Abstract
Cytomegalovirus (CMV) has successfully established a long-lasting latent infection in humans due to its ability to counteract the host antiviral innate immune response. During coevolution with the host, the virus has evolved various evasion techniques to evade the host’s innate immune surveillance. At present, there is still no vaccine available for the prevention and treatment of CMV infection, and the interaction between CMV infection and host antiviral innate immunity is still not well understood. However, ongoing studies will offer new insights into how to treat and prevent CMV infection and its related diseases. Here, we update recent studies on how CMV evades antiviral innate immunity, with a focus on how CMV proteins target and disrupt critical adaptors of antiviral innate immune signaling pathways. This review also discusses some classic intrinsic cellular defences that are crucial to the fight against viral invasion. A comprehensive review of the evasion mechanisms of antiviral innate immunity by CMV will help investigators identify new therapeutic targets and develop vaccines against CMV infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Based on a survey of CMV seropositivity, approximately 83% of the world’s population is infected with the virus. In comparison, in developing countries, it is almost 100% [1]. It will become a lifelong chronic infection in the infected host. Most patients have no symptoms or only show mild disease status, but under immunosuppressive conditions, such as AIDS patients and transplant recipients, the virus can be reactivated, resulting in mental retardation, seizures, and encephalitis [2, 3]. In addition to affecting nervous system diseases, it also affects the occurrence and progression of atherosclerosis, tumors, and other diseases, even endangering life [4,5,6]. As a result, it remains a serious public health issue.
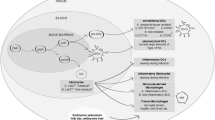
CMV belongs to the β Herpesvirus subfamily and is a double-stranded DNA virus [7]. CMV can replicate in various cells, including epithelial cells, fibroblasts, smooth muscle cells, endothelial cells, macrophages, dendritic cells, and hepatocytes, facilitating systemic transmission, efficient proliferation, and host-to-host transmission of the virus [8, 9]. CMV relies on direct fusion or endocytic pathways to enter human cells [10]. When the viruses invade the host, the viral envelope glycoprotein trimeric complex (gH/gL/gO) binds to platelet-derived growth factor receptor α (PDGFRα), which in turn infects fibroblasts, whereas the pentameric complex (PC) (gH/gL/pUL128-pUL130-pUL131A) is required for the infection of endothelial, epithelial, and medullary-like cells [11]; in parallel, neurofibrillary protein 2, the olfactory receptor family member OR14I1, is thought to be the PC receptor that mediates viral entry into endothelial/epithelial cells [12]. After the viral envelope fuses with the cell membrane, the nucleocapsids are released into the cytoplasm and then transported through the microtubule system to the nucleus, where they release viral DNA [13]. The HCMV genome is expressed in a temporally sequential manner and is categorized into immediate early (IE) genes, early (E) genes, and late (L) genes. IE1 and IE2 are the earliest encoded and most abundantly expressed IE genes that can affect viral promoter activity and initiate transcription from viral E genes [14]. Typical early viral proteins include DNA polymerase (pUL54), phosphotransferase (pUL97), and terminal enzymes (pUL51, pUL52, pUL56, pUL77, pUL89, pUL93, and pUL104), which facilitate viral DNA replication and packaging processes [15]. L genes primarily encode structural proteins required for virion assembly and expulsion that make up the capsid, envelope, or tegument, such as pUL77, pUL93, pUL115, pp28, and pp150 [16]. After DNA replication is complete, it is encapsulated into a capsid, and the nucleocapsid first forms a primary envelope at the inner nuclear membrane, then crosses the nuclear membrane, subsequently unwraps at the outer nuclear membrane, and finally reaches the cytoplasm and coils in the middle compartment of the endoplasmic reticulum (ER)-Golgi apparatus, after which the viral particles are released by budding [17, 18] (Fig. 1).Viral particles can infect various tissues and organs, and in certain cell types, such as hematopoietic stem cells and undifferentiated myeloid cells, the virus can remain silent and latent [19].
CMV life cycle. (1) CMV glycoproteins interact with specific cellular receptors, leading to (2) endocytosis and plasma membrane fusion. (3) The nucleocapsid enters the nucleus through microtubules, and linear DNA is released into the nucleus. (4) (5) Transcription and translation of viral IE genes. (6) (7) Transcription and translation of viral E genes. (8) The viral genome replicates within the nucleus. (9) (10) L proteins, mainly capsid proteins, are expressed and assembled into a new nucleocapsid (11). (12) Membrane proteins and glycoproteins are assembled in the endoplasmic reticulum and Golgi apparatus to form mature viral particles. (14) (15) The virus is released into the extracellular space by budding
The innate immune system is the first line of defence against pathogens invading the body. When viruses enter a host cell, host-pattern recognition receptors (PRRs) detect pathogen-associated molecular patterns (PAMPs), which activate intracellular signal transduction and gene expression programs, leading to the production of a series of mediators, such as inflammatory factors, cytokines, and chemokines, ultimately promoting the body’s natural immune response [20, 21]. However, in the long-term coevolution process, viruses, such as herpes simplex virus-1 (HSV-1), have developed various mechanisms to evade host antiviral innate immunity, including the Toll-like receptor (TLR) signaling pathway, the retinoic acid-induced gene I-like receptor (RLR) signaling pathway, the DNA sensing signaling pathway and intrinsic cellular defences [22,23,24,25,26,27]. To date, CMV has coevolved with mammalian hosts for millions of years; the battle between them has never stopped, and multiple strategies have been developed to evade innate immunity [28].
Among PRRs, the most fully characterized are TLRs [29]. During CMV infection, viral proteins and nucleic acids, including single- or double-stranded RNA (ssRNA or dsRNA), CpG-rich genomes, and envelope glycoproteins, are the true PAMPs recognized by TLRs [30, 31]. The second subfamily comprises RLRs, including retinoic acid-inducible gene 1 (RIG-1), melanoma differentiation-associated gene 5 (MDA5), and laboratory of genetics and physiology 2 (LGP2), which can sense the viral replication intermediate dsRNA and trigger the signal cascade reaction of interferon transcription [32, 33]. The third subfamily is the nucleotide-binding and oligomerization domain (NOD)-like receptor (NLR) family, which includes NOD1, absent in melanoma 2 (AIM2), NLRP1 and NLRP3 [34]. Finally, some DNA sensors, such as stimulators of interferon genes (STING), also recognize CMV and induce an antiviral innate immune response, helping to control early CMV infection and triggering the IFN-I response, thus exerting antiviral effects [35, 36]. Accordingly, it is speculated that CMV has the opportunity to promote autoimmune evasion by blocking the above PRR-related transduction pathways.
When viruses infect the body, host cells are in a state of stress, which triggers a series of stress reactions (autophagy, apoptosis, necrosis, and ER stress), and these stress responses also act as an important part of the innate immune response process [37,38,39]. Therefore, it is not surprising that the cunning virus may also create intracellular environments conducive to its replication and transmission by interfering with these stress pathways [40,41,42].
This review summarizes and updates strategies employed by CMV to evade host innate immune surveillance, including typical PRR-mediated induction of IFN-I and its downstream interferon-stimulated genes (ISGs), as well as intrinsic cellular defences, such as apoptosis, autophagy and ER stress-mediated antiviral innate immunity. A comprehensive evaluation of the battle between host innate antiviral immunity and viral immune evasion is beneficial for familiarizing ourselves with the pathogenesis of the virus and for identifying new drug targets and potential immunotherapies to combat CMV infection-associated diseases.
CMV evades the PRR-mediated antiviral innate immune response
The innate immune system utilizes PRRs to detect the invasion of pathogenic microorganisms, thereby achieving precise recognition and immune response to harmful stimuli [43]. Once PAMPs bind to the corresponding ligands, they activate multiple adapter molecules, ultimately triggering the activation of interferon regulatory factor (IRF) and NF-κB, leading to the expression of proinflammatory cells and chemokines. Meanwhile, IFN-I also induces hundreds of ISGs, thereby endowing the host with strong innate antiviral ability [44, 45]. Nonetheless, viruses evolve various strategies to evade host antiviral innate immunity [46,47,48]. Here, we will summarize how CMV evades PRRs, such as TLRs, RLRs, NLRs, DNA sensors, and their downstream signaling pathways.
TLR signaling pathway
In humans, the TLR family consists of ten members [49]: TLR1, TLR2, TLR4, TLR5, and TLR6 are located on the cell surface, while TLR3, TLR7, TLR8, and TLR9 are located inside the cell [50] and contain a ligand recognition domain rich in leucine repeat sequences, a transmembrane domain, and a Toll/interleukin-1 receptor (TIR) homologous domain [51]. TLRs activate a series of innate immune signaling pathways by recognizing PAMPs, leading to the production of interferons, proinflammatory cytokines, and chemokines, such as IL-1, IL-6, and TNFα, ultimately leading to early identification and defence against foreign pathogens [52,53,54].
The activation status of TLRs in the host plays a crucial role in initial virus replication and persistence [55], and TLRs are involved in almost the entire process of the virus life cycle and are able to recognize virus envelope glycoproteins to promote virus entry into host cells and recognize nucleic acid to induce the production of inflammatory factors [20, 56]. Studies have shown that TLR2 and TLR4 can recognize and participate in the detection of CMV envelope proteins such as glycoproteins gB and gH [57, 58]. Moreover, other studies have suggested that the envelope glycoproteins gB and gH interact with TLR2 and TLR1 and that TLR2/1 heterodimers are functional sensors of HCMV [59, 60]. It is believed that PRRs may initiate the innate immune response before CMV enters cells. In pregnant women with early-onset preeclampsia, the percentage of CMV IgG-positive serum significantly increases, while the expression level of TLR2/4 mRNA is upregulated [61]. During the differentiation of monocytes into macrophages, infection with HCMV promotes the expression of TLR4 and TLR5 to enhance macrophage inflammation significantly [62]. In addition, TLR3 and 9 located within cells can induce the production of IFN-I and proinflammatory cytokines by recognizing dsRNA and CpG-rich genomes [63].
Single nucleotide polymorphisms (SNPs) of TLRs can also affect the immunogenicity and disease status of CMV infection. A study suggested that TLR9 SNPs are associated with CMV reactivation and disease and that genetic polymorphisms may downregulate TLR9 signaling and lead to reduced antiviral response efficiency [64]. TLR2 and TLR7 genetic polymorphisms are also associated with CMV status in late pregnancy in women and regulate the immune response to CMV [65]. However, some studies have found no statistically significant association between SNPs in the TLR gene and congenital CMV infection or disease status [66]. Therefore, more studies are needed to determine the relationship between TLR SNPs and CMV infection.
To date, various viruses, such as HSV and hepatitis B virus, have evolved specific proteins targeting TLRs that interfere with the innate immune response [56, 67]. To respond to immune system attacks, CMV has also developed various evasion mechanisms (Fig. 2). US7 utilizes the ER-related degradation components Derlin-1 and Sect. 61 to promote the ubiquitination of TLR3 and TLR4. US8 not only disrupts the binding of TLR3-UNC93B1 but also targets TLR4 to lysosomes, leading to its rapid degradation. Therefore, US7 and US8 are considered key inhibitors of TLR3 and TLR4 in human foreskin fibroblasts (HFFs) [68]. HCMV encodes miR-UL112-3p, which effectively inhibits endogenous TLR2 protein levels during CMV infection of normal human dermal fibroblasts (NHDFs), thereby significantly downregulating the TLR/IRAK1/NF-κB signaling pathway [69]. Currently, few studies investigated the correlation between CMV and TLRs in terms of immune evasion, and there is still a significant gap in knowledge of the mechanism by which viral protein targeting blocks the antiviral signaling pathway of TLRs. More studies on the immune evasion mechanism by which CMV targets TLRs in the future may provide new insights for disease prevention and control.
