different environments depended on ecotypes, we investigated the relationship between the FTR consistency and the ecotypes. As shown in Table 4 and Figure S12, the proportions of each ecotype in the consistent categories and moderately fluctuated categories (Fig. S12d and e) were not much different from those in the whole set of GWAS accessions (the 292 accessions) (Fig. S12c). However, the proportions of the winter ecotype and semi-winter type were over and insufficiently represented, respectively, in the drastically fluctuated category (Fig. S12f), considering their proportions in the whole set. A probable reason is that semi-winter accessions were more adaptable to all three geographic locations, as most of them were native cultivars/lines. But, most winter accessions were from Europe, where the eco-conditions are very different from the three experimental locations. They were, therefore, more sensitive to flowering conditions. Overall, the consistency of FTR depends more on individual genotypes. We assume that each flowering pathway might not be equally important for the individuals, and the coordination of homologous genes between the subgenomes of the polyploidy might be unique for a specific genotype. However, we need further experiments to verify this assumption.
GWAS-SNP revealed that JX and HZ shared a total of 152 genes that were associated with FT. In contrast, XY and HZ, XY, and JX only shared 40 and 18 genes that were associated with FT, respectively (Fig. 7a; Table S15), suggesting that different FTRG networks would control FT in varying locations. The greater the distance between two locations, the more different the genes controlling FT. GWAS-PAV and GWAS-SV revealed a significantly lower number of FT-associated genes but did not reveal any FTRGs that were shared by all three locations (Fig. 7b, c). Moreover, relatively fewer associated genes were identified at XY than at HZ and JX (Fig. 7b, c). We enriched the number of FTRGs and provided ID and annotations of the genes (Tables S15–S18). The GWAS-SNP confirmed the importance of FLC in determining FT, which was also reported in our previous study (Wu et al. 2019) and another study (Song et al. 2020). Interestingly, homologous copies of a gene appeared to play different roles in various environments. For example, both BnaA03G0144400ZS and BnaC02G0039100ZS are FLC homologous genes and were identified as loci associated with FT; however, BnaFLC-A03 (BnaA03G0144400ZS) was identified at HZ, whereas BnaFLC-C02 (BnaC02G0039100ZS) was found at JX. Similarly, both BnaC03G0129200ZS and BnaA02G0118200ZS were VIN3 homologs; however, BnaVIN3-C03 (BnaC03G0129200ZS) was identified at JX, whereas BnaVIN3-A02 (BnaA02G0118200ZS) was found at XY (Tables S15 and S16). This may be ascribed to epigenetic activation or repression of a locus (such as temporary methylation or demethylation) in a given environment. However, the role of epigenetic modifications in such phenomena requires further investigation. The GWAS revealed a PHYTOCHROME-INTERACTING FACTOR 6 (PIF6) ortholog (BnaA09G0562200ZS) at XY, but not at HZ and JX (Tables S15 and S16), implying that the role of a certain flowering pathway (e.g., the photoperiod pathway) might not be equally important for different locations. The daily sunny hours could be an important factor limiting the transition from the vegetative stage to flowering at XY (Noda and Ozawa 2018). On the other hand, genes in the vernalization pathway could be dominantly important for flowering at HZ and JX, as the winter of the two locations might not have been cold enough to meet the vernalization requirements (Fig. S11).
In conclusion, we provide the highest resolution graphics of FTRGs in B. napus so far on a pangenome-wide scale based on SNP and SV analyses. We gain novel insights into the genetic basis of flowering time QTL and the G × E effect on flowering. We discover that the FTO of plants in a genetic population changes a lot across various environments, and homolog copies of some key FTRGs played different roles in different locations. The results of our study can be of reference in selecting and/or manipulating candidate FTRGs for the introduction and domestication of varieties to a new environment.
Data availability
The supporting data of Figures and Tables are available in Supplemental Tables S1–S23. The raw reads of the rapeseed accessions have been deposited in the public database of National Center of Biotechnology Information under SRP155312 (https://www.ncbi.nlm.nih.gov /sra/SRP155312) and China National Center for Bioinformation (NGDC) (https://ngdc.cncb.ac.cn/gsa/browse/CRA001854).
