Abstract
Purpose
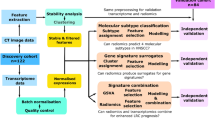
Genetic tumour profiles and radiomic features can be used to complement clinical information in head and neck squamous cell carcinoma (HNSCC) patients. Radiogenomics imply the potential to investigate complementarity or interrelations of radiomic and genomic features, and prognostic factors might be determined. The aim of our study was to explore radiogenomics in HNSCC.
Methods
For 20 HNSCC patients treated with primary radiochemotherapy, next-generation sequencing (NGS) of tumour and corresponding normal tissue was performed. In total, 327 genes were investigated by panel sequencing. Radiomic features were extracted from computed tomography data. A hypothesis-driven approach was used for radiogenomic correlations of selected image-based heterogeneity features and well-known driver gene mutations in HNSCC.
Results
The most frequently mutated driver genes in our cohort were TP53 (involved in cell cycle control), FAT1 (Wnt signalling, cell–cell contacts, migration) and KMT2D (chromatin modification). Radiomic features of heterogeneity did not correlate significantly with somatic mutations in TP53 or KMT2D. However, somatic mutations in FAT1 and smaller primary tumour volumes were associated with reduced radiomic intra-tumour heterogeneity.
Conclusion
The landscape of somatic variants in our cohort is well in line with previous reports. An association of somatic mutations in FAT1 with reduced radiomic tumour heterogeneity could potentially elucidate the previously described favourable outcomes of these patients. Larger studies are needed to validate this exploratory data in the future.
Zusammenfassung
Hintergrund
Genetische Tumorprofile und Radiomics können potenziell als ergänzende Informationen genutzt werden, um die Behandlung von Patienten mit einem Kopf-Hals-Tumor zu personalisieren. Radiogenomics – die Kombination aus genetischen und bildgebenden Informationen – könnten Komplementarität oder Kausalzusammenhänge evaluieren und möglicherweise prognostischen Nutzen haben. Ziel der Studie war es, Radiogenomics bei Patienten mit Kopf-Hals-Tumoren zu untersuchen.
Methoden
Bei 20 Patienten mit Kopf-Hals-Tumoren, die eine primäre Radiochemotherapie erhielten, wurde Tumor- und Normalgewebe sequenziert (Next-Generation Sequencing, NGS). Per Panel wurden hierbei 327 Gene untersucht. Radiomic-basierte Parameter wurden aus Computertomographiedatensätzen extrahiert. Im Sinne eines hypothesengetriebenen Ansatzes wurden selektierte Heterogenitätsparameter mit etablierten Treibermutationen korreliert.
Ergebnisse
Die am häufigsten mutierten Treibergene unserer Kohorte waren TP53 (Zellzyklus), FAT1 (Wnt-Signalweg, Zell-Zell-Kontakte, Migration) und KMT2D (Chromatinmodifikation). Die untersuchten bildgebenden Heterogenitätsparameter korrelierten nicht signifikant mit somatischen Mutationen von TP53 oder KMT2D. Bei FAT1 und kleineren Primärtumorvolumina zeigte sich hingegen eine Assoziation mit einer verminderten bildgebenden Tumorheterogenität.
Schlussfolgerung
Die gefundenen somatischen Tumorvarianten unserer Kohorte stimmen gut mit den bekannten, häufigen Treibermutationen in Kopf-Hals-Tumoren überein. Die Assoziation von somatischen FAT1-Mutationen mit reduzierter bildgebender Heterogenität könnte zur Erklärung der vorbeschriebenen verbesserten Prognose dieser Patientengruppe beitragen. Künftige Studien sind jedoch nötig, um diese Pilotdaten zu validieren.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Locally advanced head and neck squamous cell carcinomas (HNSCCs) are commonly treated with surgery and adjuvant radio(chemo)therapy or with definitive radiotherapy [1]. In definitive radiotherapy, outcome can be enhanced by concomitant chemotherapy [2]. However, overall survival (OS) and loco-regional control (LRC) still need to be improved. In this regard, recent data of the German Cancer Consortium Radiation Oncology Group (DKTK-ROG) report an LRC rate of 62.6% and OS of 59.6% after 2 years of follow-up in HNSCC [3].
