Abstract
Axillary meristems (AMs) are located in the leaf axil and can establish new growth axes. Whereas their neighboring cells are differentiated, the undifferentiated cells in the AM endow the AM with the same developmental potential as the shoot apical meristem. The AM is, therefore, an excellent system to study stem cell fate maintenance in plants. In this review, we summarize the current knowledge of AM initiation. Recent findings have shown that AMs derive from a stem cell lineage that is maintained in the leaf axil. This review covers AM progenitor cell fate maintenance, reactivation, and meristem establishment. We also highlight recent work that links transcription factors, phytohormones, and epigenetic regulation to AM initiation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Unlike most animals, plants are ramifying systems with new cycles of growth throughout their lifespan. At one level, this constant growth is achieved by postembryonic organogenesis. The shoot apical meristem (SAM), which is formed during embryogenesis and harbors stem cells in its center, forms lateral organs such as leaves at its periphery. Thus, the SAM creates the shoot, i.e., the aboveground portion of a plant. Similarly, the root apical meristem gives rise to the entire underground root system. However, these two meristems only establish a single growth axis, and the periodic formation of branching meristems is needed to initiate new growth axes. In seed plants, branching is achieved by the axillary meristem (AM) formed at the leaf axil, where the boundary region separates the leaf from the stem. In the model plant Arabidopsis thaliana and many other species, an AM is associated with each leaf, which, together with an internode section, forms a developmental unit called a phytomere. AMs form axillary buds, which can either develop into a branch or remain dormant for a certain amount of time or even permanently. Axillary bud dormancy is strongly promoted by the SAM, and this phenomenon is termed apical dominance. Extensive studies have shown that auxin, strigolactone, and sucrose as well as light and other environmental signals all affect apical dominance. A number of excellent recent reviews have focused on axillary bud outgrowth [1,2,3]. In this review, we focus on AM initiation, which is an ideal system to study stem cell fate determination.
AM initiation is informative about cell fate, stem cells, and meristem organization
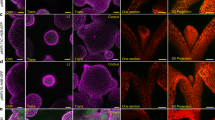
The origin of AMs has long been a matter of debate. Based on morphology, two theories have been proposed. The de novo theory posits that differentiated cells dedifferentiate to become stem cells and form AMs. The alternative detached theory proposes that some leaf axil cells remain undifferentiated and later form AMs [4]. Recent works identified a meristematic cell population in the leaf axil that is necessary for AM initiation [5]. However, this leaf axil population differs from the SAM cell population, as it does not express the stem cell markers CLAVATA3 (CLV3) and WUSCHEL (WUS) [6,7,8,9]. These leaf axil meristematic cells undergo a maintenance phase, which is marked by low expression levels of the meristem gene SHOOTMERISTEMLESS (STM); a subsequent activation phase, in which STM expression increases; and finally an emergence phase, which is characterized by the establishment of the WUS-CLV3 feedback loop (Fig. 1). This leaf axil meristematic lineage and AM initiation represent an ideal conceptual and technical framework for the study of cell fate, stem cells, and meristem organization [10]. Recent findings have shed light on the molecular regulation of each phase, and the following sections cover the regulation of stem cell fate during AM initiation. While this review focuses on vegetative stage branching, readers can refer to recent reviews on reproductive-stage branching, especially spike branching in grasses [11,12,13,14,15,16].
Conceptual models of the different stages of AM initiation with a focus on the expression profiles of STM and WUS. a According to the expression levels of STM and WUS, AM initiation can be divided into three stages: maintenance, activation, and initiation (detailed in b–d). Arrows and inhibition symbols indicate activation and repression, respectively
Distinct gene regulatory networks control postembryonic AM initiation
Because AMs have the same developmental potential as the SAM, a natural line of inquiry is whether AM initiation is under the same developmental regulation as embryonic SAM formation. There are indeed many similarities between the AM and SAM, particularly after the morphogenesis phase. In contrast, the maintenance and activation phases are largely specific to AM initiation and are not associated with SAM formation. Genes such as LATERAL SUPRESSOR (LAS) and REGULATOR OF AXILLARY MERISTEMS (RAX) in Arabidopsis have expression patterns highly specific to AM initiation. On the other hand, mutations in STM and WUS, both of which are required for SAM maintenance, also affect AM initiation.
STM regulates SAM maintenance, and is broadly expressed in the SAM since globular stage of embryogenesis [17, 18]. In the SAM, STM prevents meristematic cell differentiation. As mentioned earlier, STM is also profoundly involved in regulating AM initiation. Whereas strong stm alleles lack leaves, the intermediate/weak stm-bum1 allele exhibits significantly reduced axillary buds formation [5]. STM expression is maintained in a small population of leaf axil cells, and these cells retain their meristematic identity and do not form vacuoles. Laser ablation of these STM-expressing cells abolishes bud formation, indicating that they are required to initiate axillary buds. The STM expression pattern in the leaf axil dynamically changes over the course of AM initiation (Fig. 1). During the earlier maintenance phase, the expression of STM is maintained but with a gradual decline. In the subsequent and activation phase, STM expression is significantly upregulated, which is accompanied by enhanced cell divisions [5]. It is important to note that only cells with maintained STM expression can have high levels of STM expression during the activation phase to initiate AMs. In other words, there is no de novo establishment of STM expression. Based on its expression patterns, STM is a convenient marker for meristematic cell fate.
WUS is another crucial factor for SAM maintenance [19, 20]. During embryogenesis, WUS expression starts at the 16-cell stage, and its expression is constrained to the organizing center below the stem cells [19]. WUS is diffusible and directly activates the expression of the stem cell marker gene CLV3 in the above stem cells [21,22,23,24]. In the organizing center, the expression of WUS-interacting HAIRY MERISTEM proteins prohibits WUS activation of CLV3 expression [8, 9]. CLV3 encodes a polypeptide hormone that represses WUS expression [23, 24]. The WUS-CLV3 feedback loop maintains stem cell niche homeostasis and according wus mutants are unable to maintain the SAM [20, 25]. WUS is also involved in AM initiation, as axillary buds are often completely absent or converted into one or a few leaf-like structures in wus mutants [6]. In contrast to STM, WUS expression in the leaf axil is not maintained. WUS expression is only detectable in mature leaf axils, around P13 (the thirteenth earliest leaf primordium) leaves in the Col-0 ecotype, when the AM has formed a visible bulge (Fig. 1a, d) [6]. WUS expression is initially scattered in a few cells, and gradually restrict to the organization center after CLV3 expression becomes detectable [6,7,8,9]. In rice (Oryza sativa), mutation in the WUS orthologous gene MONOCULM3 (MOC3, also named TILLERS ABSENT1 and STERILE AND REDUCED TILLERING1) causes AM initiation defects and female fertility, with minimal effects on SAM maintenance [26,27,28].
Transcriptional regulation
To date, most of the identified regulators of AM initiation encode transcription factors (Fig. 2). Many of these genes are expressed specifically in the leaf axil and form a leaf axil-enriched gene regulatory network (GRN). Several key regulators of the GRN were initially identified by forward genetics, and have been used as ‘anchor points’ to pull out additional ones [29, 30].
Summary diagram of transcription factors and phytohormones controlling AM initiation. This diagram includes factors that function at different temporal stages. The black boxes denote the key meristematic factors STM and WUS. Green arrows and red inhibition symbols represent positive regulation and negative regulation, respectively
LAS/MOC1
The Arabidopsis LAS gene specifically affects AM initiation and not the SAM. LAS encodes a GRAS family transcription factor, and is specifically expressed in the leaf axil [31]. Lateral suppressor (Ls), the LAS ortholog in tomato (Solanum lycopersicum), was first identified by forward genetic analysis. Tomato ls mutant plants lack axillary buds and also have petal development defects [32]. Using reverse genetics, Arabidopsis las mutants were found to have severe defects in axillary bud formation but normal flower development [31]. Large-scale GRN analysis indicated that the LAS promoter is bound by many transcription factors and is likely to be a “regulatory hub” [30]. An independent forward genetic research identified MONOCULM1 (MOC1), the rice LAS ortholog, which also exhibits boundary-specific expression. Rice moc1 mutants have significantly reduced tiller number and inflorescence branching [33], suggesting that MOC1 is involved in both vegetative and reproductive development. In rice, a co-activator of the cell cycle anaphase-promoting complex, TILLERING AND DWARF 1 (also named TILLER ENHANCER), targets MOC1 to the 26S proteasome for degradation in a cell cycle-dependent manner [34, 35]. At the protein level, rice MOC1 physically interacts with MOC3 and enhances MOC3 activation of FLORAL ORGAN NUMBER1, the CLV1 ortholog in rice. Rice MOC1 promotes tiller bud outgrowth [36], which is distinct from the functions of its orthologs in Arabidopsis and tomato.
