Abstract
Key message
We report the map-based cloning and functional characterization of SNG1, which encodes OsHXK3, a hexokinase-like protein that plays a pivotal role in controlling grain size in rice.
Abstract
Grain size is an important agronomic trait determining grain yield and appearance quality in rice. Here, we report the discovery of rice mutant short and narrow grain1 (sng1) with reduced grain length, width and weight. Map-based cloning revealed that the mutant phenotype was caused by loss of function of gene OsHXK3 that encodes a hexokinase-like (HKL) protein. OsHXK3 was associated with the mitochondria and was ubiquitously distributed in various organs, predominately in younger organs. Analysis of glucose (Glc) phosphorylation activities in young panicles and protoplasts showed that OsHXK3 was a non-catalytic hexokinase (HXK). Overexpression of OsHXK3 could not complement the Arabidopsis glucose insensitive2-1 (gin2-1) mutant, indicating that OsHXK3 lacked Glc signaling activity. Scanning electron microscopy analysis revealed that OsHXK3 affects grain size by promoting spikelet husk cell expansion. Knockout of other nine OsHXK genes except OsHXK3 individually did not change grain size, indicating that functions of OsHXKs have differentiated in rice. OsHXK3 influences gibberellin (GA) biosynthesis and homeostasis. Compared with wild type, OsGA3ox2 was significantly up-regulated and OsGA2ox1 was significantly down-regulated in young panicle of sng1, and concentrations of biologically active GAs were significantly decreased in young panicles of the mutants. The yield per plant of OsHXK3 overexpression lines (OE-4 and OE-35) was increased by 10.91% and 7.62%, respectively, compared to that of wild type. Our results provide evidence that an HXK lacking catalytic and sensory functions plays an important role in grain size and has the potential to increase yield in rice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Grain size in rice specified by grain length, width, and thickness determines grain weight and is an important component of grain yield. Grain size is also an important quality attribute with consumer preferences differing in different parts of the world (Zuo and Li 2014). About 20 quantitative trait loci (QTLs) regulating grain size have been validated over the past two decades. These genes act in diverse pathways such as phytohormone, proteasomal degradation, G-protein signaling and transcriptional regulation of regulate cell division and elongation (Fan and Li 2019; Li and Li 2016). However, the molecular basis of grain size in rice remains elusive. Identification of new genes controlling grain size may help to elucidate the molecular regulation of this trait.
Hexokinase (HXK) catalyzes ATP-dependent phosphorylation of hexoses. It is the first enzyme in glycolysis with an important rate-limiting role in Glc metabolism and homeostasis. HXK also has an important role in sugar signaling. Plant HXK family contains 5–11 genes, and functional diversification has occurred within particular species (Karve et al. 2010). The Arabidopsis HXK family contains three proteins that can phosphorylate Glc and three hexokinase-like (HKL) proteins that lack Glc metabolic activity (Karve et al. 2008). AtHXK1 and AtHXK2 are both hexokinase and Glc signaling proteins; AtHKL1, AtHKL2 and AtHKL3 might have only regulatory functions; and AtHXK3 likely has only a catalytic function (Jang et al. 1997; Karve et al. 2008). The most studied HXK in plants is AtHXK1. The AtHXK1 loss-of-function mutant glucose-insensitive2-1 (gin2-1) has a reduced root system, small leaves, and short petioles and inflorescences at the flowering stage (Moore et al. 2003). It also exhibits a Glc-insensitive phenotype in seedling development and decreased Glc -dependent responses in gene expression. The role of AtHXK1 in sugar signaling is independent of its metabolic function, as transgenic plants overexpressing catalytically inactive AtHXK1 mutant alleles in the gin2-1 mutant background complement gin2-1 and restore Glc dependency (Moore et al. 2003). AtHXK1 is mainly present in the mitochondria and can enter the nuclei through an unknown mechanism to affect sugar sensing and signaling (Cho et al. 2006b).
The HXK family in rice includes ten members. Cho et al. (2006a) reported that all the OsHXKs have hexokinase activity from a yeast complementation test. However, another study indicated that non-catalytic HXK might occur in rice, as OsHXK3 lacked apparent glucokinase activity in a transient expression assay in maize protoplasts (Karve et al. 2010). OsHXK5, OsHXK6 and OsHXK7 can complement the gin2-1 mutant, resulting in the wild-type phenotypes of seedling development, Glc-dependent repression, and plant growth, indicating that these members function as Glc sensors (Cho et al. 2009; Kim et al. 2016). The catalytic and sensory functions of OsHXK5 and OsHXK6 are uncoupled (Cho et al. 2009). OsHXK5, OsHXK6 and OsHXK10 affect pollen fertility (Lee et al. 2020; Huang et al. 2015; Xu et al. 2008). OsHXK6 physically associates with RF6 (restorer-of-fertility6) to control pollen development and rescue cytoplasmic male sterility (Huang et al. 2015). OsHXK10 has a role in anther dehiscence, pollen germination and controls seed setting rate (Xu et al. 2008). OsHXK1 positively regulates leaf senescence, and knocking-out of OsHXK1 potentially improves the photosynthetic efficiency and yield (Zheng et al. 2021a, b).
In this paper, we report the map-based cloning and characterization of SNG1 that positively influences grain size in rice. SNG1 encodes OsHXK3, an HKL protein localized on mitochondria lacking catalytic and sensory functions. Loss of function of OsHXK3 results in decreased bioactive GA levels in young panicles. Elevated transcript levels of OsHXK3 lead to enhanced grain size and weight.
Materials and methods
Field planting and trait measurement
The field trial was carried out under natural field conditions at the experimental stations of Huazhong Agricultural University at Wuhan (114.36° E, 30.49° N) and Lingshui (110.04° E, 18.51° N), China. Planting density was 16.5 cm between plants within rows; rows were 26 cm apart. Field management followed normal local practices.
All grains and productive panicles from each plant were used to measure the yield per plant, panicle number per plant, grains per panicle, panicle length and seed setting rate. Three months after harvest, about 500 fully filled grains were used for measuring grain length, width and weight via a digital yield traits scorer (Yang et al. 2014).
RNA preparation and real-time PCR
Total RNA was extracted from plant tissues using TRIzol reagent (Invitrogen). The first-strand cDNA was synthesized in 20 µl volumes containing 2 µg RNA and 200 U M-MLV reverse transcriptase (Promega Kit). The full-length OsHXK3 cDNA was amplified from first-strand cDNA of Hwayoung (HW) and confirmed by sequencing. Real-time PCR was carried out on a QuantStudio6 Flex machine using SYBR Green PCR (Takara) according to the manufacturer’s protocol.
