Abstract
A library of N-benzylbenzamide, N-phenethylbenzamide and N-benzyloxybenzamide derivatives were designed, synthesized and evaluated as amyloid beta (Aβ42) aggregation inhibitors. These compounds were designed by replacing the α,β-unsaturated linker region of chalcone with an amide bioisostere. The Aβ42 aggregation inhibition properties of these 27 benzamide derivatives were evaluated by the thioflavin T (ThT)-based fluorescence aggregation kinetics assay, transmission electron microscopy (TEM) studies, Aβ42-induced cytotoxicity assay in mouse hippocampal neuronal HT22 cell lines, fluorescence live cell imaging, and computational modelling studies using a pentamer model of Aβ42. These studies led to the identification of N-benzylbenzamides 3a and 3f, N-phenethylbenzamide 5a and N-benzyloxybenzamide 7a as promising compounds that were able to exhibit anti-aggregation properties in the ThT-based fluorescence experiments, TEM studies and more significantly were able to rescue the hippocampal neuronal HT22 cells from Aβ42-induced cytotoxicity (91–96% cell viability at 25 µM). These results demonstrate the usefulness of these benzamide-based templates in the design and development of novel small molecules as chemical tools and therapeutics to study and treat Alzheimer’s disease.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alzheimer’s disease (AD) is the most common type of dementia. With an increase in the aging population worldwide, the incidence and prevalence of AD are predicted to rise in the coming years. AD is a neurodegenerative disorder that significantly hampers the function of the central nervous system, especially the brain regions of learning and memory [1,2,3,4]. Unfortunately, the diverse, complex, yet unified neurodegenerative pathophysiological pathways, have stalled the discovery of novel drug candidates to treat and cure AD. One of the known factors of AD is the aggregation of amyloid-beta (Aβ) peptides into neurotoxic forms [4, 5]. Recent evidence have shown that Aβ aggregation can trigger other AD pathological pathways either directly or indirectly to promote neurodegeneration and cognitive decline [6,7,8,9]. The Aβ peptide formed after the proteolytic degradation of the amyloid precursor protein, undergoes rapid self-assembly and aggregation to form the senile plaques found in AD patient brain [4, 5, 10]. Based on these findings, the inhibition of Aβ aggregation would provide therapeutic benefits to AD patients. Recent years saw an uptick in the discovery and development of novel anti-Aβ therapies as a novel class of disease-modifying agents to treat AD [11, 12]. In this regard, two monoclonal antibodies: aducanumab (Aduhelm®) and lecanemab (Leqembi®) received approval from the US Food and Drug Administration with the later receiving full approval recently. Another monoclonal antibody, donanemab, is poised to receive FDA approval soon [13,14,15,16]. These developments in the AD drug discovery field demonstrate that targeting the amyloid cascade holds promise as a strategy to treat AD.
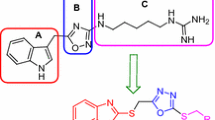
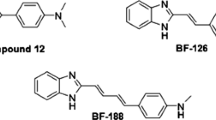
The field is now ripe to discover and develop novel anti-Aβ therapies based on small molecules since they are less expensive to manufacture, exhibit better stability on storage and can be administered orally which is beneficial to AD patients [17]. In this regard, we are working on designing novel small molecules that can bind to Aβ and prevent its aggregation into toxic forms [18,19,20]. We focused our attention on the small molecule chalcone (1) considered as privileged structure (Fig. 1) and is known to demonstrate anti-aggregation properties toward Aβ [21,22,23,24,25,26]. However, chalcone has been identified as pan-assay interfering compound (PAINS) [25,26,27,28,29,30]. They interact with their molecular targets and biological thiols with no specificity, thus providing false positives and interfere with either UV- or fluorescence-based screening assays. The presence of the α,β-unsaturated system in their chemical structure makes them function as Michael acceptors, and the chalcone template can undergo covalent bonding with biological thiols [25, 26, 30]. Therefore, small molecules with these templates are flagged as PAINS in drug development. However, these still represent privileged structures in medicinal chemistry that can be optimized to design and develop novel agents. In this regard, previous studies have demonstrated that molecules bearing flat and planar C6-Cn-C6 carbon chain template are favoured to bind and prevent Aβ self-assembly (Fig. 1), where C6 refers to the presence of two aromatic rings and Cn refers to the type of linkage present between two aromatic ring systems [31].
Based on these findings, we adopted the C6-Cn-C6 template and replaced the Michael acceptor α,β-unsaturated system with an amide bioisostere to obtain the benzamide template (Fig. 1). This template has the following advantages: i) amide bond is not considered as PAINS [30] and does not undergo Michael addition with biological molecules; ii) the resonance delocalization in amide bond provides double bond characteristic and can act as a bioisostere of the α,β-unsaturated system present in chalcone. Thus, the flexibility of the designed ligands would be comparable to chalcone which fulfils the requirements to bind and prevent Aβ aggregation; iii) furthermore, from a synthetic chemistry point of view, amide bond eliminates the need to evaluate cis/trans or (Z)-/(E) isomers. Based on these hypotheses, we synthesized a library of 27 N-benzyl, N-phenethyl and N-benzyloxybenzamide derivatives, with various aromatic substituents and linkers containing either carbon spacer (3a–i and 5a–i), or oxygen spacer (7a–i). Their bioactivity toward Aβ42 aggregation was investigated by carrying out thioflavin T (ThT) fluorescence-based aggregation kinetic study, transmission electron microscopy (TEM), computational modelling and cell viability studies in mouse hippocampal HT22 neuronal cell lines against Aβ42 induced cytotoxicity. These studies demonstrate that the benzamide derivatives exhibit anti-Aβ42 activity in the thioflavin T (ThT) based fluorescent spectroscopy and electron microscopy experiments. Strikingly, these compounds demonstrate excellent neuroprotection against Aβ42-induced neurotoxicity in HT22 mouse hippocampal neuronal cells.
Results and discussion
Chemistry
The synthesis of target benzamide derivatives (27 compounds), was accomplished through direct coupling of primary amines (phenylmethanamine 2, or 2-phenylethan-1-amine 4, or O-benzylhydroxylamine 6) with the corresponding acid chlorides (RCOCl) using triethylamine (TEA) either at room temperature or under reflux using known chemistry (Scheme 1). Alternatively, the benzamide derivatives were synthesized by coupling the corresponding acids (RCOOH) with primary amines using standard amide coupling agent EDC•HCl and HOBt in the presence of TEA (Scheme 1) [32, 33]. The target compounds 3a–i, 5a–i and 7a–i were obtained in good-to-excellent yields ranging from 72–93%. Coupling the acids with amines generally provided greater yields. Analytical data of final compounds 3a–i, 5a–i and 7a–i is provided in the Supplementary Material file.
