Abstract
Key message
Introgressing one-eighth of synthetic hexaploid wheat genome through a double top-cross plus a two-phase selection is an effective strategy to develop high-yielding wheat varieties.
Abstract
The continued expansion of the world population and the likely onset of climate change combine to form a major crop breeding challenge. Genetic advances in most crop species to date have largely relied on recombination and reassortment within a relatively narrow gene pool. Here, we demonstrate an efficient wheat breeding strategy for improving yield potentials by introgression of multiple genomic regions of de novo synthesized wheat. The method relies on an initial double top-cross (DTC), in which one parent is synthetic hexaploid wheat (SHW), followed by a two-phase selection procedure. A genotypic analysis of three varieties (Shumai 580, Shumai 969 and Shumai 830) released from this program showed that each harbors a unique set of genomic regions inherited from the SHW parent. The first two varieties were generated from very small populations, whereas the third used a more conventional scale of selection since one of bread wheat parents was a pre-breeding material. The three varieties had remarkably enhanced yield potential compared to those developed by conventional breeding. A widely accepted consensus among crop breeders holds that introducing unadapted germplasm, such as landraces, as parents into a breeding program is a risky proposition, since the size of the breeding population required to overcome linkage drag becomes too daunting. However, the success of the proposed DTC strategy has demonstrated that novel variation harbored by SHWs can be accessed in a straightforward, effective manner. The strategy is in principle generalizable to any allopolyploid crop species where the identity of the progenitor species is known.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bread wheat (Triticum aestivum) arose from an alloploidization event that occurred around 10,000 years ago and involved the tetraploid species T. turgidum and the diploid species Aegilops tauschii (Kihara 1944; McFadden and Sears 1944). Since that time, the widespread dispersal from the site of origin in the Fertile Crescent, in conjunction with intensive selection, first by early farmers and later through deliberate breeding, resulted in adaptation to a wider range of environments than occupied by any other crop species (Dubcovsky and Dvorak 2007). While breeding over the past century has delivered a notable increase in yield potential, a yield plateau now seems to have been reached, leading to concerns that the expanding global demand for wheat will not be easily met without breeding innovations (Hawkesford et al. 2013). The consensus is that it will be necessary to expand the germplasm base to meet this challenge (Tester and Langridge 2010; Moore 2015).
It has been recognized for some time that use of the bread wheat secondary gene pool, represented by the progenitor species, has the potential to introduce a substantial amount of novel genetic variation for exploitation by wheat breeders. Given that the species is derived from a relatively small number of founder amphiploids, recreating these hybrids by crossing T. turgidum with Ae. tauschii to produce so-called synthetic hexaploid wheats (SHWs) should offer a major opportunity to widen the genetic base of the crop (McFadden and Sears 1944; Mujeeb-Kazi et al. 1996). Efforts in this direction have confirmed that use of SHWs as breeding parents can enhance yields across a diverse range of environments (Hoisington et al. 1999; Coghlan 2006; Warburton et al. 2006; Dreisigacker et al. 2008; Trethowan and Mujeeb-Kazi 2008). The major disadvantage of SHWs is that they inevitably harbor genes that negatively impact key domestication traits (Zhang et al. 2013). As a result, the contribution of SHWs to elite commercial varieties remains relatively minor (Yang et al. 2009; Ogbonnaya et al. 2013; Börner et al. 2015). Here, we present a simple, reproducible breeding method that is designed to enhance yield based on the idea of selection of genomic segments inherited from an SHW parent.
Materials and methods
Plant materials
SHW-L1 was bred from a chromosome-doubled amphiploid between Chinese T. turgidum ssp. turgidum landrace AS2255 as female parent and Iranian Ae. tauschii ssp. tauschii accession AS60 as the male parent (Zhang et al. 2004). AS60 is quite distinct from bread wheat D genome, as shown by previous phylogenetic (Wang et al. 2013) and gene expression analyses (Hao et al. 2017; Ramírez-González et al. 2018). An F7 generation recombinant inbred line (RIL) population from the cross SHW-L1 × Chuanmai 32 was previously described by Yu et al. (2014).