Vector construction and rice transformation
To prepare the overexpression construct (OE), full-length OsHXK3 cDNA of HW was inserted into the plant binary vector pU1301, which was modified based on pCAMIA1301 (CAMBIA) and contained the maize Ubiquitin promoter (Huang et al. 2007). The 3.6 kb promoter fragment of OsHXK3 from HW fused with its cDNA was inserted into vector pCAMBIA1301 and used to prepare the complementation construct (Com). CRISPR/Cas9-related vectors were constructed based on the vector pCXUN-Cas9 according to a previously described method (Sun et al. 2017). Specific sequences of guide RNAs were designed according to the web tool CRISPR-P 2.0 (Liu et al. 2017).
The OE and CRISPR/Cas9-related constructs were introduced into HW, and the Com construct was introduced into sng1 mutant, by Agrobacterium-mediated transformation (Hiei et al. 1994).
GUS staining
The 3.6 kb promoter fragment of OsHXK3 was inserted into vector pDX2181 to drive the GUS expression. The construct was introduced into HW. Various tissues of transgene-positive plants were stained in a GUS buffer, and photographs were taken using a dissecting microscope (Jefferson et al. 1987).
Subcellular localization
The OsHXK3 cDNA (without the stop codon) was inserted into a pM999-YFP vector for subcellular localization analysis. Transient gene expression in rice protoplasts was carried out as described (Zhang et al. 2011). Imaging of fluorescent proteins was conducted using a confocal laser scanning microscope (Leica TCS SP2) with excitation wavelengths for YFP and RFP of 514 and 561 nm, respectively.
Scanning electron microscopy (SEM)
Samples were fixed in an FAA solution (90% ethanol, 5% formaldehyde and 5% glacial acetic acid) for SEM. After dehydration in a graded ethanol series and substitution with isoamyl acetate, samples were dried by carbon dioxide, coated with gold, and examined under a scanning electron microscope (JSM-6390LV, JEOL). The lengths and widths of cells were determined using ImageJ software (US National Institute of Health).
Measures of glycolytic enzyme activity
The activities of HXK, PFK and PK were assayed as described previously (Zhang et al. 2017), using kits purchased from SuZhou Keming Bioengineer Company, China. Young panicles of different genotype plants were sampled around 5:00 pm in the field. 0.1 g of plant material was ground to a fine powder in liquid nitrogen and incubated in 1 ml related enzyme extract buffers. The enzymatic activities were measured following the manufacturer’s instructions of the kits from SuZhou Keming Bioengineer Company.
To determine the HXK activities in protoplasts, constructs containing OsHXK5, OsHXK6, OsHXK3 and the mutant alleles driven by the 35S promoter and the internal control (construct containing the Renilla luciferase gene) were transformed into rice protoplast (Zhang et al. 2011). The relative HXK activities were expressed as the ratio of HXK activities to the Renilla luciferase.
Quantitative analysis of sugar content
Tissues including young panicles and developing endosperms were sampled around 5:00 pm in the field. The contents of glucose, fructose, sucrose and pyruvate were assayed spectrophotometrically (Stitt et al. 1989), using kits purchased from the Suzhou Keming Bioengineer Company, China. Three biological replicates were performed, with each replicate being measured by three technical repeats.
Arabidopsis transformation
OsHXK3 and sng1 cDNA were inserted into the pCHF3 vector under control of the 35S promoter to produce transgenic Arabidopsis plants overexpressing OsHXK3 and sng1. Agrobacterium tumefaciens EHA105 strains were used for transformation. gin2-1 mutant plants were transformed using the floral dip method as previously described (Clough and Bent 1998).
GAs measurements
Quantification of endogenous GAs in young panicles was performed as described (Ma et al. 2011). As a single 8 cm young panicle was insufficient to measure GA content, three young panicles (about 0.3 g) were pooled as a sample.
Primers
Primers used in this study are listed in Table S4 and S5.
Results
Isolation of the sng1 mutant
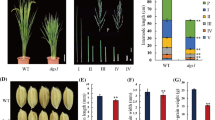
To understand the molecular basis of grain size, we isolated a small grain mutant sng1 (short and narrow grain 1-1) in the genetic background of japonica accession Hwayoung (HW) from a T-DNA insertion library (Jeon et al. 2000). The clone number was PFG_1C-12303. However, no T-DNA flanking sequence could be obtained. Compared with HW, the grain length and width of the sng1 mutant were reduced by 5.1% and 12.8%, respectively, and the 1000-grain weight was decreased by 18.8% (Fig. 1a–d). We also compared the grain filling rates of sng1 and HW and found that the fresh and dry grain weights of the mutant at 13 days post-fertilization were markedly decreased relative to HW, and differences reached a maximum at 22 days post-fertilization (Fig. 1e, f). Thus, the significant decrease in grain weight resulted from decreases in both hull size and rate of grain filling.
sng1 phenotypes and map-based cloning of SNG1. a Grain phenotypes of wild type (HW), sng1 and ZS97. Scale bar, 3 mm. b–d Grain length, width and weight of HW and sng1. Data are means ± SD (n = 36). Student’s t test was used to generate P values. e, f Time-course for grain fresh weight and grain dry weight (n = 100 grains for each data point). g Primary mapping of SNG1. h Fine mapping of the SNG1 region using a BC3F3 population with 7680 plants. i High-resolution linkage analysis of phenotypes and marker genotypes. Genotypes at the SNG1 locus were confirmed by progeny tests. j Sequence variations among HW, sng1 and ZS97 in the 5.5 kb region. No. of Recs, number of recombinants between SNG1 and indicated molecular marker
Map-based cloning of SNG1
An F2 population derived from a cross between sng1 and HW segregated 156 normal to 51 mutant, suggestive of single gene inheritance (χ23:1 = 0.01, P > 0.05) (Fig. S1). Map-based cloning was carried out to locate the mutant gene. An F2 population derived from a cross of sng1 mutant and Zhenshan 97 (ZS97, an indica cultivar) was used for preliminary mapping, and a QTL controlling grain length and width was located between insertion/deletion (InDel) markers Y11 and Y91 at the end of chromosome 1 by bulked segregant analysis (BSA), with the allele from ZS97 contributing to higher trait values (Table S1). The QTL was supposed to be SNG1, and its genetic effect was further validated in a BC3F2 population of 192 plants (Fig. S2 and Table S2).
