Abstract
Key message
We mapped three adult plant resistance (APR) loci on oat chromosomes 4D and 6C and developed flanking KASP/PACE markers for marker-assisted selection and gene pyramiding. Using sequence orthology search and the available oat genomic and transcriptomic data, we surveyed these genomic regions for genes that may control disease resistance.
Abstract
Sources of durable disease resistance are needed to minimize yield losses in cultivated oat caused by crown rust (Puccinia coronata f. sp. avenae). In this study, we developed five oat recombinant inbred line mapping populations to identify sources of adult plant resistance from crosses between five APR donors and Otana, a susceptible variety. The preliminary bulk segregant mapping based on allele frequencies showed two regions in linkage group Mrg21 (Chr4D) that are associated with the APR phenotype in all five populations. Six markers from these regions in Chr4D were converted to high-throughput allele specific PCR assays and were used to genotype all individuals in each population. Simple interval mapping showed two peaks in Chr4D, named QPc.APR-4D.1 and QPc.APR-4D.2, which were detected in the OtanaA/CI4706-2 and OtanaA/CI9416-2 and in the Otana/PI189733, OtanaD/PI260616, and OtanaA/CI8000-4 populations, respectively. These results were validated by mapping two entire populations, Otana/PI189733 and OtanaA/CI9416, genotyped using Illumina HiSeq, in which polymorphisms were called against the OT3098 oat reference genome. Composite interval mapping results confirmed the presence of the two quantitative trait loci (QTL) located on oat chromosome 4D and an additional QTL with a smaller effect located on chromosome 6C. This mapping approach also narrowed down the physical intervals to between 5 and 19 Mb, and indicated that QPc.APR-4D.1, QPc.APR-4D.2, and QPc.APR-6C explained 43.4%, 38.5%, and 21.5% of the phenotypic variation, respectively. In a survey of the gene content of each QTL, several clusters of disease resistance genes that may contribute to APR were found. The allele specific PCR markers developed for these QTL regions would be beneficial for marker-assisted breeding, gene pyramiding, and future cloning of resistance genes from oat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The use of genetic resistance is considered the most important and effective strategy to manage oat crown rust, a destructive disease caused by the fungus Puccinia coronata f. sp. avenae (Simons 1985; Nazareno et al. 2017). Resistance to crown rust, or to cereal rusts in general, can either be race-specific or non-race specific, depending on the number of genes involved and the nature of resistance (Periyannan et al. 2017). Race-specific resistance (also called seedling resistance) is usually controlled by a single gene that encodes nucleotide-binding site leucine-rich repeat (NBS-LRR) proteins (Dodds et al. 2006). This type of resistance provides protection all throughout plant development, but it is vulnerable to the emergence of new pathogen races through pathogen evolution. Race-specific resistance to oat crown rust is conferred by Pc genes but these identified genes have not proven durable; and most are only effective for less than five years (Carson 2008). There are more than 100 known Pc genes, and the majority of these are race-specific and exhibit a hypersensitive reaction phenotype (Klos et al. 2017). Race-specific resistance results from the recognition of the avirulence (Avr) protein from the pathogen by the corresponding R protein in the plant (Flor 1971; Dodds and Rathjen 2010). Effective sources of seedling resistance have been identified from A. sterilis, A. strigosa and A. magna, and successfully introgressed into commercial oat varieties (Simons 1985; McMullen et al. 2005; Rines et al. 2007, 2018). Some of the race-specific R genes that have been identified from wild relatives are still used today but often in combination with other genes (Brown et al. 2001) or in production areas where virulence to those genes has not been reported (Sowa and Paczos-Grzęda 2020). To date, only a few race-specific genes have been genetically mapped with available molecular markers, including Pc38, Pc48, Pc54, Pc58a, Pc59, Pc68, Pc71, Pc91, Pc92, Pc94, and PcKM (Penner et al. 1993; Rooney et al. 1994; Bush and Wise 1998; Wight et al. 2004; Chong et al. 2004; McCartney et al. 2011; Klos et al. 2017). The recent mapping of Pc39, Pc45, and Pc53 (Admassu-Yimer et al. 2018; Kebede et al. 2019; Sowa and Paczos-Grzęda 2020; Zhao et al. 2020) adds to the growing list of mapped Pc genes.
Non-race specific resistance, also termed partial or adult plant resistance (APR), does not prevent all pathogen infection and tends to be more durable than race-specific resistance (Leonard 2002; Carson 2009). Partial resistance is thought to be polygenic, although it may be controlled by major loci as suggested by recent studies (Fu et al. 2009; Krattinger et al. 2009; Lin et al. 2014; Moore et al. 2015). Non-race specific resistance is generally characterized by a reduction in pathogen multiplication and colonization, and a lower disease severity (Pilet-Nayel et al. 2017). This is often associated with reduced infection frequency, longer latency period, smaller uredinium size, and early telia formation (Leonard 2002; Díaz-Lago et al. 2003). Furthermore, the known genes for partial resistance are only effective in the adult plant stages; thus, seedlings are moderately susceptible (Rinaldo et al. 2017; Rines et al. 2018). Some examples of APR genes in wheat include Lr34, which encodes an ATP‐binding cassette (ABC) transporter that induces flag leaf senescence and simultaneously provides resistance to leaf rust, stripe rust, and powdery mildew (Krattinger et al. 2009). Another senescence-associated rust resistance gene is Yr36, a kinase-START protein whose activity reduces the ability of the cells to detoxify reactive oxygen species (ROS), which leads to host cell death that confers resistance to stripe rust (Fu et al. 2009; Gou et al. 2015). In addition, Lr67 is a mutated hexose transporter APR gene that causes leaf tip necrosis and limits colonization of biotrophic pathogens (Moore et al. 2015).
The mechanism of APR in oat is still poorly understood, though a number of oat lines have been identified to be effective sources of APR. Notably, MN841801, a line developed by the University of Minnesota, has exhibited a resistant phenotype in over 30 years of testing under high disease pressure (Leonard 2002). The resistance in MN841801 is controlled by a major QTL on oat chromosome 14D (Lin et al. 2014); however the causal gene has not been identified so far. Recently, two lines derived from diploid A. strigosa, MNBT1020-1 and MNBT1021-1, were identified to be useful sources of APR to crown rust after over 8 years of testing. These lines carry two dominant genes in the oat linkage group Mrg20 conferring the resistance (Rines et al. 2018). Not only do these lines provide durable sources of resistance but they also offer opportunities to understand the mechanism of APR in oat through identification and future cloning of APR genes. In light of the recent crown rust epidemics in the United States (Nazareno et al. 2017), more sources of APR are needed to develop oat varieties with broad spectrum crown rust resistance. In this study, five recombinant-inbred mapping populations were generated and mapped with molecular markers to identify APR loci and developed PCR markers linked to the QTL for marker-assisted breeding.
Materials and methods
Selection of oat donors with adult plant resistance (APR) to crown rust
To select for donors of APR based on the phenotype, thirty-three A. sativa lines were screened at the University of Minnesota Matt Moore Buckthorn Plots at the Minnesota Agricultural Experiment Station, in St. Paul, MN (buckthorn nursery), where a sexually recombining crown rust population is present. In addition, bulk urediniospores were collected from these field plots, stored, and then used to inoculate individual entries at the seedling stage under controlled conditions in the greenhouse (24 °C, 45% relative humidity). The urediniospores from susceptible pustules observed on these seedlings were collected from each oat entry. At adult plant stage, the bulk urediniospores were then used to inoculate the same entry from which the spores were collected, in a subsequent greenhouse test with two replicates (Fig. 1A). Lines with moderate to low severity reactions in both buckthorn and inoculated nurseries were selected as parents for biparental mapping (Fig. 1B).