To improve outcomes, much effort is made to establish personalised treatment strategies in radiation oncology [4, 5]. Precision medicine implies the potential to individualise therapy by the integration of multimodal data including genomics and radiomics.
A prominent publication investigated 440 computed tomography (CT)-based radiomic features including intensity, shape, texture and multiscale wavelet in lung cancer and HNSCC for prognostic value [6]. After training and validation, the best performing prognostic features of each category were identified. A worsened survival rate was associated with increasing radiomic heterogeneity. In addition, in one lung cancer cohort, gene expression profiles were correlated with radiomic features [6]. An association between upregulated cell cycle pathways and increased intra-tumour heterogeneity features (texture and wavelet feature) has been found.
In addition, with regard to immunotherapy as an upcoming therapeutic option in HNSCC, genetic prognosticators like the tumour mutational burden (TMB) are discussed [7] and some genetic variants might have predictive and prognostic value for therapy response. Tumour genome sequencing facilitates the determination of functional changes and risk groups of HNSCC patients [8]. The Cancer Genome Atlas (TCGA) characterised several frequent somatic variants in HNSCC including TP53 (cell cycle control and survival), KMT2D (chromatin modification) and FAT1 (Wnt/ß-catenin signalling, cell–cell contacts, cell orientation, cell fate) [9, 10], and the main signalling pathways in HNSCC are visualised [9]. As a cross-link to clinical features, variants in TP53 and FAT1 were predominantly found in human papillomavirus (HPV)-negative tumours [9]. In HPV-negative patients, variants in FAT1 were reported to be associated with beneficial outcome in surgically treated HNSCC patients [11]. The authors of this study found mutations in FAT1 as a strong, independent prognostic factor for overall survival in the TCGA cohort and could validate these findings in data of the International Cancer Genome Consortium (ICGC).
The aim of our study was to investigate radiomic tumour heterogeneity according to particular previously reported features and their associations with recurrent somatic driver mutations in HNSCC. With this hypothesis-driven approach of radiogenomic associations, we intended to find correlations that might refer to functional relationships or complemental characteristics of imaging features and genetic aberrations.
Methods
Patients and diagnostics
Twenty patients with locally advanced HNSCC were recruited for this prospective biomarker study. All declared their written informed consent. The study was approved by the local ethics committee (reference number 577/2014BO2) and conducted in accordance with the Helsinki Declaration. All patients were treated with definitive radiochemotherapy up to 70–77 Gy. HPV association was investigated by immunohistochemical staining for p16 or PCR-based assays. Clinical data was extracted from the medical reports.
Radiomics
Due to our limited cohort, we followed a hypothesis-driven approach for finding associations between radiomic heterogeneity and driver gene mutations. Based on the report by Aerts et al. [6], our first tested hypothesis postulated that cell cycle-related somatic mutations (i.e. driver gene mutations in TP53) might correspond with increased radiomic heterogeneity. As a second hypothesis, we investigated if other frequently mutated driver genes correlate with heterogeneity features of the tumour.
Based on our unenhanced planning CT scans (Somatom Sensation Open, Siemens Healthineers, Erlangen, Germany; slice thickness of 3 mm, in-plane pixel size of 1.27 mm, ordered subset expectation maximization [OSEM] 3D [4 iterations, 8 subsets] with a 3D Gaussian filtering for imaging reconstruction), we analysed the two best performing radiomic features for measuring intra-tumour heterogeneity that were described by Aerts et al. [6], namely “Run Length Nonuniformity” (Aerts et al.: Textural Feature 48) and “wavelet Grey Level Nonuniformity HLH” (Aerts et al.: Feature Group 4; decomposition of the image in mid-frequencies). Furthermore, we included “Grey Level Nonuniformity” (Aerts et al.: Textural Feature 47), as a complemental feature, as the authors also reported on this feature in the reference publication. Therefore, in total, three particular heterogeneity features were investigated, and the features were calculated following the previous report of Aerts et al. [6] for confirmability and standardisation.