References
Allaire JJ, Gandrud C, Russell K, Yetman C (2017) networkD3: D3 JavaScript Network Graphs from R., R package version 0.4 edn
Bayer PE, Hurgobin B, Golicz AA, Chan CKK, Yuan Y, Lee H, Renton M, Meng J, Li R, Long Y, Zou J, Bancroft I, Chalhoub B, King GJ, Batley J, Edwards D (2017) Assembly and comparison of two closely related Brassica napus genomes. PLANT BIOTECHNOL J 15:1602–1610
Bayer PE, Golicz AA, Scheben A, Batley J, Edwards D (2020) Plant pan-genomes are the new reference. NAT PLANTS 6:914–920
Bayer PE, Scheben A, Golicz AA, Yuan Y, Faure S, Lee H, Chawla HS, Anderson R, Bancroft I, Raman H, Lim YP, Robbens S, Jiang L, Liu S, Barker MS, Schranz ME, Wang X, King GJ, Pires JC, Chalhoub B, Snowdon RJ, Batley J, Edwards D (2021) Modelling of gene loss propensity in the pangenomes of three Brassica species suggests different mechanisms between polyploids and diploids. Plant Biotechnol J 19:2488–2500
Berry S, Dean C (2015) Environmental perception and epigenetic memory: mechanistic insight through FLC. Plant J 83:133–148
Bouché F, Lobet G, Tocquin P, Périlleux C (2016) FLOR-ID: an interactive database of flowering-time gene networks in Arabidopsis thaliana. Nucleic Acids Res 44:D1167–D1171
Chakrabortee S, Kayatekin C, Newby GA, Mendillo ML, Lancaster A, Lindquist S (2016) Luminidependens (LD) is an Arabidopsis protein with prion behavior. Proc Natl Acad Sci USA 113:6065–6070
Chalhoub B, Denoeud F, Liu S, Parkin IAP, Tang H, Wang X, Chiquet J, Belcram H, Tong C, Samans B, Corréa M, Silva CD, Just J, Falentin C, Koh CS, Le Clainche I, Bernard M, Bento P, Noel B, Labadie K, Alberti A, Charles M, Arnaud D, Guo H, Daviaud C, Alamery S, Jabbari K, Zhao M, Edger PP, Chelaifa H, Tack D, Lassalle G, Mestiri I, Schnel N, Le Paslier M, Fan G, Renault V, Bayer PE, Golicz AA, Manoli S, Lee T, Thi VHD, Chalabi S, Hu Q, Fan C, Tollenaere R, Lu Y, Battail C, Shen J, Sidebottom CHD, Wang X, Canaguier A, Chauveau A, Bérard A, Deniot G, Guan M, Liu Z, Sun F, Lim YP, Lyons E, Town CD, Bancroft I, Wang X, Meng J, Ma J, Pires JC, King GJ, Brunel D, Delourme R, Renard M, Aury J, Adams KL, Batley J, Snowdon RJ, Tost J, Edwards D, Zhou Y, Hua W, Sharpe AG, Paterson AH, Guan C, Wincker P (2014) Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345:950–953
Chen K, Wallis JW, McLellan MD, Larson DE, Kalicki JM, Pohl CS, McGrath SD, Wendl MC, Zhang Q, Locke DP, Shi X, Fulton RS, Ley TJ, Wilson RK, Ding L, Mardis ER (2009) BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nat Methods 6:677–681
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890
Chen X, Tong C, Zhang X, Song A, Hu M, Dong W, Chen F, Wang Y, Tu J, Liu S, Tang H, Zhang L (2021) A high-quality Brassica napus genome reveals expansion of transposable elements, subgenome evolution and disease resistance. Plant Biotechnol J 19:615–630
Danecek P, Bonfield JK, Liddle J, Marshall J, Ohan V, Pollard MO, Whitwham A, Keane T, McCarthy SA, Davies RM, Li H (2021) Twelve years of SAMtools and BCFtools. Gigascience 10:1–4
Dolatabadian A, Bayer PE, Tirnaz S, Hurgobin B, Edwards D, Batley J (2020) Characterization of disease resistance genes in the Brassica napus pangenome reveals significant structural variation. Plant Biotechnol J 18:969–982
Freytes SN, Canelo M, Cerdán PD (2021) Regulation of flowering time: When and where? Curr Opin Plant Biol 6:102049
Fussell GE (1955) History of Cole (Brassica sp.). Nature 176:48–51
Gendall AR, Levy YY, Wilson A, Dean C (2001) The VERNALIZATION 2 Gene Mediates the Epigenetic Regulation of Vernalization in Arabidopsis. Cell 107:525–535
Gu Z, Eils R, Schlesner M (2016) Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32:2847–2849
Guo Y, Hans H, Christian J, Molina C (2014) Mutations in single FT- and TFL1-paralogs of rapeseed (Brassica napus L.) and their impact on flowering time and yield components. Front Plant Sci 5:282
Hyun Y, Richter R, Coupland G (2017) Competence to Flower: age-controlled sensitivity to environmental cues. Plant Physiol 173:36–46
