Abstract
Key message
A FT/TFL1 subfamily gene, rice CENTRORADIALIS 2, also known as RCN1, regulates seed germination and increase salt tolerance via ABA-mediated pathway. The ABA synthesis and metabolism related genes were changed relative expression levels.
Abstract
Seed germination is a complex biological process that is affected by many factors. Although a number of germination-related genes have been reported, the molecular mechanism of germination regulation has not yet been fully elucidated. Here, we reported that the rice OsCEN2 gene can negatively regulate seed germination. The germination speed of OsCEN2-RNAi seeds was significantly faster while that of OsCEN2-overexpression (OE) seeds was slower than that of the wild type (WT). The results of qRT-PCR showed that the OsCEN2 expression was increased in the early stage of seed germination. Exogenous application of abscisic acid (ABA) on seeds and seedlings showed that OsCEN2-OE seeds and seedlings were highly sensitive to ABA during germination and post-germination growth, respectively. The determination of endogenous ABA content in seeds also showed that the ABA content of OsCEN2-RNAi seeds was lower, while that of OsCEN2-OE seeds was higher. Moreover, the transgenic plants changed salt tolerance because of the altered ABA level. In addition, differences were also observed in the expression of genes related to ABA synthesis and metabolism in the seeds of OsCEN2-transgenic lines. This study reveals that OsCEN2 regulates the germination speed by affecting the content of ABA during seed germination and provides a theoretical basis for research on rice direct seeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
During seed germination, vigorous seeds absorb water under suitable environmental conditions, expand their volume, release dormancy, until the embryo breaks through the seed coat, which is key to the life cycle of all flowering plants that propagate through seeds (Nonogaki et al. 2010). The difference in germination speed of crop seeds affects the uniformity of emergence and the regularity of seedlings, which in turn affects crop yields (Gubler et al. 2005). In rice, seed germination speed and regularity also affect mechanical direct seeding, leading to problems with sparse seedling of direct seeding rice, as well as low or unstable yields (Mahender et al. 2015).
In the process of seed germination, environmental factors play a vital role. In addition, endogenous factors such as seed developmental stage, hormone levels and seed structure are also of importance. Studies have indicated that gibberellin (GA) and abscisic acid (ABA) are mutually antagonistic during seed dormancy and germination; ABA can inhibit while GA can promote seed germination (Corbineau et al. 2014). Seeds can maintain the dormancy state through accumulating ABA during development, avoiding preharvest sprouting. During seed germination, the ABA content would gradually decrease, and exogenous application of ABA would inhibit seed germination to a certain extent (Gubler et al. 2005; Jiang et al. 2019; Song et al. 2020).
ABA signal transduction is mainly composed of three core components, including the ABA receptor (REGULATORY COMPONENT OF ABA RECEPTOR 1/PYRABACTIN RESISTANCE 1/PYR1-RELATED HOMOLOGUES, RCAR/PYR1/PYL), protein phosphatase (PROTEIN PHOSPHATASE 2C, PP2C) and SNF1-related kinase 2 (SNF1- RELATED PROTEIN KINASE 2, SnRK2) (Miyazono et al. 2009). Under normal growth conditions, ABA is sensed by receptors such as RCAR/PYR1/PYL, which interacts with the free-state PP2C (with protein phosphatase activity) and forms a complex, PYR/RCAR-PP2C, to inhibit PP2C activity. Afterward, SnRK2 is phosphorylated to activate downstream transcription factors responding to ABA signals (Finkelstein 2013; Koramutla et al. 2021). ABA INSENSITIVE 5 (ABI5) is a major transcription factor in the ABA signaling pathway. In rice, the abi5 mutant was not sensitive to ABA during germination, but plants overexpressing ABI5 were highly sensitive to ABA (Zou et al. 2008). ABI5 can also bind to the ABA response element (ABRE) on the promoter of its target genes to promote or inhibit their expression. In Arabidopsis, ABI5 can bind to the promoters of EARLY METHIONINE-LABELED 1 (EM1) and EM6 to regulate seed germination (Carles et al. 2002). ABI3/OsVP1 is mainly involved in signal transduction during plant seed development and thus affects seed germination. Rice OsVP1 can interact with Rc and another regulatory factor, OsC1, to enhance the sensitivity of seeds to ABA by promoting the biosynthesis of proanthocyanidins and the perception of ABA signals, ultimately inhibiting early germination (Wang et al. 2020).
OsCEN2 belongs to the FLOWERING LOCUS T (FT)/TERMINAL FLOWER 1 (TFL1) subfamily within the phosphatidylethanolamine binding protein (PEBP) family (Coelho et al. 2014). In addition to playing a role in regulating flowering time and plant structure, FT/TFL1 is also associated with plant seed germination (Karlgrenet et al. 2011; Jin et al. 2021). A study has revealed that the Arabidopsis FT gene can inhibit the expression of the FLOWERING LOCUS C (FLC) gene by activating the RNA-COOLAIR of FLC and then release seed dormancy by regulating the chromatin state (Chen et al. 2018). However, another study reported that FT inhibits the FLC expression through a 5′ promoter region of FLC (Luo et al. 2019). The potato (Solanum tuberosum) FT/TFL1 family gene StCEN plays a regulatory role in the process of potato tuberization. The expression level of StCEN is related to the growth rate of buds, which is lower in StCEN overexpression lines compared to the wild type (Morris et al. 2019). In poplars (Populus spp.), reducing the expression of PopCEN1 shortens the dormancy time and accelerates the flowering time, while the trees overexpressing PopCEN1 show the opposite phenotype and the flowering of which is completely inhibited (Mohamed et al. 2010). The PcFT2 gene of pear can promote plant vegetative growth, and delay seed dormancy and leaf senescence (Freiman et al. 2015). In Arabidopsis, AtMFT regulates seed germination through ABA and GA signaling pathways (Xi et al. 2010). Many studies have shown that the loss-of-function mft mutant exhibits sensitivity to ABA during seed germination. In the process of seed germination, the expression of MFT is directly regulated by two key transcription factors, ABI3 and ABI5, in the ABA signaling pathway. At the same time, MFT directly inhibits ABI5, negatively regulating ABA signaling to promote embryo growth (Xi et al. 2010). In wheat (Triticum aestivum), the increased TaMFT expression is related to low germination index. Transient expression of TaMFT in immature embryos can inhibit the premature germination of immature embryos, indicating that TaMFT is an inhibitor of seed germination (Nakamura et al. 2011). In rice, OsMFT2-knockout lines show preharvest sprouting, while OsMFT2-overexpression lines exhibit delayed sprouting. It is revealed that OsbZIP23, OsbZIP66 and OsbZIP72 can interact with OsMFT2; in addition, OsbZIP23/66/72 can bind to the promoter of the ABA response element gene Rab16A. Such binding is enhanced by OsMFT2, indicating that OsMFT2 negatively regulates seed germination (Song et al. 2020). The functions of multiple FT/TFL1 family genes have been reported; however, whether OsCEN2 can affect rice seed germination remains elucidated.
