Abstract
Stem cells are endowed with the awesome power of self-renewal and multi-lineage differentiation that allows them to be major contributors to tissue homeostasis. Owing to their longevity and self-renewal capacity, they are also faced with a higher risk of genomic damage compared to differentiated cells. Damage on the genome, if not prevented or repaired properly, will threaten the survival of stem cells and culminate in organ failure, premature aging, or cancer formation. It is therefore of paramount importance that stem cells remain genomically stable throughout life. Given their unique biological and functional requirement, stem cells are thought to manage genotoxic stress somewhat differently from non-stem cells. The focus of this article is to review the current knowledge on how stem cells escape the barrage of oxidative and replicative DNA damage to stay in self-renewal. A clear statement on this subject should help us better understand tissue regeneration, aging, and cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Adult animals have developed distinct mechanisms to replenish the inevitable loss of cells that occurs daily as a result of normal attrition or accidental injury. In lower organisms with simple body plan, tissue regeneration may be carried out by some differentiated cells that can be reversed back to an undifferentiated state and then redirected into cell fates other than their original ones. This process is known as de-differentiation. In higher vertebrates, de-differentiation may cause undesired consequences of incomplete or even faulty differentiation due to the changing environment and the complexity of tissue organization. To avoid such aberrancy, a small population of cells with tissue-restricted potential (known as somatic, tissue-specific, or tissue-resident stem cells) are embedded within the organs that require constant renewal. Those tissue-specific stem cells persist throughout life and retain the ability to: (1) become all cell types cognate to their resident tissues (i.e. multipotential), and (2) maintain their population throughout continuous division (i.e. self-renewal) [1]. A vast amount of studies done in the 1990s have concluded that such stem-like cells not only exist in organs that turn over regularly but also can be isolated from those with no apparent regenerative capability, which even include the regeneration-unfriendly neural tissue [2, 3]. This revelation inspires the idea that stem or stem-like cells may be embedded in every organ and rekindled if necessary. So begins the era of stem cells!
The power of stem cells can work as a double-edged sword. On the bright side, self-renewal ensures that stem cell populations are not depleted over time so that they can provide an inexhaustible source for cell replacement in vivo and in therapeutics [4]. To date, scientists are still trying to unleash the de novo regenerative power of stem cells in tissues that are either non-regenerative by nature or capable of regeneration but decompensated by diseases, injuries, or the aging process. On the dark side, the self-renewal-driving machinery may be hijacked by transformed cells to achieve replicative immortality [5]. On this conceptual ground thus rises the cancer stem cell (CSC) theory, which postulates that there are stem-like cells in tumors that are tumorigenic and sit atop the tumor cellular hierarchy [6, 7]. In the 2000s, the existence of CSCs has been extensively researched and experimentally shown in acute myeloid leukemia [8, 9], breast cancers [10], brain tumors [11], and other types of solid tumors [12–16].
Because of their long lifespan and self-renewal properties, normal stem cells and CSCs (hereafter collectively referred to as stem cells unless otherwise specified) are thought to be uniquely equipped to deal with the occurrence and consequence of genomic damage by ways different from short-term dividing or non-dividing cells. This notion starts to gain a stronger foothold when more and more studies are conducted that deepen our understanding of how stem cells balance between self-renewal and genome preservation [17–20]. Indeed, embryonic stem (ES) cells display a lower mutation rate compared to somatic cells despite their robust mitotic activity [21]. In support, it has been demonstrated that mice deficient in one or more components in the DNA repair pathways, such as ataxia telangiectasia mutated (ATM), DNA ligase 4 (LIG4) [22], DNA-dependent protein kinase catalytic subunit (DNA-PKcs), mutS homolog 2 (MSH2), and Fanconi anemia complementation group D1 (FANCD1), show limited stem cell functions in various tissues [23–32]. Although the exact mechanisms by which stem cells preserve their genome integrity throughout self-renewal may not be entirely clear as yet, I believe that a focused review on this subject will help gather the much needed interest and momentum to this field to inspire new research directions in the future. This article will take on this task from four broad angles that cover damage prevention, stalled replication restart, damage repair, and outcome selection in stem versus non-stem cells (Fig. 1). It will not emphasize as much on how perturbation of these various pathways may affect stem cell functions as on how stem cells differ from other dividing cells in their ways of dealing with genomic stress.