The double top-cross and two-phase selection strategy
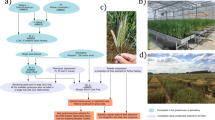
Double top-cross (DTC) was used to establish F1 populations that retain only theoretical 12.5% of the nucleus genome of the synthetic wheat SHW-L1 (Fig. 1). Hybrid seeds for each of the first top-cross combinations were planted in one 2 m row, and one to three hybrid plants resistant to stripe rust were randomly chosen for the second top-cross. The bulked hybrid seeds formed DTC F1 populations; those grown in seasonal nurseries were selected for stripe rust resistance but there was no selection in off-season nurseries. DTC F2 bulks were harvested from all retained F1s.
Double top-cross (DTC) strategy used to introgress genes from the SHW selection SHW-L1. The expected outcome of the crosses (DTC F1) is a genetic background in which 87.5% of the sequence derives from three bread wheat parents indicated by BW1–BW3. Selection in the early generations (F2–F3) focused on short stature, early maturity, resistance to stripe rust and soft glumes. In the later generations (F4–Fn), selection was imposed on yield and its components. Elite selections were used in yield tests. The spikes were taken from SHW-L1 (6x) and its T. turgidum (4x) and Ae. tauschii (2x) parents
Two-phase selection (2PS) was used to reduce population size. The first phase was used in F2–F3 generations to eliminate serious defects of synthetic wheat, while the second phase was used in F4–Fn to improve yields (Fig. 1). The full selection process is outlined in Table S1. All F2 plants were planted in a seasonal nursery, with selection aimed at eliminating four SHW-L1 traits: tough glumes, late maturity, tall stature and stripe rust susceptibility. 100–200 F2 individuals were typically screened, with larger populations for crosses involving less established varieties. F3 bulks were planted in either seasonal or off-season nurseries, with selection repeated against the same SHW-L1 traits in seasonal nurseries. In addition, plants with low fertility, non-uniform tiller length and short upper internodes were discarded, and all retained plants bulked. From F4 onward, planting was restricted to seasonal nurseries. Individual plant selections focused on yield components including spikes per plant, grains per spike and 1000 grain weight (TGW).
Seasonal plantings were sown at the Dujiangyan Experimental Station (31°01′N, 103°32′W) and at the Wenjiang Experimental Station (30°36′N, 103°41′W) in 2 m rows containing 20 plants (Table S1), with each row separated from its neighbor by 30 cm. The stripe rust susceptible variety SY95-71 was planted either side of each experimental row and inoculated with a mixture of locally predominant races. Off-season (summer) plantings of the F1 and F3 materials were conducted at Maerkang (31°92′N, 102°13′W, ~ 2600 m a.s.l): There, ~ 50 plants were sown in each 2.0 m row, with an inter-row separation of 30 cm.
Yield testing
Elite selections were tested at Wenjiang Experimental Station, with management according to local methods. A small number of lines were entered into official uniformity variety trials, including regional variety tests (RVTs) run over two cropping seasons and province productivity trials (PTs) over one cropping season. The PT trial required a plot size ten times the size of the RVT trial and was intended to replicate typical farmer conditions. The official experiments involving Shumai 580 were conducted in Yunnan province in 2013–2015, while the experiments of Shumai 969 and Shumai 830 were carried out in Sichuan province in 2010–2013 and 2014–2017, respectively.
Fluorescence in situ hybridization (FISH) karyotyping
FISH was used to identify gross chromosomal variants and to investigate the transmission of certain specific chromosomal segments. The FISH methodology used followed Komuro et al. (2013) and Zhao et al. (2018), utilizing as probes FAM- or TAMRA-labeled (CTT)10, pSc119.2, pTa-535, pTa71 and pTa-713, all synthesized by TSINGKE Biological Technology Company (Chengdu, Sichuan, China).
Single-nucleotide polymorphism (SNP) genotyping
Genomic DNA was isolated from seedling leaves using a Plant Genomic DNA Kit (TIANGEN Biotech (Beijing) Co., Ltd.). Chip-based genotyping was performed by CapitalBio Corporation (www.capitalbio.com) using a wheat 55 K SNP array, following the Affymetrix Axiom 2.0 Assay Manual Workflow protocol. The flanking sequences for each SNP allowed the location of most of the assays to be mapped onto the bread wheat reference sequence (https://urgi.versailles.inra.fr/download/iwgsc/IWGSC_RefSeq_Assemblies/v1.0/), by imposing a BLASTN E-value threshold of 10−10 and allowing a maximum mismatch of one base. The SNP genotyping was used in phylogenetic analysis following Suzuki and Shimodaira (2006). The distribution of missing markers on each chromosome was analyzed. The ratios of missing markers within neighboring 10 Mb intervals along individual chromosomes were calculated.
