Abstract
Immune checkpoint inhibitors (ICIs) have achieved impressive success in lung adenocarcinoma (LUAD). However, the response to ICIs varies among patients, and predictive biomarkers are urgently needed. PCDH11X is frequently mutated in LUAD, while its role in ICI treatment is unclear. In this study, we curated genomic and clinical data of 151 LUAD patients receiving ICIs from three independent cohorts. Relations between PCDH11X and treatment outcomes of ICIs were examined. A melanoma cohort collected from five published studies, a pan-cancer cohort, and non-ICI-treated TCGA-LUAD cohort were also examined to investigate whether PCDH11X mutation is a specific predictive biomarker for LUAD ICI treatment. Among the three ICI-treated LUAD cohorts, PCDH11X mutation (PCDH11X-MUT) was associated with better clinical response compared to wild-type PCDH11X (PCDH11X-WT). While in ICI-treated melanoma cohort, the pan-cancer cohort excluding LUAD, and the non-ICI-treated TCGA-LUAD cohort, no significant differences in overall survival (OS) were observed between the PCDH11X-MUT and PCDH11X-WT groups. PCDH11X mutation was associated with increased PD-L1 expression, tumor mutation burden (TMB), neoantigen load, DNA damage repair (DDR) mutations, and hot tumor microenvironment in TCGA-LUAD cohort. Our findings suggested that the PCDH11X mutation might serve as a specific biomarker to predict the efficacy of ICIs for LUAD patients. Considering the relatively small sample size of ICI-treated cohorts, future research with larger cohorts and prospective clinical trials will be essential for validating and further exploring the role of PCDH11X mutation in the context of immunotherapy outcomes in LUAD.
Key messages
-
PCDH11X mutation is associated with better clinical response compared to wild type PCDH11X in three ICIs-treated LUAD cohorts.
-
In ICIs-treated melanoma cohort, the pan-cancer cohort excluding LUAD, and non-ICIs-treated TCGA-LUAD cohorts PCDH11X mutation is not associated with better clinical response, suggesting PCDH11X mutation might be a specific biomarker to predict the efficacy of ICIs treatment for LUAD patients.
-
PCDH11X mutation is associated with increased PD-L1 expression, tumor mutation burden, and neoantigen load in TCGA-LUAD cohort.
-
PCDH11X mutation is associated with hot tumor microenvironment in TCGA-LUAD cohort.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is the most commonly diagnosed cancer and the leading cause of cancer-related death worldwide, causing approximately 1.8 million deaths each year [1]. Lung cancer is divided into two broad histological categories: non-small cell lung carcinoma (NSCLC) and small-cell lung carcinoma (SCLC). NSCLC represents more than 80 to 85% of lung cancers. Most common histological subtype of NSCLC is adenocarcinoma (LUAD, 40%), followed by squamous cell carcinoma (LUSC, 25%) [2, 3]. It is widely speculated that LUAD and LUSC arise from distinct cells of origin: LUAD is considered to arise mainly from alveolar epithelial cells, whereas LUSC is from basal cells [4]. However, mixed histology non-small cell lung cancer and histologic transdifferentiation of LUAD to LUSC have been observed [4, 5]. Also, transition of LUSC to LUAD has been reported, although the evidences are limited [6]. Treatment options for LUAD usually include surgery, radiotherapy, chemotherapy, molecularly targeted therapy, and immunotherapy, based on the stage, histology, genetic alterations, and patient’s condition. According to the study in the United States, survival after diagnosis with LUAD has improved substantially along with treatment advances: among men, incidence-based mortality from NSCLC decreased 6.3% annually from 2013 through 2016 [7]. In recent years, immune checkpoint inhibitors (ICIs) targeting programmed cell death (ligand)-1 (PD-1/PD-L1) and cytotoxic T lymphocyte-antigen 4 (CTLA-4) have substantially improved outcomes of NSCLC treatment [8, 9]. However, despite the promising antitumor effects exhibited by ICIs, the objective response rate (ORR) of ICI treatment is only about 20% [10]. Currently, several biomarkers have showed potential to identify patients who response better to ICI treatment. These biomarkers include PD-L1 expression, neoantigen load (NAL), TMB, and DDR pathway. However, it is important to note that the predictive utility of these biomarkers still has some limitations [11]. In light of this, there is an urgent need to discover more potential biomarkers that can effectively screen patients who would derive the greatest therapeutic benefit from ICIs.
Protocadherin 11 X-linked (PCDH11X), a non-clustered δ-protocadherin belonging to the cadherin superfamily, is frequently mutated in lung cancer [12]. Some studies have shown that PCDHs regulate immune processes. For example, PCDH15 is expressed in cytotoxic tumor-derived T- and NK-cell lines instead of normal T, B, or NK lymphocytes [13]. PCDH18 has been reported to interact with p56lck kinase to block proximal TCR signaling and plays a key role to regulate CD8+ tumor-infiltrating T cell function [14, 15]. In addition, cadherin superfamily members FAT1/2/3/4 are proposed to be potential immunological biomarkers to screen non-small cell lung cancer (NSCLC) patients who may benefit from ICI treatment [16, 17]. However, the role of PCDH11X mutation in LUAD patients is not clear.
In this study, we found that PCDH11X mutation was associated with better clinical benefits in three independent ICI-treated LUAD cohorts. Subsequently, we assessed immunogenic features based on PCDH11X status to explore the possible underlying mechanism using TCGA datasets. We found that PCDH11X mutation is associated with higher PD-L1 expression, TMB, neoantigen load, DNA damage repair (DDR) mutations, and hot microenvironment in TCGA-LUAD cohort. Our current evidence suggests that PCDH11X mutation yields promising predictive value for ICI treatment in LUAD patients.
Materials and methods
Clinical cohort
The present study used four ICI-treated cohorts and one non-ICI-treated cohort for analysis (Tables S1 and S2). The ICI-treated cohorts included the following: (1) the Hellmann cohort [18] included 75 LUAD patients received combined anti-PD-1 and CTLA-4 treatment; (2) the Miao cohort [19] included 249 patients (47 LUAD, 202 non-LUAD) with various types of cancer received anti-CTLA-4 and/or anti-PD-L1 or PD-L1 treatment; (3) the Rizvi cohort [20] included 34 NSCLC (29 LUAD, 5 LUSC) received pembrolizumab treatment; and (4) melanoma cohort included 424 patients with melanoma collected from five published studies [21,22,23,24,25]. All these cohorts included response data and mutation data obtained from exome sequencing. TCGA-LUAD cohort was a non-ICI-treated cohort. It included whole exome sequencing data of 567 patients and RNA-seq data of 509 patients. TCGA-LUAD cohort was used to explore the possible underlying mechanism. Data from ICI-treated cohorts was downloaded from cBioPortal (https://www.cbioportal.org/) or supplementary materials from published articles. Data from TCGA-LUAD cohort was downloaded from UCSC-XENA (https://xenabrowser.net/datapages/).
