Abstract
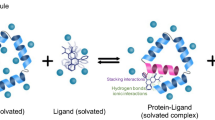
Visualizing post-translational modifications, conformations, and interaction surfaces of protein structures at atomic resolution underpins the development of novel therapeutics to combat disease. As computational resources expand, in silico calculations coupled with experimentally derived structures and functional assays have led to an explosion in structure-based drug design (SBDD) with several compounds in clinical trials. It is increasingly clear that “hidden” transition-state structures along activation trajectories can be harnessed to develop novel classes of allosteric inhibitors. The goal of this mini-review is to empower the clinical researcher with a general knowledge of the strengths and weaknesses of nuclear magnetic resonance (NMR) spectroscopy in molecular medicine. Although NMR can determine protein structures at atomic resolution, its unrivaled strength lies in sensing subtle changes in a nuclei’s chemical environment as a result of intrinsic conformational dynamics, solution conditions, and binding interactions. These can be recorded at atomic resolution, without explicit structure determination, and then incorporated with static structures or molecular dynamics simulations to produce a complete biological picture.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Clinical significance—The awesome power of NMR
Visualizing post-translational modifications, conformations, and interaction surfaces of protein structures at atomic resolution underpins the development of novel therapeutics to combat disease. As computational resources expand, in silico calculations coupled with experimentally derived structures and functional assays have led to an explosion in structure-based drug design (SBDD) with several compounds in clinical trials. It is increasingly clear that “hidden” transition-state structures along activation trajectories can be harnessed to develop novel classes of allosteric inhibitors. The goal of this mini-review is to empower the clinical researcher with a general knowledge of the strengths and weaknesses of nuclear magnetic resonance (NMR) spectroscopy in molecular medicine. Although NMR can determine protein structures at atomic resolution, its unrivaled strength lies in sensing subtle changes in a nuclei’s chemical environment as a result of intrinsic conformational dynamics, solution conditions, and binding interactions. These can be recorded at atomic resolution, without explicit structure determination, and then incorporated with static structures or molecular dynamics simulations to produce a complete biological picture.
The niche of NMR in modern structural biology
Static structural representations subconsciously train scientists to envision a rigid protein architecture. However, macromolecules are exceptionally acrobatic possessing fast, local fluctuations and slow, concerted structural motions (Fig. 1). Therefore, a more accurate depiction is a series of structures where the kinetics, thermodynamics, and function of each conformation varies significantly within a statistical ensemble. Understanding how these dynamic molecules behave in solution with atomic resolution is essential for deciphering their roles in biological processes such as ligand binding and protein-protein interactions. A classic example is how oxygen accesses the cavity of myoglobin—whose crystal structures reveal no channels for the ligand to enter the heme-binding site [1]. NMR spectroscopy is the only technique available to probe these dynamics, at atomic resolution, on timescales ranging over 15 orders of magnitude. As illustrated in Fig. 1, there are NMR experiments to look at backbone vibrations, side-chain motions, secondary structure movements, domain rearrangements, and even folding all under physiological solution-state conditions [2,3,4,5]. This unique perspective from NMR spectroscopy lends major contributions to understanding the biological impact of inherently flexible systems such as intrinsically disordered proteins (IDPs) and integral membrane proteins (IMPs). It should be emphasized that this mini-review is written for the NMR novice. We aim to give an introduction for modern NMR applications to clinically relevant systems such as structure-based drug design, IDPs, and IMPs. Recommendations to in-depth reviews are found throughout the text for the interested researcher.