The immune evasion of the TLR signaling pathway by CMV. virus-encoded products antagonizes the TLR signaling pathway to promote immune evasion. US7 promotes the ubiquitination of TLR3 and TLR4, and US8 promotes their degradation. miR-UL112-3p decreases endogenous TLR2 protein levels. CMV proteins can also interfere with multiple other steps in the TLR pathway, such as TBKI, IRF3, IKK, p65, and p50, to promote immune evasion
RLR signaling pathway
The three members of the RLR family are highly homologous, with the same DExD/H-box RNA helicase domain at the center and the same zinc-binding domain at the C-terminus [70]. However, LGP2 lacks a signal domain (CRAD) and needs to interact with MDA5 to play a role in viral sensing [71]. After RLRs recognize and bind to the corresponding PAMPs, their conformation changes and a pair of CRADs are released, recruiting and activating downstream mitochondrial antiviral signaling proteins (MAVSs) containing CRADs. Subsequently, the downstream protein TANK binding kinase 1/IkappaB kinase (TBK1/IKK) is recruited to activate the transcription factors IRF3/7 and nuclear factor κB (NF-κB), which produce IFN and proinflammatory cytokines [72, 73].
Undoubtedly, RLRs have become key cytoplasmic receptors for detecting RNA viruses [74]. However, it has been shown that both viral and host RNA can also act as RLR ligands during DNA virus infection [75]. In 2017, for the first time, the immunoreactivity of RIG-I, MDA5, and LGP2 in placental, chorionic, and amniotic tissues was reported and might be related to CMV sensing in pregnancy-related tissues [76]. When the placenta was infected with CMV, the secretion of IFN-β increased, further enhancing the expression of RIG-I and MDA5 at the mRNA level, and it also upregulated the expression of DDX58 and IFIH1. This study suggested that RLRs may be involved in CMV sensing, but more studies are needed [77].
NLR signaling pathway
The NLR family consists of more than 20 members, among which NLRP3 is one of the most studied DNA sensors related to inflammasomes. Activation of the NLRP3 inflammasome by PAMPs and damage-associated molecular patterns (DAMPs) leads to the secretion of the proinflammatory cytokines IL-1β and IL-18, which play crucial roles in the antiviral innate immune response [78]. Available evidence suggests that the viral glycoprotein components of CMV, as well as increased levels of mtDNA released into the cytoplasm during CMV infection, can activate the NLRP3 inflammasome [34, 79]. NLRC5 is also upregulated in human fibroblasts after CMV infection and activates IFN-γ-mediated antiviral signaling pathways [80]. In addition, activating NOD1 also induces an IFN response and is suppressed by HCMV [81]. Due to their role in the antiviral response, these inflammasomes seem to play a protective role, but the inflammatory response is also involved in the occurrence and development of many diseases. Inhibiting the inflammatory response is a key mechanism for treating certain diseases, so it may also cause serious pathological damage to the body. For example, MCMV infection can cause neuronal death and hearing loss by activating the NLRP3 inflammasome [82]. Downregulation of NLRP3 is beneficial for hypertensive vascular and myocardial remodeling [83]. Therefore, during infection, the roles of these inflammasomes are diverse and worthy of further investigation.
AIM2 can also detect the genetic material of CMV entering the cytoplasm to promote the activation of inflammatory factors [84]. Compared to those in wild-type cells, the ability of AIM2-deficient macrophages to induce IL-1β expression and cell death is reduced, and the transcription of the viral DNA polymerase gene UL54 and the capsid protein-encoding gene UL83 is greater, which indicates that AIM2-deficient cells cannot effectively control HCMV infection [85]. In a CMV-induced sensorineural hearing loss (SNHL) mouse model, a significant increase in AIM2 levels was detected, and the expression of IL-1, IL-18, IL-6, and TNF-α increased to resist CMV infection [86]. Interestingly, the expression of AIM2 is time-dependent, with an increase in AIM2 levels in the early stages of HCMV infection and a decrease in AIM2 levels 24 h after infection. This decrease may be related to the viral immune evasion mechanism [87]. pUL83 interacts with AIM2 in the cytoplasm during the early stages of HCMV infection, thereby interfering with the activation of AIM2 [87]. Subsequently, the immediate early 86 kDa protein (IE86) of HCMV was shown to block the transcription of immature interleukin-1β and induce its degradation during the activation of AIM2 [88].
DNA-sensing signaling pathway
TLR9 was the first DNA sensor described in 2000 [89]. Since then, an increasing number of DNA sensors, including cGAS, STING, DHX9, DHX36, DDX41, IFI16, and ZBP1, have been identified [90, 91]. Among various cell types, cGAS has been identified as the only universal cytoplasmic DNA sensor [92]. When PAMPs recognize and activate cGAS, their conformation changes, which in turn catalyzes the synthesis of cGAMP and activates STING. Subsequently, STING in the ER rapidly dimerizes and undergoes typical COPII-dependent anterograde transport, passing through the ER Golgi intermediate compartment and transferring to the Golgi apparatus. STING located in the Golgi apparatus undergoes palmitoylation and recruits TBK1 for phosphorylation. Then, STING recruits and phosphorylates IRF3, which dimerizes and translocates to the nucleus to activate IFN-I, TBK1 and its homolog IKK β, which enter the nucleus to activate NF-κB signal transduction, inducing the generation of inflammatory factors and participating in antiviral immune responses [93,94,95,96].
During HCMV infection, the nucleocapsid is first delivered to the host cell through membrane fusion. Then, the nucleocapsid is transported to the nuclear pore through the cytoskeleton system, releasing viral genomes into the nucleus in a pressure-dependent manner [97,98,99]. A portion of the viral capsid is degraded by the ubiquitin-proteasome pathway, which then releases the viral DNA into the cytoplasm, where it is detected by a cytoplasmic DNA sensor [100]. The innate immune pathway mediated by DNA sensing has a vital impact on limiting CMV infection. Many viral proteins also counteract immune evasion from the host at the cGAS, STING, MTITA, and IFI16 levels and their downstream signaling adaptors, such as the TBK1-IRF3 axis and NF-κB (Fig. 3).
The immune evasion of the DNA-sensing signaling pathway by CMV. CMV proteins can target multiple adapters and transcription factors in the DNA-sensing signaling pathway for immune evasion. UL31, UL42, and PP65 act on cGAS to inhibit cGAS activity, cGAMP synthesis or interaction with STING. UL138 and UL122 degrade STING, and pUL48 inhibits the ubiquitination of STING. UL97 binds to IFI16, causing it to detach from the nucleus. m152, UL82, UL94 and UL42 inhibit the translocation of STING from the ER to the Golgi. USP13, UL37 × 1, UL36, US9, UL82 and UL35 inhibit the interaction between IRF3, TBK1 and the STING complex. UL48, M48, and M45 inhibit the ubiquitination of RIP1. UL44 binds the p65 and p50 complex, US28 inhibits the nuclear localization of p65, and UL26 prevents the degradation of IKB. UL35, UL36, UL138 and PP65 inhibit the phosphorylation of IRF3. UL44 and M35 inhibit the binding of IRF3 to the promoters of antiviral genes
cGAS
cGAS was first discovered in 2013 as a dsDNA sensor that can activate the second messenger cGAMP, thereby activating STING to form the main pathway for perceiving nonself DNA in animals and triggering effective immune responses [25]. However, CMV likely utilizes multiple strategies to inhibit cGAS enzyme activity and interfere with host immune function. A previous study proposed that the HCMV protein pUL31 can directly interact with cGAS to inhibit the enzyme activity of cGAS, prevent DNA from cGAS sensing, and reduce the production of cGAMP and IFN-I and downstream antiviral genes [101]. The IFN levels produced by primary HFFs infected with mutant strains that cannot express pp65 were significantly greater than those infected with wild-type strains, indicating that pp65 may be a viral evasion protein and selectively binds to cGAS, which competitively inhibits the interaction of cGAS with STING, thereby blocking the signal transmission of the cGAS/STING/IRF3 axis [102]. In addition, another study has revealed the interaction between UL42 and cGAS through immunoprecipitation technology and further revealed that UL42 reduces the synthesis of cGAMP by inhibiting DNA binding and oligomerization of cGAS in HFFs [103].
STING
After cGAS senses dsDNA and synthesizes its second messenger, cGAMP binds to STING and initiates signal transduction. During this process, the activity of STING is regulated by its dimerization, translocation from the ER through the Golgi apparatus to the perinuclear site, and posttranslational modifications such as phosphorylation or ubiquitination [104, 105]. Investigators have shown that UL138 or UL122 could induce proteasome-mediated degradation of STING through lysosomal and proteasome-dependent pathways [106, 107]. Another study further confirmed that amino acids (aa) 136–289 in IE86 are responsible for STING degradation [108]. Additionally, UL138 colocalizes and interacts with STING and TBK1 to inhibit cGAS/STING/TBK1 signaling pathway-induced IRF3 phosphorylation and IFN-β promoter activation [106]. The UL82 gene encodes the capsid protein (pp71), and mutated viruses lacking UL82 have a reduced number of viral production after HDF infection, indicating that UL82 is crucial for HCMV replication [109]. UL82 and UL42 disrupt the process of STING translocation from the ER, resulting in the blockade of STING activation [103, 110]; in parallel, UL94 also disrupts this process by inhibiting the dimerization of STING when HCMV infects HFFs [111]. HCMV pUL48 encodes a deubiquitinating enzyme and downregulates the ubiquitination of STING, TRAF6, TRAF3, IRAK1, and IRF7 to counteract the PRR-mediated IFN-I antiviral response in human diploid fibroblasts [112]. Another study has revealed that the MCMV protein m152 delays its transport to the Golgi compartment by binding to STING. Interestingly, m152 inhibits STING-mediated IRF signaling, but not STING-mediated NF-κB signal transduction [113]. In 2018, a study revealed that glycoprotein US9 blocks STING dimerization by binding to STING and TBK1, thereby inhibiting the interaction between STING and TBK1 [114].
IFI16
The IFI16 protein is a member of the pyrin and hin domain (PYHIN) family, which consists of one PYRIN domain and two HIN domains composed of 200 amino acids; this protein has been proven to bind to intracellular viral DNA and trigger the expression of antiviral cytokines through the STING/TBK1/IRF3 signaling pathway [115]. IFI16 binds to viral DNA in the early stages of HCMV infection in human embryonic lung fibroblasts, whereas in the late stages, IFI16 binds to pUL97, which contributes to the translocation of IFI16 from the nucleus into the cytoplasm and the integration of pUL97 into newly assembled viral particles, thereby effectively expelling IFI16 from the infected host to hijack its antiviral activity [116]. Thus, the mislocalization of IFI16 to the cytoplasm by viral proteins is critical for CMV to evade IFI16’s antiviral activity [117].
TBK1-IRF3
cGAMP binds to STING to induce STING protein transformation. Subsequently, TBK1 and IRF3 are recruited to the STING complex, where TBK1 is a serine/threonine protein kinase that phosphorylates and activates IRF3, which then translocates to the nucleus and initiates type I IFN production [118]. Given that the TBK1-IRF3 axis is at the core of the antiviral innate immune response, CMV will also evolve various mechanisms to block the TBK1-IRF3 axis signal transduction.
CMV-encoded UL37 exon-1 and UL82 bind directly to TBK1 and inhibit the recruitment of TBKI and IRF3 to the STING complex to eliminate TBK1-STING-IRF3 interactions [110, 119]. One study used WT or UL35-deficient recombinant HCMV to infect primary human foreskin fibroblasts (HFF-1) and revealed that UL35 reduced the phosphorylation of TBK1, STING, IRF3, and STAT1 [120]. USP13 can uncouple multiubiquitin chains connected to K27, thereby reducing the recruitment of TBK1 and its downstream signaling [121]. These findings emphasize the importance of viral protein to evade innate immune sensor TBKI.
UL44 inhibits the binding of IRF3 to downstream antiviral gene promoters, and its overexpression also inhibits the activation of the interferon-stimulated response element (ISRE) induced by IRF3 in HFFs [122]. Investigators have shown that UL36 and UL138 can directly interact on TBK1 or IRF3 to inhibit the phosphorylation of IRF3 induced by the cGAS/STING/TBK1 pathway when HCMV is transfected into 293T cells or HFFs [73, 106]. pp65 is distributed between the nucleus and the cytoplasm and is important for producing viral particles and inducing dense bodies [123]. This protein regulates the IFN-I response by inhibiting the accumulation of IRF-3 in the nucleus and reducing its phosphorylation [124]. The reduction of antiviral activity by M35 during MCMV infection is dependent on its dimerization, where its homodimer binds directly to specific DNA sequences in the Ifnb1 promoter, thereby antagonizing the recruitment of IRF3 to the promoter of antiviral genes [125].