RAX
There are three paralogous RAX genes in Arabidopsis, all encoding R2R3 MYB transcription factors that redundantly regulate AM initiation. The RAX ortholog in tomato, Blind (Bl), was identified by forward genetic analysis of bl mutants lacking side shoots [37]. The three Arabidopsis RAX genes similarly regulate early AM initiation. RAX1 and RAX3 expression domains are restricted to the center of the leaf axils and mark the positions where AMs initiate [38, 39]. In contrast, RAX2 is more broadly expressed. Among the RAX gene mutants, rax1 mutants have the strongest AM initiation defects, which are enhanced by rax2 and rax3 mutations. rax1 rax2 rax3 triple mutants seldom develop axillary buds in either the rosette or cauline leaves [38].
CUC
In Arabidopsis, there are three NAC transcription factor-encoding CUP-SHAPED COTYLEDON (CUC) genes that redundantly regulate boundary formation. Two of them, CUC1 and CUC2, are under miR164 regulation. CUC1 and CUC2 were first isolated for their redundant functions in embryonic SAM formation and cotyledon separation during embryogenesis, and the double mutant phenotype is characterized by occasionally fused cup-shaped cotyledons [40]. Similar mutant phenotypes were identified in petunia, Antirrhinum, and tomato [41,42,43]. CUC genes and their orthologous genes in other species redundantly affect phyllotaxis, leaf-stem separation, compound leaf formation or leaf margin serration, and carpel margin development [44,45,46,47,48,49,50,51,52], In Arabidopsis, CUC genes also redundantly promote AM initiation, with CUC3, and to a lesser extent CUC2, playing the leading roles. cuc3 single mutants have dramatic AM initiation defects in rosette leaves, and both cuc1 and cuc2 mutations enhance the cuc3 mutant phenotype. In cuc1 cuc2 cuc3 triple mutant plants, almost all rosette and cauline leaves have barren leaf axils [53, 54]. In terms of their expression patterns, all three Arabidopsis CUC genes are restricted to the boundary regions. During embryogenesis, CUC1 and CUC2 are excluded from the uppermost layer, while CUC3 expression is detected in this layer [40, 54, 55]. Consistent with this differential expression, only CUC1 and CUC2 transcripts are degraded by miR164. Among the three MIR164 genes, MIR164c also exhibits boundary-specific expression [53]. Similar to LAS, the CUC2 promotor is bound by a large number of transcription factors, including RAX1 [30]. It remains unclear if this RAX1 binding represents primary regulation or secondary feedback, as CUC2 expression precedes that of RAX1. In terms of downstream regulation, CUC genes regulate STM and LAS [30, 40, 53, 56]. STM expression is lost in cuc1 cuc2 double mutant heart-stage embryos, and CUC1 protein binds to the STM promoter to induce its expression [40, 55, 56]. In turn, STM protein binds to the CUC1 promotor to activate CUC1 expression, forming a positive feedback loop [56, 57]. The same regulatory mechanism may also function during AM initiation. In cuc triple mutant plants, LAS expression in the leaf axil is lost [53]. Consistently, CUC2 proteins directly bind to the LAS promotor and upregulate its expression [30].
REV
REVOLUTA (REV) has pleiotropic effects on shoot and root development, including SAM and floral meristem activity, leave and stem patterning and growth, vascular development, and root development [58, 59]. Notably, rev mutant plants lack axillary buds in both rosette and cauline leaf axils [58, 60, 61]. REV is broadly expressed in the SAM, leaf adaxial domain facing the SAM, center of the leaf axil proceeding AM initiation, and vascular tissues [5, 60,61,62]. REV encodes an HD-ZIPIII family transcription factor, which also includes PHABULOSA (PHB), PHAVOLUTA (PHV), CORONA, and ATHB8 [60, 61]. The expressions of these HD-ZIPIII genes are regulated by MIR165/6 [63, 64]. Nevertheless, only rev mutants have AM initiation defects, but not other single mutants for HD-ZIPIII genes [60]. On the other hand, dominant mutants of PHB, PHV, and REV, which escape from MIR165/6 regulation, similarly exhibit adaxialized leaves and ectopic axillary buds surrounding the adaxialized leaves [61, 62, 65, 66].
REV is a downstream factor during AM initiation, and LAS promotes REV expression in the leaf axil [31]. REV in turn upregulates STM expression immediately prior to AM initiation by directly binding to the STM promoter region [5]. REV upregulation of STM requires prior STM expression and a permissive local epigenetic environment, which ensures that STM is not ectopically expressed in other tissues where REV is active [5]. Similar to the findings in Arabidopsis, rice HD-ZIPIII transcription factor LATERAL FLORET1 upregulates the expression of OSH1, a homolog of STM, to promote spikelet branching and to form extra florets [67].
Although the REV-related PHB and PHV proteins can similarly bind to STM and promote its expression [5, 68], the observed AM initiation defects are unique to rev mutants [60]. It is likely that among these genes, only REV is strongly expressed in mature leaf axils. On the other hand, PHB, PHV, and REV alleles that lead to ectopic expression of these genes all give rise to ectopic AMs on the abaxial leaf base, providing support for the de novo theory [65]. It is conceivable that ectopic activity of these HD-ZIPIII transcription factors could maintain STM expression in the leaf abaxial side and promote ectopic AM initiation.
LAX1/BA1/ROX
The LAX PANICLE1 (LAX1) bHLH protein gene in rice was identified by forward genetic analysis, and mutation of this gene leads to defects in both vegetative and reproductive-stage branching [69]. Stage- and direction-specific LAX1 protein trafficking is essential for its function during AM initiation [70]. An independent forward genetic analysis showed that mutations in the orthologous barren stalk1 (ba1) gene in maize similarly lead to vegetative and reproductive branching defects [71]. LAX1 and ba1 orthologs broadly exist in many species [72]. Reverse genetic analysis showed that the Arabidopsis ortholog, REGULATOR OF AXILLARY MERISTEM FORMATION (ROX), also participates in AM initiation, but specifically affects early vegetative development [73]. ROX expression is restricted to the leaf axil, and is upregulated by both RAX and LAS [73]. The wild sunflower (Helianthus annuus) plants are highly branched with many small flowering heads. By contrast, domesticated sunflowers commonly produce a single large head. Recent studies showed that the ROX ortholog, ROX-LIKE (Ha-ROXL), was responsible for the plant architecture changes associated with domestication [74, 75]. Like rice and maize, but different from Arabidopsis, sunflower Ha-ROXL is responsible for both vegetative and reproductive-stage AM initiation. Furthermore, HA-ROXL promotes the expressions of RAX, LAS, and CUC2 orthologs [74].
LAX2/BA2
Rice LAX2 participates in AM initiation during both the vegetative and reproductive stages [76]. LAX2 is broadly expressed in meristems and encodes a novel nuclear protein that contains a plant-specific conserved domain. BA2 is the LAX2 ortholog in maize, and it affects tiller bud formation, ear row number, and tassel branching [77]. The physical interactions of LAX2 with LAX1 in rice and BA2 with BA1 in maize are both known to regulate AM formation. However, both the lax1 lax2 and ba1 ba2 double mutants have much more severe bud formation defects than the single mutants [76, 77], suggesting the existence of additional interacting factors.
