Abstract
Animal regeneration, the ability to restore a lost body part, is a process that has fascinated scientists for centuries. In this review, we first present what regeneration is and how it relates to development, as well as the widespread and diverse nature of regeneration in animals. Despite this diversity, animal regeneration includes three common mechanistic steps: initiation, induction and activation of progenitors, and morphogenesis. In this review article, we summarize and discuss, from an evolutionary perspective, the recent data obtained for a variety of regeneration models which have allowed to identify key shared mechanisms that control these main steps of animal regeneration. This review also synthesizes the wealth of high-throughput mRNA sequencing data (bulk mRNA-seq) concerning regeneration which have been obtained in recent years, highlighting the major advances in the regeneration field that these studies have revealed. We stress out that, through a comparative approach, these data provide opportunities to further shed light on the evolution of regeneration in animals. Finally, we point out how the use of single-cell mRNA-seq technology and integration with epigenomic approaches may further help researchers to decipher mechanisms controlling regeneration and their evolution in animals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
What is regeneration?
Concept and definition
From the fantastical features of mythic beasts to the routine tasks of less remarkable organisms, the description of species’ restorative properties has only increased our collective fascination for regeneration. Seemingly every civilization has built folklore as well as some basic knowledge on the topic, like the regenerating eye of the ancient Egyptians’ god Horus, or the newt-like regenerative feats of the Aztec deity Xolotl, without forgetting the regenerating liver of the ancient Greeks’ Prometheus [1]. Regeneration has also drawn the attention of great minds and scientific communities throughout history. Starting from antiquity with Aristotle and Pliny the Elder [1], regeneration studies were especially successful with the works of Abraham Trembley on hydras [2] and Lazzaro Spallanzani’s work on salamanders [3], with many important concepts and biological principles discovered in their wake [4, 5]. However, it seems that no one attempted to sketch a clear definition of regeneration until the seminal work of Thomas Hunt Morgan in 1901 [6]. Morgan juxtaposed three different phenomena that could be covered by the notion of regeneration: egg and embryo regeneration, physiological regeneration, and restorative regeneration. These are better known today as “embryonic regulation” (the compensation of cell or tissue loss during embryogenesis), “homeostatic regeneration” (periodic loss and replacement of cells and tissues, such as the gut lining, epidermis, hair and feathers turn over) and “restorative regeneration” (replacement of lost body parts or the entire body), respectively [7, 8]. It is noteworthy that the term “regeneration” is also used for both unicellular organisms and microbial communities, or even ecosystems [9]. We will, however, restrict our discussion to restorative regeneration processes of individual metazoans in this review. Some authors [10] define this phenomenon as “the restoration, the replacement of a lost body part through traumatic injury (either amputation or autotomy)”. It is, however, not a sufficiently precise definition and may raise potential confusion between healing or scarring and regeneration. It also does not properly clarify the overlap of regeneration and asexual reproduction in some species. Furthermore, one of the main disagreements in the domain of regeneration is whether it should be considered as a part of development [8].
Regeneration versus development
For many authors, the striking similarities between embryonic development and regeneration make the case for considering the restorative capabilities found in adults as a part of development, which therefore would need to be extended throughout the life of animals. Yet, there are major differences between these processes that would, by contrast, suggest that embryonic development and regeneration are distinct phenomena with an overlapping use of common cellular and genetic mechanisms. There are, for instance, at least eight restorative regeneration-specific processes found in some or all metazoans [8, 11].
-
1.
Wound healing: the formation of an epithelial barrier against environmental insults necessary for regeneration to take place [12].
-
2.
Immune response: the deployment of the animal immune system at the site of injury seems crucial for normal regeneration in some metazoans [13].
-
3.
Induced reprogramming: resident or distant adult cells—either stem cells or differentiated cells—are activated by the injury and may require unique pathways to recover embryonic-like cell behaviors [11].
-
4.
Transdifferentiation: regeneration depends in some cases on the redifferentiation of dedifferentiated adult cells into different cell types than their original ones [12].
-
5.
System integration: regenerated cells and tissues are integrated and organized in existing differentiated structures, with an emphasis with the continuation of existing systems, such as the vascular and nervous systems [11].
-
6.
Nerve dependency: many regeneration events seem to depend on the activity of the established nervous system [14].
-
7.
Size recognition and termination: positional identity and positional memory in regenerating structures allow for a harmonious growth until an adequate morphology is reconstructed [15].
-
8.
Proliferation of differentiated cells: only during the regeneration of some organs or structures will fully differentiated cells start dividing again [16].
Despite these specificities, it remains unclear whether these regeneration-specific responses could justify their exclusion from bona fide developmental mechanisms. One way to advance the “regeneration versus development” discussion is to rigorously classify the different types of regeneration living organisms can perform.
Different types of regeneration
T.H. Morgan proposed a dualistic grouping of varieties of regeneration based on the requirement of active cell proliferation [6] that to date is still used. At the time regarded as a regeneration expert [17], Morgan compiled a thorough synthesis on earlier regenerative studies, leading him to propose a basic subdivision of regeneration into two general types called epimorphosis and morphallaxis. In epimorphosis, an organism, such as a salamander restoring its limb, regenerates by recruiting dividing cells, whereas morphallaxis refers to the remodeling of existing tissue, exemplified by a regenerating Hydra.
Further precisions and discussion points followed and researchers also frequently try to discriminate blastemal regeneration (where the process requires the apparition of a mass of undifferentiated cells or blastema) from other types of regeneration. Yet some authors warn that blastemas, while they look alike in many species, may have wildly different origins, ontogenically or evolutionarily [8]. It is also possible to differentiate regeneration processes involving pre-existing stem cells (such as in the regenerative prowess of planarians relying on migrating stem cells called neoblasts), dedifferentiating or transdifferentiating cells, and compensatory regeneration involving dividing differentiated cells (displayed during human liver regeneration for instance).
Finally, regeneration processes can also be classified depending on the extent of their restorative power. A hierarchical regeneration ladder can therefore be established depending on the structures that can be restored, starting with cell regeneration, such as axonal regeneration (which we will exclude from the present review), organ (O) or tissue regeneration, such as the restoration of a newt lens, complex structure (CS) regeneration, such as that of limbs, and finally whole-body (WB) regeneration, such as restoration of a full flatworm from a small body fragment.
All these previous types of regeneration are displayed by at least one member of the multicellular organisms in the tree of life [18], especially in the metazoan lineage [10], which will be our sole point of focus during the remainder of this review.
Regeneration at the metazoan scale
Mapping up-to-date data on the regeneration abilities of different metazoan lineages can be a crucial tool to understanding the evolution of regeneration, and discussing whether a single or independent origins of regeneration scenario is more likely [19, 20]. We therefore generated such a tree (Fig. 1) encompassing the main metazoan lineages. Some broad observations can be made, such as the lack of WB regeneration capacity in ecdysozoans and vertebrates, or the almost ubiquitous organ/tissue regeneration ability shared by metazoans (with the notable exception of nematodes, and uncertainties regarding tardigrades, onychophorans, and priapulids due to the scarcity of experimental tests). It is noteworthy that this kind of mapping (Fig. 1) highlights the relatively frequent occurrence of regenerative abilities among metazoans. An experimental bias towards ecdysozoan and vertebrate models may have hindered regenerative research so far, but a recent slew of new and unconventional model systems offers an opportunity to reduce the paucity of our basic knowledge on regeneration [21].

adapted from PhyloPic.org (not to scale) [Illustrators: Hillewaert Keesey, Duygu Özpolat, Michelle Site, Jake Warner, Yan Wong]
Mapping regeneration types on the metazoan phylogenetic tree. Most metazoan phyla include representative species with documented abilities to regenerate organs (O, yellow circles) and complex structures (CS, green circles). While absent (struck-through circles) in all vertebrates, ecdysozoans, and some other scattered lineages, whole-body regeneration (WB, blue circles) is a widespread phenomenon among Metazoa. Lack of substantiated data (gray circles) is mostly observed in Brachiopoda and Ecdysozoa. Phylogenetic tree topology is derived from recent metazoans’ phylogenies [175, 176], with still-controversial lineage positioning depicted with dotted lines. Porifera, Hydra, flatworm, octopus, Platynereis, Drosophila, sea-star, enteropneust, Ciona, zebrafish, and axolotl silhouettes are
Mechanisms of regeneration
Despite the high variability of regenerative types and processes displayed by metazoan species, a careful mechanistic analysis revealed what appears to be a conserved, tri-modular organization of the restorative regeneration of complex structures, such as appendages and the whole body [20, 22] (Fig. 2). These modules are, by chronological order:
-
1.
Wound closure through the formation of an epithelium and general body response to injury
-
2.
Regeneration induction, usually by the recruitment of progenitors and the formation of a blastema
-
3.