Biological activity studies
Evaluation of the anti-Aβ42 aggregation activity of 3a–i, 5a–i and 7a–i in the thioflavin T (ThT) based fluorescence spectroscopy
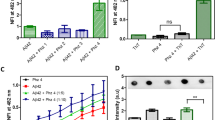
The structure-activity relationship (SAR) of benzamide derivatives to inhibit Aβ42 aggregation was evaluated by ThT-based fluorescence kinetic assay [19]. The benzamide derivatives 3a–i, 5a–i and 7a–i (25 µM) were incubated with Aβ42 (10 μM) at 37 °C, pH 7.4 and the ThT fluorescence intensity was measured after 24 h incubation period to determine the anti-aggregation activity. The results are summarized in Fig. 2. In the N-benzylbenzamide series (3a–i), the presence of an unsubstituted benzene ring in compound 3a (R=benzene) provided 27.1 ± 2.3% inhibition of Aβ42 aggregation (Fig. 2). Replacing the benzene ring with the corresponding pyridine bioisostere in compounds 3b and 3c, led to a significant reduction in their anti-aggregation activity (~ 19% and 13% inhibition, Fig. 2). In contrast, incorporating five-member heterocycles (eg: pyrrole, furan and thiophene) restored the anti-Aβ42 activity with compounds 3d, 3e and 3 f exhibiting 23.8–27.8% inhibition range (Fig. 2). The SAR was further investigated by incorporating planar, bicyclic aromatic rings (eg: naphthyl, quinoline and indole). These changes led to mixed results with compound 3 h (R=quinoline), demonstrating comparable inhibition (~ 27%, Fig. 2), similar to compound 3a (R=benzene, 27% inhibition, Fig. 2). Incorporating the planar, bicyclic indole ring led to a significant loss of anti-aggregation activity in compound 3i (~7% inhibition of Aβ42 aggregation, Fig. 2), whereas the presence of a naphthyl ring in compound 3 g, provided moderate inhibition (~ 16%, Fig. 2). In the N-phenethylbenzamide series (5a–i), the addition of an extra carbon spacer did not change their ability to prevent Aβ42 aggregation (Fig. 2) and similar pattern was seen. In this series, compound 5a (R = benzene, ~ 28% inhibition, Fig. 2), was identified as the best with comparable activity as 3a (~27% inhibition). The next best compound was 5 f possessing a five-member thiophene ring (~ 27% inhibition, Fig. 2). Similar to the N-benzylbenzamide series, the addition of a planar, bicyclic indole ring led to a significant loss in the Aβ42 inhibition activity (5i, ~ 7% inhibition, Fig. 2). In the N-benzyloxybenzamide series (7a–i), the SAR was investigated by adding an oxygen linker next to the amide nitrogen. In this series, compound 7a (R=benzene, 26% inhibition) was identified as the best compound, followed by compounds 7i, 7f, 7d and 7 g with activity ranging from 23–25% (Fig. 2). These studies demonstrate that the N-benzyl, N-phenethyl and N-benzyloxybenzamide derivatives have the potential to be developed as Aβ42 aggregation inhibitors.
Inhibition activity of benzamide derivatives (3a–i, 5a–i and 7a–i, 25 μM each) against Aβ42 aggregation at 24 h time period. The compounds (25 µM) were incubated with Aβ42 (10 μM) for 24 h at 37 °C, pH 7.4 and the ThT fluorescence was monitored (excitation=440 nm, emission=490 nm). The results shown are average±standard deviation of triplicate readings based on three independent experiments
Transmission electron microscopy studies of 3a, 3f, 3h, 5a, 5f and 7a in the presence of Aβ42
Next, we investigated the Aβ42 aggregation inhibition properties of representative benzamide derivatives (3a, 3f, 3h, 5a, 5f and 7a, Fig. 3) by carrying out transmission electron microscopy (TEM) experiments [19]. The morphology of Aβ42 (10 µM) shows a combination of long, twisted, unbranched mature fibrils. In the presence of benzamide derivatives, there was a significant reduction in the formation of Aβ42 fibrils which was characterized by the presence of sparse, thinner, and lower-ordered fibrils (Fig. 3). TEM studies further demonstrate the Aβ42 aggregation inhibition potential of these benzamides.
Effect of 3a, 3f, 3h, 5a, 5f, and 7a on Aβ42-induced cytotoxicity in mouse hippocampal HT22 neuronal cells
In order to determine if the benzamide derivatives were capable of modifying the Aβ42 aggregation pathway in cellular environment, we investigated the ability of representative benzamide derivatives to prevent Aβ42-mediated cytotoxicity in mouse hippocampal HT22 neuronal cell lines. The cytotoxicity of benzamide derivatives (3a, 3f, 3h, 5a, 5f, and 7a) alone, was evaluated in HT22 cells using the cell counting kit 8 (CCK-8)-based colorimetric assay [34]. These studies show that at 25 µM, these benzamide derivatives were not toxic to cells (83–110% viability) compared to untreated cells (UC) as shown in Fig. 4A. Then, HT22 cells were incubated with Aβ42 (5 μM) in the presence and absence of benzamide derivatives (25 μM) for 48 h. The cell viability reduced dramatically in the presence of Aβ42 (~ 39%, Fig. 4B). In contrast, benzamides 3a, 3f, 3h, 5a, 5f, and 7a demonstrated significant neuroprotection to HT22 cells against Aβ42-induced neurotoxicity, and demonstrated cell viability ranging from 81–97% (Fig. 4B). For example, benzamides 3a (R=benzene), 3f (R=thiophene), 5a (R=benzene), and 7a (R=benzene) exhibited excellent neuroprotection against Aβ42-induced neurotoxicity (94.9 ± 5.9%, 95.9 ± 13.7%, 96.7 ± 14.9%, and 91.5 ± 8.3% cell viability respectively). These striking neuroprotection observed suggests that benzamide derivatives have the potential to convert toxic Aβ assemblies into nontoxic forms thereby reducing the cytotoxic effects of Aβ42 assemblies [34]. This provides further evidence that N-benzyl, N-phenethyl and N-benzyloxybenzamides are novel templates that can be employed by researchers to design and discover anti-AD agents.
Neuroprotective effect of benzamides 3a, 3d, 3f, 5a, 5 f, 7a on HT22 hippocampal cells. Panel A (cell viability on y-axis) shows the effect of compounds alone (25 μM each) on HT22 hippocampal neuronal cells after 48 h incubation and Panel B shows the effect of compounds (25 μM each) on HT22 cells treated with Aβ42 (5 μM) after 48 h incubation. Cell viability was determined by CCK-8 based colorimetric assay. The cell viability data in Panel B (left y-axis) was correlated with the Aβ42 aggregation percent inhibition of compounds obtained from the ThT assay (right y-axis). The results shown are average of quadruplicate readings based on three independent experiments. *p < 0.01 compared to Aβ42-treated group (One-way ANOVA followed by Bonferroni post hoc test)
ProteoStat® based fluorescence imaging studies in mouse hippocampal (HT22) neuronal cells
To further validate the anti-aggregation activities of compounds 3a, 5a, and 7a in the cellular environment, live cell fluorescence imaging studies were carried out. The dye ProteoStat® which selectively stains amyloidogenic protein aggregates [35] was used to quantitatively measure and visualize the formation of Aβ42 fibrils in the mouse hippocampal HT22 cell lines in the presence and absence of compounds 3a, 5a, and 7a (25 µM each). When the cells were exposed to Aβ42 alone or the presence of test compounds, the presence of mature β sheet-rich structures was marked by the red ProteoStat® stain (Fig. 5A). The stain density was quantified by summing up the pixel values as the integrated gray value (IGV) using the software ImageJ. These studies show that there was a clear decrease in the amount of Aβ42 fibrils formed in the presence of derivatives 3a, 5a, and 7a (Fig. 5B). The highest IGV was observed for the Aβ42 control group, with a value of 3.5 × 107 (Fig. 5B). When the cells were co-incubated with aggregation inhibitors 3a, 5a, and 7a, there was a reduction in the IGV which demonstrates the ability of these compounds to reduce the formation of Aβ42 fibrils. For compound 3a, around 24% decrease in IGV (2.67 × 107) was observed whereas for compounds 5a and 7a, around 28% and 29% decreases in Aβ42 fibril density was observed which further provides evidence on their ability to inhibit the aggregation of Aβ42 in the cellular environment.
Fluorescence live cell imaging with ProteoStat®. A Images of HT22 cells treated with Aβ42 (10 µM) in the presence and absence of compounds 3a, 5a, and 7a at 25 µM. The images were visualized with the RFP light filter and the binary masks were created to measure the integrated gray value (IGV) of the original fluorescence images. B Quantitative analysis of ProteoStat® measured by IGV in the presence and absence of compounds 3a, 5a, and 7a. The results shown are averages of three independent experiments from three randomized views. *p < 0.01 compared to the Aβ42 control group (One-way ANOVA followed by Bonferroni post hoc test)
LDH release cytotoxicity assay in mouse hippocampal HT22 neuronal cells
One of the mechanisms of Aβ42 cytotoxicity is related to its ability to interact with the lipid membrane which leads to the formation of pores and eventual cell lysis [36]. In this regard, the ability of compounds 3a, 5a, and 7a to rescue the HT22 cells from Aβ42-induced neurotoxicity was evaluated by the lactate dehydrogenase (LDH) assay. Cell membranes can become damaged and leaky due to Aβ42-induced toxicity and consequently, cytosolic LDH can get released which can be measured to determine cytotoxicity [37]. Our studies show that the exposure of Aβ42 to HT22 cell lines led to a 1.29-fold increase in the LDH release compared to untreated cells (Fig. 6), indicating that Aβ42 can cause damage to cell membranes. When the cells were co-incubated with Aβ42 and compounds 3a, 5a, and 7a (25 µM each), the amount of LDH released was reduced significantly (to 0.93, 0.98, and 1.04-fold respectively for compounds 3a, 5a and 7a as shown in Fig. 6). These results further provide evidence that N-benzyl, N-phenethyl and N-benzyloxybenzamides are able to protect HT22 cells from Aβ42-induced cytotoxicity.