The business end of NMR: Chemical shifts
Spectroscopy concerns the interaction between light and matter. NMR spectroscopy probes nuclei that possess non-zero quantum spin angular momentum, or simply spin. Biomolecular NMR typically focuses on spin ½ nuclei which produce easily interpretable spectra. The primary atomic constituents of biomolecules carbon, nitrogen, and hydrogen, all possess a stable spin ½ isotope (e.g., −13C, 15N, and 1H). When placed in an external magnetic field, electromagnetically irradiated nuclei resonate at a frequency dictated by their chemical environment—what atoms is it bonded to or spatially nearby. In other words, the resonance frequency of each nuclei is modulated by its primary, secondary, tertiary, and quaternary structure. This unique chemical dependence enables structure determination as well as the ability to monitor changes in motion, and even map atom-specific changes upon ligand binding. For practical purposes, spectroscopists express resonance frequency in terms of chemical shift (units of parts per million; ppm)—which is the resonance frequency scaled by magnetic field strength. Chemical shifts are the most easily accessible NMR parameter that can be measured with high accuracy. As illustrated in Fig. 2a, the unique chemistry of each functional group causes them to occupy a particular region of the spectrum. Subsequently, each amino acid gives rise to distinct chemical shift ranges assuming the residue exists in isolation, or an unstructured conformation, known as a random coil chemical shift (Fig. 2b). These chemical shifts are further altered by the nuclei’s oxidation state, orientation of covalent bonds, isotope of neighboring atoms, hydrogen bonding, and proximity to groups with high magnetic susceptibility such as carbonyl, aromatic, or paramagnetic ions (metals, lanthanides, etc.). In fact, a common technique involves intentional incorporation of paramagnetic ions to modulate the chemical shift for unique structure and dynamic information (as reviewed in [6]).
NMR is highly sensitive to chemical environment based on primary, secondary, tertiary, and quaternary protein structure. a Each atom within a functional group possesses a distinct chemical shift range as illustrated for 1H nuclei. b This translates to unique values for each atoms on a per residue basis as illustrated for 13C groups of select amino acids. A complete list of predicted chemical shift values is available online at the Biological Magnetic Resonance Data Bank [53]. c A series of two-dimensional spectra collected with increasing ligand concentrations (colored from black to magenta); only L83 is specifically located at the binding interface as demonstrated by its large chemical shift perturbations. Inset: the complex affinity can be quantified by fitting changes in chemical shift, and/or peak intensity, as a function of ligand concentration to an appropriate binding equation [28]. d Finally, the residues that experience chemical shift changes can be highlighted on the surface of structure or homology model to map the binding interface
It was not until the early 1990s that the NMR structural database reached a sufficient sample size for “chemical shift homology” searches to predict secondary structures for sequence alignment, fold classification, protein visualization, and homology modeling. Qualitative secondary structure predictions first measured the difference between a residue’s chemical shift from the random coil value, known as the secondary chemical shift, and plotted it as a function of primary sequence. Initial estimates focused on how 13Cα and 13Cβ secondary chemical shifts could discriminate the electronic distribution of beta-strand and alpha-helical regions [7] that formed the chemical shift index (CSI) method [8]. CSI then paved the way for more robust methods that utilize 15N, 1Hα, 13C′, 13Cα, and 13Cβ chemical shifts in numerous web-based or downloadable programs to estimate secondary structure based on chemical shifts, with DANGLE and PsiCSI reporting the highest fidelity [9, 10]. The random coil index (RCI), developed by the Wishart group, goes further by using chemical shifts alone to accurately quantify protein flexibility [11]. Without explicit knowledge of the 3D structure or additional NMR experiments, the chemical shift values of nuclei that sample multiple conformations on a sub-millisecond scale are population averaged. Comparison of these data with molecular dynamics simulations or crystallographic B-factors can quickly identify whether the B-factor is a result of static conformational heterogeneity or true conformational dynamics. Backbone chemical shift “homology searches” are also exploited to predict backbone ϕ and ψ dihedral angles within ≤30° for at least 85% of sites; the most popular programs are PREDITOR, TALOS+, and DANGLE [10, 12, 13]. Stereospecific methyl labeling has led to the development of a simple formula for side-chain rotamers of Val (χ1), Leu (χ2), Ile (χ2) and calculations of further polar and aromatic residues should be possible with larger structural datasets [14, 15].