The gross tumour volumes (GTVs) of the primary tumours were delineated for treatment planning by experienced radiation oncologists. These delineations were subsequently used for radiomic analyses. Due to concerns regarding the influence of dental artefacts [12], we investigated both the data of all 20 patients and, as a subgroup, the patients that had no CT artefacts in the area of interest. Texture features were preprocessed in a 3D fashion regardless of the in-plane, in-slice difference, and we categorised the intensity values in 64 different bins due to the sparse range of intensity values (between −250 to 120 Hounsfield units) across the GTV. For wavelet estimations we used the undecimated wavelet filter. If air or bony structures were included in the GTV, the delineations were adapted and extreme Hounsfield units were excluded for radiomics. Thereby, solely in one patient, the GTV was considerably modified due to massive air and bone involvement (oropharyngeal HNSCC with infiltration of the maxillary sinus).
Genetic analyses
Formalin-fixed paraffin-embedded (FFPE) tumour tissue (obtained at primary diagnosis) was provided by the pathology department and ethylenediaminetetraacetic acid (EDTA) blood samples were collected as normal tissue. We used a particular HNSCC cancer panel containing 327 genes which was originally designed by the DKTK-ROG partner site in Berlin. The library preparation and in-solution capture of the exonic regions were performed using the Agilent HaloplexHS technology (Agilent, Santa Clara, CA, USA). The samples were paired-end sequenced using the HiSeq2500 instrument (Illumina, San Diego, CA, USA). An in-house developed pipeline, called “megSAP”, was used for data analysis (version 0.1-755-g54185f9, https://github.com/imgag/megSAP). In brief, sequencing reads were aligned to the human genome reference sequence (GRCh37) using Burrows–Wheeler Aligner (BWA, version 0.7.17) [13]. Reads aligned to the same chromosomal position and with identical unique molecular identifiers were deduplicated by creating a consensus read constructed per position by choosing the most frequent base (with at least 75% frequency) or by replacing with “N”. Variants were called using Strelka2 (version 2.8.4) [14] and annotated with SnpEff/SnpSift (version 4.37) [15]. All variants were visually validated with the Integrative Genomics Viewer (version 2.3.97) [16], and quality control (QC) parameters were collected during all analysis steps [17]. For further interpretation, we uploaded all somatic variants to the Cancer Genome Interpreter (CGI) [18]. Somatic nucleotide variants were annotated as drivers based on the classification tier 1 and tier 2 (predicted driver mutations) for the single-nucleotide variants (SNVs).
Due to the limited cohort size, only mutations in the most recurrently mutated known driver genes with predicted driver variants according to the CGI database [18], namely TP53, FAT1 and KMT2D, were correlated with radiomic measures of tumour heterogeneity. One patient was excluded from genetic correlations since he showed a hypermutated genotype; thus, the functional impact of single variants remained unclear. For validity and clinical relevance, an allele frequency (AF) of ≥5% was required for reported mutations. However, we recorded driver mutations with lower frequency (<5%) in TP53, FAT1 and KMT2D, as the tumour content of some samples was comparably low. If these variants were annotated by the CGI database, the discrepancy between predicted function and low AF was considered debatable and therefore these variants were marked and excluded from analysis.
Statistics
For the statistical analyses, we used R [19] and SPSS (IBM Corp., Armonk, NY, USA). The Mann–Whitney U test and robust linear regression (M-estimator from MASS R package, log2-transformed values) were used for calculations. Significance estimations of regression coefficients were calculated by the robust F‑test (Wald test, sfsmisc R package). A p-value <0.05 was considered significant.
Results
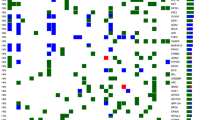
Clinical data of the patients are summarized in Table 1. The most frequently detected driver gene variants and the HPV status are shown in Fig. 1. TP53, FAT1 and KMT2D were the most frequently mutated genes we found in our cohort. Therefore, we correlated the mutation status of these three genes with the image-based heterogeneity features.