Selection of APR donors (1–9 severity scale). A Screening procedure for potential APR sources in the buckthorn nursery and inoculated greenhouse nursery. B Regression plot of mean crown rust severity in the buckthorn nursery (BN) and mean crown rust severity in the inoculated nursery (IN) for the 33 oat lines. Recombinant inbred mapping populations were developed for the oat lines are shown in red font; populations that were mapped in this paper are boxed. The oat differential line postulated to contain the seedling resistance gene Pc94 was included as experimental control
Mapping populations and crown rust phenotyping
Five recombinant inbred mapping populations derived from crosses between the susceptible cultivar Otana and five APR donors were developed using the single seed descent method (Table 1). Recombinant inbred lines (RILs) from each mapping population were sown as head rows (1–3 m long, 0.3 m apart) in the buckthorn nursery in May during three seasons (2017, 2018, and 2020). The populations were phenotyped for APR around the milk to dough stages on a per row basis using the modified Cobb’s scale (Peterson et al. 1948). Crown rust reaction was assessed by assigning each plot a disease severity, which is the estimate of the percentage of leaf area covered by pustules, and a reaction class (resistant (R), moderately resistant (MR), moderately susceptible (MS), and susceptible (S), that was then converted to a reaction score from 0 to 1 (R = 0, RMR = 0.1667, MR = 0.3333, MRMS = 0.5, MS = 0.6667, MSS = 0.8333, S = 1). The disease severity (DS) was then multiplied to the reaction class (RC) to obtain the coefficient of infection (CI) (Lin et al. 2014). The CI (%) was used in the rank analysis to determine lines that consistently exhibit extreme phenotypes (highly resistant and highly susceptible) (Table 1). For trait analysis of the CI phenotype, a pairwise correlation was performed in Microsoft Excel v16.48 using the CORREL function. The CI, DS, and RC data were used in simple and composite interval mapping.
DNA extraction, selective genotyping, and allele frequency calculation
Genomic DNA was extracted from silica-dried leaves of plants from each population, including the parents, using BioSprint 96 DNA Plant Kit (Qiagen, MD, USA), following the manufacturer’s protocol. The DNA concentration was measured using a Biotek plate reader (Biotek Instruments, Inc., VT, USA) and normalized to 50 ng/ul for genotyping assays. From each population, 12 to 24 highly resistant and 12 to 24 highly susceptible lines were selected for genotyping (Table 1). Each line was individually assayed using the 6 K Oat SNP Chip (Illumina, CA, USA). Allele calling was conducted using the Illumina GenomeStudio v2.0 polyploid clustering module. Genotypes were clustered using both Polygentrain and DBSCAN algorithms and markers with allele frequencies of either 1 or 0 were discarded. After filtering, the allele calls of the remaining markers were manually inspected to only include polymorphic markers (Table 1), prior to their use for mapping resistance loci based on allele frequencies. In both resistant and susceptible allele groups, the frequency of the resistant allele was computed by counting the number of individuals possessing the allele and dividing it by the total number of individuals in the group. The allele frequency difference (AFD) was calculated by obtaining the absolute value of the difference in allele frequency between resistant and susceptible groups. The Wilcoxon rank sum test imbedded in the compare_means function in R v4.0.5 was used to determine linkage groups with significantly different AFD per population. AFD values for all five populations were plotted using ggplot2 v3.3.3. To determine the relatedness among APR donors, a neighbor-joining tree was created in TASSEL v5 (Bradbury et al. 2007) using 3015 SNP markers from the Infinium 6 K Oat SNP Chip (Tinker et al. 2014) (Supplementary Fig. 1).
Kompetitive allele-specific PCR (KASP) and PCR allelic competitive extension (PACE) marker development and assays
The KASP markers were designed by inputting the flanking sequence of the desired SNP marker in the Web-based Allele Specific Primer (WASP) design tool (https://bioinfo.biotec.or.th/WASP) (Wangkumhang et al. 2007) with default parameters. The sequences of the HEX (5’ GAAGGTCGGAGTCAACGGATT 3’) and FAM (5’ GAAGGTGACCAAGTTCATGCT 3’) dyes were then added to the 5’ end of the forward primers (Supplementary Table 1). Meanwhile, the PACE markers were designed using the free assay design service by 3CR Bioscience (Essex, UK) (www.3crbio.com/free-assay-design). The forward and reverse primers were maintained at stock concentrations of 100 µM. In a 100 µl assay mix, 12 µl each of the forward primers was mixed with 30 µl of the reverse primer and 46 µl of DEPC water. The final 10 µl reaction mix per well is composed of the following: 5 µl 2X KASP buffer (LGC Genomics, MA, USA) or 2X PACE Genotyping Master Mix (3CR Bioscience, Essex, UK), 2.86 µl DEPC water, 0.14 assay mix, and 2 µl template DNA. The PCR assay in LightCycler 480 II (Roche, IN, USA) was performed under the following conditions: Pre-incubation- 1 cycle, 95 °C target, 15 min hold, 4.4 °C/s ramp rate; Amplification- 40–50 cycles, 94 °C target, 20 s hold, 2.2 °C/s ramp rate; Annealing- 40–50 cycles, 61–63 °C target, 1 min hold, 2.2 °C/s ramp rate, 55 °C sec target, 5 °C step size; Endpoint read- 37 °C target, single acquisition mode, 2 min hold, 2.2 °C/s ramp rate. The KASP/PACE assays were optimized by adjusting the number of cycles, annealing temperature, and concentrations of the forward primers (Supplementary Table 1). Genotype data from each population were exported from LightCycler 480 II and visualized in R v4.0.5 using ggplot2 v3.3.3.
Simple interval mapping using KASP/PACE genotyping data
Six KASP/PACE markers were developed from the two QTL regions in Chr4D that were initially identified using bulk segregant analysis (BSA) mapping. The markers were used to genotype all individuals of the five populations and the genotype data were converted into ABH format using a custom R script. The genetic map was constructed using the MAP functionality of the IcIMapping software v4.2 (Meng et al. 2015) with grouping by threshold value (REC = 0.30), ordering by k-Optimality (by REC, 2-OptMap, and NN initials of 10), and rippling by REC (window size = 5) options. The phenotype (DS, RC, and CI), marker, and genetic map data were converted to qdf format for the QTL analysis in QGene v4 (Joehanes and Nelson 2008). Simple interval mapping was performed to detect loci associated with APR and logarithm of odds (LOD) scores of 3 or higher were considered significant (Bernardo 2020). Mapping statistics were exported from QGene and loaded into ggplot2 v3.3.3 for visualization in R v4.0.5. Linkage maps were visualized using the R package LinkageMapView v2.1.2 (Ouellette et al. 2017).