Jacobsen SE, Olszewski NE (1993) Mutations at the SPINDLY locus of Arabidopsis alter gibberellin signal transduction. Plant Cell 5:887–896
Jeffares DC, Jolly C, Hoti M, Speed D, Shaw L, Rallis C, Balloux F, Dessimoz C, Bähler J, Sedlazeck FJ (2017) Transient structural variations have strong effects on quantitative traits and reproductive isolation in fission yeast. Nat Commun 8:14061
Jian H, Zhang A, Ma J, Wang T, Yang B, Shuang LS, Liu M, Li J, Xu X, Paterson AH, Liu L (2019) Joint QTL mapping and transcriptome sequencing analysis reveal candidate flowering time genes in Brassica napus L. BMC Genom 20:21
King GJ, Chanson AH, McCallum EJ, Ohme-Takagi M, Byriel K, Hill JM, Martin JL, Mylne JS (2013) The Arabidopsis B3 Domain Protein VERNALIZATION1 (VRN1) Is Involved in Processes Essential for Development, with Structural and Mutational Studies Revealing Its DNA-binding Surface. J Biol Chem 288:3198–3207
Kinoshita A, Richter R (2020) Genetic and molecular basis of floral induction in Arabidopsis thaliana. J Exp Bot 71:2490–2504
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359
Lee H, Chawla HS, Obermeier C, Dreyer F, Abbadi A, Snowdon R (2020) Chromosome-scale assembly of winter oilseed rape Brassica napus. Front Plant Sci 11:496
Li K, Gao Z, He H, Terzaghi W, Fan L, Deng XW, Chen H (2015) Arabidopsis DET1 represses photomorphogenesis in part by negatively regulating DELLA protein abundance in darkness. Mol Plant 8:622–630
Liu X, Huang M, Fan B, Buckler ES, Zhang Z (2016) Iterative usage of fixed and random effect models for powerful and efficient genome-wide association studies. Plos Genet 12:e1005767
Liu Z, Dong X, Zheng G, Xu C, Wei J, Cui J, Cao X, Li H, Fang X, Wang Y, Tian H (2022) Integrate QTL mapping and transcription profiles reveal candidate genes regulating flowering time in Brassica napus. Front Plant Sci 13:904198
Lu K, Wei L, Li X, Wang Y, Wu J, Liu M, Zhang C, Chen Z, Xiao Z, Jian H, Cheng F, Zhang K, Du H, Cheng X, Qu C, Qian W, Liu L, Wang R, Zou Q, Ying J, Xu X, Mei J, Liang Y, Chai Y, Tang Z, Wan H, Ni Y, He Y, Lin N, Fan Y, Sun W, Li N, Zhou G, Zheng H, Wang X, Paterson AH, Li J (2019) Whole-genome resequencing reveals Brassica napus origin and genetic loci involved in its improvement. Nat Commun 10
Macknight R, Bancroft I, Page T, Lister C, Schmidt R, Love K, Westphal L, Murphy G, Sherson S, Cobbett C, Dean C (1997) FCA, a gene controlling flowering time in arabidopsis, encodes a protein containing RNA-binding domains. Cell 89:737–745
McLaren W, Gil L, Hunt SE, Riat HS, Ritchie GRS, Thormann A, Flicek P, Cunningham F (2016) The ensembl variant effect predictor. Genome Biol 17:122
Nagaharu U (1935) Genomic analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jpn J Bot 7:389–452
Noda N, Ozawa T (2018) Light-controllable transcription system by nucleocytoplasmic shuttling of a truncated phytochrome B. Photochem Photobiol 94:1071–1076
Parkin IAP, Gulden SM, Sharpe AG, Lukens L, Trick M, Osborn TC, Lydiate DJ (2005) Segmental structure of the Brassica napus genome based on comparative analysis with Arabidopsis thaliana. Genetics 171:765–781
Peng J, Harberd NP (1993) Derivative alleles of the arabidopsis gibberellin-insensitive (gai) mutation confer a wild-type phenotype. Plant Cell 5:351–360
Putterill J, Robson F, Lee K, Simon R, Coupland G (1995) The CONSTANS gene of Arabidopsis promotes flowering and encodes a protein showing similarities to zinc finger transcription factors. Cell 80:847–857
Scheben A, Severn-Ellis AA, Patel D, Pradhan A, Rae SJ, Batley J, Edwards D (2020) Linkage mapping and QTL analysis of flowering time using ddRAD sequencing with genotype error correction in Brassica napus. BMC Plant Biol 20:546
Schiessl S, Huettel B, Kuehn D, Reinhardt R, Snowdon R (2017a) Post-polyploidisation morphotype diversification associates with gene copy number variation. Sci Rep-UK 7:41845
Schiessl S, Huettel B, Kuehn D, Reinhardt R, Snowdon RJ (2017b) Targeted deep sequencing of flowering regulators in Brassica napus reveals extensive copy number variation. Sci Data 4:170013