Other transcription factors
There are additional regulatory genes of AM initiation, and the majority of them encode transcription factors. A genome-scale yeast one-hybrid (Y1H) screen identified new transcription factors that bind to the promotor regions of LAS, CUC2, RAX1, and STM [30]. LAS and CUC2 are putative regulatory hubs based on the abundance of transcription factors that bind to their promoters. SQUAMOSA PROMOTER-BINDING PROTEIN-LIKE 9 and 15 (SPL9/15) bind to the LAS promotor to suppress its expression, and mutations in SPL9/15 lead to the formation of extra AMs, termed accessory meristems, in cauline leaf axils [30]. DORNRÖSCHEN (DRN, also known as ENHANCER OF SHOOT REGENERATION1, ESR1) and its homolog DRN-LIKE (DRNL, also known as ESR2) are required for AM initiation. In single and double drn and drnl mutants, a significant portion of the leaf axils are barren [30]. DRN and DRNL regulate AM initiation through various factors: both proteins bind to the CUC2 promoter to promote its expression [30], and they also upregulate CUC1 expression [78, 79].
Recent studies identified regulators of RAX1. EXCESSIVE BRANCHES1 (EXB1, also named WRKY71) encodes a boundary-specific WRKY transcription factor, and EXB1 overexpression leads to excessive AM initiation. Fusing EXB1 with an EAR repression domain to suppress downstream gene expression leads to inhibition of AM initiation. However, exb1 mutant plants exhibit normal axillary bud formation, suggesting genetic redundancy. EXB1 also directly activates the expression of RAX genes [80]. LEAFY (LFY) is a master regulator of the reproductive-stage transition. An LFY allele with reduced floral function was used to show that LFY directly activates RAX1 expression to promote AM initiation [81]. This finding also explains why AM initiation is much faster in cauline leaves, which are formed during reproductive-stage transition [82], compared to rosette leaves.
Genes with boundary-specific expression are often involved in AM initiation. One example is LATERAL ORGAN FUSION1 (LOF1), which encodes an MYB transcription factor. LOF1 weakly promotes AM initiation together with its homolog LOF2 [83]. The tomato ortholog Trifoliate promotes compound leaf formation and also weakly affects AM initiation [84]. A recent leaf axil-specific translatome analysis also identified new regulators of AM initiation [29]. HANABA TARANU, which encodes a GATA transcription factor, promotes AM initiation in addition to its known roles in embryogenesis and meristem maintenance [85,86,87,88]. The translatome analysis also identified RABBIT EARS, which encodes a C2H2 family zinc-finger transcriptional repressor, as a suppressor of AM initiation.
A recent cell domain-specific transcriptome analysis showed that leaf axil boundary genes were also expressed in maize ligules, which separate grass leaf sheath and leaf blade [89]. This finding is supported by genetic analyses of the BLADE-ON-PETIOLE (BOP) genes in several species. The BOP genes were first identified in Arabidopsis as suppressors of lamina formation on the petiole [90, 91], and have later been associated with abscission zone formation, bract suppression, and floral patterning [92,93,94]. Notably, the barley BOP gene UNICULME4 (CUL4) is expressed in the boundary and promotes axillary bud and ligule formation, as well as inflorescence development [95]. Furthermore, Eligulum-a (ELI-a) another related barley gene expressed in the preligular band and various other tissues is related to AM initiation. Mutations in ELI-a in combination with the uniculm2 mutant, which lacks tillers, result in plants that restore tiller formation [96]. The functions of BOP appear to be highly diverged among different species. They control maize axillary bud outgrowth [97] and tomato inflorescence development [98]. However, BOP function in rice is restricted to leaf development [99]. Nevertheless, the connection between AM initiation and ligule formation point to new approaches to identify novel genes that regulate AM development.
Phytohormones
Phytohormones play critical roles in various plant developmental process, and recent studies have linked some of them to AM initiation (Fig. 2).
Auxin
Whereas auxin maxima are often associated with organogenesis, such as the formation of leaves, flowers, and lateral roots, a low auxin environment is necessary for vegetative AM initiation in Arabidopsis and tomato [100, 101]. The DII auxin sensor, whose signal negatively correlates with auxin levels [102, 103], has strong signals in the leaf axil boundary region starting at leaf primordium emergence in Arabidopsis. DR5 signal, which correlates with auxin signaling, is excluded from the leaf axils in Arabidopsis and tomato. Consistent with these observed patterns, ectopic expression of the iaaM auxin biosynthesis enzyme in the leaf axil led to dramatically compromised AM initiation in Arabidopsis [100, 101]. Local auxin application in the leaf axil in tomato also blocks AM initiation [100]. The leaf axil auxin minimum depends on polar auxin transport, and both auxin efflux and auxin influx mutants exhibit varying degrees of axillary bud formation defects [100, 101]. Similar AM phenotype can also be obtained by treating Arabidopsis or tomato with auxin efflux inhibitor N-1-naphthylphthalamic acid [101]. In pCUC2 ≫ iaaM plants with ectopic auxin biosynthesis in the leaf axil, STM expression is significantly reduced, disrupting STM maintenance [5]. Consistently, auxin signaling suppresses STM expression in the floral meristem [104].
In contrast to its effect on vegetative AM initiation, auxin promotes the formation of the floral meristem, which is considered a specialized AM [4], and probably also cauline leaf AMs, which form during the floral transition [82]. The formation of Arabidopsis floral primordia is further regulated by a reciprocal feedback loop between auxin levels and LFY expression. Auxin promotes LFY expression, and LFY feeds back onto multiple components of the auxin pathway [105,106,107]. A live-imaging analysis of Arabidopsis cauline leaf AMs showed high DR5 expression, which was similarly observed in tomato AMs, likely also after the floral transition [108]. The role of auxin in maize and rice spike branching has been more extensively studied [109].
In maize, auxin biosynthesis is required for AM initiation during vegetative development, in addition to promoting reproductive-stage AM initiation [110, 111]. Given that auxin is required for SAM homeostasis and maintenance [112], it is possible that auxin functions in a later step of AM initiation or bud maintenance. It remains to be determined if auxin is required for vegetative stage bud formation in eudicots, such as Arabidopsis.
Cytokinins
Cytokinins regulate SAM patterning and homeostasis [113,114,115,116], and are broadly involved in shoot meristem functions [117]. During AM initiation in Arabidopsis, a cytokinin signaling pulse, which can be monitored using the TCS cytokinin signaling sensor [118], is detectable just a few days prior to AM initiation in the center of the leaf axil [100]. Cytokinin signaling requires cytokinins, receptors, and signaling pathway components. STM promotes cytokinin biosynthesis in the SAM [119,120,121], and may function similarly in the leaf axil. AHK4, one of the three cytokinin receptors in Arabidopsis, is specifically detected in the leaf axil center [100]. Several type-B ARR transcription factors, which are downstream cytokinin signaling, are expressed or even enriched in the leaf axil [6, 100]. Cytokinin signaling in the leaf axil is causally related to AM initiation, and several combinations of cytokinin perception mutations lead to AM initiation defects. Mutants of type-B ARR transcription factor genes also show compromised AM initiation, with arr1 having the strongest phenotype [100]. The leaf axil cytokinin signaling pulse depends on LAS, RAX, and REV. Moreover, ectopic cytokinin production in the leaf axil can rescue rax mutant phenotypes [100].
Cytokinin de novo activation of WUS expression in the leaf axil helps to establish functional AMs. Unlike STM, WUS expression is not maintained in the leaf axil. Soon after the leaf axil TCS signal, WUS expression is activated in TCS-expressing cells. At the molecular level, type-B ARR transcription factors, especially ARR1, directly bind to the WUS promoter to activate its expression [6]. Consistent with this activity, ectopic WUS expression rescues the AM initiation defect of the arr1-4 mutant [6]. Similar to their function in AM initiation, type-B ARRs also activate WUS expression de novo during shoot regeneration from callus [122, 123]. In addition to binding to WUS, ARR1 also binds to the LAS promotor and activates its expression [30]. Cytokinins may also promote STM expression, which has been shown in the SAM [124].
Other phytohormones
Additional hormones may also contribute to AM initiation. For example, a recent study reported that gibberellin negatively regulates AM initiation [125]. DELLA proteins negatively regulate gibberellin signaling, and della pentuple mutant plants have slightly reduced axillary bud numbers. Additionally, ectopic expression of the GA20ox2 gibberellin biosynthesis gene in the leaf axil inhibits AM initiation [125]. DELLA proteins interact with SPL9 [126], which bind to the LAS promoter to repress its expression [30, 125]. These molecular links may contribute to AM initiation and warrant further confirmation.