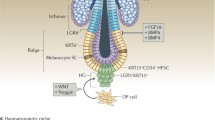
Genomic stress management in stem cells. DNA damage can occur spontaneously as a result of genome replication and cell metabolism or reactively to exogenous insults (gray boxes). Avoiding genomic damage and instability is critical for stem cells to maintain their longevity and self-renewal potential. Stem cells may achieve this goal by employing four strategies that: (1) prevent damage at first sight (pink boxes); (2) restart stalled replication forks (blue box); (3) repair/cosegregate genomic damage (green boxes); or (4) select the least harmful outcome out of many (yellow boxes). The green and red arrows indicate a causative effect or a suppressive effect, respectively. ABC ATP-binding cassette transporter, ROS reactive oxygen species, TLS translesion synthesis, TS template switching, HR homologous recombination, NHEJ non-homologous end-joining, BER base excision repair
Nip in the bud: controlling damage at first sight
No cell can avoid the risk of genotoxic damage, especially those that enjoy the benefit of a long and productive life. Not only is the genome under constant attack from extrinsic sources of insults, but it is also faced with damages that arise internally as a result of genome replication, hydrolytic cleavage, which causes DNA deamination or depurination, or reaction to reactive oxygen, nitrogen, or carbonyl species produced during mitochondrial respiration [33]. It is estimated that there may be up to 106 DNA damage events occurring in a single cell on a daily basis [34]. The two major cell-extrinsic sources of genome-damaging insults are chemoreagents and ultraviolet (UV) radiation. It has been shown that stem cells express higher levels of ATP-binding cassette (ABC) transporters or multidrug resistance (MDR) genes, which pump out intracellular drugs and lower their amounts inside the cell [35]. So far, there is no evidence indicating that stem cells are less likely exposed to UV or ionizing radiation (IR) than their neighboring non-stem cells.
Genome replication itself is an intrinsic source of double-stranded breaks (DSBs). Replicative DNA damage can happen as a result of three naturally occurring events. First of all, the movement of the replication machinery (also known as the replisome) may be stalled at the sites that are: (1) previously damaged and unrepaired, (2) forming complex secondary structure, (3) bound by protein complexes, or (4) containing fragile DNA or repeat sequences (e.g. ribosomal DNA, telomere, and Alu). It may also be triggered by exogenous chemicals or drugs that block the activity of DNA replication machinery or deplete the endogenous nucleotide pool [36–38]. Stalled replication forks, if unresolved in time, may collapse and result in DSBs. Alternatively, DSBs may be generated spontaneously during genome replication when replisomes encounter single-stranded breaks (SSBs). Lastly, replication of the chromosomal ends will introduce telomere attrition as a result of the end-replication problem. As a repeat sequence itself, the telomere is also subjected to DNA damage during the genome replication process. Mitotic quiescence offers stem cells the first line of defense against issues arising from genome replication [39]. Many adult tissues harbor a subset of stem cells held in a mitotically quiescent state by their microenvironment for most of the time. However, mitotic quiescence is not a one-size-fits-all solution. Embryonic stem (ES) cells, induced pluripotent stem (iPS) cells, embryonic tissue-specific stem cells, some adult tissue-specific stem cells during injury, and CSCs undergo active self-renewal. Mouse ES cells, in particular, have a shortened G1 phase and an inactive G1/S checkpoint control [40]. Similar cell cycle profiles have also been observed in human ES cells and iPS cells [41]. Consequently, a majority (50–70 %) of these pluripotent stem cells are in the S-phase and hence are susceptible to replication-induced DNA damage [40]. For those cells, a second or third line of defense is needed.