NF-κB
NF-κB plays a crucial regulatory role in the innate immune response and includes a series of important nuclear transcription factors. The NF-κB family in mammals consists of five members: p65/RelA, RelB, p105/p50/NF-κB1, p100/p52/NF-κB2, and c-Rel [126]. Although the p50, p52, and Rel proteins can form multiple homologs and heterodimers, the main form of NF-κB is the heterodimer of p65 and p50 [127]. In a resting state, NF-κB in the cytoplasm combines with the inhibitory factor IκB to form the trimeric p50-p65-IκB. When IκB kinase (IKK) is activated, it catalyzes the phosphorylation of two conserved serine residues of IκB and degrades them through the ubiquitin-proteasome pathway connected by K48, and activated NF-κB translocates into the nucleus and binds to its corresponding DNA sequence to induce target gene transcription, leading to the production of IFN and inflammatory factors [128,129,130].
The results revealed that both HCMV and MCMV regulate NF-κB activity in a time-dependent manner, activating NF-κB in the first few hours of infection but inhibiting its activation afterward, and the potential molecular mechanisms are yet to be determined [131]. In vitro cell experiments confirmed the interaction between the HCMV proteins pUL44 and p65 or p50, thereby inhibiting the activation of NF-κB induced by IKKβ and p65 overexpression, as well as inhibiting the binding of p65 to the promoter of CXCL2 [122]. HCMV UL26 blocks the nuclear translocation of NF-κB, phosphorylation of the IKK complex and degradation of IκB in MRC5 fibroblasts [132]. US31 may act as an E3-like ubiquitin ligase that promotes the ubiquitination of phosphorylated p100, thereby activating the NF-κB2 pathway [133]. Compared to infection with the mutant virus lacking pp65, infection with the wild-type virus downregulates IL-1β levels by inhibiting NF-κB activity [134]. HCMV US28 expression in undifferentiated monocytic THP-1 cells inhibits the nuclear translocalization of p65, attenuating the NF-κB pathway to facilitate viral entry into latency [135]. Later, it was revealed that MCMV M35 shuttles to the nucleus before p65 translocation and inhibits type I IFN transcription regulated by NF-κB [136].
NF-κB activation requires the ubiquitination of receptor-interacting protein 1 (RIP1) and NF-κB essential modulator (NEMO) [137]. Moreover, the M45 protein also targets NEMO to autophagosomes for degradation in lysosomes in mouse embryonic fibroblasts infected with MCMV [138]. M45 also blocks the ubiquitination of RIP1, thereby counteracting the innate immune signaling pathway [139]. CMV-encoded UL48 and M48 interact with RIP1 and rely on its deubiquitinating enzyme activity to cleave K48- and K63-linked polyubiquitin chains to inhibit NF-κB activation [140, 141].
The IFNAR-JAK-STAT signaling pathway and its downstream ISGs
IFNAR-JAK-STAT signaling pathway
IFN occupies an absolute core position in antiviral innate immunity, and IFN-I binds to IFNAR1 and IFNAR2 to promote downstream signaling cascades [142]. When IFN binds to IFNAR1 and IFNAR2, it recruits and phosphorylates tyrosine kinase 2 (TYK2) and JAK1, followed by further phosphorylation and activation of STAT1/STAT2 [143, 144]. Activated STAT1 and STAT2 form heterodimers and bind to IRF9 in the cytoplasm, forming heterotrimeric complexes of IFN-stimulating gene factor 3 (ISGF3) [145]. Furthermore, the complex undergoes nuclear translocation and binds to the ISG promoter to initiate the expression of ISGs [146].
In patients with chronic HCV infection, by comparing the transcriptional levels of JAK-STAT pathway mediators between the CMV-positive and CMV-negative groups, it was found that the expression of STAT2 and IRF7 in the CMV-positive group was downregulated, and it was speculated that the elevated risk of CMV in liver cirrhosis patients after HCV infection may be related to dysfunction of the JAK-STAT pathway, which weakens the antiviral effect [147].
Viral proteins reducing ISG expression and evading innate immune system attacks by manipulating the IFNAR-JAK-STAT signaling pathway are shown in Fig. 4. Promoting the degradation of STAT1/STAT2 and interfering with the tyrosine phosphorylation of STAT1/STAT2 are two major evasion mechanisms [148]. The replication of the M27 deletion virus is highly reduced, and its virulence is weakened in mice, indicating that pM27 is a key protein affecting CMV replication and virulence [149]. pM27 binds and downregulates STAT2 in mouse embryonic fibroblasts, while IFN-γ induces an increase in STAT2 tyrosine phosphorylation in M27(-) MCMV-infected cells, which means that pM27 expression also downregulates STAT2 tyrosine phosphorylation [150]. Moreover, pM27 recruits DDB1-containing ubiquitin ligase complexes and directly induces the ubiquitination and degradation of STAT2 through the ubiquitin-proteasome pathway [151]. Similar to pM27, both isoforms of the pUL145 protein (pUL145-Long, pUL145-Short) were found to recruit DDB1-containing ubiquitin ligases to ubiquitinate and degrade STAT2 in HCMV-infected U251 cells [152]. There is conclusive evidence that HCMV UL23 directly interacts with human N-myc-interacting factor (Nmi) or blocks STAT1 phosphorylation in U251 cells to inhibit the nuclear translocation and expression of STAT1 and attenuate the antiviral effects of IFN [153, 154]. UL26 was recently found to form complexes with several STAT family members and various PIAS proteins (repressor proteins that activate STAT transcriptional activity) to block STAT phosphorylation in MRC5 cells infected with HCMV [155]. CMV encodes miR-US33as-5p, which inhibits the phosphorylation of STAT1, STAT2, Tyk2, and Jak1 by binding to the 3′-UTR of IFNAR1, resulting in inactivation of the JAK-STAT signaling pathway and decreasing the expression of ISGs (Mx1, RSAD2, DDX58, BST2, IFIT2, and ISG20) [156].
The immune evasion of the IFNAR-JAK-STAT signaling pathway by CMV. CMV has evolved multiple tricks to evade immune killing via the IFNAR-JAK-STAT signaling pathway. pM27 and pUL145 induce the ubiquitination and degradation of STAT2. UL23 and UL26 block the phosphorylation of STAT1. MiR-US33as-5p inhibits the phosphorylation of STAT1, STAT2, Tyk2, and Jak1. IE1 and UL26 downregulate the levels of ISG15. IE1 induces the citrullination of IFIT1, IFIT2 and IFIT3. TRS1, IRS1, m142, m143, and US22 blocks PKR to exert antiviral effects, and IE1 induces TRIM desumoylation
ISGs
ISGs can be induced in an interferon-dependent or interferon-independent manner to limit viral infection and replication after CMV infection of HFFs, and as a result, elevated expression of IFIT1, IFIT2, IFIT3, Mx1, Mx2, CXCL10, and ISG15 has been detected [157, 158]. In response to these viral defences, CMV has also evolved a series of immune evasion strategies against ISGs.
ISG15
ISG15 belongs to the ubiquitin-like protein family, and its protein modification (ISGylation) is closely related to innate immunity, which can inhibit viral replication [159]. Viral dsDNA could bind to specific receptors to increase ISG15 protein. When HCMV infects primary human fibroblasts, IE1 downregulates ISG15 by inhibiting dsDNA-induced ISGylation [160]. The expression of HCMV UL26 restricts the expression of ISG15 and protein ISGylation in MRC5 fibroblasts [161]. Other studies have shown that UL26 interacts with ISG15 and evades ISG15-related antiviral immune responses by inhibiting ISGylation [162]. Additionally, pUL50 could inhibit protein ISGylation by proteasomal degradation of UBE1L, an E1-activating enzyme for ISGylation [163].
ZAP
Zinc-finger antiviral protein (ZAP) is an effective antiviral protein. The expression of ZAP may disrupt the transcription of UL4-UL6, reducing viral mRNA and protein [164]. Moreover, ZAP’s antiviral activity exhibits HCMV strain-dependent differences [165]. However, one study revealed that HCMV can evade the antiviral activity of ZAP by inhibiting CpG dinucleotides in the IE1 gene [166].
IFIT1 and ITIT3
Investigators discovered that virally infected astrocytes markedly reduce viral titers in IFITI-overexpressing cells and increase viral titers in IFITI-knockdown cells, suggesting that IFIT1 has crucial antiviral functions [167]. IFIT3 knockout mice showed increased susceptibility to CMV infection, further indicating that IFIT3 plays an important antiviral role during CMV infection [168]. Upon HCMV infection of primary human fibroblasts, IE1 induces the expression of the protein arginine deiminase gene (PAD), which contributes to the significant citrullination of certain ISGs, such as IFIT1, IFIT2, IFIT3, and Mx1, to evade host immune defences [169].
Viperin
Viperin is an antiviral protein that can inhibit the replication of various DNA and RNA viruses. HCMV infection directly induces the expression of Viperin [170]. Viperin is transmitted to the ER, mitochondria, and viral assembly chambers and exhibits antiviral activity when located in the ER, which enhances TLR7- and TLR9-mediated IFN production to combat CMV infection [171, 172]. However, another study found that HCMV infection does not interfere with viperin mRNA and protein and viperin promotes HCMV infection in human trophoblast cells [173]. These contradictory results indicate that viperin plays an extremely complicated role in CMV infection, and more studies are needed to investigate the potential underlying mechanisms.
PKR
PKR first binds to CMV dsRNA, which then dimerizes and activates autophosphorylation. Activated PKR, in turn, inhibits mRNA translation by phosphorylating its substrate eukaryotic initiation factor 2α (eIF2α), thereby hindering virus translation initiation and replication [174]. Studies have shown that viral proteins TRS1 and IRS1 can bind to both dsRNA and PKR simultaneously, blocking PKR kinase activity and inhibiting eIF2α phosphorylation by binding to conserved amino acids in the PKR-eIF2α binding site [175,176,177,178]. In mouse embryonic fibroblasts, MCMV m142 and m143 also effectively selectively antagonize PKR by binding to dsRNA, blocking the phosphorylation of PKR and eIF2α, and relocating PKR to the nucleus and insoluble cytoplasmic compartments [179, 180]. In addition, some investigators have attempted to identify a PKR antagonist encoded by GPCMV and ultimately found that gp145, a member of the US22 protein family, can target dsRNA to block the PKR pathway [181].
TRIMs
The TRIM protein family is a large class of E3 ubiquitin ligases that primarily play an antiviral role by promoting the ubiquitination of key protein molecules [182]. Promyelocytic leukemia-nuclear body (PML-NB), also known as TRIM19-NB, is a key mediator of intrinsic immunity against pathogens, and HCMV IE1 is composed of a spherical core (IE1CORE) region and an inherently disordered region on both sides. Studies have shown that IE1CORE effectively binds to the coiled-coil domain of TRIM19, selectively preventing SUMOylation at specific lysine residues, such as K160, thus inducing deSUMOylation of TRIM to counteract the intrinsic immunity of TRIM-NB [183, 184]. Interestingly, the core area of IE1 is integrated not only with TRIM19 but also with TRIM5α [184]. In 2013, a study identified cellular proteins that interact with IE1 and revealed that IE1 also interacts with TRIM33 [185]. Therefore, given the interaction between IE1 and TRIM5α or TRIM33, IE1 might also antagonize the antiviral effects of TRIM5α and TRIM33.
CMV evades apoptosis/necrosis, autophagy, and ER stress
When CMV infects the host, viral proteins or nucleic acids effectively induce a series of cellular stress responses, including apoptosis, necrosis, autophagy, and ER stress, to kill and clear the virus and maintain cellular homeostasis (Fig. 5).