Morphogenesis involving patterning, differentiation, and growth
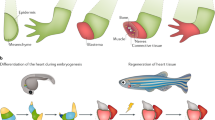
Key common steps in eumetazoan regeneration. a Initiation of regeneration. Within minutes after amputation, reactive oxygen species (ROS) are produced at the wound site. Their accumulation promotes apoptosis, and activates innate immunity. Innate immunity is also activated by apoptotic cells and produces pro-inflammatory cytokines (green squares) that sustain apoptosis. Surrounding cells enter mitosis through apoptosis-induced proliferation (AiP) and the diffusion of mitogenic pro-inflammatory cytokines. b Blastema formation. A blastema, whatever is its origin (migration of resident stem cells or local dedifferentiation) is always composed of undifferentiated mesenchymal cells covered by a wound epidermis. Cell proliferation within the blastema as well as its innervation are crucial for a successful regeneration. c Morphogenesis. Four important aspects of this complex step are depicted: role of innervation, positional memory of the remaining tissue, patterning of the regenerated region in particular in term of axes, and cell differentiation. More details can be found in the main text
Bound to overlap in some instances, these three successive modules are nevertheless distinct in their nature. A thorough exploration of the processes and genetic pathways involved in these modules in diverse animal models might therefore highlight specificities or conserved similarities at the metazoan level. We will review here the common signals and pathways that have been described as crucial for the transition between wound healing and blastema formation, which constitutes the initiation of regeneration per se. Following that, we will discuss the diversity of both the origins and potency of the progenitors populating the blastema. Lastly, we will highlight the mechanisms that allow the regeneration of functional structures across metazoans.
Initiation of regeneration in metazoans
Three major pathways are emerging as key factors for the initiation of regeneration in various animal species: ROS (reactive oxygen species) signaling, apoptosis, and innate immunity (Fig. 2a).
ROS
ROS is an umbrella term used for a wide variety of oxidizers containing oxygen, such as hydrogen peroxide (H2O2) and superoxide anion radical (O2.−), both of which are produced as metabolic by-products [23]. They are the cause of oxidative stress and damage, and can lead to cell death; however, they are also involved in cell signaling during various processes, especially regeneration. In vertebrates, notably during zebrafish fin or Xenopus tadpole tail regeneration, ROS are produced very early at the wound site and their production is sustained after wound closure [24, 25]. In both species, lowering the level of ROS by inhibiting NADPH oxidase (NOX) activity—which is responsible for their production—strongly reduces the size of the regenerated structure, notably by reducing the number of proliferative cells in the blastema. A similar role has been demonstrated during planarian regeneration, as inhibiting ROS production affects stem cell early differentiation, resulting in an improper regeneration and shortening of the blastema [26]. The importance of ROS has been also acknowledged during Drosophila imaginal disc regeneration, where they activate JNK and p38 signaling pathways which in turn trigger the JAK/STAT pathway required for regeneration [27]. Finally, in the cnidarian Hydra, ROS are produced at the wound site briefly after amputation and might therefore be required for proper regeneration in this species as well [28].
Cell death
ROS are known to trigger cell death [29], especially by apoptosis, and there is now clear evidence that apoptosis promotes regeneration through the so-called apoptosis-induced compensatory proliferation (AiP) [30]. During Xenopus tadpole tail regeneration, apoptosis does not follow amputation immediately, but triggers cell proliferation later on [31]. Two rounds of apoptosis happen quite early during zebrafish fin regeneration; the second round co-occurs with the peak of ROS production and triggers subsequent cell proliferation which is crucial for blastema formation [24]. In both species, blocking apoptosis at early time points after amputation impairs regeneration. In planarians, two rounds of apoptosis happen as well, co-occurring with increased cell divisions: the first happens early after amputation and is restricted to the wound region, while the second spreads throughout the organism and happens later during regeneration [32]. While apoptosis is required for tissue remodeling during planarian regeneration [33], a causal link between apoptosis and cell division has not been demonstrated. Lastly, Hydra head regeneration and the sea anemone Nematostella vectensis oral regeneration provide another example of AiP, within cnidarians. In Nematostella, a burst of apoptotic activity appears at the cut site shortly after amputation. Pan-caspase inhibitors hamper both apoptosis and proliferation which subsequently block regeneration [34]. In Hydra, there is an early round of apoptosis followed by cell proliferation induced by the diffusion of Wnt molecules from apoptotic bodies [35]. Wnt-containing apoptotic bodies are also found in regenerating zebrafish epidermis and shown to promote proliferation of adjacent stem cells through Wnt signaling [36]. Thus, the chain of events, starting with ROS production which triggers apoptosis and in turn leads to cell proliferation, is emerging as a widely distributed, and possibly conserved, key pathway involved in regeneration initiation in Eumetazoa (Cnidaria plus Bilateria).
Innate immunity
Innate immunity appears to have an important role during the first step of regeneration, especially in vertebrates. In regenerating zebrafish fins and salamander limbs, both neutrophils and macrophages populate the wound region quickly after amputation, and the depletion of macrophages shortly after amputation tilts the balance in favor of inflammation, which results in a significant reduction of blastema size and impaired regeneration [37]. Yet, pro-inflammatory cytokines are secreted after amputation as well [13] and were shown to be important for early regeneration: knockdown (KD) of interleukin-8 (IL-8; also known as CXCL8) prevents the recruitment of neutrophils at the wound site and the normal expression of inflammatory genes during the first phases of zebrafish fin regeneration [38]. IL-8 KD delays regeneration and leads to defective recruitment of monocytes and granulocytes during salamander limb regeneration [39]. The implication of innate immunity during regeneration in other phyla is less obvious as the available data are scarcer, but growing evidence hints at a similar importance. In both planarians and Hydra, mRNA-seq data for regenerative tissues show an up-regulation of numerous genes annotated as innate immunity-related, notably lectins, metalloproteinases, and putative members of Toll-like Receptor (TLR) signaling [40, 41].
It is now well-established that ROS signaling, apoptosis, and immune response to injury play major roles in the first steps of regeneration in many animals. ROS production is the earliest signal, being detectable in some species within minutes after the injury, and could serve as a wound sensor. Apoptosis and immune response to injury are shown to start acting slightly later to control activation of progenitors and correct patterning of the reformed tissues. It must be noted that those three pathways are extremely intricate and can trigger each other easily, in any direction [29, 42, 43].
Recruitment and activation of progenitors
We will describe two main aspects of this process, the source or origin of the pool of undifferentiated cells that constitute the blastema, and the fate and potency of these blastemal cells (Fig. 2b).
Origin of blastemal cells
In many animals, the blastema is formed by migratory cells that are recruited upon injury. The best and prime instance of such is WB regeneration in planarians. The source of their blastemal cells has triggered a long scientific controversy [44], and it is now fully confirmed that those cells arise from undifferentiated resident stem cells called neoblasts. Indeed, it has been shown in the planarian Schmidtea mediterranea that upon amputation of the anterior-most part of the worm, neoblasts rapidly migrate to the wound and form the blastema [45]. The identification of the distant origin of planarian blastemal cells in turn triggered the search for migratory cells involved in regeneration in other animals. In annelids, such migratory cells were observed in a regeneration context in three species, either through an indirect cell lineage approach by S-phase labelling in Enchytraeus japonensis [46] and Capitella teleta [47], or using long-term live imaging in Pristina leidyi [48].
Among deuterostomes, regeneration in the colonial ascidians Botrylloides leachi requires the activation of non-circulatory “inner cells” lining the epithelium of blood vessels [49]. Upon injury, those cells detach from the epithelium and migrate towards the wound where they form aggregates [49]. Similarly, in the solitary ascidian Ciona intestinalis, precursors of the regenerating distal oral siphon originate from a proximal organ, the branchial sac, whose cells migrate upon injury towards the wound [50]. There are examples of cell migration in vertebrate’s regeneration as well [51]. During zebrafish fin regeneration, the tracking of Di-I-labelled cells proved that migration can occur from distant epithelial and connective tissues towards the blastema in a short time window (up to 2 days after amputation) [52]. In particular, it was shown that, upon amputation, osteoblast progenitor cells migrate from their niche in the joints and contribute significantly to the regenerated bones [53]. Migration of regenerative cells also happens in non-bilaterian species. In Hydra, in vivo tracing of Green Fluorescent Protein-producing (GFP +) interstitial stem cells grafted onto wild-type (WT) animals showed that those cells migrate upon amputation towards the wound and replace the lost tissues and stem cells [54]. In Hydractinia echinata, both direct and indirect cell tracking techniques showed that cells forming the head blastema originate from more posterior tissues in the body column [55]. Migration is thus the primary source of regenerative cells in several organisms scattered across eumetazoans, which advocates for the potent ancestrality and conservation of this mechanism in various lineages of regenerative animals. Yet, convergence cannot be excluded.
