Abstract
Key message
Leaf width was correlated with plant-level transpiration efficiency and associated with 19 QTL in sorghum, suggesting it could be a surrogate for transpiration efficiency in large breeding program.
Abstract
Enhancing plant transpiration efficiency (TE) by reducing transpiration without compromising photosynthesis and yield is a desirable selection target in crop improvement programs. While narrow individual leaf width has been correlated with greater intrinsic water use efficiency in C4 species, the extent to which this translates to greater plant TE has not been investigated. The aims of this study were to evaluate the correlation of leaf width with TE at the whole-plant scale and investigate the genetic control of leaf width in sorghum. Two lysimetry experiments using 16 genotypes varying for stomatal conductance and three field trials using a large sorghum diversity panel (n = 701 lines) were conducted. Negative associations of leaf width with plant TE were found in the lysimetry experiments, suggesting narrow leaves may result in reduced plant transpiration without trade-offs in biomass accumulation. A wide range in width of the largest leaf was found in the sorghum diversity panel with consistent ranking among sorghum races, suggesting that environmental adaptation may have a role in modifying leaf width. Nineteen QTL were identified by genome-wide association studies on leaf width adjusted for flowering time. The QTL identified showed high levels of correspondence with those in maize and rice, suggesting similarities in the genetic control of leaf width across cereals. Three a priori candidate genes for leaf width, previously found to regulate dorsoventrality, were identified based on a 1-cM threshold. This study provides useful physiological and genetic insights for potential manipulation of leaf width to improve plant adaptation to diverse environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Leaf size is an important determinant of canopy leaf area development, thus influencing crop photosynthesis, transpiration and yield (Duncan et al. 1967; Kholová et al. 2014; Zhu et al. 2012). In sorghum (Sorghum bicolor (L.) Moench) and maize (Zea mays L.), with their typical elongated and narrow monocot leaves, the area of individual leaves is linearly related to the product of length and maximum width (Stickler et al. 1961; Dwyer and Stewart 1986; Birch et al. 1998). For sorghum, each emerging leaf is longer and wider than the previous one, in most cases reaching a maximum around the third or fourth leaf below the flag leaf (Borrell et al. 2014a). Length and width of the largest leaf are indicative of total plant leaf area which, in turn, is associated with the water demand of plants via canopy transpiration (Stickler et al. 1961; Duncan et al. 1967; Borrell et al. 2014b; Kholová et al. 2014; George-Jaeggli et al. 2017). At the whole-plant level, leaf area index is commonly used as a critical variable when simulating photosynthesis- and/or transpiration-related crop productivity in cereals (Duncan et al. 1967; Wu et al. 2018; Chenu et al. 2018; Hammer et al. 2019). In combination with leaf angle, the size of the largest leaf is also a key component of canopy structure and influences light capture and canopy photosynthesis (Duncan 1971; Zhi et al. 2022a; Birch et al. 1998).
At the leaf level, leaf size is correlated with many physiological traits and their interplay can affect functional aspects of leaf gas exchange (Parkhurst and Loucks 1972; Chatterjee et al. 2016; Cano et al. 2019; Pan et al. 2021). As a component of leaf size, leaf width is closely associated with variation in leaf structural properties (e.g. vein density or patterning), which impacts the access of CO2 to chloroplasts and also water transport efficiency (Evans 1999; Giuliani et al. 2013; Baird et al. 2021). Similarly, leaf width in C4 crops has recently been found to be positively correlated with stomatal conductance, while being negatively associated with both mesophyll conductance and intrinsic water use efficiency (iWUE, defined as the ratio of CO2 assimilation rate to stomatal conductance at the leaf segment level) (Cano et al. 2019; Pan et al. 2021). This could be attributed to the negative association between leaf width and vein density, with the latter usually being positively correlated with stomatal density (Fiorin et al. 2016; Baird et al. 2021). Greater interveinal distance (smaller vein density) observed in wider leaves (Baird et al. 2021) results in reduced mesophyll conductance and a greater requirement for increased stomatal conductance to CO2 (greater stomatal opening) to maintain high photosynthetic rates (Pan et al. 2021), which at the same time leads to an increase in transpiration, and hence decreased iWUE (Crookston and Moss 1974; Farquhar and Sharkey 1982; Evans et al. 2009). Wider leaves also have a greater boundary layer resistance, which influences vapour transfer and heat exchange, and induces increased stomatal conductance (Parkhurst and Loucks 1972), which further contributes to reduced iWUE (Baird et al. 2021). A recent detailed study into the effects of leaf width on gas exchange, boundary-layer effects and leaf anatomy using 48 diverse field-grown genotypes in sorghum, corroborated these relationships between leaf width and iWUE (Pan et al. 2021). Therefore, leaf width may represent a potential breeding target to improve adaptation of sorghum to the hotter and drier climates of the future (Haussmann et al. 2012; Menamo et al. 2021), particularly if the leaf-level effects of leaf width on iWUE can scale up to transpiration efficiency (TE) at the plant level. Whole-plant TE has previously been identified as an important target trait for advancing adaptation relevant to future climates (Hammer et al. 2020), and useful genetic variation has been reported in sorghum (Hammer et al. 1997; Geetika et al. 2019; van Oosterom et al. 2021).
In order to efficiently select for optimal leaf width in a plant breeding program, it is important to understand the genetic basis of the trait and its interactions with other traits. The genetic control of leaf width has been investigated in rice (Oryza sativa) (Li et al. 1998; Yue et al. 2006; Zhao et al. 2011; Wang et al. 2012; Tang et al. 2018; Wen et al. 2020) and maize (Tian et al. 2011; Ku et al. 2012; Yang et al. 2016; Zhao et al. 2018; Fu et al. 2019), demonstrating that leaf width is under complex genetic control. For example, a comprehensive maize genome-wide association study (GWAS) using a Nested Association Mapping population (n = 4892) detected 295 significant single nucleotide polymorphisms (SNPs) associated with variation in leaf width, suggesting that leaf width is controlled by multiple QTL, most with effect sizes of less than 3 mm (Tian et al. 2011). In a recent mapping study in rice using a chromosomal segment substitution line population, nine leaf width QTL explaining 55.8% of phenotypic variation were detected (Tang et al. 2018). In addition, a narrow leaf gene (Nal1 in rice and ns1 in maize) was identified, which decreased leaf width in maize (Nardmann et al. 2004) and was associated with increased photosynthetic capacity in rice (Tang et al. 2018). The changes in leaf width of Nal1 mutants were attributed to changes in the number of longitudinal leaf veins in rice (Qi et al. 2008; Takai et al. 2013), indicating a close linkage between leaf width and leaf vein or stomatal features as reported recently (Pan et al. 2021). Despite a number of genetic mapping studies in other cereals and the growing evidence that leaf width and stomatal features are tightly linked (Pan et al. 2021), work in sorghum has previously been limited to a few studies (Feltus et al. 2006; Sakhi et al. 2013; Kapanigowda et al. 2014; Shehzad and Okuno 2015; McCormick 2017). These previous sorghum studies have typically been of small population size (ranging from 107 to 370 individuals) and only detected eleven leaf-width QTL. Therefore, a more comprehensive study was needed to dissect the genetic basis of leaf width in sorghum.