Validation of BSA mapping results with genotyping-by-sequencing (GBS) data
DNA extractions and library preparation for the double digest (PstI-MspI) of OtanaA/CI9416-2 and Otana/PI189733 populations sequencing libraries were performed as described in previous work (Huang et al. 2014). The DNA libraries were pooled (96 samples per flow-cell lane) and sequenced on an Illumina HiSeq4000 instrument in PE 150 mode. Paired end reads were quality filtered and adapter trimmed with bbduk v37.02 (https://jgi.doe.gov/data-and-tools/bbtools/bb-tools-user-guide/bbduk-guide/) and demultiplexed with sabre (https://github.com/najoshi/sabre). Reads were aligned to the RefSeqV1 oat genome (PepsiCo 2020) with bowtie2 using the -very-sensitive-local option. SNPs were called with bcftools v1.9 from reads with a mapQ score greater than 20. Using VCFtools v1.16, SNPs were converted to missing if the read depth was less than 4 and SNPs were removed if they have more than 50% missing data or have a minor allele frequency (MAF) of less than 0.01 within a given population. The genotype data was converted into ABH format and loaded in the IcIMapping software v4.2 to remove markers that are present in the same genetic position using the BIN functionality. For binning, markers with 100% missing rate and distortion P value of 0 were deleted; the option ‘consider anchor info’ was selected and redundant markers were deleted by missing rate (%). The genetic map was then constructed using the MAP functionality with the sequence-anchor option, and the phenotype (CI, DS, and RC), genetic map, and marker data were loaded to QGene v4 for linkage mapping, as described above. In QGene v4, composite interval mapping was selected with default settings of forward cofactor selection for QTL analysis. The option that allows QTL as cofactor, instead of markers, was also selected to compare the results. To declare significant QTL, a permutation analysis for LOD score of CI was conducted with 1000 iterations (P < 0.05) to compute for the LOD threshold. Mapping statistics were exported from QGene and loaded into ggplot2 v3.3.3 for visualization in R v4.0.5. To determine the linkage disequilibrium (LD) in the QTL regions, the genotype data (HapMap format) was loaded to TASSEL v5, where pairwise LD was analyzed after KNNi imputation (Money et al. 2015). The results were exported from TASSEL v5 and visualized in R v4.0.5 using the package LDheatmap v1.0.4 (Shin et al. 2006).
Survey of putative candidate genes in QTL regions
A survey of orthologous genes within the physical interval of the three APR loci was conducted by performing BLAST alignments of the available oat OT3098 v1 mRNA transcripts (PepsiCo 2020; Hu et al. 2020) with the non-redundant NCBI database using default parameters of BLASTN. An alignment E value of less than 1e−10, query coverage of at least 30%, and sequence identity of at least 50% were used to declare orthologous genes. The mRNA sequences and coordinates were obtained from the oat genome browser accessible at https://wheat.pw.usda.gov/GG3/graingenes_downloads/oat-ot3098-pepsico.
Results
Selection of oat donors that contain APR against crown rust
Thirty-three oat lines were screened to determine potential sources of APR (Carson, unpublished). These lines were planted in the buckthorn nursery for initial screening and then bulk urediniospores from that nursery were collected to inoculate seedlings in the greenhouse. Urediniospores from all pustules with scores of 3, 3 + , and 4 in the modified Murphy’s scale (Murphy 1935) that developed in each entry were collected and used to inoculate the same entry in the greenhouse (Fig. 1A). Nine potential sources of APR were identified (Fig. 1B). Recombinant inbred mapping populations were developed from five donors, namely: CI4706, PI260616, CI9416, CI8000, and PI189733 (Table 1). Oat lines were considered appropriate sources of APR if low to moderate severity of crown rust was observed in adult plants in both inoculated and buckthorn nurseries (Fig. 1B). In Fig. 1B, oat line Pc94 is a distinct outlier that shows low mean severity in the buckthorn nursery but high mean severity in the inoculated nursery, which is expected for an oat differential line that contains only one major Pc gene, while Lodi and Portage cultivars were more susceptible than the remainder of the entries. The total number of recombinant inbred lines, number of selected highly resistant and highly susceptible lines, and number of markers used for bulk segregant analysis (BSA) mapping and QTL validation in each of the mapping populations are reported in Table 1.
Phenotypic analysis of recombinant inbred mapping populations
Phenotypic data for disease severity (DS), reaction coefficient (RC), and coefficient of infection (CI) were collected in 2017 for OtanaD/PI260616-1 (OTD/PI260616-1) and OtanaA/CI8000-4 (OTA/CI8000-4), in 2018 for OtanaA/CI4706-2 (OTA/CI4706-2) and OtanaA/CI9416-2 (OTA/CI9416-2), and in 2020 for Otana/PI189733 (OT/PI189733) (Fig. 2). All five populations showed a normal distribution for DS. However, OTA/CI4706-2 and OTA/CI9416-2 exhibited a distribution skewed towards susceptible phenotypes for RC, in contrast to OT/PI189733 population, which had more resistant lines, and OTD/PI260616-1 and OTA/CI8000-4 which displayed more even distributions. For CI, generally bimodal distributions were observed for the OTA/CI4706-2, OTA/CI9416-2, and OTA/CI8000-4 populations. The susceptible parent, Otana and its reselections, OtanaA and OtanaD, did not differ considerably for DS, RC, and CI. The range of phenotypic values for the five APR parents were 30–40 for DS, 0.33 to 0.5 for RC, and 9.9 to 20 for CI. The pairwise correlation analysis of the CI phenotype suggested that OTA/CI4706-2 and OTA/CI9416-2 were more closely correlated, as were OTA/CI8000-4 to OTD/PI260616-1. However, OT/PI189733 was correlated to both OTA/CI9416-2 and OTD/PI260616-1 (Supplementary Table 2).
Phenotypic scores for disease severity (DS), reaction class (RC), and coefficient of infection (CI) for each of the five mapping populations. The y-axis in each plot represents the number of lines from each population that display scores shown in the x-axis, while population ID is shown on the right y-axis; OT = Otana, OTA = OtanaA, OTD = OtanaD
Identifying APR loci based on allele frequencies
The number of polymorphic SNP markers used for BSA mapping of APR loci are shown in Table 1. Differences in marker allele frequencies were computed for each population and the results of the Wilcoxon rank sum test (P < 0.05) revealed a significant difference in AFD for linkage group Mrg21 (Chr4D) in all five populations examined (Supplementary Fig. 2). Changes in allele frequencies between the resistant and susceptible groups also showed the presence of two peaks in Chr4D. A total of six SNP markers spanning these peaks were converted for KASP/PACE assays. The optimized PCR markers were polymorphic and differentiated marker genotypes across five populations (Supplementary Fig. 3, Supplementary Table 1).
