Abstract
Key message
Fifteen and eleven loci, with most loci being novel, were identified to associate with seedling and adult resistances, respectively, to the durum-specific races of leaf rust pathogen in cultivated emmer.
Abstract
Leaf rust, caused by Puccinia triticina (Pt), constantly threatens durum (Triticum turgidum ssp. durum) and bread wheat (Triticum aestivum) production worldwide. A Pt race BBBQD detected in California in 2009 poses a potential threat to durum production in North America because resistance source to this race is rare in durum germplasm. To find new resistance sources, we assessed a panel of 180 cultivated emmer wheat (Triticum turgidum ssp. dicoccum) accessions for seedling resistance to BBBQD and for adult resistance to a mixture of durum-specific races BBBQJ, CCMSS, and MCDSS in the field, and genotyped the panel using genotype-by-sequencing (GBS) and the 9 K SNP (Single Nucleotide Polymorphism) Infinium array. The results showed 24 and nine accessions consistently exhibited seedling and adult resistance, respectively, with two accessions providing resistance at both stages. We performed genome-wide association studies using 46,383 GBS and 4,331 9 K SNP markers and identified 15 quantitative trait loci (QTL) for seedling resistance located mostly on chromosomes 2B and 6B, and 11 QTL for adult resistance on 2B, 3B and 6A. Of these QTL, one might be associated with leaf rust resistance (Lr) gene Lr53, and two with the QTL previously reported in durum or hexaploid wheat. The remaining QTL are potentially associated with new Lr genes. Further linkage analysis and gene cloning are necessary to identify the causal genes underlying these QTL. The emmer accessions with high levels of resistance will be useful for developing mapping populations and adapted durum germplasm and varieties with resistance to the durum-specific races.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat (Triticum aestivum L., 2n = 6x = 42) leaf (brown) rust, caused by the fungus Puccinia triticina Eriks. (Pt), is one of the three most important rust diseases of wheat (Anikster et al. 1997). Although leaf rust is less damaging than stem rust and stripe rust, it causes higher annual yield losses due to its frequent and widespread occurrence in most of the wheat-growing areas around the world (Chai et al. 2020). Among the wheat diseases, leaf rust is now ranked as the most devastating disease, affecting 94.4% of global wheat production and causing 3.25% yield loss (a value of $5.6 billion) annually at the global level (Savary et al. 2019; Chai et al. 2020). In some hotspots such as Indo-Gangetic Plain and Mainland China, annual yield losses were estimated at 4.25% and 4.38%, respectively (Savary et al. 2019). Although leaf rust caused the lowest yield loss annually in North America (0.54%) compared to other hotspots investigated by Savary et al. (2019), its regular outbreaks also had a major impact on wheat production in this region (Aboukhaddour et al. 2020; Kolmer et al. 2022). For instance, the losses caused by leaf rust in Canada reached up to 10% of total yield per year between 2000 and 2009 (Huerta-Espino et al. 2011). In the United States, a leaf rust outbreak in 2007 caused yield losses up to 14% in Kansas (Kolmer et al. 2009). A recent estimation indicated that leaf rust caused yield losses at 8% in Oklahoma and 3% in Texas and Kansas in 2019 (Kolmer et al. 2022).
Leaf rust will be a continuous threat to global wheat production. Chai et al. (2020) estimated that leaf rust could cause annual grain losses of 8.6–18.3 million metric tons, which are equivalent to economic values of $1.5–$3.3 billion, projected from 2000 to 2050. To minimize the occurrence and spread of leaf rust disease, the wheat community has placed a major effort on the utilization of host plant resistance. For rust diseases, resistance has been broadly classified at seedling and adult stages. Seedling resistance is pathogen race-specific and is controlled by single resistance genes with major effects that confer a hypersensitive response to prevent the spread of infection (Ellis et al. 2014; Mondal et al. 2016). In contrast, adult plant resistance is mostly race non-specific and effective at later stages of plant growth, and it is often controlled by multiple genes with minor and additive effects and found to be more durable (Zhang et al. 2021). To date, 81 leaf rust resistance (Lr) genes have been identified from common wheat and its wild and cultivated relatives (McIntosh et al. 2013, 2014, 2016, 2018, 2020; Xu et al. 2022). Most of the reported Lr genes confer race-specific resistance, except for Lr34 (Singh et al. 2012), Lr46 (Singh et al. 2013), Lr67 (Herrera-Foessel et al. 2014) and Lr68 (Herrera-Foessel et al. 2012). It has been commonly accepted that combinations of certain race-specific and race-non-specific resistance genes can enhance the level and durability of resistance against different Pt races at both seedling and adult stages. Thus far, six Lr genes have been cloned, including Lr1 (Cloutier et al. 2007), Lr10 (Feuillet et al. 2003), Lr21 (Huang et al. 2003), Lr22a (Thind et al. 2017), Lr34 (Krattinger et al. 2009) and Lr67 (Moore et al. 2015).
The Pt pathogen displays high genetic diversity, with new virulent races constantly emerging and evolving in nature (Huerta–Espino 1992; Kolmer 2005; Omara et al. 2021; Komer et al. 2022). For example, Pt race BBG/BN, detected in northwestern Mexico in 2001 with virulence to Lr72, overcame the resistance of widely adapted durum wheat [T. turgidum ssp. durum (Desf.) Husn., 2n = 4x = 28, AABB] cultivars from the CIMMYT (International Maize and Wheat Improvement Center) breeding program, which had been effective for more than 25 years (Singh et al. 2004; Aoun et al. 2016). This resulted in severe epidemics of leaf rust in durum wheat between 2000 and 2003, causing heavy yield loss worth $32 million (Singh et al. 2004). In addition, a new variant, BBG/BP, with virulence to Lr12, Lr27 and Lr31, was detected in durum cultivars between 2007 and 2008 (Huerta–Espino et al. 2008). In the US, a highly virulent race similar to BBG/BN, designated as BBBQD, was detected in durum fields of California in 2009 and Kansas in 2013 (Kolmer 2013, 2015). This poses a threat to the major durum-producing regions of the US such as North Dakota, which produces 58% of the total US durum wheat (Aoun et al. 2016). Therefore, it is necessary to identify resistance genes against race BBBQD among the existing wheat gene pools.
Cultivated emmer wheat [T. turgidum ssp. dicoccum (Schrank ex Schübler) Thell., 2n = 4x = 28, AABB] is one of the oldest domesticated crops, and an important genetic resource for the improvement of durum and bread wheat. Several breeding programs have utilized cultivated emmer to develop wheat cultivars with good resistance to rust and other fungal diseases (Zaharieva et al. 2010; Sharma et al. 2019). Wild emmer wheat [T. turgidum ssp. dicoccoides (Körn. ex Asch. and Graebner) Thell.], the wild progenitor of cultivated emmer, served as the source of Lr53 and Lr64 for leaf rust resistance (Marais et al. 2005; McIntosh et al. 2013). However, the presence of Lr genes in the germplasm collection of cultivated emmer has not been explored. Therefore, we conducted this study with aims to find sources of resistance to the durum-specific Pt races in a collection of cultivated emmer accessions and to identify quantitative trait loci (QTL) contributing to leaf rust resistance.