Oxidative stress is another endogenous source of genotoxic insult and is the leading cause of DNA damage in quiescent stem cells. Quiescent stem cells exist in some adult tissues (e.g. hematopoietic stem cells in the bone marrow, bulge stem cells in the hair follicle, and crypt stem cells in the intestinal epithelium), remain mitotically inactive, and undergo asymmetric cell division only when needed. As quiescent stem cells remain largely in the G0 phase, their genome is exempt from replicative DNA damage. Despite that, they are still faced with oxidative stress produced by endogenous mitochondrial respiration and exposure to exogenous UV or chemicals. It is estimated that reactive oxygen species (ROS), which include superoxide radicals, hydroxyl free radicals, and hydrogen peroxide, damage 104 bases per day in a human cell [42]. Oxidative stress can create oxidized nucleotides (e.g. 8-oxyguanine), SSBs, and DNA hydrolysis (which leads to abasic or deaminated lesions). It has been shown that ROS have a profound effect in limiting the lifespan of hematopoietic stem cells [43–45]. Fortunately, the ROS level in hematopoietic stem cells is 100-fold less than that in myeloid progenitors. Reduction of ROS in stem cells may be controlled by their high levels of FoxO transcription factors, which operate downstream of the PI3K-AKT pathway and regulate the expression of ROS detoxication genes, including superoxide dismutase 2 (SOD2) and catalase [46–49]. In consistence, the FoxO pathway is required for the maintenance of leukemic CSCs in chronic myeloid leukemia [50]. Bmi1 (a polycomb RING-finger protein) can also help to reduce the generation of ROS from the inside of stem cells [51]. Other than those cell-intrinsic programs, cell-extrinsic factors may play a role as well. Some stem cells are known to reside in a hypoxic microenvironment. The hypoxic stem cell niche has been shown for hematopoietic stem cells [52, 53], intestinal stem cells [54], and breast CSCs [55], and supported by the low oxygen culture condition for neural and hematopoietic stem cells [56, 57]. Within the hypoxic niches, stem cells employ anaerobic glycolysis instead of mitochondrial oxidative phosphorylation for energy metabolism—a decision that may be driven by their low mitochondrial mass and a HIF-1α-controlled mechanism [58–61]. The preferential use of a selective energy pathway also helps to lower the intracellular ROS level of stem cells. However, some studies seem to contradict the link between stem cells and hypoxia. One study showed that the self-renewal of proliferating neural stem cells is propelled by a high ROS level [62]. Another study showed that breast CSCs contain abundant mitochondria, and the enrichment of mitochondria in breast CSCs is driven by the Wnt1/FGF3 pathway [63]. Counter-intuitively, high mitochondrial mass appears to promote the resistance of breast CSCs to DNA damage [64].
A window of opportunity: restarting stalled replication forks before they collapse
As prolonged replication stalling may lead to replication fork collapse, DSBs, and chromosomal rearrangement, all dividing cells must learn how to resolve replication stalling efficiently as their next line of defense [65, 66]. The resolution of stalled replication forks consists of a sensing step and a bypass step. To date, the molecular mechanisms underlying each of these two events have just begun to emerge. As a result, it has yet to be determined whether and how these events operate in a stem cell-specific setting. For this reason, this review will discuss the current knowledge on how cells in general manage to restart the stalled replication fork, with the anticipation that stem cell-unique regulation on some of the pathways may be uncovered in the future.