Analysis of introgressed segments
SNP markers that were heterozygous in either the parents or the final varieties were removed, and the remaining markers used to calculate parental contributions to the three varieties and to characterize SHW-L1 introgressions. Introgressions were estimated by computing the ratios of same SNP to the total SNPs scored between a variety and its four parents using a sliding window of 10 Mb and step length of 1 Mb and considered to run from the first window to the start of the last consecutive window that had the highest ratio of SHW-L1 SNPs. Graphical representations were constructed using the R package ggplot2 (v.2.2.1) (Wickham 2016). The gene sequences of known major genes determining yield-related traits (Nadolska-Orczyk et al. 2017) and aspects of flour quality were downloaded from GenBank and located on the whole-genome reference sequence. A subset was PCR-amplified from genomic DNA and amplicons sequenced by the TSINGKE Biological Technology Company; primer information is in Table S2. SDS-PAGE was used to identify the composition of high molecular weight glutenin subunits (HWM-GSs), as described by Wan et al. (2005).
Quantitative trait loci (QTL) analysis
The RIL population created from the cross SHW-L1 × Chuanmai 32 was used in a QTL analysis. The relevant phenotypic data were gathered from experiments carried out over the period 2008–2011 (Yu et al. 2014) and were supplemented by fresh data collected in the 2014–2017 at Wenjiang and 2017 at Beijing. QTL were identified using QTL IciMapping v4.1 software (Li et al. 2007) (www.isbreeding.net). Threshold values were calculated using 1000 permutations, assuming a 0.05 type I error. QTL lying within a 50 Mb region defined by flanking markers were used to estimate possible QTL introgressions in the new varieties. A introgression was considered if the QTL region overlapped with an introgressed SHW-L1 region detected by sliding window analysis.
Statistical analysis
All statistical analyses were performed using programs implemented in either R (v3.4.4) software or in Microsoft Excel 2007.
Results
Yield performance of the new varieties
A total of 67 DTC F1 combinations, involving 51 bread wheat lines (ten in 2003–2004 and 57 in 2005–2006), were made. Three commercial varieties have been released from three combinations (Table S1): These have been named Shumai 580, Shumai 969 and Shumai 830. The first two of these were generated from relatively small populations (respectively, ca. 650 and 1150 F1–F4 plants), whereas the third required a more conventional scale of selection (ca. 6360 plants). The three varieties differed with respect to spike size and their grain characteristics (Fig. 2a, b) and were superior in yield to other leading genotypes (Fig. 2c–e).
Spikes and grain of the three Shumai varieties and their yield potential compared to current elite genotypes. a Shumai 830 formed long spikes. b Shumai 830 set large grains. c Shumai 580 outperformed the yield of five check varieties by at least 30% in a Yunnan province productivity trial conducted in 2015–2016, according to the official trial data provided by the Yunnan Seed Administration Station. d Shumai 969 out-yielded 99 current varieties in the Sichuan provincial productivity trials conducted over the period 2007–2017, according to the official trial data provided by the Sichuan Seed Administration Station. e Shumai 830 produced a larger grain mass per spike compared to 99 current varieties in Sichuan provincial regional variety trials conducted over the period 2007–2017
Yield superiority of Shumai 580 was particularly clear in the Yunnan province productivity trial (PT), where it outperformed five locally bred varieties by at least 30% and the commercial check cultivar Yunmai 54 by 56% (Fig. 2c). Shumai 969 was the only variety yielding over 6 t/ha in Sichuan province productivity trials conducted during 2007–2017 (Fig. 2d). The high yield potential of Shumai 830 was mainly attributed to high grain weight. The Sichuan regional variety test (RVT) data collected for 100 varieties released over the past decade showed that Shumai 830 spikes have been consistently heavier than those of other varieties (Fig. 2e). The high yield potential of the other two varieties also relates to improvements in TGW (Table S3). Shumai 969 and Shumai 830 produced grain with a relatively high TGW, even though the anthesis to maturity period was relatively short, indicating rapid grain filling.
The genotypic composition of the three Shumai varieties
FISH karyotyping results (Fig. S1) indicate that the 1BL/1RS wheat/rye translocation present in some of the parents was not transmitted to these varieties, but that chromosome segments from SHW-L1 were present in some (2DS) or all (5DS).