Somatic variant analysis and genomics characteristics
We analyzed the somatic mutation distribution of PCDH11X gene in the Hellmann and TCGA-LUAD cohorts using the R package Maftools [26]. The MAF file was download from cBioPortal and UCSC-XENA database, respectively. The ComplexHeatmap function was used to draw a oncoPrint plot that displayed the top 10 most frequently mutated genes and clinical features for two cohorts [27].
Copy number variation analysis in TCGA
The copy number variation (CNV) data of TCGA-LUAD cohort were downloaded from the GDC portal using R package TCGAbiolinks [28]. GSITIC 2.0 was adopted to detect significant amplified and deletion in genomic region with default parameter (confidence level was 0.9) [29]. Regions with an FDR value less than 0.01 were visualized using R package Maftools.
Assessment of clinical response
The clinical indicators used to assess clinical response of immunotherapy included overall survival (OS), progression-free survival rate (PFS) and objective response rate (ORR). For cohorts with OS available (Miao cohort, Melanoma cohort, TCGA-LUAD cohort), OS was regarded as the main endpoint; otherwise, PFS was regarded as the main endpoint (Hellmann cohort, Rizvi cohort). ORR was defined as the percentage of patients who have confirmed complete response (CR) or partial response (PR) according to RECIST V.1.1. For the ICI-treated cohorts, PFS and OS were calculated from the start date of treatment. For non-ICI-treated cohorts, namely, TCGA-LUAD, OS were calculated from the date of the first diagnosis.
TMB, neoantigen load, PD-L1 gene expression, and DDR pathway gene mutation frequency
Tumor mutation burden (TMB) was defined as the number of mutations (SNV or indels) per million bases (MB) of interrogated genomic sequencing. The TMB of the two groups of PCDH11X-MUT and PCDH11X-WT was compared by Mann–Whitney-Wilcoxon test. Predicted neoantigen load was extracted from a previous TCGA pan-cancer study conducted by Thorsson et al. [30]. PD-L1 expression (FPKM) between PCDH11X-MUT and PCDH11X-WT groups was compared by Mann–Whitney-Wilcoxon test in TCGA-LUAD cohort.
We used the combined ICI-treated dataset (combing Hellmann cohort, Miao cohort, and Rizvi cohort) to perform multivariable Cox regression analysis. TMB above the median was defined as high TMB; otherwise, it was defined as low TMB. PD-L1 expression level data (negative, weak, strong) was available in Rizvi (29 samples) and Hellmann (68 of 75 samples) cohorts. We obtained the data from the supplementary material of the published papers, where PD-L1 expression was assessed by immunohistochemistry, and strong staining represented ≥ 50% PD-L1 expression, weak represented 1 to 49%, and negative represented < 1%.
The gene set related to the DDR pathway came from the Broad Institute MSigDB database (https://www.gsea-msigdb.org/gsea/msigdb/, Table S3). For each sample, we counted the mutation occurred in each DDR pathway. The Mann–Whitney-Wilcoxon test was used to compare the mutation counts of each pathway between the PCDH11X-MUT and PCDH11X-WT groups.
Tumor microenvironment analysis of TGCA-LUAD cohort
The immune infiltration scores of 29 cell types estimated using CIBERSORT were extracted from a previous TCGA pan-cancer study conducted by Thorsson et al. [31]. Eighteen immune signatures collected from published articles were calculated using FPKM values and compared by Mann–Whitney-Wilcoxon test (Table S4). R package edgeR [32] was used to perform differentially expressed gene (DEG) analysis between PCDH11X-WT (408 samples) and PCDH11X-MUT (101 samples) groups in TCGA-LUAD cohort using htseq-count data. Genes with FDR value less than 0.05 were considered as DGEs. The differences in the expression levels of 78 immune-related genes defined by Thorsson et al. [31] were also studied.
Gene set variation analysis (GSVA)
Hallmark and KEGG pathway gene sets were downloaded from the molecular signature database MSigDB (http://software.broadinstitute.org/gsea/msigdb). The gene set variance analysis (GSVA) score of each gene set for each sample in TCGA-LUAD cohort was obtained using the R package GSVA. The GSVA score could represent the degree of enrichment of gene sets. Then, R package limma was used to compare the GSVA score between PCDH11X-MUT and PCDH11X-WT groups. Gene sets with adjusted P value lower than 0.05 were considered as significantly different between groups.
Statistical analysis
Associations between the PCDH11X mutation status and OS or PFS were analyzed via the Kaplan–Meier method using R packages survminer; survival curves were compared via the log-rank test. Multivariate Cox regression analysis was performed in the combined ICI-treated cohort, adjusting factors including age, gender, smoking history, TMB, and PD-L1 level. Statistical analysis for comparisons between two groups was conducted using the Wilcoxon test. R software (version 3.6.3) was applied to perform all statistical analyses, and P values were two-tailed. A P value < 0.05 was considered to indicate significance.
Results
Landscapes of PCDH11X gene mutations in LUAD
We first explored the mutation distribution located in the protein structures and somatic mutation rate of PCDH11X gene in the Hellmann and TCGA cohort (Fig. 1A, B). The somatic mutation rates in the patients from Hellmann cohort were 13.33% and 14.64% in the TCGA-LUAD cohort. The most frequent somatic mutation of PCDH11X gene in both two cohorts was missense mutations spanned entire gene, suggesting that it may function as a tumor suppressor in LUAD as previously reported [12]. Then, we divided patients into PCDH11X-MUT (mutant type) and PCDH11X-WT (wild type) groups and compared the mutational events and clinical characters between the two groups. Figure 1C, D shows the summary of the top 10 most commonly mutated genes and clinical information of LUAD patients from Hellmann and TCGA cohorts using oncoPrint plot. Eight genes overlapped between Hellmann and TCGA cohorts, including TP53, TTN, MUC16, KRAS, RYR2, LRP1B, XIRP2, and ZFHX4. The mutation rates of them in the patients with PCDH11X mutation were higher than those in patients with wild-type PCDH11X, suggesting PCDH11X mutation was associated with higher tumor mutation burden (TMB). For clinical characters, we found in Hellmann cohort, ORR and the proportion of patients with progression free in the PCDH11X-MUT group were higher than those in the PCDH11X-WT group, suggesting that PCDH11X mutation may be related to the response to ICIs in patients with LUAD.