Current structure determination procedures rely primarily on cataloging short-range (typically ≤5 Å) 1H-1H distance constraints to define the polypeptide architecture. The holy grail of NMR would be tertiary and quaternary structure calculations from chemical shifts alone. Since the year 2000, international structural genomics initiatives rapidly added datasets to the point that high-resolution structures can now be calculated solely from chemical shifts. The dominant methods, CHESSIRE, CS-ROSETTA, and CS23D, rely on backbone chemical shift assignments alone but can faithfully calculate model structures with backbone RMSD <1.5 Å from reference structures [16,17,18]. These model structures are currently limited to ~110 amino acids but inclusion of side-chain chemical shifts, residual dipolar couplings (RDCs), and unassigned nuclear Overhauser effect (NOE) restraints has been successfully applied to proteins up to 200 amino acid proteins [19]. More recent iterations model oligomeric structures from chemical shift perturbations upon complex formation. Programs such as HADDOCK, CamDock, and CS-ROSETTA sequentially, or simultaneously, combine docking and structure determination algorithms to produce tertiary structures from inputs of the free-state structures [20,21,22].
Applications
Solution NMR in drug discovery
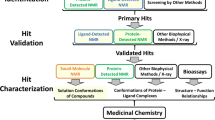
In the last two decades, drug discovery’s focus has shifted to screening fragment molecules (typically containing less than 20 atoms) for development into high-affinity lead compounds. Initial fragments with optimal binding efficiencies (affinity vs. number of atoms) can be further enhanced into lead-like molecules through the addition of chemical functionalities, and/or the linking of ligands that target disparate sites. However, fragments are inherently low-affinity (micromolar affinity) molecules that do not easily crystalize and are difficult to identify in functional assays. In contrast, NMR is not affinity limited and can directly determine binding constants during compound screening to rank them for optimization and lead selection. Binding kinetics directly determine accurate quantification of drug affinity by NMR (for detailed texts on NMR kinetics, see [23,24,25]). The particular exchange regime of a binding reaction can be readily identified by observing each peak throughout a titration; it is important to note that each atom can exhibit its own rate of exchange for a particular binding reaction.
First spearheaded by Fesik and colleagues, structure-activity relationship (SAR) by NMR [26] has matured into a moderate-throughput screening technology. Titrations can be performed in two different formats: (1) ligand-based or (2) target-based. Ligand-based screening relies on excess concentrations of molecular fragments and monitors chemical shift changes through one-dimensional spectra collected in minutes. A positive binding interaction alters the chemical shift and/or the line shape of a given resonance. Details of various ligand-based experiments are reviewed here [27], for example, pulse sequences that monitor changes in the ligands’ diffusion coefficient (diffusion-ordered spectroscopy, DOSY), or only allow visualization of atoms within a particular distance of the target protein (such as saturation transfer difference spectroscopy, STD). The high resolution of small molecules allows batch screening of 5–10 molecules to improve throughput (for details, we recommend this excellent review [27]). Ligand-based screening is further advantageous as the target protein does not need to be isotopically labeled and usually only studied in small concentrations. Conversely, target-based screening follows changes to on the protein side. Once the chemical shift assignments of the target protein are known, isotopically enriched protein can be titrated with a putative ligand or binding partner and the resulting chemical shift changes in monitored by two-dimensional correlation spectroscopy (Fig. 2c) to determine binding constants and stoichiometry [28]. Highlighting perturbed residues on the surface of a known protein structure, called chemical shift mapping, can be used to identify the binding pocket (Fig. 2d). Two-dimensional 1H-15 N heteronuclear single-quantum coherence (HSQC) spectroscopy is the most common method for mapping backbone chemical shifts because every residue type (except proline) contains a highly sensitive probe to changes in local magnetic environment. HMQC [29] and TROSY [30] are alternative pulse schemes that supply equivalent spectral information and provide advantages in certain situations such as large molecular weight proteins. As most protein-ligand interactions are side-chain mediated, two-dimensional 1H-13C HSQCs can provide additional information, but side-chain signals typically suffer from overlap in large target proteins. Thus, methyl-selective 1H, 13C-labeling of Ile, Leu, Ala, and Val residues can significantly improve resolution without compromising sensitivity [31].