In all, 51,160 SNP sequences were mapped onto Chinese Spring reference sequence: 17,879 mapped to A genome sites, 18,203 to B genome sites and 15,078 to D genome sites. Except for chromosome 4D, which was marked by 1087 SNP loci, each chromosome had at least 2000 markers. On the basis of allelic status at the 51,160 SNP sites, it was clear that SHW-L1 is genetically very distant from any of the bread wheat varieties used as parents in the DTC (Fig. S2). This is particularly applied to the D genome, as the Axiom assay returned missing data for a large number of SNP loci in SHW-L1 (Fig. S3).
The number of SHW-L1 alleles present in the three varieties represent 17.2% (Shumai 580), 13,4% (Shumai 830) and 13.8% (Shumai 969) of the scored SNP loci (Table S4). These were unevenly distributed across the wheat genome, with only 69 loci, including a block of 56 on chromosome 5D, present in all three varieties (Fig. 3a, b).
SHW-L1 SNP alleles and graphical genotypes of three new Shumai varieties. a Number of SHW-L1 SNP alleles presented in Shumai 580 (red circle), Shumai 830 (green circle) and Shumai 969 (blue) according to the A (left), B (middle), D (right) genomes. b SHW-L1 introgressed segments in the three varieties. The red arrowheads indicate the centromeres. The black arrowheads indicate major genes carrying alleles inherited from the SHW-L1 parent. The colored boxes refer to introgressions from the donor SHW-L1 into Shumai varieties, for instance green for Shumai 580 and black for the shared introgression by three varieties (color figure online)
The number of introgressed segments per variety varied from 57 to 83 (Fig. 3b; Table S5), most of which were shorter than 30 Mb in length. Assuming an overall genome size of 14.05 Gb (IWGSC 2018), the overall ratio of SHW-L1 DNA retained by Shumai 969 was 12.4%, and the ratios for Shumai 830 and Shumai 580 were, respectively, 13.5 and 15.0% (Table S5).
QTL analysis on a SHW-L1/Chuanmai 32 RIL population
We detected 86 QTL influencing 12 yield-related traits from an analysis of a SHW-L1/Chuanmai 32 RIL population (Table S6). The 86 QTL mapped to 19 of the 21 chromosomes (the exceptions were chromosomes 5D and 7D) (Table S7) were particularly frequent on chromosomes 5A (17 loci) and 2D (11 loci). On the basis of a box plot analysis, the SHW-L1 Vrn-A1 allele on 5A appeared to hasten anthesis, whereas the Ppd-D1 allele on 2D delayed it (Fig. S4). In addition, the presence of Vrn-B1 allele in SHW-L1 was also associated with a QTL for early heading (Table S7). One half of the QTL mapped to sites on A genome chromosomes. Interesting, 40 out of 86 favorable yield-related QTL alleles were inherited from SHW-L1.
Major genes and QTL introgressed from SHW-L1
All three Shumai varieties inherited Ppd-D1, Rht-B1 and Rht-D1 alleles from a bread wheat parent (Table 1). However, Shumai 969 inherited Ppd-A1 and Vrn-B1 alleles from SHW-L1, whereas Shumai 830 inherited a Vrn-A1 allele from the synthetic parent (Fig. 3b). Shumai 830 inherited TaTEF and GPC-2 alleles from SHW-L1, while Shumai 969 inherited TaGASR7, TaCKX6 and TaSus1 and Shumai 580 inherited TaGW2 and TaGASR7 alleles from SHW-L1. In addition, Shumai 830 and Shumai 580 retained the SHW-L1 Glu-B1 allele and Shumai 969 harbored the SHW-L1 Glu-D1 allele, which encode endosperm proteins affecting flour quality.
The analysis was then extended to the 86 QTL detected from the SHW-L1/Chuanmai 32 RIL population (Table S7). Of these, 37 were introgressed into the three Shumai varieties. The presence of each of the Vrn-A1 and Vrn-B1 alleles in SHW-L1 was associated with a QTL for early heading (Table S7), and its Glu-B1 allele was associated with a QTL for early heading. There were no favorable QTL associated with any of the other introgressed major genes referred to above (Table 1), probably caused by the interference of SHW-L1 traits such as tough glumes on QTL detection.
The numbers of SHW QTL contributing negatively to traits as predicted from the RIL analysis were nine of 20 in Shumai 580, ten of 20 in Shumai 830 and four of seven in Shumai 969 (Table S7). Three positive SHW-L1 QTL for spike length (on chromosomes 2A and 5A) and one for spikelet number on chromosome 2D were inherited by Shumai 830.