Sensing replication stalling
Replication fork stalling takes place routinely during genome replication [67, 68]. To finish self-renewal and preserve genome integrity efficiently, mitotically active stem cells need to learn how to reinitiate or bypass the stalled site when they encounter one. Failure to do so may result in the collapse of replication forks that ultimately leads to DSBs. The signaling cascade triggered by replication stalling begins with the uncoupling of DNA polymerase and helicase and the formation of replication protein A (RPA)-coated single-stranded DNAs (ssDNA). DNA-bound RPA then recruits the ataxia telangiectasia and Rad3 related (ATR), Rad17-Rfc2-5, the 9-1-1 clamp, and topoisomerase II binding protein 1 (TopBP1) to the stalled replication site, which serves the function of triggering G2/M arrest and stabilizing the stalled replication fork [69–72]. TopBP1 also plays a role in DNA replication initiation and is needed for neural progenitors to maintain their genome integrity and reduce replication-associated DNA damage during neural development [73]. Although not completely characterized as yet, the mechanisms by which cells reengage the stalled replication machinery, bypass the lesion site, and restart processive replication can follow either an error-prone translesion synthesis (TLS) pathway or an error-free template-switching pathway [74].
Translesion synthesis (TLS)
Compared to the replicative DNA polymerases [e.g. Pol δ (delta)], those specialized in TLS [e.g. Pol η (eta) and Pol ζ (zeta)] have broader active sites, exhibit lower processivity, and lack 3′-to-5′ exonuclease editing. The type of TLS polymerases recruited to the stalled site will also determine the bypass fidelity. For example, UV-induced pyrimidine dimers can be bypassed by using Pol η in a relatively error-free mode or by using Pol ζ and Rev1 in an error-prone mode [75–77]. A recent study showed that ovarian CSCs express high levels of Pol η, which may contribute to their cisplatin resistance [78]. The TLS mechanism can be viewed as a cycle of DNA polymerase switching, orchestrated by the monoubiquitinylation and deubiquitinylation of proliferating cell nuclear antigen (PCNA) (Fig. 2). During regular DNA replication, PCNA works as a sliding clamp around DNAs and plays an important role in the switching of replicative polymerase from the primase-Pol α [alpha] complex to the processive Pol δ. The DNA loading and unloading of PCNA is controlled by the arc-shaped replication factor C (RFC) complex (Rfc1-5) [79]. At the UV-damaged site, PCNA is monoubiquitinylated on K164 by the RAD6 (E2) and RAD18 (E3) heterodimer [80–83]. Monoubiquitinylated PCNA promotes direct lesion bypass by recruiting TLS polymerases (Pol η and Pol ζ) to the stalled replication fork through the ubiquitin-binding domain present on all Y-family polymerases [84–88]. In addition to PCNA monoubiquitination, Pol η can also be recruited to the UV-damaged site by its direct interaction with RAD18 [85]. Switching between different polymerases allows cells to use Pol η to add the first adenine across the TT dimer, Pol ζ to extend the mismatch, and Pol δ to continue with the rest of DNA replication. Another molecule involved in TLS is C1orf24. C1orf24 is a PCNA-binding protein that stabilizes RAD18 localization, promotes PCNA monoubiquitinylation, and performs polymerase switching from Pol δ to Pol η at the UV-induced DNA damage site by a Valosin-containing protein-dependent mechanism [89]. BAF180 also participates in TLS. BAF180 is the human ortholog of yeast RSC1–RSC2–RSC4 fusion and a component in the chromatin-remodeling complex that consists of BAF57, BAF200, and BRG1 (SWI/SNF core complex) [90]. Depletion of BAF180 has been shown to reduce PCNA ubiquitinylation as well as the chromatin-bound unmodified PCNA after UV radiation. As depletion of BAF180 does not diminish chromatin-bound RPA, it may promote the bypass by remodeling the chromatin structure to support the switching of TLS polymerases and the repriming of PCNA.