BSA mapping using KASP/PACE markers
The six PCR markers were used to genotype all individuals in all five populations and generate data for validation and Mrg21 linkage map construction (Fig. 3). For simplicity, the markers were arranged according to their positions in the consensus linkage map (Chaffin et al. 2016). However, analysis of the marker order per population showed that the markers were linked in three pairs: GMI_ES17_c5923_221 with GMI_DS_LB_3908, GMI_ES05_c8031_345 with GMI_DS_LB_4204, and GMI_ES05_c15948_428 with GMI_ES03_c2344_498 (Supplementary Fig. 4). The genetic distances among the six markers in these populations also deviated considerably from the consensus map. Simple interval mapping revealed the presence of two QTL regions in Chr4D (Fig. 3). The first significant region was detected in the OTA/CI4706-2 and OTA/CI9416-2 populations flanked by GMI_DS_LB_4204 (238,688,556 bp) and GMI_ES05_c15948_428 (294,423,672 bp), while the second significant region was mapped in the OT/PI189733, OTD/PI260616-1, and OTA/CI8000-4 populations, and was flanked by GMI_ES03_c2344_498 (332,650,198 bp) and GMI_ES17_c5923_221 (414,098,082 bp). The physical positions of the KASP/PACE markers in the oat genome were obtained from the T3 Oat Database (https://triticeaetoolbox.org/oat/) and through aligning the flanking sequence of the markers to the OT3098 reference genome, which indicated that the significant regions in Mrg21 were physically located on oat chromosome 4D (Supplementary Table 1). The six KASP/PACE markers were also used to genotype a set of oat germplasm for validation (Supplementary Table 4).
Simple interval mapping using markers identified from the bulk segregant analysis (BSA) in Chr4D (Mrg21). The markers were ordered based on their positions in the consensus linkage map and regressed against the disease severity (DS), reaction class (RC), and coefficient of infection (CI) data per population. The physical positions of the KASP/PACE markers in Chr4D are shown in parenthesis
Composite interval mapping using GBS data
Composite interval mapping of two populations, OTA/CI9416-2 and OT/PI189733, which were fully genotyped by Illumina HiSeq GBS, was conducted to verify the results of the BSA mapping and narrow down the significant regions. The genetic mapping results confirmed the presence of two distinct regions on chromosome 4D (Fig. 4), named QPc.APR-4D.1 and QPc.APR-4D.2, while an additional minor QTL on chromosome 6C also was detected in OT/PI189733. Two methods of composite interval mapping were compared: selecting QTL as cofactor vs. selecting markers as cofactor. The results suggest that the forward selection of markers as cofactor (in auto mode) narrowed down the significant regions into 19 Mb, 5 Mb, and 16 Mb sizes for QPc.APR-4D.1, QPc.APR-4D.2, and QPc.APR-6C, respectively (Fig. 5A–C). The mapping statistics for the two populations are shown in Table 2. The maximum LOD score for CI was 21.37 for QPc.APR-4D.1, 17.97 for QPc.APR-4D.2, and 8.95 for QPc.APR-6C, while the R2 values were 43.4%, 38.5%, and 21.5% respectively. Based on the physical positions of markers linked to the QTL, QPc.APR-4D.1 and QPc.APR-4D.2 were about 150 Mb apart. Further, GMI_DS_LB_4204 and GMI_ES03_c2344_498 could be used for haplotype selection of QPc.APR-4D.1, GMI_ES17_c5923_221 and GMI_DS_LB_3908 for QPc.APR-4D.2, and S6C_515268054 and S6C_531401236 for QPc.APR-6C (Table 3). Pairwise LD between markers in chromosome 4D was also computed to determine the extent of linkage within the QTL regions. LD blocks are evident in chromosome 4D in both mapping populations, as shown by high R2 values (Fig. 5A–C). Meanwhile, the physical order of markers coincides with the genetic order in OT/PI189733, while a deviation from the physical order was observed in OTA/CI9416-2.
Comparison of composite interval mapping of A QPc.APR-4D.1, B QPc.APR-4D.2, and C) QPc.APR-6C using markers as cofactor vs. using QTL as cofactor. The genetic positions of markers were anchor-ordered based on recombination fraction estimates in each population. Pairwise linkage disequilibrium (LD) was computed and is shown based on the physical order of markers in chromosome 4D. Higher R2 indicate higher LD; dotted lines denote relative positions of LD blocks in the linkage map and the physical map. Markers in bold have been converted to KASP/PACE; those in italic-bold were used for haplotype analysis
Results of the additive effect analysis from composite interval mapping showed that the per copy effect of resistant SNP alleles could explain the decrease in CI by up to 4 to 8.7 (Table 2), equivalent to 8 to 17.4 in lines homozygous to these alleles. Haplotype analysis of flanking markers for QPc.APR-4D.1 and QPc.APR-4D.2 showed significant differences in lines containing the resistant alleles compared to those that possess the susceptible alleles (Table 3). Among lines that contain the resistant allele haplotypes, the mean CI was lowered by 14.3 to 17.7 units for QPc.APR-4D.1, and 10.1 to 22.9 units for QPc.APR-4D.2, and 7.8 units for QPc.APR-6C. Additive effects of for QPc.APR-4D.2 and QPc.APR-6C also significantly reduced the CI in lines with the two QTL, compared to having just one of these QTL (Table 3). All the susceptible allele haplotypes originated from Otana, which contributed to a significant increase in CI.
Survey of putative candidate genes in QTL regions
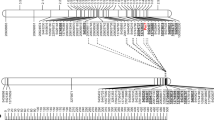
Using the recently released oat genome and transcriptome assemblies (PepsiCo 2020; Hu et al. 2020), we performed a sequence orthology search on NCBI, which identified genes within and near the physical interval of the three APR loci that may contribute to resistance (Fig. 6A–C). Based on the genome assembly, chromosome 4D has a length of 425 Mb out of the total oat genome size of 11 Gb (PepsiCo 2020). The 4D chromosome has an estimated total of 9300 expressed genes, 758 of which are within the 19 Mb region of QPc.APR-4D.1 and 124 genes are within the 5 Mb interval of QPc.APR-4D.2. By contrast, oat chromosome 6C has a size of 625 Mb and 8612 expressed genes, 596 of which are within the 16 Mb significant region of QPc.APR-6C. For the two major QTL, QPc.APR-4D.1 and QPc.APR-4D.2, the gene content of the region flanking the APR locus was also surveyed (Fig. 6A–C).
For QPc.APR-4D.1, gene such as those that encode for protein kinases, WRKY-like protein, and disease resistance proteins were detected in the significant region. Nearby genes outside the interval include orthologs of barley serine/threonine kinase rust resistance gene Rpg1 (ABC1041), Triticum dicoccoides powdery mildew resistance gene Pm41 with coiled-coil, nucleotide-binding site and leucine-rich repeat domains, and Brachypodium distachyon protein kinase (brassinosteroid-signaling kinase 5, BSK5). For QPc.APR-4D.2, distinct clusters of resistance genes were detected near the telomere of chromosome 4D. The 5 Mb significant region contained an ABC transporter, a disease resistance gene analog (RGA), a defensin, and other genes that may contribute to plant immunity. Proximate to the significant region, a cluster including a receptor kinase, ABC transporter, and RGA was present, and further upstream, orthologs of the wheat Yr36 (WKS1 and WKS2) and a receptor-like protein coding gene were detected. For QPc.APR-6C, the 16 Mb significant region includes genes for kinases, ABC and sugar transporters, disease resistance, and other genes that code for F-box/leucine-rich repeat (LRR) protein, along with an mRNA decapping enzyme, aquaporin, and ethylene sensor genes (Fig. 6C). Near the peak of the QTL, we found genes encoding an RGA, a LRR-receptor kinase, a phosphatase, and several ethylene receptor genes.