Thirteen recombinants were identified between Y11 and Y91 in the BC3F2 population, and their genotypes at the SNG1 locus were determined by progeny testing. Based on newly developed markers, SNG1 was mapped between markers Y6 and Y54 (Fig. 1g). Further analysis of 7,680 plants from a BC3F3 population narrowed the position of SNG1 to an interval of 5.5 kb between markers Y1 and Y7 (Fig. 1h, i). Two open reading frames, LOC_Os01g71310 and LOC_Os01g71320, were present in this region (Fig. 1j).
Comparative sequence analysis of the entire region revealed a single nucleotide polymorphism (SNP) between HW and sng1 in the ninth exon of LOC_Os01g71320, changing an arginine codon (AGA) in HW to a termination codon (TGA) in sng1 (Fig. 1j). This premature termination resulted in a 103 amino acids truncation in the C-terminus of the predicted protein. The genotype of this SNP in ZS97 was the same as that of HW. There were 14 SNPs between HW and ZS97 in the region (Fig. 1j). Thus, LOC_Os01g71320 was considered the candidate for SNG1.
Database searches indicated that SNG1 encodes OsHXK3. Phylogenetic analysis produced similar results to previously published studies (Cho et al. 2006a; Karve et al. 2010). OsHXK3 and OsHXK10 are evolutionarily closely related to Arabidopsis AtHKL1 and AtHKL2, but not the Glc sensor AtHXK1 (Fig. S3). AtHKL1 and AtHKL2 encode HKL proteins that have potential regulatory functions, but do not phosphorylate Glc or affect Glc signaling (Karve et al. 2008; Karve and Moore 2009).
Confirmation of OsHXK3 as SNG1
To test whether OsHXK3 corresponds to the candidate gene, a complementation vector (Com) bearing the HW OsHXK3 coding region driven by its native promoter was constructed and introduced into sng1. Transgene-positive plants showed increased grain length, width and weight compared to transgene-negative plants. Grain size in some transgene-positive plants was restored to the wild-type level (Fig. 2a–d). Co-segregation in T1 progenies from independent T0 plants showed that grain length, width and weight co-segregated with the introduction of OsHXK3 (Fig. S4).
Verification of OsHXK3 function. a Grain shape of the complementation (Com) transgenic plants in T0 progeny. B–d Analysis of grain length, width and weight between Com transgene negative (−) and transgene positive (+) T0 progeny. e Grain shape of overexpressing (OE) transgenic T0 plants. f–h Analysis of grain length, width and weight between OE transgene negative (−) and transgene positive (+) plants in T0 progeny. i sng1c-1 and sng1c-2 knockout mutants generated by CRISPR-Cas9. Upper panel, a gene model of OsHXK3 to show positions of target sites. Lower panel, mutation sites in sng1c-1 and sng1c-2, compared to the HW sequence. Sequences in bold, target sites; underlined, protospacer-adjacent motif sequences (PAMs). Arrowheads indicate insertion sites. j Grain shapes of sng1c-1 and sng1c-2 transgenic plants in T0 progeny. K–m Analysis of grain length, width and weight in HW and T2 progeny of mutants without T-DNA insertions. Scale bars in a, e and j, 3 mm. Data in b-d, f–h and k–m are means ± SD (n = 24). P values were produced by two-tailed t tests
We constructed an overexpression vector (OE) carrying the OsHXK3 coding region driven by the maize Ubiquitin promoter and introduced it into HW. Transgene-positive plants showed increased grain size and weight (Fig. 2e–h). Again, grain size and weight co-segregated with the genotype in T1 lines (Fig. S5). Overexpression of OsHXK3 had little effect on plant morphology and flowering time (Fig. S6).
Two CRISPR-Cas9 constructs expressing guide RNAs targeting regions in the fifth and ninth exons of OsHXK3 were made and separately transformed into HW. Sequencing of target regions from transgenic plants allowed us to identify two knockout mutants: sng1c-1 with a 4-nt (AGAC) deletion in the ninth exon, and sng1c-2 with a 1-nt (G) insertion in the fifth exon (Fig. 2i), predicted to cause frameshifts and produce truncated proteins (Fig. S7). Gross morphologies of sng1, sng1c-1 and sng1c-2 plants were similar (Fig. S8), and grains produced from homozygous sng1c-1 and sng1c-2 plants without T-DNA insertions were markedly smaller than those of HW (Fig. 2j–m).
These results confirmed that the mutant sng1 was caused by loss-of function of OsHXK3. And OsHXK3 positively affects grain length, grain width and grain weight.
OsHXK3 expression pattern and subcellular localization of its gene product
Twelve tissues from HW were assayed for temporal and spatial expression patterns of OsHXK3 by real-time PCR. OsHXK3 was expressed in all tissues examined (Fig. 3a), consistent with a previous study (Cho et al. 2006a). We next examined the tissue expression specificity of OsHXK3 in transgenic plants harboring a OsHXK3 promoter::β-glucuronidase (GUS) construct. GUS expression was observed in all tissues, and the GUS signal was particularly strong in young tissues such as shoots of seedlings 7 days after germination, tiller buds and endosperm 7 days after fertilization, as well as in interconnecting tissues such as pulvini and nodes (Fig. 3b–m).
Expression pattern of OsHXK3. a Expression pattern of OsHXK3 in HW determined by real-time PCR. Tissues: R, S and FL, root, stem, and flag leaf at the heading stage; 2P, 4P, 6P, 8P, 10P and 15P, young panicles of 2, 4, 6, 8, 10 and 15 cm in length; H2B, hulls 2 days before heading; 7E and 14E, endosperm 7 and 14 days post-fertilization. All data are based on four biological replications, each with three technical replications. Error bars, SD. b–m Representative histochemical analysis of tissue expression of GUS transgene under the control of the OsHXK3 promoter. b Seedlings 7 days post-germination (T1 progeny); c Stems; d Leaf sheaths; e Flag leaves; f Pulvinus; g Node; h Tiller bud; i Young panicle; j Seed 7 days post-fertilization; k Endosperm 7 days post-fertilization; l Seed 20 days post-fertilization; m Endosperm 20 days post-fertilization. n Subcellular-localization of OsHXK3 protein in rice protoplasts. The Arabidopsis thaliana F1-ATPase protein fused with RFP is a mitochondrial specific marker. Scale bars, 5 μm
OsHXK3 is a type-B HXK characterized by a common hydrophobic membrane anchor domain (Olsson et al. 2003). Type-B HXKs are associated with mitochondria in several plant species (Cheng et al. 2011; Kandel-Kfir et al. 2006; Karve and Moore 2009; Kim et al. 2013). We transiently co-expressed OsHXK3 fused to yellow fluorescent protein (YFP) with F1-ATPase-γ:RFP, a mitochondrial marker in rice protoplasts (Jin et al. 2003), to determine the subcellular localization of OsHXK3. Yellow fluorescent signals of OsHXK3-YFP overlapped with red fluorescent signals of F1-ATPase-γ:RFP (Fig. 3n). Therefore, OsHXK3 was associated with the mitochondria.