Sorghum, the world’s fifth most important cereal, is morphologically and genetically diverse (Reddy and Patil 2015; Tao et al. 2021a) and can hence be adapted to a wide range of environments. Cultivated sorghum can be classified into five basic races, including guinea, caudatum, kafir and durra, which are traditionally grown in different agro-ecological zones (Harlan and Wet 1972). Evaluating differences in leaf width among the sorghum races might provide information about the importance of the trait for environmental adaptation (Lasky et al. 2015; Menamo et al. 2021) and hence its potential in breeding sorghum varieties with improved adaptation to variable climates (Hammer et al. 2020).
The objectives of the present study were to (i) investigate whether leaf width affects plant-level TE using lysimetry experiments; (ii) quantify the natural variation in leaf width using a sorghum diversity panel; (iii) examine whether there are differences in leaf width among sorghum races to understand its role in adaptation to different environments; and (iv) detect the genetic basis of leaf width using GWAS and identify potential candidate genes.
Materials and methods
Experimental design and genotypes used in lysimetry experiments
Two experiments (Exp1 and Exp2) were conducted using a fully automated lysimetry platform inside a semi-controlled enclosure at the University of Queensland, Gatton Campus (27°33´ S, 152°20´ E) in southeast Queensland, Australia. The automated lysimetry platform has been described previously (Chenu et al. 2018). Briefly, the enclosure has a solar weave roof and meshed poly weave sides, plus a gable fan that provides additional airflow when air temperature inside the enclosure exceeds 38 °C. Temperature and humidity inside the enclosure are automatically recorded every ten minutes (Table S1). Single plants were grown in large pots (at least 50 L capacity) to provide non-stressed growing conditions for sorghum. The amount of water used per plant was monitored every ten minutes by automatic weighing of the pots, which were set up on individual load cells mounted on trolleys. Each trolley held two rows of four pots and trolleys were setup such that the overall experiments had six rows and six columns. Individual pots were watered automatically via a PVC access tube embedded in the soil. In both experiments, pots were rewatered to their initial weight as soon as they dropped below a threshold value of 65 kg, at which time the soil moisture content was at approximately 50% of its drained upper limit.
Six diverse grain sorghum genotypes, which were known to cover a range of stomatal conductance and leaf widths (unpublished data), were grown in Exp1. In Exp2, twelve genotypes were grown with known differences in C4 photosynthetic parameters determined via fitting CO2 response curves (Zhi et al. 2022b), in addition to known variability in leaf width; two genotypes were in common across both experiments (Table S2). Genotypes were assigned to lysimetry pots using a completely randomised block design with six (Exp1) and three (Exp2) replicates. Exp1 was sown on 29 October 2018 and harvested on 9 January 2019 (anthesis) and Exp2 was sown on 25 March 2019 and harvested on 13 May 2019 (prior to anthesis). Before sowing, each pot was lined with a plastic bag to facilitate removal of soil at harvesting. Additionally, a 30-cm PVC tube with a volume of 750 ml was embedded vertically into the soil for watering. Pots were filled with air-dried black vertisol soil from a field near Dalby, a sorghum-growing area in southeast Queensland. For each pot, 20 g of Scotts Osmocote Plus controlled release fertiliser (16%N, 3.5%P, 10%K) (Scotts Australia, Baulkham Hills, NSW, Australia) and 10 g of Dolomite were applied to the top thirty centimetres of soil. Five seeds were sown per lysimeter, and seedlings were gradually thinned until one seedling per pot was left once the plants were well established (17 DAS in Exp1 and 14 DAS in Exp2). At that time, the soil surface of each pot was completely sealed using plastic sheets, leaving only a small opening for the plants, thereby minimising water losses from soil evaporation. One teaspoon of Thrive (Yates Australia, Padstow, NSW, Australia) water soluble fertiliser (25%N, 5%P, 8.8%K) was added fortnightly to the watering tubes to ensure plants were adequately fertilised.
Leaf width and plant water use data collection in lysimetry experiments
The number of fully expanded and senesced leaves on mainstems and tillers were recorded weekly. A leaf was considered fully expanded when its ligule was visible above the ligule of the previous leaf and considered senesced once less than 50% of its area was green. Once fully expanded, each leaf was measured to determine its width (measured across the widest part of the leaf). Water use data was aggregated hourly based on the weight differences of each pot. Water use per plant for the entire experimental period was determined by calculating overall water use of each pot from the time of being sealed until harvest date, including the fresh weight of each plant. At harvest, plants were cut at the base of the stem and dried in a fan-forced dehydrator at 80 °C until a stable dry weight was reached. Plant TE (g kg−1) was taken as final dry mass divided by total water use of each plant.
Experimental design and plant material used in field trials
To explore the genetic basis of leaf width, 701 lines from a sorghum diversity panel were used. The diversity panel has been described previously (Tao et al. 2020) and consists primarily of lines from the Sorghum Conversion Program, in which genes for standard height and maturity have been introgressed into diverse sorghum lines and landraces from all over the world (Rosenow et al. 1997). Three field experiments were conducted during three consecutive summer cropping seasons at Hermitage Research Facility (HRF), Warwick, Queensland, Australia (28°12’S, 152°5’E, 470 m above sea level), and Gatton Research Station (GAT), Gatton, Queensland, Australia (27°33’S, 152°20’E, 94 m above sea level). At HRF, two experiments (HRF1 and HRF2) were sown on 6 December 2016 and 8 January 2018, respectively. The third experiment was sown on 14 January 2019 at GAT. In all experiments, genotypes were assigned to plots using randomised row-column designs with partial replications (around 30% of genotypes being replicated). Each plot planted to one genotype comprised four rows and was 3 m wide and 4.5 m long. Experiments were planted with a target population density of 50,000 plants ha−1 at HRF and 108,000 plants ha−1 at GAT. Experimental areas were fertilised sufficiently to provide nutrient non-limiting conditions. HRF1 and HRF2 were sown on full sub-soil moisture profiles and only irrigated at sowing to enable establishment, whereas GAT was irrigated regularly to minimise the risk of water limitation during summer.
Phenotyping and genotyping in field trials
Around two weeks after flowering, one plant per plot was randomly chosen from the middle two rows to measure leaf width. Measurements were made on the largest mainstem leaf, as the largest leaf has been found to be most closely correlated with total plant leaf area in maize plants (Stickler et al. 1961; Birch et al. 1998). The largest leaf in the genotypes from the diversity panel was found to be generally the third or fourth leaf below the flag leaf (Borrell et al. 2014a), however, the final number of leaves varies depending on maturity of each genotype. Width of the largest leaf was therefore found to be affected by final leaf number, which in turn is largely driven by flowering time in sorghum (Hesketh et al. 1969). Both traits were also measured; the date when more than 50% of the plants in a plot reached anthesis was recorded as days to flower for that plot. Final leaf number was determined by consecutively numbering leaves with a permanent marker as they appeared on one tagged plant per plot. To minimise the influence of flowering time, leaf width of the largest leaf was adjusted for flowering time. Additionally, the association between days to flower and final leaf number for 625 lines was evaluated in HRF2.