Then, we performed the analysis of copy number variation events in TCGA-LUAD dataset. As shown in Fig. 1E, F, the number of significantly amplified and deletion peaks in the PCDH11X-WT group was notably more compared to the PCDH11X-MUT group. In the PCDH11X-MUT group, the top five significant genomic events were composed of four amplified regions and one deleted region. They were located in 1q21.3, 12p12.1, 12p11.21, 14q13.3, and 9p21.3, respectively. In the PCDH11X-WT group, three most significantly amplified events were located in 5p15.33, 8q24.21, and 14q13.3; the two deleted events were located in 9p21.3 and 9p23.
Genomic variation landscapes associated with PCDH11X mutations in the Hellmann and TCGA cohorts. A, B Lollipop plots display the amino acid changes induced by somatic mutation in the longest isoform of PCDH11X in both Hellmann and TCGA cohort. Different color boxes are shown for protein structures, and the lollipops indicate the types and location of mutation. C, D Top 10 genes with the highest number of mutation in the Hellmann (n = 75) and TCGA cohorts (n = 567). The PCDH11X mutational status and clinical features are annotated together. E, F CNV analysis using GISTIC 2.0 for TCGA LUAD dataset grouping into PCDH11X-MUT and PCDH11X-WT. Peaks colored by red and blue represent the amplifications and deletions in the copy number segment, respectively. The top 5 most significantly mutational sites have been labeled. The G-score means the probability of copy number alterations in a chromosomal location
PCDH11X mutation is associated with better clinical outcomes in ICI-treated lung adenocarcinoma
We investigated the association between PCDH11X mutation and ICI treatment benefits in Hellmann cohort. In the cohort, the mutation frequency of PCDH11X was 20% (15/75). Patients in the PCDH11X-MUT group experienced longer PFS than those in the PCDH11X-WT group (median: not reached versus 6.8 months, P = 0.02; Fig. 2A). The ORR of patients with PCDH11X-MUT was also higher than that of patients with PCDH11X-WT, although the P value is slightly higher than 0.05 (Fig. 2E; 53.33% (8/15) versus 26.66% (16/60), odds ratio = 3.089, P = 0.065).
To further evaluate the predictive value of PCDH11X, Miao and Rizvi cohorts were analyzed. In Miao cohort, mutation frequency of PCDH11X was 12.5% (6/48) in LUAD patients. The OS benefit was more prominent in the PCDH11X-MUT group than that in the PCDH11X-WT group (Fig. 2B; median: not reached versus 12.6 months, P = 0.033). The ORR of patients with PCDH11X-MUT was also higher than that of patients with PCDH11X-WT, although the P value is slightly higher than 0.05 (Fig. 2E; 66.67% (4/6) versus 26.19% (11/42), odds ratio = 5.40, P = 0.07). In Rizvi cohort, mutation frequency of PCDH11X was 13.8%. The PFS was longer in the PCDH11X-MUT group than that in the PCDH11X-WT group, although the P value is not significant (Fig. 2C; median: 8.6 months versus 6.3 months, P = 0.27). In terms of ORR, the rate in the PCDH11X-MUT group is also higher than that in the PCDH11X-WT group (Fig. 2E; 50% (2/4) versus 22.9% (8/25), odds ratio = 5.40, P = 0.59).
Association of PCDH11X mutation and ICI treatment response in LUAD. A Kaplan–Meier curves comparing PFS of patients with or without PCDH11X mutation in Hellmann cohort. B Kaplan–Meier curves comparing OS of patients with or without PCDH11X mutation in Miao cohort. C, D Kaplan–Meier curves comparing PFS of patients with or without PCDH11X mutation in Rizvi cohort and combined ICI treated cohort. E ORR were compared between the PCDH11X-MUT and PCDH11X-WT groups in Hellmann, Miao-LUAD, Rizvi, and combined cohorts
In the combined dataset, the PFS of the PCDH11X-MUT group was longer than that of the PCDH11X-WT group (Fig. 2D; 21.71 months versus 7.82 months, P = 0.034). In terms of ORR, the rate of the PCDH11X-MUT group is nearly twice than that of the PCDH11X-WT group (Fig. 2E; 54.55% (12/22) versus 27.10% (29/107), odds ratio = 3.19, P = 0.02).
PCDH11X mutation is an independent predictive biomarker
We used multivariable Cox regression to assess the effect of multiple factors, including PCDH11X mutational status, age, gender, smoking history, TMB, and PD-L1 expression for predicting PFS of ICI therapy in the combined dataset. The results showed that PCDH11X mutational status remained an independent predictive factor for PFS (Table 1).
PCDH11X mutation is a specific biomarker for ICI treatment response in lung adenocarcinoma
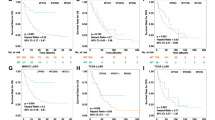
We assessed whether PCDH11X mutation was a specific biomarker for ICI treatment response in LUAD. We first assessed the value of PCDH11X mutation in predicting ICI treatment response in melanoma data combined from five cohorts and in Miao cohort excluding LUAD. The association of PCDH11X mutation and better OS was not observed in melanoma (Fig. 3A). Also, in pan-cancer (Miao) cohort excluding LUAD, no significant difference was found in OS between the PCDH11X-WT and PCDH11X-MUT groups (Fig. 3B). Then, we assessed the potential prognosis value of PCDH11X mutation in TCGA-LUAD cohort, which is a non-ICI-treated cohort. OS between the PCDH11X-WT and PCDH11X-MUT groups was not significantly different in TCGA-LUAD cohort (Fig. 3C). These results suggested that PCDH11X mutation was a specific biomarker for predicting ICI treatment response in LUAD.