OsHXK3 influences cell expansion in spikelet hulls
The spikelet hulls limit grain growth and determine grain size in rice. The final size of a spikelet hull is coordinately determined by cell proliferation and cell expansion (Li and Li 2016). To understand how OsHXK3 influences grain size in cellular level, we investigated cells in HW and sng1 spikelet hulls by scanning electron microscopy (SEM). Both length and width of outer epidermal cells in sng1 spikelet hulls were dramatically decreased compared with that in HW spikelet hulls (Fig. 4a–d). Similarly, sng1 hulls contained smaller cells in the inner epidermis compared with HW hulls (Fig. 4e–h). We further counted cell numbers in HW and sng1 spikelet hulls. The outer epidermal cell numbers in both the grain width direction and the grain length direction showed no differences between HW and sng1 (Fig. 4i, f). We also examined the expression of 25 genes involved in the cell cycle in 8-cm young panicles (Li et al. 2011b) and observed that none was significantly down-regulated in sng1 relative to HW (Fig. S9), suggesting that decreased seed size in sng1 did not result from repression of cell division. These results indicate that OsHXK3 positively influences grain size by increasing cell size, sharing a similar mechanism to AtHXK1 in controlling organ size in Arabidopsis (Moore et al. 2003).
OsHXK3 regulates grain size by affecting cell expansion. a, b The outer surfaces of HW and sng1 grains observed by scanning electron microscope (SEM). Bars = 100 μm. c, d Outer epidermal cell length and width of HW and sng1 lemmas (n ≥ 120). e, f The inner surfaces of HW and sng1 glumes observed by SEM. Bars = 100 μm. g, h Inner epidermal cell length and width of HW and sng1 lemmas (n ≥ 120). i, j Outer epidermal cell number in the grain length direction of the lemma and in the grain-width direction (n ≥ 10). Data in c, d and g, f are means ± SD. **P < 0.01, *P < 0.05, based on Student’s two-tailed t test
OsHXK3 lacks Glc phosphorylation and Glc signaling activities
To examine the HXK activity of OsHXK3, we measured Glc phosphorylation activities in 8-cm young panicles, which is an important stage affecting glume development and grain size. Compared with HW, Glc phosphorylation activity was slightly decreased in sng1, while no change was observed in sng1c-1, sng1c-2 and two homozygous OsHXK3 overexpression lines (OE-4 and OE-35) (Fig. 5a). Dosage of Glc, fructose (Fru) and sucrose (Suc) in the mutants and overexpression lines showed no significant differences with those in HW (Fig. 5b–d). In order to determine whether OsHXK3 affects sugar metabolism, enzymatic activities of 6-phosphofructokinase and pyruvate kinase were analyzed, which are rate-limiting enzymes in glycolysis, and no consistent changes were observed (Fig. 5e, f). And there were no differences in the contents of pyruvate, which is the product of glycolysis (Fig. 5g). Glc, Fru and Suc contents in developing endosperms of the mutants were not influenced (Fig. S10). We also confirmed that OsHXK3 is catalytically inactive in rice protoplasts. Protoplasts overexpressing OsHXK5 and OsHXK6 were used as positive controls, and their measured HXK activities after 12 h of culturing were enhanced about sevenfold over background levels of endogenous HXK activities. In contrast, expression of wild-type OsHXK3 and the mutant alleles could not increase Glc phosphorylation activity in rice protoplasts (Fig. 5h). These results indicated that OsHXK3 lacks HXK activity.
OsHXK3 lacks hexokinase activity. a Glucose phosphorylation activities, b glucose contents, c fructose contents, d sucrose contents, e 6-phosphofructokinase activities, f pyruvate kinase activities, and g pyruvate contents in 8-cm young panicles of the mutants (sng1, sng1c-1 and sng1c-2), and overexpression transgenic (OE-4 and OE-35) compared to HW. h Relative glucose phosphorylation activity in rice protoplasts expressing OsHXK5, OsHXK6, OsHXK3 and the mutant alleles under the control of the 35S promoter. A construct containing the Renilla luciferase gene was transformed as an internal control. Control protoplasts (CK) were transfected with empty vector. Glucose phosphorylation activity in the CK was arbitrarily defined as 1. Bars are means ± SD from three biological replicates and each replicate with three technical repeats. Significant differences are indicated (Student’s two-tailed t test, n = 3), *P < 0.05, **P < 0.01
To examine the functions of OsHXK3 in sugar sensing and signaling, we transformed the wild-type OsHXK3 and sng1 mutant allele into Arabidopsis gin2-1 mutant under the control of the 35S promoter. After obtaining homozygous transgenic lines with relatively high transgene expression levels, we germinated the progenies on 1/2 MS media supplemented with 6% Glc. Growth of wild type and sng1 mutant transgenic plants was similar to gin2-1 mutant and was not drastically suppressed by 6% Glc (Fig. S11a). All tested lines showed no difference when grown with 6% mannitol, indicating that the high Glc effects in transgenic plants were not caused by osmotic stress (Fig. S11b). In Arabidopsis, carbonic anhydrase (CAA) is a gene downstream of Glc signaling, and its expression is repressed by high Glc treatment with the existence of wild-type AtHXK1 (Moore et al. 2003; Cho et al. 2009). Therefore, we measured the expression of the CAA gene in transgenic plants by RT-PCR. Consistently, Glc repressed CAA expression in wild-type Arabidopsis (Ler), but the transgenic plants overexpressing OsHXK3 and sng1 maintained CAA expression at comparable levels to that of the gin2-1 mutant (Fig. S11a). These findings indicated that OsHXK3 could not complement the role of AtHXK1 in Glc sensing and signaling in Arabidopsis.
Rice seedlings overexpressing Glc sensor OsHXK5 or OsHXK6 exhibited hypersensitive growth retardation in response to Glc treatment (Cho et al. 2009). To further investigate the function of OsHXK3 in Glc sensing, seeds of HW, the mutants and two overexpression lines were germinated on water agar media containing 60 mM Glc. The growth of all seedling plants was more severely inhibited on Glc-containing media than on media containing 60 mM sorbitol. Compared with HW controls, seedlings of the three mutants did not display weakened Glc-dependent growth inhibition, and the two overexpression lines seedlings did not have enhanced Glc-dependent growth inhibition, (Fig. S12). These results showed that OsHXK3 does not act as a Glc sensor in rice.