Translesion synthesis (TLS) represents a cycle of polymerase switching driven by the ubiquitinylation of PCNA. TLS allows replication forks to bypass the stalled sites. The key event underlying TLS is the monoubiquitinylation of PCNA, mediated by Rad6 and Rad18. Monoubiquitinylation of PCNA and its deubiquitinylation by USP1 trigger a cycle of polymerase (Pol) switching between the processive Pol δ and the permissive Pol η and Pol ζ. Red asterisks mutations, U monoubiquitinylation, 5′ primase/RNAs, purple circles replication protein A (RPA)
Two regulatory events have been reported that negatively control TLS. One involves FbH1, an UvrD DNA repair helicase and the human functional analog of Srs2. It was shown that FbH1 overexpression weakens blocked replication-induced homologous recombination (HR) and reduces nuclear RAD51 foci, suggesting that FbH1 may prevent HR repair by restraining RAD51 localization at the stalled replication site [91]. Moreover, FbH1-deficient cells are hyposensitive to replication stress induced by hydroxyurea, show a reduced activation of ATM and p53, and exhibit better survival with decreased apoptosis [92, 93]. The negative function of FbH1 at the stalled replication site is regulated by its interaction with RPA and PCNA [93, 94] as well as by the impaired recruitment of Pol η to UV-damaged chromatin [94]. Another negative regulator of TLS is the SUMOylation of PCNA. It has been shown that SUMO modification of yeast PCNA negatively affects HR by granting access to the Srs2 helicase (the functional equivalent of human FbH1) to disrupt the RAD51 nucleoprotein filament [95] and by interfering with Eco1-dependent sister chromatid cohesion [96]. In human cells, SUMO modification of PCNA is facilitated by RFC and functions to prevent replication fork collapse into DSBs [97]. If the replication fork stalls at DNA lesion sites, SUMOylated PCNA exhibits the ability to inhibit HR repair [97].
Template switching
Stalled replication can be restarted by an error-free mechanism that involves the polyubiquitinylation of PCNA and template switching. Addition of K63-linked polyubiquitin chains on PCNA requires RAD5 and the MMS2–UBC13 complex, but may or may not take place directly on the monoubiquitinylated PCNA [98, 99]. RAD5 is a member of the SWI/SNF family. It interacts with a heterodimeric E2 enzyme, MMS2–UBC13, to promote methyl methanesulfonate-induced PCNA polyubiquitinylation [100, 101]. Human RAD5 homolog, SNF2 histone-linker PHD ring helicase (SHPRH), is located on chromosome 6q24 and acts as a tumor suppressor in addition to its E2 ubiquitin ligase role [102]. RAD5 also works with RAD6–RAD18 to promote PCNA monoubiquitinylation. Helicase-like transcription factor (HLTF) is another human RAD5 homolog that shares similar domains and functions with SHPRH in binding UBC13 and PCNA and facilitating PCNA polyubiquitinylation [103, 104]. Inactivation of SHPRH or HLTF elevates spontaneous mutagenesis and genome instability [103, 105]. Hence, SHPRH and HLTF cooperatively regulate PCNA polyubiquitinylation to activate the template-switching pathway to protect the replicating genome from DNA lesion-associated mutagenesis, genome instability, and subsequent carcinogenesis. The K164 mutation on PCNA impairs the Pol η- and Pol ζ-dependent TLS defense against UV lesions, and yet imparts a higher RAD51-mediated recombination activity [106]. These findings support that K164 ubiquitinylation is a critical posttranslational modification of PCNA that determines which of the two PPR pathways, i.e. TLS and template switching, will be chosen. Finally, how polyubiquitinylated PCNA turns on the template-switching pathway still remains speculative. One hypothesis is that PCNA polyubiquitinylation may induce template switching via a recombination-like mechanism. In mammalian cells, DSBs and other lesions associated with DNA replication are, for the most part, repaired by HR [107, 108]. It has been shown that RAD51 deficiency can also lead to the accumulation of DSBs at the sites of stalled replication forks, suggesting that RAD51-mediated HR may help resolve the stalling of replication forks [109]. The HR mechanism required for repairing two-ended DSBs has been extensively researched in the past. In contrast, the HR event occurring in response to replication stalling in mammalian cells is different from that seen in the two-ended DSB repair and is much less understood [65, 110, 111].