Association of PCDH11X mutation and prognosis in ICI-treated non-LUAD patients or non-ICI-treated LUAD patients. A–C Kaplan–Meier curves comparing OS of patients with or without PCDH11X mutation in ICI-treated melanoma cohort, ICI-treated Miao cohort excluding LUAD, and TCGA non-ICI-treated LUAD cohort
PCDH11X mutation is associated with increased PD-L1 expression, TMB, neoantigen load, and DDR mutations in lung adenocarcinoma
PD-L1 expression is currently the most widely validated and accepted biomarker of ICI treatment. To investigate the possible mechanism underlying the predictive role of PCDH11X mutation, we firstly compared the expression of PD-L1 between the PCDH11X-WT and PCDH11X-MUT groups. We found that PCDH11X mutation was associated with higher PD-L1 expression (Fig. 4A). Tumors with higher TMB are thought to produce more neoantigens, be more immunogenic, and consequently have better response to ICIs [33, 34]. Neoantigen load, which represents the number of tumor mutations actually targeted by T cells, has also been reported to be related to the response to ICIs [35]. So, we next compared TMB and neoantigen load between the PCDH11X-WT and PCDH11X-MUT groups. We found that PCDH11X-MUT was associated with both higher TMB (Fig. 4B) and neoantigen load (Fig. 4C).
Association of PCDH11X mutation and established predictors for ICI treatment response. A–D Comparison of PD-L1 expression (A), TMB (B), neoantigen load (C), and mutation rate in the DDR-related pathways (D) between the PCDH11X-MUT and PCDH11X-WT groups in the TCGA cohort. *P < 0.05; ***P < 0.001; ns, no significance
Considering that DDR gene alterations may explain the higher level of TMB and was reported to associated with the response to ICIs in tumors [36], we further examined the association of the PCDH11X mutation status and DDR gene mutations. We observed a trend towards an enrichment of DDR gene alterations in the PCDH11X-MUT group. Seven pathways (i.e., base excision repair (BER), homologous recombination (HR), mismatch repair (MMR), Fanconi anemia (FA), non-homologous end join (NHEJ), DNA repair (DR), and nucleotide excision repair (NER) demonstrated enrichment for mutations in the PCDH11X-MUT group versus the PCDH11X-WT group (Fig. 4D).
PCDH11X mutation is associated with hot tumor microenvironment in TCGA-LUAD cohort
High immunogenicity caused by high TMB and neoantigen load may lead to a hot tumor microenvironment (TME), which is characterized by T cell infiltration, molecular signatures of immune activation and association with better response to ICIs [37, 38]. To investigate whether PCDH11X mutation is associated with hot TME and further investigate the possible mechanism underlying the predictive role of PCDH11X mutation, we first calculated 18 reported immune signatures in TCGA-LUAD cohort and compared the scores between the PCDH11X-MUT and PCDH11X-WT groups. We found that multiple T cell or CD8 T cell–related signatures, including T cell activation, T-effector (Teff), CD8 T cytotoxic, and cytotoxic T lymphocyte (CTL) level, were significantly higher in the PCDH11X-MUT group (Fig. 5A, B). Besides, IFNγ signature, which is a critical driver of programmed death ligand-1 (PD-L1) expression in cancer and host cells, was also higher in the PCDH11X-MUT group (Fig. 5A, B).
PCDH11X mutations are associated with activated antitumor immunity. A Comparison of the 18 immune signatures between the PCDH11X-MUT and PCDH11X-WT groups. B Comparison of the expression levels of genes used for immune signature calculation between the PCDH11X-MUT and PCDH11X-WT groups. C Comparison of the infiltration levels of 22 types of immune cells between the PCDH11X-MUT and PCDH11X-WT groups. The blue color represents the PCDH11X-WT group, and the pink color represents the PCDH11X-MUT group. *P < 0.05; ***P < 0.001; ns, no significance. D Differences in the mRNA expression levels of immune-related genes between the PCDH11X-MUT and PCDH11X-WT groups
Tumor-infiltrating immune cells, especially CD8 T cells, have an important effect on the prognosis of patients receiving ICI treatment. We thus continued to survey the relationships between PCDH11X mutation and immune cell infiltration in TCGA-LUAD cohort. As shown in Fig. 5C, in consistent with the higher CD8 T cell–related signature scores in the PCDH11X-MUT group, infiltration level of CD8 T cell was significantly higher in the PCDH11X-MUT group. Besides, follicular helper T cells were higher in the PCDH11X-MUT group. For CD4 memory cells, activated CD4 memory T cell was higher in the PCDH11X-MUT group, while resting CD4 memory T cell was lower in the PCDH11X-MUT group. M0 and M1 macrophage cells were higher in the PCDH11X-MUT group. Infiltration levels of dendritic cell, mast cells, monocytes, and neutrophils were lower in the PCDH11X-MUT group.
The relative expressions of 78 immune-related genes in the TCGA-LUAD cohort in the PCDH11X-WT and PCDH11X-MUT groups were analyzed (Fig. 5D). The result showed that the expression levels of several ligands that function as stimulatory immune checkpoints, including CXCL10, CXCL9, IFNG, IL1B, TNF, and LI12A, were significantly higher in the PCDH11X-MUT group. In addition, EDNRB, an immune inhibitor highly expressed in immunologically quiet subtype of cancers [31], was down-regulated in the PCDH11X-MUT group.
Overall, these results suggested that PCDH11X mutation was associated with hot TME.
PCDH11X mutation affect the tumor-related biological pathways
The mechanism of how PCDH11X mutations positively affect the immunogenicity and tumor microenvironment is unknown. In order to understand this mechanism, GSVA was carried out in TCGA-LUAD cohort. As shown in Fig. 6A, GSVA for hallmark gene signatures showed that “SPERMATOGENESIS” and cell cycle–related gene sets, including “E2F TARGETS” and “G2M CHECKPOINT,” were significantly upregulated in the PCDH11X-MUT group, while “HEDGEHOG SIGNALING,” which is responsible for tumorigenesis and interplays with autophagy in multiple cancers, was significantly down-regulated in the PCDH11X-MUT group. Besides, “BILE ACID METABOLISM,” which was associated with migration of LUAD [39], was also significantly down-regulated in the PCDH11X-MUT group. Figure 6B shows GSVA results for KEGG pathway. “CELL CYCLE” and DDR-related pathways, including “HOMOLOGOUS RECOMBINATION” and “MISMATCH REPAIR,” were significantly upregulated in the PCDH11X-MUT group. In the other hand, multiple metabolism-related pathways, including “ABC TRANSPOTERS,” “BILE ACID BIOSYNTHESIS,” “FATTY ACID METABOLISM,” and “DRUG METABOLISM CYTOCHROME P450,” were significantly down-regulated in the PCDH11X-MUT group. Besides, PPAR signaling pathway, which was identified as critical controllers for several key enzymes that catalyze the oxidation of fatty acids [40], and GnRH signaling pathway, which plays an important role in the control of tumorigenesis and progression in human cancers [41], were down-regulated in the PCDH11X-MUT group.