Knockout of other OsHXK genes did not change grain size
The rice genome encodes nine other OsHXK3 homologs. To test whether they influence grain size, we first examined the expression profiles of those genes by real-time PCR. Compared with the control gene, Actin1, OsHXK1 and OsHXK10 were almost not expressed in most tissues (data not shown), consistent with previous studies (Cho et al. 2006a; Xu et al. 2008). OsHXK2, OsHXK4, OsHXK5, OsHXK6, OsHXK7, OsHXK8 and OsHXK9 showed similar expression patterns; they were mainly expressed in flag leaves, stems and roots, and their expression levels in young panicles were relatively low (Fig. S13), whereas OsHXK3 was relatively highly expressed (Fig. 3a). We then knocked out other OsHXK genes in HW using the CRISPR-Cas9 system. According to the homology of their sequences, OsHXK1, OsHXK7 and OsHXK8 were edited simultaneously using a single construct. OsHXK2 and OsHXK9, OsHXK3 and OsHXK10, OsHXK5 and OsHXK6 were also edited simultaneously, respectively. Only OsHXK4 was edited alone. After genotyping, transgenic positive plants were backcrossed with HW. For each construct, two independent F1 plants without T-DNA insertions were chosen to develop F2 populations for assessment of whether OsHXK genes affected grain size. Grain size and weight of hxk1hxk7hxk8 triple mutants, hxk2hxk9 double mutants and hxk4 mutants showed no differences from wild-type plants segregated from corresponding F2 populations, respectively (Fig. 6 and Figs. S14 and S15). hxk5hxk6 double mutants were sterile, and their husk size was unchanged compared with HW (Figs. S14 and S16). Only knockout of OsHXK3, but not OsHXK10, its closest homolog, significantly decreased grain length, width and weight (Fig. 6 and Figs. S14 and S15). These results indicated that the functions of OsHXKs have differentiated and that OsHXK3 has a unique role in determining grain size.
Effects of knocking out OsHXKs on grain size and weight in rice. a–c Statistical data for grain length, width and weight of OsHXK mutants and corresponding controls. Uppercase HXKs represent wild-type genotypes, and lowercase hxks represent mutant genotypes. Data are from BC1F2 populations and are shown as means ± SD (n ≥ 4). CKs are wild-type plants segregated from corresponding BC1F2 populations. Student’s t tests were used to generate the P values. Different letters (in HXK3 and HXK10) indicate significant differences at the 1% level, determined by the Tukey–Kramer test
OsHXK3 affects GA biosynthesis
Gibberellins (GAs) with biological activity are hormones that stimulate plant organ growth mainly through enhancement of cell elongation. Several studies indicate that GAs affect grain size in rice (Ashikari et al. 1999; Li et al. 2011a; Liu et al. 2011). The observed defect in cell expansion in spikelet hulls of sng1 mutant suggested a potential link between OsHXK3 and plant growth hormones involved in cell elongation. And the sng1, sng1c-1 and sng1c-2 mutants all showed some GA deficient or insensitive phenotypes, such as delayed seed germination and retarded seedling growth (Fig. S17), and small and dark green flag leaves (Fig. S18). These results suggested that OsHXK3 might be related to GA biosynthesis or signaling.
We measured the expression of genes involved in GA biosynthesis and signal transduction in 8-cm young panicles of HW and sng1 by real-time PCR. The expression level of OsGA3ox2 was eightfold higher in sng1 than in HW, whereas expression level of OsGA2ox1 was tenfold lower in sng1 (Fig. 7b). GA 3-oxidase (GA3ox) catalyzed the formation of bioactive GAs, and GA 2-oxidase (GA2ox) deactivated bioactive GAs and their precursors (Fig. 7a). Compared with HW, the expression level of CPS was 2.5-fold higher in sng1. Expression levels of genes involved in GA signal transduction showed no differences between HW and sng1 (Fig. 7b).
Expression levels of genes involved in GA biosynthesis and signal transduction in 8-cm young panicles of HW and sng1 mutant. a Schematic of the GA metabolism pathway in rice. Bioactive GAs are shown in black rectangles. The figure is modified from Ma et al. (2011). b Expression levels of genes involved in GA biosynthesis and signal transduction. Expression levels were determined by real-time PCR with four biological replications, each with three technical replications. Black and white bars represent HW and sng1 mutant, respectively. Data are means ± SD. Significant differences are indicated (Student’s two-tailed t test, n = 4), *P < 0.05, **P < 0.01
We then measured GA concentrations in 8-cm young panicles of HW, sng1, sng1c-1 and sng1c-2. Concentrations of biologically active GAs (GA1, GA3 and GA4) were all significantly decreased, and concentrations of upstream precursors of biologically active GAs (GA12 and GA53) were also significantly decreased in all the mutants (Table 1).
Plants maintain optimal bioactive GA levels for coordinated plant growth and development. This homoeostatic regulation is mainly mediated through transcriptional control of specific dioxygenases (GA20oxs, GA3oxs and GA2oxs) in the GA metabolic pathway (Hedden and Thomas 2012; Yamaguchi 2008). For example, elevated levels of bioactive GA in Arabidopsis suppress the expression of some GA20ox and GA3ox genes while stimulating the expression of GA2ox. By contrast, reduced bioactive GA levels up-regulate expression of GA20ox and GA3ox and down-regulate expression of GA2ox (Mitchum et al. 2006; Phillips et al. 1995; Rieu et al. 2008). Up-regulation of OsGA3ox2 and down-regulation of OsGA2ox1 in 8-cm young panicle of sng1 mutant indicated a homeostatic regulation mechanism to balance the decreased levels of bioactive GAs in the mutants.
Finally, we analyzed second leaf sheath lengths in response to exogenous GA3 treatment. The second leaf sheaths of the mutants were shorter than those of HW with concentrations of exogenous GA3 below 10−7 M. With GA3 treatment up to 10−5 M, the mutant sheaths were almost as long as HW sheaths (Table S3). This finding suggests that the mutants could respond to exogenous GA3 to rescue the dwarf phenotype, and the mutants might be deficient in active GAs. Taken together, loss of function of OsHXK3 in rice disrupts gibberellin biosynthesis.