Genetic profiles and according human papillomavirus (HPV) infection status of 19 patients are shown in the heatmap. Each column represents a different patient. One patient was excluded from genetic analysis due to a hypermutated genotype. The labelled (asterisk) TP53 and FAT1 mutations were below the 5% cut-off and therefore excluded from statistics. On the right side, the most frequently mutated driver genes are shown. On the y‑axis the frequency of the respective drivers is provided (%). At the top, the overall driver mutations are shown
Regarding the three selected radiomic features indicating tumour heterogeneity, there was no significant correlation found with variants in TP53 or KMT2D. Therefore, the data are not shown. However, a significant association with FAT1 was found, as variants in FAT1 corresponded with reduced radiomic heterogeneity of the primary tumour (Grey Level Nonuniformity: p = 0.019; Fig. 2a; Run Length Nonuniformity: p = 0.046; Fig. 2b and wavelet Grey Level Nonuniformity HLH: p = 0.035; Fig. 2c). This association was found in all three selected heterogeneity features and the observation remained significant in two features when patients who had dental artefacts in the area of the primary tumour were excluded (Fig. 3). Two FAT1 mutations were excluded for reliability due to an AF <5% in one patient (marked with * in Fig. 1) and a hypermutated genotype in another patient (who was therefore excluded from genetic analysis). However, both of the variants were reproducible in the raw data. If these variants were included in the correlations, the association between FAT1 variants and image-based heterogeneity improved for all three features (Grey Level Nonuniformity: p = 0.005; Run Length Nonuniformity: p = 0.024 and wavelet Grey Level Nonuniformity HLH: p = 0.011, data not shown).
Association between FAT1 mutations and radiomic heterogeneity features for patients without dental artefacts. Patients with dental artefacts in the area of interest (gross tumour volume) were excluded. Correlations are shown for a Grey Level Nonuniformity, b Run Length Nonuniformity and c wavelet Grey Level Nonuniformity HLH
The association of somatic mutations in FAT1 and smaller GTVs of the primary tumour was not significant (p = 0.059; Fig. 4a). However, smaller GTVs of the primary tumour corresponded with reduced radiomic heterogeneity (Grey Level Nonuniformity: p < 0.001; Fig. 4b; Run Length Nonuniformity: p < 0.001; Fig. 4c and wavelet Grey Level Nonuniformity HLH: p < 0.001; Fig. 4d).
Gross tumour volume (GTV) and a associated FAT1 mutation status, b Grey Level Nonuniformity, c Run Length Nonuniformity and d wavelet Grey Level Nonuniformity HLH. For correlations between GTVs and radiomic heterogeneity features, a linear regression model was used and data are shown on a log2 scale
Detailed information about the TP53, FAT1 and KMT2D variants in our cohort is visualized in the supplement (Suppl. Fig.).
Discussion
Imaging biomarkers and genetic variants are considered promising features to inform personalised therapeutic decisions. However, reports on correlations of radiomics and genomics remain sparse.
Regarding somatic mutations, our findings are well in line with previously reported mutation profiles in HNSCC [9]. For the investigation of correlations between radiomic data and somatic mutations, we used a hypothesis-driven approach. Our first hypothesis of a correlation between alterations in TP53 as a cell cycle regulator and increased tumour heterogeneity ascertained by radiomic features was not confirmed in our cohort. The previously described gene expression data indicating that increased activity of cell cycle pathways and enhanced proliferation correlate with tumour heterogeneity [6] did not translate into a significant association with somatic mutations of TP53. One could speculate that TP53 has broad effect on tumour development and treatment response, and a unidimensional correlation cannot be found.
For our second hypothesis, we investigated the association of tumour heterogeneity with two other frequently mutated driver genes of our cohort, FAT1 and KMT2D, which had also been identified as recurrently mutated in HNSCC in previous studies [9]. Interestingly, variants in FAT1 were associated with reduced tumour heterogeneity according to all three investigated radiomic heterogeneity features.