In this study, a rice CENTRORADIALIS gene OsCEN2, also known as RCN1, is responsible for regulating seed germination in rice. We found that OsCEN2-RNAi seeds showed increased germination speed, while OsCEN2-overexpression (OE) seeds showed delayed germination. Exogenous application of ABA to the seeds and seedlings revealed that OsCEN2 affected seed germination and the sensitivity of seedlings to ABA. In summary, OsCEN2 affects the ABA content during seed germination by affecting the expression of genes related to ABA synthesis and metabolism, which in turn affects the seed germination speed.
Materials and methods
Plant material and growth conditions
Zhonghua 11 (ZH11, O. sativa ssp. Japonica) used as WT and recipient for genetic transformation in this study. The plants were grown in the paddy field at South China Agricultural University in Guangzhou. Rice cultivation is managed according to local common practices.
To construct the OsCEN2 overexpression vector, 522-bp fragment of the OsCEN2 cDNA sequence driven by the Ubiquitin (Ubi) promoter was inserted into the pYLox vector (Yu et al. 2010).
To construct the OsCEN2 RNAi vector, two fragments of OsCEN2 were isolated by PCR using the primers RNAi1 and RNAi2 from the rice ZH11 genome DNA into the EcoR I–Hind III sites and from immature panicle cDNA with primers RNAi2 and RNAi3 into the Hind III–Sal I sites of the cloning vector of pBluescript II. The RNAi construct was driven by the native promoter of OsCEN2. The OsCEN2 promoter was amplified from the rice genome using the primers PRNAiF and PRNAiR and cloned into the BamH I–EcoR I sites of the cloning vector pBluescript II with RNAi fragments. After sequencing, the total fragments containing the native promoter and RNAi fragments were digested with BamH I and Sal I and cloned into the binary vector pCAMBIA1380. The cloning was performed following the previously described methods (Li et al. 2011; Zhou et al. 2014).
To generate the CRISPR/Cas9 vector, the CEN2Cas were constructed using pYLgRNA-OsU3, as described previously (Ma et al. 2015). The sequence of the target site was 5′- TGTGTCTAAACCAAGAGTTG-3′, which contained a protospacer adjacent motif (PAM) AGG at the 3′ end.
All constructs were confirmed by sequencing, introduced into Agrobacterium tumefaciens EHA105 cells and transformed into ZH11 by Agrobacterium-mediated transformation according our previously reported method (Jiang et al. 2018). All primers used for vector construction are listed in Table S1.
RNA isolation and qRT-PCR
Total RNA was isolated from the seed using fruit-mate (TaKaRa, Dalian, China) and RNAiso Plus (TaKaRa, Dalian, China). The seeds were ground into powder (100 mg) in liquid nitrogen, then transferred to 500 μL fruit-mate and mixed thoroughly. After centrifugation, the supernatant fluid was transferred and mixed with 500 μL RNAiso Plus reagent and mixed thoroughly and incubated for 5 min. Then 200 μL chloroform was added and mixed thoroughly. After centrifugation, the aqueous phase was transferred and mixed with an equal volume of isopropanol. After gently inversions, the mixture was chilled at −20 °C for 20 min and then centrifuged. The supernatant fluid should be removed and washed using 75% ethanol with diethyl pyrocarbonate-treated ddH2O (DEPC-ddH2O). The precipitate is dissolved with DEPC-ddH2O and quantified with a DU730 spectrophotometer (Beckman Coulter, Germany).
cDNA was synthesized with a 5 × iScript RT Supermix kit (TaKaRa, Dalian, China) using about 2 μg of total RNA.
qRT-PCR was performed with SYBR premix Ex Taq II (TaKaRa, Dalian, China) in a total volume of 20 μL on the Bio-Rad CFX 96 following the manufacturer’s protocol. Data were normalized to the internal rice UBIQUITIN (UBI) gene, and relative quantification was used for data analysis. All primers for qRT-PCR are shown in Supporting Information Table S1.
Measurement of ABA level
Seeds freshly harvested were dehulled and soaked in distilled water at 28 °C. Seeds (100 mg) at 0, 6 and 12 HAI (hours after imbibition) were ground into fine powder in liquid nitrogen. They were used for endogenous ABA measurement by plant ABA ELISA Kit (Kejing Biological Technology Co. Ltd.) following the manufacturer’s protocol.
Seed germination assay and ABA or FLU treatment
To treat with ABA or Fluridone (FLU), the fresh harvest grains were soaked with distilled water, 5 μM ABA or 100 μmol/L FLU solution for 24 h. A total of 180 seeds were spread onto plates covered with wet filter papers for three biological replicates of each line. The plates were placed in a chamber at 28 °C.
For counting the germination rates of grains, 180 seeds for each line were soaked in distilled water for 24 h and then spread onto plates covered with wet filter papers for three biological replicates.
To medium culture with water and agar in plate, 60 husked seeds for each line were sterilized with 1.5% NaClO for 20 min and washed with sterilized water five times. The seeds were spread onto the medium for three biological replicates of each line.
Germination was defined as the emergence of the radical, and the number of germinated seeds was counted every 3 or 6 h. The germination rate was calculated as the number of total germinated seeds divided by the number of total seeds. The photographs were taken by camera and the length of roots and shoots were measured by calipers and ruler.
To treat with ABA, the 10 d-seedlings were cultured with hydroponics with or without 5 μM ABA solution. To determine the ABA sensitivity of post-gemination growth, 45 husked seeds of each line were sterilized with 1.5% NaClO and washed with sterilized water for three times. The seeds were spread onto 1/2 MS medium. We conducted the germination assay every 12 h to select the seeds with the same growth vigor and state. Then, they were transferred onto 1/2 MS medium containing a gradient concentration of ABA (0, 1, 3, 5, 10 μM). For every treatment, three biological replicates were conducted. The seeds were grown in the chamber at 28 ℃. The length of roots and shoots was measured at 10 days after transplanting.
RNA-seq and data analysis
Total RNA was extracted from seeds in triplicate at 0, 6 and 12 HAI (hour after imbibition) using fruit-mate and RNAiso Plus (TaKaRa, Dalian, China), following to the protocol. RNA was used for RNA-seq by BGI Genomics (Wuhan, China), using the DNBSEQ platform.
Salt treatments
WT and transgenic homozygous seeds were germinated in water and transferred to Kimura B nutrient solution after germination 4 days. For salt stress conditions, 2-week-old seedlings were treated with Kimura B complete nutrient solution supplemented with 150 mM sodium chloride. After stress treatments for 6 days, seedlings were transplanted to Kimura B nutrient solution for 6 days.