Application potential of OsHXK3 in rice breeding
To determine the effects of OsHXK3 on yield components, we grew HW and the overexpression lines OE-4 and OE-35 in the field and measured their agronomic characteristics. The results showed that, compared with HW, the panicle number per plant of OE-4 and OE-35 was increased by 17.61% and 15.12%, respectively. The grain size and weight of the overexpression lines were also significantly increased. While compared to the grains per panicle of HW, those of OE-4 and OE-35 were decreased by 8.47% and 7.62%, respectively. And the yield per plant of OE-4 and OE-35 was increased by 10.91% and 7.62%, respectively, compared with that of HW (Table 2). Thus, overexpression of OsHXK3 might have a practical role in improving grain productivity.
We also investigated yield components of the mutants. All of the mutants showed significantly decreased grain size, grain weight and yield per plant in comparison with HW (Table 2). The seed setting rates of sng1, sng1c-1 and sng1c-2 mutants were 32.8%, 27.2% and 29.5% lower than that of HW, respectively. Indicating that OsHXK3 has a role in fertility control, similar to those of OsHXK5, OsHXK6 and OsHXK10 (Lee et al. 2020; Huang et al. 2015; Xu et al. 2008). In addition, the days to heading and the panicle number per plant were slightly increased, whereas the plant height, the grains per panicle and the panicle length were slightly decreased (Table 2). The overall impacts of OsHXK3 on these traits corresponded to its consecutive expressing pattern.
Discussion
A number of genes and QTLs for grain size have been identified and characterized in recent years. However, our understanding of the underlying mechanisms is still fragmentary and requires cloning of more genes controlling this important trait. In this study, we report map-based cloning and functional characterization of OsHXK3 that positively influences grain length, width and weight by promoting cell expansion. OsHXK3 encodes an HKL protein without Glc phosphorylation and signaling activities. OsHXK3 controls GA biosynthesis and affects biologically active GA levels, leading to changes in the expression levels of specific dioxygenase genes possibly via homeostatic regulation mechanisms of GAs. We concluded that OsHXK3 positively controls glume cell expansion partially by affecting GA biosynthesis, as a consequence influencing grain size and weight.
HXKs are ancient and conserved metabolic enzymes that phosphorylate hexoses. Several HXKs act as Glc sensors that affect sugar signaling and sensing in plants (Cho et al. 2009, 2006b; Hu et al. 2016; Karve et al. 2010; Kim et al. 2016; Moore et al. 2003; Veramendi et al. 2002). It has been suggested that HXKs control organ size, programmed cell death, seed development, starch content, and pollen germination (Granot et al. 2013). However, which of the above roles are related to sugar sensing or metabolic catalytic functions of HXK remain to be determined. The catalytically inactive alleles of AtHXK1, OsHXK5 and OsHXK6 complemented Glc-sensitive gin2-1 seedlings under high Glc conditions and significantly enhanced Glc repression of the sugar-responsive genes in mesophyll protoplasts, indicating that sensory and catalytic functions of the HXKs are uncoupled (Cho et al. 2009; Moore et al. 2003). OsHXK7 overexpression rescued the Glc-sensitive phenotype in gin2-1 mutant, but the catalytically inactive mutant alleles did not, suggesting that OsHXK7 functions in sugar signaling via a glycolysis-dependent pathway (Kim et al. 2016). OsHXK3 is evolutionarily closely related to AtHKL1 and AtHKL2, which do not phosphorylate Glc or affect Glc signaling (Karve et al. 2008; Karve and Moore 2009). And AtHKL1 negatively regulates plant growth under conditions of excessive Glc availability (Karve and Moore 2009). Analysis of Glc phosphorylation activities in young panicles and protoplasts showed that OsHXK3 was a non-catalytic HXK. OsHXK3 could neither recover the Glc-sensitive response in Arabidopsis gin2-1 mutant, nor affect sensitivity to Glc in rice seedlings. These results indicated that OsHXK3 lacks Glc phosphorylation and signaling activities. However, OsHXK3 has an important role in controlling growth and development in rice.
AtHXK1-mediated Glc signaling affects several hormonal signaling pathways in Arabidopsis, such as abscisic acid, auxin, cytokinin and ethylene (Ramon et al. 2008; Rolland et al. 2006). A study in Arabidopsis showed that sugar metabolism modulates GA synthesis and determines plant size (Paparelli et al. 2013). Arabidopsis mutants defective in either starch synthesis or degradation have dwarf stature, and the dwarf phenotype can be rescued by exogenous GA application. Additionally, carbon starvation represses the expression of ent-kaurene synthase (KS), leading to low levels of kaurene, a precursor of GA. It seems that synthesis of GA depends on the availability of sugars in order to coordinate growth (Paparelli et al. 2013). Biologically active GA (GA1, GA3 and GA4) levels were significantly decreased in three independent mutants (OsHXK3, OsHXK3c-1 and OsHXK3c-2). As the major ent-kaurene synthase gene (KS1) was not down-regulated in young panicles of OsHXK3 mutant (Fig. 7b), OsHXK3 apparently controls GA biosynthesis through a mechanism different from that reported in Arabidopsis by Paparelli et al. (2013). OsGA3ox2 was up-regulated, and OsGA2ox1 was down-regulated in young panicle of OsHXK3 mutant, possibly by homeostatic regulation of GAs. However, the detailed molecular mechanism of how OsHXK3 controls GA biosynthesis remains to be investigated.