Sharma N, Geuten K, Giri BS, Varma A (2020) The molecular mechanism of vernalization in Arabidopsis and cereals: role of Flowering Locus C and its homologs. Physiol Plantarum 170:373–383
Skidmore ZL, Wagner AH, Lesurf R, Campbell KM, Kunisaki J, Griffith OL, Griffith M (2016) GenVisR: genomic visualizations in R. Bioinformatics 32:3012–3014
Song J, Guan Z, Hu J, Guo C, Yang Z, Wang S, Liu D, Wang B, Lu S, Zhou R, Xie W, Cheng Y, Zhang Y, Liu K, Yang Q, Chen L, Guo L (2020) Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat Plants 6:34–45
Song JM, Liu DX, Xie WZ, Yang Z, Guo L, Liu K, Yang QY, Chen LL (2021) BnPIR: Brassica napus pan-genome information resource for 1689 accessions. Plant Biotechnol J 19:412–414
Tettelin H, Masignani V, Cieslewicz MJ, Donati C, Medini D, Ward NL, Angiuoli SV, Crabtree J, Jones AL, Durkin AS, DeBoy RT, Davidsen TM, Mora M, Scarselli M, Ros IMY, Peterson JD, Hauser CR, Sundaram JP, Nelson WC, Madupu R, Brinkac LM, Dodson RJ, Rosovitz MJ, Sullivan SA, Daugherty SC, Haft DH, Selengut J, Gwinn ML, Zhou L, Zafar N, Khouri H, Radune D, Dimitrov G, Watkins K, Connor KJBO, Smith S, Utterback TR, White O, Rubens CE, Grandi G, Madoff LC, Kasper DL, Telford JL, Wessels MR, Rappuoli R, Fraser CM (2005) Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial “pan-genome.” Proc Natl Acad Sci USA 102:13950–13955
Tyler L, Thomas SG, Hu J, Dill A, Alonso JM, Ecker JR, Sun T (2004) DELLA proteins and gibberellin-regulated seed germination and floral development in arabidopsis. Plant Physiol 135:1008–1019
Villanueva RAM, Chen ZJ (2019) ggplot2: elegant graphics for data analysis (2nd ed.). Measurement: interdisciplinary research and perspectives 17:160–167
Wang B, Wu Z, Li Z, Zhang Q, Hu J, Xiao Y, Cai D, Wu J, King GJ, Li H, Liu K (2018) Dissection of the genetic architecture of three seed-quality traits and consequences for breeding in Brassica napus. Plant Biotechnol J 16:1336–1348
Whitelam GC, Johnson E, Peng J, Carol P, Anderson ML, CowI JS, Harberd NP (1993) Phytochrome A null mutants of Arabidopsis display a wild-type phenotype in white light. Plant Cell 5:757–768
Wu D, Liang Z, Yan T, Xu Y, Xuan L, Tang J, Zhou G, Lohwasser U, Hua S, Wang H, Chen X, Wang Q, Zhu L, Maodzeka A, Hussain N, Li Z, Li X, Shamsi IH, Jilani G, Wu L, Zheng H, Zhang G, Chalhoub B, Shen L, Yu H, Jiang L (2019) Whole-genome resequencing of a worldwide collection of rapeseed accessions reveals the genetic basis of ecotype divergence. Mol Plant 12:30–43
Xu Y, Zhang B, Ma N, Liu X, Qin M, Zhang Y, Wang K, Guo N, Zuo K, Liu X, Zhang M, Huang Z, Xu A (2021) Quantitative trait locus mapping and identification of candidate genes controlling flowering time in Brassica napus L. Front Plant Sci 11:626205
Yan T, Yao Y, Wu D, Jiang L (2021) BnaGVD: a genomic variation database of rapeseed (Brassica napus). Plant Cell Physiol 62:378–383
Yang J, Lee SH, Goddard ME, Visscher PM (2011) GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet 88:76–82
Yoo SK, Chung KS, Kim J, Lee JH, Hong SM, Yoo SJ, Yoo SY, Lee JS, Ahn JH (2005) CONSTANS activates suppressor of overexpression of constans 1 through flowering locus T to promote flowering in arabidopsis. Plant Physiol 139:770–778
Zou J, Mao L, Qiu J, Wang M, Jia L, Wu D, He Z, Chen M, Shen Y, Shen E, Huang Y, Li R, Hu D, Shi L, Wang K, Zhu Q, Ye C, Bancroft I, King GJ, Meng J, Fan L (2019) Genome-wide selection footprints and deleterious variations in young Asian allotetraploid rapeseed. Plant Biotechnol J 17:1998–2010
Acknowledgements
We thank Dr. Ulrike Lohwasser from Leibniz Institute of Plant Genetics and Crop Plant Research, Germany, for providing a part of the rapeseed accessions for this study, Dr. Li Ruimin from Zhejiang Meteorological Bureau, for providing the meteorological data of the experimental sites, and Mr. Rui Sun from Agricultural Experiment Station of Zhejiang University for the management of field experiments.