FAT1 was found to act as a tumour suppressor by binding ß‑catenin and subsequently decreasing ß‑catenin translocation to the nucleus [20]. Thereby, FAT1 indirectly inhibits cell proliferation and tumour growth. Inactivating mutations of FAT1 are therefore thought to promote the Wnt/ß-catenin signalling pathway [20]. Furthermore, FAT1 is linked to cell–cell contacts and seems to be required for tight cell–cell adhesions [20, 21] and cell polarity [21], as well as for control of cell migration [22] and invasion [22]. In addition, FAT1 contributes to the regulation of epithelial–mesenchymal transition (EMT) [23], which is thought to be associated with tumour aggressiveness. However, the role of FAT1 in tumourigenesis is discussed controversially as both tumour suppressive and oncogenic. These attributes might be dependent on different tumour types. The knockdown of FAT1 was found to reduce cell migration and invasiveness in oral squamous cell carcinoma, glioblastoma and colon cancer [22, 24, 25]. On the contrary, Hu et al. report on accelerated cell migration and EMT after FAT1 knockdown in oesophageal squamous carcinoma [26].
We found an association between FAT1-mutated tumours and reduced heterogeneity of the primary tumours according to radiomic features. Reduced heterogeneity corresponded to smaller primary tumour volumes, and in FAT1-mutated tumours, a trend towards reduced primary tumour volumes was observed, although significance levels were not reached. Postulating FAT1 to be a tumour suppressor in HNSCC [10], inactivating mutations would be expected to cause rather extensive volumes. However, the influence of FAT1 on proliferation in oral squamous cell carcinomas was described to be rather limited [22]. In this way, other oncogenes and tumour suppressors might have a comparably stronger influence on tumour growth and GTV extent.
As discussed above, inactivating/missense variants of FAT1 might result in reduced invasiveness, attenuated migration and looser cell–cell contacts. One could speculate that this translates into less radiomic heterogeneity and smaller tumour volumes as indicated in our cohort. As increased heterogeneity correlated with poor outcome [6] and smaller GTVs [27, 28] are associated with a good prognosis, our findings might support a recent publication reporting favourable outcomes of HPV-negative, surgically treated HNSCC patients, if they presented with mutant FAT1 [11]. Thus, our data suggest a possible interrelation between FAT1 mutations, reduced heterogeneity and smaller GTVs.
Our study revealed interesting preliminary findings in HNSCC patients. The limitation of our study is the small cohort and resulting limited effect size. Therefore, our radiogenomic observations remain solely descriptive and need to be confirmed in larger studies. A further limitation is the location of the biopsy. We sequenced FFPE material that was collected at primary diagnosis. The exact localisation within the primary tumour is therefore unknown. In case of intra-tumour genetic heterogeneity, some variants might be missed and others overestimated. However, we chose only the most frequently mutated driver genes for correlations. As these are well in line with previously reported drivers of HNSCC, these mutations are thought to determine relevant biological and functional variations that might translate into radiomic features.
Conclusion
The collection and integration of omic data for decision-making in precision medicine is essential. Furthermore, tumour characteristics can be investigated and correlated by radiogenomics. Here, we found that reduced radiomic tumour heterogeneity correlated with mutations in FAT1 and with smaller gross tumour volumes, possibly elucidating the cause of the previously described improved overall survival of HPV-negative, FAT1-mutated HNSCC patients.