As such, regeneration does not always rely on migratory cells. During urodele limb regeneration, blastema formation does not depend on distant cells but rather on dedifferentiation of local mature tissues abutting the amputation plane [56]. For instance, it was shown in transgenic lines of adult newts in which myofibers were labelled with fluorescent markers that myocytes do dedifferentiate during regeneration and proliferate within the blastema [57]. Likewise, grafting experiments showed that nerve cell dedifferentiation contributes to blastema formation in axolotls [58]. During zebrafish fin regeneration, labelling of mature fibroblasts and osteoblasts showed that both cell types dedifferentiate and actively populate the blastema, as well as the newly regenerated tissues [59]. Similarly in mice, which have very limited regenerative abilities but are still able to regenerate their digit tip after amputation, axons close to the wound degenerate, which triggers the dedifferentiation of their associated Schwann cells [60]. Those cells participate in the blastema formation, and are crucial for its proper growth [60]. In the annelid Platynereis dumerilii, indirect evidence by S-phase labelling also suggests that posterior body regeneration mainly occurs through local dedifferentiation [61]. Histological analyses also support the presence of dedifferentiation during regeneration in various species, such as the amphioxus Branchiostoma lanceolatum [62], some echinoderms [63, 64] and the jellyfish Polyorchis penicillatus [65]. Lastly, the blastema can also originate from the activation of local resident stem cells. In Xenopus tadpoles and zebrafish larvae, muscle satellite cells close to the wound are activated and populate the blastema [66]. Similarly, satellite cells participate in regeneration in axolotls [57] and larval newts [67]. In zebrafish, both dedifferentiated osteoblasts and undifferentiated cells, the osteoblast progenitors contribute to bone regeneration [51, 53]. In mice digit tips, most of the blastema is formed thanks to activation of several types of resident stem/progenitor cells [68]. Finally, outside vertebrates, during leg regeneration of the crustacean Parhyale hawaiensis, the blastema is partly formed by normally quiescent cells located in the muscles, resembling vertebrate satellite cells [69].
Fate and potency of blastemal cells
A second important aspect is to define the identities of blastemal cells, in particular whether a blastema contains pluripotent and/or tissue-specific (multipotent or unipotent) stem/progenitor cells. As advanced techniques of cell tracking and fate mapping are required for these identities to be properly investigated, data are still scarce. However, two opposite models are emerging. It was hypothesized for a long time that morphologically indistinct blastemal cells constitute a pool of multi/pluripotent progenitors [70]. This assertion was properly proven in the planarian Schmidtea mediterranea in which a subset of neoblasts called ‘cNeoblasts’ can restore regenerative abilities when injected in small numbers into lethally irradiated hosts [71]. A single cNeoblast can even replenish an entire irradiated animal. Acoels worms (Acoelomorpha) harbor neoblasts as well, even though it is still not clear whether they correspond to a homologous cell type and have the exact same properties [72, 73]. This led to the hypothesis that the involvement of multi/pluripotent stem cells during regeneration could be conserved across bilaterians [74]. As such, in the ascidian Ciona intestinalis, regeneration occurs from a unique stem cell niche, the branchial sac, whose progenitors can give rise to both muscle and nervous tissues [75]. Outside bilaterians, the existence in Hydra of multipotent stem cells, the interstitial cells or i-cells, was well-known for a long time [76] and it has been shown that those stem cells could migrate upon injury and participate in regeneration [54]. Similarly, in Hydractinia, regeneration of the head of a polyp involves the migration of multipotent i-cells [55].
In contrast, in vertebrates, amphibians, zebrafish, or mammals, blastemal cells are lineage-restricted stem/progenitor cells. In Xenopus, grafts of GFP + tissues onto WT animals proved that during tadpole tail regeneration, the spinal cord and notochord are regenerated by their own remnants which dedifferentiate and proliferate at the amputation stump, while muscles are reformed only through the activation of satellite cells [77]. Similar grafting experiments, recently supported by data obtained with CRIPSR/Cas9 technology, led to the same conclusions in axolotls in which cartilage, muscle, or nervous cells are unable to switch to another lineage during regeneration [58, 78]. The conservation of cell lineages during regeneration is also documented for zebrafish fin regeneration, with a strict separation of fate between osteoblasts, fibroblasts and epidermal cells [79]. Likewise, cell tracing in mice showed a strict restriction of fate during digit tip regeneration between germ layers, and that bone and cartilage on the one hand, and tendons and vasculature on the other hand do not transdifferentiate into one another [68]. Thus, strict lineage restriction during regeneration appears to be a conserved feature among vertebrates. It must however still be noted that there are examples in which mesodermal progenitors, during regeneration, can expand their progeny compared to the one produced in homeostatic conditions [80, 81].
Redifferentiation of the progenitors and morphogenesis
Another major question is how differentiation of blastemal cells is controlled, especially in terms of size and patterning of regenerated structures, to produce a fully functional structure which usually displays shape, size and function in accordance with its position in the body (e.g., along anterior–posterior and dorsal–ventral axes), and similar to the one that was amputated. Two main points will be discussed here: how the amputated tissues keep in memory what the structure to be regenerated is, as well as the importance of nervous tissues and reinnervation for proper regeneration (Fig. 2c).
Positional memory during regeneration in bilaterians: role of mesodermal tissues
Planarians, given their outstanding regenerative abilities, are particularly relevant models to study how positional information is encoded, said information being required for the blastema to regenerate the proper structure depending on its position along the body of the animal. Witchley and colleagues [82] searched for genes exhibiting a regionalized expression pattern that they refer to as Position Control Genes (PCG). Strikingly, it appears that expression of these genes is almost entirely restricted to the muscles of the body wall. Upon any kind of amputation, their overall pattern is rapidly re-established in the amputated fragments, which allows for the correct specialization of the activated neoblasts forming the blastema [82]. A similar expression of PCG and similar role of muscles have been shown during Hofstenia regeneration [83]. In vertebrates, positional information seems to be encoded in mesodermal tissues as well. In axolotl, grafts of GFP + tissues into WT animals showed that cartilage-derived cells possess proximo-distal identity [58]. Intercalation assays where distal blastemas are grafted upon proximal stumps confirmed that connective tissue cells contain proximo-distal information that allows proper regeneration of the limb [84]. Zebrafish posterior fins are also able to maintain their shape through regeneration with longer peripheral fin rays and shorter central fin rays. Fin ray grafts that are displaced along the proximo-distal axis retain memory of their origin and will not regenerate according to their new position [85]. Given that epidermal tissues of the graft are quickly replaced by the host epidermis, this memory is most likely also carried by the mesoderm [85].
Importance of innervation
Although proper functional and mechanistic evidence is still scarce in most animals, nerve-dependent regeneration appears to be shared among metazoans (reviewed in [14]). The role of innervation during regeneration was particularly highlighted in the salamander limb in which denervation impedes a blastema to grow and differentiate [86] and rerouting a peripheral nerve underneath an epidermal wound triggers the formation of an ectopic limb [87]. Re-innervation induces a gradient of the growth factor nAG, a ligand for the receptor Prod1 which is crucial for the establishment of the proximo-distal axis during regeneration [88]. Furthermore, it was shown that nerves induce the up-regulation of a histone deacetylase (HDAC) in the blastema just before the differentiation stages. HDAC expression could modify the epigenetic landscape of the blastema and allows its differentiation as the inhibition of HDAC activity results in a severely delayed regeneration [89]. In planarians, nerves were shown to be crucial for the patterning of regenerative parts along with cell–cell communication through the gap junctions. Severing the ventral nerve cord and blocking gap junctions lead to the formation of ectopic heads in place of tails [90]. More recently, modeling of planarian regeneration suggested that the diffusion of small molecules alone cannot explain the quick and stereotypical reestablishment of the PCG pattern in body fragments of various sizes [91]. The authors of this study proposed that nerves may provide a scaffold for the directional transport of morphogens, and therefore that initial nerve polarity would determine the antero-posterior axis of regenerating fragments.
As for development, many entangled developmental genes and signaling pathways exert crucial functions in the later stages of regeneration, i.e. the patterning and morphogenesis of the regenerated structures. In the next section, we review how genome-wide transcriptomic analyses allowed to identify such genes and pathways involved in regeneration in diverse animals and how these approaches may help to better understand the evolution of regeneration in animals.
Insights from regeneration bulk transcriptomic data
While having been intensively studied morphologically during the first part of the twentieth century, animal regeneration withdrew from developmental biology’s center stage with the rise of genetic and molecular developmental studies in the 70s [10, 92]. Indeed, the main developmental biology models that emerged at that time (i.e. Drosophila, C. elegans, Mus musculus), while having been instrumental in providing insights into many biological questions, were of limited interest to study restorative regeneration due to their restricted regenerative abilities (Fig. 1) [10]. Conversely, the majority of key regeneration model species were not easily amenable for functional analysis [93]. We are, however, currently witnessing a strong revival of interest from the scientific community for regeneration [92, 93]. New mechanistic studies of regeneration are mainly fostered by technical advances, notably high-throughput sequencing technologies that are currently widely and successfully applied to less conventional regeneration models.