Intrinsically disordered proteins
Since the initial discovery of functional proteins without a stable architecture, it is been estimated that >30% of the human proteome constitute partially, or completely, unstructured polypeptides [32]. Intrinsically disordered proteins (IDPs) exist in an ensemble of high-energy states that may kinetically sample globular structures and often only in the presence of specific binding partners. They evolved to maintain a level of solubility required for optimal function and have significant biological roles in signaling and regulation—in part due to their promiscuous binding activity. A large proportion of the peptides and proteins responsible for aberrant misfolding diseases are intrinsically disordered in their free soluble forms—amyloid-β in Alzheimer’s [33], α-synuclein in Parkinson’s [34], and amylin in type II diabetes [35]. Whereas X-ray crystallography may obtain structures when IDPs adopt well-ordered bound states, NMR spectroscopy can report on both the bound-state and the apo, high-energy ensemble. Besides NMR, small-angle X-ray scattering (SAXS) is now increasingly used to characterize IDPs. Their high flexibility results in spectroscopic properties that are characteristic of small molecules, allowing for backbone chemical shift assignment even for high molecular weight proteins. Once assigned, one can monitor changes upon interaction with a particular ligand using chemical shift mapping as described in the SBDD section. Although NMR has monopolized their structural characterization, IDPs do present unique technical challenges that are beginning to be addressed by low-gamma nuclei (13C and 15N) detection experiments. In brief, using low-gamma nuclei instead of traditional 1H-detected experiments potentially results in larger signal dispersion, sensitivity, and resolution. Interested readers are directed towards an excellent review by Takeuchi and colleagues [36].
Integral membrane proteins and nanodisc technology
Proteins embedded into plasma membranes are critical to the molecular mechanisms of life, responding to extracellular stimuli and communicating changes in the external environment to intracellular machinery. Some estimates suggest that integral membrane proteins (IMPs) comprise one third of the genome and nearly half of the current pharmacology targets [37, 38]. Despite being ubiquitous in healthy and pathological systems, structural studies of IMPs have been limited. Until recently, X-ray crystallography was unable to handle the fluid-like environment of lipid membranes. In the past decade, lipidic cubic phase-based studies have revolutionized crystallography of membrane proteins with more than 200 structures, nearly half of which were determined in the last 5 years [39]. In contrast, NMR spectroscopy has steadily contributed to IMP structural information—being limited primarily by isotope labeling, molecular weight, and detergent stability. However, the size limitations have decreased significantly following the invention of TROSY methods and improved isotopic labeling schemes. At this point, a major limitation is the loss of structure and function that accompanies purification in detergent and removal of the native lipid environment. If a suitable detergent environment can be found, an additional challenge arises in characterizing protein-protein interactions which are frequently compromised by detergents. A promising strategy is to incorporate IMPs into discrete phospholipid-bilayer mimetics called nanodiscs.
Nanodiscs are an elegant solution created by Sligar and colleagues [40]. Modeled after high-density lipoprotein (HDL) particles, they comprise of a discoidal lipid bilayer stabilized by a polypeptide belt, termed a membrane scaffold protein (MSP). In 2013, Hagn et al. modified the MSP primary sequence to reduce the disc diameter and subsequently rotational correlation times—critical to making nanodiscs accessible to the NMR community [41]. The nanodiscs range from 4 to 10 nm (approximately equivalent to 52–124 kDa) and dominate the IMP’s relaxation properties assuming there are minimal extramembrane portions. In 2017, an empty nanodisc structure was determined by NMR spectroscopy [42]. As suggested by previous bioinformatics and biochemical experiments, MSP dimerizes and adopts a series of alpha-helical repeats that wrap around the lipids in an anti-parallel alignment. One suggested limitation of nanodisc technology is inherent heterogeneity of the disc diameter and difficulty in controlling the stoichiometry of embedded IMPs. Recently, Wagner and colleagues have modified the MSP construct to be covalently circularized using sortase technology to control 95% of discs to within a 1 nm size distribution [43]. MSP-based nanodiscs rely on detergent molecules to solubilize protein: lipid mixtures prior to disc formation. An alternative styrene maleic acid polymer is capable of directly incorporating IMPs without detergent solubilization and may gain traction in the future once disc diameters can be faithfully formed below 15 nm [44, 45].