All 701 diversity panel entries were genotyped by Diversity Arrays Technology Pty Ltd (https://www.diversityarrays.com/technology-and-resources/dartreseq/). DNA was extracted from young leaf tissue of five plants in each plot using a modified cetyl trimethyl ammonium bromide (CTAB) method (Doyle 1987). The samples were digested with methylation-sensitive restriction enzymes (HpaII, MseI) to remove repetitive sequences prior to sequencing of the most informative representations of genomic DNA on next-generation sequencing platforms (Illumina, HiSeq 2500). The sequence data generated was aligned to version v3.1 of the sorghum reference genome sequence (McCormick et al. 2018) to identify SNPs (Single Nucleotide Polymorphisms).
Statistical analyses
Phenotypic data
All statistical analyses were conducted using the R open-source statistical programming language (Team 2018). The relationship between plant TE and leaf width was evaluated by linear regression in Exp1 and Exp2 (Table S2). Averaged leaf width of leaf 10–12 in Exp1 and that of leaf 7–9 in Exp2 were used to determine the association with plant TE to avoid the confounding effect of total leaf number on the genotypic differences in leaf width, given these leaves were in the middle of increasing leaf size and active vegetation growth.
In the field trials, best linear unbiased predictors (BLUPs) of width of the largest leaf were calculated to minimise environmental effects within and across experiments using a restricted maximum likelihood (REML) by fitting a linear mixed model using the ASReml-R package (equation [1]) (Gilmour et al. 1997; Butler et al. 2018). To remove the variation in leaf width of the largest leaf caused by differences in days to flower which is closely associated with final leaf number, leaf width of the largest leaf adjusted for days to flower of each line was predicted by including days to flower as a fixed effect within each trial in the model [1]. Data from all trials was appended to allow for a multi-environment analysis using the following model.
where the response vector y is modelled by all the fixed effects β, random effects u and all the residual effects ε. The matrix X represents the design matrix for the fixed effects, and the matrix Z is the design matrix for the random effects. The fixed effects were composed of main effects for every trial plus any effects associated with linear changes along the rows and columns for every trial. The random effects contained sources of error within each trial including replication and any trial specific random row and column effects. The residual effects included trial specific residuals effects and first order auto-regressive (AR1) effects in both the row and column directions for each trial. This model included genotype as a random effect to predict genotype BLUPs within trials. The G × E was included in the model with an experiment by genotype interaction with a correlated variance structure with a separate genetic variance for each trial and pairwise correlations between each pair of trials. All possible sources of variation in the BLUPs were allowed for in the linear mixed model (Gilmour et al. 1997). A generalised measure of heritability was calculated due to the complex variance structure of the fitted model. The calculation of generalised heritability for each trial is given by equation [2].
where \({\varvec{H}}^{2}\) is the generalised heritability, \({\varvec{\sigma}}_{{\varvec{g}}}^{2}\) represents the genetic variance and \(\overline{{{\text{SED}}}}\) is the average standard error of difference (Cullis et al. 2006).
The 701 genotypes in the diversity panel were assigned to racial groups, based on population structure results reported previously (Tao et al. 2020). However, the cut-off of genetic identity to select racial representatives (374 genotypes in total) was adjusted to 70% within a racial group for valid inter-group comparisons. The racial differences in leaf width were analysed independently for HRF1, HRF2 and GAT, using Wald tests.
Genome-wide association studies and identification of QTL and candidate genes for leaf width
The GWAS analyses were performed using the software FarmCPU (Liu et al. 2016). A Bonferroni-corrected significant threshold was calculated based on 0.05/number of effective tests (Duggal et al. 2008; Li et al. 2012, 2021). The significant SNPs detected across locations were further clustered into unique QTL based on a 1-cM window, in accordance with the linkage disequilibrium decay previously identified in the diversity panel (Tao et al. 2020), 200 kb, which is equivalent to ~ 1 cM (Mace and Jordan 2011).
Leaf width QTL previously identified in sorghum (Feltus et al. 2006; Sakhi et al. 2013; Kapanigowda et al. 2014; Shehzad and Okuno 2015; McCormick 2017) were compared with the QTL identified in the current study, using the predicted locations of the QTL CI from the Sorghum QTL Atlas (Mace et al. 2019). In addition, inter-species QTL comparisons for leaf width were conducted, specifically in maize (Tian et al. 2011) and rice (Tang et al. 2018), where the maize and rice QTL were projected onto the sorghum consensus map following the same methodology as described previously (Mace et al. 2019). Thirty-two sorghum orthologues of a priori candidate genes (Table S3), compiled previously (Tian et al. 2011), were used to evaluate the proximity of the QTL detected in the sorghum diversity panel based on 1 cM window. To further validate the function of a priori candidate genes, haplotype analysis (Minor allele frequency > 5%) was conducted in R using packages “ape” (Paradis et al. 2004) and “pegas” (Paradis 2010), as previously described (Tao et al. 2021b).
Results
Correlation between plant water use efficiency (TE) and leaf width in the lysimetry experiments
In the semi-controlled lysimetry experiments, a negative correlation between plant TE and average leaf width (across leaves 10–12 and leaves 7–9 in Exp1 and Exp2, respectively) was observed in both Exp1 and Exp2 (Fig. 1). This correlation was particularly strong and significant among the twelve genotypes in Exp2 (Fig. 1B: R2 = 0.6, p = 0.003). The correlation was not significant across the six genotypes in Exp1 (Fig. 1A: R2 = 0.23, p = 0.33); however, a consistent trend was shown in the regression of plant TE versus leaf width.
Relationship between plant transpiration efficiency and leaf width across genotypes in Exp1 (A) and Exp2 (B) Note: leaf width: the mean of leaves 10–12 in Exp1 (A) and leaves 7–9 in Exp2 (B); R2: coefficient of determination of the relationship; p: significance level; error bars on each data point indicate standard errors of plant transpiration efficiency and leaf width
Association of leaf width with days to flower in the sorghum diversity panel
To evaluate the effect of flowering time on width of the largest leaf, correlation analysis was conducted among leaf width, days to flower and final leaf number in HRF2 (n = 625 lines). Days to flower and final leaf number were highly correlated with each other as expected (r = 0.58). Significant correlations of width of the largest leaf with days to flower (r = 0.32) and final leaf number (r = 0.29) were also found, suggesting that about 10% of variation in width of the largest leaf could be driven by differences in flowering time (Fig. 2). Therefore, to minimise the confounding effects of differences in flowering time on width of the largest leaf, GWAS analyses were performed based on BLUPs of leaf width adjusted for days to flower.
Correlation among leaf width of the largest leaf within a plant (mm), days to flower and final leaf number in HRF2 (n = 625 lines) Note: Data shown is leaf width, days to flower and final leaf number BLUPs. Leaf width in mm is shown on the x-axes of all three panels on the left, days to flower on the panels in the middle and final leaf number on the panels on the right. Final leaf number is shown on the y-axes of all panels in the bottom row, days to flower on the panels in the middle and leaf width in mm on all panels in the top row. The variable names are displayed on the outer edges of the matrix. The boxes along the diagonals display the density plot for each variable. The boxes in the lower left corner display the scatterplot between each variable. The boxes in the upper right corner display the Pearson correlation coefficient between each variable plus significance levels as stars (***, **, * correspond to p < 0.001, 0.01, 0.05, respectively). Axis labels are for the bottom and left panels
Phenotypic variation in leaf width in the sorghum diversity panel
Leaf width varied from 43.1 to 116.8 mm across three trials (Table 1). Strong correlations (~ 0.80) among trials and high heritability (on average 0.85) were observed, suggesting high levels of genetic effects on leaf width across trials with low levels of interactions between genotypes and environments (G × E) (Table 1). This suggests that a large proportion of leaf width variation is attributed to genetic differences among sorghum lines in the diversity panel (Table 1).