Percentage LDH release from HT22 cell lines when exposed to Aβ42 (10 µM) in the presence and absence of lead derivatives 3a, 5a, and 7a at 25 µM. The results shown are averages of quadruplicate readings based on three independent experiments. *p < 0.001 compared to the Aβ42 control group; **p < 0.0001 compared to the Aβ42 control group (One-way ANOVA followed by Bonferroni post hoc test)
In silico physicochemical and ADME analysis
The physiochemical properties, blood-brain barrier (BBB) penetration and presence of structural alerts were predicted for representative compounds 3a, 3f, 5a and 7a using the web tool SwissADME [38]. These results show that benzamide derivatives demonstrate drug-like properties with Log P values ranging from 2.28–2.93, and are predicted to cross into BBB, without the liability of possessing any structural alerts (Table 1).
Molecular docking studies of 3a, 5a and 7a in the Aβ42 pentamer model
The binding interactions of benzamide derivatives from each series (3a, 5a, and 7a), were investigated in the Aβ42 pentamer assembly by carrying out molecular docking studies [34]. The Aβ42 pentamer model was prepared using the solved 3D structure of Aβ42 fibril (PDB: 5KK3) [39]. The two regions of Aβ42, the KLVFFA and the C-terminal IGLMVGGVVIA hydrophobic tail region are of great interest due to their role in the self-assembly process. Molecular docking studies show that the benzamide derivatives (3a, 5a and 7a) were able to interact with both the KLVFFA and IGLMVGGVVIA regions (Fig. 7). The ribbon diagram of 3a, 5a and 7a docked to Aβ42 pentamer in Fig. 7A and the 2D interaction maps are shown in Fig. 7B, C. It is interesting to note that all three derivatives exhibited similar binding modes and were in the same binding region between the C- and N-termini (Fig. 7). The aromatic rings at either ends underwent hydrophobic and van der Waal’s interactions with Leu17 and Leu34 (distance ~ 5.0 Å) respectively. This study also shows that the aromatic rings were the major contributor to their interactions with the C- and N-termini regions of Aβ42. Furthermore, the linker regions in 3a, 5a and 7a also underwent van der Waal’s interactions with Gln15, Leu17 and Leu34 amino acid residues (distance ~ 5.0 Å). This suggests that the N-benzylbenzamide, N-phenethylbenzamide and N-benzyloxybenzamide templates can undergo hydrophobic and van der Waal’s contact with the Leu17 and Leu34 to stabilize the conformation and prevent further aggregation. The CDOCKER energy and the CDOCKER interaction energies for 3a, 5a and 7a are given in (Table 2) which shows that they all form stable complexes with the Aβ42 pentamer.
Binding modes of 3a, 5a and 7a in the Aβ42 pentamer model (PDB: 5KK3). A The binding mode of 3a in the Aβ42 pentamer model (red and blue regions highlighting the C- and N-termini), and close-up view of 3a (purple), 5a (black), and 7a (green) bound to the pentamer assembly. B–D 2D interaction map of 3a, 5a, and 7a. The atoms are color-coded (carbon in grey, nitrogen in blue, and oxygen in red). The hydrophobic interactions are shown in pink and the van der Waals interactions are shown in green. The hydrogen atoms were removed to enhance clarity
Conclusion
In summary, we designed a library of 27 small molecules by replacing the α,β-unsaturated system present in the chalcone template with an amide bioisostere to obtain benzamide derivatives and evaluated them as inhibitors of Aβ42 aggregation. These derivatives possess either N-benzylbenzamide, N-phenethylbenzamide or N-benzyloxybenzamide templates. The ThT-based anti-aggregation studies show that the compounds 3a, 3f, 3h, 5a, 6f and 8a exhibit ~ 27–28% inhibition of Aβ42 aggregation. Their anti-aggregation activity was also confirmed by TEM studies which showed a reduction in the formation of Aβ42 fibrils. Strikingly, these benzamide derivatives demonstrated excellent neuroprotection against Aβ42-induced cytotoxicity in HT22 mouse hippocampal neuronal cells (80–97% cell viability at 25 µM) and were not toxic. Quantification of the Aβ42 fibril load in the ProteoStat-based live cell imaging confirmed the ability of compounds 3a, 5a, and 7a to reduce Aβ42 fibril load in the cellular environment (27–29% inhibition at 25 µM). In addition, these compounds were also able to offer neuroprotection to HT22 cells exposed to Aβ42 in the LDH assay.
These studies suggest that benzamide derivatives can bind to Aβ42 assemblies and have the potential to induce changes in their conformation and reduce their cytotoxic effects in the cellular environment. They also exhibit drug-like properties and do not possess any structural alerts, unlike the chalcone template that possesses the reactive α,β-unsaturated system which is considered as PAINS [25, 30]. Our studies show that benzamides possessing N-benzylbenzamide, N-phenethylbenzamide or N-benzyloxybenzamide templates, hold promise in the design and development of novel small molecules as Aβ42 aggregation inhibitors.
Experimental
General
All chemicals and reagents were purchased from various vendors (Sigma Aldrich USA, Alfa Aesar USA, Bio Basic Canada, AA Blocks USA, Cayman Chemical USA, Thermo Fisher USA, and Corning USA), were > 95% pure and used without further purification. The Aβ42 1,1,3,3,3-hexafluoroisopropanol (HFIP) > 95% pure, was purchased from Anaspec USA and rPeptide USA. The chemical reactions were monitored by thin-layer chromatography (TLC), Merck silica gel 60, F254. Column chromatography was carried out using Merck silica gel 230-400 mesh. The spots were visualized with short or long wavelengths (254 nm or 365 nm). The melting points of compounds were determined using a digital melting point apparatus (REACH Devices, USA). The proton NMR (1H NMR) spectra of compounds were obtained using a 300 MHz Bruker Avance spectrometer. Either CDCl3 or DMSO-d6 were used as the solvents. Coupling constants (J values) were recorded in Hertz (Hz). Abbreviations used to represent 1H NMR signals were s – singlet, d – doublet, t – triplet, m – multiplet, br s – broad singlet. The mass and purity were confirmed on Agilent 1260 Infinity liquid chromatography (LC) module equipped with 6130 Quadrupole mass spectrometry (Agilent 6100 series LCMS). The column used on LCMS was ZORBAX Eclipse AAA, 4.6 × 75 mm, 3.5 micron (Agilent Technologies, Canada). A mixture of water and acetonitrile 1:1 v/v with 0.1% formic acid (1.0 mL/min flow rate), was used as the solvent system to assess compound purity and mass. The compounds were of ≥ 95% pure as determined by LCMS analysis before testing in various biological screening assays.
General procedure for the synthesis of 3a-h, 5a-h and 7a-h
Phenylmethanamine (2) (1 mmol), or 2-phenylethan-1-amine (4) (1 mmol), or O-benzylhydroxylamine (6) (1 mmol) was added into a 50 ml round bottom flask containing 15 ml anhydrous tetrahydrofuran (THF). To this mixture, 1.2 equivalent of triethylamine (TEA) and 1.0 equivalent of corresponding acid chlorides with various R groups were added (Scheme 1). The reaction was kept at either room temperature or under reflux for 6–24 h and monitored by TLC. After the reaction was complete, the solvent was evaporated in vacuo and the resulting reaction mixture was dissolved in 15 mL ethyl acetate (EtOAc), and then washed s with 15 mL brine solution (x 3). The organic layer was collected and dried over anhydrous MgSO4 and filtered. The organic solvent was removed in vacuo and the residue/oil obtained was further purified by silica gel column chromatography using a combination of solvent systems (1:1 n-hexanes: EtOAc or 5:2 n-hexanes: EtOAc or 9:1 dichloromethane (DCM): Methanol (MeOH) or 4:1 DCM: n-hexanes) to afford the final compounds 3a–h, 5a–h and 7a–h. The yields ranged from 72–93%. The analytical data for these compounds are given in the following section. The 1H NMR spectra and LCMS chromatograms are provided in the Supplementary Material document.