Bulk mRNA-seq data uncover the transcriptional profiles of various types of regeneration in many non-model species throughout metazoans
High-throughput mRNA sequencing, commonly called bulk mRNA-seq, is a popular technology used nowadays to monitor changes in gene expression that accompany complex and dynamic biological processes [94]. This unbiased approach provides a timed in-depth overall view of the transcriptomic landscape of cells participating in regeneration. By estimating and comparing relative gene expression levels with a high degree of accuracy, identifying thousands of differentially expressed genes that may play crucial roles during regeneration steps is now a fairly straightforward process. In the last ten years, a huge amount of mRNA-seq data was gathered, with more than 100 studies on species (around 50) belonging to almost all metazoan lineages with regenerative abilities (Figs. 1 and 3, Supplementary Table 1). Unsurprisingly, the main bilaterian model systems for regeneration, namely flatworms (Platyhelminthes [95]), salamanders and Xenopus (Lissamphibia [96]), and the zebrafish (Actinopterygii [97]), encompass the majority of bulk mRNA-seq studies (Fig. 3, Supp. Table 1). In addition, many studies investigated regeneration in Mammalia, although their regenerative potential is more limited, driven by possible applications for regenerative medicine [98]. In contrast, while non-bilaterian lineages, namely Porifera [99], Placozoa, Ctenophora, and Cnidaria, harbor extensive regeneration capabilities [10], very few studies have investigated their transcriptomic changes during regeneration, with the noticeable exceptions of Cnidaria [100, 101]. In terms of phylogenetic distribution, information from important regenerative groups, i.e., Ctenophora [102], Nemertea [103], Cephalochordata [62], and Mollusca [104] (as well as more under-looked ones: Brachiopoda, Phoronida, Ectoprocta, Gastotricha, Entoprocta, Chaetognatha, Rotifera, Priapulida, Xenoturbellida [10]) is still missing in current existing datasets.
For the other main clades (Acoelomorpha, Annelida, Arthropoda, Hemichordata, Tunicata, Mammalia), data come mostly from one or two species, which prevents any generalization to the entire group. A single species cannot be considered as representative of its whole lineage, especially concerning a character as labile as regeneration. Regeneration has a rather rich evolutionary history, as species-specific innovations are frequent among closely related species often exhibiting very different regenerative abilities [12, 103, 105]. In contrast, in Platyhelminthes, up to seven species displaying various regeneration features have been molecularly studied, uncovering the regulatory network at the origin of head regeneration (see below) [106,107,108,109,110,111]. So far, our knowledge of changes in transcription during regeneration concerns more than 50 species and various types of regeneration, including whole-body regeneration of Cnidaria, Acoelomorpha, Tunicata, and Planaria, regeneration of complex structures, such as body axis (Annelida, Hemichordata), appendages (Actinopterygii, Lissamphibia and Arthropoda), and organ regeneration (Vertebrata) (Fig. 1, Supp. Table1).
The overwhelming majority of these studies has been performed on adult animals, with only a couple of studies investigating larval stages (in sea stars [112]), juvenile (Xenopus tadpoles [113]) and newborn (rats [114] and mice [115]), which leaves a gap in our greater comprehension of regeneration processes. The majority of available data are series of mRNA-seq data at different time points after an amputation or a wound, which cover the three main aforementioned steps of regeneration and therefore allow us to finely highlight the dynamics of the process. In a couple of earlier studies [116, 117], the data provided were restricted to one specific regenerative stage (usually a blastema stage) or time point, and are consequently much less informative. Altogether, these bulk mRNA-seq studies made significant strides toward our understanding of regeneration and have given us the opportunity to further shed light on its evolution using a comparative approach [98, 112, 118].
Major advances in the regeneration field revealed by bulk mRNA-seq
New questions on regeneration can be addressed thanks to bulk mRNA-seq data and their comparative analyses. Four of those major topics will be discussed below.
Regeneration versus development
Whether adult injury-induced regeneration is a post-embryonic developmental process or a phenomenon distinct from development is a long-standing, unresolved scientific question. On the one hand, regeneration has its proper evolutionary history and contains specific features not found during development [8]. On the other hand, restoring complex tissue structures upon injury involves the reactivation of developmental processes that are specific to the regenerated structure and that have to remain available in adult stages [119]. Transcriptomic data allowed to address this question by defining, in a couple of species, to which extent regeneration and developmental programs share molecular commonalities and present specificities. A recent study on the cnidarian Nematostella vectensis (sea anemone) sought to answer this question for the first time on a greater scale, thanks to the comparison of a large time series of mRNA-seq data for development and regeneration [34, 101]. This study revealed that regeneration uses a partial and rewired embryonic gene regulatory network (GRN), with regeneration‐specific modules driving cellular events unique to regeneration, such as apoptosis [34]. Along the same line, another recent study investigated the process of skeletogenesis in the echinoderm Amphiura filiformis (brittle star) during both embryogenesis and arm regeneration [120]. This analysis focused mainly on the role of the FGF signaling pathway during skeletogenesis and supports the hypothesis that regeneration recapitulates development in this species. While these seminal analyses pointed out similarities between development and regeneration in both the investigated species, more studies are required to fully address the question of the relationships between development and regeneration in Metazoa.
Patterning
Like an embryo, a regenerating body region must be meticulously patterned to ensure that only the appropriate structures are formed, at the proper location, and with the correct size and shape [7]. Patterning during regeneration is regulated by positional memory, a cellular property that involves adult cells spared from injury and maintains necessary information for structures’ replacement [121]. Positional memory has been studied mainly in the context of appendages and main body axis regeneration. Comparison of tissue transcriptomes at different anatomical locations was performed during appendage regeneration in both zebrafish [121] and axolotls [122]. This enabled the identification of several candidate transcripts involved in positional memory along the proximo-distal axis of the limb (such as the RNA-binding protein cirbp, which plays a cytoprotective role, and kazald1, whose knockout in blastema impairs regeneration) and caudal fin (such as transcription factors of the dlx and msx families, known to be involved in the patterning of developing appendages), paving the way for future mechanistic studies [121, 122]. Which programs drive anterior/posterior (AP) regeneration in bilaterians, but also apical/basal (or oral/aboral) regeneration in cnidarians, is a prominent question in the field. In Hydra, while a similar initial transcriptional response is observed early after amputation on either side of the oral/aboral axis, dramatically different programs are set up later on, involving BMP signaling on the basal part and Wnt-β-catenin pathway on the apical side [123]. The important role of the latter during apical regeneration has been described in both Hydra and Hydractinia in which knockdown (KD) of Wnt3 impedes regeneration upon decapitation [35, 124]. Furthermore, incubation of Wnt3 protein induces the formation of ectopic “heads” on body columns [125]. Similarly, in Nematostella, Wnt-β-catenin members are highly expressed in the oral pole compared to the aboral one [126]. Strikingly, the Wnt-β-catenin pathway is also a central regulator of axial polarity during regeneration of a variety of bilaterian species. In the larva of the sea star Patiria miniata, Wnt ligands and Frizzled receptors are up-regulated posteriorly during regeneration, suggesting that Wnt-β-catenin may have a role in AP patterning [112]. In the annelid Aeolosoma viride, inhibition or overactivation of the Wnt-β-catenin pathway impairs head regeneration [127]. In the planarian Schmidtea, Wnt-β-catenin activation specifies posterior regeneration while its inhibition (through NOTUM) specifies anterior regeneration [107, 128]. KD of β-catenin results in the regeneration of a head in place of a tail [128], and KD of notum in the opposite outcome [129]. Similarly, in the Acoelomorpha Hofstenia, Wnt-β-catenin is activated during posterior regeneration [130, 131]: KD of Wnt-β-catenin positive mediators results in anterior-like structures in place of the tail, and KD of Wnt inhibitors in tail structures in place of the head [73]. A gene regulatory network for posterior regeneration initiation has recently been described, with a Wnt ligand (Wnt-3) being transcriptionally regulated by an early wound response factor (Egr) [131]. The Wnt-β-catenin pathway is also responsible for other crucial functions in later stages of regeneration. In axolotls, its inhibition prevents normal regeneration and results in the formation of a spiky outgrowth [132], while its global activation results in malformations of regenerated skeletal tissues [133]. Similarly, in zebrafish fin regeneration, inhibition of Wnt-β-catenin signaling early after amputation prevents blastema formation [132], while later on during blastema differentiation, its inhibition hinders proper bone calcification [134]. During mouse digit tip regeneration, Wnt-β-catenin pathway activation in the epidermal progenitors is crucial for the differentiation of the regenerated nail, but also for blastema growth and overall regeneration [135]. Besides highlighting the role of the Wnt-β-catenin in regeneration success, the comparison of head and tail regeneration transcriptomes in Schmidtea led to another important and unanticipated observation. While their transcriptome profiles were initially very different, they later converged to a shared core regenerative program [107]. In stunning contrast, in the Syllidae Sphaerosyllis hystrix and Syllis gracilis (annelid worms), anterior and posterior regenerations do not rely on a common transcriptomic landscape: the posterior regeneration program is indeed more related to the one used during posterior growth (growth by addition of segments in the posterior body part of uninjured juvenile animals) than to the head regeneration program [136].