Although there is no evidence that the growth-promoting brassinosteroid phytohormones interfere with AM initiation, they do affect organ boundary formation [127, 128]. Additionally, the expression levels of several AM initiation-related genes are altered in brassinosteroid signaling and biosynthesis mutants. Since many of these affected genes, such as CUC and LOF1, are involved in both boundary separation and AM initiation, it remains to be determined if brassinosteroids substantially affect AM initiation.
Epigenetic regulation
Epigenetic regulation allows differential expression of the same gene in different cell types from the same genome. Thus, epigenetic regulation is involved in virtually all developmental processes, including AM initiation. The above-mentioned transcriptional regulation of WUS and STM also depends on epigenetic regulation to ensure that their spatiotemporal expression patterns are precisely controlled. Neither REV, which activates STM expression, nor ARR1, which activates WUS expression, is limited to the leaf axil. Epigenetic regulation restricts the expression of both genes. The Polycomb group (PcG) complexes mediate histone 3 lysine 27 trimethylation (H3K27me3), which is a repressive mark. In differentiated tissues, such as mature leaves, both STM and WUS have high H3K27me3 levels. In contrast, these genes have a low level of H3K27me3 and a high level of H3Ac, a mark associated with active chromatin, in tissues containing the leaf axil STM-expressing cells [5]. It is likely that an open chromatin status is associated with the maintenance of STM expression. In PcG-related mutants, both WUS and STM expression levels are upregulated [5, 6, 129, 130]. Moreover, treating mature leaf cells with the histone deacetylation inhibitor trichostatin A leads to ectopic WUS expression [6].
Epigenetic regulation is more dynamic for WUS, whose expression is terminated in the leaf axil and then subsequently activated de novo [6]. In the leaf axil tissues of young leaf primordia, where WUS is not expressed, the WUS locus has abundant H3K27me3, but is depleted of H3/4Ac. Prior to WUS activation, the levels of the H3K27me3 repressive mark decrease, while the levels of the H3/4Ac active mark increase [6].
Perspectives
Phenotypic characterization of changes in AM initiation is often difficult due to strong apical dominance in many plant species, including the model plant Arabidopsis. Despite the identification of key transcription factors and phytohormones that regulate AM initiation (Fig. 2), there are clearly many “unknowns” in this research area. First, the leaf axil stem cell lineage warrants further characterization. Given that plant cells in general have higher plasticity than animal cells, this stem cell lineage provides an excellent system to study cell fate determination. It remains unknown how this stem cell lineage is maintained, how the epigenetic status is maintained, and to what extent the fate of this stem cell lineage is reversible. To answer these questions, new cellular resolution technologies are needed. Second, there are likely many unknown AM initiation regulators are awaiting to be uncovered. We need new tools for forward genetics and system biology to dissect the gene regulatory networks underlying AM initiation. Third, we need to better connect the known AM initiation regulators to obtain a plausible regulatory network. Finally, regulatory mechanisms other than transcriptional regulation need to be identified affecting AM initiation.
References
Domagalska MA, Leyser O (2011) Signal integration in the control of shoot branching. Nat Rev Mol Cell Biol 12(4):211–221. https://doi.org/10.1038/nrm3088
Wang B, Smith SM, Li J (2018) Genetic regulation of shoot architecture. Annu Rev Plant Biol 69:437–468. https://doi.org/10.1146/annurev-arplant-042817-040422
Barbier FF, Dun EA, Kerr SC, Chabikwa TG, Beveridge CA (2019) An update on the signals controlling shoot branching. Trends Plant Sci 24(3):220–236. https://doi.org/10.1016/j.tplants.2018.12.001
Long J, Barton MK (2000) Initiation of axillary and floral meristems in Arabidopsis. Dev Biol 218(2):341–353. https://doi.org/10.1006/dbio.1999.9572
Shi B, Zhang C, Tian C, Wang J, Wang Q, Xu T, Xu Y, Ohno C, Sablowski R, Heisler MG, Theres K, Wang Y, Jiao Y (2016) Two-step regulation of a meristematic cell population acting in shoot branching in Arabidopsis. PLoS Genet 12(7):e1006168. https://doi.org/10.1371/journal.pgen.1006168
Wang J, Tian C, Zhang C, Shi B, Cao X, Zhang TQ, Zhao Z, Wang JW, Jiao Y (2017) Cytokinin signaling activates WUSCHEL expression during axillary meristem initiation. Plant Cell 29(6):1373–1387. https://doi.org/10.1105/tpc.16.00579
Xin W, Wang Z, Liang Y, Wang Y, Hu Y (2017) Dynamic expression reveals a two-step patterning of WUS and CLV3 during axillary shoot meristem formation in Arabidopsis. J Plant Physiol 214:1–6. https://doi.org/10.1016/j.jplph.2017.03.017
Zhou Y, Yan A, Han H, Li T, Geng Y, Liu X, Meyerowitz EM (2018) HAIRY MERISTEM with WUSCHEL confines CLAVATA3 expression to the outer apical meristem layers. Science 361(6401):502–506. https://doi.org/10.1126/science.aar8638
Gruel J, Deichmann J, Landrein B, Hitchcock T, Jonsson H (2018) The interaction of transcription factors controls the spatial layout of plant aerial stem cell niches. NPJ Syst Biol Appl 4:36. https://doi.org/10.1038/s41540-018-0072-1
Wang Y, Jiao Y (2018) Axillary meristem initiation — a way to branch out. Curr Opin Plant Biol 41:61–66. https://doi.org/10.1016/j.pbi.2017.09.001
Koppolu R, Schnurbusch T (2019) Developmental pathways for shaping spike inflorescence architecture in barley and wheat. J Integr Plant Biol 61(3):278–295. https://doi.org/10.1111/jipb.12771
Gauley A, Boden SA (2019) Genetic pathways controlling inflorescence architecture and development in wheat and barley. J Integr Plant Biol 61(3):296–309. https://doi.org/10.1111/jipb.12732
Kyozuka J, Tokunaga H, Yoshida A (2014) Control of grass inflorescence form by the fine-tuning of meristem phase change. Curr Opin Plant Biol 17:110–115. https://doi.org/10.1016/j.pbi.2013.11.010
Tanaka W, Pautler M, Jackson D, Hirano H-Y (2013) Grass meristems II: inflorescence architecture, flower development and meristem fate. Plant Cell Physiol 54(3):313–324. https://doi.org/10.1093/pcp/pct016
Pautler M, Tanaka W, Hirano H-Y, Jackson D (2013) Grass meristems I: shoot apical meristem maintenance, axillary meristem determinacy and the floral transition. Plant Cell Physiol 54(3):302–312. https://doi.org/10.1093/pcp/pct025
McSteen P (2009) Hormonal regulation of branching in grasses. Plant Physiol 149(1):46–55. https://doi.org/10.1104/pp.108.129056
Clark SE, Jacobsen SE, Levin JZ, Meyerowitz EM (1996) The CLAVATA and SHOOT MERISTEMLESS loci competitively regulate meristem activity in Arabidopsis. Development 122(5):1567–1575