Technical considerations in brief
Sensitivity
The magnitude of a NMR signal is dependent on the population difference between high and low-energy states. The population difference is directly proportional to the static magnetic field strength—which is the primary reason for building higher-field spectrometers. For a given field strength, signals scale linearly with sample concentration and signal-to-noise (S/N) ratio scales as a √2. It is important to recognize that signal strength reflects the total mole fraction of NMR-active spins rather than the molar concentration, which means the spectra of a 200 μl sample at 100 μM is effectively equivalent to a 400 μl sample at 50 μM (in the absence aggregation, oligomerization, etc.). Sample volumes are typically around 300–500 μl. However, smaller tubes are available that reduces sample volume to 175–200 μl. Modern spectrometer hardware has reduced the minimum protein concentration to ≤15 μM (in 500 μl) for collecting a two-dimensional spectrum within a couple of hours, although higher concentrations will directly result in shorter experimental times. Samples are tolerant to a wide range of solution conditions and temperatures—limited mainly by high salt concentrations (special tubes should be used at >200 mM salt). Temperatures are only limited by the solvent’s freezing point and the spectrometer’s hardware boundary; modern cold probes operate at temperatures between −40 °C and 80 °C.
Molecular weight
The details of spin physics are beyond the scope of this review; however, it is important for any scientists to have an idea of the limitations of each technique. For an accessible, yet thorough text, we recommend understanding NMR spectroscopy by James Keeler [46]. The maximum theoretical signal is proportional to the number of NMR-active spins within a sample. This signal is then modulated by exponential decay functions that reflect relaxation, or the return of a spin system to thermodynamic equilibrium. The relaxation source divides signal decay into two types: T1 (aka longitudinal or spin-lattice relaxation) and T2 (aka transverse or spin-spin relaxation). T2 relaxation is particularly important in biomolecular NMR spectroscopy—with the decay rate increasing as a function of molecular weight. Many researchers are quick to note that T2 relaxation limits the application of NMR to low molecular weight systems. However, as we have detailed in this review—although de novo structure determination becomes challenging at higher molecular weights—the sensitivity of each nuclei to changes in protein structure and dynamics can provide invaluable information without requiring complete structure determination.
Isotope labeling
The major limitation of NMR spectroscopy is the requirement of NMR-active atoms, specifically spin-1/2 nuclei. Luckily for biological systems, hydrogen’s most common isotope is spin-1/2 (1H; 99.9% natural abundance); however, samples must be enriched with 15N and 13C-atoms as their natural abundance is ≤1%. The details of isotope enrichment depend on the specific recombinant protein expression system. In brief, Escherichia coli cultures can be cheaply grown with simple isotopically labeled carbon and nitrogen sources (typically, glucose and ammonium chloride, respectively) whereas eukaryotic systems require supplementation with costlier labeled amino acids. Depending on yields, isotopic labeling can be prohibitively expensive. We recommend these articles for E. coli [47], yeast [47], insect cell [48], or mammalian expression systems [48]. As molecular weights increase (typically ≥30 kDa), non-exchangable hydrogens must be partially or fully deuterated (2H) to improve T2 relaxation properties (as reviewed in [47]).