Constitutive overexpression of AtHXK1 in Arabidopsis and tomato has growth-inhibiting effects (Dai et al. 1999; Jang et al. 1997; Kelly et al. 2012), which might be caused by continuous unnatural reduction of photosynthesis by sugar-sensing effects of AtHXK1. Growth-inhibiting effects were not observed in OsHXK3 overexpression lines in rice (Table 2 and Fig. S6). In a recent study, knocking out of OsHXK1 in three different indica varieties improved rice yield, mainly by increasing the panicle number per plant and the grains per panicle (Zheng et al. 2021b). However, in our study, the yield of OsHXK3 mutants were significantly reduced. And overexpression of OsHXK3 increased the panicle number per plant and the grain weight and decreased the grains per panicle. Compared with HW, the yield per plant of OE-4 and OE-35 was increased by 10.91% and 7.62%, respectively (Table 2). Obviously, OsHXK3 acts differently from OsHXK1 in controlling growth and development of rice. Our findings advance the understanding of the molecular basis of regulation of grain size and may help to improve rice yield and quality.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Ashikari M, Wu J, Yano M, Sasaki T, Yoshimura A (1999) Rice gibberellin-insensitive dwarf mutant gene Dwarf 1 encodes the alpha-subunit of GTP-binding protein. Proc Natl Acad Sci 96:10284–10289
Chen W, Gao Y, Xie W, Gong L, Lu K, Wang W, Li Y, Liu X, Zhang H, Dong H, Zhang W, Zhang L, Yu S, Wang G, Lian X, Luo J (2014) Genome-wide association analyses provide genetic and biochemical insights into natural variation in rice metabolism. Nat Genet 46:714–721
Cheng W, Zhang H, Zhou X, Liu H, Liu Y, Li J, Han S, Wang Y (2011) Subcellular localization of rice hexokinase (OsHXK) family members in the mesophyll protoplasts of tobacco. Biol Plantarum 55:173–177
Cho JI, Ryoo N, Ko S, Lee SK, Lee J, Jung KH, Lee YH, Bhoo SH, Winderickx J, An G, Hahn TR, Jeon JS (2006a) Structure, expression, and functional analysis of the hexokinase gene family in rice (Oryza sativa L.). Planta 224:598–611
Cho YH, Yoo SD, Sheen J (2006b) Regulatory functions of nuclear hexokinase1 complex in glucose signaling. Cell 127:579–589
Cho JI, Ryoo N, Eom JS, Lee DW, Kim HB, Jeong SW, Lee YH, Kwon YK, Cho MH, Bhoo SH, Hahn TR, Park YI, Hwang I, Sheen J, Jeon JS (2009) Role of the rice hexokinases OsHXK5 and OsHXK6 as glucose sensors. Plant Physiol 149:745–759
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Dai N, Schaffer A, Petreikov M, Shahak Y, Giller Y, Ratner K, Levine A, Granot D (1999) Overexpression of Arabidopsis hexokinase in tomato plants inhibits growth, reduces photosynthesis, and induces rapid senescence. Plant Cell 11:1253–1266
Fan Y, Li Y (2019) Molecular, cellular and Yin-Yang regulation of grain size and number in rice. Mol Breed 39:1–25
Granot D, David-Schwartz R, Kelly G (2013) Hexose kinases and their role in sugar-sensing and plant development. Front Plant Sci 4:44
Hedden P, Thomas SG (2012) Gibberellin biosynthesis and its regulation. Biochem J 444:11–25
Hiei Y, Ohta S, Komari T, Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6:271–282
Hu DG, Sun CH, Zhang QY, An JP, You CX, Hao YJ (2016) Glucose sensor MdHXK1 phosphorylates and stabilizes MdbHLH3 to promote anthocyanin biosynthesis in apple. PLoS Genet 12:e1006273
Huang L, Sun Q, Qin F, Li C, Zhao Y, Zhou DX (2007) Down-regulation of a SILENT INFORMATION REGULATOR2-related histone deacetylase gene, OsSRT1, induces DNA fragmentation and cell death in rice. Plant Physiol 144:1508–1519
Huang W, Yu C, Hu J, Wang L, Dan Z, Zhou W, He C, Zeng Y, Yao G, Qi J, Zhang Z, Zhu R, Chen X, Zhu Y (2015) Pentatricopeptide-repeat family protein RF6 functions with hexokinase 6 to rescue rice cytoplasmic male sterility. Proc Natl Acad Sci 112:14984–14989
Jang JC, Leon P, Zhou L, Sheen J (1997) Hexokinase as a sugar sensor in higher plants. Plant Cell 9:5–19
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Jeon JS, Lee S, Jung KH, Jun SH, Jeong DH, Lee J, Kim C, Jang S, Yang K, Nam J et al (2000) T-DNA insertional mutagenesis for functional genomics in rice. Plant J 22:561–570
Jin JB, Bae H, Kim SJ, Jin YH, Goh CH, Kim DH, Lee YJ, Tse YC, Jiang L, Hwang I (2003) The Arabidopsis dynamin-like proteins ADL1C and ADL1E play a critical role in mitochondrial morphogenesis. Plant Cell 15:2357–2369
Kandel-Kfir M, Damari-Weissler H, German MA, Gidoni D, Mett A, Belausov E, Petreikov M, Adir N, Granot D (2006) Two newly identified membrane-associated and plastidic tomato HXKs: characteristics, predicted structure and intracellular localization. Planta 224:1341–1352
Karve A, Moore BD (2009) Function of Arabidopsis hexokinase-like1 as a negative regulator of plant growth. J Exp Bot 60:4137–4149
Karve A, Rauh BL, Xia X, Kandasamy M, Meagher RB, Sheen J, Moore BD (2008) Expression and evolutionary features of the hexokinase gene family in Arabidopsis. Planta 228:411–425
Karve R, Lauria M, Virnig A, Xia X, Rauh BL, Moore B (2010) Evolutionary lineages and functional diversification of plant hexokinases. Mol Plant 3:334–346
Kelly G, David-Schwartz R, Sade N, Moshelion M, Levi A, Alchanatis V, Granot D (2012) The pitfalls of transgenic selection and new roles of AtHXK1: a high level of AtHXK1 expression uncouples hexokinase1-dependent sugar signaling from exogenous sugar. Plant Physiol 159:47–51
Kim YM, Heinzel N, Giese JO, Koeber J, Melzer M, Rutten T, Von Wiren N, Sonnewald U, Hajirezaei MR (2013) A dual role of tobacco hexokinase 1 in primary metabolism and sugar sensing. Plant Cell Environ 36:1311–1327
Kim HB, Cho JI, Ryoo N, Shin DH, Park YI, Hwang YS, Lee SK, An G, Jeon JS (2016) Role of rice cytosolic hexokinase OsHXK7 in sugar signaling and metabolism. J Integr Plant Biol 58:127–135
Lee SK, Kim H, Cho JI, Nguyen CD, Jeon JS (2020) Deficiency of rice hexokinase HXK5 impairs synthesis and utilization of starch in pollen grains and causes male sterility. J Exp Bot 71:116–125
Li N, Li Y (2016) Signaling pathways of seed size control in plants. Curr Opin Plant Biol 33:23–32