The immune evasion of ER stress, autophagy, and cell apoptosis by CMV. CMV counteracts stress responses such as ER stress, apoptosis and autophagy to increase survival. Inhibition of apoptosis/necrosis: UL36 degrades MLKL; M36 inhibits CASP8, CASP3, and UL37 × 1 binding and inhibits Bak and Bax; UL141 binds to TRAIL death receptors; M45 inhibits the RIPK3-dependent necrosis pathway; and IE1 and IE2 inhibit TNF-α-induced apoptosis. IE1 induces the ubiquitination of RIPK3, and US21 regulates the intracellular Ca2+ balance. Inhibition of autophagy: TRS1 and IRS1 bind to BECN1 to inhibit autophagy. US12 upregulates ULK1 phosphorylation and LC3-II conversion. Inhibition of ER stress: UL38 induces the accumulation of ATF4 and limits the phosphorylation of JNK. UL148 retains CD58 in the ER, and UL50 and M50 significantly decrease the IRE1 protein level
Apoptosis/Necrosis
Apoptosis is usually considered an innate cellular response to invading pathogens that can clear cells infected by pathogens and limit and resist pathogen invasion [186]. Apoptosis can be initiated through internal or external pathways, which include three main pathways: the mitochondrial pathway, cell death receptor pathways (such as the Fas/FasL pathway), and the ER stress pathway [187]. When exogenous and endogenous cell apoptosis pathways converge, they cleave and activate Caspase family proteases such as Caspase 3, Caspase 8, and Caspase 9, subsequently activating or inhibiting substrates that promote morphological changes related to cell apoptosis, such as chromatin concentration, cell contraction, and nuclear and plasma membrane foaming [96, 188]. In addition to cell apoptosis, necrosis, characterized by swelling of cytoplasmic organelles, moderate chromatin condensation, loss of membrane integrity, and leakage of cellular components into intercellular spaces, has also received increasing attention [189]. Necrosis is also considered a strategy for host cells to resist the invasion of pathogenic microorganisms such as viruses and bacteria [190].
Host cells can directly or indirectly activate multiple cell death pathways by detecting CMV components or by activating stress pathways through viral replication [191], but CMV also encodes various inhibitors of cell apoptosis and necrosis to ensure the persistence of infection. The interfereing function of UL36 in apoptosis or necrosis has been fully studied. Notably, UL36 or M36 is a multifunctional inhibitor of apoptosis and necrosis and promotes viral growth, and the growth of MCMV deficient in M36 is severely attenuated both in macrophages and in vivo [192]. UL36 targets the degradation of mixed lineage kinase domain-like (MLKL), a key protein involved in cell necrosis, and inhibits the apoptotic program in HCMV-infected primary human fetal foreskin fibroblasts [193]. However, UL36 does not inhibit apoptosis in mouse cells, suggesting that the inhibition of apoptosis by this protein is species-specific [194]. Consistent with these results, MCMV M36 encodes an apoptosis inhibitor that binds to caspase-8 and blocks the death receptor pathway in macrophages [195]. Bak and Bax are core regulatory factors for the intrinsic pathways of cell apoptosis [196]. The protein product of HCMV UL37 × 1 can target mitochondria and inhibit cell death by binding and inhibiting Bak and Bax [197]. HCMV UL141 encodes a glycoprotein that can bind to proteins such as TRAIL death receptors, downregulate the cell surface expression of members of the tumor necrosis factor receptor superfamily (TNFRSF), and thus resist host defence [198].
Receptor-interacting protein kinase-1 and − 3 (RIPK1 and RIPK3) are key mediators of necrosis and strictly regulate the process of necrosis. The MCMV protein M45 contains the RIP homologous interaction motif (RHIM), which is sufficient to protect infected cells from necrotic cell death [199]. Recently, M45 was shown to inhibit the RIPK3-dependent necrosis pathway through Z-DNA binding protein 1 (ZBP1)-dependent (in non-myeloid cells) and ZBP1-independent (in macrophages) pathways. M36 suppresses the TNF signaling-dependent CASP8-CASP3 activation axis, as well as CASP8-independent cleavage activation of CASP9 and CASP3 [200]. Therefore, MCMV proteins ensure their adaptability by blocking different host proteins involved in programmed cell death pathways. IE1 and IE2 are early characterized antiapoptotic proteins that are crucial for regulating host cell death to ensure HCMV replication [201], and the relevant mechanisms include inhibiting TNF-α-induced cell apoptosis and inhibiting cell apoptosis by activating phosphatidylinositol kinase (PI3K) to promote survival pathways [202]. A recent study demonstrated that IE1 interacts with RIPK3 in an RHIM domain-independent manner in HFFs, resulting in substantial upregulation of posttranslational modifications of RIPK3, particularly ubiquitination, to prevent apoptosis [203]. The US21 protein is an ion channel encoded by viruses that regulates intracellular Ca2+ homeostasis and protects cells from apoptosis [204].
Autophagy
Viral infections usually result in autophagy, which is an inherent cellular defence mechanism [40]. Autophagy typically controls infection by delivering viral particles to lysosomes for degradation and further coupling with PRR signaling pathways to induce interferon-mediated viral clearance [205, 206]. Currently, little is known about the interaction between CMV and autophagy. It is generally believed that inducing autophagy can damage viral DNA replication and interfere with viral particle release through different mechanisms [207]. Some autophagy inducers, such as trehalose and SMER28, can induce autophagy to reduce the accumulation of viral proteins in HFFs and HAECs, interfere with viral particle transport pathways, redirect viral particles to acidification compartments for degradation, and delay viral genome replication [208, 209]. However, knocking out autophagy-related genes and inhibiting the activity of autophagy-related proteins can downregulate the replication of HCMV, indicating that autophagy-related factors are required for effective HCMV replication and induced autophagy will enhance viral replication [210, 211]. The sharp contrast in autophagy may be due to the fine requirement and balance of autophagy during its replication cycle.
Not surprisingly, various strategies have been developed to block the autophagy process in CMV. The binding of the viral protein TRS1 to BECN1 could inhibit autophagy [212], which could be completely inhibited 24 h after HCMV infection in human primary fibroblasts. IRS1, homologous to TRS1, can also bind to BECN1 and inhibit autophagy. When these two proteins are expressed simultaneously, they hinder the development of autophagosomes, but when they are expressed separately, they block the onset of autophagy [211]. US12 interacts with regulatory proteins necessary for inducing autophagy, thereby upregulating the phosphorylation of the autophagy initiation protein ULK1 and the transformation of LC3-II to induce and accelerate autophagic flux [213]. However, additional studies are needed to determine whether US12 increases viral replication through autophagy.
ER stress
The ER is a crucial organelle involved in physiological processes such as protein processing and the regulation of Ca2+ homeostasis [214]. The synthesis and folding of proteins are precisely regulated and sensitive to changes in ER homeostasis. When the ER is subjected to stress, such as Ca2+ depletion, hypoxia, and viral infections, it produces a large amount of unfolded proteins [215]. The ER enters a stress state once its folding capacity is exceeded or when there is an excessive accumulation of misfolded proteins. These proteins are generally detected by three ER-resident sensor molecules, activating transcription factor 6 (ATF6), inositol-requiring enzyme 1α (IRE1α) and protein kinase R-like ER kinase (PERK), which further activate the unfolded protein response (UPR). Moreover, the phosphorylation of the PERK translation initiation factor eIF2α reduces protein translation [216]. Eventually, ER homeostasis is restored by increasing protein folding ability and reducing protein load.
Some studies have shown that HCMV can manipulate the UPR to promote infection. HCMV with a mutated UL38 has severe growth defects, but its growth is restored in HF cells expressing wild-type pUL38, possibly because pUL38 protects cells from ER stress during HCMV infection by inducing the accumulation of activated transcription factor 4 (ATF4) and limiting c-Jun N-terminal kinase (JNK) phosphorylation [217]. HCMV UL148 retains CD58 within the ER, preventing it from reaching the cell surface to weaken the CD58-mediated antiviral response. However, UL148 also promotes the activation of PERK and IER1 and reshapes the ER, but these two effects can be separated as the virus mutates and evolves [218, 219].
RE1/X box binding protein 1 (XBP1) is an important component of unfolded protein response elements. The expression of M50 in MCMV-infected NIH-3T3 cells leads to a significant decrease of IRE1 protein, while mRNA splicing and the expression of XBP1 are also inhibited in infected cells. Similarly, the homolog of M50, HCMV UL50, also degrades IRE1 [220]. Another study further revealed that the M50 protein inhibits the IRE1 branch of the UPR by associating IRE1 with SEL1L, thereby promoting its degradation via the ER-related degradation (ERAD) mechanism [221].
Conclusion
CMV is ubiquitous and can establish lifelong infection in the host. It can cause serious complications in individuals with a suppressed or dampened immune system and is also the most common cause of congenital viral infections. CMV infection can trigger a wide range of defence mechanisms, among which the innate immune defence mechanism is the first line of defence against viral invasion, mainly limiting virus infection and transmission by producing IFN-I and ISGs. In the long-term struggle between the virus and the host, CMV has developed various strategies to evade innate immune attacks and facilitate its growth and reproduction. Multiple viral proteins are involved, especially some envelope proteins, IE proteins, and other functional proteins that target key adapters to antagonize antiviral signaling pathways, such as TLR2/4, cGAS, STING, NF-KB, TBKI, and IRF3. Moreover, some viral proteins can also interfere with intracellular processes, such as apoptosis, necrosis, autophagy, and ER stress pathways, to evade the innate immune system and result in persistent infections.
Accumulating evidence suggests that TLR agonists have great potential as antibacterial, anticancer, and vaccine adjuvants and are also of great significance for vaccine development and immunotherapy against HSV, HBV, HIV, and other viruses [222,223,224]. Moreover, TLR antagonists have also been developed as reagents and drugs to suppress immune responses [225]. Therefore, this article reviews the immune evasion mechanism by CMV to target PRRs and various cellular processes, which may also provide great insights into developing vaccines and antiviral drugs.
Data availability
Not applicable.