Conclusion
NMR has always been chemists’ default technique for robust small-molecule structure determination, but its role in the biomolecular arena has been slightly more contested. X-ray crystallography remains the workhorse of structural biology—providing 90% of the Protein Data Bank’s (PDB) entries. Although crystallization has contributed priceless amounts of architectural information, it inherently stabilizes the lowest-energy conformation at the expense of eliminating protein conformational dynamics. It requires a variety of schemes to trap the reaction substrates or coax all molecules within the crystal to adopt the same substrate conformation. Even then, the crystallization procedure itself has the potential to modify the structure, or introduce artifacts in the form of oligomerization, non-specific protein-ligand interactions, or occlusion of binding interfaces. In contrast, solution-state NMR spectroscopy can be performed at physiological temperatures and buffer conditions. Experiments exist to quantitatively probe motions over all timescales from picoseconds to days, and most of these experiments provide atomic resolution information about conformations without the need to determine the complete structure. One glaring absence in this review is the description of NMR experiments probing conformational motions. Even cursory explanation of NMR-based dynamics requires in-depth discussion of spin physics for numerous experiments (Fig. 1). Interested readers are directed to the texts Spin Dynamics [25], Structural Biology: Practical NMR Applications [49], and Protein NMR Spectroscopy [50]. Protein structure determination methods have improved considerably over the past two decades as a result of improved hardware, software, and isotopic labeling strategies. Hardware advancements such as cryogenic probes increase the sensitivity allowing for low concentration studies. Data processing advancements such as non-uniform sampling (NUS) allow faster data collection or the allocation of a given time to collect more data for higher resolution or signal: noise ratio. Novel pulse sequences, especially transverse relaxation-optimized spectroscopy (TROSY) continue to push the molecular weight limits to nearly 1-megadalton complexes [51, 52]. NMR spectroscopy will continue to be a valuable tool for not only exploring fundamental biological principles, but also understanding the molecular details of a variety of diseases and provide critical aid for designing robust medical therapies.
References
Kendrew JC, Dickerson RE, Strandberg BE, Hart RG, Davies DR, Phillips DC, Shore VC (1960) Structure of myoglobin: a three-dimensional Fourier synthesis at 2 A. resolution. Nature 185:422–427
Igumenova TI, Frederick KK, Wand AJ (2006) Characterization of the fast dynamics of protein amino acid side chains using NMR relaxation in solution. Chem Rev 106:1672–1699
Tolman JR, Ruan K (2006) NMR residual dipolar couplings as probes of biomolecular dynamics. Chem Rev 106:1720–1736
Palmer AG 3rd, Kroenke CD, Loria JP (2001) Nuclear magnetic resonance methods for quantifying microsecond-to-millisecond motions in biological macromolecules. Methods Enzymol 339:204–238
Yuwen T, Kay LE (2017) Longitudinal relaxation optimized amide 1H-CEST experiments for studying slow chemical exchange processes in fully protonated proteins. J Biomol NMR. doi:10.1007/s10858-017-0104-y
Otting G (2010) Protein NMR using paramagnetic ions. Annu Rev Biophys 39:387–405
Spera S, Bax A (1991) Empirical correlation between protein backbone conformation and C.alpha. and C.beta. 13C nuclear magnetic resonance chemical shifts. J Am Chem Soc 113:5490–5492
Wishart DS, Sykes BD, Richards FM (1992) The chemical shift index: a fast and simple method for the assignment of protein secondary structure through NMR spectroscopy. Biochemistry 31:1647–1651
Hung LH, Samudrala R (2003) Accurate and automated classification of protein secondary structure with PsiCSI. Protein Sci 12:288–295
Cheung MS, Maguire ML, Stevens TJ, Broadhurst RW (2010) DANGLE: a Bayesian inferential method for predicting protein backbone dihedral angles and secondary structure. J Magn Reson 202:223–233