Materials and methods
Plant materials
A diverse panel of 180 cultivated emmer wheat accessions, originating from 32 countries and covering 5 continents, was used in this study (Table S1). These cultivated emmer wheat accessions are currently maintained at the National Small Grains Collection (NSGC, Aberdeen, ID), a component of the National Plant Germplasm System (NPGS) of the United States Department of Agriculture–Agriculture Research Service (USDA-ARS). The detailed source information was retrieved from USDA-ARS GRIN-Global (https://npgsweb.ars-grin.gov/gringlobal/search).
Evaluation of seedling resistance
Leaf rust resistance was evaluated at both seedling and adult stages. At the seedling stage, the 180 emmer accessions were evaluated for their reactions to Pt race BBBQD in the greenhouse. BBBQD is virulent to Lr10, Lr39, LrB and it is avirulent to Lr1, Lr2a, Lr2c, Lr3, Lr3ka, Lr9, Lr11, Lr14a, Lr16, Lr17, Lr18, Lr21, Lr24, Lr26, Lr28, Lr30, and Lr42 (Table 1). All the seedlings were screened at the North Dakota Agricultural Experiment Station (AES) Research Greenhouse Complex, Fargo, ND in 2013 and 2014 with three replicates each. Five seeds per accession were planted in a single cell in 50-cell trays containing sunshine mix #1 (Sungro Horticulture Distribution Inc., Quincy, MI, USA) and slow-release commercial fertilizer Osmocote 15-9-12, N-P-K (Everris NA Inc., OH, USA) in a rust-free greenhouse set to 22 °C/18 °C (day/night) with 16-h photoperiod. The leaf rust susceptible durum line RL6089 was included in each tray as the susceptible check. After seedling emergence, foliar fertilizer, Peat Lite 20-20-20, was applied once a week. Seedlings at the two-leaf stage were inoculated with fresh rust spores suspended in Soltrol-170 solvent (Phillips Petroleum, Bartlesville, OK, USA) at a rate of 0.01 g/mL and then left to air dry. After inoculation, plants were placed in a dark dew chamber for 16–18 h at 20 °C and were then relocated to the greenhouse. About 12–14 days after inoculation, the infection types (ITs) were scored using a 0–4 scale (McIntosh et al. 1995), where IT with 0 represents no visible sign or symptom, 1 represents small uredinia with necrosis, 2 represents small to medium sized uredinia with green islands and surrounded by necrosis or chlorosis, 3 represents medium sized uredinia with or without chlorosis, and 4 represents large uredinia without chlorosis. Two additional signs “−” and “+” indicate smaller and larger uredinia, respectively, for each basic level. For genome-wide association study (GWAS), the IT scores were converted to a linearized 0–9 scale (referred as infection response, IR) using the method described by Zhang et al. (2014). The conversion was performed in R using the scripts for ‘seedling rust score conversion’ provided by Gao et al. (2016) (https://github.com/umngao/rust_scores_conversion). The mean IR scores of six replicates were used for marker-trait association analysis. Genotypes with IR scores of 1–3 were considered as resistant, 4–6 as moderately resistant, and 7–9 as susceptible.
Evaluation of adult plant resistance
Leaf rust resistance at the adult stage was evaluated in a field nursery of the Debre Zeit Research Center, Ethiopian Institute of Agricultural Research, Ethiopia. The nursery site was located at 08° 46′ N, 39° 00′ E, and 1,900 m in elevation in Debre Zeit, Ethiopia. The evaluation experiments for the emmer panel were conducted with one replicate at the main-season (June-to-October growing season) of 2013 and two replicates at the off-season (December-to-April growing season) of 2014. The accessions were planted in single 1-m row plots and were exposed to natural Pt inoculum collected from the nursery. Common wheat variety ‘Thatcher’ was used as the susceptible check. The races of Pt population present in the collected spores were identified as BBBQJ, CCMSS and MCDSS (Table 1) by the USDA-ARS Cereal Disease Laboratory (St. Paul, MN).
The leaf rust reactions of each accession were scored as disease severity (DS) and infection response (IR) at the soft dough stage when the disease was fully developed and the symptoms had not been affected by host senescence (Newcomb et al. 2013). DS was represented as the percentage of tissue infected, and it was determined based on the modified Cobb Scale (Peterson et al. 1948). IR was represented as ‘R’ for resistant, ‘MR’ for moderately resistant, ‘MS’ for moderately susceptible, and ‘S’ for susceptible. ‘RMR’, ‘MRMS’, or ‘MSS’ represented mixed responses, with the predominant response being listed first (Newcomb et al. 2013). The IR evaluations were converted to a numeric 0–1 scale (Roelfs et al. 1992), and genotypes with the range of 0–0.3 were considered resistant, 0.4–0.6 as moderately resistant, and 0.7–1 as susceptible. The mean of DS and IR were multiplied to obtain the coefficient of infection (COI) (Yu et al. 2012), which was used for the association analysis for adult plant resistance. These phenotypic data of field studies were automatically converted to three numerical measures (DS, IR and COI) using the R scripts for ‘field rust score conversion’ provided by Gao et al. (2016) (https://github.com/umngao/rust_scores_conversion).
Welch’s t-test was performed for multiple replicates of each growing seasons and between 2013 and 2014 replicates at both seedling and adult stages using the ggbetweenstats() function in R package, ‘ggstatsplot’ (Patil 2021). Because no significant difference (p > 0.05) was observed among replicates for each growing season and between yearly replicates, the average of all replicates for seedlings and adult plants were taken and used for further analysis. The broad-sense heritability of traits was performed using the R package ‘variability’ with gen.var() function (Popat et al. 2020). Pearson's correlation between different disease traits of seedlings and adult plants was calculated using the ‘corrplot’ R package (Wei and Simko 2021).
Plant genotyping
The emmer panel was genotyped using the genotyping-by-sequencing (GBS) method as described in Poland et al. (2012) and Liu et al. (2020) and the wheat 9 K SNP Infinium iSelect array (Cavanagh et al. 2013) as per manufacturer’s guidelines (Illumina, San Diego, CA). For the GBS dataset, SNP discovery genotype calling was performed using the TASSEL-GBS pipeline (Glaubitz et al. 2014) with Bowtie2 alignment (Langmead and Salzberg 2012) to the IWGSC Chinese Spring RefSeq v1.0 reference genome (IWGSC 2018). SNP markers aligned on D genome were removed for further population structure and association analysis. The LD k-nearest neighbor (LD KNNi) imputation for missing genotypes (Money et al. 2015) and SNP filtering were performed using TASSEL 5.0 (Bradbury et al. 2007). SNP markers with > 5% minor allele frequency (MAF), genotypes with < 10% missing SNP calls, and markers with < 20% missingness were retained and used in GWAS analysis.
Population structure and principal component analyses
The population structure of the emmer panel was analyzed by randomly selecting 5000 SNPs from the GBS data. The genotype file was converted to population genetic data (PGD) format using PGDSpider 2.1.1.5 (Lischer and Excoffier 2012). The population structure was analyzed using STRUCTURE 2.3.4 (Pritchard et al. 2000), with both the burn-in period and Markov Chain Monte Carlo (MCMC) reps after burn-in set at 10,000. The constant alpha value was achieved over this period of time. Three iterations were performed for each simulated value of K, ranging from 1 to 10. The optimal K values for the population and subpopulation structures of the emmer panel were determined by structure harvester (http://taylor0.biology.ucla.edu/structureHarvester/) (Earl and vonHoldt 2012), using the log probability of the data [LnP(D)] and delta K (ΔK) based on the rate of change in [LnP(D)] between successive K values (Evanno et al. 2005). These population/subpopulation structures were also reflected as clusters in the principal component (PC) analyses performed using the R packages, ‘GAPIT3’ (Wang and Zhang 2021) and ‘stats’ with prcomp function (Team 2013). The number of PCs was chosen based on the scree plot. PCA plots were generated using the ‘ggplot2’ package in R (Wickham 2016).