References
Adland E et al (2015) Ongoing burden of disease and mortality from HIV/CMV coinfection in Africa in the antiretroviral therapy era. Front Microbiol 6:1016. https://doi.org/10.3389/fmicb.2015.01016
Nishida K et al (2020) Prediction of neurodevelopmental impairment in congenital cytomegalovirus infection by early postnatal magnetic resonance imaging. Neonatology 117(4):460–466. https://doi.org/10.1159/000508218
Marais B et al (2023) Cytomegalovirus haemorrhagic cystitis in a pregnant patient with AIDS. Ther Adv Urol 15:17562872231159531. https://doi.org/10.1177/17562872231159531
Fonseca Brito L et al (2019) Cytomegalovirus (CMV) pneumonitis: cell tropism, inflammation, and immunity. Int J Mol Sci 20(16). https://doi.org/10.3390/ijms20163865
Bonavita CM, Cardin RD (2021) Don’t Go Breaking My Heart: MCMV as a Model for HCMV-Associated Cardiovascular Diseases. Pathogens 10(5). https://doi.org/10.3390/pathogens10050619
Branch KM et al (2021) Productive infection of human breast Cancer cell lines with human cytomegalovirus (HCMV). Pathogens 10(6). https://doi.org/10.3390/pathogens10060641
Nukui M et al (2018) The natural flavonoid compound deguelin inhibits HCMV Lytic replication within fibroblasts. Viruses 10(11). https://doi.org/10.3390/v10110614
Jean Beltran PM, Cristea IM (2014) The life cycle and pathogenesis of human cytomegalovirus infection: lessons from proteomics. Expert Rev Proteomics 11(6):697–711. https://doi.org/10.1586/14789450.2014.971116
Krstanović F et al (2021) Cytomegalovirus infection and inflammation in developing brain. Viruses 13(6). https://doi.org/10.3390/v13061078
Nguyen CC, Kamil JP (2018) Pathogen at the Gates: human cytomegalovirus entry and cell tropism. Viruses 10(12). https://doi.org/10.3390/v10120704
Paradowska E et al (2019) Distribution of the CMV glycoprotein gH/gL/gO and gH/gL/pUL128/pUL130/pUL131A complex variants and associated clinical manifestations in infants infected congenitally or postnatally. Sci Rep 9(1):16352. https://doi.org/10.1038/s41598-019-52906-y
Gerna G et al (2019) Human cytomegalovirus cell tropism and host cell receptors. Vaccines 7(3). https://doi.org/10.3390/vaccines7030070
Döhner K et al (2021) The journey of herpesvirus capsids and genomes to the host cell nucleus. Curr Opin Virol 50:147–158. https://doi.org/10.1016/j.coviro.2021.08.005
Isomura H et al (2011) The human cytomegalovirus gene products essential for late viral gene expression assemble into prereplication complexes before viral DNA replication. J Virol 85(13):6629–6644. https://doi.org/10.1128/jvi.00384-11
Adamson CS, Nevels MM (2020) Bright and Early: Inhibiting Human Cytomegalovirus by Targeting Major Immediate-Early Gene expression or protein function. Viruses 12(1). https://doi.org/10.3390/v12010110
Rozman B et al (2022) Temporal dynamics of HCMV gene expression in lytic and latent infections. Cell Rep 39(2):110653. https://doi.org/10.1016/j.celrep.2022.110653
Mitra D et al (2021) Tegument protein pp150 sequence-specific peptide blocks cytomegalovirus infection. Viruses 13(11). https://doi.org/10.3390/v13112277
Leigh KE et al (2015) Structure of a herpesvirus nuclear egress complex subunit reveals an interaction groove that is essential for viral replication. Proc Natl Acad Sci USA 112(29):9010–9015. https://doi.org/10.1073/pnas.1511140112
Poole E et al (2021) Using primary human cells to Analyze Human Cytomegalovirus Biology. Methods in molecular biology. (Clifton NJ) 2244:51–81. https://doi.org/10.1007/978-1-0716-1111-1_4
Zhu H, Zheng C (2020) The race between host antiviral innate immunity and the Immune Evasion Strategies of Herpes Simplex Virus 1. Microbiol Mol Biology Reviews: MMBR 84(4). https://doi.org/10.1128/mmbr.00099-20
Woo Y et al (2024) Hepatocyte intrinsic innate antiviral immunity against Hepatitis Delta Virus infection: the voices of Bona Fide Human hepatocytes. Viruses 16(5). https://doi.org/10.3390/v16050740
Xing J et al (2012) Herpes simplex virus 1 tegument protein US11 downmodulates the RLR signaling pathway via direct interaction with RIG-I and MDA-5. J Virol 86(7):3528–3540. https://doi.org/10.1128/jvi.06713-11
You H et al (2019) Herpes Simplex Virus 1 Tegument Protein UL46 Inhibits TANK-Binding Kinase 1-Mediated Signaling. mBio 10(3) https://doi.org/10.1128/mBio.00919-19
You H et al (2020) β-Catenin is required for the cGAS/STING signaling pathway but antagonized by the Herpes Simplex Virus 1 US3 protein. J Virol 94(5). https://doi.org/10.1128/jvi.01847-19
Zhu H et al (2022) UNC93B1 attenuates the cGAS-STING signaling pathway by targeting STING for autophagy-lysosome degradation. J Med Virol 94(9):4490–4501. https://doi.org/10.1002/jmv.27860
Xu H et al (2017) Herpes Simplex Virus 1 UL24 abrogates the DNA sensing Signal Pathway by inhibiting NF-κB activation. J Virol 91(7). https://doi.org/10.1128/jvi.00025-17
Zhu H et al (2011) Varicella-Zoster virus immediate-early protein ORF61 abrogates the IRF3-mediated innate immune response through degradation of activated IRF3. J Virol 85(21):11079–11089. https://doi.org/10.1128/jvi.05098-11
Akar-Ghibril N (2022) Defects of the Innate Immune System and Related Immune deficiencies. Clin Rev Allergy Immunol 63(1):36–54. https://doi.org/10.1007/s12016-021-08885-y
Takeda K, Akira S (2015) Toll-like receptors. Current protocols in immunology 109:14.12. 11–14.12.10
Full F, Ensser A (2019) Early nuclear events after herpesviral infection. J Clin Med 8(9). https://doi.org/10.3390/jcm8091408
Cui J et al (2014) Mechanisms and pathways of innate immune activation and regulation in health and cancer. Hum Vaccines Immunotherapeutics 10(11):3270–3285. https://doi.org/10.4161/21645515.2014.979640
Thoresen D et al (2021) The molecular mechanism of RIG-I activation and signaling. Immunol Rev 304(1):154–168. https://doi.org/10.1111/imr.13022
Stok JE et al (2022) RNA sensing via the RIG-I-like receptor LGP2 is essential for the induction of a type I IFN response in ADAR1 deficiency. EMBO J 41(6):e109760. https://doi.org/10.15252/embj.2021109760
Xu X et al (2023) Human cytomegalovirus infection activates NLRP3 inflammasome by releasing mtDNA into the cytosol in human THP-1 cells. Microbiol Immunol 67(6):303–313. https://doi.org/10.1111/1348-0421.13063
Lio CW et al (2016) cGAS-STING signaling regulates initial Innate Control of Cytomegalovirus infection. J Virol 90(17):7789–7797. https://doi.org/10.1128/jvi.01040-16
Jin X et al (2023) The battle between the innate immune cGAS-STING signaling pathway and human herpesvirus infection. Front Immunol 14:1235590. https://doi.org/10.3389/fimmu.2023.1235590
Carlsen L, El-Deiry WS (2022) Anti-cancer immune responses to DNA damage response inhibitors: molecular mechanisms and progress toward clinical translation. Front Oncol 12:998388. https://doi.org/10.3389/fonc.2022.998388
Germic N et al (2019) Regulation of the innate immune system by autophagy: monocytes, macrophages, dendritic cells and antigen presentation. Cell Death Differ 26(4):715–727. https://doi.org/10.1038/s41418-019-0297-6
Shi M et al (2021) Endoplasmic reticulum stress-Associated neuronal death and Innate Immune Response in Neurological diseases. Front Immunol 12:794580. https://doi.org/10.3389/fimmu.2021.794580
Yin HC et al (2019) Interactions between Autophagy and DNA viruses. Viruses 11(9). https://doi.org/10.3390/v11090776
Verdonck S et al (2022) Viral manipulation of host cell necroptosis and pyroptosis. Trends Microbiol 30(6):593–605. https://doi.org/10.1016/j.tim.2021.11.011
Zhang P et al (2017) Herpes Simplex Virus 1 UL41 protein suppresses the IRE1/XBP1 Signal Pathway of the unfolded protein response via its RNase activity. J Virol 91(4). https://doi.org/10.1128/jvi.02056-16
Zheng C (2018) Evasion of cytosolic DNA-Stimulated Innate Immune responses by Herpes Simplex Virus 1. J Virol 92. https://doi.org/10.1128/jvi.00099-17
Carty M et al (2021) Detection of viral infections by Innate Immunity. Biochem Pharmacol 183:114316. https://doi.org/10.1016/j.bcp.2020.114316
Wang H et al (2023) Human cytomegalovirus UL23 antagonizes the antiviral effect of Interferon-γ by restraining the expression of specific IFN-Stimulated genes. Viruses 15(4). https://doi.org/10.3390/v15041014
Bowie AG, Unterholzner L (2008) Viral evasion and subversion of pattern-recognition receptor signalling. Nat Rev Immunol 8(12):911–922. https://doi.org/10.1038/nri2436
Goodier MR et al (2018) CMV and natural killer cells: shaping the response to vaccination. Eur J Immunol 48(1):50–65. https://doi.org/10.1002/eji.201646762
El Baba R, Herbein G (2021) Immune Landscape of CMV infection in Cancer patients: from Canonical diseases toward Virus-Elicited Oncomodulation. Front Immunol 12:730765. https://doi.org/10.3389/fimmu.2021.730765
Fernandes-Santos C, Azeredo EL (2022) Innate Immune response to Dengue Virus: toll-like receptors and antiviral response. Viruses 14(5). https://doi.org/10.3390/v14050992
Liu ZM et al (2022) Toll-like receptor (TLRs) agonists and antagonists for COVID-19 treatments. Front Pharmacol 13:989664. https://doi.org/10.3389/fphar.2022.989664
Gay NJ et al (2014) Assembly and localization of toll-like receptor signalling complexes. Nat Rev Immunol 14(8):546–558. https://doi.org/10.1038/nri3713
Kim CU et al (2022) Extracellular nucleoprotein exacerbates influenza virus pathogenesis by activating toll-like receptor 4 and the NLRP3 inflammasome. Cell Mol Immunol 19(6):715–725. https://doi.org/10.1038/s41423-022-00862-5
Cai M et al (2013) The herpes simplex virus 1-encoded envelope glycoprotein B activates NF-κB through the toll-like receptor 2 and MyD88/TRAF6-dependent signaling pathway. PLoS ONE 8(1):e54586. https://doi.org/10.1371/journal.pone.0054586
Cai MS et al (2012) Herpesviral infection and toll-like receptor 2. Protein Cell 3(8):590–601. https://doi.org/10.1007/s13238-012-2059-9
Kim BS (2023) Critical role of TLR activation in viral replication, persistence, and pathogenicity of Theiler’s virus. Front Immunol 14:1167972. https://doi.org/10.3389/fimmu.2023.1167972
Jahanban-Esfahlan R et al (2019) Toll-like receptors as novel therapeutic targets for herpes simplex virus infection. Rev Med Virol 29(4):e2048. https://doi.org/10.1002/rmv.2048
Compton T et al (2003) Human cytomegalovirus activates inflammatory cytokine responses via CD14 and toll-like receptor 2. J Virol 77(8):4588–4596. https://doi.org/10.1128/jvi.77.8.4588-4596.2003
Delale T et al (2005) MyD88-dependent and -independent murine cytomegalovirus sensing for IFN-alpha release and initiation of immune responses in vivo. J Immunol (Baltimore Md: 1950) 175(10):6723–6732. https://doi.org/10.4049/jimmunol.175.10.6723
Arav-Boger R et al (2012) Polymorphisms in toll-like receptor genes influence antibody responses to cytomegalovirus glycoprotein B vaccine. BMC Res Notes 5:140. https://doi.org/10.1186/1756-0500-5-140
Boehme KW et al (2006) Human cytomegalovirus envelope glycoproteins B and H are necessary for TLR2 activation in permissive cells. J Immunol (Baltimore Md: 1950) 177(10):7094–7102. https://doi.org/10.4049/jimmunol.177.10.7094
Xie F et al (2014) CMV infection, TLR-2 and – 4 expression, and cytokine profiles in early-onset preeclampsia with HELLP syndrome. Am J Reproductive Immunol (New York NY: 1989) 71(4):379–386. https://doi.org/10.1111/aji.12199