Discussion
Understanding the nature of the genes involved in disease resistance is important in supporting crop breeding programs. This is particularly relevant for pathogens like P. coronata f. sp. avenae, which display broadly virulent, diverse, and fast-evolving populations (Menzies et al. 2019; Miller et al. 2021). In these cases, finding sources of APR (quantitative resistance or partial resistance) provides a mechanism to slow down pathogen evolution and the emergence of new races by reducing inoculum levels in the field (Leonard 2002). Differentiating quantitative from qualitative resistance (seedling resistance) in field trials presents challenges, particularly when potential donors of APR are screened with uncharacterized pools of pathogen races and seedling resistance have masking effects. To increase the chances of detecting quantitative resistance and reduce interference from seedling resistance, it is best to conduct donor screenings under high pathogen diversity and complement such trials with controlled seedling infection assays. A screening procedure was employed that involved collecting bulk urediniospores from the buckthorn nursery and using these spores to inoculate oat lines at seedling stage under controlled conditions to identify APR sources. By collecting only virulent pustules from each oat entry from the greenhouse tests and inoculating each oat entry with its specifically virulent population as adult plants in the greenhouse, APR was differentiated from seedling resistance. Putative donors of APR were also validated in the buckthorn nursery in several trials across multiple years. Given that buckthorn hedges facilitate sexual reproduction of P. coronata f. sp. avenae, oat entries were screened against a high number of pathogen genotypes under maximum disease pressure. This scheme has been used previously to identify oat lines with effective APR (Leonard 2002; Carson 2009). Based on this approach, five donors of APR were selected for downstream analysis in this study.
Similarity among the oat populations was evident based on the phenotype and BSA mapping data on Chr4D. Based on the trait correlation analysis of CI, OTA/CI4706-2 and OTA/CI9416-2 were similar, as were OTD/PI260616-1 and OTA/CI8000-4, while OT/PI189733 has similarity to both OTA/CI9416-2 and OTD/PI260616-1. BSA mapping results also showed that OTA/CI4706-2 and OTA/CI9416-2 contain the same APR region with peak while OT/PI189733, OTD/PI260616-1, and OTA/CI8000-4 possess the same QTL (Fig. 3). To determine the relatedness between APR donors, we conducted a phylogenetic analysis using polymorphic SNP markers, which showed that PI189733 belonged to a different clade compared to the rest of the APR parental lines (Supplementary Fig. 1). This supports the results of the phenotypic analysis and indicated that PI189733 is genetically unique compared to the other parents. Based on the passport data from GRIN-Global (https://npgsweb.ars-grin.gov/gringlobal), PI189733 was a collection from Pichincha, Ecuador, while the other parents are either breeding lines or varieties (Supplementary Table 3), which might explain the genetic differences among these lines.
Normally distributed phenotypic values for a quantitative trait implies that the phenotype is controlled by several minor genes (Li et al. 2015). This is not the case for our populations, particularly with the CI trait, suggesting that a few major QTL with large effects confer the resistance. While APR is thought to be a polygenic trait, detecting major QTL is a common finding when BSA is used to map RIL populations (Liu et al. 2014; Zou et al. 2016). Due to the simplicity of BSA with regards to sampling and reduced cost compared to genotyping all individuals of each population, this approach was favored to achieve the primary goal of quickly mapping APR loci with large effects. The modified BSA method involved individual genotyping of selected lines, which allowed scoring for allele frequency for each marker. To statistically determine the effects of the APR loci, SNP markers in putative regions of Chr4D (Supplementary Fig. 2) were converted into KASP/PACE markers and used to genotype all individuals in each population for linkage mapping. With this, the APR-associated loci were rapidly narrowed down, and two genomic regions of interest were detected in the linkage group in all five populations (Fig. 3). These KASP/PACE markers will be useful for germplasm selection and future gene pyramiding to extend the lifetime of disease resistance genes in oat varieties. To demonstrate this, we used the six KASP/PACE markers to genotype a set of 16 oat germplasm and found that they could detect the presence/absence of the two QTL (Supplementary Table 4). It should be noted, however, that other QTL for rust resistance may be present in these lines and caution must be exercised when using the markers with diverse materials.
To corroborate the BSA mapping results, two representative populations that possess two different QTL in Chr4D were selected for whole population genotyping. We utilized an Illumina HiSeq genotyping platform to generate genotyping-by-sequencing (GBS) data from all lines in OTA/CI9416-2 and OT/PI189733 populations and SNPs were called against the OT3098 oat genome. This enabled anchoring of the genetic markers onto the physical oat map for linkage mapping, enhancing the resolution and estimates of the QTL effects, and determine the relationships between the linkage map and the physical map. The results of the interval mapping confirmed the BSA mapping results (Fig. 4) and implied that Mrg21 corresponds to oat chromosome 4D, as also suggested previously (Maughan et al. 2019). Further, a minor QTL in OT/PI189733, named QPc.APR-6C, was detected in Chr6C (Mrg17). The correspondence of the Mrg groups and physical chromosomes in this study was in accordance with the recently approved nomenclature by the International Oat Nomenclature Committee (IONC, 2021).
The marker cofactor selection improved the mapping resolution and estimates of QTL effects (Fig. 5A–C). Composite interval mapping uses both interval mapping, which defines the location of the QTL, and multiple regression, which reduces the background noise caused by presence of other QTL through selection of a few markers as cofactors (Zeng 1994; Bernardo 2020). This enabled the narrowing down of the significant regions into 5 to 19 Mb sized regions and indicated that QPc.APR-4D.1 and QPc.APR-4D.2 are QTL with large effects, while QPc.APR-6C is a QTL with small effect. QPc.APR-4D.1 and QPc.APR-4D.2 were found in different positions of the chromosome 4D linkage map, which in parallel, are also observed in positions that are 150 Mb apart in the physical map, suggesting that they are different QTL.
Analysis of linkage disequilibrium in the APR loci revealed high LD and presence of LD blocks on the chromosome, implying that these regions are not subject to frequent recombination. However, recombination between LD blocks likely occurs, as shown in OTA/CI9416-2 (Fig. 5A). This further implies that the KASP/PACE markers within a LD block will have a high chance of staying linked to the QTL, even if situated a few cM or Mb away from the peak, which was the case for the PCR markers that were developed for both QPc.APR-4D.1 and QPc.APR-4D.2. This highlighted one of the disadvantages of BSA and simple interval mapping, which is their inability to detect the exact QTL peak in a significant region. After examining the KASP/PACE genotyping data to determine the additive effects of the QTL, we found that CI could be lowered by up to 17.7 with QPc.APR-4D.1 and 22.9 with QPc.APR-4D.2, while CI was decreased by 7.8 with QPc.APR-6C (Table 3). Overall, this is a large reduction of the disease coefficient, considering that the maximum CI observed was 70. Selecting based on the haplotype of the flanking markers also increased the chance of selecting the entire QTL, compared to using only one marker for selection. The additive effect of QPc.APR-4D.2 and QPc.APR-6C in OT/PI189733 RILs also showed that the combination of the two QTL could slightly reduce CI compared to having just one of the QTL. It was also observed that the genetic distances of the KASP/PACE markers were not comparable to the ones indicated in the consensus map, which may be attributed to chromosomal rearrangements, such as inversions and translocations (Nikoloudakis and Katsiotis 2015; Chaffin et al. 2016; Latta et al. 2019) or other genetic differences between our populations and those used in the consensus map development.