Relative water content
The relative water content measurement was performed according to the reported methods (Khan et al. 2017). Three mature leaves at 0 and 3rd day of salt stress were harvested, and the fresh weight (FW) was recorded. These leaf samples were kept 2 h in water to attain turgidity and measured turgid weight (TW). These leaves were kept in oven at 60 °C for 24 h and measured the dry weight (DW). The relative water content was calculated as the following formula, RWC (%) = [(FW − DW) / (TW − DW)] × 100%
The electrolyte leakage
The electrolyte leakage measurement was performed following the reported methods (Lv et al. 2017). In brief, three leaves from three plants at 0 and 6th day of salt stress were cut into segments of the same size and immersed in 20 mL of double distilled water in a 250-mL test tube for 6 h with shaking. The initial conductivity (R1) was measured with a conductivity meter (Model DDS-11A, Shanghai Hongyi Instruments and Apparatuses Co. Ltd.). Then, the test tubes with leaf segments were placed in boiling water for 30 min and cooled naturally to room temperature, and the conductivity (R2) was determined. The relative electrolyte leakage was calculated as the ratio of R1 to R2.
Data analysis
To test statistically significant differences in the results, the one-way ANOVA (LSD analysis) test was conducted pairwise comparisons between transgenic line and wild type. These tests were performed using SPSS version 20. We judged the data significance level according to the P value. In all statistical tests, P value < 0.05 was considered statistically significant. *: P < 0.05, **: P < 0.01, ***: P < 0.001.
Accession numbers
Sequence data from this article can be found in the Rice Annotation Project database (RAP-DB), under the following accession numbers: OsCEN2 (Os11g0152500), OsNCED3 (Os03g0645900), OsNCED5 (Os12g0617400), OsZEP1 (Os04g0448900), ABA8ox1 (Os02g0703600).
Results
OsCEN2 affects seed germination speed
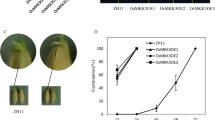
To study the function of OsCEN2 during seed germination, we obtained OsCEN2 overexpression (OsCEN2-OE) plants and RNAi (OsCEN2-RNAi) plants of rice cultivar ‘Zhonghua 11’. Seed germination experiments showed that the germination speed of OsCEN2 transgenic seeds was significantly different from that of wild type (WT). The OsCEN2-RNAi plants (RNAi12 and RNAi2) germinated significantly faster while the OsCEN2-OE plants (OE1 and OE2) germinated slower compared to WT at 48 h after imbibition (HAI; Fig. 1a). To further explore the effect of OsCEN2 on seed germination, we calculated the germination speed of these seeds. At 24 HAI, the WT and OsCEN2-RNAi seeds began to germinate, while the OsCEN2-OE seeds did not. At 36 HAI, the OE1 seeds began to germinate, while the OE2 seeds begin to germinate until 45 HAI. The germination rate of WT seeds had reached 18.9%, while the germination rates of the two OsCEN2-RNAi lines were 29.8% and 28.9%, respectively, at 45 HAI. The germination rates of RNAi lines were 57.05%, WT was 38.01%, while those of OE1 and OE2 were 23.71% and 15.08% at 57 HAI. The germination rate of OsCEN2-RNAi and WT seeds exceeded 90% at 99 HAI, while those of OE1 and OE2 were 78.74 and 83.34%, respectively (Fig. 1b). In addition, we conducted germination experiments under sterile conditions. The germination speed of OsCEN2-RNAi seeds was faster than that of WT, while that of OsCEN2-OE lines was the slowest (Fig. S1), which is consistent with the trend of germination speed of seeds with husks (Fig. 1b). The germination experiments conducted on OsCEN2-knockout seeds also showed that the germination speed of transgenic seeds was significantly faster than that of WT seeds (Fig. S2a, c). This is consistent with the results obtained from OsCEN2-RNAi seeds.
OsCEN2 negatively regulates seed germination. a Phenotype of WT, overexpression and RNAi lines at 48 h after imbibition. Scale bars: 1 cm. b Grains germination rate of WT, overexpression and RNAi lines. The grains were soaked in distilled water and the germination grains were counted every 3 h after 24 h imbibition. Values are presented as means ± SD of three biological replicates (n = 60). c Seedling phenotypes of WT, overexpression and RNAi lines. Scale bar: 10 cm. d The plant height of WT, overexpression and RNAi lines. Values are presented as means ± SD (n = 30), **P < 0.01; one-way ANOVA (LSD analysis) test. WT: wild type Zhonghua 11; OE 1, OE 2: overexpression lines; RNAi 12, RNAi 2: RNAi lines
At 23 d post-germination, a significant difference was observed in plant height between WT and transgenic seedlings (Fig. 1c). The plant height of the two OsCEN2-RNAi lines was 9.53 and 9.96 cm, respectively, which were significantly higher than that of 8.34 cm of WT. On the contrary, the plant height of the two OsCEN2-OE lines was 5.85 and 6.34 cm, respectively, which were significantly lower than that of WT (Fig. 1d). Moreover, the OsCEN2-knockout seedlings were also significantly higher than WT (Fig. S2b, d).
Expression pattern of OsCEN2
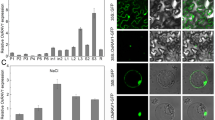
We used qRT-PCR to investigate the expression level of OsCEN2 during seed germination. The results showed that OsCEN2 was expressed throughout the germination process of rice seeds (Fig. 2a). With seed germination proceeded, the expression level of OsCEN2 was gradually increased and reached the peak at 18 HAI. Then, the expression level of OsCEN2 began to decline sharply at 24 HAI and remained a low level at 30–48 HAI (Fig. 2a).
Expression pattern analysis of OsCEN2. a Expression levels of OsCEN2 at different time during seed germination. b Expression levels of OsCEN2 at 3, 6 and 12 h time points of germinated seeds after 5 μM ABA solution treatment. c Expression levels of OsCEN2 at different time of 10 d-seedling after 5 μM ABA treatment. Values are presented as means ± SD (n = 3)
ABA is a key hormone that regulates seed germination. To study the effect of ABA on OsCEN2, we examined the OsCEN2 expression in germinated seeds after 5 μM ABA treatment. The results of qRT-PCR showed that the expression level of OsCEN2 increased significantly after 12 h of ABA treatment (Fig. 2b). In addition, we applied ABA on 10-day-old WT rice seedlings, and the expression level of OsCEN2 increased with the increase in treatment duration, reached the highest level at 24 h and then began to decline (Fig. 2c). The above results indicate that exogenous ABA application can induce the expression of OsCEN2.