Long JA, Moan EI, Medford JI, Barton MK (1996) A member of the KNOTTED class of homeodomain proteins encoded by the STM gene of Arabidopsis. Nature 379(6560):66–69. https://doi.org/10.1038/379066a0
Mayer KF, Schoof H, Haecker A, Lenhard M, Jurgens G, Laux T (1998) Role of WUSCHEL in regulating stem cell fate in the Arabidopsis shoot meristem. Cell 95(6):805–815
Laux T, Mayer KF, Berger J, Jurgens G (1996) The WUSCHEL gene is required for shoot and floral meristem integrity in Arabidopsis. Development 122(1):87–96
Daum G, Medzihradszky A, Suzaki T, Lohmann JU (2014) A mechanistic framework for noncell autonomous stem cell induction in Arabidopsis. Proc Natl Acad Sci USA 111(40):14619–14624. https://doi.org/10.1073/pnas.1406446111
Yadav RK, Perales M, Gruel J, Girke T, Jonsson H, Reddy GV (2011) WUSCHEL protein movement mediates stem cell homeostasis in the Arabidopsis shoot apex. Genes Dev 25(19):2025–2030. https://doi.org/10.1101/gad.17258511
Brand U, Fletcher JC, Hobe M, Meyerowitz EM, Simon R (2000) Dependence of stem cell fate in Arabidopsis on a feedback loop regulated by CLV3 activity. Science 289(5479):617–619
Schoof H, Lenhard M, Haecker A, Mayer KF, Jurgens G, Laux T (2000) The stem cell population of Arabidopsis shoot meristems is maintained by a regulatory loop between the CLAVATA and WUSCHEL genes. Cell 100(6):635–644
Fletcher JC, Brand U, Running MP, Simon R, Meyerowitz EM (1999) Signaling of cell fate decisions by CLAVATA3 in Arabidopsis shoot meristems. Science 283(5409):1911–1914
Lu Z, Shao G, Xiong J, Jiao Y, Wang J, Liu G, Meng X, Liang Y, Xiong G, Wang Y, Li J (2015) MONOCULM 3, an ortholog of WUSCHEL in rice, is required for tiller bud formation. J Genet Genomics 42(2):71–78. https://doi.org/10.1016/j.jgg.2014.12.005
Tanaka W, Ohmori Y, Ushijima T, Matsusaka H, Matsushita T, Kumamaru T, Kawano S, Hirano H-Y (2015) Axillary meristem formation in rice requires the WUSCHEL ortholog TILLERS ABSENT1. Plant Cell 27(4):1173–1184. https://doi.org/10.1105/tpc.15.00074
Mjomba FM, Zheng Y, Liu H, Tang W, Hong Z, Wang F, Wu W (2016) Homeobox is pivotal for OsWUS controlling tiller development and female fertility in rice G36(7):2013–2021. https://doi.org/10.1534/g3.116.028837
Tian C, Wang Y, Yu H, He J, Wang J, Shi B, Du Q, Provart NJ, Meyerowitz EM, Jiao Y (2019) A gene expression map of shoot domains reveals regulatory mechanisms. Nat Commun 10(1):141. https://doi.org/10.1038/s41467-018-08083-z
Tian C, Zhang X, He J, Yu H, Wang Y, Shi B, Han Y, Wang G, Feng X, Zhang C, Wang J, Qi J, Yu R, Jiao Y (2014) An organ boundary-enriched gene regulatory network uncovers regulatory hierarchies underlying axillary meristem initiation. Mol Syst Biol 10:755. https://doi.org/10.15252/msb.20145470
Greb T, Clarenz O, Schafer E, Müller D, Herrero R, Schmitz G, Theres K (2003) Molecular analysis of the LATERAL SUPPRESSOR gene in Arabidopsis reveals a conserved control mechanism for axillary meristem formation. Genes Dev 17(9):1175–1187. https://doi.org/10.1101/gad.260703
Schumacher K, Schmitt T, Rossberg M, Schmitz G, Theres K (1999) The Lateral suppressor (Ls) gene of tomato encodes a new member of the VHIID protein family. Proc Natl Acad Sci USA 96(1):290–295
Li X, Qian Q, Fu Z, Wang Y, Xiong G, Zeng D, Wang X, Liu X, Teng S, Hiroshi F, Yuan M, Luo D, Han B, Li J (2003) Control of tillering in rice. Nature 422(6932):618–621. https://doi.org/10.1038/nature01518
Xu C, Wang Y, Yu Y, Duan J, Liao Z, Xiong G, Meng X, Liu G, Qian Q, Li J (2012) Degradation of MONOCULM 1 by APC/CTAD1 regulates rice tillering. Nat Commun 3:750. https://doi.org/10.1038/ncomms1743
Lin Q, Wang D, Dong H, Gu S, Cheng Z, Gong J, Qin R, Jiang L, Li G, Wang J-L, Wu F, Guo X, Zhang X, Lei C, Wang H, Wan J (2012) Rice APC/CTE controls tillering by mediating the degradation of MONOCULM 1. Nat Commun 3:752. https://doi.org/10.1038/ncomms1716
Shao G, Lu Z, Xiong J, Wang B, Jing Y, Meng X, Liu G, Ma H, Liang Y, Chen F, Wang Y, Li J, Yu H (2019) Tiller bud formation regulators MOC1 and MOC3 cooperatively promote tiller bud outgrowth by activating FON1 expression in rice. Mol Plant 12(8):1090–1102. https://doi.org/10.1016/j.molp.2019.04.008
Schmitz G, Tillmann E, Carriero F, Fiore C, Cellini F, Theres K (2002) The tomato Blind gene encodes a MYB transcription factor that controls the formation of lateral meristems. Proc Natl Acad Sci USA 99(2):1064–1069. https://doi.org/10.1073/pnas.022516199
Müller D, Schmitz G, Theres K (2006) Blind homologous R2R3 Myb genes control the pattern of lateral meristem initiation in Arabidopsis. Plant Cell 18(3):586–597. https://doi.org/10.1105/tpc.105.038745
Keller T, Abbott J, Moritz T, Doerner P (2006) Arabidopsis REGULATOR OF AXILLARY MERISTEMS1 controls a leaf axil stem cell niche and modulates vegetative development. Plant Cell 18(3):598–611. https://doi.org/10.1105/tpc.105.038588
Aida M, Ishida T, Tasaka M (1999) Shoot apical meristem and cotyledon formation during Arabidopsis embryogenesis: interaction among the CUP-SHAPED COTYLEDON and SHOOT MERISTEMLESS genes. Development 126(8):1563–1570
Weir I, Lu J, Cook H, Causier B, Schwarz-Sommer Z, Davies B (2004) CUPULIFORMIS establishes lateral organ boundaries in Antirrhinum. Development 131(4):915–922. https://doi.org/10.1242/dev.00993
Souer E, van Houwelingen A, Kloos D, Mol J, Koes R (1996) The no apical meristem gene of petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 85(2):159–170. https://doi.org/10.1016/s0092-8674(00)81093-4
Berger Y, Harpaz-Saad S, Brand A, Melnik H, Sirding N, Alvarez JP, Zinder M, Samach A, Eshed Y, Ori N (2009) The NAC-domain transcription factor GOBLET specifies leaflet boundaries in compound tomato leaves. Development 136(5):823–832. https://doi.org/10.1242/dev.031625
Peaucelle A, Morin H, Traas J, Laufs P (2007) Plants expressing a miR164-resistant CUC2 gene reveal the importance of post-meristematic maintenance of phyllotaxy in Arabidopsis. Development 134(6):1045–1050. https://doi.org/10.1242/dev.02774
Nikovics K, Blein T, Peaucelle A, Ishida T, Morin H, Aida M, Laufs P (2006) The balance between the MIR164A and CUC2 genes controls leaf margin serration in Arabidopsis. Plant Cell 18(11):2929–2945. https://doi.org/10.1105/tpc.106.045617
Laufs P, Peaucelle A, Morin H, Traas J (2004) MicroRNA regulation of the CUC genes is required for boundary size control in Arabidopsis meristems. Development 131(17):4311–4322. https://doi.org/10.1242/dev.01320
Blein T, Pulido A, Vialette-Guiraud A, Nikovics K, Morin H, Hay A, Johansen IE, Tsiantis M, Laufs P (2008) A conserved molecular framework for compound leaf development. Science 322(5909):1835–1839. https://doi.org/10.1126/science.1166168
Nahar MA, Ishida T, Smyth DR, Tasaka M, Aida M (2012) Interactions of CUP-SHAPED COTYLEDON and SPATULA genes control carpel margin development in Arabidopsis thaliana. Plant Cell Physiol 53(6):1134–1143. https://doi.org/10.1093/pcp/pcs057