N-Benzylbenzamide (3a): White solid, 92.8% [40, 41]. 1H NMR (300 MHz, DMSO-d6) δ 9.00 (t, J = 5.6 Hz, 1H), 7.90–7.81 (m, 2H), 7.53–7.39 (m, 3H), 7.29–7.10 (m, 5H), 4.45 (d, J = 6.0 Hz, 2H). mp: 105–107 °C. ESI-MS, m/z for C14H14NO [M + H]+ 212.1. Purity: 99.9% (LCMS).
N-Benzylisonicotinamide (3b): White solid, 72% [41, 42]. 1H NMR (300 MHz, DMSO-d6) δ 9.33 (t, J = 5.3 Hz, 1H), 8.79–8.67 (m, 2H), 7.85–7.75 (m, 2H), 7.34–7.27 (m, 5H), 4.51 (d, J = 6.0 Hz, 2H). mp: 95–98 °C. ESI-MS, m/z for C13H13N2O [M + H]+ 213.1. Purity: 99.0% (LCMS).
N-Benzylpicolinamide (3c): White solid, 74.3% [42]. 1H NMR (300 MHz, DMSO-d6) δ 9.32 (br s, 1H), 8.67–8.65 (m, 1H), 8.10–7.96 (m, 2H), 7.64–7.59 (m,1H), 7.35– 7.20 (m, 5H), 4.50 (d, J = 6.4 Hz, 2H). mp: 94–97 °C. ESI-MS, m/z for C13H13N2O [M + H]+ 213.1. Purity: 99.6% (LCMS).
N-Benzyl-1H-pyrrole-2-carboxamide (3d): White solid, 79.6% [43]. 1H NMR (300 MHz, CDCl3) δ 9.50 (br s, 1H), 7.51–7.12 (m, 5H), 6.92–6.90 (m, 1H), 6.53–6.51 (m, 1H), 6.23–6.20 (m, 1H), 6.13 (br s, 1H), 4.60 (d, J = 5.9 Hz, 2H). mp:122–125 °C. ESI-MS, m/z for C12H13N2O [M + H]+ 201.1. Purity: 96.1% (LCMS).
N-Benzylfuran-2-carboxamide (3e): White solid, 93% [44]. 1H NMR (300 MHz, DMSO-d6) δ 9.04 (t, J = 5.8 Hz, 1H), 7.84 (d, J = 3.7 Hz, 1H), 7.36–7.21 (m, 5H), 7.25 (d, J = 6.0 Hz, 1H), 6.62 (d, J = 5.0 Hz, 1H), 4.46 (d, J = 6.0 Hz, 2H). mp: 120–123 °C. ESI-MS, m/z for C12H12NO2 [M + H]+ 202.1. Purity: 99.6% (LCMS).
N-Benzylthiophene-2-carboxamide (3 f): White solid, 78.7% [42]. 1H NMR (300 MHz, DMSO-d6) δ 9.04 (t, J = 5.7 Hz, 1H), 7.88–7.81 (m, 1H), 7.39–7.19 (m, 5H), 7.15–7.05 (m, 2H), 4.45 (d, J = 6.2 Hz, 2H). mp: 115–117 °C. ESI-MS, m/z for C12H12NOS [M + H]+ 218.1. Purity: 99.7% (LCMS).
N-Benzyl-1-naphthamide (3 g): White solid, 87.6% [45]. 1H NMR (300 MHz, DMSO-d6) δ 9.09 (t, J = 5.7 Hz, 1H), 8.22–8.19 (m, 1H), 8.06–7.95 (m, 2H), 7.66–7.53 (m, 4H), 7.38–7.26 (m, 5H), 4.55 (d, J = 6.2 Hz, 2H). mp: 127–129 °C. ESI-MS, m/z for C18H16NO [M + H]+ 262.1. Purity: 98.3% (LCMS).
N-Benzylquinoline-2-carboxamide (3 h): White solid, 93.6% [46]. 1H NMR (300 MHz, DMSO-d6) δ 9.47 (t, J = 6.2 Hz, 1H), 8.58 (d, J = 8.5 Hz, 1H), 8.20–8.04 (m, 3H), 7.88 (t, J = 7.6 Hz, 1H), 7.73 (t, J = 7.5 Hz, 1H), 7.40–7.22 (m, 5H), 4.58 (d, J = 6.2 Hz, 2H). mp: 128–130 °C. ESI-MS, m/z for C17H15N2O [M + H]+ 263.1. Purity: 99.5% (LCMS).
N-Phenethylbenzamide (5a): White solid, [47]. 1H NMR (300 MHz, CDCl3) δ 7.78–7.59 (m, 2H), 7.52–7.28 (m, 5H), 7.24 (q, J = 4.9, 4.4 Hz, 3H), 6.10 (s, 1H), 3.71 (q, J = 6.9 Hz, 2H), 2.93 (t, J = 6.9 Hz, 2H). mp: 122–125 °C. ESI-MS, m/z for C15H16NO [M + H]+ 226.1. Purity: 99.9% (LCMS).
N-Phenethylisonicotinamide (5b): White solid, 76% [42]. 1H NMR (300 MHz, CDCl3) δ 8.71–8.68 (m, 2H), 7.50–7.48 (m, 2H), 7.36–7.20 (m, 5H), 6.15 (br s, 1H), 3.72 (q, J = 6.6 Hz, 2H), 2.93 (t, J = 6.8 Hz, 2H). mp: 129-130 °C. ESI-MS, m/z for C14H15N2O [M + H]+ 227.1. Purity: 98.7% (LCMS).
N-Phenethylpicolinamide (5c): Yellowish oil, 74.5% [48]. 1H NMR (300 MHz, CDCl3) δ 8.53 (d, J = 4.5 Hz, 1H), 8.22 (d, J = 7.8 Hz, 1H), 8.16 (s, 1H), 7.85 (td, J = 7.7, 1.6 Hz, 1H), 7.44–7.23 (m, 6H), 3.72 (q, J = 7.0 Hz, 2H), 2.97 (t, J = 7.3 Hz, 2H). ESI-MS, m/z for C14H15N2O [M + H]+ 227.1. Purity: 99.9% (LCMS).
N-Phenethyl-1H-pyrrole-2-carboxamide (5d): White solid, 82.8% [49]. 1H NMR (300 MHz, CDCl3) δ 9.38 (br s, 1H), 7.37–7.26 (m, 2H), 7.26–7.17 (m, 3H), 6.91–6.88 (m, 1H), 6.41–6.38 (m, 1H), 6.20–6.17 (m, 1H), 5.84 (br s, 1H), 3.66 (q, J = 6.7 Hz, 2H), 2.88 (t, J = 6.9 Hz, 2H). mp: 134–136 °C. ESI-MS, m/z for C13H15N2O [M + H]+ 215.1. Purity: 98.6% (LCMS).
N-Phenethylfuran-2-carboxamide (5e): White solid, 75.5% [47]. 1H NMR (300 MHz, CDCl3) δ 7.38–7.20 (m, 5H), 7.08 (d, J = 3.5 Hz, 1H), 6.46–6.36 (m, 2H), 3.67 (q, J = 7.0 Hz, 2H), 2.90 (t, J = 7.0 Hz, 2H). mp: 115–117 °C. ESI-MS, m/z for C13H14NO2 [M + H]+ 216.1. Purity: 99.7% (LCMS).
N-Phenethylthiophene-2-carboxamide (5 f): White solid, 73.2% [50]. 1H NMR (300 MHz, CDCl3) δ 7.41–7.18 (m, 7H), 7.08–7.01 (m, 1H), 6.00 (br s, 1H), 3.67 (q, J = 7.0 Hz, 2H), 2.90 (t, J = 7.0 Hz, 2H). mp: 116–119 °C. ESI-MS, m/z for C13H14NOS [M + H]+ 232.1. Purity: 99.8% (LCMS).