OsCEN2 affects rice sensitivity to ABA
To study the effect of exogenous ABA application on seed germination, we treated the seeds with water and 5 μM ABA solution, respectively. The results showed that after ABA treatment, the seed germination speed exhibited delay (Fig. 3a, S3b). Under ABA treatment conditions, the RNAi12 seeds showed a relative lower sensitivity compared with WT seeds. While the OE2 seeds showed a higher sensitivity compared to WT seeds (Fig. 3a). For example, at time point 112 HAI, the germination rate of WT seeds decreased from 68.3% to 51.7% after ABA treatment, while the RNAi12 seeds decreased from 78.9% to 76.7%, the OE1 seeds decreased from 27.2% to 5.6% (Fig. 3a). The above results suggest that ABA has a greater impact on the germination speed of OsCEN2 OE seeds compared to WT and RNAi seeds. As for the final germination rate, WT and OsCEN2-RNAi seeds changed little, while that of OsCEN2-OE seeds decreased by 21.1% after ABA treatment (Fig. 3a), suggesting that the OsCEN2-OE seeds are more sensitive to ABA.
ABA treatment influences seed germination and seedling growth. a Dynamic seed germination rate of WT, OE 1 and RNAi 12. The grains were soaked in distilled water, with or without 5 μM ABA, which were harvested at the same season. The germination rate was counted every 6 h after 48 h-imbibition. Values are presented as means ± SD of three biological replicates (n = 60). b Phenotypes of WT, OE 1, OE 2, RNAi 12 and RNAi 2 of germinated seeds under different ABA level treatments for 10 days. Scale bar: 10 cm. c, d The plant height (c) and root length (d) of WT, OE 1, OE 2, RNAi 12 and RNAi 2 of germinated seeds under different ABA level treatments for 10 days in b. Values are presented as means ± SD (n = 30). *: P < 0.05; **: P < 0.01; one-way ANOVA (LSD analysis) test. WT: wild type Zhonghua 11; OE 1, OE 2: overexpression lines; RNAi 12, RNAi 2: RNAi lines
We further observed the effect of exogenous ABA application on the post-germination growth of seedlings. In this study, seeds were treated with water and ABA solution with different concentrations, respectively. The results showed that the decrease in plant height of OsCEN2-OE seedlings was significantly greater than that of WT and OsCEN2-RNAi seedlings (Fig. 3b, c), and the root length also showed similar patterns (Fig. 3b, d). These results indicate that in the process of post-germination growth, OsCEN2-RNAi seedlings are less sensitive, while OsCEN2-OE seedlings are more sensitive to ABA compared with WT.
Changes in endogenous ABA content of transgenic rice seeds lead to varied seed germination speeds
ABA in plants is mainly synthesized through the carotenoid pathway. Fluridone (FLU) can inhibit the biosynthesis of carotenoids, thereby affecting the ABA content in plants. After germination, the growth of OsCEN2-OE seedlings was significantly slower than WT seedlings. However, no significant difference was observed between OsCEN2-OE and WT seedlings after OE seeds treatment with 100 μmol/L FLU (Fig. 4a). At 8 DAI, there was no obvious difference between FLU-treated OsCEN2-OE and WT seedlings from phenotype (Fig. 4a). The plant height was consistent among them too (Fig. 4b). However, significant difference was found in the growth rate between the OsCEN2-OE seedlings without FLU treatment and WT (Fig. 4a, b), indicating that the inhibition of endogenous ABA synthesis can accelerate OsCEN2-OE seed germination.
Effects of FLU on rice seed germination and seedling growth. a Germinated seeds phenotypes with water or FLU treated for 5 and 8 days separately. Scale bars: 1 cm. b The seedling height and root length of treatment in a for 8 days. Values are presented as means ± SD (n = 30), **P < 0.01; the one-way ANOVA (LSD analysis) test. WT: wild-type Zhonghua 11; OE 1, OE 2: overexpression lines. FLU: fluridone
To further explore the reasons for the difference in seed germination, we examined the ABA content in the seeds. Before water soaking, the ABA content in OsCEN2-RNAi seeds was significantly lower than that in WT, while the ABA content in OsCEN2-OE seeds was the highest. At 6 HAI, the ABA content in all these lines were decreased compared to before imbibition, respectively. At this time, the ABA content in OsCEN2-OE seeds was still significantly higher than that of WT and OsCEN2-RNAi seeds (Fig. 5a). At 12 HAI, the ABA content of OsCEN2-OE seeds kept decreasing compared to at 6 HAI. However, the ABA content in OsCEN2-OE seeds was the highest among WT, overexpression and RNAi lines (Fig. 5a). The above results indicate that OsCEN2 affects the ABA content in seeds, and we speculate that the change in ABA content is the main factor affecting seed germination.
ABA levels and relative expression levels of ABA relative genes during seed germination. a ABA content in the seeds after imbibition in water for 0, 6 and 12 h. Values are presented as means ± SD ( n = 3), *: P < 0.05, **P < 0.01; the one-way ANOVA (LSD analysis) test. (b-c) Relative expression levels analysis of ABA relative genes in seeds before imbibition (b) and after imbibition for 12 h (c). WT: wild-type Zhonghua 11; OE 1, OE 2: overexpression lines; RNAi 12, RNAi 2: RNAi lines
Changes in the expression level of genes related to ABA synthesis and metabolism
To study the reasons for the changes in ABA content in seeds, we investigated the expression levels of ABA-related genes. The expression levels of the ABA synthesis-related genes OsNCED3, OsNCED5 and OsZEP1 were higher in OsCEN2-OE seeds, while lower in OsCEN2-RNAi seeds compared to WT before water soaking. The ABA degradation gene ABA8ox3 was expressed at a low level in OsCEN2-OE seeds, but it was highly expressed in OsCEN2-RNAi seeds (Fig. 5b). At 12 HAI, the expression levels of OsNCED3 and OsZEP1 in OsCEN2-OE seeds were decreased and that of ABA8ox3 was also lower compared to WT. The expression levels of OsNCED5 and ABA8ox3 in OsCEN2-RNAi seeds were higher than those of WT (Fig. 5c). These results indicate that OsCEN2 can regulate the expression level of ABA-related genes, resulting in the change of ABA content.
Changes in the resistance of OsCEN2-transgenic plants to salt stress
The change of ABA content in plants affects their resistance to abiotic stress, such as drought and salt conditions. To confirm the effect of changes in ABA content on the resistance of OsCEN2-transgenic plants, the rice seedlings were treated with 150 mM NaCl solution. The results showed that the phenotype of OsCEN2-OE plants was no obvious different from WT before NaCl treatment, and no visible difference too in the phenotype after NaCl treatment (Fig. S4a-c). Although the relative electrolyte leakage (EL) also showed no significant difference between the OsCEN2-OE and WT plants before NaCl treatment, after salt treatment the EL value of WT was 22.91%, while that of the two OsCEN2-OE lines was 6.47% and 6.33%, respectively (Fig. S4d), indicating that OsCEN2-OE lines are more resistant to salt stress than WT. In addition, the OsCEN2-knockout plants all died at 7 d of recovery after NaCl treatment, while most of WT plants resumed growth (Fig. 6a–c). The OsCEN2-knockout plants had significantly higher relative EL than WT, and they also exhibited a significantly lower relative water content compared to WT (Fig. 6d, e). This indicated that the knockout plants are more sensitive to salt tolerance than WT. The above results showed that OsCEN2 expression level leads to change tolerance of rice to salt treatment. These results suggest OsCEN2 maybe regulate ABA content and then influence the seedling resistance to salt stress.