Marker-trait associations and gene annotations
GWAS analyses were conducted to identify significant markers associated with seedling and adult plant resistance to leaf rust. These involve association of filtered SNPs of either the GBS or the 9 K SNP array with seedling (IR) or adult plant (COI) responses of the 180 emmer accessions. Four statistical models were implemented for GWAS analysis in GAPIT3 (Wang and Zhang 2021) with population structure and kinship accounted for. These included the single-locus mixed linear model (MLM) (Zhang et al. 2010), and the three higher-power, multi-locus models multiple-locus MLM (MLMM), Fixed and random model circulating probability unification (FarmCPU) (Liu et al. 2016), and Bayesian-information and linkage-disequilibrium iteratively nested keyway (Blink) (Huang et al. 2018). Phenotypic variances explained (R2, % PVE) by the individual SNPs were derived from the MLM model. The Q-Q plots were used to evaluate the best fit model for seedling and adult plant resistance using both genotypic datasets. Based on the Q-Q plots, the significance threshold for marker-trait associations was set at p ≤ 0.0001 [−log10(p) ≥ 4]. Manhattan plots were generated using the ‘CMplot’ package in R (https://github.com/YinLiLin/CMplot) (Yin et al. 2021).
Annotation for the associated markers for seedling and adult plant resistance were assigned based on the closest high-confidence annotated gene identified near the SNP physical location using the T. aestivum IWGSC RefSeq v1.0 reference genome (IWGSC 2018). To identify novel resistance genes in the cultivated emmer panel, the physical positions of significant SNPs were compared to the markers associated with known Lr genes, which were previously mapped to the IWGSC RefSeq v1.0 by two earlier studies (Fatima et al. 2020; Zhang et al. 2021).
Results
Distribution and genotyping of the emmer panel
A diverse panel of 180 cultivated emmer accessions from USDA-ARS National Small Grains Collection were evaluated to identify loci associated with leaf rust resistance in the emmer population. The emmer accessions were originally collected from 32 different countries around the world, with the majority from Ethiopia, Spain, Russia, India and Serbia (Table S1).
The emmer panel was genotyped by GBS and the 9 K SNP Infinium array, resulting in 66,215 and 5911 polymorphic SNPs, respectively. After LD KNNi imputation and filtration, 46,383 SNPs from the GBS dataset (Table S2) and 4331 SNPs from the 9 K SNP array (Table S3) were retained with > 5% minor allele frequency (MAF) and < 20% missingness. The distribution of the filtered SNPs on the 14 chromosomes ranged between 2060 and 4553 SNPs from the GBS dataset (Fig. 1a) and 121–513 SNPs from the 9 K SNP array (Fig. 1b). In both cases, chromosome 2B was the most densely populated with SNPs and 4B was the least.
Chromosomal distribution of filtered SNPs on the A and B genomes of cultivated emmer using a GBS dataset and b 9 K SNP array based on the alignment to the IWGSC Chinese Spring RefSeq v1.0 (IWGSC 2018) and the consensus wheat SNP map in Cavanagh et al. (2013), respectively
Population structure and principal component analyses
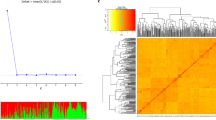
To determine the population structure of emmer panel, 5000 SNPs from the GBS dataset were randomly selected and analyzed in STRUCTURE 2.3.4 (Pritchard et al. 2000). Distinct peaks were observed at K = 2 and K = 7, suggesting two major populations and seven subpopulations in the emmer panel (Fig. 2a). The population clusters based on the optimal K values were also assessed in principal component analyses. The scree plot revealed the first three PCs to explain most of the genetic variation (57%) present in the emmer panel (Fig. 2b). At K = 2, the first two PCs revealed the majority of emmer accessions from Africa were clustered in one group, whereas Europe, North and South America in a separate group (Fig. 2c, d). Emmer accessions from Asia were divided into both groups (Fig. 2c, d), with India and Iran as the two major countries that clustered with Africa and Europe, respectively (Table S1). At K = 7, the second and third PCs clustered the emmer accessions from Ethiopia (Africa), India (Asia) and Spain (Europe) into the respective Subpopulation 1, 2 and 3 (Fig. 2e, f, Table S1). The formation of other clusters was based on a close geographic proximity. For example, most of the emmer accessions from Former Yugoslavia and Serbia (Asia and Europe) were clustered into Subpopulation 5, and Iran and Georgia (Asia and Europe) into Subpopulation 6. Emmer accessions belonging to the rest of European countries were clustered into Subpopulation 4, whereas no specific cluster was observed for Subpopulation 7.
Population structure analysis of the emmer panel based on 5000 randomly selected SNPs from the GBS dataset. a The plot shows delta K (ΔK) for different number of populations (K) assumed in the analysis. b The scree plot displaying the percent variance explained by the principal component (PC) analyses. The first two PCs clustering the emmer panel based on c continents and d K = 2 populations. The second and third PCs clustering the emmer panel based on e continents and f K = 7 subpopulations
Phenotyping of the emmer panel
The emmer panel was evaluated for leaf rust responses at seedling and adult stages. The raw phenotypic data and calculated infection response (IR) values at the seedling stage and coefficient of infection (COI) values at the adult stage of all emmer accessions are presented in Table S4. The infection response against Pt race BBBQD at the seedling stage in greenhouse had a mean of 6.3 based on the 0–9 scale (Table 2). In the field nurseries, adult plants inoculated with the mixture of Pt races BBBQJ, CCMSS and MCDSS had mean leaf rust responses of 11.2, 0.7 and 9.1% for DS, IR and COI, respectively (Table 2). These mean values were derived from both 2013 and 2014 replicates. No significant differences were observed between the yearly replicates for seedlings (p = 0.294) and adult plants (p = 0.054) based on Welch's t-test (Fig. S1).
Significant phenotypic variations were observed for all the leaf rust responses at both seedling and adult stages (Fig. 3). However, IR at both stages were skewed more toward susceptibility (Fig. 3a, b), whereas DS was skewed toward low severity (Fig. 3c). COI represented the product of IR and DS (Fig. 3d); thus, these traits were positively correlated. A stronger correlation was detected between COI and DS (0.92) than COI and IR (0.45) (Fig. 3e). The leaf rust responses of seedlings and adult plants exhibited positive, but weak correlations.