The GSVA results. A GSVA for hallmark gene sets. B GSVA for KEGG pathway gene sets. A, B t value < 0 (blue) represents that the scores of corresponding gene sets are higher in the PCDH11X-WT group; t value > 0 (pink) represents that the scores of corresponding gene sets are higher in the PCDH11X-MUT group
Discussion
The discovery of predictive biomarkers is imperatively needed in developing clinical LUAD immunotherapy strategy. In this study, we presented PCDH11X, a member of cadherin superfamily, as a biomarker indicating better clinical outcomes of the ICI treatment in LAUD. Three ICI-treated LAUD cohorts were used to investigate the association between PCDH11X mutation and treatment benefits. Survival analysis revealed that survival time (OS or PFS) was consistently longer in the PCDH11X-MUT group, especially for Hellmann cohort and Miao cohort, which showed significant differences. Then, the three cohorts were combined to one dataset, and the analysis for PFS also confirmed the conclusion. ORR was compared respectively in three cohorts. The trends were consistent that the PCDH11X-MUT group presented higher ORR, and the not significant P values may result from the limited sample size of each cohort. But the combined dataset showed a significantly higher ORR in the PCDH11X-MUT group than that in the PCDH11X-WT group, almost twice as much.
Furthermore, our study also suggested the specificity of PCDH11X mutation for predicting ICI treatment response in LUAD. Firstly, we found that in Miao cohort, the patients with non-LUAD carcinoma did not exhibit different clinical benefits depending on PCDH11X status. Secondly, melanoma is a type of cancer with relatively good response to ICIs, and datasets with mutation data based on whole exome sequencing are available. So, we include melanoma cohort for analysis. We found that in melanoma, the patients also did not exhibit different clinical benefits depending on PCDH11X status. Thirdly, the LAUD patients without ICI treatment from TCGA cohort also did not exhibit different clinical benefits depending on PCDH11X status.
Based on these findings, we further assessed the relevance between PCDH11X mutation and several known predictors for ICI treatment in LUAD. PD-L1 expression, TMB, and neoantigen load are widely reported predictor for ICI treatment benefits [42,43,44]. Therefore, we speculated whether the PCDH11X mutation was associated with PD-L1 expression, TMB, and neoantigen load and tried to find a clue for the prediction mechanism of PCDH11X. The statistical results confirmed our speculation and suggested that the higher expression of PD-L1 in the PCDH11X-MUT group might be responsible for the observed benefits of ICI treatment. Furthermore, the PCDH11X-MUT group was observed with higher mutation levels in DDR pathways, which could explain its high TMB [45]. And it was plausible to infer that the high neoantigen load of the mutant type was a result of genomic instability caused by gene mutations in DDR pathways [46]. These molecular characteristics collectively contribute to the therapeutic benefits of ICI treatment in patients with PCDH11X mutation.
Since it is generally accepted that tumors with hot tumor microenvironment (TME), which is characterized by heightened immune activity, response better to ICI treatment [47], we next investigated whether PCDH11X mutation was associated with hot TME. Most of the immune signatures displayed higher scores in the PCDH11X-MUT group, and five of 18 signatures showed statistical significance. The T cell–related signatures, T cell activation, Teff signature, CD8 T cytotoxic, and CTL levels indicated the immune response status of tumor-infiltrated T cell. Activated T cell immunity facilitates the ICIs working through T cell cytotoxic effects and cytokine secretion. IFNγ, which is a critical driver of PD-L1 expression in cancer and host cells, was also higher in the PCDH11X-MUT group [48, 49]. Higher expression of IFNγ may provide an explanation for the higher expression of PD-L1 in the PCDH11-MUT group. Immune cell infiltration analysis showed that the immune cells directly participating the antitumor immune response, such as CD8 T cell and macrophage (including M0 and M1 macrophage) were increased in mutant type. T cell follicular helper, which provides essential help to B cells for potent antibody responses and associates with better response in ICI treatment [50], was also increased, indicating that the antibody producing was enhanced in the PCDH11X-MUT group. For CD4 memory cells, activated CD4 memory T cell was higher in the PCDH11X-MUT group while resting CD4 memory T cell was lower in the PCDH11X-MUT group. It was reported in gastric cancer that lower levels of resting CD4 + memory T cells and higher levels of activated CD4 + memory T cells were associated with better prognosis [51]. Infiltration of monocytes, which serves as precursors of macrophages, was lower in the PCDH11X-MUT group. In contrast to the increase of immune cells with immune-activating functions, immune cells with immune-suppressing functions, including dendritic cells [52, 53], resting mast cells [54], and neutrophils [55], were decreased in the PCDH11-MUT group. The 78-gene expression detection showed 11 genes significantly higher in the PCDH11X-MUT group, and 7/11 of them are stimulatory immune checkpoint. These genes stimulated immune response contribute to hot immune microenvironment and make the patients sensible to ICI treatment. The increased expression of PD-L1, LAG3, and PD-1 immune checkpoints in the PCDH11X-MUT group indicates an exhausted state of T cells within the pre-treatment tumors. This exhausted T cell phenotype is associated with a better clinical response to ICI treatment [56].
Although researchers have noticed that PCDH11X was frequently mutated in multiple types of cancers including LUAD [57], there have been limited reports on its function. We observed that the expression of PCDH11X was lower in LUAD tumors compared to normal samples, suggesting that PCDH11X may function as a tumor suppressor (Fig. S1). To further understand the function of PCDH11X and how its mutation affects immunogenicity and antitumor immunity, we performed gene set variation analysis (GSVA). We found that PCDH11X mutation was associated with multiple well-known gene sets related to cancer occurrence and progression, such as “E2F TARGETS” and “G2M CHECKPOINT” and “HEDGEHOG SIGNALING,” confirming the role of PCDH11X mutation in cancer occurrence and progression. Interestingly, we also found that multiple metabolism-related gene sets were down-regulated in the PCDH11X-MUT group. It is widely recognized that metabolic changes occur in tumor cells and immune cells during cancer occurrence and development [58]. Recent studies have shown that metabolic changes may influence the response to ICI treatment by altering the tumor microenvironment [58]. We speculate that PCDH11X mutation may lead to metabolic reprogramming, thereby affecting ICI treatment response.