Discussion
Unadapted germplasm including SHWs have been used as resources of new variations for wheat improvement. A few large-scale pre-breeding programs have been launched for this goal, such as the Wheat Pre-breeding Project (2011), the Wheat Improvement Strategic Programme (Moore 2015) and CIMMYT’s Seeds of Discovery (Singh et al. 2018). Typically, SHWs have been crossed and then backcrossed (A/B//B) or top-crossed (A/B//C) to wheat varieties or elite lines to produce breeding populations from which new elite lines, or in a few cases varieties, have been selected (Yang et al. 2009; Ogbonnaya et al. 2013). For example, this strategy has been successfully used in China, and four SHW-derived varieties were released in 2003–2005. One of these (Chuanmai 42) has proven to be both a high yielding variety and an outstanding crossing parent, from which 12 commercially released varieties (including Shumai 969) have been bred (Li et al. 2014).
Compared to the top-cross (A/B/C), DTC (A/B/C/D) populations with one more cross have an reduction of synthetic wheat genome from theoretical 25.0 to 12.5%. However, our breeding using a small population size demonstrates that DTC is highly efficient to enhance high yielding potentials. A factor beneficial to the success is that synthetic wheat has a high frequency of favorable alleles across the genome. In our analysis on a RIL population between synthetic wheat SHW-L1 and a variety, although the expectation was that favorable yield-related QTL alleles would be less likely associated with SHW-L1, almost half (40 out of 86) of the loci behaved in this way. Similar phenomenon was also found in synthetic wheat using the advanced backcross populations (Huang et al. 2003, 2004). Meanwhile, DTC populations are involved in three modern varieties, so that progenies have high possibility to combine different allelic variations between common wheat and synthetic wheat. Each of the released Shumai varieties harbors a unique set of genomic regions inherited from the synthetic wheat parent, highlighting the significance of synergistic recombination of yield-related alleles/traits.
The major contribution made by SHW-L1 to the high yield potential of three Shumai varieties relates to improvements in TGW. Additionally, some genes directly affecting TGW (Table 1) and phenology (and therefore the genes that underlie it) also have an impact. The most well-documented genes influencing growth duration are the Vrn (response to vernalization), Ppd (response to day length) and Eps (earliness per se, or the developmental rate) genes (Snape et al. 2001; Li et al. 2017). Growth duration genes act pleiotropically on a number of yield-related traits (Snape et al. 1985; Börner et al. 1993). The grain filling duration exhibited by SHW-L1 was relatively rapid (Yu et al. 2015). The three Shumai varieties reached maturity as early as, if not earlier, than conventional varieties, but produced heavier grains (Table S3).
The signature trait of Shumai 830—the ability to form heavy spikes—was achieved by combining high TGW with a large number of grains per spike. This outcome suggests a pathway for producing varieties with heavy spikes, in which plant phenology is coordinated with other key traits (Fig. 4). The presence of the SHW-L1 Vrn-A1 allele was associated with earlier flowering, the introduction of which probably reduced the requirement for vernalization, thereby resulting in a genotype producing a less than the optimal number of spikes per unit area. On the other hand, more time would then be available for the development and growth of individual spikes. Shumai 830 also harbored QTL, mapping to chromosomes 2A, 2D and 5A, which acted to increase both the length of the spike and number of spikelets per spike. The high TGW reflected not only the presence of TaTEF-7A and GPC-2 alleles from SHW-L1, but also an ability to fill the grains rapidly.
A model devised to explain the basis of formation of heavy spikes. The high TGW achieved by Shumai 830 was associated with earliness per se (Eps) genes, and grain size genes TaTEF and GPC-2. A high grain number per spike was conferred by the presence of Vrn-A1 allele from the synthetic wheat SHW-L1, along with QTL that increased the spike length and number of spikelets per spike. The presence of a Vrn-A1 from SHW-L1 probably acted to reduce the number of spikes formed per unit area
The three Shumai varieties had remarkably enhanced yield potential compared to those developed by conventional breeding. SNP-based genotyping showed that each had retained a spectrum of both beneficial and deleterious alleles from SHW-L1. Studies in tomato (Tanksley et al. 1996; Tanksley and Nelson 1996), rice (Xiao et al. 1998) and wheat (Huang et al. 2003, 2004; Pestsova et al. 2006) have confirmed that wild materials can harbor beneficial alleles, along with deleterious ones, as was also clearly the case for SHW-L1. The distinct spectrum of SHW-L1 alleles harbored by the three varieties suggests that progeny of crosses between them could enable combination of a greater number of beneficial SHW-L1 alleles, offering the prospect of further genetic advancement. The positive outcomes delivered by applying the DTC strategy imply that the secondary gene pool has great potential to contribute to wheat improvement.