Differences in leaf width among sorghum races in the diversity panel
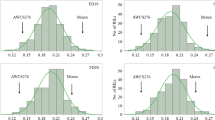
Lines from the diversity panel used in these trials were assigned to five racial groups. Significant differences in leaf width among these groups were observed and groups ranked consistently across the three field trials (Fig. 3, p < 0.001, Wald tests). Racial groups explained ~ 70% of the observed variation in leaf width. The guinea racial types exhibited significantly narrower leaves compared with the remaining four racial groups in all trials, while the East African durra types showed wider leaves than the other racial types (Fig. 3).
Leaf width for genotypes assigned to five sorghum racial groups in the trials of HRF1 (A), HRF2 (B) and GAT (C) Note: Data shown is leaf width BLUPs of the largest leaf adjusted for days to flower; Min: the minimum leaf width; Max: the maximum leaf width; Mean: the mean leaf width; std.error: standard error; H2: generalised heritability; HRF1: diversity panel grown at Hermitage Research Facility in Warwick QLD in 2017; HRF2: the diversity panel grown at Hermitage Research Facility in Warwick QLD in 2018; GAT: the diversity panel grown at Gatton Research Facility in Gatton, QLD in 2019
GWAS for leaf width in the diversity panel
In total, 414,899 SNPs were identified in the diversity panel and genetic (centimorgan, cM) positions of the SNPs were predicted using a sorghum consensus map (Mace et al. 2019). The GWAS analyses were performed with 354,715 SNPs with minor allele frequency (MAF) > 0.01 and a Bonferroni-corrected significant threshold of p < 2e-6. Using BLUPs for leaf width of the largest leaf adjusted for days to flower in HRF1, HRF2 and GAT, 20 significant SNPs were detected (Fig. 4), resulting in 19 leaf width QTL based on a 1-cM window (Table 2 and Table S4). The leaf width QTL were distributed across all ten chromosomes in sorghum. Eleven leaf width QTL have been identified in sorghum from five previous studies (Feltus et al. 2006; Sakhi et al. 2013; Kapanigowda et al. 2014; Shehzad and Okuno 2015; McCormick 2017), which were conducted in relatively small populations incorporating 107 to 370 individuals. Based on a 1-cM overlap window (Table 2 and Table S4), three (16%) QTL in the diversity panel were found to be co-located with leaf width QTL identified previously, representing a significant enrichment (p < 0.05, Chi-square test). The remaining 16 were firstly reported in this study, representing additional genomic regions controlling leaf width that are available for further studies.
Manhattan plot of leaf width in HRF1, HRF2 and GAT Note: HRF1: diversity panel grown at Hermitage Research Facility in Warwick QLD in 2017; HRF2: diversity panel grown at Hermitage Research Facility in Warwick QLD in 2018; GAT: diversity panel grown at Gatton Research Facility in Gatton, QLD in 2019; the SNPs in red are significant ones detected using the p-value < 2e-6
In addition, compared with projected locations of previously identified maize leaf width QTL (Tian et al. 2011) on the sorghum consensus map (Mace et al. 2019), eight (42%) sorghum QTL overlapped with maize QTL based on a 1 cM (Table 2 and Table S4). This represented a significant enrichment with maize leaf width QTL (p < 0.05, Chi-square test). Comparing the sorghum QTL for leaf width identified in this study with the nine leaf width QTL identified from a recent rice study (Tang et al. 2018), four (21%) were less than 5 cM from the projected locations of leaf width QTL in rice (Tang et al. 2018; Mace et al. 2019) (Table 2 and Table S4). Enrichment analysis found that the level of correspondence was significant (p < 0.05, Chi-square test), based on a 5-cM window, indicating similarities between the genetic bases of leaf width in sorghum and rice.
A priori candidate genes for leaf width in the diversity panel
In total, three a priori candidate genes overlapped with leaf width QTL identified in this study based on a 1-cM threshold (Table 3). These three a priori candidate genes have been reported to be involved in leaf patterning either via cell specification and differentiation, such as abaxial or adaxial cell specification, or expansion and proliferation of cell groups that affect leaf architectural traits including leaf length and width (Nogueira et al. 2007; Strable et al. 2017; Zhong et al. 2021).
To further explore effect of the three candidate genes on leaf width, haplotype analysis was conducted. Two major haplotypes were identified in Sobic.008G070600 (homolog of leafbladeless1 gene (lbl, GRMZM2G020187), co-located with qLW_dtf8.1 based on 0 cM), with haplotype I comprising mainly caudatum types and haplotype II comprising mainly Asian durra types (Fig. 5A). Significant difference in leaf width between these two major haplotypes was detected (Fig. 5B), consistent with the role of Sobic.008G070600 in controlling leaf width. Moreover, a haplotype analysis for another a priori candidate genes, Sobic.001G199200 (homolog of yab11 (GRMZM2G167824) gene, 0.1 cM away from qLW_dtf1.1), showed five major haplotypes (Fig. 5C). Haplotype I mainly comprised caudatum types and haplotype II dominated in guinea types. Leaf width was significantly different among these five major haplotypes of Sobic.001G199200 (Fig. 5 D). Regarding to Sobic.003G298600 (0.9 cM away from qLW_dtf3.1), no SNPs were detected within the gene. In summary, the haplotype analyses for Sobic.008G070600 and Sobic.001G199200 further supported their strong association with leaf width in sorghum.
Haplotype network of Sb008G070600 (A) and Sb001G199200 (C), and boxplots showing effects of major haplotypes of Sb008G070600 (B) and Sb001G199200 (D) on leaf width Note: P value in plot B indicates the difference in leaf width between two haplotypes by t-test; different letters over the boxes in plot D mean statistically significant differences in leaf width determined through Tukey-pairwise comparison among the five major haplotypes of Sb001G199200
Discussion
Leaf width is suggested to be an important component trait affecting plant-level TE in sorghum, as indicated by the negative association of plant-level TE with leaf width in both lysimetry experiments in this study. This has not been previously reported. The wide range of leaf widths observed in the diversity panel facilitated the dissection of the underpinning genetic control via GWAS. In addition, consistent and significant differences in leaf width among the five sorghum races suggested an association of leaf width with environmental adaptation. Across the three trials, 19 unique leaf width QTL were identified with high levels of overlap among leaf width QTL in sorghum, maize (Tian et al. 2011) and rice (Tang et al. 2018). This suggests likely similarities in the genetic basis of leaf width in cereals. Furthermore, three a priori candidate genes were confirmed based on a 1-cM threshold, which were characterized as regulating leaf patterning including dorsoventrality and expansion of the leaf blade.