Variations in regenerative capabilities
Regenerative capacity varies greatly across animals but also across the life cycle of a given species [10]. Why do certain animals possess substantial regenerative capacities (at least during a specific period of their life cycle), while others lack such amazing abilities? Answering this crucial question is fundamental to implementing medical strategies aiming to unlock the regenerative potential of humans. To start answering this question, researchers have exploited the vast richness of regeneration patterns in animals. Xenopus is an amphibian species that does not display life-long regenerative capacity, as this ability is restricted to the pre-metamorphic larval stages (with the exception of the retina) [137]. Lee-Liu and colleagues investigated Xenopus spinal cord regeneration during its regenerative and non-regenerative stages, revealing differences in the timing of the transcriptional response and in the inventory of regulated transcripts involved in broad biological processes including neurogenesis, metabolism, immune response and inflammation, cell cycle, development, and response to stress [113]. The axolotl Ambystoma mexicanum is another amphibian which can regenerate its appendages in a nerve-dependent manner: regeneration does not occur in denervated limbs [96]. Interestingly, repeated limb amputations lead to regeneration defects and failure [138]. Two studies explored the transcriptomic landscape of those impaired regenerations in comparison to normal regenerating limbs, laying down a detailed blueprint of mis-regulated genes hindering regeneration, such as amphiregulin, an EGF-like ligand [138, 139]. Similarly, comparison between regenerating tails versus non-regenerating limbs were performed in the lizard Podarcis muralis, revealing the importance of small nucleolar RNAs and Wnt signaling pathways for tail regeneration [140]. In planarians, while some species, such as Schmidtea, are able to reform a full animal from a single pluripotent stem cell [71], many others lack (or have limited) regenerative abilities and notably are unable to regenerate their anterior part upon amputation [106, 110]. Procotyla fluviatilis and Dendrocoelum lacteum, for example, fail to regenerate their head if amputated too posteriorly along their body axis. Comparative transcriptomic analysis between regeneration-competent and non-competent tissues revealed that the Wnt signaling pathway is aberrantly activated in non-competent tissues and that downregulation of this pathway is sufficient to restore head regeneration from regeneration-non-competent tissues [106, 110]. These two seminal papers revealed that manipulating a single signaling pathway can be sufficient to reverse the evolutionary loss of regenerative potential in planarians.
Evolutionary history of regeneration
The origin and evolution of regeneration in animals is a long-standing debate, and while several ecological reasons and evolutionary hypotheses have been proposed, why and how regeneration abilities evolved remain a mystery [10]. Nevertheless, as regeneration is found in species that belong to all of the main branches of the metazoan tree, the possibility that this ability is an ancestral feature of animals and might therefore rely on homologous mechanisms and genetic networks is a tempting hypothesis that requires careful examination [21]. A broad comparative approach, using the tremendously increasing quantity of mRNA-seq data obtained in the recent years for a large spectrum of metazoan species, will be a compelling strategy to address this question. A couple of attempts have already been made, leading to a patchy inventory of similar, homologous, or co-opted components of regeneration networks in distantly related species [112, 118, 126]. Hence, comparisons between Hydra, Schmidtea, and Patiria whole-body regeneration revealed the common involvement, during the immediate-early phase of regeneration, of cell death/apoptosis-related genes, MAPK signaling-associated genes (such as Jnk), and the transcription factor-encoding gene Egr. At later stages of regeneration, the Wnt signaling pathway is mandatory to specify the axis in those three species before the occurrence of a massive cell proliferation event which is associated with the expression of different genes. Another computational and broad comparison between Hydra, Schmidtea and the echinoderm Apostichopus japonicus (sea cucumber) identified 18 common differentially expressed genes with functions related to metabolic processes and signaling pathways, such as Wnt and cadherin [118]. Our own analysis of the extensive bibliography related to bulk mRNA-seq data during regeneration (Supp. Table 1) also points out several major signaling pathway components in addition to the Wnt-β-catenin pathway already mentioned, notably Jak/STAT [113, 141, 142], Notch [126, 143,144,145], MAPK [108, 142, 143, 146], and FGF/FGFR signaling [120, 123, 147, 148], which are dynamically expressed in various regeneration contexts and steps (Fig. 4). This analysis also unveils the importance of less-studied regulators, such as those linked to epigenetic modifications [112, 143, 149], non-coding RNAs [146, 147], and importantly many unknown novel or species-specific regeneration genes [107, 113, 122, 141, 147, 150].
Majors signaling pathways’ components are dynamically expressed in various metazoan regeneration contexts. Transcriptomic data highlight the potential importance of 13 major signaling pathways during regeneration of 15 main metazoan lineages. Circles indicate that at least one transcriptomic study reports differential expression of those pathway components. Cnidaria—A Anthozoa; Cnidaria—M Medusozoa; Echinodermata—H Holothuroidea; Echinodermata—O Ophiuroidea; Echinodermata—A Asteroidea
In summary, we outlined in this section that a comparative evolutionary study of regeneration is a powerful strategy to tackle key regeneration-linked questions. There is clearly much more to learn from the wealth of information gathered so far from mRNA-seq analyses, and this will undoubtedly be pursued in the next coming years.
Perspectives
While bulk transcriptomics has duly demonstrated its usefulness to identify genes and pathways involved in complex phenomena, such as development or regeneration, it is not applicable for the identification and characterization of cellular states nor for the comprehension of how those states change over the course of such processes [151]. Bulk analysis eliminates crucial information by averaging signals from individual cells and does not allow to discriminate between changes due to gene regulation and those due to modifications in cell type composition. In recent years, NGS-based technologies for transcriptomics have been exploring a new direction for characterizing individual cells, i.e., single-cell RNA sequencing (scRNA-seq) (e.g., [151, 152]). Methods have been developed to use scRNA-seq to characterize cell populations and track distinct cell lineages during embryonic development (e.g., [153, 154]). These approaches have also been used to understand regeneration processes in different species across the animal kingdom, including WB in the planarian Schmidtea [155,156,157,158,159,160], anterior body regeneration in the earthworm Eisenia [161], tail regeneration in Xenopus [162, 163], limb regeneration in the axolotl Ambystoma [164,165,166], fin and heart regeneration in zebrafish [167,168,169], and digit regeneration in mice [170,171,172,173].
These studies revealed the diversity of cell populations during regeneration and unveiled previously unrecognized cell types in the blastema, such as multipotent mesenchymal-like progenitors producing various connective tissue lineages during axolotl limb regeneration [164, 165], regeneration-organizing cells which belong to the wound epidermis and act as a signaling center during Xenopus tail regeneration [162], or rare and transient cell types required for WB in Schmidtea [160]. Another very promising path to further unravel regeneration mechanisms is to combine mRNA-seq (both bulk and scRNA-seq) with epigenomic approaches. In a recently published seminal study, bulk mRNA-seq, scRNA-seq, and chromatin immunoprecipitation sequencing (ChIP-seq) were used to compare regeneration in two distantly related teleosts, zebrafish and African killifish Nothobranchius. This study revealed an evolutionarily conserved regeneration response program involving specific regeneration-responsive enhancers, some of which may have been repurposed for other functions in vertebrates with poor regeneration abilities, such as mammals [168]. Application of these transcriptomic and epigenomic approaches, and more generally of multi-omics approaches [174] to an increasing number of regeneration models will undoubtedly help decipher fundamental regeneration mechanisms and their evolution in animals.