Many of our leading crop species are allopolyploids and have become genetically isolated from their progenitor species; the effect of domestication and subsequent selection and breeding has been to drastically erode their genetic base (Stebbins 1950; Buckler et al. 2001). The present research has shown that a relatively simple crossing strategy can unlock the secondary gene pool of wheat, and by inference, a similar strategy could be applied to other allopolyploid crops species (such as oat, cotton, groundnut and canola) to exploit diversity that is currently unexploited.
References
Börner A, Worland AJ, Plaschke J, Schumann E, Law CN (1993) Pleiotropic effects of genes for reduced height (Rht) and day length insensitivity (Ppd) on yield and its components for wheat grown in middle Europe. Plant Breed 111:204–216
Börner A, Ogbonnaya FC, Röder MS, Rasheed A, Periyannan S, Lagudah ES (2015) Aegilops tauschii introgressions in wheat. In: Molnár-Láng M, Ceoloni C, Doležel J (eds) Alien introgression in wheat. Springer, Cham, pp 245–271
Buckler ES, Thornsberry JM, Kresovich S (2001) Molecular diversity, structure and domestication of grasses. Genet Res 77:213
Coghlan A (2006) Synthetic wheat offers hope to the world. New Scientist 2538
Dreisigacker S, Kishii M, Lage J, Warburton M (2008) Use of synthetic hexaploid wheat to increase diversity for CIMMYT bread wheat improvement. Aust J Agric Res 59:413–420
Dubcovsky J, Dvorak J (2007) Genome plasticity a key factor in the success of polyploid wheat under domestication. Science 316:1862–1866
Hao M, Li A, Shi T, Luo J, Zhang L, Zhang X, Ning S, Yuan Z, Zeng D, Kong X, Li X, Zheng H, Lan X, Zhang H, Zheng Y, Mao L, Liu D (2017) The abundance of homoeologue transcripts is disrupted by hybridization and is partially restored by genome doubling in synthetic hexaploid wheat. BMC Genom 18:149
Hawkesford MJ, Araus JL, Park R, Calderini D, Miralles D, Shen T, Zhang J, Parry MAJ (2013) Prospects of doubling global wheat yields. Food Energy Secur 2:34–48
Hoisington D, Khairallah M, Reeves T, Ribaut JM, Skovmand B, Taba S, Warburton M (1999) Plant genetic resources: what can they contribute toward increased crop productivity? Proc Natl Acad Sci USA 96:5937–5943
Huang XQ, Coster H, Ganal MW, Roder MS (2003) Advanced backcross QTL analysis for the identification of quantitative trait loci alleles from wild relatives of wheat (Triticum aestivum L.). Theor Appl Genet 106:1379–1389
Huang XQ, Kempf H, Ganal MW, Roder MS (2004) Advanced backcross QTL analysis in progenies derived from a cross between a German elite winter wheat variety and a synthetic wheat (Triticum aestivum L.). Theor Appl Genet 109:933–943
International Wheat Genome Sequencing Consortium (IWGSC) (2018) Shifting the limits in wheat 401 research and breeding using a fully annotated reference genome. Science 361:7191
Kihara H (1944) Discovery of the DD-analyser, one of the ancestors of Triticum vulgare. Agric Hortic 19:889–890
Komuro S, Endo R, Shikata K, Kato A (2013) Genomic and chromosomal distribution patterns of various repeated DNA sequences in wheat revealed by a fluorescence in situ hybridization procedure. Genome 56:131–137
Li H, Ye G, Wang J (2007) A modified algorithm for the improvement of composite interval mapping. Genetics 175:361–374
Li J, Wan HS, Yang WY (2014) Synthetic hexaploid wheat enhances variation and adaptive evolution of bread wheat in breeding processes. J Syst Evol 52:735–742
Li G, Boontung R, Powers C, Belamkar V, Huang T, Miao F, Baenziger PS, Yan L (2017) Genetic basis of the very short life cycle of ‘Apogee’ wheat. BMC Genom 18:838
Liu DC, Hao M, Li AL, Zhang LQ, Zheng YL, Mao L (2016) Allopolyploidy and interspecific hybridization for wheat improvement. In: Mason AS (ed) Polyploidy and hybridization for crop improvement. CRC Press, Cambridge, pp 27–52