Smith PD et al (2014) Cytomegalovirus enhances macrophage TLR expression and MyD88-mediated signal transduction to potentiate inducible inflammatory responses. J Immunol (Baltimore Md: 1950) 193(11):5604–5612. https://doi.org/10.4049/jimmunol.1302608
Redondo N et al (2022) Influence of single-nucleotide polymorphisms in TLR3 (rs3775291) and TLR9 (rs352139) on the risk of CMV infection in kidney transplant recipients. Front Immunol 13:929995. https://doi.org/10.3389/fimmu.2022.929995
Boo KH, Yang JS (2010) Intrinsic cellular defenses against virus infection by antiviral type I interferon. Yonsei Med J 51(1):9–17. https://doi.org/10.3349/ymj.2010.51.1.9
Mhandire DZ et al (2020) Genetic variation in toll like receptors 2, 7, 9 and interleukin-6 is associated with cytomegalovirus infection in late pregnancy. BMC Med Genet 21(1):113. https://doi.org/10.1186/s12881-020-01044-8
Taniguchi R et al (2013) Polymorphisms in TLR-2 are associated with congenital cytomegalovirus (CMV) infection but not with congenital CMV disease. Int J Infect Diseases: IJID : Official Publication Int Soc Infect Dis 17(12):e1092–1097. https://doi.org/10.1016/j.ijid.2013.06.004
Pei RJ et al (2014) Control of hepatitis B virus replication by interferons and toll-like receptor signaling pathways. World J Gastroenterol 20(33):11618–11629. https://doi.org/10.3748/wjg.v20.i33.11618
Park A et al (2019) HCMV-encoded US7 and US8 act as antagonists of innate immunity by distinctively targeting TLR-signaling pathways. Nat Commun 10(1):4670. https://doi.org/10.1038/s41467-019-12641-4
Landais I et al (2015) Human cytomegalovirus miR-UL112-3p targets TLR2 and modulates the TLR2/IRAK1/NFκB signaling pathway. PLoS Pathog 11(5):e1004881. https://doi.org/10.1371/journal.ppat.1004881
Zheng J et al (2023) RIG-I-like receptors: molecular mechanism of activation and signaling. Adv Immunol 158:1–74. https://doi.org/10.1016/bs.ai.2023.03.001
Duic I et al (2020) Viral RNA recognition by LGP2 and MDA5, and activation of signaling through step-by-step conformational changes. Nucleic Acids Res 48(20):11664–11674. https://doi.org/10.1093/nar/gkaa935
Di Giorgio E, Xodo LE (2022) Endogenous retroviruses (ERVs): does RLR (RIG-I-Like Receptors)-MAVS Pathway directly Control Senescence and Aging as a consequence of ERV De-repression? Front Immunol 13:917998. https://doi.org/10.3389/fimmu.2022.917998
Ren Y et al (2023) Human cytomegalovirus UL36 inhibits IRF3-dependent immune signaling to counterbalance its immunoenhancement as apoptotic inhibitor. Sci Adv 9(40):eadi6586. https://doi.org/10.1126/sciadv.adi6586
Da Silva MHM et al (2019) Innate immune response in patients with acute Zika virus infection. Med Microbiol Immunol 208(6):703–714. https://doi.org/10.1007/s00430-019-00588-8
Zhao Y, Karijolich J (2019) Know Thyself: RIG-I-Like receptor sensing of DNA virus infection. J Virol 93(23). https://doi.org/10.1128/jvi.01085-19
Bryant AH et al (2017) Human gestation-associated tissues express functional cytosolic nucleic acid sensing pattern recognition receptors. Clin Exp Immunol 189(1):36–46. https://doi.org/10.1111/cei.12960
Jabłońska A et al (2020) Insight into the expression of RIG-I-like receptors in human third trimester placentas following ex vivo cytomegalovirus or vesicular stomatitis virus infection. Mol Immunol 126:143–152. https://doi.org/10.1016/j.molimm.2020.08.001
Elliott EI, Sutterwala FS (2015) Initiation and perpetuation of NLRP3 inflammasome activation and assembly. Immunol Rev 265(1):35–52. https://doi.org/10.1111/imr.12286
Eisfeld HS et al (2021) Viral glycoproteins induce NLRP3 inflammasome activation and Pyroptosis in macrophages. Viruses 13(10). https://doi.org/10.3390/v13102076
Kuenzel S et al (2010) The nucleotide-binding oligomerization domain-like receptor NLRC5 is involved in IFN-dependent antiviral immune responses. J Immunol (Baltimore Md: 1950) 184(4):1990–2000. https://doi.org/10.4049/jimmunol.0900557
Fan YH et al (2016) Role of nucleotide-binding oligomerization domain 1 (NOD1) and its variants in human cytomegalovirus control in vitro and in vivo. Proc Natl Acad Sci USA 113(48):E7818–e7827. https://doi.org/10.1073/pnas.1611711113
Li M et al (2023) Murine cytomegalovirus employs the mixed lineage kinases family to regulate the spiral ganglion neuron cell death and hearing loss. Neurosci Lett 793:136990. https://doi.org/10.1016/j.neulet.2022.136990
Wang Y et al (2020) Mir-1929-3p overexpression alleviates Murine Cytomegalovirus-Induced Hypertensive Myocardial remodeling by suppressing Ednra/NLRP3 inflammasome activation. Biomed Res Int 2020:6653819. https://doi.org/10.1155/2020/6653819
Man SM et al (2016) AIM2 inflammasome in infection, cancer, and autoimmunity: role in DNA sensing, inflammation, and innate immunity. Eur J Immunol 46(2):269–280. https://doi.org/10.1002/eji.201545839
Huang Y et al (2017) Human cytomegalovirus triggers the assembly of AIM2 inflammasome in THP-1-derived macrophages. J Med Virol 89(12):2188–2195. https://doi.org/10.1002/jmv.24846
Shi X et al (2015) Inflammasome activation in mouse inner ear in response to MCMV induced hearing loss. J Otology 10(4):143–149. https://doi.org/10.1016/j.joto.2015.12.001
Huang Y et al (2017) Interaction between HCMV pUL83 and human AIM2 disrupts the activation of the AIM2 inflammasome. Virol J 14(1):34. https://doi.org/10.1186/s12985-016-0673-5
Botto S et al (2019) Human cytomegalovirus Immediate Early 86-kDa protein blocks transcription and induces degradation of the immature Interleukin-1β protein during virion-mediated activation of the AIM2 inflammasome. mBio 10(1). https://doi.org/10.1128/mBio.02510-18
Hemmi H et al (2000) A toll-like receptor recognizes bacterial DNA. Nature 408(6813):740–745. https://doi.org/10.1038/35047123
Briard B et al (2020) DNA sensing in the Innate Immune Response. Physiol (Bethesda Md) 35(2):112–124. https://doi.org/10.1152/physiol.00022.2019
Huérfano S et al (2022) The interplay between viruses and host DNA sensors. Viruses 14(4). https://doi.org/10.3390/v14040666
Sun L et al (2013) Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339(6121):786–791. https://doi.org/10.1126/science.1232458
Kwon J, Bakhoum SF (2020) The cytosolic DNA-Sensing cGAS-STING pathway in Cancer. Cancer Discov 10(1):26–39. https://doi.org/10.1158/2159-8290.Cd-19-0761
Decout A et al (2021) The cGAS-STING pathway as a therapeutic target in inflammatory diseases. Nat Rev Immunol 21(9):548–569. https://doi.org/10.1038/s41577-021-00524-z
Erttmann SF et al (2022) The gut microbiota prime systemic antiviral immunity via the cGAS-STING-IFN-I axis. Immunity 55(5):847–861e810. https://doi.org/10.1016/j.immuni.2022.04.006
Wang S et al (2022) PEBP balances apoptosis and autophagy in whitefly upon arbovirus infection. Nat Commun 13(1):846. https://doi.org/10.1038/s41467-022-28500-8
Döhner K et al (2005) Viral stop-and-go along microtubules: taking a ride with dynein and kinesins. Trends Microbiol 13(7):320–327. https://doi.org/10.1016/j.tim.2005.05.010
Bauer DW et al (2013) Herpes virus genome, the pressure is on. J Am Chem Soc 135(30):11216–11221. https://doi.org/10.1021/ja404008r
Brandariz-Nuñez A et al (2019) Pressure-driven release of viral genome into a host nucleus is a mechanism leading to herpes infection. eLife 8. https://doi.org/10.7554/eLife.47212
Horan KA et al (2013) Proteasomal degradation of herpes simplex virus capsids in macrophages releases DNA to the cytosol for recognition by DNA sensors. J Immunol (Baltimore Md: 1950) 190(5):2311–2319. https://doi.org/10.4049/jimmunol.1202749
Huang ZF et al (2018) Human cytomegalovirus protein UL31 inhibits DNA sensing of cGAS to Mediate Immune Evasion. Cell Host Microbe 24(1):69–80e64. https://doi.org/10.1016/j.chom.2018.05.007
Biolatti M et al (2018) Human cytomegalovirus tegument protein pp65 (pUL83) dampens type I Interferon Production by inactivating the DNA sensor cGAS without affecting STING. J Virol 92(6). https://doi.org/10.1128/jvi.01774-17
Fu YZ et al (2019) Human cytomegalovirus protein UL42 antagonizes cGAS/MITA-mediated innate antiviral response. PLoS Pathog 15(5):e1007691. https://doi.org/10.1371/journal.ppat.1007691
Burdette DL, Vance RE (2013) STING and the innate immune response to nucleic acids in the cytosol. Nat Immunol 14(1):19–26. https://doi.org/10.1038/ni.2491
Zhang ZD, Zhong B (2022) Regulation and function of the cGAS-MITA/STING axis in health and disease. Cell Insight 1(1):100001. https://doi.org/10.1016/j.cellin.2021.100001
Albright ER et al (2021) Human cytomegalovirus UL138 protein inhibits the STING pathway and reduces Interferon Beta mRNA Accumulation during Lytic and latent infections. mBio 12(6):e0226721. https://doi.org/10.1128/mBio.02267-21
Kim JE et al (2018) Corrigendum: human cytomegalovirus IE2 86 kDa protein induces STING degradation and inhibits cGAMP-Mediated IFN-β induction. Front Microbiol 9:350. https://doi.org/10.3389/fmicb.2018.00350
Lee JK et al (2020) Human cytomegalovirus IE86 protein aa 136–289 mediates STING degradation and blocks the cGAS-STING pathway. J Microbiol 58(1):54–60. https://doi.org/10.1007/s12275-020-9577-6
Bresnahan WA, Shenk TE (2000) UL82 virion protein activates expression of immediate early viral genes in human cytomegalovirus-infected cells. Proc Natl Acad Sci USA 97(26):14506–14511. https://doi.org/10.1073/pnas.97.26.14506
Fu YZ et al (2017) Human cytomegalovirus tegument protein UL82 inhibits STING-Mediated signaling to evade antiviral immunity. Cell Host Microbe 21(2):231–243. https://doi.org/10.1016/j.chom.2017.01.001
Zou HM et al (2020) Human cytomegalovirus protein UL94 targets MITA to evade the antiviral Immune response. J Virol 94. https://doi.org/10.1128/jvi.00022-20
Kwon KM et al (2021) Human cytomegalovirus UL48 deubiquitinase primarily targets Innermost Tegument proteins pp150 and itself to regulate their Stability and protects virions from inclusion of Ubiquitin conjugates. J Virol 95(23):e0099121. https://doi.org/10.1128/jvi.00991-21
Stempel M et al (2019) The herpesviral antagonist m152 reveals differential activation of STING-dependent IRF and NF-κB signaling and STING’s dual role during MCMV infection. EMBO J 38(5). https://doi.org/10.15252/embj.2018100983
Choi HJ et al (2018) Human cytomegalovirus-encoded US9 targets MAVS and STING signaling to evade type I interferon immune responses. Nat Commun 9(1):125. https://doi.org/10.1038/s41467-017-02624-8
Fu Y et al (2023) USP12 promotes antiviral responses by deubiquitinating and stabilizing IFI16. PLoS Pathog 19(7):e1011480. https://doi.org/10.1371/journal.ppat.1011480
Dell’oste V et al (2014) Innate nuclear sensor IFI16 translocates into the cytoplasm during the early stage of in vitro human cytomegalovirus infection and is entrapped in the egressing virions during the late stage. J Virol 88(12):6970–6982. https://doi.org/10.1128/jvi.00384-14
Landolfo S et al (2016) Intrinsic host restriction factors of human cytomegalovirus replication and mechanisms of viral escape. World J Virol 5(3):87–96. https://doi.org/10.5501/wjv.v5.i3.87
Yum S et al (2021) TBK1 recruitment to STING activates both IRF3 and NF-κB that mediate immune defense against tumors and viral infections. Proc Natl Acad Sci USA 118(14). https://doi.org/10.1073/pnas.2100225118