References
Elliott SA, Alvarado AS (2018) Planarians and the history of animal regeneration: paradigm shifts and key concepts in biology. Methods Mol Biol 1774:207–239
Trembley A et al (1744) Mémoires pour servir à l’histoire d’un genre de polypes d’eau douce, à bras en forme de cornes. Chez Jean & Herman Verbeek, Leyde
Spallanzani L (1768) Prodromo di un’opera da imprimersi sopra le riproduzioni animali dato in luce da Spallanzani. Montanari, Modena
Dinsmore CE (1991) A history of regeneration research: milestones in the evolution of a science. Cambridge University Press, Cambridge
Keller J (1894) Die ungeschlechtliche Fortpflanzung der Süsswasserturbellarien. Jen Zeit Naturw 94:3823–3827
Morgan TH (1901) Regeneration. Macmillan, New York
Poss KD (2010) Advances in understanding tissue regenerative capacity and mechanisms in animals. Nat Rev Genet 11(10):710–722
Vervoort M (2011) Regeneration and development in animals. Biol Theory 6(1):25–35
MacCord K, Maienschein J (2019) Understanding regeneration at different scales. Elife 8:e46569. https://doi.org/10.7554/eLife.46569
Bely AE, Nyberg KG (2010) Evolution of animal regeneration: re-emergence of a field. Trends Ecol Evol 25(3):161–170
Barresi MJF, Gilbert SF (2020) Developmental biology. Oxford University Press, Oxford
Brockes JP, Kumar A (2008) Comparative aspects of animal regeneration. Annu Rev Cell Dev Biol 24:525–549
Godwin JW, Pinto AR, Rosenthal NA (2013) Macrophages are required for adult salamander limb regeneration. Proc Natl Acad Sci U S A 110(23):9415–9420
Sinigaglia C, Averof M (2019) The multifaceted role of nerves in animal regeneration. Curr Opin Genet Dev 57:98–105
Uemoto T, Abe G, Tamura K (2020) Regrowth of zebrafish caudal fin regeneration is determined by the amputated length. Sci Rep 10(1):649
Sanchez Alvarado A, Tsonis PA (2006) Bridging the regeneration gap: genetic insights from diverse animal models. Nat Rev Genet 7(11):873–884
Sunderland ME (2010) Regeneration: Thomas Hunt Morgan’s window into development. J Hist Biol 43(2):325–361
Coffman JA (2019) Regenerative potential across species: an eco-evo-devo perspective. In: Palacios D (ed) Epigenetics and regeneration. Academic Press, Elsevier, pp 197–214
Slack JM (2017) Animal regeneration: ancestral character or evolutionary novelty? EMBO Rep 18(9):1497–1508
Tiozzo S, Copley RR (2015) Reconsidering regeneration in metazoans: an evo-devo approach. Front Ecol Evol 3:67. https://doi.org/10.3389/fevo.2015.00067
Lai AG, Aboobaker AA (2018) EvoRegen in animals: time to uncover deep conservation or convergence of adult stem cell evolution and regenerative processes. Dev Biol 433(2):118–131
Galliot B, Chera S (2010) The Hydra model: disclosing an apoptosis-driven generator of Wnt-based regeneration. Trends Cell Biol 20(9):514–523
Sies H, Jones DP (2020) Reactive oxygen species (ROS) as pleiotropic physiological signalling agents. Nat Rev Mol Cell Biol 21(7):363–383
Gauron C et al (2013) Sustained production of ROS triggers compensatory proliferation and is required for regeneration to proceed. Sci Rep 3:2084
Love NR et al (2013) Amputation-induced reactive oxygen species are required for successful Xenopus tadpole tail regeneration. Nat Cell Biol 15(2):222–228
Pirotte N et al (2015) Reactive oxygen species in planarian regeneration: an upstream necessity for correct patterning and brain formation. Oxid Med Cell Longev. https://doi.org/10.1155/2015/392476
Santabarbara-Ruiz P et al (2015) ROS-induced JNK and p38 signaling is required for unpaired cytokine activation during drosophila regeneration. PLoS Genet 11(10):e1005595
Vriz S, Reiter S, Galliot B (2014) Cell death: a program to regenerate. Curr Top Dev Biol 108:121–151
Mittal M et al (2014) Reactive oxygen species in inflammation and tissue injury. Antioxid Redox Signal 20(7):1126–1167
Fan Y, Bergmann A (2008) Apoptosis-induced compensatory proliferation. The Cell is dead. Long live the Cell! Trends Cell Biol 18(10):467–473
Tseng AS et al (2007) Apoptosis is required during early stages of tail regeneration in Xenopus laevis. Dev Biol 301(1):62–69
Pellettieri J et al (2010) Cell death and tissue remodeling in planarian regeneration. Dev Biol 338(1):76–85
Beane WS et al (2013) Bioelectric signaling regulates head and organ size during planarian regeneration. Development 140(2):313–322
Warner JF et al (2019) Regeneration is a partial redeployment of the embryonic gene network. BioRxiv. https://doi.org/10.1101/658930
Chera S et al (2009) Apoptotic cells provide an unexpected source of Wnt3 signaling to drive hydra head regeneration. Dev Cell 17(2):279–289
Brock CK et al (2019) Stem cell proliferation is induced by apoptotic bodies from dying cells during epithelial tissue maintenance. Nat Commun 10(1):1044
Petrie TA et al (2014) Macrophages modulate adult zebrafish tail fin regeneration. Development 141(13):2581–2591
de Oliveira S et al (2013) Cxcl8 (IL-8) mediates neutrophil recruitment and behavior in the zebrafish inflammatory response. J Immunol 190(8):4349–4359
Tsai SL, Baselga-Garriga C, Melton DA (2019) Blastemal progenitors modulate immune signaling during early limb regeneration. Development 146(1):dev169128. https://doi.org/10.1242/dev.169128
Peiris TH, Hoyer KK, Oviedo NJ (2014) Innate immune system and tissue regeneration in planarians: an area ripe for exploration. Semin Immunol 26(4):295–302
Wenger Y et al (2014) Injury-induced immune responses in Hydra. Semin Immunol 26(4):277–294
Yang Y et al (2015) Programmed cell death and its role in inflammation. Mil Med Res 2:12
Fogarty CE, Bergmann A (2017) Killers creating new life: caspases drive apoptosis-induced proliferation in tissue repair and disease. Cell Death Differ 24(8):1390–1400
Baguna J (2012) The planarian neoblast: the rambling history of its origin and some current black boxes. Int J Dev Biol 56(1–3):19–37
Wenemoser D, Reddien PW (2010) Planarian regeneration involves distinct stem cell responses to wounds and tissue absence. Dev Biol 344(2):979–991
Sugio M et al (2012) Stem cells in asexual reproduction of Enchytraeus japonensis (Oligochaeta, Annelid): proliferation and migration of neoblasts. Dev Growth Differ 54(4):439–450
de Jong DM, Seaver EC (2018) Investigation into the cellular origins of posterior regeneration in the annelid Capitella teleta. Regeneration (Oxf) 5(1):61–77
Zattara EE, Turlington KW, Bely AE (2016) Long-term time-lapse live imaging reveals extensive cell migration during annelid regeneration. BMC Dev Biol 16:6
Rinkevich Y et al (2010) Piwi positive cells that line the vasculature epithelium, underlie whole body regeneration in a basal chordate. Dev Biol 345(1):94–104
Jeffery WR (2015) Distal regeneration involves the age dependent activity of branchial sac stem cells in the Ascidian Ciona intestinalis. Regeneration (Oxf) 2(1):1–18
Sehring IM, Weidinger G (2020) Recent advancements in understanding fin regeneration in zebrafish. Wiley Interdiscip Rev Dev Biol 9(1):e367
Poleo G et al (2001) Cell proliferation and movement during early fin regeneration in zebrafish. Dev Dyn 221(4):380–390
Ando K et al (2017) Osteoblast production by reserved progenitor cells in zebrafish bone regeneration and maintenance. Dev Cell 43(5):643-650.e3
Boehm AM, Bosch TC (2012) Migration of multipotent interstitial stem cells in Hydra. Zoology (Jena) 115(5):275–282
Bradshaw B, Thompson K, Frank U (2015) Distinct mechanisms underlie oral vs aboral regeneration in the cnidarian Hydractinia echinata. Elife 4:e05506
Stocum DL, Cameron JA (2011) Looking proximally and distally: 100 years of limb regeneration and beyond. Dev Dyn 240(5):943–968
Sandoval-Guzman T et al (2014) Fundamental differences in dedifferentiation and stem cell recruitment during skeletal muscle regeneration in two salamander species. Cell Stem Cell 14(2):174–187
Kragl M et al (2009) Cells keep a memory of their tissue origin during axolotl limb regeneration. Nature 460(7251):60–65
Stewart S, Stankunas K (2012) Limited dedifferentiation provides replacement tissue during zebrafish fin regeneration. Dev Biol 365(2):339–349
Johnston AP et al (2016) Dedifferentiated schwann cell precursors secreting paracrine factors are required for regeneration of the mammalian digit tip. Cell Stem Cell 19(4):433–448
Planques A et al (2019) Morphological, cellular and molecular characterization of posterior regeneration in the marine annelid Platynereis dumerilii. Dev Biol 445(2):189–210
Somorjai IM et al (2012) Vertebrate-like regeneration in the invertebrate chordate amphioxus. Proc Natl Acad Sci U S A 109(2):517–522