McFadden ES, Sears ER (1944) The artificial synthesis of Triticum spelta. Rec Genet Soc Am 13:26–27
Moore G (2015) Strategic pre-breeding for wheat improvement. Nat Plant 1:15018
Mujeeb-Kazi A, Rosas V, Roldan S (1996) Conservation of the genetic variation of Triticum tauschii (Coss.) Schmalh. (Aegilops squarrosa auct. non L.) in synthetic hexaploid wheats (T. turgidum L. s. lat. × T. tauschii; 2n = 6x = 42, AABBDD) and its potential utilization for wheat improvement. Genet Resour Crop Evolut 43:129–134
Nadolska-Orczyk A, Rajchel IK, Orczyk W, Gasparis S (2017) Major genes determining yield-related traits in wheat and barley. Theor Appl Genet 130:1081–1098
Ogbonnaya FC, Abdalla OS, Mujeeb-Kazi A, Kazi AG, Xu SS, Gosman N, Lagudah ES, Bonnett D, Sorrells ME, Tsujimoto H (2013) Synthetic hexaploids: harnessing species of the primary gene pool for wheat improvement. In: Janick J (ed) Plant breeding reviews. Wiley, New York, pp 35–122
Pestsova EG, Börner A, Röder MS (2006) Development and QTL assessment of Triticum aestivum–Aegilops tauschii introgression lines. Theor Appl Genet 112:634–647
Ramírez-González RH, Borrill P, Lang D et al (2018) The transcriptional landscape of polyploid wheat. Science 361:662
Singh S, Vikram P, Sehgal D et al (2018) Harnessing genetic potential of wheat germplasm banks through impact-oriented-prebreeding for future food and nutritional security. Sci Rep 8:12527
Snape JW, Law CN, Parker BB, Worland AJ (1985) Genetical analysis of chromosome 5A of wheat and its influence on important agronomic characters. Theor Appl Genet 71:518–526
Snape JW, Butterworth K, Whitechurch E, Worland AJ (2001) Waiting for fine times: genetics of flowering time in wheat. Euphytica 119:185–190
Stebbins GL (1950) Variation and evolution in plants. Columbia University Press, New York
Suzuki R, Shimodaira H (2006) Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics 22:1540–1542
Tanksley SD, Nelson JC (1996) Advanced backcross QTL analysis: a method for the simultaneous discovery and transfer of valuable QTLs from unadapted germplasm into elite breeding lines. Theor Appl Genet 92:191–203
Tanksley SD, Grandillo S, Fulton TM, Zamir D, Eshed Y, Petiard V, Lopez J, Beck-Bunn T (1996) Advanced backcross QTL analysis in a cross between an elite processing line of tomato and its wild relative L. pimpinellifolium. Theor Appl Genet 92:213–224
Tester M, Langridge P (2010) Breeding technologies to increase crop production in a changing world. Science 327:818–822
The Wheat Pro-Breeding Project, Crop Wild Relatives (CWR) (2011) Global Crop Diversity Trust (GCDT). http://www.cwrdiversity.org/partnership/wheat-pre-breeding-project/
Trethowan RM, Mujeeb-Kazi A (2008) A novel germplasm resources for improving environmental stress tolerance of hexaploid wheat. Crop Sci 48:1255–1265
Wan Y, Yan Z, Liu K, Zheng Y, D’Ovidio R, Shewry PR, Halford NG, Wang D (2005) Comparative analysis of the D genome-encoded high-molecular weight subunits of glutenin. Theor Appl Genet 111:1183–1190
Wang J, Luo MC, Chen Z, You FM, Wei Y, Zheng Y, Dvorak J (2013) Aegilops tauschii single nucleotide polymorphisms shed light on the origins of wheat D-genome genetic diversity and pinpoint the geographic origin of hexaploid wheat. New Phytol 198:925–937
Warburton ML, Crossa J, Franco J, Kazi M, Trethowan R, Rajaram S, Pfeiffer W, Zhang P, Dreisigacker S, Van Ginkel M (2006) Bringing wild relatives back into the family: recovering genetic diversity in CIMMYT improved wheat germplasm. Euphytica 149:289–301
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer, New York