Leaf width was negatively associated with plant-level TE
Leaf width in C4 species has recently been found to be positively associated with stomatal conductance, while being negatively correlated with water use efficiency at the leaf level (iWUE) (Cano et al. 2019; Pan et al. 2021). However, it has not previously been reported whether this relationship scales up to the plant level in sorghum. In this study, a negative association between whole-plant TE and leaf width was shown in two lysimetry experiments, suggesting transpiration losses could potentially be reduced via decreased leaf width without trade-offs in biomass. While the association was moderately strong and significant in Exp2 which comprised twelve genotypes, the correlation between leaf width and plant TE was not significant in Exp1 (in which six genotypes were tested). However, the direction was consistent across both experiments. This is a significant finding, particularly since the negative relationship between water use efficiency and leaf width has only been investigated at the leaf level (Cano et al. 2019; Pan et al. 2021). Hence, dissecting the genetic basis of leaf width might be beneficial for improving water use efficiency in breeding, particularly, given the fact that small differences in plant-level TE are likely to have major effects on crop yield and adaptation (Hammer et al. 2020). However, a follow-up study is needed to further explore the extent of the relationship between leaf width and plant-level TE.
One of the underlying mechanisms associated with the negative relationship between water use efficiency and leaf width may be the strongly negative relationship between leaf width and the 1st and 2nd order vein density in C4 species (Baird et al. 2021). Narrow leaves are associated with high vein density and, in turn, with high density of bundle sheath cells, which increases the capacity for CO2 concentration and hence rates of photosynthetic assimilation (Dengler et al. 1994; Christin et al. 2013). Moreover, high vein density with small interveinal distance results in reduced stomata opening and thus lower stomatal conductance which, in turn, contributes to high iWUE (Pan et al. 2021). Apart from the low stomatal density, wide leaves are also associated with a thicker boundary layer, which reduces vapour flux from the leaf to the ambient air, thereby reducing stomatal resistance compared to narrower leaves with lower boundary layer resistance. The consequent slight increase in leaf temperature causes higher internal vapour pressure, which enhances transpiration. This reduces iWUE in the wider leaf type, especially at low wind speeds (Parkhurst and Loucks 1972). Narrow leaf types with thin boundary layers, which facilitate more direct vapour and heat exchange between leaves and the atmosphere, more easily maintain leaf temperature close to ambient air temperature while having higher stomatal resistance (Nobel 1999). Additionally, this association between leaf width and leaf temperature may be important to photosynthetic performance by preventing leaves from reaching damagingly high temperatures (Gates 1968; Parkhurst and Loucks 1972; Moore et al. 2021). In summary, the negative linkage between leaf width and water use efficiency (both iWUE and plant-level TE) could be attributed partly to its coordination with vein density and boundary layer, both of which are associated with stomatal features (e.g. density and opening) and thus with water use efficiency (Pan et al. 2021; Baird et al. 2021).
Using leaf width in breeding to improve plant water use efficiency in different environments
The large natural variation, relative ease of measurement and high heritability of leaf width make it amenable to selection in breeding programs, as a potential surrogate for transpiration efficiency. However, the complex genetic basis underpinned by multiple small-effect QTL makes marker-assisted selection more difficult. A breeding approach using whole-genome information (e.g. genomic selection) could be an effective strategy (Fernandes et al. 2018). The complex genetic architecture and existence of many small-effect QTL also indicate that large populations are required to ensure sufficient power to identify the majority of QTL by GWAS. The fact that only eleven leaf width QTL have previously been detected is partly due to the small population sizes used (Feltus et al. 2006; Sakhi et al. 2013; Kapanigowda et al. 2014; Shehzad and Okuno 2015; McCormick 2017). The larger population used in the current study enabled detection of additional QTL and improved dissection of the genetic basis of leaf width in sorghum. The cross-species comparisons of leaf width QTL suggest conserved genetic control of leaf width across cereals. This highlights the important information that can be provided by using sorghum, which is genetically diverse and adapted to a range of environments, as a model for the genetic dissection of traits that will contribute to greater adaptation of cereals to future climates and a range of production systems.
Furthermore, the consistent differences in leaf width among sorghum races found in this study potentially reflects specific adaptation to their agro-ecological environments of origin (Morris et al. 2013; Menamo et al. 2021). However, given the interactions and linkages between leaf width, stomatal conductance, photosynthesis and water use efficiency, the extent to which there is any advantage in water use efficiency relative to leaf width will depend on the canopy and environment contexts, especially with regard to water availability (Tsukaya 2004). Under low resource availability, narrower leaves associated with higher vein density and thinner boundary layers, are expected to provide advantages in water use efficiency, as suggested in the present study. Narrow leaf types would also be advantageous in dry conditions because of reduced total plant leaf area, and therefore lower canopy transpiration over time, provided that narrow leaf width is not accompanied by a greater number of leaves or increased tiller number (Tardieu et al. 2018; Hammer et al. 2019). In addition to the reduced transpiration at both leaf and plant levels, narrow leaves might also be coordinated with leaf angle to improve radiation penetration throughout the canopy in water-stressed environments (Duncan 1971; Witkowski and Lamont 1991). In the absence of water limitation, the situation likely differs. At the leaf level, wider leaves with greater interveinal distance will have greater stomatal aperture, increased transpiration, and decreased water use efficiency (Baldocchi et al. 1985; Pan et al. 2021; Baird et al. 2021). However, at the canopy level, in such potential growth conditions, the greater stomatal conductance is conducive to enhanced photosynthesis and crop growth. A simulation analysis for sorghum quantified this trade-off and indicated a yield disadvantage of reduced plant-level TE in high-yielding environments (Hammer et al. 2019). Hence, the optimal leaf width is dependent on the environmental context.
Regulating dorsoventrality may play a role in varying leaf width
The a priori candidate genes identified in the diversity panel, based on a 1-cM threshold, have previously been found to regulate dorsoventrality (Nogueira et al. 2007; Strable et al. 2017; Zhong et al. 2021). The dorsoventral axis is a prerequisite to lateral outgrowth of the leaf lamina thus playing an important role in establishing leaf blades (Waites and Hudson 1995), resulting in the adaxial surface for light capture and the abaxial surface for gas exchange (Tsukaya 2004; Yang and Hwa 2008). One of the a priori candidate genes identified here, Sobic.008G070600 the orthologue of maize leafbladeless1 (lbl1, GRMZM2G020187), has been shown to specify adaxial and abaxial organ polarity (Nogueira et al. 2007). The recessive mutations of lbl1, mainly expressed in the shoot apical meristem, vasculature and adaxially in leaf primordia, exhibited alteration in abaxialization and width of leaves (Nogueira et al. 2007). It has also been reported that KANADI and YABBY gene families regulate leaf width through controlling abaxialization (Candela et al. 2008; Ku et al. 2012). In the current study, Sobic.001G199200, orthologous to GRMZM2G167824, which is one of the YABBY family genes (Strable et al. 2017), was identified to be 0.1 cM away from qLW_dtf1.1. GRMZM2G167824 has been shown to affect leaf width likely through regulating cell specification and differentiation during leaf development in maize (Strable et al. 2017). Moreover, Sobic.003G298600, the orthologue of GRMZM5G874163, has been reported to control leaf width and regulate the formation of distal tissues by coordinating with KANAD1 family genes in Arabidopsis (Arabidopsis thaliana) as orthologue of AT5G60450 (Pekker et al. 2005). In summary, processes associated with dorsoventrality are likely one of the molecular mechanisms underlying leaf width regulation in sorghum.