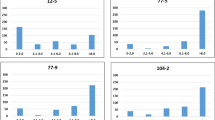
Leaf rust responses of the emmer panel. Histograms of a infection response (IR) at the seedling stage; b IR, c disease severity (DS) and d coefficient of infection (COI) at the adult stage; e Correlation between different leaf rust responses at the seedling and adult stages. f The susceptibility of emmer accessions at the seedling and adult stages based on linearized infection responses (a, b)
With regards to leaf rust responses, 16% of the emmer accessions were resistant, 16% were moderately resistant, and 68% were susceptible at the seedling stage (Fig. 3f). At the adult stage, 2%, 31% and 66% of emmer accessions were resistant, moderately resistant, and susceptible, respectively. As evident, the majority of emmer accessions were susceptible at both stages. However, more accessions were found to be resistant at the seedling stage and moderately resistant at the adult stage. Among 180 accessions, 24 showed consistent resistance to Pt race BBBQD at seedling stage across two test experiments (Table 3) and nine had consistent adult resistance in the field nursey in 2013 and 2014 (Table 4). Two accessions (PI 94674–1 and PI 94738–1) showed consistent resistance among replicates at both stages, whereas 79 accessions were susceptible at both stages (Table S4).
Marker-trait associations
Four statistical models were primarily implemented for GWAS analyses to find marker-trait associations for seedling and adult plant resistance using both the GBS dataset (Table S5-S6) and the 9 K SNP array (Table S7, S8). The assessment of Q-Q plots revealed FarmCPU to be the best fit model for the GBS dataset (Fig. S2, S3) and MLMM for the 9 K SNP array (Fig. S3, S4). Using the GBS dataset, 13 SNPs were identified to significantly (p < 0.0001) associate with seedling resistance (Fig. 4a) and 15 SNPs with adult plant resistance (Fig. 4b). Using the 9 K SNP array, 9 SNPs were found to significantly (p < 0.0001) associate with seedling resistance (Fig. 5a) and 2 SNPs with adult plant resistance (Fig. 5b). No overlap in significant SNPs were observed between the seedling and adult plant resistance as well as between the GBS and 9 K SNP array datasets.
Manhattan plots displaying SNPs from the GBS dataset that are significantly associated with a infection response of cultivated emmer at the seedling stage, and b coefficient of infection at the adult stage, based on the FarmCPU model. The significance threshold was set at [−log10(p) ≥ 4] (dashed line). The y-axis indicates −log10(p) for GBS markers. The x-axis indicates the physical distribution of all the GBS markers on the 14 chromosomes of cultivated emmer based on the alignment to the IWGSC RefSeq v1.0 (IWGSC 2018)
Manhattan plots displaying SNPs from the 9 K SNP array that are significantly associated with a infection response of cultivated emmer at the seedling stage, and b coefficient of infection at the adult stage, based on the MLMM model. The significance threshold was set at [−log10(p) ≥ 4] (dashed line). The y-axis indicates −log10(p) for SNP array markers. The x-axis indicates the positions of all the SNP array markers on the 14 chromosomes of cultivated emmer based on the consensus wheat SNP map in Cavanagh et al. (2013)
We further performed the Welch’s t-test to retain only the associated SNPs that displayed a significant phenotypic difference (p ≤ 0.05) between the alternative alleles. This resulted in a total of 15 QTL for seedling resistance using both genotypic datasets, which were found on chromosomes 2A, 2B, 3B, 4A, 5A, 6B, 7A and 7B (Table 5). Among these, wsnp_Ex_c5072_9006666 on chromosome 4A and wsnp_Ex_c1996_3754394 on 2B contributed the most to the phenotypic variance (> 15%). These markers were found near the genomic regions of genes encoding DNA methyltransferase and cytochrome P450. The remaining loci have minor (< 5%) to moderate (< 10%) effects on the phenotypic variance. On the other hand, 11 QTL were identified for adult plant resistance that display significant marker-trait associations between the alternative alleles (Table 5). These QTL were located on chromosomes 1A, 2B, 3B, 4A, 5B, 6A and 6B, and the majority of them exhibited minor effects on the phenotypic variance. Markers on chromosome 3B such as S3B_330595912 and S3B_583036386 displayed moderate effects and mapped near genes that encode protein MIZU-KUSSEI 1 and RING/U-box superfamily protein, respectively.
Discussion
Constant evolution of P. triticina population frequently renders the deployed Lr genes ineffective, necessitating new sources of leaf rust resistance. In the primary gene pool of wheat, cultivated emmer is an excellent source of useful genes for improving modern wheat germplasm. Resistance genes for stem rust such as Sr2/Yr30 and Sr13 were originally derived from cultivated emmer, which are now widely deployed in modern durum and/or bread wheat cultivars (McFadden 1930; Williams and Gough 1965; Zaharieva et al. 2010; Sharma et al. 2019). Stripe/yellow rust resistance genes such as Yr15, Yr35, Yr36, and YrH52, and leaf rust resistance genes such as Lr53 and Lr64 were discovered in the wild progenitor of cultivated emmer, and successfully transferred to modern wheat cultivars (Gerechter–Amitai et al. 1989; Peng et al. 1999; Uauy et al. 2005; Fu et al. 2009; Dadkhodaie et al. 2011; McIntosh et al. 2013). Despite their importance as resistance sources for rust diseases, the large collections of cultivated emmer have not been explored for leaf rust resistance.
The study herein evaluated the diverse panel of 180 cultivated emmer against the highly virulent Pt race BBBQD at the seedling stage and mixed races BBBQJ, CCMSS and MCDSS at the adult stage in the nursery field. Among these accessions, 24 (13.3%) had consistent seedling resistance to Pt race BBBQD at seedling stage and nine (5.0%) had consistent adult resistance. In addition, approximately 31 (17.2%) accessions had variable expressions for reactions to BBBQD at seedling stage among the replications in the two experiments, with most of the plants in these accessions having resistant reactions. This result indicated that cultivated emmer is good source for resistance to leaf rust. Aoun et al. (2016) evaluated 496 durum wheat accessions for resistance to the durum wheat-specific races including BBBQD, BBBQJ, and MCDSS used in this study. They found that only 4.9% (24) of 496 durum accessions showed resistance to BBBQD, indicating that source of resistance to this race is rare in durum germplasm. Therefore, the cultivated emmer accessions with resistance to BBBQD may carry novel Lr genes that could be used to improve modern durum crops for resistance to this newly emerged Pt race. In addition, a few of emmer wheat accessions from Africa with adult resistance to mixed races BBBQJ, CCMSS and MCDSS may also have unique Lr genes. To detect the genomic regions harboring the Lr genes, we performed GWAS analyses on the emmer panel and identified few overlapping loci with previously reported Lr genes or QTL in addition to finding new loci.
Seedling resistance is considered effective at all stages of plant growth, and it is often race-specific and controlled by major effect resistance genes (Ellis et al. 2014; Zetzsche et al. 2019). In the cultivated emmer panel, we identified 15 QTL that contributed to seedling resistance, of which, 2 QTL were found near the marker regions of two known Lr genes such as Lr28 and Lr53 (Table 5, S5). The moderate effect S-QLr4A.1 was mapped to the similar genomic regions where the markers (barc343-wmc219) associated with Lr28 were earlier detected (McIntosh and Te 1982; Sohail et al. 2014). However, Lr28 was reported to originate from chromosome 7S#2 of Aegilops speltoides, and translocated to chromosome 4AL of bread wheat (Naik et al. 1998). Therefore, finding Lr28 in the cultivated emmer genome might be unlikely and their shared genomic regions might have occurred by chance. On the other hand, a moderate effect QTL, S-QLr6B.2, was found < 1 Mb upstream of the Lr53-associated marker, CFD1 (Dadkhodaie et al. 2011; McIntosh et al. 2013). These QTL of cultivated emmer may represent the same loci contributing to resistance conferred by Lr53, the gene derived from its progenitor, wild emmer wheat.