Bilsborough GD, Runions A, Barkoulas M, Jenkins HW, Hasson A, Galinha C, Laufs P, Hay A, Prusinkiewicz P, Tsiantis M (2011) Model for the regulation of Arabidopsis thaliana leaf margin development. Proc Natl Acad Sci USA 108(8):3424–3429. https://doi.org/10.1073/pnas.1015162108
Sieber P, Wellmer F, Gheyselinck J, Riechmann JL, Meyerowitz EM (2007) Redundancy and specialization among plant microRNAs: role of the MIR164 family in developmental robustness. Development 134(6):1051–1060. https://doi.org/10.1242/dev.02817
Furutani M, Vernoux T, Traas J, Kato T, Tasaka M, Aida M (2004) PIN-FORMED1 and PINOID regulate boundary formation and cotyledon development in Arabidopsis embryogenesis. Development 131(20):5021–5030. https://doi.org/10.1242/dev.01388
Vroemen CW, Mordhorst AP, Albrecht C, Kwaaitaal MA, de Vries SC (2003) The CUP-SHAPED COTYLEDON3 gene is required for boundary and shoot meristem formation in Arabidopsis. Plant Cell 15(7):1563–1577
Raman S, Greb T, Peaucelle A, Blein T, Laufs P, Theres K (2008) Interplay of miR164, CUP-SHAPED COTYLEDON genes and LATERAL SUPPRESSOR controls axillary meristem formation in Arabidopsis thaliana. Plant J 55(1):65–76. https://doi.org/10.1111/j.1365-313X.2008.03483.x
Hibara K, Karim MR, Takada S, Taoka K, Furutani M, Aida M, Tasaka M (2006) Arabidopsis CUP-SHAPED COTYLEDON3 regulates postembryonic shoot meristem and organ boundary formation. Plant Cell 18(11):2946–2957. https://doi.org/10.1105/tpc.106.045716
Takada S, Hibara K, Ishida T, Tasaka M (2001) The CUP-SHAPED COTYLEDON1 gene of Arabidopsis regulates shoot apical meristem formation. Development 128(7):1127–1135
Scofield S, Murison A, Jones A, Fozard J, Aida M, Band LR, Bennett M, Murray JAH (2018) Coordination of meristem and boundary functions by transcription factors in the SHOOT MERISTEMLESS regulatory network. Development 145(9):dev157081. https://doi.org/10.1242/dev.157081
Spinelli SV, Martin AP, Viola IL, Gonzalez DH, Palatnik JF (2011) A mechanistic link between STM and CUC1 during Arabidopsis development. Plant Physiol 156(4):1894–1904. https://doi.org/10.1104/pp.111.177709
Talbert PB, Adler HT, Parks DW, Comai L (1995) The REVOLUTA gene is necessary for apical meristem development and for limiting cell divisions in the leaves and stems of Arabidopsis thaliana. Development 121(9):2723–2735
Zhong R, Ye Z-H (1999) IFL1, a gene regulating interfascicular fiber differentiation in Arabidopsis, encodes a homeodomain-leucine zipper protein. Plant Cell 11(11):2139–2152
Prigge MJ, Otsuga D, Alonso JM, Ecker JR, Drews GN, Clark SE (2005) Class III homeodomain-leucine zipper gene family members have overlapping, antagonistic, and distinct roles in Arabidopsis development. Plant Cell 17(1):61–76. https://doi.org/10.1105/tpc.104.026161
Otsuga D, DeGuzman B, Prigge MJ, Drews GN, Clark SE (2001) REVOLUTA regulates meristem initiation at lateral positions. Plant J 25(2):223–236
McConnell JR, Emery J, Eshed Y, Bao N, Bowman J, Barton MK (2001) Role of PHABULOSA and PHAVOLUTA in determining radial patterning in shoots. Nature 411(6838):709–713. https://doi.org/10.1038/35079635
Mallory AC, Reinhart BJ, Jones-Rhoades MW, Tang G, Zamore PD, Barton MK, Bartel DP (2004) MicroRNA control of PHABULOSA in leaf development: importance of pairing to the microRNA 5′ region. EMBO J 23(16):3356–3364. https://doi.org/10.1038/sj.emboj.7600340
Williams L, Grigg SP, Xie M, Christensen S, Fletcher JC (2005) Regulation of Arabidopsis shoot apical meristem and lateral organ formation by microRNA miR166g and its AtHD-ZIP target genes. Development 132(16):3657–3668. https://doi.org/10.1242/dev.01942
McConnell JR, Barton MK (1998) Leaf polarity and meristem formation in Arabidopsis. Development 125(15):2935–2942
Eshed Y, Baum SF, Perea JV, Bowman JL (2001) Establishment of polarity in lateral organs of plants. Curr Biol 11(16):1251–1260
Zhang T, Li Y, Ma L, Sang X, Ling Y, Wang Y, Yu P, Zhuang H, Huang J, Wang N, Zhao F, Zhang C, Yang Z, Fang L, He G (2017) LATERAL FLORET 1 induced the three-florets spikelet in rice. Proc Natl Acad Sci USA 114(37):9984–9989. https://doi.org/10.1073/pnas.1700504114
Tian C, Jiao Y (2015) A systems approach to understand shoot branching. Curr Plant Biol 3–4:13–19. https://doi.org/10.1016/j.cpb.2015.08.001
Komatsu K, Maekawa M, Ujiie S, Satake Y, Furutani I, Okamoto H, Shimamoto K, Kyozuka J (2003) LAX and SPA: major regulators of shoot branching in rice. Proc Natl Acad Sci USA 100(20):11765–11770. https://doi.org/10.1073/pnas.1932414100
Oikawa T, Kyozuka J (2009) Two-step regulation of LAX PANICLE1 protein accumulation in axillary meristem formation in rice. Plant Cell 21(4):1095–1108. https://doi.org/10.1105/tpc.108.065425
Gallavotti A, Zhao Q, Kyozuka J, Meeley RB, Ritter MK, Doebley JF, Pe ME, Schmidt RJ (2004) The role of barren stalk1 in the architecture of maize. Nature 432(7017):630–635. https://doi.org/10.1038/nature03148
Woods DP, Hope CL, Malcomber ST (2011) Phylogenomic analyses of the BARREN STALK1/LAX PANICLE1 (BA1/LAX1) genes and evidence for their roles during axillary meristem development. Mol Biol Evol 28(7):2147–2159. https://doi.org/10.1093/molbev/msr036
Yang F, Wang Q, Schmitz G, Müller D, Theres K (2012) The bHLH protein ROX acts in concert with RAX1 and LAS to modulate axillary meristem formation in Arabidopsis. Plant J 71(1):61–70. https://doi.org/10.1111/j.1365-313X.2012.04970.x
Basile A, Fambrini M, Tani C, Shukla V, Licausi F, Pugliesi C (2019) The Ha-ROXL gene is required for initiation of axillary and floral meristems in sunflower. Genesis 57(9):e23307. https://doi.org/10.1002/dvg.23307
Fambrini M, Salvini M, Pugliesi C (2017) Molecular cloning, phylogenetic analysis, and expression patterns of LATERAL SUPPRESSOR-LIKE and REGULATOR OF AXILLARY MERISTEM FORMATION-LIKE genes in sunflower (Helianthus annuus L.). Dev Genes Evol 227(2):159–170. https://doi.org/10.1007/s00427-016-0571-2
Tabuchi H, Zhang Y, Hattori S, Omae M, Shimizu-Sato S, Oikawa T, Qian Q, Nishimura M, Kitano H, Xie H, Fang X, Yoshida H, Kyozuka J, Chen F, Sato Y (2011) LAX PANICLE2 of rice encodes a novel nuclear protein and regulates the formation of axillary meristems. Plant Cell 23(9):3276–3287. https://doi.org/10.1105/tpc.111.088765
Yao H, Skirpan A, Wardell B, Matthes MS, Best NB, McCubbin T, Durbak A, Smith T, Malcomber S, McSteen P (2019) The barren stalk2 gene is required for axillary meristem development in maize. Mol Plant 12(3):374–389. https://doi.org/10.1016/j.molp.2018.12.024
Matsuo N, Mase H, Makino M, Takahashi H, Banno H (2009) Identification of ENHANCER OF SHOOT REGENERATION 1-upregulated genes during in vitro shoot regeneration. Plant Biotechnol 26(4):385–393
Ikeda Y, Banno H, Niu Q-W, Howell SH, Chua N-H (2006) The ENHANCER OFSHOOT REGENERATION 2 gene in Arabidopsis regulates CUP-SHAPED COTYLEDON 1 at the transcriptional level and controls cotyledon development. Plant Cell Physiol 47(11):1443–1456. https://doi.org/10.1093/Pcp/Pcl023