OsCEN2 knockout lines decreases salt stress tolerance. a–c Gross morphology of OsCEN2 knockout lines and WT rice plants before treatment (a), after 150 mM NaCl treatment for 6 days (b) and recovered for 6 days c are shown. d–e Electrolyte leakage (d) and relative water content (e) of WT, Cas9-2 and Cas9-7 lines before and after salt treatment for 6 days and 3 days. Values are presented as means ± SD (n = 9), *: P < 0.05; one-way ANOVA (LSD analysis) test. WT: wild-type Zhonghua 11; Cas9-2, Cas9-7: OsCEN2 knockout lines
OsCEN2 regulates plant hormone and MAPK signaling pathways during seed germination
Transcriptome (RNA-seq) helps to analyze the molecular regulation pathways of genes. We collected transcriptome data of rice seeds at 6, 12 and 24 HAI to investigate the changes in rice seeds responding to OsCEN2 expression. Venn diagram analysis showed that a total of 1,104 differentially expressed genes (DEGs) were found at 12 HAI, while at 6 and 24 HAI, only 279 and 181 DEGs were identified (Fig. 7a); 58 and 97 DEGs were presented in OE and RNAi lines compared with WT at all the three time points, respectively (Fig. 7b). Gene ontology (GO) analysis showed that DEGs were enriched in starch and sugar metabolism, MAPK signaling pathway and glycolysis pathway at 6 HAI (Fig. 7c); DEGs were enriched in MAPK signaling pathway, plant hormone signal transduction, fatty acid metabolism and amino acid synthesis pathway at 12 HAI (Fig. 7d); and DEGs were enriched in phenylpropanoid biosynthesis, MAPK signaling pathway and amino sugar and nucleotide sugar metabolism pathway at 24 HAI (Fig. 7e).
DEGs at the 6, 12 and 24 h of seed imbibition are involved in plant hormone signal transduction and MAPK signal pathway. a Venn diagram showing DEGs overlap between OE lines (green plate) and RNAi lines (orange plate) relative to WT at the 6, 12 and 24 h after seed imbibition. b Venn diagram showing DEGs which expression levels were different from WT in OsCEN2 overexpression lines or RNAi lines overlap at 6 h (orange plate), 12 h (green plate) and 24 h (pink plate) after seed imbibition. c–e Gene ontology enrichment pathways predicted to be involved in seed germination at the 6 h (c), 12 h (d) and 24 h (e) after seed imbibition. WT: wild-type Zhonghua 11; OE: overexpression plants; RNAi: RNAi plants
Among these DEGs, the expression trends of 73 DEGs were opposite between OsCEN2-OE and OsCEN2-RNAi lines compared with WT at 6 HAI, which were involved in plant hormone signaling, MAPK signaling pathway and sugar metabolism (Fig. S5a, S6). At 12 HAI, 81 DEGs that were involved in plant hormone signaling and starch and sugar metabolism showed opposite expression trends between OsCEN2-OE and OsCEN2-RNAi lines (Fig. S5b, S7). At 24 HAI, 92 DEGs that participated in plant hormone signaling and ABC transport exhibited different expression patterns between OsCEN2-OE and OsCEN2-RNAi lines (Fig. S5c, S8). The above results indicate that OsCEN2 may regulate plant hormone signal transduction, sugar metabolism and MAPK signal pathways to regulate seed germination in the early stage of germination.
ABA and GA play different key roles in the regulation of seed dormancy and germination, and the metabolism and signaling by both phytohormones also changes during seed development (Shu et al. 2015). To illustrate the changes of ABA- and GA-related genes during germination, we analyzed the expression in transcriptome data and performed the heat map analysis of ABA and GA-related genes at 6, 12 and 24 h after imbibition. The results showed the expression levels of several genes showed in opposite direction in OsCEN2 overexpression and RNAi lines (Fig. S9). At 6 HAI, the expression level of OsNF-YC5 (LOC112936059) and ABA8ox1 (LOC9267503) decreased in overexpression plants, while increased in RNAi plants. At 12 HAI, OsNF-YC5 (LOC112936059) and AAA-ATPase ASD (LOC4352916) decreased in overexpression plants, while increased in RNAi plants. In addition, ethylene-responsive transcription factor ERF110 (LOC4351606), PP2C37 (LOC9268838) and zinc finger protein 2 (LOC4347837) were decreased in overexpression plants, while increased in RNAi plants (Fig. S9). These results suggested that OsCEN2 affected the expression levels of ABA-related genes during germination.
We pay attention to the genes relative to GA, a heat map analysis was conducted at different timepoints. At 6 HAI, the expression level of OsCPS2 (LOC9266189) was increased in overexpression plants, while decreased in RNAi plants. In contrary, myb-related protein 308 (LOC4349938) was decreased in overexpression plants while increased in RNAi plants. Senescence-specific cysteine protease SAG39 (LOC4335170) increased in overexpression plants while decreased in RNAi plants at 12 HAI. In addition, gibberellin 2-beta-dioxygenase 6-like, GA2ox6 (LOC9266251) increased in overexpression plants while decreased in RNAi plants at 24 HAI. However, cytochrome P450 714D1-like (LOC4339131) and OsGA20ox2 (LOC4325003) were decreased in overexpression plants while increased in RNAi plants. These genes show the opposite trend in overexpression plants and RNAi plants (Fig. S10). These results suggested that OsCEN2 affected the expression levels of GA-related genes during germination.
Discussion
OsCEN2 belongs to the FT/TFL1 subfamily within the PEBP gene family, the members of which have been extensively studied (Coelho et al. 2014). Their functions involve dormancy (Chen et al. 2018), leaf senescence (Freiman et al. 2015) and flowering (Nakagawa et al. 2002). In this study, we found that OsCEN2 could affect the germination of rice seeds. The germination of OsCEN2-OE seeds was delayed, while that of OsCEN2-RNAi seeds was accelerated, which is consistent with the results of other studies. For example, the PcFT2 gene of pear can delay seed dormancy and leaf senescence (Freiman et al. 2015); Arabidopsis AtMFT regulates seed germination through ABA and GA signaling pathways (Xi et al. 2010); and OsMFT2 negatively regulates seed germination by affecting the sensitivity of seeds to ABA during germination and post-germination growth in rice (Song et al. 2020).