In this study, two APR QTL on chromosome 4D and one in chromosome 6C are reported: QPc.APR-4D.1 with a physical interval of 19 Mb, QPc.APR-4D.2 with a physical interval of 5 Mb, and QPc.APR-6C. The Chr4D intervals can be further refined for future fine mapping of APR genes, considering the size of the chromosome, which is 425 Mb. The same can be inferred for the QPc.APR-6C interval, with Chr6C having a size of 625 Mb. However, these intervals were comparable to some recent coarse mapping work in wheat (Yang et al. 2020; Lin et al. 2021), which has a larger genome size (17 Gb) than oat. These QTL are also believed to be novel, as no crown rust QTL have been reported in these linkage groups thus far based on Pc genes and QTL that have been resolved to the latest oat consensus map (Chaffin et al. 2016; Klos et al. 2017), however, other crown rust resistance QTL have been reported in linkage groups Mrg02, Mrg03, Mrg06, Mrg08, Mrg12, Mrg17, Mrg20, and Mrg23 (Zhu and Kaeppler 2003; Portyanko et al. 2005; Acevedo et al. 2010; Lin et al. 2014; Babiker et al. 2015; Klos et al. 2017). Based on BLAST analysis of the marker sequences, neither of our two QTL on Chr4D are the same with the highly effective QTL reported by Rines et al. 2018 on the Chr4A homeolog. There were, however, five crown rust QTL that have been reported on Chr4A, in addition to Pc48 (Klos et al. 2017).
Access to the newly released oat genome and transcriptome (PepsiCo 2020; Hu et al. 2020) enabled us to survey the gene content of the three APR loci. The gene survey for QPc.APR-4D.1 revealed interesting genes; notably, a WRKY1-like gene, a mitogen activated protein kinase (MAPK)-kinase, a Pik-2 ortholog, an A. strigosa putative resistance gene, an RGA, and a BAM1 ortholog (Fig. 5B). WRKY transcription factors and MAPK kinases are known to be involved in PAMP-triggered immunity (PTI) (Arthur and Ley 2013; Bakshi and Oelmüller 2014). The resistance genes Pik-2 and Pik-1 confer resistance in rice to rice blast (Zhai et al. 2011), while BAM1 works with a receptor-like protein kinase and is involved in a signaling pathway in Arabidopsis (Shimizu et al. 2015). RGAs can either be NLRs, receptor-like kinases, or receptor-like proteins (Sekhwal et al. 2015), while the A. strigosa L7M2.1 putative resistance was predicted to be a NLR (GenBank: FJ829742.1). For QPc.APR-4D.2, a narrower interval led to fewer putative candidates, including ABC transporter B, defensin, RGA5-like, and Pik6-NP-like genes. ABC transporters have been shown to be engaged in a multitude of plant processes, including resistance to diseases (Kang et al. 2011), while defensins are antimicrobial compounds primarily involved in basal plant immunity (Parisi et al. 2019). On the other hand, RGA5 and Pik6 genes both encode for immune receptors in rice that detect avirulence proteins from Magnaporthe grisea (Ashikawa et al. 2008; Cesari et al. 2013). For QPc.APR-6C, genes identified closest to the peak include those that code for RGA, LRR-receptor kinase, phosphatase, and an ethylene receptor, any of which could play a role in PTI (Gallie 2015; Durian et al. 2016). Based on the gene surveys from both QTL, it appears that both regions contain genes that are involved in PAMP-triggered and effector-triggered immunity (ETI), and thus, it would be beneficial to fine map these regions to determine if there is only one gene affecting the resistance phenotype, or if multiple genes are working together to confer the resistance. By definition, NLRs cannot confer APR and if these genes are responsible for the phenotypes, we would not expect the resistance to be long term durable. Isolation of the genes conferring the resistance phenotype at the adult stage is critical to determine if they are bonafide APR sources.
In summary, we identified several sources of resistance against oat crown rust that act at the adult plant stage. We subjected these donors to a modified bulked segregant analysis to rapidly map major loci contributing to the resistance phenotype. The QTL were validated using genotype data from GBS and markers linked to them were developed for marker-assisted selection and tested in a set of oat germplasm. Finally, we identified genes in the QTL regions that may be responsible for the resistance phenotypes. This study lays a foundation for future fine mapping and cloning of genes that control APR and provides insights for further investigations of oat-crown rust interactions.
Data availability
The raw QTL mapping files in qdf format are available at https://github.umn.edu/nazar047/TAG-AdultPlantResistance. Marker information are provided in the supplementary files.
References
Acevedo M, Jackson EW, Chong J et al (2010) Identification and validation of quantitative trait loci for partial resistance to crown rust in oat. Phytopathology 100:511–521
Admassu-Yimer B, Bonman JM, Esvelt Klos K (2018) Mapping of crown rust resistance gene Pc53 in oat (Avena sativa). PLoS ONE 13:e0209105
Arthur JSC, Ley SC (2013) Mitogen-activated protein kinases in innate immunity. Nat Rev Immunol 13:679–692. https://doi.org/10.1038/nri3495
Ashikawa I, Hayashi N, Yamane H et al (2008) Two adjacent nucleotide-binding site–leucine-rich repeat class genes are required to confer Pikm-specific rice blast resistance. Genetics 180:2267–2276. https://doi.org/10.1534/genetics.108.095034
Babiker EM, Gordon TC, Jackson EW et al (2015) Quantitative trait loci from two genotypes of oat (Avena sativa) conditioning resistance to Puccinia coronata. Phytopathology 105:239–245. https://doi.org/10.1094/PHYTO-04-14-0114-R
Bakshi M, Oelmüller R (2014) WRKY transcription factors: jack of many trades in plants. Plant Signal Behav 9:e27700–e27700. https://doi.org/10.4161/psb.27700
Bernardo RN (2020) Breeding for quantitative traits in plants. Stemma Press, Woodbury
Bradbury PJ, Zhang Z, Kroon DE et al (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Brown PD, Duguid SD, Haber S et al (2001) AC Assiniboia oat. Can J Plant Sci 81:77–79. https://doi.org/10.4141/P00-036
Bush AL, Wise RP (1998) High-resolution mapping adjacent to the Pc71 crown-rust resistance locus in hexaploid oat. Mol Breed 4:13–21. https://doi.org/10.1023/A:1009652222382
Carson ML (2008) Virulence frequencies in oat crown rust in the United States from 2001 through 2005. Plant Dis 92:379–384. https://doi.org/10.1094/PDIS-92-3-0379
Carson ML (2009) Broad-spectrum resistance to crown rust, Puccinia coronata f. sp. avenae, in accessions of the tetraploid slender oat Avena Barbata. Plant Dis 93:363–366