N-Phenethyl-1-naphthamide (5 g): White solid, 72.8% [47]. 1H NMR (300 MHz, CDCl3) δ 8.20–8.16 (m, 1H), 7.81–7.75 (m, 2H), 7.50–7.44 (m, 3H), 7.40–7.23 (m, 6H), 5.96 (br s, 1H), 3.82 (q, J = 6.6 Hz, 2H), 2.99 (t, J = 6.8 Hz, 2H). mp: 125–128 °C. ESI-MS, m/z for C19H18NO [M + H]+ 276.1. Purity: 99.2% (LCMS).
N-Phenethylquinoline-2-carboxamide (5 h): White solid, 79.9% [51]. 1H NMR (300 MHz, CDCl3) δ 8.36–8.27 (m, 1H), 8.30 (s, 2H), 8.04 (d, J = 8.5 Hz, 1H), 7.86 (dd, J = 8.1, 1.5 Hz, 1H), 7.74 (ddd, J = 8.5, 6.9, 1.5 Hz, 1H), 7.60 (dd, J = 8.2, 6.8 Hz, 1H), 7.36–7.19 (m, 5H), 3.78 (q, J = 7.6 Hz, 2H), 2.99 (t, J = 7.3 Hz, 2H). mp: 81–84 °C. ESI-MS, m/z for C18H17N2O [M + H]+ 277.1. Purity: 99.56% (LCMS).
N-(Benzyloxy)benzamide (7a): White solid, 77.9% [33]. 1H NMR (300 MHz, DMSO-d6) δ 11.76 (s, 1H), 7.75 (d, J = 7.6 Hz, 2H), 7.55–7.32 (m, 8H), 4.93 (s, 2H). mp: 104–106 °C. ESI-MS, m/z for C14H14NO2 [M + H]+ 228.1. Purity: 99.6% (LCMS).
N-(Benzyloxy)isonicotinamide (7b): White solid, 86.1%. 1H NMR (300 MHz, DMSO-d6) δ 12.07 (br s, 1H), 8.72 (d, J = 4.9 Hz, 2H), 7.65 (d, J = 5.3 Hz, 2H), 7.43 (dd, J = 19.5, 6.8 Hz, 5H), 4.96 (s, 2H). mp: 117–120 °C. ESI-HRMS, m/z for C13H14N2O2 [M + H]+ 229.0977, found 229.0966. Purity: 99.4% (LCMS).
N-(Benzyloxy)picolinamide (7c): Yellowish oil, 77.0% [52]. 1H NMR (300 MHz, DMSO-d6) δ 12.04 (s, 1H), 8.61 (d, J = 4.7 Hz, 1H), 8.00 (d, J = 4.0 Hz, 2H), 7.62–7.37 (m, 6H), 4.95 (s, 2H). ESI-MS, m/z for C13H13N2O2 [M + H]+ 229.1. Purity: 99.7% (LCMS).
N-(Benzyloxy)-1H-pyrrole-2-carboxamide (7d): White solid, [53]. 1H NMR (300 MHz, CDCl3) δ 9.51 (br s, 1H), 7.39–7.24 (m, 5H), 6.91 (td, J = 2.7, 1.3 Hz, 1H), 6.52 (dd, J = 3.8, 2.5 Hz, 1H), 6.21 (dt, J = 3.7, 2.6 Hz, 1H), 6.13 (br s, 1H), 4.60 (d, J = 5.9 Hz, 2H). mp: 77–80 °C. ESI-MS, m/z for C12H13N2O2 [M + H]+ 217.1. Purity: 96.5% (LCMS).
N-(Benzyloxy)furan-2-carboxamide (7e): White solid, 77.9% [54]. 1H NMR (300 MHz, DMSO-d6) δ 11.74 (s, 1H), 7.86 (s, 1H), 7.46–7.36 (m, 5H), 7.10 (d, J = 4.3 Hz, 1H), 6.63 (s, 1H), 4.90 (s, 2H). mp: 82–85 °C. ESI-MS, m/z for C12H12NO3 [M + H]+ 218.1. Purity: 99.9% (LCMS).
N-(Benzyloxy)thiophene-2-carboxamide (7 f): White solid, 92.9% [55]. 1H NMR (300 MHz, DMSO-d6) δ 11.79 (s, 1H), 7.83–7.81 (m, 1H), 7.65–7.62 (m, 1H), 7.46–7.34 (m, 5H), 7.15 (d, J = 4.3 Hz, 1H), 4.92 (s, 2H). mp: 103–106 °C. ESI-MS, m/z for C12H12NO2S [M + H]+ 234.1. Purity: 98.7% (LCMS).
N-(Benzyloxy)-1-naphthamide (7 g): White solid, 72.1% [56]. 1H NMR (300 MHz, CDCl3) δ 8.30–8.21 (m, 2H), 7.90 (d, J = 8.2 Hz, 1H), 7.87–7.79 (m, 1H), 7.58–7.32 (m, 8H), 5.13 (s, 2H). mp: 132–135 °C. ESI-MS, m/z for C18H16NO2 [M + H]+ 278.1. Purity: 98.9% (LCMS).
N-(Benzyloxy)quinoline-2-carboxamide (7 h): White solid, 80.1% [57]. 1H NMR (300 MHz, CDCl3) δ 10.35 (s, 1H), 8.26 (dd, J = 8.5 1.5 Hz, 2H), 8.02 (d, J = 8.5 Hz, 1H), 7.9 (dd, J = 8.2, 1.4 Hz, 1H), 7.74 (dd, J = 8.5, 6.9 Hz, 1H), 7.61 (dd, J = 8.1, 6.8 Hz, 1H), 7.52–7.49 (m, 2H), 7.41–7.38 (m, 2H), 5.11 (s, 2H). ESI-MS, m/z for C17H18N2O2 [M + H]+ 279.1. Purity: 99.7% (LCMS).
General procedure for the synthesis of 3i, 5i, 7i
Phenylmethanamine (2) (1 mmol), or 2-phenylethylamine (4) (1 mmol), or O-benzyloxyamine (6) (1 mmol) was added into a 50 mL round bottom flask containing 15 mL anhydrous THF. Next, 1.0 equivalent 1H-indole-2-carboxylic acid, 1.2 equivalent EDC hydrochloride, 1.5 equiv 1-hydroxybenzotriazole (HOBt), and 2.2 equivalent TEA were added in sequence. The reaction was kept at room temperature overnight. The solvent was evaporated in vacuo and the resulting reaction mixture was dissolved in 15 ml EtOAc and washed with 15 ml brine solution (x 3). The organic layer was collected, dried over anhydrous MgSO4 and filtered. The organic solvent was then removed in vacuo to obtain a residue/oil which was further purified by silica gel column chromatography using either 1:1 n-hexanes: EtOAc or 5:2 n-hexanes: EtOAc as eluents to afford 3i, 5i and 7i in yields ranging from 80–92%. Their analytical data is given below. The 1H NMR spectra and LCMS chromatograms are provided in the Supporting Information.
N-Benzyl-1H-indole-2-carboxamide (3i): White solid, 90.6% [45]. 1H NMR (300 MHz, DMSO-d6) δ 11.56 (s, 1H), 9.00 (t, J = 6.1 Hz, 1H), 7.57 (d, J = 8.0 Hz, 1H), 7.41 (d, J = 8.2 Hz, 1H), 7.38–7.27 (m, 4H), 7.17–7.12 (m, 3H), 7.03–6.98 (m, 1H), 4.49 (d, J = 6.0 Hz, 2H). mp: 229–232 °C. ESI-MS, m/z for C16H15N2O [M + H]+ 251.1.Purity: 95.4% (LCMS).
N-(Phenethyl)-1H-indole-2-carboxamide (5i): White solid, 87.5% [58]. 1H NMR (300 MHz, DMSO-d6) δ 11.51 (br s, 1H), 8.57 (t, J = 5.6 Hz, 1H), 7.60 (d, J = 7.9 Hz, 1H), 7.43 (d, J = 8.2 Hz, 1H), 7.34–7.01 (m, 6H), 3.49–3.55 (m, 2H), 2.87 (t, J = 7.4 Hz, 2H). mp: 191–194 °C. ESI-MS, m/z for C17H17N2O [M + H]+ 265.1. Purity: 97.6% (LCMS).