There are many factors that affect the germination of rice seeds, including the inherent genetic characteristics of seeds such as α-amylase activity, endogenous plant hormones and soluble sugars, and external environmental factors such as temperature, humidity and light (An et al. 2018). In this study, the expression of OsCEN2 was induced by ABA in both seeds and seedlings. OsCEN2-OE seeds were more sensitive to ABA during germination, while OsCEN2-RNAi seeds showed lower sensitivity to ABA, which indicates that OsCEN2 plays an important role in ABA signal transduction. ABA and GA play different key roles in the regulation of seed dormancy and germination, and the metabolism and signaling by both phytohormones also changes during seed development (Shu et al. 2015). The effects of these hormones depend on both the amounts of one relative to the other, resulting from the rates of synthesis and catabolism, as well as the sensitivity of the tissues to these hormones (Finkelstein et al. 2008). Studies have shown that the sensitivity of plant seeds to ABA is the key to determine whether seeds can break dormancy and turn to the germination stage (Schmitz et al. 2002). In Arabidopsis, there are several ABA-insensitive (ABI) genes: ABI1 and ABI2 encode PP2C, ABI3 encodes B3 transcription factor, ABI4 encodes APETALA2-like transcription factor, and ABI5 encodes bZIP transcription factor (Finkelstein and Lynch 2000; Merlot et al. 2001; Shu et al. 2013; Feng et al. 2014). These are all key genes in the plant ABA signaling pathway and play an important role in the biological processes of seed germination and dormancy and in response to adverse environments (Piskurewicz et al. 2008; Lopez-Molina et al. 2001, 2002). The abi3, abi4 and abi5 mutants are insensitive to ABA during seed germination and early seedling development and are lack of dormancy (Söderman et al. 2000; Zou et al. 2007; Feng et al. 2014). Studies have found that the sapk2 mutant was ABA-insensitive in both germination and post-germination stages, indicating SAPK2 plays a key role in ABA-mediated seed dormancy. Therefore, the seed germination process is highly associated with the sensitivity of seeds to ABA, which indicates that OsCEN2 may regulate seed germination by affecting the sensitivity of seeds to ABA.
Studies have demonstrated that ABA content can affect seed germination speed (Jiang et al. 2019; Song et al. 2020). In the present study, the slow germination speed and low germination rate of OsCEN2-OE seeds maybe related to the ABA content in seeds. The ABA content in OsCEN2-OE dry seeds was higher than that in WT and OsCEN2-RNAi dry seeds. In addition, the germination rate and germination speed of OsCEN2-OE seeds were reduced more after treatment with exogenous ABA. In plants, ABA 8’-hydroxylation is the main pathway of ABA catabolism. In Arabidopsis, the CYP707A family members CYP707A1-CYP707A4 encode ABA 8’-hydroxylase; the seeds of cyp707a2 mutant are highly dormant due to the high accumulation of ABA (Kushiro et al. 2004). ABI4 can positively regulate the synthesis of ABA in seeds to maintain the dormancy state of seeds (Shu et al. 2013). In this study, the expression levels of ABA synthesis-related genes in OsCEN2-OE dry seeds were higher while in OsCEN2-RNAi seeds were lower than those in WT; however, the expression pattern of the degradation gene ABA8ox3 was the opposite. These results are consistent with those previously reported. Therefore, OsCEN2 affects the endogenous ABA content by affecting the expression of ABA synthesis-related genes, thereby altering the germination rate and germination speed of seeds.
Published literatures reported that ABA modulates plant adaptation to osmotic stress mainly through increasing cellular dehydration tolerance and reducing water loss (Lee et al. 2012). The nine-cis-epoxycarotenoid dehydrogenases (NCEDs) are the key enzymes in ABA biosynthesis (Zhu et al. 2009). Some NCED proteins have been reported to contribute to increasing ABA levels and abiotic stress tolerance in plants (Sun et al. 2012). OsMADS23 was reported to confer drought and salt tolerance by regulating ABA biosynthesis in rice. More importantly, in parallel to osmads23 mutant, osnced2 mutants had reduced ABA accumulation and increased sensitivity to drought and oxidative stress (Li et al. 2021). Previous results showed that the OsNAC2 overexpression plants with higher ABA contents exhibited increased drought and salt tolerance (Jiang et al. 2019). Consistent with these findings, our results also showed that the knockout lines with lower ABA levels are more sensitive to salt tolerance than WT, and the overexpression lines with higher ABA levels are more resistant to drought and salt stress than WT. These results suggest OsCEN2 maybe regulate ABA content and then influence the seedling’s resistance to drought and salt stress.
This study provides a basis for altering the germination rate and speed of rice seeds as well as their resistance to salt stress through regulation of ABA content. It is of great significance for variety breeding suit for mechanical direct seeding with appropriate germination speed and increased salt tolerance.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
An JP, Li R, Qu FJ, You CX, Wang XF, Hao YJ (2018) An apple NAC transcription factor negatively regulates cold tolerance via CBF-dependent pathway. Journal of Plant Physiol 221:74–80. https://doi.org/10.1016/j.jplph.2017.12.009
Carles C, Bies-Etheve N, Aspart L, LeÂon-Kloosterziel KM, Koornneef M, Echeverria M, Delseny M (2002) Regulation of Arabidopsis thaliana Em genes: role of ABI5. Plant J 30:373–383. https://doi.org/10.1038/s41598-021-88874-5
Chen M, Penfield S (2018) Feedback regulation of COOLAIR expression controls seed dormancy and flowering time. Science 360:1014–1017. https://doi.org/10.1126/science.aar7361