Besides the catalogued Lr genes, many QTL contributing to leaf rust resistance have been identified in bread and durum wheat using meta-QTL analysis (Amo and Soriano 2022). The moderate effect S-QLr7A of the emmer was present close to durum wheat QTL, QLr.locus-7A.2 (~ 30 kb) (Table S7) (Wang et al. 2014; Aoun et al. 2016). These might represent the same QTL since the AB genomes of durum wheat are derived from emmer wheat. The remaining QTL (13/15) may represent novel loci in the emmer panel that potentially contribute to seedling resistance. A majority of these novel QTL exhibited moderate effects, among which, S-QLr4A.1 and S-QLr5A were significantly detected by three statistical models, and S-QLr3B.1 and S-QLr6B.3 by two models (Table S5). These QTL are strong candidates for novel Lr genes for seedling resistance. In addition, S-QLr4A.2 and S-QLr2B.1 had major effects, and transfer of such QTL might enhance seedling resistance in modern wheat cultivars.
Adult plant resistance is effective at later stages of plant growth and often race non-specific and controlled by multiple minor effect resistance genes (Zhang et al. 2021). In the emmer panel, 11 unique QTL were associated with adult plant resistance to leaf rust. The moderate effect A-QLr5B was present within the marker regions flanking Lr18 (Table 5, S6) (Carpenter et al. 2018; Fatima et al. 2020). Since Lr18 was originally derived from G genome of T. timopheevi and translocated to chromosome arm 5BL of bread wheat (Friebe et al. 1996), it is unlikely for the cultivated emmer to acquire the same Lr18 gene without translocation. Hence, the QTL found near the region of Lr18 might be a novel locus.
However, the minor effect A-QLr6B mapped to the overlapping regions as the markers (Xbarc5–Xgwm469.2) of QLr.fcu-6BL, which was previously identified to contribute to adult plant resistance in synthetic hexaploid wheat (SHW) (Table S6) (Chu et al. 2009). Hence, these QTL may represent the same loci in hexaploid wheat and emmer wheat. The remaining QTL (10/11) are potentially novel loci that confer adult plant resistance to leaf rust in the emmer panel. These QTL mostly provide minor to moderate effects and exist mainly on chromosomes 2B, 3B and 6A. Among these, A-QLr3B.2 and A-QLr3B.3 that contributed highest to the phenotypic variance were conferred significant by three or more statistical models (Table S6). In addition, A-QLr5B and A-QLr6A.1 were detected significant by two models. These QTL are strong candidates for adult plant resistance.
In this study, we identified 24 and nine cultivated emmer accessions having consistent seedling resistance to Pt race BBBQD and adult plant resistance to mixed races BBBQJ, CCMSS and MCDSS, respectively. These accessions should be a valuable resource for improving modern durum and bread wheat for resistance to leaf rust. A total of 15 and 11 QTL were identified to be associated with seedling and adult resistance, respectively. Interestingly, most of these QTL are in the genomic regions where no known Lr genes or QTL were previously reported. Furthermore, the cultivated emmer germplasm collections were not systematically explored for leaf rust resistance and none of the 79 catalogued Lr genes in wheat are derived from cultivated emmer. Therefore, some QTL might be potentially associated with new Lr genes. Many of the resistant emmer accessions (Tables 3 and 4) contain the resistant alleles of QTL identified herein especially S-QLr2B.2 and S-QLr4A.2 for seedling resistance (Table S9), and A-QLr-2B.1, A-QLr-2B.2, A-QLr-3B.2, A-QLr-3B.3, A-QLr-4A and A-QLr-5B for adult plant resistance (Table S10). These QTL likely represent novel Lr genes derived from the cultivated emmer.
However, the GWAS results presented in this study only provides preliminary information on the genomic regions potentially harboring Lr genes. The QTL identified need to be confirmed and validated through further linkage analysis, cloning, and functional genetics. The phenotypic and genotypic data and GWAS results presented in this study provide guidance to select certain emmer accessions as the parents for developing mapping populations that can be used to identify the causal genes underlying the QTL. Several emmer accessions, such as CItr 14916–1, CItr 14917–1, PI 273981–1, PI 275996–1, PI 349046–1, CItr 14639, CItr 14751, PI 193878, PI 361833, PI 384318, PI 479957, PI 480307, and PI 480313 consistently exhibited a high level of seedling resistance to Pt race BBBQD. Because Pt race BBBQD poses a potential threat to durum crop in North America and sources of resistance is rare in durum germplasm, the emmer accessions with a high level of resistance can be used as resistance donors in developing adapted durum germplasm and varieties with resistance to this race. Furthermore, the cultivated emmer panel and its associated genotypic data sets presented in this study will be useful for identifying and mapping the genes for resistance to other fungal diseases.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
Change history
23 March 2023
A Correction to this paper has been published: https://doi.org/10.1007/s00122-023-04323-z
References
Aboukhaddour R, Fetch T, McCallum BD, Harding MW, Beres BL, Graf RJ (2020) Wheat diseases on the prairies: a Canadian story. Plant Pathol 69:418–432. https://doi.org/10.1111/ppa.13147
Amo A, Soriano JM (2022) Unravelling consensus genomic regions conferring leaf rust resistance in wheat via meta-QTL analysis. Plant Genome 15(1):e20185. https://doi.org/10.1002/tpg2.20185
Anikster Y, Bushnell WR, Roelfs AP, Eilam T, Manisterski J (1997) Puccinia recondita causing leaf rust on cultivated wheats, wild wheats, and rye. Can J Bot 75:2082–2096. https://doi.org/10.1139/b97-919
Aoun M, Breiland M, Turner MK, Loladze A, Chao S, Xu SS, Ammar K, Anderson JA, Kolmer JA, Acevedo M (2016) Genome-wide association mapping of leaf rust response in a durum wheat worldwide germplasm collection. Plant Genome 9(3):plantgenome2016-01. https://doi.org/10.3835/plantgenome2016.01.0008
Aoun M, Kolmer JA, Rouse MN, Chao S, Bulbula WD, Elias EM, Acevedo M (2017) Inheritance and bulked segregant analysis of leaf rust and stem rust resistance in durum wheat genotypes. Phytopathology 107:1496–1506. https://doi.org/10.1094/PHYTO-12-16-0444-R
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) Tassel: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Carpenter NR, Griffey CA, Rosso L, Malla S, Chao S, Brown-Guedira GL (2018) Mapping Lr18: a leaf rust resistance gene widely deployed in soft red winter wheat. J Plant Dis Biomark 1(1):4–10