Funding
The work was financially sponsored by Natural Science Foundation of China (code nos. 32130076 and 31961143008), Key Science and Technology Project of Zhejiang Province (2021C02057), and Jiangsu Collaborative Innovation Centre for Modern Crop Production.
Author information
Authors and Affiliations
Contributions
LJ conceived the experiments. YX carried out the field experiments and data analyses. XK contributed a lot to programming and data analyses. YG, RW, XY, XC, YL, JD, YZ, MC, and HC participated in field experiments in multiple locations and years. TY and DW were involved in data analyses and many constructive discussions. YX and LJ wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Communicated by Isobel AP Parkin.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
122_2023_4326_MOESM1_ESM.pdf
Fig. S1 Venn diagrams showing the presence-frequency difference (PFD) number of flowering time-related genes (FTRGs) in winter and semi-winter (blue color circle), spring and semi-winter (yellow color circle), spring and winter (red color circle) ecotypes, and the overlapped number of FTRGs (PDF 93 KB)
122_2023_4326_MOESM2_ESM.pdf
Fig. S2 The number of structural variations (SVs) in different frequency intervals in the 991 germplasms population (PDF 113 KB)
122_2023_4326_MOESM3_ESM.pdf
Fig. S3 Scatter plot showing structural variation (SV) frequency difference between the spring and semi-winter ecotypes (PDF 2029 KB)
122_2023_4326_MOESM4_ESM.pdf
Fig. S4 Line graph showing the frequencies of flowering time-related structural variations (SVs) that are significantly different between the spring and semi-winter ecotypes. The IDs of the genes are listed in Table S20 (PDF 116 KB)
122_2023_4326_MOESM5_ESM.pdf
Fig. S5 Scatter plot showing structural variation (SV) frequency difference between the spring and winter ecotypes (PDF 3065 KB)
122_2023_4326_MOESM6_ESM.pdf
Fig. S6 Line graph showing the frequencies of flowering time-related structural variations (SVs) that are significantly different between the spring and winter ecotypes. The IDs of the genes are listed in Table S21 (PDF 136 KB)
122_2023_4326_MOESM7_ESM.pdf
Fig. S7 Scatter plot showing structural variation (SV) frequency difference between the winter and semi-winter ecotypes (PDF 3359 KB)
122_2023_4326_MOESM8_ESM.pdf
Fig. S8 Line graph showing the frequencies of flowering time-related structural variations (SVs) that are significantly different between the winter and semi-winter ecotypes. The IDs of the genes are listed in Table S22 (PDF 141 KB)
122_2023_4326_MOESM9_ESM.pdf
Fig. S9 Histograms of the days from sowing to flowering (DSF) in the three locations in two successive years (i.e., six environments) (PDF 508 KB)
122_2023_4326_MOESM10_ESM.pdf
Fig. S10 Manhattan plots of GWAS-SNP on the days from sowing to flowering (DSF) in three locations in two successive years (i.e., six environments) (PDF 4637 KB)
122_2023_4326_MOESM11_ESM.pdf
Fig. S11 Accumulated temperature (a), precipitation (b), and sunshine hours (c) during the vegetative growth stage in the three locations in two successive years (i.e., six environments) (PDF 209 KB)
122_2023_4326_MOESM12_ESM.pdf
Fig. S12 Number of accessions and proportions of each ecotype in three categories differing from each other in the degree of flowering time rank (FTR) change across six environments. (a) Bar chart showing the number of accessions of each ecotype, (b)–(f) pie charts showing the number of accessions in each category (b), proportions of each ecotype in the whole GWAS population (c), in the FTR-consistent category (d), the FTR-moderately fluctuated category (e), and the FTR-drastically fluctuated category (f) (PDF 161 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, Y., Kong, X., Guo, Y. et al. Structural variations and environmental specificities of flowering time-related genes in Brassica napus. Theor Appl Genet 136, 42 (2023). https://doi.org/10.1007/s00122-023-04326-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-023-04326-w