N-(Benzyloxy)-1H-indole-2-carboxamide (7i): White solid, 82.7% [59]. 1H NMR (300 MHz, DMSO-d6) δ 11.81 (s, 1H), 11.66 (s, 1H), 7.60 (d, J = 7.9 Hz, 1H), 7.42 (dq, J = 14.0, 7.0, 6.4 Hz, 6H), 7.20 (t, J = 7.5 Hz, 1H), 7.09–6.96 (m, 2H), 4.96 (s, 2H). mp: 152–155 °C. ESI-MS, m/z for C16H15N2O2 [M + H]+ 267.1. Purity: 95.6% (LCMS).
Biological activity assay
ThT-based Aβ42 aggregation inhibition assay
The ability of benzamide derivatives 3a–i, 5a–i and 7a–i to inhibit the Aβ42 aggregation was evaluated using the thioflavin T (ThT) based fluorescence kinetic assay [34]. A 15 μM ThT solution was prepared in ultra-pure-water (UPW) in 50 mM glycine buffer. The pH of the solution was adjusted to 7.4 with 50 mM sodium hydroxide solution. The aggregation assay was carried out using 215 mM sodium phosphate dibasic heptahydrate buffer prepared in UPW and the pH was adjusted to 7.4 by using 50 mM hydrochloric acid solution. The Aβ42•HFIP > 95% pure (Anaspec, USA) was treated with 1% ammonium hydroxide solution to afford 1 mg/mL stock solution. The stock solution was then vortexed and sonicated for 5 min before diluted to 50 μM working solution using the assay buffer. The stock solutions of test compounds 3a–i, 5a–i, 7a–i and resveratrol (RVT), were prepared in assay buffer with DMSO as the solubilizing agent (final concentration per well was less than 2%). The 24 h aggregation kinetics assay was carried out using 384-well plates (Costar, black, clear-bottom), by adding 44 μL of ThT, 12 μL of assay buffer, 16 μL of Aβ42 solution (10 µM final well concentration), and 8 μL of test compound dilution (25 µM) into each well. The Aβ42 control wells contained ThT, Aβ42 solution and the assay buffer without any test compound. The plates were covered with a transparent plate cover and were incubated at 37 °C for 24 h with shaking at 300 cpm between readings for 30 s. The readings were taken every 10 min (bottom reading), with an excitation wavelength of 440 nm and an emission wavelength of 490 nm using the BioTek Synergy H1 microplate reader. Each sample was measured in triplicate readings and the results were obtained based on three independent experiments.
TEM studies to determine Aβ42 morphology
The TEM assay was carried out by first running the Aβ42 aggregation assay (as described above in the ThT assay) [34] by incubating test compounds 3a, 3f, 3h, 5a, 5f and 7a (25 µM each) with Aβ42 (10 μM) for 24 h at 37 °C. The TEM grids were prepared by aliquoting 20 μL each sample from the 384-well plate after the 24 h incubation period. These samples were loaded onto 400-mesh formvar-coated copper grid (Electron Microscopy Sciences, USA). The grids were air-dried overnight before washing with 40 μL of UPW for three times. After washing, the grids were further air-dried for another 4 h. Then, grids were stained with 20 μL 2% phosphotungstic acid solution for 30 s. The imaging studies were carried out using a Philips CM 10 TEM (Dept. of Biology, University of Waterloo) at 60 kV and the micrographs were obtained through 14-megapixel AMT camera at 60,000X magnification.
Aβ42-induced cytotoxicity assay in mouse hippocampal HT22 neuronal cell line
The cytotoxicity and neuroprotective effects of benzamide derivatives 3a, 3f, 3h, 5a, 5f and 7a toward HT22 cells was carried out by using the UV-based cell metabolism assay kit (cell counting kit 8, CCK-8, TargetMol, USA). The HT22 cells were cultured in DMEM/F12 1:1 with the addition of glutamate supplemented with 10% FBS and 1% penicillin and streptomycin at 37 °C in 5% CO2. The cells were seeded at a density of 50,000 cells per 1 mL. After 24 h, the cells were treated with various compounds (25 μM) prepared in phosphate buffer pH 7.4 and DMSO (less than 2%) and incubated for 48 h. After that, the CCK-8 reagent was added to each well, and incubated for another 2 h, before measuring the UV absorbance at 450 nm as per the manufacturer’s protocol. The neuroprotective effects of test compounds (25 µM) toward Aβ42-induced cytotoxicity was determined by treating the HT22 cells with 5 µM of Aβ42•HFIP (> 95%, rPeptide USA), and then incubating for 48 h at 37 °C [34]. The cell viability assay was determined by adding the CCK-8 reagent and incubating for another 2 h to measure the absorbance at 450 nm. The cell viability was calculated as average percent cell viability based on quadruplicate readings and based on three independent experiments.
Fluorescence live cell imaging in mouse hippocampal HT22 cell line
The Aβ42 used in the assay was purchased from Bachem Americas Inc, Torrance, CA, USA. The HT22 cells were cultured and treated as per Aβ42-induced cytotoxicity assay. After treatment, the cells were washed with PBS and then stained with ProteoStat® working solution (prepared according to the vendor’s manual, ENZO Life Sciences, USA) for 20 min under dark at 37 °C [35]. Then the cells were washed with PBS three times with 5 min incubation at 37 °C to rinse out the excessive amount of dye. The cells were then visualized with an EVOS® FL Auto imaging system with an RFP light cube at fixed light intensity. The binary masks were generated by thresholding the original fluorescence images using the Huang algorithm in ImageJ software and used to measure the integrated gray value (IGV) of ProteoStat®-positive stains. The results presented are averages of three independent experiments with three randomized views.
LDH release cytotoxicity assay in mouse hippocampal HT22 cell line
The CytoScan® LDH assay kit was used to determine the LDH release (G-Biosciences, St. Louis, MO, USA) [37]. The reagents and solutions were prepared according to vendor’s manual. The HT22 cells were cultured and treated with 10 µM Aβ42 (Bachem, USA) as per Aβ42-induced cytotoxicity assay protocol. After treatment, 50 µL of the cell medium was transferred to a new well plate followed by the addition of 50 µL reaction mixture and further incubated for 20 min at 37 °C. This was followed by the addition of 50 µL stop solution before taking the readings at 490 nm and 680 nm. The percentage LDH release was calculated by subtracting the absorbance reading at 680 nm from that of 490 nm, and compared to Aβ42 control group. The results shown are averages of quadruplicate readings based on three independent experiments.
Molecular docking studies of compounds in Aβ42
The binding interactions 3a (N-benzylbenzamide), 5a (N-phenethylbenzamide) and 7a (N-benzyloxybenzamide) Aβ42 were investigated by carrying out molecular docking studies conducted on Discovery Studio software program v20.1.0.19295, Structure-Based-Design program (BIOIA Inc. San Diego, USA). The Aβ42 pentamer model was prepared by extracting the coordinates from the solved 3D structure of Aβ42 fibril (PDB id: 5KK3) [34, 39]. A binding sphere of 20 Å radius was selected and defined as the ligand binding site which covers the KLVFFA and the C-terminal IGLMVGGVVIA hydrophobic region of the Aβ42. Compounds 3a, 5a and 7a were built in 3D using the Small Molecules module in the software and were subjected to energy minimization protocol (1000 and 2000 steps of steepest descent followed by conjugate gradient method) using CHARMm force field. The CDOCKER algorithm was used to carry out the docking studies. The Receptor-Ligand Interactions module in the software was used to perform CDOCKER simulations. The docking simulation included 2000 heating steps, a heating target temperature of 700 K and 5000 cooling steps and a cooling target temperature of 300 K. The top ligand binding modes of 3a, 5a and 7a with Aβ42 pentamer assembly was analyzed by evaluating the CDOCKER energy and CDOCKER interaction energies (in kcal/mol). Further analysis was carried out by determining the polar and nonpolar contacts of test compounds in the Aβ42 pentamer assembly.
In silico physicochemical and ADME analysis
The physicochemical properties including H-bond acceptors, H-bond donors, topological polar surface area (TPSA), blood-brain barrier permeability (BBB), lipophilicity parameter Log P, and structural alert flags were calculated for representative compounds (3a, 3f, 5a, 7a and chalcone) using the SwissADME web tool [38].