Conclusions
The negative association between whole-plant TE and leaf width suggests that the effect of leaf width on iWUE at the leaf level could translate to the whole-plant level. The wide range and consistent differences in leaf width among the five sorghum races suggest that it might be associated with environmental adaptation. The QTL and candidate genes for leaf width identified in the diversity panel provide valuable information, facilitating modification of leaf width to improve plant TE and canopy characteristics to improve cereal adaptation to different agro-ecological environments and changing climates.
Data availability
The data used in the current study is available from the corresponding author upon request.
References
Baird AS, Taylor SH, Pasquet-Kok J et al (2021) Developmental and biophysical determinants of grass leaf size worldwide. Nature 592:242–247. https://doi.org/10.1038/s41586-021-03370-0
Baldocchi DD, Verma SB, Rosenberg NJ et al (1985) Microclimate-plant architectural interactions: Influence of leaf width on the mass and energy exchange of a soybean canopy. Agric for Meteorol 35:1–20. https://doi.org/10.1016/0168-1923(85)90070-X
Birch CJ, Hammer GL, Rickert KG (1998) Improved methods for predicting individual leaf area and leaf senescence in maize (Zea mays). Aust J Agric Res 49:249. https://doi.org/10.1071/A97010
Borrell AK, Mullet JE, George-Jaeggli B et al (2014a) Drought adaptation of stay-green sorghum is associated with canopy development, leaf anatomy, root growth, and water uptake. J Exp Bot 65:6251–6263. https://doi.org/10.1093/jxb/eru232
Borrell AK, van Oosterom EJ, Mullet JE et al (2014b) Stay-green alleles individually enhance grain yield in sorghum under drought by modifying canopy development and water uptake patterns. New Phytol 203:817–830. https://doi.org/10.1111/nph.12869
Butler DG, Cullis BR, Gilmour AR, Gogel BJ (2018) ASReml‐R 4 reference manual: Mixed models for S language environments: Queensland Department of Primary Industries and Fisheries
Candela H, Johnston R, Gerhold A et al (2008) The milkweed pod1 Gene Encodes a KANADI Protein That Is Required for Abaxial/Adaxial Patterning in Maize Leaves. Plant Cell 20:2073–2087. https://doi.org/10.1105/tpc.108.059709
Cano FJ, Sharwood RE, Cousins AB, Ghannoum O (2019) The role of leaf width and conductances to CO2 in determining water use efficiency in C4 grasses. New Phytol 223:1280–1295. https://doi.org/10.1111/nph.15920
Chatterjee J, Dionora J, Elmido-Mabilangan A et al (2016) The evolutionary basis of naturally diverse rice leaves anatomy. PLoS ONE 11:e0164532. https://doi.org/10.1371/journal.pone.0164532
Chenu K, Van Oosterom EJ, McLean G et al (2018) Integrating modelling and phenotyping approaches to identify and screen complex traits: transpiration efficiency in cereals. J Exp Bot 69:3181–3194. https://doi.org/10.1093/jxb/ery059
Christin P-A, Osborne CP, Chatelet DS et al (2013) Anatomical enablers and the evolution of C4 photosynthesis in grasses. Proc Natl Acad Sci 110:1381–1386. https://doi.org/10.1073/pnas.1216777110
Crookston RK, Moss DN (1974) Interveinal distance for carbohydrate transport in leaves of C3 and C4 grasses. Crop Sci 14(1):123. https://doi.org/10.2135/cropsci1974.0011183X001400010038x
Cullis BR, Smith AB, Coombes NE (2006) On the design of early generation variety trials with correlated data. J Agric Biol Environ Stat 11:381–393
Dengler NG, Dengler RE, Donnelly PM, Hattersley PW (1994) Quantitative leaf anatomy of C3 and C4 grasses (Poaceae): bundle sheath and mesophyll surface area relationships. Ann Bot 73:241–255. https://doi.org/10.1006/anbo.1994.1029
Doyle JJ, J. L. (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull
Duggal P, Gillanders EM, Holmes TN, Bailey-Wilson JE (2008) Establishing an adjusted p-value threshold to control the family-wide type 1 error in genome wide association studies. BMC Genomics 9:516. https://doi.org/10.1186/1471-2164-9-516
Duncan WG (1971) Leaf angles, leaf area, and canopy photosynthesis. Crop Sci 11:482–485. https://doi.org/10.2135/cropsci1971.0011183X001100040006x
Duncan W, Loomis R, Williams W, Hanau R (1967) A model for simulating photosynthesis in plant communities. Hilgardia 38:181–205
Dwyer LM, Stewart DW (1986) Leaf area development in field-grown maize. Agron J 78:334–343. https://doi.org/10.2134/agronj1986.00021962007800020024x
Evans JR (1999) Leaf anatomy enables more equal access to light and CO2 between chloroplasts. New Phytol 143:93–104. https://doi.org/10.1046/j.1469-8137.1999.00440.x
Evans JR, Kaldenhoff R, Genty B, Terashima I (2009) Resistances along the CO2 diffusion pathway inside leaves. J Exp Bot 60:2235–2248. https://doi.org/10.1093/jxb/erp117
Farquhar GD, Sharkey TD (1982) Stomatal conductance and photosynthesis. Annu Rev Plant Physiol 33:317–345. https://doi.org/10.1146/annurev.pp.33.060182.001533
Feltus FA, Hart GE, Schertz KF et al (2006) Alignment of genetic maps and QTLs between inter- and intra-specific sorghum populations. Theor Appl Genet 112:1295. https://doi.org/10.1007/s00122-006-0232-3
Fernandes SB, Dias KOG, Ferreira DF, Brown PJ (2018) Efficiency of multi-trait, indirect, and trait-assisted genomic selection for improvement of biomass sorghum. Theor Appl Genet 131:747–755. https://doi.org/10.1007/s00122-017-3033-y
Fiorin L, Brodribb TJ, Anfodillo T (2016) Transport efficiency through uniformity: organization of veins and stomata in angiosperm leaves. New Phytol 209:216–227. https://doi.org/10.1111/nph.13577
Fu Y, Xu G, Chen H et al (2019) QTL mapping for leaf morphology traits in a large maize-teosinte population. Mol Breed 39:103. https://doi.org/10.1007/s11032-019-1012-5
Gates DM (1968) Transpiration and leaf temperature. Annu Rev Plant Physiol 19:211–238. https://doi.org/10.1146/annurev.pp.19.060168.001235
Geetika G, van Oosterom EJ, George-Jaeggli B et al (2019) Genotypic variation in whole-plant transpiration efficiency in sorghum only partly aligns with variation in stomatal conductance. Funct Plant Biol 46:1072–1089
George-Jaeggli B, Mortlock MY, Borrell AK (2017) Bigger is not always better: Reducing leaf area helps stay-green sorghum use soil water more slowly. Environ Exp Bot 138:119–129. https://doi.org/10.1016/j.envexpbot.2017.03.002
Gilmour AR, Cullis BR, Verbyla AP (1997) Accounting for natural and extraneous variation in the analysis of field experiments. J Agric Biol Environ Stat 2:269–293. https://doi.org/10.2307/1400446