Cesari S, Thilliez G, Ribot C et al (2013) The rice resistance protein pair RGA4/RGA5 recognizes the Magnaporthe oryzae effectors AVR-Pia and AVR1-CO39 by direct binding. Plant Cell 25:1463–1481. https://doi.org/10.1105/tpc.112.107201
Chaffin AS, Huang Y-F, Smith S et al (2016) A consensus map in cultivated hexaploid oat reveals conserved grass synteny with substantial subgenome rearrangement. Plant Genome. https://doi.org/10.3835/plantgenome2015.10.0102
Chong J, Reimer E, Somers D et al (2004) Development of sequence-characterized amplified region (SCAR) markers for resistance gene Pc94 to crown rust in oat. Can J Plant Pathol 26:89–96. https://doi.org/10.1080/07060660409507118
Díaz-Lago JE, Stuthman DD, Leonard KJ (2003) Evaluation of components of partial resistance to oat crown rust using digital image analysis. Plant Dis 87:667–674. https://doi.org/10.1094/PDIS.2003.87.6.667
Dodds PN, Rathjen JP (2010) Plant immunity: towards an integrated view of plant–pathogen interactions. Nat Rev Genet 11:539–548
Dodds PN, Lawrence GJ, Catanzariti A-M et al (2006) Direct protein interaction underlies gene-for-gene specificity and coevolution of the flax resistance genes and flax rust avirulence genes. Proc Natl Acad Sci 103:8888–8893. https://doi.org/10.1073/pnas.0602577103
Durian G, Rahikainen M, Alegre S et al (2016) Protein phosphatase 2A in the regulatory network underlying biotic stress resistance in plants. Front Plant Sci 7:812. https://doi.org/10.3389/fpls.2016.00812
Flor H (1971) Current status of the gene-for-gene concept. Annu Rev Phytopathol 9:275–296. https://doi.org/10.1146/annurev.py.09.090171.001423
Fu D, Uauy C, Distelfeld A et al (2009) A kinase-START gene confers temperature-dependent resistance to wheat stripe rust. Science 323:1357–1360. https://doi.org/10.1126/science.1166289
Gallie DR (2015) Ethylene receptors in plants-why so much complexity? F1000Prime Reports. https://doi.org/10.12703/P7-39
Gou J-Y, Li K, Wu K et al (2015) Wheat stripe rust resistance protein WKS1 reduces the ability of the thylakoid-associated ascorbate peroxidase to detoxify reactive oxygen species. Plant Cell 27:1755–1770. https://doi.org/10.1105/tpc.114.134296
Hu H, Gutierrez-Gonzalez JJ, Liu X et al (2020) Heritable temporal gene expression patterns correlate with metabolomic seed content in developing hexaploid oat seed. Plant Biotechnol J 18:1211–1222. https://doi.org/10.1111/pbi.13286
Huang Y-F, Poland JA, Wight CP et al (2014) Using genotyping-by-sequencing (GBS) for genomic discovery in cultivated oat. PLoS ONE 9:e102448
IONC (2021) International Oat Nomenclature Committee Meeting Minutes. Resolution on oat chromosome nomenclature. https://wheat.pw.usda.gov/GG3/oatnomenclature
Joehanes R, Nelson JC (2008) QGene 4.0, an extensible Java QTL-analysis platform. Bioinformatics 24:2788–2789. https://doi.org/10.1093/bioinformatics/btn523
Kang J, Park J, Choi H et al (2011) Plant ABC transporters. Arab B 9:e0153–e0153. https://doi.org/10.1199/tab.0153
Kebede AZ, Friesen-Enns J, Gnanesh BN et al (2019) Mapping oat crown rust resistance gene Pc45 confirms association wit PcKM. G3 Genes|genomes|genetics 9(2):505–511. https://doi.org/10.1534/g3.118.200757
Klos KE, Yimer BA, Babiker EM et al (2017) Genome-wide association mapping of crown rust resistance in oat elite germplasm. Plant Genome 10:1–13. https://doi.org/10.3835/plantgenome2016.10.0107
Krattinger SG, Lagudah ES, Spielmeyer W et al (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science (80-) 323:1360–1363
Latta RG, Bekele WA, Wight CP, Tinker NA (2019) Comparative linkage mapping of diploid, tetraploid, and hexaploid Avena species suggests extensive chromosome rearrangement in ancestral diploids. Sci Rep 9:12298. https://doi.org/10.1038/s41598-019-48639-7
Leonard KJ (2002) Oat lines with effective adult plant resistance to crown rust. Plant Dis 86:593–598. https://doi.org/10.1094/PDIS.2002.86.6.593
Li Z, Möttönen J, Sillanpää MJ (2015) A robust multiple-locus method for quantitative trait locus analysis of non-normally distributed multiple traits. Heredity (edinb) 115:556–564. https://doi.org/10.1038/hdy.2015.61
Lin Y, Gnanesh BN, Chong J et al (2014) A major quantitative trait locus conferring adult plant partial resistance to crown rust in oat. BMC Plant Biol 14:250. https://doi.org/10.1186/s12870-014-0250-2
Lin Y, Jiang X, Hu H et al (2021) QTL mapping for grain number per spikelet in wheat using a high-density genetic map. Crop J. https://doi.org/10.1016/j.cj.2020.12.006
Liu W-Y, Kang J-H, Jeong H-S et al (2014) Combined use of bulked segregant analysis and microarrays reveals SNP markers pinpointing a major QTL for resistance to Phytophthora capsici in pepper. Theor Appl Genet 127:2503–2513. https://doi.org/10.1007/s00122-014-2394-8
Maughan PJ, Lee R, Walstead R et al (2019) Genomic insights from the first chromosome-scale assemblies of oat (Avena spp.) diploid species. BMC Biol. https://doi.org/10.1186/s12915-019-0712-y
McCartney CA, Stonehouse RG, Rossnagel BG et al (2011) Mapping of the oat crown rust resistance gene Pc91. Theor Appl Genet 122:317–325. https://doi.org/10.1007/s00122-010-1448-9
McMullen MS, Doehlert DC, Miller JD (2005) Registration of ‘HiFi’ oat. Crop Sci 45:1664. https://doi.org/10.2135/cropsci2005.003
Meng L, Li H, Zhang L, Wang J (2015) QTL IciMapping: Integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop J 3:269–283. https://doi.org/10.1016/j.cj.2015.01.001
Menzies JG, Xue A, Gruenke J et al (2019) Virulence of Puccinia coronata var avenae f. sp. avenae (oat crown rust) in Canada during 2010 to 2015. Can J Plant Pathol 41:379–391. https://doi.org/10.1080/07060661.2019.1577300
Miller ME, Nazareno ES, Rottschaefer SM et al (2021) Increased virulence of Puccinia coronata f. sp. avenae populations through allele frequency changes at multiple putative Avr loci. PLOS Genet 16:e1009291
Money D, Gardner K, Migicovsky Z et al (2015) LinkImpute: fast and accurate genotype imputation for nonmodel organisms. G3 Genes|genomes|genetics 5(11):2383–2390. https://doi.org/10.1534/g3.115.021667
Moore J, Herrera-Foessel S, Lan C et al (2015) A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat Genet 47:1494–1498
Murphy HC (1935) Physiologic specialization in Puccinia coronata avenae. US Dep Agric Tech Bull 433
Nazareno E, Feng L, Madeleine S et al (2017) Puccinia coronata f. sp. avenae: a threat to global oat production. Mol Plant Pathol 19:1047–1060. https://doi.org/10.1111/mpp.12608
Nikoloudakis N, Katsiotis A (2015) Comparative molecular and cytogenetic methods can clarify meiotic incongruities in Avena allopolyploid hybrids. Caryologia 68:84–91. https://doi.org/10.1080/00087114.2015.1021170