References
Lyketsos CG, Carrillo MC, Ryan JM, Khachaturian AS, Trzepacz P, Amatneik J, Miller SM. Neuropsychiatric symptoms in Alzheimer’s disease. Alzheimers Dement. 2011;7:532–9. https://doi.org/10.1016/J.JALZ.2011.05.2410
Atri A. The Alzheimer’s disease clinical spectrum: diagnosis and management. Med Clin North Am. 2019;103:263–93. https://doi.org/10.1016/J.MCNA.2018.10.009
Porsteinsson AP, Isaacson RS, Knox S, Sabbagh MN, Rubino I. Diagnosis of early Alzheimer’s disease: clinical practice in 2021. J Prev Alzheimers Dis. 2021;8:371–86. https://doi.org/10.14283/JPAD.2021.23
Knopman DS, Amieva H, Petersen RC, Chetelat G, Holtzman DM, Hyman BT, Jones DT. Alzheimer disease. Nat Rev Dis Primers. 2021;7:33. https://doi.org/10.1038/S41572-021-00269-Y
Selkoe DJ, Hardy J. The amyloid hypothesis of Alzheimer’s disease at 25 years. EMBO Mol Med. 2016;8:595–608. https://doi.org/10.15252/EMMM.201606210
Kumar A, Singh A, Ekavali. A review on Alzheimer’s disease pathophysiology and its management: An update. Pharmacol. Rep. 2015;67:195–203. https://doi.org/10.1016/j.pharep.2014.09.004
Deture MA, Dickson DW. The neuropathological diagnosis of Alzheimer’s disease. Mol Neurodegener. 2019;14:32. https://doi.org/10.1186/S13024-019-0333-5
Kametani F, Hasegawa M. Reconsideration of amyloid hypothesis and tau hypothesis in Alzheimer’s disease. Front Neurosci. 2018;12:25. https://doi.org/10.3389/FNINS.2018.00025
Aisen PS, Cummings J, Jack CR, Morris JC, Sperling R, Frolich L, Dubois B. On the path to 2025: understanding the Alzheimer’s disease continuum. Alzheimers Res Ther. 2017;9:60. https://doi.org/10.1186/S13195-017-0283-5
Zhang H, Ma Q, Zhang YW, Xu H. Proteolytic processing of Alzheimer’s β-amyloid precursor protein. J Neurochem. 2012;120:9–21. https://doi.org/10.1111/J.1471-4159.2011.07519.X
Cummings J, Lee G, Nahed P, Kambar MEZN, Zhong K, Fonesca J, Taghva K Alzheimer’s disease drug development pipeline: 2022. Alzheimers Dement. 2022;8:e12295. https://doi.org/10.1002/TRC2.12295
Cummings J, Lee G, Ritter A, Sabbagh M, Zhong K. 2020. Alzheimer’s disease drug development pipeline: 2020. Alzheimers and Dement. 2020;6:e12050. https://doi.org/10.1002/trc2.12050
Sevigny J, Chiao P, Bussière T. The antibody aducanumab reduces Aβ plaques in Alzheimer’s disease. Nature. 2016;537:50–56. https://doi.org/10.1038/nature19323
Salloway S, Chalkias S, Barkhof F, Barakos J, Purcell D, Suhy J, Smirnakis K. Amyloid-related imaging abnormalities in 2 Phase 3 studies evaluating aducanumab in patients with early Alzheimer disease. JAMA Neurol. 2022;79:13–21. https://doi.org/10.1001/JAMANEUROL.2021.4161
Van Dyck CH, Swanson CJ, Aisen P, Bateman RJ, Chen C, Gee M, Iwatsubo T. Lecanemab in early Alzheimer’s disease. N Engl J Med. 2023;388:142–3. https://doi.org/10.1056/NEJMOA2212948
Reardon S. Alzheimer’s drug donanemab: what promising trial means for treatments. Nature. 2023;617:232–3. https://doi.org/10.1038/D41586-023-01537-5
Makurvet FD. Biologics vs. small molecules: Drug costs and patient access. Med Drug Discov. 2021;9:100075. https://doi.org/10.1016/J.MEDIDD.2020.100075
Mohamed T, Yeung JCK, Vasefi MS, Beazely MA, Rao PPN. Development and evaluation of multifunctional agents for potential treatment of Alzheimer’s disease: Application to a pyrimidine-2,4-diamine template. Bioorg Med Chem Lett. 2012;22:4707–12. https://doi.org/10.1016/j.bmcl.2012.05.077
Mohamed T, Shakeri A, Tin G, Rao PPN. Structure-activity relationship studies of isomeric 2,4-diaminoquinazolines on β-amyloid aggregation kinetics. ACS Med Chem Lett. 2016;7:502–7. https://doi.org/10.1021/acsmedchemlett.6b00039
Mohamed T, Shakeri A, Rao PPN. Amyloid cascade in Alzheimer’s disease: recent advances in medicinal chemistry. Eur J Med Chem. 2016;113:258–72. https://doi.org/10.1016/J.EJMECH.2016.02.049
Sashidhara KV, Kumar M, Modukuri RK, Sonkar R, Bhatia G, Khanna AK, Shukla R. Synthesis and anti-inflammatory activity of novel biscoumarin–chalcone hybrids. Bioorg Med Chem Lett. 2011;21:4480–4. https://doi.org/10.1016/j.bmcl.2011.06.002
Kim MJ, Lee YH, Kwak J, Na Y, Yoon HG. Protective effects of a chalcone derivative against Aβ-induced oxidative stress and neuronal damage. BMB Rep. 2011;44:730–4. https://doi.org/10.5483/BMBREP.2011.44.11.730
Cao Z, Yang J, Xu R, Song Q, Zhang X, Liu H, Deng Y. Design, synthesis and evaluation of 4’-OH-flurbiprofen-chalcone hybrids as potential multifunctional agents for Alzheimer’s disease treatment. Bioorg Med Chem. 2018;26:1102–15. https://doi.org/10.1016/J.BMC.2018.01.030
Liu HR, Liu XJ, Fan HQ, Tang JJ, Gao XH, Liu WK. Design, synthesis and pharmacological evaluation of chalcone derivatives as acetylcholinesterase inhibitors. Bioorg Med Chem. 2014;22:6124–33. https://doi.org/10.1016/J.BMC.2014.08.033
Jackson PA, Widen JC, Harki DA, Brummond KM. Covalent modifiers: a chemical perspective on the reactivity of α,β-unsaturated carbonyls with thiols via hetero-Michael addition reactions. J Med Chem. 2017;60:839–85. https://doi.org/10.1021/ACS.JMEDCHEM.6B00788
Zhuang C, Zhang W, Sheng C, Zhang W, Xing C, Miao Z. Chalcone: a privileged structure in medicinal chemistry. Chem Rev. 2017;117:762–7810. https://doi.org/10.1021/acs.chemrev.7b00020
Baell J, Walters MA. Chemistry: chemical con artists foil drug discovery. Nature. 2014;513:481–3. https://doi.org/10.1038/513481A
Baell JB. Redox-active nuisance screening compounds and their classification. Drug Discov Today. 2011;16:840–1. https://doi.org/10.1016/j.drudis.2011.06.011
Burgos-Morón E, Calderón-Montaño JM, Salvador J, Robles A, Lazaro ML. The dark side of curcumin. Int J Cancer. 2010;126:1771–5. https://doi.org/10.1002/IJC.24967
Baell JB, Holloway GA. New substructure filters for removal of pan assay interference compounds (PAINS) from screening libraries and for their exclusion in bioassays. J Med Chem. 2010;53:2719–40. https://doi.org/10.1021/JM901137J
Brahmachari S, Paul A, Segal D, Gazit E. Inhibition of amyloid oligomerization into different supramolecular architectures by small molecules: mechanistic insights and design rules. Future Med Chem. 2017;9:797–810. https://doi.org/10.4155/FMC-2017-0026
Katritzky AR, Kirichenko N, Rogovoy BV Efficient conversions of carboxylic acids into O-alkyl, N-alkyl and O,N-dialkylhydroxamic acids. Synthesis 2003; 2777-80. https://doi.org/10.1055/S-2003-42488
Majewski MW, Cho S, Miller PA, Franzblau SG, Miller MJ. Syntheses and evaluation of substituted aromatic hydroxamates and hydroxamic acids that target Mycobacterium tuberculosis. Bioorg Med Chem Lett. 2015;25:4933–6. https://doi.org/10.1016/J.BMCL.2015.04.099