Better late than never: repairing damaged chromosomes
Base excision repair (BER)
Oxidized nucleotides are removed by the BER mechanism, which corrects oxidized and alkylated bases as well as SSBs. The BER pathway may work through a short-patch mechanism, which replaces single nucleotides, or a long-patch pathway, which replaces 2–13 nucleotides [112]. BER is initiated by one of several DNA glycosylases [e.g. OGG1 (8-oxoguanine glycosylase), UDG (uracil DNA glycosylase), and AAG (3-alkyladenine DNA glycosylase)] that recognizes and removes specific modified bases. For example, OGG1 is specialized in recognizing 8-oxoG, one of the most common lesions caused by oxidative damage. The resulting abasic or apyrimidinic/apurinic (AP) lesions are then excised by AP endonuclease (e.g. APE1) to create SSB. As an intermediate product of BER, SSB is recognized by poly(ADP-ribose) polymerase 1 (PARP1), which participates in the subsequent recruitment of Pol β [beta] and XRCC1-DNA ligase 3 for gap filling and closure, respectively. It has been shown that human ES cells in general show higher expression levels of BER genes, such as OGG1 and APE1, and that they also exhibit more efficient BER and lower 8-oxoG lesions compared to differentiated cells [113, 114]. Similarly, mouse neural stem and progenitor cells express higher levels of OGG1 and Neil1 than do differentiated neurons [115]. Some BER genes, such as XRCC1, DNA ligase 3, and DNA ligase 1, were found to be down-regulated during the differentiation of mouse myoblasts [116].
Mismatch repair (MMR), nucleotide excision repair (NER), and Fanconi anemia (FA)
The pathway that negotiates base mismatch, insertion loops, and deletion loops created during genome replication is MMR. In this pathway, mismatched bases are sensed by the MSH2–MSH3 or MSH2–MSH6 complex. The MSH complex then recruits MLH-1 and PMS2 to coordinate mismatch removal by exonuclease, gap filling by DNA polymerase, and gap closure by DNA ligase [117]. It has been shown that human pluripotent stem cells again express higher expression levels of MMR-related genes (e.g. MSH2, MSH5, MSH6, MLH-1, and PMS2) and display more efficient MMR repair compared to differentiated cells [113, 114, 118, 119]. UV radiation, environmental pollutants (e.g. aldehydes), and cross-linking reagents (e.g. platinum-related chemotherapeutic agents) can cause helix-distorting DNA lesions. This type of lesions requires the NER pathway for repair [120]. NER can be carried out by a global genome repair (GGR) mechanism, which senses and repairs damage occurring throughout the entire genome and depends on the functions of XPA and XPC, or by a transcription-coupled repair (TCR) mechanism, which repairs lesions on the transcribed strands of transcriptionally active genes and depends on the functions of XPA, Cockayne syndrome A (CSA), and Cockayne syndrome B (CSB) proteins. NER involves the sequential recruitment of a group of proteins that sense and prepare the DNA lesion [i.e. XPA, XPC-RAD23B, CSA, CSB, and transcription factor IIH (TFIIH, including XPB and XPD helicases)], remove damaged nucleotides (i.e. ERCC1-XPF, and XPG), synthesize DNAs (i.e. Pol δ, Pol ε, and accessory proteins such as PCNA and RPA), and close the gap (i.e. DNA ligase 3 and 1) [121–123]. The GGR activity is found to be attenuated upon differentiation of neural and macrophage precursors, whereas the TCR activity remains unchanged during the differentiation of these cells [124, 125]. Another mechanism specialized in the repair of intrastrand crosslinks is the FA pathway. The detail of this pathway can be found in several published reviews, in which readers may find more detail information [126–128]. For the interest of this review, it is worth noting that most of the components in the FA pathway are decreased during macrophage differentiation [129]. Interstrand crosslinks (ICLs), on the other hand, are caused by platinum-related chemotherapeutic agents and mitomycin C. They are repaired by a combination of pathways that include NER, HR, TLS, and FA, and, therefore, will have the same stem cell connotation as described previously [130].