Xiao J, Li J, Grandillo S, Ahn SN, Yuan L, Tanksley SD, McCouch SR (1998) Identification of trait-improving quantitative trait loci alleles from a wild rice relative, Oryza rufipogon. Genetics 150:899–909
Yang W, Liu D, Li J, Zhang L, Wei H, Hu X, Zheng Y, He Z, Zou Y (2009) Synthetic hexaploid wheat and its utilization for wheat genetic improvement in China. J Genet Genom 36:539–546
Yu M, Chen GY, Zhang LQ, Liu YX, Liu DC, Wang JR, Pu ZE, Zhang L, Lan XJ, Wei YM, Liu CJ, Zheng YL (2014) QTL mapping for important agronomic traits in synthetic hexaploid wheat derived from Aegiliops tauschii ssp. tauschii. J Inter Agric 13:1835–1844
Yu M, Chen GY, Pu ZE, Zhang LQ, Liu DC, Lan XJ, Wei YM, Zheng YL (2015) Quantitative trait locus mapping for growth duration and its timing components in wheat. Mol Breed 35:44
Zhang L, Liu D, Yan Z, Lan X, Zheng Y, Zhou Y (2004) Rapid changes of microsatellite flanking sequence in the allopolyploidization of new synthesized hexaploid wheat. Sci China C Life Sci 47:553–561
Zhang H, Bian Y, Gou X, Zhu B, Xu C, Qi B, Li N, Rustgi S, Zhou H, Han F, Jiang J, Van Wettstein D, Liu B (2013) Persistent whole-chromosome aneuploidy is generally associated with nascent allohexaploid wheat. Proc Natl Acad Sci USA 110:3447–3452
Zhao L, Ning S, Yi Y, Zhang L, Yuan Z, Wang J, Zheng Y, Hao M, Liu D (2018) Fluorescence in situ hybridization karyotyping reveals the presence of two distinct genomes in the taxon Aegilops tauschii. BMC Genom 19:3
Acknowledgements
The authors thank Chi Yen and Junliang Yang (Sichuan Agricultural University) for suggestions on the use of synthetic wheat; Robert McIntosh (University of Sydney) and Robert Koebner (smartenglish2008@gmail.com) for revising the article; the International Wheat Genome Sequencing Consortium for providing pre-publication access to the RefSeq v1.0 assembly and its annotation; Jizeng Jia (Chinese Academy of Agricultural Sciences) for providing SNP flanking sequences; Peng Qin (Yunnan Agricultural University) for running trials in Yunnan province; and Qiuzhen Jia (Gansu Academy of Agricultural Sciences) for providing Chinese Puccinia striiformis. f. sp. tritici races. This research was financially supported by the Chinese Government National Key Research and Development Program (2016YFD0102000), the National Natural Science Foundation of China (31071420, 30700495, 31671689, 31071418, 30270804, 31601300 and 31661143007), the Sichuan Provincial Agricultural Department Innovative Research Team (wheat-10) and the Sichuan Province Science and Technology Department Crops Breeding Project (2016NYZ0030). MJH and Rothamsted Research is supported via the Designing Future Wheat project (BB/P016855/1) by the UK Biotechnology and Biological Sciences Research Council.
Author information
Authors and Affiliations
Contributions
D.L, L.Q.Z., M.H. and Y.Z. designed the project; D.L., L.Q.Z., M.H., Z.W.Y, S.N., S.D., Z.H.Y., B.W., Y.Z., X.L., H.Z. and L.H. produced the new elite lines; L.B.Z., D.X., Q.L., W.C. and K.Z. performed the FISH and SDS-PAGE analyses, gene isolation and plant-type comparison. M.H, M.Y., Y.W., L.B.Z., A.L. and W.Y. performed the phenotypic and QTL analyses. M.H., D.L., J.W., M.C. and X.C. performed the SNP genotyping and statistical analyses. D.L., M.H., M.K., M.J.H. and L.M wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author declares no competing financial interests.
Additional information
Communicated by Ian Mackay.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hao, M., Zhang, L., Zhao, L. et al. A breeding strategy targeting the secondary gene pool of bread wheat: introgression from a synthetic hexaploid wheat. Theor Appl Genet 132, 2285–2294 (2019). https://doi.org/10.1007/s00122-019-03354-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-019-03354-9