Liu Y et al (2022) TRIM56 positively regulates TNFα-induced NF-κB signaling by enhancing the ubiquitination of TAK1. Int J Biol Macromol 219:571–578. https://doi.org/10.1016/j.ijbiomac.2022.08.019
Fabits M et al (2020) The Cytomegalovirus tegument protein UL35 antagonizes pattern recognition receptor-mediated type I IFN transcription. Microorganisms 8(6). https://doi.org/10.3390/microorganisms8060790
Sun H et al (2017) USP13 negatively regulates antiviral responses by deubiquitinating STING. Nat Commun 8:15534. https://doi.org/10.1038/ncomms15534
Fu YZ et al (2019) Human cytomegalovirus DNA polymerase subunit UL44 antagonizes antiviral Immune responses by suppressing IRF3- and NF-κB-Mediated transcription. J Virol 93(11). https://doi.org/10.1128/jvi.00181-19
García-Ríos E et al (2022) Identification and characterization of epithelial cell-derived dense bodies produced upon cytomegalovirus infection. Vaccines 10(8). https://doi.org/10.3390/vaccines10081308
Abate DA et al (2004) Major human cytomegalovirus structural protein pp65 (ppUL83) prevents interferon response factor 3 activation in the interferon response. J Virol 78(20):10995–11006. https://doi.org/10.1128/jvi.78.20.10995-11006.2004
Schwanke H et al (2023) The Cytomegalovirus M35 protein directly binds to the Interferon-β enhancer and modulates transcription of Ifnb1 and other IRF3-Driven genes. J Virol 97(6):e0040023. https://doi.org/10.1128/jvi.00400-23
Alharbi KS et al (2021) Nuclear factor-kappa B and its role in inflammatory lung disease. Chemico-Biol Interact 345:109568. https://doi.org/10.1016/j.cbi.2021.109568
Zhang Y et al (2014) Krüppel-like factor 6 is a co-activator of NF-κB that mediates p65-dependent transcription of selected downstream genes. J Biol Chem 289(18):12876–12885. https://doi.org/10.1074/jbc.M113.535831
Nennig SE, Schank JR (2017) The role of NFkB in drug addiction: beyond inflammation. Alcohol and alcoholism. (Oxford Oxfordshire) 52(2):172–179. https://doi.org/10.1093/alcalc/agw098
Lai JL et al (2017) Indirubin inhibits LPS-Induced inflammation via TLR4 abrogation mediated by the NF-kB and MAPK signaling pathways. Inflammation 40(1):1–12. https://doi.org/10.1007/s10753-016-0447-7
Iwai K (2014) Diverse roles of the ubiquitin system in NF-κB activation. Biochim Biophys Acta 1843(1):129–136. https://doi.org/10.1016/j.bbamcr.2013.03.011
Krause E et al (2014) Murine cytomegalovirus virion-associated protein M45 mediates rapid NF-κB activation after infection. J Virol 88(17):9963–9975. https://doi.org/10.1128/jvi.00684-14
Mathers C et al (2014) The human cytomegalovirus UL26 protein antagonizes NF-κB activation. J Virol 88(24):14289–14300. https://doi.org/10.1128/jvi.02552-14
Guo G et al (2018) The cytomegalovirus protein US31 induces inflammation through mono-macrophages in systemic lupus erythematosus by promoting NF-κB2 activation. Cell Death Dis 9(2):104. https://doi.org/10.1038/s41419-017-0122-4
Biolatti M et al (2018) The viral tegument protein pp65 impairs transcriptional upregulation of IL-1β by human cytomegalovirus through inhibition of NF-kB activity. Viruses 10(10). https://doi.org/10.3390/v10100567
Krishna BA et al (2017) Latency-Associated Expression of Human Cytomegalovirus US28 Attenuates Cell Signaling Pathways To Maintain Latent Infection. mBio 8(6) https://doi.org/10.1128/mBio.01754-17
Chan B et al (2017) The murine cytomegalovirus M35 protein antagonizes type I IFN induction downstream of pattern recognition receptors by targeting NF-κB mediated transcription. PLoS Pathog 13(5):e1006382. https://doi.org/10.1371/journal.ppat.1006382
Chen H et al (2024) Arteannuin B, a sesquiterpene lactone from Artemisia annua, attenuates inflammatory response by inhibiting the ubiquitin-conjugating enzyme UBE2D3-mediated NF-κB activation. Phytomedicine: Int J Phytotherapy Phytopharmacology 124:155263. https://doi.org/10.1016/j.phymed.2023.155263
Fliss PM et al (2012) Viral mediated redirection of NEMO/IKKγ to autophagosomes curtails the inflammatory cascade. PLoS Pathog 8(2):e1002517. https://doi.org/10.1371/journal.ppat.1002517
Mack C et al (2008) Inhibition of proinflammatory and innate immune signaling pathways by a cytomegalovirus RIP1-interacting protein. Proc Natl Acad Sci USA 105(8):3094–3099. https://doi.org/10.1073/pnas.0800168105
Kwon KM et al (2017) Cooperative inhibition of RIP1-mediated NF-κB signaling by cytomegalovirus-encoded deubiquitinase and inactive homolog of cellular ribonucleotide reductase large subunit. PLoS Pathog 13(6):e1006423. https://doi.org/10.1371/journal.ppat.1006423
Hilterbrand AT et al (2017) Murine cytomegalovirus deubiquitinase regulates viral chemokine levels to control inflammation and Pathogenesis. mBio 8(1). https://doi.org/10.1128/mBio.01864-16
De Weerd NA et al (2007) Type I interferon receptors: biochemistry and biological functions. J Biol Chem 282(28):20053–20057. https://doi.org/10.1074/jbc.R700006200
Platanias LC (2005) Mechanisms of type-I- and type-II-interferon-mediated signalling. Nat Rev Immunol 5(5):375–386. https://doi.org/10.1038/nri1604
Stark GR, Darnell JE Jr. (2012) The JAK-STAT pathway at twenty. Immunity 36(4):503–514. https://doi.org/10.1016/j.immuni.2012.03.013
Mccormack R et al (2020) An Essential Role for Perforin-2 in Type I IFN Signaling. Journal of immunology (Baltimore, Md: 1950) 204(8):2242–2256. https://doi.org/10.4049/jimmunol.1901013
Chen Q et al (2023) MGF360-12L of ASFV-SY18 is an immune-evasion protein that inhibits host type I IFN, NF-κB, and JAK/STAT pathways. Pol J Vet Sci 26(1):119–130. https://doi.org/10.24425/pjvs.2023.145013
Ibrahim MK et al (2017) Increased incidence of cytomegalovirus coinfection in HCV-infected patients with late liver fibrosis is associated with dysregulation of JAK-STAT pathway. Sci Rep 7(1):10364. https://doi.org/10.1038/s41598-017-10604-7
Le VTK et al (2008) Human cytomegalovirus interferes with signal transducer and activator of transcription (STAT) 2 protein stability and tyrosine phosphorylation. J Gen Virol 89(Pt 102416–2426. https://doi.org/10.1099/vir.0.2008/001669-0
Abenes G et al (2001) Murine cytomegalovirus open reading frame M27 plays an important role in growth and virulence in mice. J Virol 75(4):1697–1707. https://doi.org/10.1128/jvi.75.4.1697-1707.2001
Zimmermann A et al (2005) A cytomegaloviral protein reveals a dual role for STAT2 in IFN-{gamma} signaling and antiviral responses. J Exp Med 201(10):1543–1553. https://doi.org/10.1084/jem.20041401
Trilling M et al (2011) Identification of DNA-damage DNA-binding protein 1 as a conditional essential factor for cytomegalovirus replication in interferon-γ-stimulated cells. PLoS Pathog 7(6):e1002069. https://doi.org/10.1371/journal.ppat.1002069
Le-Trilling VTK et al (2020) The human cytomegalovirus pUL145 isoforms Act as viral DDB1-Cullin-Associated factors to instruct host protein degradation to Impede Innate Immunity. Cell Rep 30(7):2248–2260e2245. https://doi.org/10.1016/j.celrep.2020.01.070
Feng L et al (2018) Human cytomegalovirus UL23 inhibits transcription of interferon-γ stimulated genes and blocks antiviral interferon-γ responses by interacting with human N-myc interactor protein. PLoS Pathog 14(1):e1006867. https://doi.org/10.1371/journal.ppat.1006867
Feng L et al (2021) Human cytomegalovirus UL23 attenuates Signal Transducer and activator of transcription 1 phosphorylation and type I Interferon Response. Front Microbiol 12:692515. https://doi.org/10.3389/fmicb.2021.692515
Ciesla J et al (2024) A UL26-PIAS1 complex antagonizes anti-viral gene expression during human cytomegalovirus infection. PLoS Pathog 20(5):e1012058. https://doi.org/10.1371/journal.ppat.1012058
Zhang Q et al (2021) Human cytomegalovirus miR-US33as-5p targets IFNAR1 to Achieve Immune Evasion during both lytic and latent infection. Front Immunol 12:628364. https://doi.org/10.3389/fimmu.2021.628364
Ashley CL et al (2019) Interferon-Independent innate responses to Cytomegalovirus. Front Immunol 10:2751. https://doi.org/10.3389/fimmu.2019.02751
Ashley CL et al (2019) Interferon-Independent Upregulation of Interferon-stimulated genes during human cytomegalovirus infection is dependent on IRF3 expression. Viruses 11(3). https://doi.org/10.3390/v11030246
Tecalco Cruz AC (2022) Free ISG15 and protein ISGylation emerging in SARS-CoV-2 infection. Curr Drug Targets 23(7):686–691. https://doi.org/10.2174/1389450123666220316094720
Bianco C, Mohr I (2017) Restriction of human cytomegalovirus replication by ISG15, a host Effector regulated by cGAS-STING double-Stranded-DNA sensing. J Virol 91(9). https://doi.org/10.1128/jvi.02483-16
Goodwin CM et al (2019) U(L)26 attenuates IKKβ-Mediated induction of Interferon-stimulated gene (ISG) expression and enhanced protein ISGylation during human cytomegalovirus infection. J Virol 93(23). https://doi.org/10.1128/jvi.01052-19
Kim YJ et al (2016) Consecutive inhibition of ISG15 expression and ISGylation by Cytomegalovirus regulators. PLoS Pathog 12(8):e1005850. https://doi.org/10.1371/journal.ppat.1005850
Lee MK et al (2018) Transmembrane protein pUL50 of human cytomegalovirus inhibits ISGylation by downregulating UBE1L. J Virol 92. https://doi.org/10.1128/jvi.00462-18
Gonzalez-Perez AC et al (2021) The Zinc Finger Antiviral Protein ZAP Restricts Human Cytomegalovirus and Selectively Binds and Destabilizes Viral UL4/UL5 Transcripts. mBio 12(3) https://doi.org/10.1128/mBio.02683-20
Lista MJ et al (2023) Strain-dependent restriction of human cytomegalovirus by Zinc Finger antiviral proteins. J Virol 97(3):e0184622. https://doi.org/10.1128/jvi.01846-22
Lin YT et al (2020) Human cytomegalovirus evades ZAP detection by suppressing CpG dinucleotides in the major immediate early 1 gene. PLoS Pathog 16(9):e1008844. https://doi.org/10.1371/journal.ppat.1008844
Zhang L et al (2017) Antiviral effects of IFIT1 in human cytomegalovirus-infected fetal astrocytes. J Med Virol 89(4):672–684. https://doi.org/10.1002/jmv.24674
Zani A, Yount JS (2018) Antiviral protection by IFITM3 in vivo. Curr Clin Microbiol Rep 5(4):229–237. https://doi.org/10.1007/s40588-018-0103-0
Griffante G et al (2021) Human cytomegalovirus-induced host protein citrullination is crucial for viral replication. Nat Commun 12(1):3910. https://doi.org/10.1038/s41467-021-24178-6
Seo JY et al (2011) Human cytomegalovirus directly induces the antiviral protein viperin to enhance infectivity. Sci (New York NY) 332(6033):1093–1097. https://doi.org/10.1126/science.1202007
Kim JJ et al (2019) Viperin differentially induces Interferon-stimulated genes in distinct cell types. Immune Netw 19(5):e33. https://doi.org/10.4110/in.2019.19.e33
Kim JJ et al (2023) A cysteine Residue of Human Cytomegalovirus vMIA protein plays a crucial role in Viperin Trafficking to control viral infectivity. J Virol 97(6):e0187422. https://doi.org/10.1128/jvi.01874-22
Wang B et al (2015) Viperin is induced following toll-like receptor 3 (TLR3) ligation and has a virus-responsive function in human trophoblast cells. Placenta 36(6):667–673. https://doi.org/10.1016/j.placenta.2015.03.002