There are several limitations in our study. Firstly, the relatively small sample size of ICI-treated cohorts limits the statistical power of our analysis. Therefore, the predictive value of PCDH11X mutation for ICI treatment needs to be confirmed in larger studies. Secondly, clinical features such as tumor stage may influence tumor microenvironment and play a role in the response of immunotherapy. Due to the low mutation frequency of PCDH11X, the sample size of the PCDH11X-MUT group is much smaller than the WT group. Therefore, we did not perform subgroup analysis to test whether PCDH11X-MUT was a robust predictor across subgroups, although we used multivariate Cox proportional hazards model to adjust potential variables. Thirdly, we used data collected from different studies to perform analysis. The retrospective design and cohort heterogeneity may introduce bias to this study. In addition, while we have obtained some hints by comparing the expression levels of immune-related genes and conducting GSVA between the PCDH11X-MUT and PCDH11X-WT groups, the underlying molecular mechanisms through which PCDH11X mutation improves the therapeutic effect of ICI are unclear and need further research.
Conclusion
Our study identifies PCDH11X mutation as a specific biomarker for predicting the benefit of ICI treatment in LUAD. PCDH11X mutation is associated with longer survival, increased PD-L1 expression, higher TMB, elevated neoantigen load, and enhanced immune activity. Furthermore, we discovered association between PCDH11X mutation and pathways related to tumorigenesis and metabolism. Overall, our study provides a novel potential biomarker for predicting the efficacy of ICI treatment in LUAD. However, further studies are needed to confirm and explore the underlying mechanisms of the predictive effect of PCDH11X.
Data availability
Sources for all data used in this study were described in the “Materials and Methods” section. These datasets were accessible freely and can be found in public databases or supplementary material of published articles.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Schabath MB, Cote ML (2019) Cancer progress and priorities: lung cancer. Cancer Epidemiol Biomark Prev 28:1563–1579. https://doi.org/10.1158/1055-9965.EPI-19-0221
Leiter A, Veluswamy RR, Wisnivesky JP (2023) The global burden of lung cancer: current status and future trends. Nat Rev Clin Oncol 20:624–639. https://doi.org/10.1038/s41571-023-00798-3
Han X, Li F, Fang Z, Gao Y, Li F, Fang R, Yao S, Sun Y, Li L, Zhang W et al (2014) Transdifferentiation of lung adenocarcinoma in mice with Lkb1 deficiency to squamous cell carcinoma. Nat commun. https://doi.org/10.1038/ncomms4261
Quintanal-Villalonga A, Taniguchi H, Zhan YA, Hasan MM, Chavan SS, Meng F, Uddin F, Allaj V, Manoj P, Shah NS et al (2021) Comprehensive molecular characterization of lung tumors implicates AKT and MYC signaling in adenocarcinoma to squamous cell transdifferentiation. J hematol oncol. https://doi.org/10.1186/s13045-021-01186-z
Zhu L, Liu Y, Gao H, Liu J, Zhou Q, Luo F (2022) Case report: partial response following nivolumab plus docetaxel in a patient with EGFR exon 20 deletion/insertion (p.N771delinsGF) mutant lung adenocarcinoma transdifferentiated from squamous cell carcinoma. Front cell dev biol. https://doi.org/10.3389/fcell.2021.755135
Howlader N, Forjaz G, Mooradian MJ, Meza R, Kong CY, Cronin KA, Mariotto AB, Lowy DR, Feuer EJ (2020) The effect of advances in lung-cancer treatment on population mortality. N Engl J Med 383:640–649. https://doi.org/10.1056/NEJMoa1916623
Paz-Ares L, Luft A, Vicente D, Tafreshi A, Gümüş M, Mazières J, Hermes B, Çay Şenler F, Csőszi T, Fülöp A et al (2018) Pembrolizumab plus chemotherapy for squamous non–small-cell lung cancer. N Engl J Med 379:2040–2051. https://doi.org/10.1056/NEJMoa1810865
Lu S, Wang J, Cheng Y, Mok T, Chang J, Zhang L, Feng J, Tu H-Y, Wu L, Zhang Y et al (2021) Nivolumab versus docetaxel in a predominantly Chinese patient population with previously treated advanced non-small cell lung cancer: 2-year follow-up from a randomized, open-label, phase 3 study (CheckMate 078). Lung Cancer 152:7–14. https://doi.org/10.1016/j.lungcan.2020.11.013
Jenkins RW, Barbie DA, Flaherty KT (2018) Mechanisms of resistance to immune checkpoint inhibitors. Br J Cancer 118:9–16. https://doi.org/10.1038/bjc.2017.434
Zeng H, Tong F, Bin Y, Peng L, Gao X, Xia X, Yi X, Dong X (2022) The predictive value of PAK7 mutation for immune checkpoint inhibitors therapy in non-small cell cancer. Front Immunol 13:834142. https://doi.org/10.3389/fimmu.2022.834142
Van Roy FJNRC (2014) Beyond E-cadherin: roles of other cadherin superfamily members in cancer. 14: 121–134
Rouget-Quermalet V, Giustiniani J, Marie-Cardine A, Beaud G, Besnard F, Loyaux D, Ferrara P, Leroy K, Shimizu N, Gaulard P et al (2006) Protocadherin 15 (PCDH15): a new secreted isoform and a potential marker for NK/T cell lymphomas. Oncogene 25:2807–2811. https://doi.org/10.1038/sj.onc.1209301
Vazquez-Cintron EJ, Monu NR, Burns JC, Blum R, Chen G, Lopez P, Ma J, Radoja S, Frey AB (2012) Protocadherin-18 is a novel differentiation marker and an inhibitory signaling receptor for CD8+ effector memory T cells. PLoS ONE 7:e36101. https://doi.org/10.1371/journal.pone.0036101
Frey AB (2017) The inhibitory signaling receptor protocadherin-18 regulates tumor-infiltrating CD8(+) T-cell function. Cancer Immunol Res 5:920–928. https://doi.org/10.1158/2326-6066.CIR-17-0187