Cavanagh CR, Chao S, Wang S, Huang BE, Stephen S, Kiani S, Forrest K, Saintenac C, Brown-Guedira GL, Akhunova A, See D, Bai G, Pumphrey M, Tomar L, Wong D, Kong S, Reynolds M, da Silva ML, Bockelman H, Talbert L, Anderson JA, Dreisigacker S, Baenziger S, Carter A, Korzun V, Morrell PL, Dubcovsky J, Morell MK, Sorrells ME, Hayden MJ, Akhunov E (2013) Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc Natl Acad Sci USA 110:8057–8062. https://doi.org/10.1073/pnas.121713311
Chai Y, Pardey PG, Hurley TM, Senay SD, Beddow JM (2020) A probabilistic bio-economic assessment of the global consequences of wheat leaf rust. Phytopathology 110:1886–1896. https://doi.org/10.1094/PHYTO-02-20-0032-R
Chu C-G, Friesen TL, Xu SS, Faris JD, Kolmer JA (2009) Identification of novel QTLs for seedling and adult plant leaf rust resistance in a wheat doubled haploid population. Theort Appl Genet 119:263–269. https://doi.org/10.1007/s00122-009-1035-0
Cloutier S, McCallum BD, Loutre C, Banks TW, Wicker T, Feuillet C, Keller B, Jordan MC (2007) Leaf rust resistance gene Lr1, isolated from bread wheat (Triticum aestivum L.) is a member of the large psr567 gene family. Plant Mol Biol 65:93–106. https://doi.org/10.1007/s11103-007-9201-8
Dadkhodaie NA, Karaoglou H, Wellings CR, Park RF (2011) Mapping genes Lr53 and Yr35 on the short arm of chromosome 6B of common wheat with microsatellite markers and studies of their association with Lr36. Theor Appl Genet 122:479–487. https://doi.org/10.1007/s00122-010-1462-y
Earl DA, vonHoldt BM (2012) Structure harvester: a website and program for visualizing structure output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Ellis JG, Lagudah ES, Spielmeyer W, Dodds PN (2014) The past, present and future of breeding rust resistant wheat. Front Plant Sci 5:641. https://doi.org/10.3389/fpls.2014.00641
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Fatima F, McCallum BD, Pozniak CJ, Hiebert CW, McCartney CA, Fedak G, You FM, Cloutier S (2020) Identification of new leaf rust resistance loci in wheat and wild relatives by array-based SNP genotyping and association genetics. Front Plant Sci 11:583738. https://doi.org/10.3389/fpls.2020.583738
Feuillet C, Travella S, Stein N, Albar L, Nublat A, Keller B (2003) Map-based isolation of the leaf rust disease resistance gene Lr10 from the hexaploid wheat (Triticum aestivum L.) genome. Proc Natl Acad Sci USA 100:15253–15258. https://doi.org/10.1073/pnas.2435133100
Friebe B, Jiang J, Raupp WJ, McIntosh RA, Gill BS (1996) Characterization of wheat-alien translocations conferring resistance to diseases and pests: current status. Euphytica 91:59–87. https://doi.org/10.1007/bf00035277
Fu D, Uauy C, Distelfeld A, Blechl A, Epstein L, Chen X, Sela H, Fahima T, Dubcovsky J (2009) A kinase-START gene confers temperature-dependent resistance to wheat stripe rust. Science 323:1357–1360. https://doi.org/10.1126/science.1166289
Gao L, Turner MK, Chao S, Kolmer J, Anderson JA (2016) Genome wide association study of seedling and adult plant leaf rust resistance in elite spring wheat breeding lines. PLoS ONE 11(2):e0148671. https://doi.org/10.1371/journal.pone.0148671
Gerechter-Amitai ZK, van Silfhout CH, Grama A, Kleitman F (1989) Yr15 - a new gene for resistance to Puccinia striiformis in Triticum dicoccoides sel. G-25. Euphytica 43:187–190. https://doi.org/10.1007/bf00037912
Glaubitz JC, Casstevens TM, Lu F, Harriman J, Elshire RJ, Sun Q, Buckler ES (2014) TASSEL-GBS: a high capacity genotyping by sequencing analysis pipeline. PLoS ONE 9(2):e90346. https://doi.org/10.1371/journal.pone.0090346
Herrera-Foessel SA, Singh RP, Huerta-Espino J, Rosewarne GM, Periyannan SK, Viccars L, Calvo-Salazar V, Lan C, Lagudah ES (2012) Lr68: a new gene conferring slow rusting resistance to leaf rust in wheat. Theort Appl Genet 124:1475–1486. https://doi.org/10.1007/s00122-012-1802-1
Herrera-Foessel SA, Singh RP, Lillemo M, Huerta-Espino J, Bhavani S, Singh S, Lan C, Calvo-Salazar V, Lagudah ES (2014) Lr67/Yr46 confers adult plant resistance to stem rust and powdery mildew in wheat. Theort Appl Genet 127:781–789. https://doi.org/10.1007/s00122-013-2256-9
Huang L, Brooks SA, Li W, Fellers JP, Trick HN, Gill BS (2003) Map-based cloning of leaf rust resistance gene Lr21 from the large and polyploid genome of bread wheat. Genetics 164:655–664. https://doi.org/10.1093/genetics/164.2.655
Huang M, Liu X, Zhou Y, Summers RM, Zhang Z (2018) Blink: a package for the next level of genome-wide association studies with both individuals and markers in the millions. Gigascience 8(2):1–12. https://doi.org/10.1093/gigascience/giy154
Huerta-Espino J, Singh RP, Herrera-Foessel SA, Pérez-López JB, Figueroa-López P (2008) First detection of virulence in Puccinia triticina to resistance genes Lr27 + Lr31 present in durum wheat in Mexico. Plant Dis 93:110. https://doi.org/10.1094/PDIS-93-1-0110C
Huerta-Espino J, Singh RP, Germán S, McCallum BD, Park RF, Chen WQ, Bhardwaj SC, Goyeau H (2011) Global status of wheat leaf rust caused by Puccinia triticina. Euphytica 179:143–160. https://doi.org/10.1007/s10681-011-0361-x
Huerta-Espino J (1992) Analysis of wheat leaf and stem rust virulence on a worldwide basis. PhD thesis, University of Minnesota, St Paul
International Wheat Genome Sequencing Consortium (IWGSC) (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361(6403):eaar7191. https://doi.org/10.1126/science.aar7191
Kolmer JA (2005) Tracking wheat rust on a continental scale. Curr Opin Plant Biol 8:441–449. https://doi.org/10.1016/j.pbi.2005.05.001
Kolmer J (2013) Leaf rust of wheat: pathogen biology, variation and host resistance. Forests 4(1):70–84. https://doi.org/10.3390/f4010070
Kolmer JA (2015) First report of a wheat leaf rust (Puccinia triticina) phenotype with high virulence to durum wheat in the Great Plains region of the United States. Plant Dis 99:156. https://doi.org/10.1094/PDIS-06-14-0667-PDN
Kolmer JA, Acevedo MA (2016) Genetically divergent types of the wheat leaf fungus Puccinia triticina in Ethiopia, a center of tetraploid wheat diversity. Phytopathology 106:380–385. https://doi.org/10.1094/PHYTO-10-15-0247-R
Kolmer JA, Long DL, Hughes ME (2009) Physiologic specialization of Puccinia triticina on wheat in the United States in 2007. Plant Dis 93:538–544. https://doi.org/10.1094/PDIS-93-5-0538