Guo D, Zhang J, Wang X, Han X, Wei B, Wang J, Li B, Yu H, Huang Q, Gu H, Qu L-J, Qin G (2015) The WRKY transcription factor WRKY71/EXB1 controls shoot branching by transcriptionally regulating RAX genes in Arabidopsis. Plant Cell 27(11):3112–3127. https://doi.org/10.1105/tpc.15.00829
Chahtane H, Vachon G, Le Masson M, Thevenon E, Perigon S, Mihajlovic N, Kalinina A, Michard R, Moyroud E, Monniaux M, Sayou C, Grbic V, Parcy F, Tichtinsky G (2013) A variant of LEAFY reveals its capacity to stimulate meristem development by inducing RAX1. Plant J 74(4):678–689. https://doi.org/10.1111/tpj.12156
Hempel FD, Feldman LJ (1994) Bidirectional inflorescence development in Arabidopsis thaliana—acropetal initiation of flowers and basipetal initiation of paraclades. Planta 192(2):276–286
Lee DK, Geisler M, Springer PS (2009) LATERAL ORGAN FUSION1 and LATERAL ORGAN FUSION2 function in lateral organ separation and axillary meristem formation in Arabidopsis. Development 136(14):2423–2432. https://doi.org/10.1242/dev.031971
Naz AA, Raman S, Martinez CC, Sinha NR, Schmitz G, Theres K (2013) Trifoliate encodes an MYB transcription factor that modulates leaf and shoot architecture in tomato. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1214300110
Zhang X, Zhou Y, Ding L, Wu Z, Liu R, Meyerowitz EM (2013) Transcription repressor HANABA TARANU controls flower development by integrating the actions of multiple hormones, floral organ specification genes, and GATA3 family genes in Arabidopsis. Plant Cell 25(1):83–101. https://doi.org/10.1105/tpc.112.107854
Kanei M, Horiguchi G, Tsukaya H (2012) Stable establishment of cotyledon identity during embryogenesis in Arabidopsis by ANGUSTIFOLIA3 and HANABA TARANU. Development 139(13):2436–2446. https://doi.org/10.1242/dev.081547
Nawy T, Bayer M, Mravec J, Friml J, Birnbaum KD, Lukowitz W (2010) The GATA factor HANABA TARANU is required to position the proembryo boundary in the early Arabidopsis embryo. Dev Cell 19(1):103–113. https://doi.org/10.1016/j.devcel.2010.06.004
Zhao Y, Medrano L, Ohashi K, Fletcher JC, Yu H, Sakai H, Meyerowitz EM (2004) HANABA TARANU is a GATA transcription factor that regulates shoot apical meristem and flower development in Arabidopsis. Plant Cell 16(10):2586–2600. https://doi.org/10.1105/tpc.104.024869
Johnston R, Wang M, Sun Q, Sylvester AW, Hake S, Scanlon MJ (2014) Transcriptomic analyses indicate that maize ligule development recapitulates gene expression patterns that occur during lateral organ initiation. Plant Cell 26(12):4718–4732. https://doi.org/10.1105/tpc.114.132688
Ha CM, Jun JH, Nam HG, Fletcher JC (2004) BLADE-ON-PETIOLE1 encodes a BTB/POZ domain protein required for leaf morphogenesis in Arabidopsis thaliana. Plant Cell Physiol 45(10):1361–1370. https://doi.org/10.1093/pcp/pch201
Ha CM, Kim GT, Kim BC, Jun JH, Soh MS, Ueno Y, Machida Y, Tsukaya H, Nam HG (2003) The BLADE-ON-PETIOLE 1 gene controls leaf pattern formation through the modulation of meristematic activity in Arabidopsis. Development 130(1):161–172
McKim SM, Stenvik GE, Butenko MA, Kristiansen W, Cho SK, Hepworth SR, Aalen RB, Haughn GW (2008) The BLADE-ON-PETIOLE genes are essential for abscission zone formation in Arabidopsis. Development 135(8):1537–1546. https://doi.org/10.1242/dev.012807
Hepworth SR, Zhang Y, McKim S, Li X, Haughn GW (2005) BLADE-ON-PETIOLE-dependent signaling controls leaf and floral patterning in Arabidopsis. Plant Cell 17(5):1434–1448. https://doi.org/10.1105/tpc.104.030536
Norberg M, Holmlund M, Nilsson O (2005) The BLADE ON PETIOLE genes act redundantly to control the growth and development of lateral organs. Development 132(9):2203–2213. https://doi.org/10.1242/dev.01815
Tavakol E, Okagaki R, Verderio G, Shariati JV, Hussien A, Bilgic H, Scanlon MJ, Todt NR, Close TJ, Druka A, Waugh R, Steuernagel B, Ariyadasa R, Himmelbach A, Stein N, Muehlbauer GJ, Rossini L (2015) The barley Uniculme4 gene encodes a BLADE-ON-PETIOLE-like protein that controls tillering and leaf patterning. Plant Physiol 168(1):164–174. https://doi.org/10.1104/pp.114.252882
Okagaki RJ, Haaning A, Bilgic H, Heinen S, Druka A, Bayer M, Waugh R, Muehlbauer GJ (2018) ELIGULUM-A regulates lateral branch and leaf development in barley. Plant Physiol 176(4):2750–2760. https://doi.org/10.1104/pp.17.01459
Dong Z, Li W, Unger-Wallace E, Yang J, Vollbrecht E, Chuck G (2017) Ideal crop plant architecture is mediated by tassels replace upper ears1, a BTB/POZ ankyrin repeat gene directly targeted by TEOSINTE BRANCHED1. Proc Natl Acad Sci USA 114(41):E8656–E8664. https://doi.org/10.1073/pnas.1714960114
Xu C, Park SJ, Van Eck J, Lippman ZB (2016) Control of inflorescence architecture in tomato by BTB/POZ transcriptional regulators. Genes Dev 30(18):2048–2061. https://doi.org/10.1101/gad.288415.116
Toriba T, Tokunaga H, Shiga T, Nie F, Naramoto S, Honda E, Tanaka K, Taji T, Itoh JI, Kyozuka J (2019) BLADE-ON-PETIOLE genes temporally and developmentally regulate the sheath to blade ratio of rice leaves. Nat Commun 10(1):619. https://doi.org/10.1038/s41467-019-08479-5
Wang Y, Wang J, Shi B, Yu T, Qi J, Meyerowitz EM, Jiao Y (2014) The stem cell niche in leaf axils is established by auxin and cytokinin in Arabidopsis. Plant Cell 26(5):2055–2067
Wang Q, Kohlen W, Rossmann S, Vernoux T, Theres K (2014) Auxin depletion from the leaf axil conditions competence for axillary meristem formation in Arabidopsis and tomato. Plant Cell 26(5):2068–2079. https://doi.org/10.1105/tpc.114.123059
Brunoud G, Wells DM, Oliva M, Larrieu A, Mirabet V, Burrow AH, Beeckman T, Kepinski S, Traas J, Bennett MJ, Vernoux T (2012) A novel sensor to map auxin response and distribution at high spatio-temporal resolution. Nature 482(7383):103–106. https://doi.org/10.1038/nature10791
Vernoux T, Brunoud G, Farcot E, Morin V, Van den Daele H, Legrand J, Oliva M, Das P, Larrieu A, Wells D, Guedon Y, Armitage L, Picard F, Guyomarc’h S, Cellier C, Parry G, Koumproglou R, Doonan JH, Estelle M, Godin C, Kepinski S, Bennett M, De Veylder L, Traas J (2011) The auxin signalling network translates dynamic input into robust patterning at the shoot apex. Mol Syst Biol 7:508. https://doi.org/10.1038/msb.2011.39
Chung Y, Zhu Y, Wu MF, Simonini S, Kuhn A, Armenta-Medina A, Jin R, Ostergaard L, Gillmor CS, Wagner D (2019) Auxin Response Factors promote organogenesis by chromatin-mediated repression of the pluripotency gene SHOOTMERISTEMLESS. Nat Commun 10(1):886. https://doi.org/10.1038/s41467-019-08861-3
Wu MF, Yamaguchi N, Xiao J, Bargmann B, Estelle M, Sang Y, Wagner D (2015) Auxin-regulated chromatin switch directs acquisition of flower primordium founder fate. eLife 4:e09269
Yamaguchi N, Wu MF, Winter CM, Berns MC, Nole-Wilson S, Yamaguchi A, Coupland G, Krizek BA, Wagner D (2013) A molecular framework for auxin-mediated initiation of flower primordia. Dev Cell 24(3):271–282. https://doi.org/10.1016/j.devcel.2012.12.017