References
Modesto A, Galissier T, Lusque A, Delord JP, Uro-Coste E, Sarini J, Mouchet F, Lopez R, Laprie A, Graff P, Vergez S, Rives M (2019) Definitive radiochemotherapy or initial surgery for oropharyngeal cancer: to what extent can p16 expression be used in the decision process? Strahlenther Onkol. https://doi.org/10.1007/s00066-019-01451-8
Pignon JP, le Maitre A, Maillard E, Bourhis J, MACH-NC Collaborative Group (2009) Meta-analysis of chemotherapy in head and neck cancer (MACH-NC): an update on 93 randomised trials and 17,346 patients. Radiother Oncol 92(1):4–14. https://doi.org/10.1016/j.radonc.2009.04.014
Linge A, Lohaus F, Lock S, Nowak A, Gudziol V, Valentini C, von Neubeck C, Jutz M, Tinhofer I, Budach V, Sak A, Stuschke M, Balermpas P, Rodel C, Grosu AL, Abdollahi A, Debus J, Ganswindt U, Belka C, Pigorsch S, Combs SE, Monnich D, Zips D, Buchholz F, Aust DE, Baretton GB, Thames HD, Dubrovska A, Alsner J, Overgaard J, Krause M, Baumann M, DKTK ROG (2016) HPV status, cancer stem cell marker expression, hypoxia gene signatures and tumour volume identify good prognosis subgroups in patients with HNSCC after primary radiochemotherapy: a multicentre retrospective study of the German Cancer Consortium Radiation Oncology Group (DKTK-ROG). Radiother Oncol 121(3):364–373. https://doi.org/10.1016/j.radonc.2016.11.008
Baumann M, Krause M, Overgaard J, Debus J, Bentzen SM, Daartz J, Richter C, Zips D, Bortfeld T (2016) Radiation oncology in the era of precision medicine. Nat Rev Cancer 16(4):234–249. https://doi.org/10.1038/nrc.2016.18
Lambin P, van Stiphout RG, Starmans MH, Rios-Velazquez E, Nalbantov G, Aerts HJ, Roelofs E, van Elmpt W, Boutros PC, Granone P, Valentini V, Begg AC, De Ruysscher D, Dekker A (2013) Predicting outcomes in radiation oncology—multifactorial decision support systems. Nat Rev Clin Oncol 10(1):27–40. https://doi.org/10.1038/nrclinonc.2012.196
Aerts HJ, Velazquez ER, Leijenaar RT, Parmar C, Grossmann P, Carvalho S, Bussink J, Monshouwer R, Haibe-Kains B, Rietveld D, Hoebers F, Rietbergen MM, Leemans CR, Dekker A, Quackenbush J, Gillies RJ, Lambin P (2014) Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat Commun 5:4006. https://doi.org/10.1038/ncomms5006
Samstein RM, Lee CH, Shoushtari AN, Hellmann MD, Shen R, Janjigian YY, Barron DA, Zehir A, Jordan EJ, Omuro A, Kaley TJ, Kendall SM, Motzer RJ, Hakimi AA, Voss MH, Russo P, Rosenberg J, Iyer G, Bochner BH, Bajorin DF, Al-Ahmadie HA, Chaft JE, Rudin CM, Riely GJ, Baxi S, Ho AL, Wong RJ, Pfister DG, Wolchok JD, Barker CA, Gutin PH, Brennan CW, Tabar V, Mellinghoff IK, DeAngelis LM, Ariyan CE, Lee N, Tap WD, Gounder MM, D’Angelo SP, Saltz L, Stadler ZK, Scher HI, Baselga J, Razavi P, Klebanoff CA, Yaeger R, Segal NH, Ku GY, DeMatteo RP, Ladanyi M, Rizvi NA, Berger MF, Riaz N, Solit DB, Chan TA, Morris LGT (2019) Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet 51(2):202–206. https://doi.org/10.1038/s41588-018-0312-8
Tinhofer I, Stenzinger A, Eder T, Konschak R, Niehr F, Endris V, Distel L, Hautmann MG, Mandic R, Stromberger C, Weichert W, Budach V (2016) Targeted next-generation sequencing identifies molecular subgroups in squamous cell carcinoma of the head and neck with distinct outcome after concurrent chemoradiation. Ann Oncol 27(12):2262–2268. https://doi.org/10.1093/annonc/mdw426
Cancer Genome Atlas Network (2015) Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 517(7536):576–582. https://doi.org/10.1038/nature14129