Berjanskii MV, Wishart DS (2005) A simple method to predict protein flexibility using secondary chemical shifts. J Am Chem Soc 127:14970–14971
Berjanskii MV, Neal S, Wishart DS (2006) PREDITOR: a web server for predicting protein torsion angle restraints. Nucleic Acids Res 34:W63–W69
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44:213–223
London RE, Wingad BD, Mueller GA (2008) Dependence of amino acid side chain 13C shifts on dihedral angle: application to conformational analysis. J Am Chem Soc 130:11097–11105
Hansen DF, Neudecker P, Kay LE (2010) Determination of isoleucine side-chain conformations in ground and excited states of proteins from chemical shifts. J Am Chem Soc 132:7589–7591
Cavalli A, Salvatella X, Dobson CM, Vendruscolo M (2007) Protein structure determination from NMR chemical shifts. Proc Natl Acad Sci U S A 104:9615–9620
Wishart DS, Arndt D, Berjanskii M, Tang P, Zhou J, Lin G (2008) CS23D: a web server for rapid protein structure generation using NMR chemical shifts and sequence data. Nucleic Acids Res 36:W496–W502
Shen Y, Lange O, Delaglio F, Rossi P, Aramini JM, Liu G, Eletsky A, Wu Y, Singarapu KK, Lemak A et al (2008) Consistent blind protein structure generation from NMR chemical shift data. Proc Natl Acad Sci U S A 105:4685–4690
Raman S, Lange OF, Rossi P, Tyka M, Wang X, Aramini J, Liu G, Ramelot TA, Eletsky A, Szyperski T et al (2010) NMR structure determination for larger proteins using backbone-only data. Science 327:1014–1018
Dominguez C, Boelens R, Bonvin AM (2003) HADDOCK: a protein-protein docking approach based on biochemical or biophysical information. J Am Chem Soc 125:1731–1737
Montalvao RW, Cavalli A, Salvatella X, Blundell TL, Vendruscolo M (2008) Structure determination of protein-protein complexes using NMR chemical shifts: case of an endonuclease colicin-immunity protein complex. J Am Chem Soc 130:15990–15996
Das R, Andre I, Shen Y, Wu Y, Lemak A, Bansal S, Arrowsmith CH, Szyperski T, Baker D (2009) Simultaneous prediction of protein folding and docking at high resolution. Proc Natl Acad Sci U S A 106:18978–18983
Bakhmutov VI (2004) Practical NMR relaxation for chemistsWiley, Chichester, West Sussex, England. Hoboken, NJ
Cavanagh J (2007) Protein NMR spectroscopy : principles and practice 2nd edn. Academic Press, Amsterdam; Boston
Levitt MH (2008) Spin dynamics : basics of nuclear magnetic resonance 2nd edn. John Wiley & Sons, Chichester, England; Hoboken, NJ
Shuker SB, Hajduk PJ, Meadows RP, Fesik SW (1996) Discovering high-affinity ligands for proteins: SAR by NMR. Science 274:1531–1534
Lepre CA, Moore JM, Peng JW (2004) Theory and applications of NMR-based screening in pharmaceutical research. Chem Rev 104:3641–3676
Fielding L (2003) NMR methods for the determination of protein-ligand dissociation constants. Curr Top Med Chem 3:39–53
Mandal PK, Majumdar A (2004) A comprehensive discussion of HSQC and HMQC pulse sequences. Concepts in magnetic resonance part a 20A: 1-23. DOI Doi 10.1002/Cmr.A.10095
Pervushin K, Riek R, Wider G, Wuthrich K (1997) Attenuated T2 relaxation by mutual cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc Natl Acad Sci U S A 94:12366–12371
Meyer B, Peters T (2003) NMR spectroscopy techniques for screening and identifying ligand binding to protein receptors. Angew Chem Int Ed Engl 42:864–890
Ward JJ, Sodhi JS, McGuffin LJ, Buxton BF, Jones DT (2004) Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. J Mol Biol 337:635–645
Shankar GM, Li S, Mehta TH, Garcia-Munoz A, Shepardson NE, Smith I, Brett FM, Farrell MA, Rowan MJ, Lemere CA et al (2008) Amyloid-beta protein dimers isolated directly from Alzheimer’s brains impair synaptic plasticity and memory. Nat Med 14:837–842
Spillantini MG, Schmidt ML, Lee VM, Trojanowski JQ, Jakes R, Goedert M (1997) Alpha-synuclein in Lewy bodies. Nature 388:839–840
Paulsson JF, Westermark GT (2005) Aberrant processing of human proislet amyloid polypeptide results in increased amyloid formation. Diabetes 54:2117–2125
Takeuchi K, Gal M, Shimada I, Wagner* G (2012) Chapter 2 low-[gamma] nuclei detection experiments for biomolecular NMR recent developments in biomolecular NMR. The Royal Society of Chemistry, pp. 25–52.