Giuliani R, Koteyeva N, Voznesenskaya E et al (2013) Coordination of leaf photosynthesis, transpiration, and structural traits in rice and wild relatives (Genus Oryza). Plant Physiol 162:1632–1651. https://doi.org/10.1104/pp.113.217497
Hammer GL, Farquhar GD, Broad IJ (1997) On the extent of genetic variation for transpiration efficiency in sorghum. Aust J Agric Res 48:649–656. https://doi.org/10.1071/a96111
Hammer GL, McLean G, van Oosterom E et al (2020) Designing crops for adaptation to the drought and high-temperature risks anticipated in future climates. Crop Sci 60:605–621. https://doi.org/10.1002/csc2.20110
Hammer G, McLean G, Doherty A, et al (2019) Sorghum crop modeling and Its utility in agronomy and breeding. In: Sorghum. John Wiley & Sons, Ltd, 215–239
Harlan JR, de Wet JMJ (1972) A simplified classification of cultivated sorghum. Crop Sci 12(2):172–176. https://doi.org/10.2135/cropsci1972.0011183X001200020005x
Haussmann BIG, Rattunde HF, Weltzien-Rattunde E et al (2012) Breeding strategies for adaptation of pearl millet and sorghum to climate variability and change in West Africa. J Agron Crop Sci 198:327–339. https://doi.org/10.1111/j.1439-037X.2012.00526.x
Hesketh JD, Chase SS, Nanda DK (1969) Environmental and genetic modification of leaf number in maize, sorghum, and hungarian millet. Crop Sci 9(4):460. https://doi.org/10.2135/cropsci1969.0011183X000900040022x
Kapanigowda MH, Payne WA, Rooney WL et al (2014) Quantitative trait locus mapping of the transpiration ratio related to preflowering drought tolerance in sorghum (Sorghum bicolor). Funct Plant Biol 41:1049–1065. https://doi.org/10.1071/FP13363
Kholová J, Murugesan T, Kaliamoorthy S et al (2014) Modelling the effect of plant water use traits on yield and stay-green expression in sorghum. Funct Plant Biol 41:1019–1034. https://doi.org/10.1071/FP13355
Ku LX, Zhang J, Guo SL et al (2012) Integrated multiple population analysis of leaf architecture traits in maize (Zea mays L.). J Exp Bot 63:261–274. https://doi.org/10.1093/jxb/err277
Lasky JR, Upadhyaya HD, Ramu P et al (2015) Genome-environment associations in sorghum landraces predict adaptive traits. Sci Adv 1:e1400218. https://doi.org/10.1126/sciadv.1400218
Li Z, Pinson SRM, Stansel JW, Paterson AH (1998) Genetic dissection of the source-sink relationship affecting fecundity and yield in rice (shape Oryza sativa L.). Mol Breed 4:419–426. https://doi.org/10.1023/A:1009608128785
Li M-X, Yeung JMY, Cherny SS, Sham PC (2012) Evaluating the effective numbers of independent tests and significant p -value thresholds in commercial genotyping arrays and public imputation reference datasets. Hum Genet 131:747–756. https://doi.org/10.1007/s00439-011-1118-2
Li L, Zhang C, Huang J et al (2021) Genomic analyses reveal the genetic basis of early maturity and identification of loci and candidate genes in upland cotton (Gossypium hirsutum L.). Plant Biotechnol J 19:109–123. https://doi.org/10.1111/pbi.13446
Liu X, Huang M, Fan B et al (2016) Iterative usage of fixed and random effect models for powerful and efficient genome-wide association studies. PLoS Genet 12:e1005767. https://doi.org/10.1371/journal.pgen.1005767
Mace ES, Jordan DR (2011) Integrating sorghum whole genome sequence information with a compendium of sorghum QTL studies reveals uneven distribution of QTL and of gene-rich regions with significant implications for crop improvement. Theor Appl Genet 123:169
Mace ES, Innes D, Hunt C et al (2019) The sorghum QTL atlas: a powerful tool for trait dissection, comparative genomics and crop improvement. Theor Appl Genet 132:751–766. https://doi.org/10.1007/s00122-018-3212-5
McCormick RF, Truong SK, Sreedasyam A et al (2018) The Sorghum bicolor reference genome: improved assembly, gene annotations, a transcriptome atlas, and signatures of genome organization. Plant J 93:338–354. https://doi.org/10.1111/tpj.13781
McCormick RF (2017) High-throughput genotyping analyses and image-based phenotyping in sorghum bicolor. Thesis, Texas A & M University
Menamo T, Kassahun B, Borrell AK et al (2021) Genetic diversity of Ethiopian sorghum reveals signatures of climatic adaptation. Theor Appl Genet 134:731–742
Moore CE, Meacham-Hensold K, Lemonnier P et al (2021) The effect of increasing temperature on crop photosynthesis: from enzymes to ecosystems. J Exp Bot 72:2822–2844. https://doi.org/10.1093/jxb/erab090
Morris GP, Ramu P, Deshpande SP et al (2013) Population genomic and genome-wide association studies of agroclimatic traits in sorghum. Proc Natl Acad Sci 110:453–458. https://doi.org/10.1073/pnas.1215985110
Nardmann J, Ji J, Werr W, Scanlon MJ (2004) The maize duplicate genes narrow sheath1 and narrow sheath2 encode a conserved homeobox gene function in a lateral domain of shoot apical meristems. Development 131:2827–2839. https://doi.org/10.1242/dev.01164
Nobel PS (1999) Physicochemical & environmental plant physiology. Academic Press
Nogueira FTS, Madi S, Chitwood DH et al (2007) Two small regulatory RNAs establish opposing fates of a developmental axis. Genes Dev 21:750–755. https://doi.org/10.1101/gad.1528607
Pan L, George-Jaeggli B, Borrell A, et al (2021) Coordination of stomata and vein patterns with leaf width underpins water use efficiency in a C4 crop. Authorea
Paradis E (2010) pegas: an r package for population genetics with an integrated–modular approach. Bioinformatics 26:419–420. https://doi.org/10.1093/bioinformatics/btp696
Paradis E, Claude J, Strimmer K (2004) APE: Analyses of phylogenetics and evolution in r language. Bioinformatics 20:289–290. https://doi.org/10.1093/bioinformatics/btg412
Parkhurst DF, Loucks OL (1972) Optimal leaf size in relation to environment. J Ecol 60:505–537. https://doi.org/10.2307/2258359
Pekker I, Alvarez JP, Eshed Y (2005) Auxin response factors mediate Arabidopsis organ asymmetry via modulation of KANADI activity. Plant Cell 17:2899–2910. https://doi.org/10.1105/tpc.105.034876
Qi J, Qian Q, Bu Q et al (2008) Mutation of the rice narrow leaf1 gene, which encodes a novel protein, affects vein patterning and polar auxin transport. Plant Physiol 147:1947–1959. https://doi.org/10.1104/pp.108.118778
Reddy S, Patil JV (2015) Genetic Enhancement of Rabi Sorghum: Adapting the Indian Durras. Academic Press