Kolmer JA, Herman AD, Fellers JP (2022) Genotype groups of the wheat leaf rust fungus Puccinia triticina in the United States as determined by genotyping by sequencing. Phytopathology 112:653–662. https://doi.org/10.1094/PHYTO-03-21-0125-R
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363. https://doi.org/10.1126/science.1166453
Langmead B, Salzberg S (2012) Fast gapped-read alignment with Bowtie2. Nat Methods 9:357–359. https://doi.org/10.1038/nmeth.1923
Lischer HEL, Excoffier L (2012) PGDSpider: an automated data conversion tool for connecting population genetics and genomics programs. Bioinformatics 28:298–299. https://doi.org/10.1093/bioinformatics/btr642
Liu X, Huang M, Fan B, Buckler ES, Zhang Z (2016) Iterative usage of fixed and random effect models for powerful and efficient genome-wide association studies. PLoS Genet 12(3):e1005767. https://doi.org/10.1371/journal.pgen.1005767
Liu Y, Zhang Q, Salsman E, Fiedler JD, Hegstad JB, Liu Z, Faris JD, Xu SS, Li X (2020) QTL mapping of resistance to tan spot induced by race 2 of Pyrenophora tritici-repentis in tetraploid wheat. Theort Appl Genet 133:433–442. https://doi.org/10.1007/s00122-019-03474-2
Marais GF, Pretorius ZA, Wellings CR, McCallum B, Marais AS (2005) Leaf rust and stripe rust resistance genes transferred to common wheat from Triticum dicoccoides. Euphytica 143:115–123
McFadden ES (1930) A successful transfer of emmer characters to vulgare wheat. J Am Soc Agron 22:1020–1034
McIntosh RA, Miller TE, Chapman V (1982) Cytogenetical studies in wheat XII. Lr28 for resistance to Puccinia recondita and Sr34 for resistance to P. graminis tritici. Z Pflanzenzucht 89:295–306
McIntosh RA, Wellings CR, Park RF (1995) Wheat rusts: an atlas of resistance genes. CSIRO Publications, Melbourne
McIntosh RA, Dubcovsky J, Rogers WJ, Morris C, Appels R, Xia XC (2013) Catalogue of gene symbols for wheat. https://shigen.nig.ac.jp/wheat/komugi/genes/symbolClassList.jsp Accessed 23 Oct 2022
McIntosh RA, Dubcovsky J, Rogers WJ, Morris C, Appels R, Xia XC (2014) Catalogue of gene symbols for wheat: 2013–2014 supplement. https://shigen.nig.ac.jp/wheat/komugi/genes/symbolClassList.jsp Accessed 23 Oct 2022
McIntosh RA, Dubcovsky J, Rogers WJ, Morris C, Appels R, Xia XC (2016) Catalogue of gene symbols for wheat: 2015–2016 supplement. https://shigen.nig.ac.jp/wheat/komugi/genes/symbolClassList.jsp Accessed 23 Oct 2022
McIntosh RA, Dubcovsky J, Rogers WJ, Xia XC, Raupp WJ (2018) Catalogue of gene symbols for wheat: 2018 supplement. https://wheat.pw.usda.gov/GG3/wgc Accessed 23 Oct 23, 2022
McIntosh RA, Dubcovsky J, Rogers WJ, Xia XC, Raupp WJ (2020) Catalogue of gene symbols for wheat: 2020 supplement. https://wheat.pw.usda.gov/GG3/wgc Accessed 23 Oct 2022
Mondal S, Rutkoski JE, Velu G, Singh PK, Crespo-Herrera LA, Guzmán C, Bhavani S, Lan C, He X, Singh RP (2016) Harnessing diversity in wheat to enhance grain yield, climate resilience, disease and insect pest resistance and nutrition through conventional and modern breeding approaches. Front Plant Sci 7:991. https://doi.org/10.3389/fpls.2016.00991
Money D, Gardner KM, Migicovsky Z, Schwaninger H, Zhong GY, Myles S (2015) LinkImpute: fast and accurate genotype imputation for nonmodel organisms. G3: Genes Genomes Genet 5:2383–2390. https://doi.org/10.1534/g3.115.021667
Moore JW, Herrera-Foessel S, Lan C, Schnippenkoetter W, Ayliffe M, Huerta-Espino J, Lillemo M, Viccars L, Milne R et al (2015) A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat Genet 47:1494–1498. https://doi.org/10.1038/ng.3439
Naik S, Gill KS, Prakasa Rao VS, Gupta VS, Tamhankar SA, Pujar S, Gill BS, Ranjekar PK (1998) Identification of a STS marker linked to the Aegilops speltoides-derived leaf rust resistance gene Lr28 in wheat. Theort Appl Genet 97:535–540. https://doi.org/10.1007/s001220050928
Newcomb M, Acevedo M, Bockelman HE, Brown-Guedira G, Goates BJ, Jackson EW, Jin Y, Njau P, Rouse MN, Singh D, Wanyera R, Bonman JM (2013) Field resistance to the Ug99 race group of the stem rust pathogen in spring wheat landraces. Plant Dis 97:882–890. https://doi.org/10.1094/PDIS-02-12-0200-RE
Omara RI, Nehela Y, Mabrouk OI, Elsharkawy MM (2021) The emergence of new aggressive leaf rust races with the potential to supplant the resistance of wheat cultivars. Biology (basel) 10(9):925. https://doi.org/10.3390/biology10090925
Patil I (2021) Visualizations with statistical details: the ‘ggstatsplot’ approach. J Open Source Software 6(61):3167. https://doi.org/10.21105/joss.03167
Peng JH, Fahima T, Röder MS, Li YC, Dahan A, Grama A, Ronin YI, Korol AB, Nevo E (1999) Microsatellite tagging of the stripe-rust resistance gene YrH52 derived from wild emmer wheat, Triticum dicoccoides, and suggestive negative crossover interference on chromosome 1B. Z Genet Breed Res 98:862–872. https://doi.org/10.1007/s001220051145
Peterson RF, Campbell AB, Hannah AE (1948) A diagrammatic scale for estimating rust intensity on leaves and stems of cereals. Can J Res 26:496–500. https://doi.org/10.1139/cjr48c-033
Poland JA, Brown PJ, Sorrells ME, Jannink J-L (2012) Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping-by-sequencing approach. PLoS ONE 7(2):e32253. https://doi.org/10.1371/journal.pone.0032253
Popat R, Patel R, Parmar D (2020) Variability: genetic variability analysis for plant breeding research. R package version 0.1.0. https://cran.r-project.org/web/packages/variability/index.html
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
Roelfs AP, Singh RP, Saari EE (1992) Rust diseases of wheat: concepts and methods of disease management. CIMMYT, Mexico
Savary S, Willocquet L, Pethybridge SJ, Esker P, McRoberts N, Nelson A (2019) The global burden of pathogens and pests on major food crops. Nat Ecol Evol 3:430–439. https://doi.org/10.1038/s41559-018-0793-y
Sharma JS, Zhang Q, Rouse MN, Klindworth DL, Friesen TL, Long Y, Olivera PD, Jin Y, McClean PE, Xu SS, Faris JD (2019) Mapping and characterization of two stem rust resistance genes derived from cultivated emmer wheat accession PI 193883. Theort Appl Genet 132:3177–3189. https://doi.org/10.1007/s00122-019-03417-x
Singh RP, Huerta-Espino J, Pfeiffer W, Figueroa-Lopez P (2004) Occurrence and impact of a new leaf rust race on durum wheat in northwestern Mexico from 2001 to 2003. Plant Dis 88:703–708. https://doi.org/10.1094/PDIS.2004.88.7.703