Gujral SS, Shakeri A, Hejazi L, Rao PPN. Design, synthesis and structure-activity relationship studies of 3-phenylpyrazino[1,2-a]indol-1(2H)-ones as amyloid aggregation and cholinesterase inhibitors with antioxidant activity. Eur J Med. Chem. Rep. 2022;6:100075. https://doi.org/10.1016/J.EJMCR.2022.100075
Navarro S, Carija A, Munoz-Torrero D, Ventura S. A fast and specific method to screen for intracellular amyloid inhibitors using bacterial model systems. Eur. J. Med. Chem. 2016;121:785–92. https://doi.org/10.1016/j.ejmech.2015.10.044
Sandler OP, Miller Y. Molecular mechanisms of membrane-associated amyloid aggregation: computational perspective and challenges. Biochem. Biophys. Acta. Biomembr. 2018;1860:1889–905. https://doi.org/10.1016/j.bbamem.2018.03.014
Kumar P, Nagarajan A, Uchil PD Cold Spring Harb. Protoc. 2018; 2018. https://doi.org/10.1101/pdb.prot095497
Daina A, Michielin O, Zoete V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep. 2017;7:42717. https://doi.org/10.1038/SREP42717
Colvin MT, Silvers R, Ni QZ, Can TV, Sergeyev I, Rosay M, Griffin RG. Atomic resolution structure of monomorphic Aβ42 amyloid fibrils. J Am Chem Soc. 2016;138:9663–74. https://doi.org/10.1021/JACS.6B05129
Ojeda-Porras A, Santana AH, Sanchez DG. Direct amidation of carboxylic acids with amines under microwave irradiation using silica gel as a solid support. Green Chem. 2015;17:3157–63. https://doi.org/10.1039/C5GC00189G
Li Y, Chen H, Liu J, Wan X, Xu Q. Clean synthesis of primary to tertiary carboxamides by CsOH-catalyzed aminolysis of nitriles in water. Green Chem. 2016;18:4865–70. https://doi.org/10.1039/C6GC01565D
Laha JK, Gulati U, Gupta A. Decarboxylative amidation of aryl/heteroarylacetic acids via activated esters through traceless α-functionalized benzylic radicals. Org Lett. 2023;25:3402–6. https://doi.org/10.1021/ACS.ORGLETT.3C00927
Darnowski MG, Lanosky TD, Paquette AR, Boddy CN. Synthesis of a constitutional Isomer of armeniaspirol A, pseudoarmeniaspirol A, via Lewis acid-mediated rearrangement. J Org Chem. 2022;87:15634–43. https://doi.org/10.1021/ACS.JOC.2C02331
Hamstra DFJ, Lenstra DC, Koenders TJ, Rutjes PJT, Mecinovic J. Poly(methylhydrosiloxane) as a green reducing agent in organophosphorus-catalysed amide bond formation. Org Biomol Chem. 2017;15:6426–32. https://doi.org/10.1039/C7OB01510K
Das J, Banerjee D. Nickel-catalyzed phosphine free direct N-alkylation of amides with alcohols. J Org Chem. 2018;83:3378–84. https://doi.org/10.1021/acs.joc.7b03215
Ren W, Yamane M. Mo(CO)(6)-mediated carbamoylation of aryl halides. J Org Chem. 2010;75:8410–5. https://doi.org/10.1021/JO101611G
Dumas AM, Molander GA, Bode JW. Amide-forming ligation of acyltrifluoroborates and hydroxyamines in water. Angew Chem Int Ed Engl. 2012;51:5683–6. https://doi.org/10.1002/anie.201201077
Xu S, Jiang D, Peng Z, Hu L, Liu T, Zhao L, Zhao J. Ynamide-mediated peptide bond formation: Mechanistic study and synthetic applications. Angew Chem Int Ed Engl. 2022;61:e202212247. https://doi.org/10.1002/ANIE.202212247
Dyson L, Wright AD, Young KA, Sakoff JA, McCluskey A. Synthesis and anticancer activity of focused compound libraries from the natural product lead, oroidin. Bioorg Med Chem. 2014;22:1690–9. https://doi.org/10.1016/J.BMC.2014.01.021
Fu R, Yang Y, Zhang J, Shao J, Xia X, Ma Y, Yuan R. Direct oxidative amidation of aldehydes with amines catalyzed by heteropolyanion-based ionic liquids under solvent-free conditions via a dual-catalysis process. Org Biomol Chem. 2016;14:1784–93. https://doi.org/10.1039/C5OB02376A
Gonec T, Bobal P, Sujan J, Pesko M, Guo J, Kralova K, Jampile J. Investigating the spectrum of biological activity of substituted quinoline-2-carboxamides and their isosteres. Molecules. 2012;17:613–44. https://doi.org/10.3390/MOLECULES17010613
Chen X, Han J, Zhu Y, Yuan C, Zhang J, Zhao Y. Transformation of masked benzyl alcohols to o-aminobenzaldehydes through C-H activation: a facile approach to quinazolines. Chem Commun. 2016;52:10241–4. https://doi.org/10.1039/C6CC05560E
Huang M, Hou J, Yang R, Zhang L, Zhu X, Wan W. A catalyst system, copper/N-methoxy-1H-pyrrole-2-carboxamide, for the synthesis of phenothiazines in poly(ethylene glycol). Synthesis. 2014;46:3356–64. https://doi.org/10.1055/S-0034-1379045
Kurz T, Pein MK, Marek L, Behrendt CT, Spanier L, Kuna K, Brucher K Microwave-assisted conversion of 4-nitrophenyl esters into 0-protected hydroxamic acids. Eur J Org Chem. 2009; 2939-42. https://doi.org/10.1002/EJOC.200900201
Sharma N, Saha R, Parveen N, Sekar G. Palladium-nanoparticles-catalyzed oxidative annulation of benzamides with alkynes for the synthesis of isoquinolones. Adv Synth Catal. 2017;359:1947–58. https://doi.org/10.1002/ADSC.201601137
Moutanea OG, Mantzourani C, Kokotou MG, Kokotos CG, Kokotos G. Sunlight- or UVA-light-mediated synthesis of hydroxamic acids from carboxylic acids. Eur. J. Org. Chem. 2023;26:e202300046. https://doi.org/10.1002/ejoc.202300046
Paolino M, Brindisi M, Vallone A, Butini S, Campiani G, Nannicini C, Brogi S. Development of potent inhibitors of the mycobacterium tuberculosis virulence factor Zmp1 and evaluation of their effect on mycobacterial survival inside macrophages. ChemMedChem. 2018;13:422–30. https://doi.org/10.1002/CMDC.201700759
Xu X, Feng H, Huang L, Liu X. Direct amidation of carboxylic acids through an active α-acyl enol ester intermediate. J. Org Chem. 2018;83:7962–9. https://doi.org/10.1021/ACS.JOC.8B00819
Schlegel M, Schneider C. Iron(III)-catalyzed (4 + 2)-cycloannulation of 2-hydroxy ketoxime ethers with indol-2-ylamides: synthesis of indole-fused 2-piperidinones. J Org Chem. 2019;84:5886–92. https://doi.org/10.1021/ACS.JOC.9B00261
Acknowledgements
The authors would like to thank the School of Pharmacy, University of Waterloo, NSERC-Discovery (RGPIN: 03830-2014 and 2020-05066), TD Pooler Charitable Foundation 2023-24, Canada Foundation for Innovation, CFI-JELF; Ontario Research Fund (ORF) and Early Researcher Award, Ministry of Research and Innovation, Government of Ontario, Canada (PR) for financial support of this research project.
Author information
Authors and Affiliations
Contributions
PPNR and YZ conceived the project and designed experiments. YZ, AS and AAF performed experiments. PPNR, YZ, AS and AAF analyzed and interpreted results. PPNR and YZ wrote the manuscript. PPNR, YZ, AS and AAF revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, Y., Shakeri, A., Hefny, A.A. et al. N-Benzyl, N-phenethyl and N-benzyloxybenzamide derivatives inhibit amyloid-beta (Aβ42) aggregation and mitigate Aβ42-induced neurotoxicity. Med Chem Res 33, 1229–1241 (2024). https://doi.org/10.1007/s00044-024-03256-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-024-03256-6