Zhu G, Ren D, Lei X, Shi R, Zhu S, Zhou N, Zu L, Mello RA, Chen J, Xu S (2021) Mutations associated with no durable clinical benefit to immune checkpoint blockade in non-S-cell lung cancer. Cancers (Basel). https://doi.org/10.3390/cancers13061397
Feng Z, Yin Y, Liu B, Zheng Y, Shi D, Zhang H, Qin J (2022) Prognostic and immunological role of FAT family genes in non-small cell lung cancer. Cancer Control 29:10732748221076682. https://doi.org/10.1177/10732748221076682
Hellmann MD, Nathanson T, Rizvi H, Creelan BC, Sanchez-Vega F, Ahuja A, Ni A, Novik JB, Mangarin LMB, Abu-Akeel M et al (2018) Genomic features of response to combination immunotherapy in patients with advanced non-small-cell lung cancer. Cancer Cell 33(843–852):e844. https://doi.org/10.1016/j.ccell.2018.03.018
Miao D, Margolis CA, Vokes NI, Liu D, Taylor-Weiner A, Wankowicz SM, Adeegbe D, Keliher D, Schilling B, Tracy A et al (2018) Genomic correlates of response to immune checkpoint blockade in microsatellite-stable solid tumors. Nat Genet 50:1271–1281. https://doi.org/10.1038/s41588-018-0200-2
Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, Lee W, Yuan J, Wong P, Ho TS et al (2015) Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348:124–128. https://doi.org/10.1126/science.aaa1348
Liu D, Schilling B, Liu D, Sucker A, Livingstone E, Jerby-Arnon L, Zimmer L, Gutzmer R, Satzger I, Loquai C et al (2019) Integrative molecular and clinical modeling of clinical outcomes to PD1 blockade in patients with metastatic melanoma. Nat Med 25:1916–1927. https://doi.org/10.1038/s41591-019-0654-5
Hugo W, Zaretsky JM, Sun L, Song C, Moreno BH, Hu-Lieskovan S, Berent-Maoz B, Pang J, Chmielowski B, Cherry G et al (2016) Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165:35–44. https://doi.org/10.1016/j.cell.2016.02.065
Riaz N, Havel JJ, Makarov V, Desrichard A, Urba WJ, Sims JS, Hodi FS, Martin-Algarra S, Mandal R, Sharfman WH et al (2017) Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171(934–949):e916. https://doi.org/10.1016/j.cell.2017.09.028
Van Allen EM, Miao D, Schilling B, Shukla SA, Blank C, Zimmer L, Sucker A, Hillen U, Foppen MHG, Goldinger SM et al (2015) Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350:207–211. https://doi.org/10.1126/science.aad0095
Snyder A, Makarov V, Merghoub T, Yuan J, Zaretsky JM, Desrichard A, Walsh LA, Postow MA, Wong P, Ho TS et al (2014) Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med 371:2189–2199. https://doi.org/10.1056/NEJMoa1406498
Mayakonda A, Lin DC, Assenov Y, Plass C, Koeffler HP (2018) Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res 28:1747–1756. https://doi.org/10.1101/gr.239244.118
Gu Z, Eils R, Schlesner M (2016) Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32:2847–2849. https://doi.org/10.1093/bioinformatics/btw313
Colaprico A, Silva TC, Olsen C, Garofano L, Cava C, Garolini D, Sabedot TS, Malta TM, Pagnotta SM, Castiglioni I et al (2016) TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res 44:e71. https://doi.org/10.1093/nar/gkv1507
Mermel CH, Schumacher SE, Hill B, Meyerson ML, Beroukhim R, Getz G (2011) GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. https://doi.org/10.1186/gb-2011-12-4-r41
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, Porta-Pardo E, Gao GF, Plaisier CL, Eddy JA et al (2018) The immune landscape of cancer. Immunity 48(812–830):e814. https://doi.org/10.1016/j.immuni.2018.03.023
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Yang T-HO, Porta-Pardo E, Gao GF, Plaisier CL, Eddy JA et al (2018) The immune landscape of cancer. Immunity. https://doi.org/10.1016/j.immuni.2018.03.023
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. https://doi.org/10.1093/bioinformatics/btp616
Ravi A, Hellmann MD, Arniella MB, Holton M, Freeman SS, Naranbhai V, Stewart C, Leshchiner I, Kim J, Akiyama Y et al (2023) Genomic and transcriptomic analysis of checkpoint blockade response in advanced non-small cell lung cancer. Nat Genet 55:807–819. https://doi.org/10.1038/s41588-023-01355-5
Choucair K, Morand S, Stanbery L, Edelman G, Dworkin L, Nemunaitis J (2020) TMB: a promising immune-response biomarker, and potential spearhead in advancing targeted therapy trials. Cancer Gene Ther 27:841–853. https://doi.org/10.1038/s41417-020-0174-y
Bai R, Lv Z, Xu D, Cui J (2020) Predictive biomarkers for cancer immunotherapy with immune checkpoint inhibitors. Biomarker research 8:34. https://doi.org/10.1186/s40364-020-00209-0
Jiang M, Jia K, Wang L, Li W, Chen B, Liu Y, Wang H, Zhao S, He Y, Zhou C (2021) Alterations of DNA damage response pathway: biomarker and therapeutic strategy for cancer immunotherapy. Acta pharmaceutica Sinica B 11:2983–2994. https://doi.org/10.1016/j.apsb.2021.01.003
Long J, Wang D, Wang A, Chen P, Lin Y, Bian J, Yang X, Zheng M, Zhang H, Zheng Y et al (2022) A mutation-based gene set predicts survival benefit after immunotherapy across multiple cancers and reveals the immune response landscape. Genome medicine 14:20. https://doi.org/10.1186/s13073-022-01024-y
Duan Q, Zhang H, Zheng J, Zhang L (2020) Turning cold into hot: firing up the tumor microenvironment. Trends in cancer 6:605–618. https://doi.org/10.1016/j.trecan.2020.02.022
Nie M, Yao K, Zhu X, Chen N, Xiao N, Wang Y, Peng B, Yao L, Li P, Zhang P et al (2021) Evolutionary metabolic landscape from preneoplasia to invasive lung adenocarcinoma. Nat Commun 12:6479. https://doi.org/10.1038/s41467-021-26685-y
Zhang X, Young HA (2002) PPAR and immune system–what do we know? Int Immunopharmacol 2:1029–1044. https://doi.org/10.1016/s1567-5769(02)00057-7