Singh R, Herrera-Foessel S, Huerta-Espino J, Bariana H, Bansal U, Mccallum B, Hiebert C, Bhavani S, Singh S, Lan C, Lagudah ES (2012) Lr34/Yr18/Sr57/Pm38/Bdv1/Ltn1 confers slow rusting, adult plant resistance to Puccinia graminis tritici. In: The 13th International Cereal Rusts and Powdery Mildews Conference, Beijing, China
Singh RP, Herrera-Foessel SA, Huerta-Espino J, Lan C, Basnet BR, Bhavani S, Singh RP, Herrera-Foessel SA, Espino J, Lan CX, Basnet BR, Lagudah ES (2013) Pleiotropic gene Lr46/Yr29/Pm39/Ltn2 confers slow rusting, adult plant resistance to wheat stem rust fungus. In: Proceedings Borlaug Global Rust Initiative, 2013 Technical Workshop. New Delhi, India
Sohail Y, Bansal U, Bariana H, Chhuneja P, Mumtaz A, Rattu A, Trethowan R (2014) Identification of an Lr28-linked co-dominant molecular marker in wheat (Triticum aestivum L.). Aust J Crop Sci 8:1210–1215. https://doi.org/10.13140/2.1.4628.5444
Team RC (2013) R: a language and environment for statistical computing. Vienna, Austria. https://www.R-project.org/
Thind AK, Wicker T, Šimková H, Fossati D, Moullet O, Brabant C, Vrána J, Doležel J, Krattinger SG (2017) Rapid cloning of genes in hexaploid wheat using cultivar-specific long-range chromosome assembly. Nat Biotechnol 35:793–796. https://doi.org/10.1038/nbt.3877
Uauy C, Brevis JC, Chen X, Khan I, Jackson L, Chicaiza O, Distelfeld A, Fahima T, Dubcovsky J (2005) High-temperature adult-plant (HTAP) stripe rust resistance gene Yr36 from Triticum turgidum ssp. dicoccoides is closely linked to the grain protein content locus Gpc-B1. Theor Appl Genet 112:97–105. https://doi.org/10.1007/s00122-005-0109-x
Wang J, Zhang Z (2021) GAPIT version 3: boosting power and accuracy for genomic association and prediction. Genomics Proteom Bioinform 19:629–640. https://doi.org/10.1016/j.gpb.2021.08.005
Wang S, Wong D, Forrest K, Allen A, Chao S, Huang BE, Maccaferri M, Salvi S, Milner SG, Cattivelli L et al (2014) Characterization of polyploid wheat genomic diversity using a high-density 90,000 single nucleotide polymorphism array. Plant Biotechnol J 12:787–796. https://doi.org/10.1111/pbi.12183
Wei T, Simko V (2021) R package “corrplot”: visualization of a correlation matrix. https://github.com/taiyun/corrplot
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer, New York
Williams ND, Gough FJ (1965) Inheritance of stem rust reaction in a khapli Emmer cross 1. Crop Sci 5:145–147. https://doi.org/10.2135/cropsci1965.0011183x000500020013x
Xu X, Kolmer J, Li G, Tan C, Carver BF, Bian R, Bernardo A, Bai G (2022) Identification and characterization of the novel leaf rust resistance gene Lr81 in wheat. Theort Appl Genet 135:2725–2734. https://doi.org/10.1007/s00122-022-04145-5
Yin L, Zhang H, Tang Z, Xu J, Yin D, Zhang Z, Yuan X, Zhu M, Zhao S, Li X, Liu X (2021) rMVP: a memory-efficient, visualization-enhanced, and parallel-accelerated tool for genome-wide association study. Genom Proteom Bioinforma 19:619–628. https://doi.org/10.1016/j.gpb.2020.10.007
Yu L-X, Morgounov A, Wanyera R, Keser M, Singh SK, Sorrells M (2012) Identification of Ug99 stem rust resistance loci in winter wheat germplasm using genome-wide association analysis. Theort Appl Genet 125:749–758. https://doi.org/10.1007/s00122-012-1867-x
Zaharieva M, Ayana NG, Hakimi AA, Misra SC, Monneveux P (2010) Cultivated emmer wheat (Triticum dicoccon Schrank), an old crop with promising future: a review. Genet Resour Crop Evol 57:937–962. https://doi.org/10.1007/s10722-010-9572-6
Zetzsche H, Serfling A, Ordon F (2019) Breeding progress in seedling resistance against various races of stripe and leaf rust in European bread wheat. Crop Breed Genet Genom 1:e190021. https://doi.org/10.20900/cbgg20190021
Zhang Z, Ersoz E, Lai CQ, Todhunter RJ, Tiwari HK, Gore MA, Bradbury PJ, Yu J, Arnett DK, Ordovas JM, Buckler ES (2010) Mixed linear model approach adapted for genome-wide association studies. Nat Genet 42:355–360. https://doi.org/10.1038/ng.546
Zhang D, Bowden RL, Yu J, Carver BF, Bai G (2014) Association analysis of stem rust resistance in US winter wheat. PLoS One 9(7):e103747. https://doi.org/10.1371/journal.pone.0103747
Zhang P, Yan X, Gebrewahid T-W, Zhou Y, Yang E, Xia X, He Z, Li Z, Liu D (2021) Genome-wide association mapping of leaf rust and stripe rust resistance in wheat accessions using the 90K SNP array. Theort Appl Genet 134:1233–1251. https://doi.org/10.1007/s00122-021-03769-3
Funding
We thank the Shiaoman Chao and Mary Osenga at USDA-ARS Small Grains Genotyping Laboratory at Fargo, ND for genotyping the emmer wheat panel using the wheat 9 K SNP iSelect assay. This research was supported in part by the USDA-ARS CRIS Project No. 3060-21000-038-00D and 2030-21430-014-00D, Non-Assistance Cooperative Agreement No. 58-3060-9-031, and an appointment to the Agricultural Research Service (ARS) Research Participation Program administered by the Oak Ridge Institute for Science and Education (ORISE) through an interagency agreement between the United States Department of Energy (DOE) and the United States Department of Agriculture (USDA). ORISE is managed by Oak Ridge Associated Universities (ORAU) under DOE contract number DE-SC0014664. QS was partially supported by China Scholarship Council. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture.
All opinions expressed in this paper are the author's and do not necessarily reflect the policies and views of USDA, ARS, USDA is an equal opportunity provider and employer.
Author information
Authors and Affiliations
Contributions
SX and MA conceived and initiated the project. QZ prepared and increased seed stocks of the emmer panel. QS and MA performed the phenotype analysis and XL and JF (Fiedler) genotyped the emmer panel with the GBS method. QS conducted preliminary analysis for dissertation research advised by SX, XC, JF (Faris) and GX and assisted in manuscript preparation. DL performed all the data and bioinformatic analyses and wrote the manuscript assisted with SX and YG. UG and MA performed gene postulations. All authors reviewed and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interest
The authors declare no competing interest.
Code availability
Not applicable.
Additional information
Communicated by Reem Aboukhaddour.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lhamo, D., Sun, Q., Zhang, Q. et al. Genome-wide association analyses of leaf rust resistance in cultivated emmer wheat. Theor Appl Genet 136, 20 (2023). https://doi.org/10.1007/s00122-023-04281-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-023-04281-6