Li J, Jiang J, Qian Q, Xu Y, Zhang C, Xiao J, Du C, Luo W, Zou G, Chen M, Huang Y, Feng Y, Cheng Z, Yuan M, Chong K (2011a) Mutation of rice BC12/GDD1, which encodes a kinesin-like protein that binds to a GA biosynthesis gene promoter, leads to dwarfism with impaired cell elongation. Plant Cell 23:628–640
Li Y, Fan C, Xing Y, Jiang Y, Luo L, Sun L, Shao D, Xu C, Li X, Xiao J, He Y, Zhang Q (2011b) Natural variation in GS5 plays an important role in regulating grain size and yield in rice. Nat Genet 43:1266–1269
Liu Y, Xu Y, Xiao J, Ma Q, Li D, Xue Z, Chong K (2011) OsDOG, a gibberellin-induced A20/AN1 zinc-finger protein, negatively regulates gibberellin-mediated cell elongation in rice. J Plant Physiol 168:1098–1105
Liu H, Ding Y, Zhou Y, Jin W, Xie K, Chen L (2017) CRISPR-P 2.0: an improved CRISPR-Cas9 tool for genome editing in plants. Mol Plant 10:530–532
Ma Q, Hedden P, Zhang Q (2011) Heterosis in rice seedlings: its relationship to gibberellin content and expression of gibberellin metabolism and signaling genes. Plant Physiol 156:1905–1920
Mitchum MG, Yamaguchi S, Hanada A, Kuwahara A, Yoshioka Y, Kato T, Tabata S, Kamiya Y, Sun TP (2006) Distinct and overlapping roles of two gibberellin 3-oxidases in Arabidopsis development. Plant J 45:804–818
Moore B, Zhou L, Rolland F, Hall Q, Cheng WH, Liu YX, Hwang I, Jones T, Sheen J (2003) Role of the Arabidopsis glucose sensor HXK1 in nutrient, light, and hormonal signaling. Science 300:332–336
Olsson T, Thelander M, Ronne H (2003) A novel type of chloroplast stromal hexokinase is the major glucose-phosphorylating enzyme in the moss Physcomitrella patens. J Biol Chem 278:44439–44447
Paparelli E, Parlanti S, Gonzali S, Novi G, Mariotti L, Ceccarelli N, van Dongen JT, Kolling K, Zeeman SC, Perata P (2013) Nighttime sugar starvation orchestrates gibberellin biosynthesis and plant growth in Arabidopsis. Plant Cell 25:3760–3769
Phillips AL, Ward DA, Uknes S, Appleford NE, Lange T, Huttly AK, Gaskin P, Graebe JE, Hedden P (1995) Isolation and expression of three gibberellin 20-oxidase cDNA clones from Arabidopsis. Plant Physiol 108:1049–1057
Ramon M, Rolland F, Sheen J (2008) Sugar Sensing and Signaling. Arabidopsis Book 6:e0117
Rieu I, Ruiz-Rivero O, Fernandez-Garcia N, Griffiths J, Powers SJ, Gong F, Linhartova T, Eriksson S, Nilsson O, Thomas SG, Phillips AL, Hedden P (2008) The gibberellin biosynthetic genes AtGA20ox1 and AtGA20ox2 act, partially redundantly, to promote growth and development throughout the Arabidopsis life cycle. Plant J 53:488–504
Rolland F, Baena-Gonzalez E, Sheen J (2006) Sugar sensing and signaling in plants: conserved and novel mechanisms. Annu Rev Plant Biol 57:675–709
Stitt M, Lilley RM, Gerhardt R, Heldt HW (1989) Metabolite levels in specific cells and subcellular compartments of plant leaves. Method Enzymol 174:518–552
Sun YW, Jiao GA, Liu ZP, Zhang X, Li JY, Guo XP, Du WM, Du JL, Francis F, Zhao YD, Xia LQ (2017) Generation of high-amylose rice through CRISPR/Cas9-mediated targeted mutagenesis of starch branching enzymes. Front Plant Sci 8:298
Veramendi J, Fernie AR, Leisse A, Willmitzer L, Trethewey RN (2002) Potato hexokinase 2 complements transgenic Arabidopsis plants deficient in hexokinase 1 but does not play a key role in tuber carbohydrate metabolism. Plant Mol Biol 49:491–501
Xu FQ, Li XR, Ruan YL (2008) RNAi-mediated suppression of hexokinase gene OsHXK10 in rice leads to non-dehiscent anther and reduction of pollen germination. Plant Sci 175:674–684
Yamaguchi S (2008) Gibberellin metabolism and its regulation. Annu Re Plant Biol 59:225–251
Yang W, Guo Z, Huang C, Duan L, Chen G, Jiang N, Fang W, Feng H, Xie W, Lian X, Wang G, Luo Q, Zhang Q, Liu Q, Xiong L (2014) Combining high-throughput phenotyping and genome-wide association studies to reveal natural genetic variation in rice. Nat Commun 5:5087
Zhang Y, Su J, Duan S, Ao Y, Dai J, Liu J, Wang P, Li Y, Liu B, Feng D, Wang J, Wang H (2011) A highly efficient rice green tissue protoplast system for transient gene expression and studying light/chloroplast-related processes. Plant Methods 7:30
Zhang H, Zhao Y, Zhou DX (2017) Rice NAD+-dependent histone deacetylase OsSRT1 represses glycolysis and regulates the moonlighting function of GAPDH as a transcriptional activator of glycolytic genes. Nucleic Acids Res 45:12241–12255
Zheng S, Lu J, Yu D, Li J, Zhou H, Jiang D, Liu Z, Zhuang C (2021a) Hexokinase gene OsHXK1 positively regulates leaf senescence in rice. BMC Plant Biol 21:580
Zheng S, Ye C, Lu J, Liufu J, Lin L, Dong Z, Li J, Zhuang C (2021b) Improving the rice photosynthetic efficiency and yield by editing OsHXK1 via CRISPR/Cas9 system. Int J Mol Sci 22:9554
Zuo J, Li J (2014) Molecular genetic dissection of quantitative trait loci regulating rice grain size. Annu Rev Genet 48:99–118
Funding
We thank Dr. Hua Zhang and Chennan Ge for Arabidopsis transformation. This work was supported by grants from the National Natural Science Foundation (U21A20211, 91935303, 31901529), the Ministry of Science and Technology (2021YFF1000200), the science and technology major program of Hubei Province (2021ABA011), the earmarked fund for the Agricultural Research System (CARS-01-01) of China, and the Natural Science Foundation of Anhui Province (2008085QC120).
Author information
Authors and Affiliations
Contributions
YP conducted most of the experiments; LYB isolated the sng1 mutant; WB, ZY and WKY carried out part of the phenotyping and expression analysis; LPB, GGJ, ZQL and LXH participated in the field management and logistic work; YP, LYB, LZF and HYQ designed the study; YP and HYQ analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflict of interest in the reported research.
Ethical approval
The authors note that this research was performed and reported in accordance with the ethical standards for scientific conduct.
Additional information
Communicated by Joshua N. Cobb.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yun, P., Li, Y., Wu, B. et al. OsHXK3 encodes a hexokinase-like protein that positively regulates grain size in rice. Theor Appl Genet 135, 3417–3431 (2022). https://doi.org/10.1007/s00122-022-04189-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-022-04189-7