Coelho CP, Minow MAA, Chalfun-Junior A, Colasanti J (2014) Putative sugarcane FT/TFL1 genes delay flowering time and alter reproductive architecture in Arabidopsis. Front Plant Sci 5:221. https://doi.org/10.3389/fpls.2014.00221
Corbineau F, Xia Q, Bailly C, El-Maarouf-Bouteau H (2014) Ethylene, a key factor in the regulation of seed dormancy. Front Plant Sci 5:539. https://doi.org/10.3389/fpls.2014.00539
Feng CZ, Chen Y, Wang C, Kong YH, Wu WH, Chen YF (2014) Arabidopsis RAV1 transcription factor, phosphorylated by SnRK2 kinases, regulates the expression of ABI3, ABI4, and ABI5 during seed germination and early seedling development. Plant J 80:654–668. https://doi.org/10.1111/tpj.12670
Finkelstein R (2013) Abscisic acid synthesis and response. Arabidopsis Book 11:e0166. https://doi.org/10.1199/tab.0166
Finkelstein R, Reeves W, Ariizumi T, Steber C (2008) Molecular aspects of seed dormancy. Annu Rev Plant Biol 59:387–415. https://doi.org/10.1146/annurev.arplant.59.032607.092740
Finkelstein RR, Lynch TJ (2000) The Arabidopsis abscisic acid response gene ABI5 encodes a basic leucine zipper transcription factor. Plant Cell 12:599–609. https://doi.org/10.1105/tpc.12.4.599
Freiman A, Golobovitch S, Yablovitz Z, Belausov E, Dahan Y, Peer R, Avraham L, Freiman Z, Evenor D, Reuveni M, Sobolev V, Edelman M, Shahak Y, Samach A, Flaishman MA (2015) Expression of flowering locus T2 transgene from Pyrus communis L. delays dormancy and leaf senescence in Malus×domestica Borkh, and causes early flowering in tobacco. Plant Sci 241:164–176. https://doi.org/10.1016/j.plantsci.2015.09.012
Gubler F, Millar AA, Jacobsen JV (2005) Dormancy release, ABA and pre-harvest sprouting. Curr Opin Plant Biol 8:183–187. https://doi.org/10.1016/j.pbi.2005.01.011
Jiang DG, Chen WT, Dong JF, Li J, Yang F, Wu ZC, Zhou H, Wang WS, Zhuang CX (2018) Overexpression of miR164b-resistant OsNAC2 improves plant architecture and grain yield in rice. J Exp Bot 69:1533–1543. https://doi.org/10.1093/jxb/ery017
Jiang DG, Zhou LY, Chen WT, Ye NH, Xia JX, Zhuang CX (2019) Overexpression of a microRNA-targeted NAC transcription factor improves drought and salt tolerance in Rice via ABA-mediated pathways. Rice 12:76. https://doi.org/10.1186/s12284-019-0334-6
Jin S, Nasim Z, Susila H, Ahn JH (2021) Evolution and functional diversification of FLOWERING LOCUS T/TERMINAL FLOWER 1 family genes in plants. Semin Cell Dev Biol 109:20–30. https://doi.org/10.1016/j.semcdb.2020.05.007
Karlgrenet A, Gyllenstrand N, Källman T, Sundström JF, Moore D, Lascoux M, Lagercrantz U (2011) Evolution of the PEBP gene family in plants: functional diversification in seed plant evolution. Plant Physiol 156:1967–1977. https://doi.org/10.1104/pp.111.176206
Khan F, Upreti P, Singh R, Shukla PK, Shirke PA (2017) Physiological performance of two contrasting rice varieties under water stress. Physiol Mol Biol Plants 23:85–97. https://doi.org/10.1007/s12298-016-0399-2
Koramutla MK, Negi M, Ayele BT (2021) Roles of glutathione in mediating abscisic acid signaling and its regulation of seed dormancy and drought tolerance. Genes 12:1620. https://doi.org/10.3390/genes12101620
Kushiro T, Okamoto M, Nakabayashi K, Yamagishi K, Kitamura S, Asami T, Hirai N, Koshiba T, Kamiya Y, Nambara E (2004) The Arabidopsis cytochrome P450 CYP707A encodes ABA 8’-hydroxylases: key enzymes in ABA catabolism. EMBO J 23:1647–1656. https://doi.org/10.1038/sj.emboj.7600121
Lee SC, Luan S (2012) ABA signal transduction at the crossroad of biotic and abiotic stress responses. Plant Cell Environ 35:53–60. https://doi.org/10.1111/j.1365-3040.2011.02426.x
Li J, Jiang DG, Zhou H, Li F, Yang JW, Hong LF, Fu X, Li ZB, Liu ZL, Li JM, Zhuang CX (2011) Expression of RNA-interference/antisense transgenes by the cognate promoters of target genes is a better gene-silencing strategy to study gene functions in rice. PLoS ONE 6:e17444. https://doi.org/10.1371/journal.pone.0017444
Li XX, Yu B, Wu Q, Min Q, Zeng RF, Xie ZZ, Huang JL (2021) OsMADS23 phosphorylated by SAPK9 confers drought and salt tolerance by regulating ABA biosynthesis in rice. PLoS Genet 17:e1009699. https://doi.org/10.1371/journal.pgen.1009699
Lopez-Molina L, Mongrand S, Chua NH (2001) A postgermination developmental arrest checkpoint is mediated by abscisic acid and requires the ABI5 transcription factor in Arabidopsis. Proc Natl Acad Sci USA 98:4782–4787. https://doi.org/10.1073/pnas.081594298
Lopez-Molina L, Mongrand S, McLachlin DT, Chait BT, Chua NH (2002) ABI5 acts downstream of ABI3 to execute an ABA-dependent growth arrest during germination. Plant J 32:317–328. https://doi.org/10.1046/j.1365-313X.2002.01430.x
Luo X, Chen T, Zeng XL, He DW, He YH (2019) Feedback regulation of FLC by FLOWERING LOCUS T (FT) and FD through a 5’ FLC promoter region in Arabidopsis. Mol Plant 12:285–288. https://doi.org/10.1016/j.molp.2019.01.013
Lv Y, Yang M, Hu D, Yang ZY, Ma SQ, Li XH, Xiong LZ (2017) The OsMYB30 transcription factor suppresses cold tolerance by interacting with a JAZ protein and suppressing β-Amylase expression. Plant Physiol 173:1475–1491. https://doi.org/10.1104/pp.16.01725
Mahender A, Anandan A, Pradhan SK (2015) Early seedling vigour, an imperative trait for direct-seeded rice: an overview on physio-morphological parameters and molecular markers. Planta 241:1027–1050. https://doi.org/10.1007/s00425-015-2273-9
Ma XL, Zhang QY, Zhu QL, Liu W, Chen Y, Qiu R, Wang B, Yang ZF, Li HY, Lin YR, Xie YY, Shen RX, Chen SF, Wang Z, Chen YL, Guo JX, Chen LT, Zhao XC, Dong ZC, Liu YG (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8:1274–1284. https://doi.org/10.1016/j.molp.2015.04.007
Merlot S, Gosti F, Guerrier D, Vavasseur A, Giraudat J (2001) The ABI1 and ABI2 protein phosphatases 2C act in a negative feedback regulatory loop of the abscisic acid signalling pathway. Plant J 25:295–303. https://doi.org/10.1046/j.1365-313x.2001.00965.x
Miyazono KI, Miyakawa T, Sawano Y, Kubota K, Kang HJ, Asano A, Miyauchi Y, Takahashi M, Zhi Y, Fujita Y, Yoshida T, Kodaira KS, Yamaguchi-Shinozaki K, Tanokura M (2009) Structural basis of abscisic acid signaling. Nature 462:609–614. https://doi.org/10.1038/nature08583