Wallin E, von Heijne G (1998) Genome-wide analysis of integral membrane proteins from eubacterial, archaean, and eukaryotic organisms. Protein Sci 7:1029–1038
Overington JP, Al-Lazikani B, Hopkins AL (2006) How many drug targets are there? Nat Rev Drug Discov 5:993–996
Caffrey M (2015) A comprehensive review of the lipid cubic phase or in meso method for crystallizing membrane and soluble proteins and complexes. Acta Crystallogr F Struct Biol Commun 71:3–18
Denisov IG, Grinkova YV, Lazarides AA, Sligar SG (2004) Directed self-assembly of monodisperse phospholipid bilayer Nanodiscs with controlled size. J Am Chem Soc 126:3477–3487
Hagn F, Etzkorn M, Raschle T, Wagner G (2013) Optimized phospholipid bilayer nanodiscs facilitate high-resolution structure determination of membrane proteins. J Am Chem Soc 135:1919–1925
Bibow S, Polyhach Y, Eichmann C, Chi CN, Kowal J, Albiez S, McLeod RA, Stahlberg H, Jeschke G, Guntert P et al (2017) Solution structure of discoidal high-density lipoprotein particles with a shortened apolipoprotein A-I. Nat Struct Mol Biol 24:187–193
Nasr ML, Baptista D, Strauss M, Sun ZJ, Grigoriu S, Huser S, Pluckthun A, Hagn F, Walz T, Hogle JM et al (2017) Covalently circularized nanodiscs for studying membrane proteins and viral entry. Nat Methods 14:49–52
Knowles TJ, Finka R, Smith C, Lin YP, Dafforn T, Overduin M (2009) Membrane proteins solubilized intact in lipid containing nanoparticles bounded by styrene maleic acid copolymer. J Am Chem Soc 131:7484–7485
Broecker J, Eger BT, Ernst OP (2017) Crystallogenesis of membrane proteins mediated by polymer-bounded lipid Nanodiscs. Structure 25:384–392
Keeler J (2010) Understanding NMR spectroscopy, 2nd edn. John Wiley and Sons, Chichester, U.K.
Downing AK (2004) Protein NMR techniques, 2nd edn. Humana Press, Totowa, N.J
Kost TA, Condreay JP, Jarvis DL (2005) Baculovirus as versatile vectors for protein expression in insect and mammalian cells. Nat Biotechnol 23:567–575
Teng Q (2005) Structural biology: practical NMR applicationsSpringer. York, New
Cavanagh J (2007) Protein NMR spectroscopy: principles and practice 2nd edn. Academic Press, Amsterdam; Boston
Fiaux J, Bertelsen EB, Horwich AL, Wuthrich K (2002) NMR analysis of a 900K GroEL-GroES complex. Nature 418:207–211
Sprangers R, Kay LE (2007) Quantitative dynamics and binding studies of the 20S proteasome by NMR. Nature 445:618–622
Ulrich EL, Akutsu H, Doreleijers JF, Harano Y, Ioannidis YE, Lin J, Livny M, Mading S, Maziuk D, Miller Z et al (2008) BioMagResBank. Nucleic Acids Res 36:D402–D408
Acknowledgements
We would like to acknowledge funding from National Institutes of Health grants K99GM115814 (JJZ), and P01 GM047467 and R01 AI037581 (GW), and a Gillian Fellowship from Howard Hughes Medical Institute (DB).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ziarek, J.J., Baptista, D. & Wagner, G. Recent developments in solution nuclear magnetic resonance (NMR)-based molecular biology. J Mol Med 96, 1–8 (2018). https://doi.org/10.1007/s00109-017-1560-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-017-1560-2