Wu Y, Liu Z, Tang D, Liu H, Luo S, Stinchcombe TE, Glass C, Su L, Lin L, Christiani DC et al (2021) Potentially functional variants of HBEGF and ITPR3 in GnRH signaling pathway genes predict survival of non-small cell lung cancer patients. Translational research : the journal of laboratory and clinical medicine 233:92–103. https://doi.org/10.1016/j.trsl.2020.12.009
Cristescu R, Mogg R, Ayers M, Albright A, Murphy E, Yearley J, Sher X, Liu XQ, Lu H, Nebozhyn M et al (2018) Pan-tumor genomic biomarkers for PD-1 checkpoint blockade-based immunotherapy. Science. https://doi.org/10.1126/science.aar3593
Hellmann MD, Ciuleanu TE, Pluzanski A, Lee JS, Otterson GA, Audigier-Valette C, Minenza E, Linardou H, Burgers S, Salman P et al (2018) Nivolumab plus ipilimumab in lung cancer with a high tumor mutational burden. N Engl J Med 378:2093–2104. https://doi.org/10.1056/NEJMoa1801946
Tunger A, Sommer U, Wehner R, Kubasch AS, Grimm MO, Bachmann MP, Platzbecker U, Bornhauser M, Baretton G, Schmitz M (2019) The evolving landscape of biomarkers for anti-PD-1 or anti-PD-L1 therapy. J clin med. https://doi.org/10.3390/jcm8101534
Knijnenburg TA, Wang L, Zimmermann MT, Chambwe N, Gao GF, Cherniack AD, Fan H, Shen H, Way GP, Greene CS et al (2018) Genomic and molecular landscape of DNA damage repair deficiency across The Cancer Genome Atlas. Cell Rep 23(239–254):e236. https://doi.org/10.1016/j.celrep.2018.03.076
Tubbs A, Nussenzweig A (2017) Endogenous DNA damage as a source of genomic instability in cancer. Cell 168:644–656. https://doi.org/10.1016/j.cell.2017.01.002
Khosravi GR, Mostafavi S, Bastan S, Ebrahimi N, Gharibvand RS, Eskandari N (2024) Immunologic tumor microenvironment modulators for turning cold tumors hot. Cancer commun. https://doi.org/10.1002/cac2.12539
Dunn GP, Bruce AT, Sheehan KC, Shankaran V, Uppaluri R, Bui JD, Diamond MS, Koebel CM, Arthur C, White JM et al (2005) A critical function for type I interferons in cancer immunoediting. Nat Immunol 6:722–729. https://doi.org/10.1038/ni1213
Ayers M, Lunceford J, Nebozhyn M, Murphy E, Loboda A, Kaufman DR, Albright A, Cheng JD, Kang SP, Shankaran V et al (2017) IFN-gamma-related mRNA profile predicts clinical response to PD-1 blockade. J Clin Invest 127:2930–2940. https://doi.org/10.1172/JCI91190
Baumjohann D, Brossart P (2021) T follicular helper cells: linking cancer immunotherapy and immune-related adverse events. J immunother cancer. https://doi.org/10.1136/jitc-2021-002588
Sun Y, Liu L, Fu Y, Liu Y, Gao X, Xia X, Zhu D, Wang X, Zhou X (2023) Metabolic reprogramming involves in transition of activated/resting CD4+ memory T cells and prognosis of gastric cancer. Front immunol. https://doi.org/10.3389/fimmu.2023.1275461
Mayoux M, Roller A, Pulko V, Sammicheli S, Chen S, Sum E, Jost C, Fransen MF, Buser RB, Kowanetz M et al (2020) Dendritic cells dictate responses to PD-L1 blockade cancer immunotherapy. Sci Transl Med. https://www.science.org/doi/10.1126/scitranslmed.aav7431
Peng Q, Qiu X, Zhang Z, Zhang S, Zhang Y, Liang Y, Guo J, Peng H, Chen M, Fu Y-X et al (2020) PD-L1 on dendritic cells attenuates T cell activation and regulates response to immune checkpoint blockade. Nat commun. https://doi.org/10.1038/s41467-020-18570-x
Somasundaram R, Connelly T, Choi R, Choi H, Samarkina A, Li L, Gregorio E, Chen Y, Thakur R, Abdel-Mohsen M et al (2021) Tumor-infiltrating mast cells are associated with resistance to anti-PD-1 therapy. Nat commun. https://doi.org/10.1038/s41467-020-20600-7
Wang C, Zheng X, Zhang J, Jiang X, Wang J, Li Y, Li X, Shen G, Peng J, Zheng P et al (2023) CD300ld on neutrophils is required for tumour-driven immune suppression. Nature 621:830–839. https://doi.org/10.1038/s41586-023-06511-9
Wherry EJ, Kurachi M (2015) Molecular and cellular insights into T cell exhaustion. Nat Rev Immunol 15:486–499. https://doi.org/10.1038/nri3862
van Roy F (2014) Beyond E-cadherin: roles of other cadherin superfamily members in cancer. Nat Rev Cancer 14:121–134. https://doi.org/10.1038/nrc3647
Bader JE, Voss K, Rathmell JC (2020) Targeting metabolism to improve the tumor microenvironment for cancer immunotherapy. Mol Cell 78:1019–1033. https://doi.org/10.1016/j.molcel.2020.05.034
Funding
This work was supported by the Collaborative Innovation Major Project of Zhengzhou (Grant No. 20XTZX08017), National Key Research and Development Program of China (Grant No. 2018YFE0102100), the UK-China Collaboration Fund to tackle AMR (Grant No. TS/S00887X/1), and National Science and Technology Major Project (Grant No. 2018ZX10305409-005).
Author information
Authors and Affiliations
Contributions
Hao Guo, Manjiao Liu, and Jie Zhao (from The First Affiliated Hospital of Zhengzhou University) contributed to the study conception and design. Data collection and analysis were performed by Manjiao Liu, Meijia Yang, Bei Zhang, Sijian Xia, Jie Zhao (from Jiangsu Simcere Diagnostics Co., Ltd.), Linlin Yan, and Yong Ren. The manuscript was written by Bei Zhang, Manjiao Liu, and Sijian Xia. The project was supervised by Jie Zhao (from The First Affiliated Hospital of Zhengzhou University) and Hao Guo. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, M., Yang, M., Zhang, B. et al. PCDH11X mutation as a potential biomarker for immune checkpoint therapies in lung adenocarcinoma. J Mol Med 102, 899–912 (2024). https://doi.org/10.1007/s00109-024-02450-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-024-02450-8