Fan T et al (2011) Patterns and cellular mechanisms of arm regeneration in adult starfish Asterias rollestoni bell. J Ocean Univ China 10(3):255–262
Di Benedetto C et al (2014) Echinoderm regeneration: an in vitro approach using the crinoid Antedon mediterranea. Cell Tissue Res 358(1):189–201
Lin YC, Grigoriev NG, Spencer AN (2000) Wound healing in jellyfish striated muscle involves rapid switching between two modes of cell motility and a change in the source of regulatory calcium. Dev Biol 225(1):87–100
Rodrigues AM et al (2012) Skeletal muscle regeneration in Xenopus tadpoles and zebrafish larvae. BMC Dev Biol 12(1):9
Tanaka EM (2016) The molecular and cellular choreography of appendage regeneration. Cell 165(7):1598–1608
Rinkevich Y et al (2011) Germ-layer and lineage-restricted stem/progenitors regenerate the mouse digit tip. Nature 476(7361):409–413
Konstantinides N, Averof M (2014) A common cellular basis for muscle regeneration in arthropods and vertebrates. Science 343(6172):788–791
Stocum DL (1984) The urodele limb regeneration blastema. Determination and organization of the morphogenetic field. Differentiation 27(1):13–28
Wagner DE, Wang IE, Reddien PW (2011) Clonogenic neoblasts are pluripotent adult stem cells that underlie planarian regeneration. Science 332(6031):811–816
De Mulder K et al (2009) Characterization of the stem cell system of the acoel Isodiametra pulchra. BMC Dev Biol 9(1):69
Srivastava M et al (2014) Whole-body acoel regeneration is controlled by Wnt and Bmp-Admp signaling. Curr Biol 24(10):1107–1113
Gehrke AR, Srivastava M (2016) Neoblasts and the evolution of whole-body regeneration. Curr Opin Genet Dev 40:131–137
Jeffery WR (2019) Progenitor targeting by adult stem cells in Ciona homeostasis, injury, and regeneration. Dev Biol 448(2):279–290
David CN (2012) Interstitial stem cells in Hydra: multipotency and decision-making. Int J Dev Biol 56(6–8):489–497
Gargioli C, Slack JM (2004) Cell lineage tracing during Xenopus tail regeneration. Development 131(11):2669–2679
Flowers GP, Sanor LD, Crews CM (2017) Lineage tracing of genome-edited alleles reveals high fidelity axolotl limb regeneration. Elife 6:e25726
Tu S, Johnson SL (2011) Fate restriction in the growing and regenerating zebrafish fin. Dev Cell 20(5):725–732
Morrison JI, Borg P, Simon A (2010) Plasticity and recovery of skeletal muscle satellite cells during limb regeneration. FASEB J 24(3):750–756
Tornini VA et al (2017) Live fate-mapping of joint-associated fibroblasts visualizes expansion of cell contributions during zebrafish fin regeneration. Development 144(16):2889–2895
Witchley JN et al (2013) Muscle cells provide instructions for planarian regeneration. Cell Rep 4(4):633–641
Raz AA et al (2017) Acoel regeneration mechanisms indicate an ancient role for muscle in regenerative patterning. Nat Commun 8(1):1260
Nacu E et al (2013) Connective tissue cells, but not muscle cells, are involved in establishing the proximo-distal outcome of limb regeneration in the axolotl. Development 140(3):513–518
Shibata E et al (2018) Robust and local positional information within a fin ray directs fin length during zebrafish regeneration. Dev Growth Differ 60(6):354–364
Brockes JP (1984) Mitogenic growth factors and nerve dependence of limb regeneration. Science 225(4668):1280–1287
Endo T et al (2015) The accessory limb model: an alternative experimental system of limb regeneration. Methods Mol Biol 1290:101–113
Kumar A et al (2007) Molecular basis for the nerve dependence of limb regeneration in an adult vertebrate. Science 318(5851):772–777
Wang MH et al (2019) Nerve-mediated expression of histone deacetylases regulates limb regeneration in axolotls. Dev Biol 449(2):122–131
Oviedo NJ et al (2010) Long-range neural and gap junction protein-mediated cues control polarity during planarian regeneration. Dev Biol 339(1):188–199
Pietak A et al (2019) Neural control of body-plan axis in regenerating planaria. PLoS Comput Biol 15(4):e1006904
Gazave E, Rottinger E (2019) 7th Euro Evo Devo meeting: Report on the "Evolution of regeneration in Metazoa" symposium. J Exp Zool B Mol Dev Evol. https://doi.org/10.1002/jez.b.22897
Grillo M, Konstantinides N, Averof M (2016) Old questions, new models: unraveling complex organ regeneration with new experimental approaches. Curr Opin Genet Dev 40:23–31
Lowe R et al (2017) Transcriptomics technologies. PLoS Comput Biol 13(5):e1005457
Ivankovic M et al (2019) Model systems for regeneration: planarians. Development 146(17). https://doi.org/10.1242/dev.167684
Joven A, Elewa A, Simon A (2019) Model systems for regeneration: salamanders. Development 146(14). https://doi.org/10.1242/dev.167700
Marques IJ, Lupi E, Mercader N (2019) Model systems for regeneration: zebrafish. Development 146(18). https://doi.org/10.1242/dev.167692
Iismaa SE et al (2018) Comparative regenerative mechanisms across different mammalian tissues. NPJ Regen Med 3:6
Kenny NJ et al (2018) Towards the identification of ancestrally shared regenerative mechanisms across the Metazoa: a transcriptomic case study in the Demosponge Halisarca caerulea. Mar Genomics 37:135–147
Vogg MC, Galliot B, Tsiairis CD (2019) Model systems for regeneration: Hydra. Development 146(21). https://doi.org/10.1242/dev.177212
Warner JF et al (2018) NvERTx: a gene expression database to compare embryogenesis and regeneration in the sea anemone Nematostella vectensis. Development 145(10). https://doi.org/10.1242/dev.162867
Ramon-Mateu J et al (2019) Regeneration in the ctenophore Mnemiopsis leidyi occurs in the absence of a blastema, requires cell division, and is temporally separable from wound healing. BMC Biol 17(1):80
Zattara EE et al (1898) A phylum-wide survey reveals multiple independent gains of head regeneration in Nemertea. Proc Biol Sci 2019(286):20182524
Imperadore P et al (2017) Nerve degeneration and regeneration in the cephalopod mollusc Octopus vulgaris: the case of the pallial nerve. Sci Rep 7:46564
Bely AE (2010) Evolutionary loss of animal regeneration: pattern and process. Integr Comp Biol 50(4):515–527
Sikes JM, Newmark PA (2013) Restoration of anterior regeneration in a planarian with limited regenerative ability. Nature 500(7460):77–80
Kao D, Felix D, Aboobaker A (2013) The planarian regeneration transcriptome reveals a shared but temporally shifted regulatory program between opposing head and tail scenarios. BMC Genomics 14:797
Qin YF et al (2011) Transcriptome profiling and digital gene expression by deep-sequencing in normal/regenerative tissues of planarian Dugesia japonica. Genomics 97(6):364–371
Almazan EMP et al (2018) Girardia dorotocephala transcriptome sequence, assembly, and validation through characterization of piwi homologs and stem cell progeny markers. Dev Biol 433(2):433–447
Liu SY et al (2013) Reactivating head regrowth in a regeneration-deficient planarian species. Nature 500(7460):81–84
Wasik K et al (2015) Genome and transcriptome of the regeneration-competent flatworm, Macrostomum lignano. Proc Natl Acad Sci U S A 112(40):12462–12467
Cary GA et al (2019) Analysis of sea star larval regeneration reveals conserved processes of whole-body regeneration across the metazoa. BMC Biol 17(1):16
Lee-Liu D et al (2014) Genome-wide expression profile of the response to spinal cord injury in Xenopus laevis reveals extensive differences between regenerative and non-regenerative stages. Neural Dev 9:12
Pibiri M et al (2015) Global gene expression profile of normal and regenerating liver in young and old mice. Age (Dordr) 37(3):9796
Wang Z et al (2019) Mechanistic basis of neonatal heart regeneration revealed by transcriptome and histone modification profiling. Proc Natl Acad Sci U S A 116(37):18455–18465
Blythe MJ et al (2010) A dual platform approach to transcript discovery for the planarian Schmidtea mediterranea to establish RNAseq for stem cell and regeneration biology. PLoS ONE 5(12):e15617
Sun L et al (2011) Large scale gene expression profiling during intestine and body wall regeneration in the sea cucumber Apostichopus japonicus. Comp Biochem Physiol D Genomics Proteomics 6(2):195–205
Fumagalli MR, Zapperi S, La Porta CAM (2018) Regeneration in distantly related species: common strategies and pathways. NPJ Syst Biol Appl 4:5
Benenati, G., J.I. Montoya-Burgos, and B. Galliot, Towards a parsimonious analysis of regeneration and self-repair in animal evolution, in Proceedings of the Seventh International Workshop on Information Processing in Cells and Tissues (IPCAT 2007), N.C.T.o. Scheper, Editor. 2007, Jesus College Oxford: Oxford, United Kingdom. p. 90–100.