Ouellette LA, Reid RW, Blanchard SG, Brouwer CR (2017) LinkageMapView—rendering high-resolution linkage and QTL maps. Bioinformatics 34:306–307. https://doi.org/10.1093/bioinformatics/btx576
Parisi K, Shafee TMA, Quimbar P et al (2019) The evolution, function and mechanisms of action for plant defensins. Semin Cell Dev Biol 88:107–118. https://doi.org/10.1016/j.semcdb.2018.02.004
Penner GA, Chong J, Wight CP et al (1993) Identification of an RAPD marker for the crown rust resistance gene Pc68 in oats. Genome 36:818–820. https://doi.org/10.1139/g93-108
PepsiCo (2020) OT3098 Hexaploid Oat v1. https://wheat.pw.usda.gov/jb/?data=/ggds/oat-ot3098-pepsico. Accessed 12 Nov 2020
Periyannan S, Milne R, Figueroa M et al (2017) An overview of genetic rust resistance: from broad to specific mechanisms. PlOS Pathog. https://doi.org/10.1371/journal.ppat.1006380
Peterson RF, Campbell AB, Hannah AE (1948) A diagrammatic scale for estimating rust intensity on leaves and stems of cereals. Can J Res 26c:496–500. https://doi.org/10.1139/cjr48c-033
Pilet-Nayel M-L, Moury B, Caffier V et al (2017) Quantitative resistance to plant pathogens in pyramiding strategies for durable crop protection. Front Plant Sci 8:1838
Portyanko VA, Chen G, Rines HW et al (2005) Quantitative trait loci for partial resistance to crown rust, Puccinia coronata, in cultivated oat, Avena sativa L. Theor Appl Genet 111:313–324. https://doi.org/10.1007/s00122-005-2024-6
Rinaldo A, Gilbert B, Boni R et al (2017) The Lr34 adult plant rust resistance gene provides seedling resistance in durum wheat without senescence. Plant Biotechnol J 15:894–905. https://doi.org/10.1111/pbi.12684
Rines HW, Porter H, Carson M, Ochocki G (2007) Introgression of crown rust resistance from diploid oat Avena strigosa into hexaploid cultivated oat A. sativa by two methods: direct crosses and through an initial 2x+4x synthetic hybrid. Euphytica 158:67–79
Rines HW, Miller ME, Carson M et al (2018) Identification, introgression, and molecular marker genetic analysis and selection of a highly effective novel oat crown rust resistance from diploid oat, Avena strigosa. Theor Appl Genet 131:721–733. https://doi.org/10.1007/s00122-017-3031-0
Rooney WL, Rines HW, Phillips RL (1994) Identification of RFLP markers linked to crown rust resistance genes Pc91 and Pc92 in oat. Crop Sci 34:940–944. https://doi.org/10.2135/cropsci1994.0011183X003400040019x
Sekhwal MK, Li P, Lam I et al (2015) Disease resistance gene analogs (RGAs) in plants. Int J Mol Sci 16:19248–19290. https://doi.org/10.3390/ijms160819248
Shimizu N, Ishida T, Yamada M et al (2015) BAM 1 and RECEPTOR-LIKE PROTEIN KINASE 2 constitute a signaling pathway and modulate CLE peptide-triggered growth inhibition in Arabidopsis root. New Phytol 208(4):1104–1113. https://doi.org/10.1111/nph.13520
Shin J-H, Blay S, Graham J, McNeney B (2006) LDheatmap: an R function for graphical display of pairwise linkage disequilibria between single nucleotide polymorphisms. J Stat Soft. https://doi.org/10.18637/jss.v016.c03
Simons M (1985) Crown rust. In: Roelfs A, Bushnell W (eds) The cereal rusts, vol 2. Academic Press Inc, Orlando
Sowa S, Paczos-Grzęda E (2020) Identification of molecular markers for the Pc39 gene conferring resistance to crown rust in oat. Theor Appl Genet 133:1081–1094. https://doi.org/10.1007/s00122-020-03533-z
Tinker NA, Chao S, Lazo GR et al (2014) A SNP genotyping array for hexaploid oat. Plant Genome. https://doi.org/10.3835/plantgenome2014.03.0010
Wangkumhang P, Chaichoompu K, Ngamphiw C et al (2007) WASP: a web-based allele-specific PCR assay designing tool for detecting SNPs and mutations. BMC Genomics 8:275. https://doi.org/10.1186/1471-2164-8-275
Wight CP, O’Donoughue LS, Chong J et al (2004) Discovery, localization, and sequence characterization of molecular markers for the crown rust resistance genes Pc38, Pc39, and Pc48 in cultivated oat (Avena sativa L.). Mol Breed 14:349–361. https://doi.org/10.1007/s11032-004-0148-z
Yang L, Zhao D, Meng Z et al (2020) QTL mapping for grain yield-related traits in bread wheat via SNP-based selective genotyping. Theor Appl Genet 133:857–872. https://doi.org/10.1007/s00122-019-03511-0
Zeng ZB (1994) Precision mapping of quantitative trait loci. Genetics 136:1457–1468
Zhai C, Lin F, Dong Z et al (2011) The isolation and characterization of Pik, a rice blast resistance gene which emerged after rice domestication. New Phytol 189:321–334. https://doi.org/10.1111/j.1469-8137.2010.03462.x
Zhao J, Kebede AZ, Bekele WA et al (2020) Mapping of the oat crown rust resistance gene Pc39 relative to single nucleotide polymorphism markers. Plant Dis 104:1507–1513. https://doi.org/10.1094/PDIS-09-19-2002-RE
Zhu S, Kaeppler H (2003) Identification of quantitative trait loci for crown rust in oat line MAM17-5. Crop Sci 43:358–366
Zou C, Wang P, Xu Y (2016) Bulked sample analysis in genetics, genomics and crop improvement. Plant Biotechnol J 14:1941–1955. https://doi.org/10.1111/pbi.12559
Acknowledgements
We thank Dr. Marty Carson for identifying the APR donors and developing the mapping populations, and Dr. Shiaoman Chao, Ms. Mary Osenga and Dr. Howard Rines for technical support in genotyping and allele calling. We also acknowledge Mr. Roger Caspers for his valuable help and assistance in the greenhouse and field experiments. This study was funded by USDA-ARS, a graduate student grant from the North Central Region-Sustainable Agriculture, Research, and Education (NCR-SARE) program, MnDRive Global Food Ventures Fellowship, and Doctoral Dissertation Fellowship from the University of Minnesota Graduate School.
Funding
This study was funded by the USDA-ARS. Partial financial support was received from North Central Region-Sustainable Agriculture, Research, and Education (NCR-SARE), MnDRive Global Food Ventures Fellowship, and Doctoral Dissertation Fellowship from the University of Minnesota Graduate School.
Author information
Authors and Affiliations
Contributions
Conceptualization: SK, MF; Methodology: EN, MM, JF; Writing–original draft preparation: Eric Nazareno; Writing–reviewing and editing: all authors; Funding acquisition: SK, MF, EN; Resources: SK, JF; Supervision: SK, MF.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare.
Additional information
Communicated by Xiaoquan Qi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nazareno, E.S., Fiedler, J., Miller, M.E. et al. A reference-anchored oat linkage map reveals quantitative trait loci conferring adult plant resistance to crown rust (Puccinia coronata f. sp. avenae). Theor Appl Genet 135, 3307–3321 (2022). https://doi.org/10.1007/s00122-022-04128-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-022-04128-6