Carpentier KS et al (2016) A single amino acid dictates protein kinase R susceptibility to unrelated viral antagonists. PLoS Pathog 12(10):e1005966. https://doi.org/10.1371/journal.ppat.1005966
Ziehr B et al (2016) Human cytomegalovirus pTRS1 and pIRS1 antagonize protein kinase R to facilitate Virus Replication. J Virol 90(8):3839–3848. https://doi.org/10.1128/jvi.02714-15
Bierle CJ et al (2013) Double-stranded RNA binding by the human cytomegalovirus PKR antagonist TRS1. Virology 442(1):28–37. https://doi.org/10.1016/j.virol.2013.03.024
Vincent HA et al (2017) Mechanism of protein kinase R inhibition by human cytomegalovirus pTRS1. J Virol 91. https://doi.org/10.1128/jvi.01574-16
Braggin JE et al (2016) Essential role of protein kinase R antagonism by TRS1 in human cytomegalovirus replication. Virology 489:75–85. https://doi.org/10.1016/j.virol.2015.11.032
Child SJ, Geballe AP (2009) Binding and relocalization of protein kinase R by murine cytomegalovirus. J Virol 83(4):1790–1799. https://doi.org/10.1128/jvi.01484-08
Ostermann E et al (2016) Knockout of the Host Resistance Gene pkr fully restores replication of murine cytomegalovirus m142 and m143 mutants in vivo. J Virol 90(2):1144–1147. https://doi.org/10.1128/jvi.02003-15
Bierle CJ et al (2012) Antagonism of the protein kinase R pathway by the guinea pig cytomegalovirus US22-family gene gp145. Virology 433(1):157–166. https://doi.org/10.1016/j.virol.2012.08.005
Giraldo MI et al (2020) TRIM proteins in Host Defense and viral pathogenesis. Curr Clin Microbiol Rep 7(4):101–114. https://doi.org/10.1007/s40588-020-00150-8
Schilling EM et al (2017) The human cytomegalovirus IE1 protein antagonizes PML nuclear body-mediated intrinsic immunity via the inhibition of PML De Novo SUMOylation. J Virol 91(4). https://doi.org/10.1128/jvi.02049-16
Scherer M et al (2014) Crystal structure of cytomegalovirus IE1 protein reveals targeting of TRIM family member PML via coiled-coil interactions. PLoS Pathog 10(11):e1004512. https://doi.org/10.1371/journal.ppat.1004512
Martínez FP, Tang Q (2013) Identification of cellular proteins that interact with human cytomegalovirus immediate-early protein 1 by protein array assay. Viruses 6(1):89–105. https://doi.org/10.3390/v6010089
Ampomah PB, Lim LHK (2020) Influenza a virus-induced apoptosis and virus propagation. Apoptosis: Int J Program cell Death 25(1–2):1–11. https://doi.org/10.1007/s10495-019-01575-3
Verburg SG et al (2022) Viral-mediated activation and inhibition of programmed cell death. PLoS Pathog 18(8):e1010718. https://doi.org/10.1371/journal.ppat.1010718
Croft SN et al (2017) Picornaviruses and apoptosis: subversion of cell death. mBio 8(5). https://doi.org/10.1128/mBio.01009-17
Khalid N, Azimpouran M (2023) Necrosis [M]. StatPearls. Treasure Island (FL) ineligible companies. Disclosure: Mahzad Azimpouran declares no relevant financial relationships with ineligible companies.; StatPearls Publishing. Copyright © 2023. StatPearls Publishing LLC
Gao J et al (2023) When ferroptosis meets pathogenic infections. Trends Microbiol 31(5):468–479. https://doi.org/10.1016/j.tim.2022.11.006
Brune W, Andoniou CE (2017) Die another day: inhibition of cell death pathways by Cytomegalovirus. Viruses 9(9). https://doi.org/10.3390/v9090249
Chaudhry MZ et al (2017) UL36 rescues apoptosis inhibition and in vivo replication of a chimeric MCMV lacking the M36 gene. Front Cell Infect Microbiol 7:312. https://doi.org/10.3389/fcimb.2017.00312
Fletcher-Etherington A et al (2020) Human cytomegalovirus protein pUL36: a dual cell death pathway inhibitor. Proc Natl Acad Sci USA 117(31):18771–18779. https://doi.org/10.1073/pnas.2001887117
Muscolino E et al (2021) Species-specific inhibition of necroptosis by HCMV UL36. Viruses 13(11). https://doi.org/10.3390/v13112134
Ebermann L et al (2012) Block of death-receptor apoptosis protects mouse cytomegalovirus from macrophages and is a determinant of virulence in immunodeficient hosts. PLoS Pathog 8(12):e1003062. https://doi.org/10.1371/journal.ppat.1003062
Peña-Blanco A, García-Sáez AJ (2018) Bax, Bak and beyond - mitochondrial performance in apoptosis. FEBS J 285(3):416–431. https://doi.org/10.1111/febs.14186
Norris KL, Youle RJ (2008) Cytomegalovirus proteins vMIA and m38.5 link mitochondrial morphogenesis to Bcl-2 family proteins. J Virol 82(13):6232–6243. https://doi.org/10.1128/jvi.02710-07
Nemčovičová I et al (2013) Structure of human cytomegalovirus UL141 binding to TRAIL-R2 reveals novel, non-canonical death receptor interactions. PLoS Pathog 9(3):e1003224. https://doi.org/10.1371/journal.ppat.1003224
Kolbrink B et al (2022) TAT-RHIM: a more complex issue than expected. Biochem J 479(3):259–272. https://doi.org/10.1042/bcj20210677
Mandal P et al (2021) Multiple Autonomous Cell Death suppression strategies ensure Cytomegalovirus Fitness. Viruses 13(9). https://doi.org/10.3390/v13091707
Zhu H et al (1995) Human cytomegalovirus IE1 and IE2 proteins block apoptosis. J Virol 69(12):7960–7970. https://doi.org/10.1128/jvi.69.12.7960-7970.1995
Paulus C, Nevels M (2009) The human cytomegalovirus major immediate-early proteins as antagonists of intrinsic and innate antiviral host responses. Viruses 1(3):760–779. https://doi.org/10.3390/v1030760
Heusel AT et al (2024) IE1 of human cytomegalovirus inhibits necroptotic cell death via Direct and Indirect Modulation of the Necrosome Complex. Viruses 16(2). https://doi.org/10.3390/v16020290
Luganini A et al (2018) Human cytomegalovirus US21 protein is a viroporin that modulates calcium homeostasis and protects cells against apoptosis. Proc Natl Acad Sci USA 115(52):E12370–e12377. https://doi.org/10.1073/pnas.1813183115
Mao J et al (2019) Autophagy and viral infection. Adv Exp Med Biol 1209:55–78. https://doi.org/10.1007/978-981-15-0606-2_5
Gerada C, Ryan KM (2020) Autophagy, the innate immune response and cancer. Mol Oncol 14(9):1913–1929. https://doi.org/10.1002/1878-0261.12774
Zimmermann C et al (2021) Autophagy interferes with human cytomegalovirus genome replication, morphogenesis, and progeny release. Autophagy 17(3):779–795. https://doi.org/10.1080/15548627.2020.1732686
Belzile JP et al (2016) Trehalose, an mTOR-Independent inducer of Autophagy, inhibits human cytomegalovirus infection in multiple cell types. J Virol 90(3):1259–1277. https://doi.org/10.1128/jvi.02651-15
Clark AE et al (2018) Human cytomegalovirus replication is inhibited by the Autophagy-Inducing compounds Trehalose and SMER28 through distinctively different mechanisms. J Virol 92(6). https://doi.org/10.1128/jvi.02015-17
König P et al (2021) The autophagy-initiating protein kinase ULK1 Phosphorylates Human Cytomegalovirus tegument protein pp28 and regulates efficient Virus Release. J Virol 95(6). https://doi.org/10.1128/jvi.02346-20
Mouna L et al (2016) Analysis of the role of autophagy inhibition by two complementary human cytomegalovirus BECN1/Beclin 1-binding proteins. Autophagy 12(2):327–342. https://doi.org/10.1080/15548627.2015.1125071
Chaumorcel M et al (2018) Correction for Chaumorcel., The Human Cytomegalovirus Protein TRS1 Inhibits Autophagy via Its Interaction with Beclin 1. Journal of virology 92(9) https://doi.org/10.1128/jvi.00252-18
Kim HJ et al (2023) HCMV-encoded viral protein US12 promotes autophagy by inducing autophagy flux. Biochem Biophys Res Commun 654:94–101. https://doi.org/10.1016/j.bbrc.2023.03.004
Merighi A, Lossi L (2022) Endoplasmic reticulum stress signaling and neuronal cell death. Int J Mol Sci 23(23). https://doi.org/10.3390/ijms232315186
Read A, Schröder M (2021) The unfolded protein response: an overview. Biology 10(5). https://doi.org/10.3390/biology10050384
Jeon JH et al (2022) Chemical chaperones to inhibit endoplasmic reticulum stress: implications in diseases. Drug design. Dev Therapy 16:4385–4397. https://doi.org/10.2147/dddt.S393816
Xuan B et al (2009) Human cytomegalovirus protein pUL38 induces ATF4 expression, inhibits persistent JNK phosphorylation, and suppresses endoplasmic reticulum stress-induced cell death. J Virol 83(8):3463–3474. https://doi.org/10.1128/jvi.02307-08
Nguyen CC et al (2020) Endoplasmic reticulum (ER) reorganization and Intracellular Retention of CD58 are functionally independent properties of the human cytomegalovirus ER-Resident glycoprotein UL148. J Virol 94(5). https://doi.org/10.1128/jvi.01435-19
Siddiquey MNA et al (2018) The Human Cytomegalovirus Endoplasmic Reticulum-Resident glycoprotein UL148 activates the unfolded protein response. J Virol 92(20). https://doi.org/10.1128/jvi.00896-18
Stahl S et al (2013) Cytomegalovirus downregulates IRE1 to repress the unfolded protein response. PLoS Pathog 9(8):e1003544. https://doi.org/10.1371/journal.ppat.1003544
Hinte F et al (2021) Viral mediated tethering to SEL1L facilitates ER-associated degradation of IRE1. J Virol 95. https://doi.org/10.1128/jvi.01990-20
Yan Y et al (2022) Novel molecular therapeutics Targeting Signaling Pathway to Control Hepatitis B viral infection. Front Cell Infect Microbiol 12:847539. https://doi.org/10.3389/fcimb.2022.847539
Rudra JS et al (2018) A combined carrier-adjuvant system of peptide nanofibers and toll-like receptor agonists potentiates robust CD8 + T cell responses. Vaccine 36(4):438–441. https://doi.org/10.1016/j.vaccine.2017.12.017
Pietrobon AJ et al (2022) Antiviral Response Induced by Toll-Like receptor (TLR) 7/TLR8 activation inhibits human immunodeficiency virus type 1 infection in Cord Blood macrophages. J Infect Dis 225(3):510–519. https://doi.org/10.1093/infdis/jiab389
Port A et al (2021) Phase 1 study in healthy participants of the safety, pharmacokinetics, and pharmacodynamics of enpatoran (M5049), a dual antagonist of toll-like receptors 7 and 8. Pharmacol Res Perspect 9(5):e00842. https://doi.org/10.1002/prp2.842
Acknowledgements
We are grateful to all the laboratory members for their valuable assistance and suggestions throughout the collection and analysis of our article.
Funding
This work was supported by grants from the Guizhou Epilepsy Basic and Clinical Research Scientific and Technological Innovation Talent Team Project (No: CXTD[2022]013), the Collaborative Innovation Center of the Chinese Ministry of Education (No: 2020-39), the Guizhou Provincial “Hundred” Level Innovative Talents Funds (No: GCC-2022-038-1), the Guizhou Provincial Science and Technology Foundation (No: ZK 2022 − 656), the Zunyi City Science and Technology Foundation (No: 2019-71 and 2021-30) and the National Natural Science Foundation of China (82072263).
Author information
Authors and Affiliations
Contributions
Shuang Li and Xie Yuanyang wrote the original manuscript. Zucai Xu, Chunfu Zheng, and Changyin Yu conceptualized the project and revised the draft. All the authors contributed to the article and agreed to its submission.
Corresponding authors
Ethics declarations
Ethical approval and consent to participate
Not applicable.
Consent to publish
All authors consented to the publication of this review.
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, S., Xie, Y., Yu, C. et al. The battle between host antiviral innate immunity and immune evasion by cytomegalovirus. Cell. Mol. Life Sci. 81, 341 (2024). https://doi.org/10.1007/s00018-024-05369-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-024-05369-y