Czarkwiani A et al (2019) FGF signalling plays similar roles in development and regeneration of the skeleton in the brittle star Amphiura filiformis. BioRxiv. https://doi.org/10.1101/632968
Rabinowitz JS et al (2017) Transcriptomic, proteomic, and metabolomic landscape of positional memory in the caudal fin of zebrafish. Proc Natl Acad Sci U S A 114(5):E717–E726
Bryant DM et al (2017) A tissue-mapped axolotl de novo transcriptome enables identification of limb regeneration factors. Cell Rep 18(3):762–776
Wenger Y et al (2019) Generic and context-dependent gene modulations during Hydra whole body regeneration. BioRxiv. https://doi.org/10.1101/587147
Duffy DJ et al (2010) Wnt signaling promotes oral but suppresses aboral structures in Hydractinia metamorphosis and regeneration. Development 137(18):3057–3066
Lengfeld T et al (2009) Multiple Wnts are involved in Hydra organizer formation and regeneration. Dev Biol 330(1):186–199
Schaffer AA et al (2016) A transcriptional time-course analysis of oral vs. aboral whole-body regeneration in the Sea anemone Nematostella vectensis. BMC Genomics 17:718
Chen C-Y, Yueh W-T, Chen J-H (2020) Canonical wnt signaling is involved in anterior regeneration of the annelid Aeolosoma viride. bioRxiv. https://doi.org/10.1101/2020.03.01.972448
Petersen CP, Reddien PW (2009) A wound-induced Wnt expression program controls planarian regeneration polarity. Proc Natl Acad Sci U S A 106(40):17061–17066
Petersen CP, Reddien PW (2011) Polarized notum activation at wounds inhibits Wnt function to promote planarian head regeneration. Science 332(6031):852–855
Gehrke AR et al (2019) Acoel genome reveals the regulatory landscape of whole-body regeneration. Science 363(6432):eaau6173. https://doi.org/10.1126/science.aau6173
Ramirez AN, Loubet-Senear K, Srivastava M (2020) A regulatory program for initiation of Wnt signaling during posterior regeneration. Cell Rep 32(9):108098
Kawakami Y et al (2006) Wnt/beta-catenin signaling regulates vertebrate limb regeneration. Genes Dev 20(23):3232–3237
Wischin S et al (2017) Chemical activation of Wnt/beta-catenin signalling inhibits innervation and causes skeletal tissue malformations during axolotl limb regeneration. Mech Dev 144(Pt B):182–190
Wehner D et al (2014) Wnt/beta-catenin signaling defines organizing centers that orchestrate growth and differentiation of the regenerating zebrafish caudal fin. Cell Rep 6(3):467–481
Takeo M et al (2013) Wnt activation in nail epithelium couples nail growth to digit regeneration. Nature 499(7457):228–232
Ribeiro RP et al (2019) Comparative transcriptomics in Syllidae (Annelida) indicates that posterior regeneration and regular growth are comparable, while anterior regeneration is a distinct process. BMC Genom 20(1):855. https://doi.org/10.1186/s12864-019-6223-y
Phipps LS et al (2020) Model systems for regeneration: Xenopus. Development 147(6). https://doi.org/10.1242/dev.180844
Bryant DM et al (2017) Identification of regenerative roadblocks via repeat deployment of limb regeneration in axolotls. NPJ Regen Med 2:30
Wu CH et al (2013) De novo transcriptome sequencing of axolotl blastema for identification of differentially expressed genes during limb regeneration. BMC Genomics 14:434
Vitulo N et al (2017) Transcriptome analysis of the regenerating tail vs. the scarring limb in lizard reveals pathways leading to successful vs. unsuccessful organ regeneration in amniotes. Dev Dyn 246(2):116–134
Petersen HO et al (2015) A comprehensive transcriptomic and proteomic analysis of hydra head regeneration. Mol Biol Evol 32(8):1928–1947
Vizcaya-Molina E et al (2018) Damage-responsive elements in Drosophila regeneration. Genome Res 28(12):1852–1866
Mashanov VS, Zueva OR, Garcia-Arraras JE (2014) Transcriptomic changes during regeneration of the central nervous system in an echinoderm. BMC Genomics 15:357
Zondag LE et al (2016) Uncovering the pathways underlying whole body regeneration in a chordate model, Botrylloides leachi using de novo transcriptome analysis. BMC Genomics 17:114
Luttrell SM et al (2016) Head regeneration in hemichordates is not a strict recapitulation of development. Dev Dyn 245(12):1159–1175
Xu C et al (2020) Transcriptional analysis of scar-free wound healing during early stages of tail regeneration in the green anole lizard, Anolis carolinensis. J Immunol Regen Med 7:100025. https://doi.org/10.1016/j.regen.2019.100025
Bhambri A et al (2018) Large scale changes in the transcriptome of Eisenia fetida during regeneration. PLoS ONE 13(9):e0204234
Hutchins ED et al (2014) Transcriptomic analysis of tail regeneration in the lizard Anolis carolinensis reveals activation of conserved vertebrate developmental and repair mechanisms. PLoS ONE 9(8):e105004
Bando T et al (2013) Analysis of RNA-Seq data reveals involvement of JAK/STAT signalling during leg regeneration in the cricket Gryllus bimaculatus. Development 140(5):959–964
Arenas Gómez CM et al (2018) Using transcriptomics to enable a plethodontid salamander (Bolitoglossa ramosi) for limb regeneration research. BMC Genomics 19(1):704
Trapnell C (2015) Defining cell types and states with single-cell genomics. Genome Res 25(10):1491–1498
Wang Y, Navin NE (2015) Advances and applications of single-cell sequencing technologies. Mol Cell 58(4):598–609
Cao J et al (2019) The single-cell transcriptional landscape of mammalian organogenesis. Nature 566(7745):496–502
Pijuan-Sala B et al (2019) A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566(7745):490–495
Scimone ML et al (2016) Two FGFRL-Wnt circuits organize the planarian anteroposterior axis. Elife. https://doi.org/10.7554/eLife.12845
Molinaro AM, Pearson BJ (2016) In silico lineage tracing through single cell transcriptomics identifies a neural stem cell population in planarians. Genome Biol 17:87
Wurtzel O, Oderberg IM, Reddien PW (2017) Planarian epidermal stem cells respond to positional cues to promote cell-type diversity. Dev Cell 40(5):491-504.e5
Plass M et al (2018) Cell type atlas and lineage tree of a whole complex animal by single-cell transcriptomics. Science 360(6391). https://doi.org/10.1126/science.aaq1723
Fincher CT et al (2018) Cell type transcriptome atlas for the planarian Schmidtea mediterranea. Science 360(6391). https://doi.org/10.1126/science.aaq1736
Benham-Pyle BW et al (2020) Identification of rare transient somatic cell states induced by injury and required for whole-body regeneration. BioRxiv. https://doi.org/10.1101/2020.06.04.132753
Shao Y et al (2020) Genome and single-cell RNA-sequencing of the earthworm Eisenia andrei identifies cellular mechanisms underlying regeneration. Nat Commun 11(1):2656
Aztekin C et al (2019) Identification of a regeneration-organizing cell in the Xenopus tail. Science 364(6441):653–658
Kakebeen AD et al (2020) Chromatin accessibility dynamics and single cell RNA-Seq reveal new regulators of regeneration in neural progenitors. Elife. https://doi.org/10.7554/eLife.52648
Gerber T et al (2018) Single-cell analysis uncovers convergence of cell identities during axolotl limb regeneration. Science 362(6413). https://doi.org/10.1126/science.aaq0681
Leigh ND et al (2018) Transcriptomic landscape of the blastema niche in regenerating adult axolotl limbs at single-cell resolution. Nat Commun 9(1):5153
Rodgers AK, Smith JJ, Voss SR (2020) Identification of immune and non-immune cells in regenerating axolotl limbs by single-cell sequencing. Exp Cell Res 394(2):112149
Cao J et al (2016) Single epicardial cell transcriptome sequencing identifies Caveolin 1 as an essential factor in zebrafish heart regeneration. Development 143(2):232–243
Wang W et al (2020) Changes in regeneration-responsive enhancers shape regenerative capacities in vertebrates. Science 369(6508). https://doi.org/10.1126/science.aaz3090
Hou Y et al (2020) Cellular diversity of the regenerating caudal fin. Sci Adv 6(33):eaba2084
Storer MA et al (2020) Acquisition of a unique mesenchymal precursor-like blastema state underlies successful adult mammalian digit tip regeneration. Dev Cell 52(4):509-524.e9
Vaughan AE et al (2015) Lineage-negative progenitors mobilize to regenerate lung epithelium after major injury. Nature 517(7536):621–625
Carr MJ et al (2019) Mesenchymal precursor cells in adult nerves contribute to mammalian tissue repair and regeneration. Cell Stem Cell 24(2):240-256.e9
Johnson GL, Masias EJ, Lehoczky JA (2020) Cellular heterogeneity and lineage restriction during mouse digit tip regeneration at single-cell resolution. Dev Cell 52(4):525-540.e5
Hasin Y, Seldin M, Lusis A (2017) Multi-omics approaches to disease. Genome Biol 18(1):83
Feuda R et al (2017) Improved modeling of compositional heterogeneity supports sponges as sister to all other animals. Curr Biol 27(24):3864
Laumer CE et al (1906) Revisiting metazoan phylogeny with genomic sampling of all phyla. Proc Biol Sci 2019(286):20190831
Acknowledgements
We are grateful to Haley Flom for her diligent proofreading of this manuscript.
Funding
This work was supported by the PROCORE—France/Hong Kong Joint Research Scheme. Work in the Vervoort team is supported by funding from the Labex “Who Am I” laboratory of excellence (No. ANR- 11-LABX-0071) funded by the French Government through its “Investments for the Future” program operated by the Agence Nationale de la Recherche under grant No. ANR-11-IDEX-0005-01, the Centre National de la Recherche Scientifique (CNRS), the INSB department (grant “Diversity of Biological Mechanisms”), the Agence Nationale de la Recherche (ANR grants TELOBLAST no. ANR-16-CE91-0007 and STEM no. ANR-19-CE27-0027-02), the “Association pour la Recherche sur le Cancer” (grant PJA 20191209482), and the “Ligue Nationale Contre le Cancer” (grant RS20/75-20). JH is funded by the TUYF Charitable Trust.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
18_2021_3760_MOESM1_ESM.docx
Supplementary file1 Supplementary table 1: Overview of bulk mRNA-seq studies during regeneration in Metazoa. Large bibliographic analysis of bulk mRNA-seq literature within Metazoa allow to address key questions in the regeneration field (see main text). For each study, model species (as well as their classification), type of regeneration (whole body, complex structures or organ, but cell regeneration excluded), and main results are listed. (DOCX 219 KB)
Rights and permissions
About this article
Cite this article
Bideau, L., Kerner, P., Hui, J. et al. Animal regeneration in the era of transcriptomics. Cell. Mol. Life Sci. 78, 3941–3956 (2021). https://doi.org/10.1007/s00018-021-03760-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-021-03760-7