Leemans CR, Snijders PJF, Brakenhoff RH (2018) The molecular landscape of head and neck cancer. Nat Rev Cancer 18(5):269–282. https://doi.org/10.1038/nrc.2018.11
Kim KT, Kim BS, Kim JH (2016) Association between FAT1 mutation and overall survival in patients with human papillomavirus-negative head and neck squamous cell carcinoma. Head Neck 38(Suppl 1):E2021–2029. https://doi.org/10.1002/hed.24372
Leijenaar RT, Carvalho S, Hoebers FJ, Aerts HJ, van Elmpt WJ, Huang SH, Chan B, Waldron JN, O’Sullivan B, Lambin P (2015) External validation of a prognostic CT-based radiomic signature in oropharyngeal squamous cell carcinoma. Acta Oncol 54(9):1423–1429. https://doi.org/10.3109/0284186X.2015.1061214
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14):1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Kim S, Scheffler K, Halpern AL, Bekritsky MA, Noh E, Kallberg M, Chen X, Kim Y, Beyter D, Krusche P, Saunders CT (2018) Strelka2: fast and accurate calling of germline and somatic variants. Nat Methods 15(8):591–594. https://doi.org/10.1038/s41592-018-0051-x
Cingolani P, Platts A, Wang LL, Coon M, Nguyen T, Wang L, Land SJ, Lu XY, Ruden DM (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w(1118); iso-2; iso-3. Fly (Austin) 6(2):80–92. https://doi.org/10.4161/fly.19695
Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP (2011) Integrative genomics viewer. Nat Biotechnol 29(1):24–26. https://doi.org/10.1038/nbt.1754
Schroeder CM, Hilke FJ, Loffler MW, Bitzer M, Lenz F, Sturm M (2017) A comprehensive quality control workflow for paired tumor-normal NGS experiments. Bioinformatics 33(11):1721–1722. https://doi.org/10.1093/bioinformatics/btx032
Tamborero D, Rubio-Perez C, Deu-Pons J, Schroeder MP, Vivancos A, Rovira A, Tusquets I, Albanell J, Rodon J, Tabernero J, de Torres C, Dienstmann R, Gonzalez-Perez A, Lopez-Bigas N (2018) Cancer Genome Interpreter annotates the biological and clinical relevance of tumor alterations. Genome Med 10(1):25. https://doi.org/10.1186/s13073-018-0531-8
R Core Team (2018). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Morris LG, Kaufman AM, Gong Y, Ramaswami D, Walsh LA, Turcan S, Eng S, Kannan K, Zou Y, Peng L, Banuchi VE, Paty P, Zeng Z, Vakiani E, Solit D, Singh B, Ganly I, Liau L, Cloughesy TC, Mischel PS, Mellinghoff IK, Chan TA (2013) Recurrent somatic mutation of FAT1 in multiple human cancers leads to aberrant Wnt activation. Nat Genet 45(3):253–261. https://doi.org/10.1038/ng.2538
Tanoue T, Takeichi M (2004) Mammalian Fat1 cadherin regulates actin dynamics and cell-cell contact. J Cell Biol 165(4):517–528. https://doi.org/10.1083/jcb.200403006
Nishikawa Y, Miyazaki T, Nakashiro K, Yamagata H, Isokane M, Goda H, Tanaka H, Oka R, Hamakawa H (2011) Human FAT1 cadherin controls cell migration and invasion of oral squamous cell carcinoma through the localization of beta-catenin. Oncol Rep 26(3):587–592. https://doi.org/10.3892/or.2011.1324
Srivastava C, Irshad K, Dikshit B, Chattopadhyay P, Sarkar C, Gupta DK, Sinha S, Chosdol K (2018) FAT1 modulates EMT and stemness genes expression in hypoxic glioblastoma. Int J Cancer 142(4):805–812. https://doi.org/10.1002/ijc.31092
Dikshit B, Irshad K, Madan E, Aggarwal N, Sarkar C, Chandra PS, Gupta DK, Chattopadhyay P, Sinha S, Chosdol K (2013) FAT1 acts as an upstream regulator of oncogenic and inflammatory pathways, via PDCD4, in glioma cells. Oncogene 32(33):3798–3808. https://doi.org/10.1038/onc.2012.393