Li W, Zhou Y, Liu X, Yu P, Cohen JD, Meyerowitz EM (2013) LEAFY controls auxin response pathways in floral primordium formation. Sci Signal 6(270):ra23. https://doi.org/10.1126/scisignal.2003937
Burian A, Barbier de Reuille P, Kuhlemeier C (2016) Patterns of stem cell divisions contribute to plant longevity. Curr Biol 26(11):1385–1394. https://doi.org/10.1016/j.cub.2016.03.067
Matthes MS, Best NB, Robil JM, Malcomber S, Gallavotti A, McSteen P (2019) Auxin EvoDevo: conservation and diversification of genes regulating auxin biosynthesis, transport, and signaling. Mol Plant 12(3):298–320. https://doi.org/10.1016/j.molp.2018.12.012
Phillips KA, Skirpan AL, Liu X, Christensen A, Slewinski TL, Hudson C, Barazesh S, Cohen JD, Malcomber S, McSteen P (2011) Vanishing tassel2 encodes a grass-specific tryptophan aminotransferase required for vegetative and reproductive development in maize. Plant Cell 23(2):550–566. https://doi.org/10.1105/tpc.110.075267
Gallavotti A, Barazesh S, Malcomber S, Hall D, Jackson D, Schmidt RJ, McSteen P (2008) Sparse inflorescence1 encodes a monocot-specific YUCCA-like gene required for vegetative and reproductive development in maize. Proc Natl Acad Sci USA 105(39):15196–15201. https://doi.org/10.1073/pnas.0805596105
Luo L, Zeng J, Wu H, Tian Z, Zhao Z (2018) A molecular framework for auxin-controlled homeostasis of shoot stem cells in Arabidopsis. Mol Plant 11(7):899–913. https://doi.org/10.1016/j.molp.2018.04.006
Gordon SP, Chickarmane VS, Ohno C, Meyerowitz EM (2009) Multiple feedback loops through cytokinin signaling control stem cell number within the Arabidopsis shoot meristem. Proc Natl Acad Sci USA 106(38):16529–16534. https://doi.org/10.1073/pnas.0908122106
Bartrina I, Otto E, Strnad M, Werner T, Schmülling T (2011) Cytokinin regulates the activity of reproductive meristems, flower organ size, ovule formation, and thus seed yield in Arabidopsis thaliana. Plant Cell 23(1):69–80. https://doi.org/10.1105/tpc.110.079079
Chickarmane VS, Gordon SP, Tarr PT, Heisler MG, Meyerowitz EM (2012) Cytokinin signaling as a positional cue for patterning the apical-basal axis of the growing Arabidopsis shoot meristem. Proc Natl Acad Sci USA 109(10):4002–4007. https://doi.org/10.1073/pnas.1200636109
Snipes SA, Rodriguez K, DeVries AE, Miyawaki KN, Perales M, Xie M, Reddy GV (2018) Cytokinin stabilizes WUSCHEL by acting on the protein domains required for nuclear enrichment and transcription. PLoS Genet 14(4):e1007351. https://doi.org/10.1371/journal.pgen.1007351
Su Y-H, Liu Y-B, Zhang X-S (2011) Auxin-cytokinin interaction regulates meristem development. Mol Plant 4(4):616–625. https://doi.org/10.1093/mp/ssr007
Müller B, Sheen J (2008) Cytokinin and auxin interaction in root stem-cell specification during early embryogenesis. Nature 453(7198):1094–1097. https://doi.org/10.1038/nature06943
Yanai O, Shani E, Dolezal K, Tarkowski P, Sablowski R, Sandberg G, Samach A, Ori N (2005) Arabidopsis KNOXI proteins activate cytokinin biosynthesis. Curr Biol 15(17):1566–1571. https://doi.org/10.1016/j.cub.2005.07.060
Sakamoto T, Sakakibara H, Kojima M, Yamamoto Y, Nagasaki H, Inukai Y, Sato Y, Matsuoka M (2006) Ectopic expression of KNOTTED1-like homeobox protein induces expression of cytokinin biosynthesis genes in rice. Plant Physiol 142(1):54–62. https://doi.org/10.1104/pp.106.085811
Jasinski S, Piazza P, Craft J, Hay A, Woolley L, Rieu I, Phillips A, Hedden P, Tsiantis M (2005) KNOX action in Arabidopsis is mediated by coordinate regulation of cytokinin and gibberellin activities. Curr Biol 15(17):1560–1565. https://doi.org/10.1016/j.cub.2005.07.023
Zhang T-Q, Lian H, Zhou C-M, Xu L, Jiao Y, Wang J-W (2017) A two-step model for de novo activation of WUSCHEL during plant shoot regeneration. Plant Cell 29(5):1073–1087. https://doi.org/10.1105/tpc.16.00863
Meng WJ, Cheng ZJ, Sang YL, Zhang MM, Rong XF, Wang ZW, Tang YY, Zhang XS (2017) Type-B ARABIDOPSIS RESPONSE REGULATORs specify the shoot stem cell niche by dual regulation of WUSCHEL. Plant Cell 29(6):1357–1372. https://doi.org/10.1105/tpc.16.00640
Rupp HM, Frank M, Werner T, Strnad M, Schmülling T (1999) Increased steady state mRNA levels of the STM and KNAT1 homeobox genes in cytokinin overproducing Arabidopsis thaliana indicate a role for cytokinins in the shoot apical meristem. Plant J 18(5):557–563
Zhang Q-Q, Wang J-G, Wang L-Y, Wang J-F, Wang Q, Yu P, Bai M-Y, Fan M (2019) Gibberellin repression of axillary bud formation in Arabidopsis by modulation of DELLA-SPL9 complex activity. J Integr Plant Biol. https://doi.org/10.1111/jipb.12818
Yu S, Galvao VC, Zhang Y-C, Horrer D, Zhang T-Q, Hao Y-H, Feng Y-Q, Wang S, Schmid M, Wang J-W (2012) Gibberellin regulates the Arabidopsis floral transition through miR156-targeted SQUAMOSA promoter binding-like transcription factors. Plant Cell 24(8):3320–3332. https://doi.org/10.1105/tpc.112.101014
Gendron JM, Liu J-S, Fan M, Bai M-Y, Wenkel S, Springer PS, Barton MK, Wang Z-Y (2012) Brassinosteroids regulate organ boundary formation in the shoot apical meristem of Arabidopsis. Proc Natl Acad Sci USA 109(51):21152–21157. https://doi.org/10.1073/pnas.1210799110
Bell EM, Lin WC, Husbands AY, Yu L, Jaganatha V, Jablonska B, Mangeon A, Neff MM, Girke T, Springer PS (2012) Arabidopsis LATERAL ORGAN BOUNDARIES negatively regulates brassinosteroid accumulation to limit growth in organ boundaries. Proc Natl Acad Sci USA 109(51):21146–21151. https://doi.org/10.1073/pnas.1210789109
Schubert D, Primavesi L, Bishopp A, Roberts G, Doonan J, Jenuwein T, Goodrich J (2006) Silencing by plant Polycomb-group genes requires dispersed trimethylation of histone H3 at lysine 27. EMBO J 25(19):4638–4649. https://doi.org/10.1038/sj.emboj.7601311
Liu X, Kim YJ, Müller R, Yumul RE, Liu C, Pan Y, Cao X, Goodrich J, Chen X (2011) AGAMOUS terminates floral stem cell maintenance in Arabidopsis by directly repressing WUSCHEL through recruitment of Polycomb group proteins. Plant Cell 23(10):3654–3670. https://doi.org/10.1105/tpc.111.091538
Acknowledgements
We apologize to those authors whose work could not be cited due to space limitations. Research in our laboratory is funded by the National Natural Science Foundation of China (Grants 31430010, 31825002, and 31861130355), the Frontier Science Key Research Project of the Chinese Academy of Sciences (Grant ZDBS-LY-SM012), the Strategic Priority Research Program of the Chinese Academy of Sciences (Grant XDA24020203), and a Royal Society Newton Advanced Fellowship (NAF\R1\180125).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cao, X., Jiao, Y. Control of cell fate during axillary meristem initiation. Cell. Mol. Life Sci. 77, 2343–2354 (2020). https://doi.org/10.1007/s00018-019-03407-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-019-03407-8