Rosenow DT, Dahlberg JA, Stephens JC et al (1997) Registration of 63 converted sorghum germplasm lines from the sorghum conversion program. Crop Sci 37:1399–1400. https://doi.org/10.2135/cropsci1997.0011183X003700040090x
Sakhi S, Shehzad T, Rehman S, Okuno K (2013) Mapping the QTLs underlying drought stress at developmental stage of sorghum (Sorghum bicolor (L.) Moench) by association analysis. Euphytica 193:433–450. https://doi.org/10.1007/s10681-013-0963-6
Shehzad T, Okuno K (2015) QTL mapping for yield and yield-contributing traits in sorghum (Sorghum bicolor (L.) Moench) with genome-based SSR markers. Euphytica 203:17–31. https://doi.org/10.1007/s10681-014-1243-9
Stickler FC, Wearden S, Pauli AW (1961) Leaf area determination in grain sorghum. Agron J 53:187–188. https://doi.org/10.2134/agronj1961.00021962005300030018x
Strable J, Wallace JG, Unger-Wallace E et al (2017) Maize YABBY genes drooping leaf1 and drooping leaf2 regulate plant architecture. Plant Cell 29:1622–1641. https://doi.org/10.1105/tpc.16.00477
Takai T, Adachi S, Taguchi-Shiobara F et al (2013) A natural variant of NAL1, selected in high-yield rice breeding programs, pleiotropically increases photosynthesis rate. Sci Rep 3:2149. https://doi.org/10.1038/srep02149
Tang X, Gong R, Sun W et al (2018) Genetic dissection and validation of candidate genes for flag leaf size in rice (Oryza sativa L.). Theor Appl Genet 131:801–815. https://doi.org/10.1007/s00122-017-3036-8
Tao Y, Zhao X, Wang X et al (2020) Large-scale GWAS in sorghum reveals common genetic control of grain size among cereals. Plant Biotechnol J 18:1093–1105. https://doi.org/10.1111/pbi.13284
Tao Y, Luo H, Xu J et al (2021a) Extensive variation within the pan-genome of cultivated and wild sorghum. Nat Plants 7:766–773. https://doi.org/10.1038/s41477-021-00925-x
Tao Y, Trusov Y, Zhao X et al (2021b) Manipulating assimilate availability provides insight into the genes controlling grain size in sorghum. Plant J 108:231–243. https://doi.org/10.1111/tpj.15437
Tardieu F, Simonneau T, Muller B (2018) The physiological basis of drought tolerance in crop plants: a scenario-dependent probabilistic approach. Annu Rev Plant Biol 69:733–759. https://doi.org/10.1146/annurev-arplant-042817-040218
Team RC (2018) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. http s. Www R-Proje Ct Org
Tian F, Bradbury PJ, Brown PJ et al (2011) Genome-wide association study of leaf architecture in the maize nested association mapping population. Nat Genet 43:159–162. https://doi.org/10.1038/ng.746
Tsukaya H (2004) Leaf shape: genetic controls and environmental factors. Int J Dev Biol 49:547–555. https://doi.org/10.1387/ijdb.041921ht
van Oosterom EJ, Kulathunga MRDL, Deifel KS et al (2021) Dissecting and modelling the comparative adaptation to water limitation of sorghum and maize: role of transpiration efficiency, transpiration rate and height. Silico Plants. https://doi.org/10.1093/insilicoplants/diaa012
Waites R, Hudson A (1995) phantastica: a gene required for dorsoventrality of leaves in Antirrhinum majus. Development 121:2143–2154. https://doi.org/10.1242/dev.121.7.2143
Wang P, Zhou G, Cui K et al (2012) Clustered QTL for source leaf size and yield traits in rice (Oryza sativa L.). Mol Breed 29:99–113. https://doi.org/10.1007/s11032-010-9529-7
Wen Y, Fang Y, Hu P et al (2020) Construction of a high-density genetic map Based on SLAF markers and QTL analysis of leaf size in rice. Front Plant Sci 11:1143. https://doi.org/10.3389/fpls.2020.01143
Witkowski ETF, Lamont BB (1991) Leaf specific mass confounds leaf density and thickness. Oecologia 88:486–493
Wu A, Doherty A, Farquhar GD, Hammer GL (2018) Simulating daily field crop canopy photosynthesis: an integrated software package. Funct Plant Biol 45:362–377. https://doi.org/10.1071/FP17225
Yang X-C, Hwa C-M (2008) Genetic modification of plant architecture and variety improvement in rice. Heredity 101:396–404. https://doi.org/10.1038/hdy.2008.90
Yang C, Tang D, Qu J et al (2016) Genetic mapping of QTL for the sizes of eight consecutive leaves below the tassel in maize (Zea mays L.). Theor Appl Genet 129:2191–2209. https://doi.org/10.1007/s00122-016-2767-2
Yue B, Xue W-Y, Luo L-J, Xing Y-Z (2006) QTL analysis for flag leaf characteristics and their relationships with yield and yield traits in rice. Acta Genet Sin 33:824–832. https://doi.org/10.1016/S0379-4172(06)60116-9
Zhao K, Tung C-W, Eizenga GC et al (2011) Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat Commun 2:467. https://doi.org/10.1038/ncomms1467
Zhao X, Fang P, Zhang J, Peng Y (2018) QTL mapping for six ear leaf architecture traits under water-stressed and well-watered conditions in maize (Zea mays L.). Plant Breed 137:60–72. https://doi.org/10.1111/pbr.12559
Zhi X, Massey-Reed SR, Wu A et al (2022a) Estimating photosynthetic attributes from high-throughput canopy hyperspectral sensing in sorghum. Plant Phenomics 2022:1–18
Zhi X, Tao Y, Jordan D et al (2022b) Genetic control of leaf angle in sorghum and its effect on light interception. J Experimental Botany 73:801–816
Zhong Y, Zhao X, Li W, Zhou W (2021) Identification and meta-analysis of QTLs for four leaf shape traits in two maize populations under contracting watering environments. Genet Resour Crop Evol 68:1557–1575. https://doi.org/10.1007/s10722-020-01085-1
Zhu X-G, Song Q, Ort DR (2012) Elements of a dynamic systems model of canopy photosynthesis. Curr Opin Plant Biol 15:237–244. https://doi.org/10.1016/j.pbi.2012.01.010
Acknowledgements
We would like to thank Glen Roulston, Kate Jordan, Janet Roberts, Jane Heron and James Heron for assistance with data collection and the farm staff at Gatton Research Facilities and staff from the Queensland pre-breeding for experiment management.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Contributions
BGJ XZ DJ EM and GH conceived the research plans; XZ SMR and BGJ performed the experiments; CH and YT provided technical assistance to XZ XZ analysed the data and wrote the article; all authors provided input into the interpretation of results and the final version of the article.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Hai-Chun Jing.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhi, X., Hammer, G., Borrell, A. et al. Genetic basis of sorghum leaf width and its potential as a surrogate for transpiration efficiency. Theor Appl Genet 135, 3057–3071 (2022). https://doi.org/10.1007/s00122-022-04167-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-022-04167-z