Pileri P, Campagnoli S, Grandi A, Parri M, De Camilli E, Song C, Ganfini L, Lacombe A, Naldi I, Sarmientos P, Cinti C, Jin B, Grandi G, Viale G, Terracciano L, Grifantini R (2016) FAT1: a potential target for monoclonal antibody therapy in colon cancer. Br J Cancer 115(1):40–51. https://doi.org/10.1038/bjc.2016.145
Hu X, Zhai Y, Kong P, Cui H, Yan T, Yang J, Qian Y, Ma Y, Wang F, Li H, Cheng C, Zhang L, Jia Z, Li Y, Yang B, Xu E, Wang J, Yang J, Bi Y, Chang L, Wang Y, Zhang Y, Song B, Li G, Shi R, Liu J, Zhang M, Cheng X, Cui Y (2017) FAT1 prevents epithelial mesenchymal transition (EMT) via MAPK/ERK signaling pathway in esophageal squamous cell cancer. Cancer Lett 397:83–93. https://doi.org/10.1016/j.canlet.2017.03.033
Knegjens JL, Hauptmann M, Pameijer FA, Balm AJ, Hoebers FJ, de Bois JA, Kaanders JH, van Herpen CM, Verhoef CG, Wijers OB, Wiggenraad RG, Buter J, Rasch CR (2011) Tumor volume as prognostic factor in chemoradiation for advanced head and neck cancer. Head Neck 33(3):375–382. https://doi.org/10.1002/hed.21459
Carpen T, Saarilahti K, Haglund C, Markkola A, Tarkkanen J, Hagstrom J, Mattila P, Makitie A (2018) Tumor volume as a prognostic marker in p16-positive and p16-negative oropharyngeal cancer patients treated with definitive intensity-modulated radiotherapy. Strahlenther Onkol 194(8):759–770. https://doi.org/10.1007/s00066-018-1309-z
Acknowledgements
We would like to gratefully acknowledge Prof. B. Sipos, Institute of Pathology and Neuropathology; Department of General and Molecular Pathology and Pathological Anatomy, Tübingen, Germany, and Prof. I. Tinhofer, Department of Radiation Oncology and Radiotherapy, Charité Berlin, Germany, for supporting this project. Furthermore, we would like to thank C. Goltermann for the language editing.
Funding
This work was supported by the Stiftung Tumorforschung Kopf-Hals in Wiesbaden, Germany. Besides, K. Zwirner is supported by the intramural Fortüne/PATE Program of the Medical Faculty, Eberhard Karls University of Tübingen (funding number: 2447-0-0) and C. Gani is supported by the Clinician Scientist Program of the Medical Faculty, Eberhard Karls University of Tübingen (funding number: 363-0-0). O. Riess receives funding from the German Research Foundation (DFG) as an NGS Competence Center (INST 37/1049-1). None of our funding sources were involved in study design, data analysis or data interpretation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
D. Thorwarth and D. Zips have research collaborations with Elekta, Philips and Siemens. K. Zwirner, F.J. Hilke, G. Demidov, J. Socarras Fernandez, S. Ossowski, C. Gani, O. Riess, C. Schroeder and S. Welz declare that they have no competing interests.
Ethical standards
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional research committee (Ethics Committee Tübingen; reference number 577/2014BO2) and with the 1964 Helsinki declaration and its later amendments. Informed consent was obtained from all individual participants included in the study.
Caption Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Zwirner, K., Hilke, F.J., Demidov, G. et al. Radiogenomics in head and neck cancer: correlation of radiomic heterogeneity and somatic mutations in TP53, FAT1 and KMT2D. Strahlenther Onkol 195, 771–779 (2019). https://doi.org/10.1007/s00066-019-01478-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00066-019-01478-x