Mohamed R, Wang CT, Ma C, Shevchenko O, Dye SJ, Puzey JR, Etherington E, Sheng XY, Meilan R, Strauss SH, Brunner AM (2010) Populus CEN/TFL1 regulates first onset of flowering, axillary meristem identity and dormancy release in Populus. Plant J 62:674–688. https://doi.org/10.1111/j.1365-313X.2010.04185.x
Morris WL, Carmen Alamar M, Lopez-Cobollo RM, Castillo Cañete J, Bennett M, Van der Kaay J, Stevens J, Kumar Sharma S, McLean K, Thompson AJ, Terry LA, Turnbull CGN, Bryan GJ, Taylor MA (2019) A member of the TERMINAL FLOWER 1/CENTRORADIALIS gene family controls sprout growth in potato tubers. J Exp Bot 70:835–843. https://doi.org/10.1093/jxb/ery387
Nakagawa M, Shimamoto K, Kyozuka J (2002) Overexpression of RCN1 and RCN2, rice TERMINAL FLOWER 1/CENTRORADIALIS homologs, confers delay of phase transition and altered panicle morphology in rice. Plant J 29:743–750. https://doi.org/10.1046/j.1365-313x.2002.01255.x
Nakamura S, Abe F, Kawahigashi H, Nakazono K, Tagiri A, Matsumoto T, Utsugi S, Ogawa T, Handa H, Ishida H, Mori M, Kawaura K, Ogihara Y, Miura H (2011) A wheat homolog of MOTHER OF FT AND TFL1 acts in the regulation of germination. Plant Cell 23:3215–3229. https://doi.org/10.1105/tpc.111.088492
Nonogaki H, Bassel GW, Derek Bewley J (2010) Germination-Still a mystery. Plant Sci 179:574–581. https://doi.org/10.1016/j.plantsci.2010.02.010
Piskurewicz U, Jikumaru Y, Nambara KN, E, Kamiya Y, Lopez-Molina L, (2008) The gibberellic acid signaling repressor RGL2 inhibits Arabidopsis seed germination by stimulating abscisic acid synthesis and ABI5 activity. Plant Cell 20:2729–2745. https://doi.org/10.1105/tpc.108.061515
Schmitz N, Abrams SR, Kermode AR (2002) Changes in ABA turnover and sensitivity that accompany dormancy termination of yellow-cedar (Chamaecyparis nootkatensis) seeds. J Exp Bot 53:89–101. https://doi.org/10.1093/jexbot/53.366.89
Song S, Wang GF, Wu H, Fan XW, Liang LW, Zhao H, Hu LSL, Y, Liu HY, Ayaad M, Xing YZ, (2020) OsMFT2 is involved in the regulation of ABA signaling-mediated seed germination through interacting with OsbZIP23/66/72 in rice. Plant J 103:532–546. https://doi.org/10.1111/tpj.14748
Shu K, Meng YJ, Shuai HW, Liu WG, Du JB, Liu J, Yang WY (2015) Dormancy and germination: How does the crop seed decide? Plant Biol 17:1104–1112. https://doi.org/10.1111/plb.12356
Shu K, Zhang HW, Wang SF, Chen ML, Wu YR, Tang SY, Liu CY, Feng YQ, Cao XF, Xie Q (2013) ABI4 regulates primary seed dormancy by regulating the biogenesis of abscisic acid and gibberellins in Arabidopsis. PLoS Genet 9:e1003577. https://doi.org/10.1371/journal.pgen.1003577
Söderman EM, Brocard IM, Lynch TJ, Finkelstein RR (2000) Regulation and function of the Arabidopsis ABA-insensitive4 gene in seed and abscisic acid response signaling networks. Plant Physiol 124:1752–1765. https://doi.org/10.1104/pp.124.4.1752
Sun L, Sun YF, Zhang M, Wang L, Ren J, Cui MM, Wang YP, Ji K, Li P, Li Q, Chen P, Dai SJ, Duan CR, Wu Y, Leng P (2012) Suppression of 9-cis-epoxycarotenoid dioxygenase, which encodes a key enzyme in abscisic acid biosynthesis, alters fruit texture in transgenic tomato. Plant Physiol 158:283–298. https://doi.org/10.1104/pp.111.186866
Wang J, Deng QW, Li YH, Yu YH, Liu X, Han YF, Luo XD, Wu XJ, Ju L, Sun JQ, Liu AH, Fang J (2020) Transcription factors Rc and OsVP1 coordinately regulate preharvest sprouting tolerance in red pericarp rice. J Agric Food Chem 68:14748–14757. https://doi.org/10.1021/acs.jafc.0c04748
Xi WY, Liu C, Liu C, Hou XL, Yu H (2010) MOTHER OF FT AND TFL1 regulates seed germination through a negative feedback loop modulating ABA Signaling in Arabidopsis. Plant Cell 22:1733–1748. https://doi.org/10.1105/tpc.109.073072
Yu L, Jiang JZ, Zhang C, Jiang LR, Ye NH, Lu YS, Yang GZ, Liu EE, Peng CL, He ZH, Peng XX (2010) Glyoxylate rather than ascorbate is an efficient precursor for oxalate biosynthesis in rice. J Exp Bot 61:1625–1634. https://doi.org/10.1093/jxb/erq028
Zhu GH, Ye NH, Zhang JH (2009) Glucose-induced delay of seed germination in rice is mediated by the suppression of ABA catabolism rather than an enhancement of ABA biosynthesis. Plant Cell Physiol 50:644–651. https://doi.org/10.1093/pcp/pcp022
Zhou H, Zhou M, Yang YZ, Li J, Zhu LY, Jiang DG, Dong JF, Liu QJ, Gu LF, Zhou LY, Feng MJ, Qin P, Hu XC, Song CL, Shi JF, Song XW, Ni ED, Wu XJ, Deng QY, Liu ZL, Chen MS, Liu YG, Cao XF, Zhuang CX (2014) RNase ZS1 processes UbL40 mRNAs and controls thermosensitive genic male sterility in rice. Nat Commun 5:4884. https://doi.org/10.1038/ncomms5884
Zou MJ, Guan YC, Ren HB, Zhang F, Chen F (2008) A bZIP transcription factor, OsABI5, is involved in rice fertility and stress tolerance. Plant Mol Biol 66:675–683. https://doi.org/10.1007/s11103-008-9298-4
Zou MJ, Guan YC, Ren HB, Zhang F, Chen F (2007) Characterization of alternative splicing products of bZIP transcription factors OsABI5. Biochem Biophys Res Commun 360:307–313. https://doi.org/10.1016/j.bbrc.2007.05.226
Acknowledgements
This work was supported by the Major Program of Guangdong Basic and Applied Research (grant no. 2019B030302006) and the National Natural Science Foundation of China (grant no. 31100872).
Author information
Authors and Affiliations
Contributions
CZ and DJ designed the research. YH and WC performed most experiments and analyzed experimental data. JT, XL, YZ, XG and JY conducted a part of experiments. HY wrote the manuscript, CZ and DJ revised the paper. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Matthias Wissuwa.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, Y., Chen, W., Tan, J. et al. Rice CENTRORADIALIS 2 regulates seed germination and salt tolerance via ABA-mediated pathway. Theor Appl Genet 135, 4245–4259 (2022). https://doi.org/10.1007/s00122-022-04215-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-022-04215-8