DSB repair choices
Double-stranded breaks can be triggered by exogenous insults and by prolonged replication stalling. Prolonged stalling causes replication forks to collapse into DSBs. In addition, any unrepaired SSB, regardless of its origin, will be converted to a DSB at the replication fork during genome replication. DSBs can lead to chromosomal loss or rearrangement, and is the most lethal threat to all dividing cells. The response of cells to DSBs begins with the recruitment of ataxia telangiectasia mutated (ATM) that eventually turns on one of the four DSB repair programs: classical NHEJ (C-NHEJ), HR, single strand annealing (SSA), and alternative NHEJ (Alt-NHEJ) (Fig. 3). The HR and SSR repair mechanisms involve the pairing of extended homologous sequences and hence take place during the S and G2 phase. Alt-NHEJ may engage microhomology between DSB ends and take place during the S/G2 phase as well. In contrast, C-NHEJ is the predominant mechanism of repair in the G0 and G1 phase but can also operate in the S and G2 phase. According to this cell cycle-dependent preference, mitotically quiescent stem cells in adult tissues use primarily the error-prone C-NHEJ as their pathway of choice for DSB repair, whereas mitotically active stem cells (e.g. ES cells, iPS cells, embryonic tissue-specific stem cells, regenerating adult stem cells, and CSCs) may use all four mechanisms. This cell cycle phase-based selection of DSB repair pathways is consistent with the developmental transition from an HR-based repair in embryos to a C-NHEJ-based repair in adult animals [131–133]. In S-phase cells, HR and C-NHEJ appear to compete against each other for DSB sites, but the balance between them differs widely between different species as well as between different cell types of the same species. The key event that decides HR over C-NHEJ is the resection of DSB ends, which involves an initial limited resection step, mediated by the complex of C-terminal binding protein (CtBP) interacting protein (CtIP) and MRN (MRE11, RAD50, and NBS1), and a second extensive resection step, mediated by the EXO1-Bloom helicase (BLM) complex or the DNA2-BLM complex. Recent studies show that C-NHEJ is stimulated by 53BP1 and RIF1, and that HR and DNA end resection are promoted by breast cancer 1 (BRCA1) and CtIP [134–137]. Another mechanism by which haploid yeasts up-regulate C-NHEJ and down-regulate HR (or vice versa in diploid yeasts) is through a MAT-dependent regulation of Nej1. In some instances, cells may also use p53 to choose between HR and C-NHEJ. For example, when rapid cell division is required, p53 suppression may serve the role of suppressing HR as well as preventing cell cycle arrest in favor of a p53-independent apoptotic pathway to get rid of cells with damaged genome. Notwithstanding the pathway choice, the abilities to repair DNA damage by either HR or C-NHEJ are both critically important for the maintenance of the stem cell population as a whole. Their individual importance is further described as follows.
Four pathways for double-stranded break (DSB) repair. Nascent DSBs are first recognized by ATM and the MRN complex that initiate a series of DNA damage response events that aim at arresting cell cycle and recruiting DSB repair proteins. The C-NHEJ pathway is triggered by direct binding of Ku70/80 protein to DSB ends, followed by the recruitment of DNA-PKcs, Artemis, DNA ligase 4, XRCC4, and XLF. For cells in the S and G2 phase, DSBs can undergo limited end resection, followed by an extended end resection that leads to primarily HR repair but sometimes SSA repair. HR repair requires a core factor, RAD51, and several cofactors, including RAD51 paralogues, BRCA2, and nucleostemin (NS). Alternatively, the exposed 3′ ends of DSBs may be directly ligated by an Alt-NHEJ mechanism that is initiated by PARP1 and followed through by WRN, DNA ligase 3/XRCC1, and DNA ligase1. See text for more abbreviations
C-NHEJ
Unprocessed DSBs can be directly bound by Ku70/80 and repaired by the C-NHEJ mechanism, which involves an orderly recruitment of DNA-dependent protein kinase catalytic subunit (DNA-PKcs), Artemis, DNA ligase 4, XRCC4, and XLF. C-NHEJ can operate throughout the cell cycle and therefore is the major pathway for DSB repair in quiescent stem cells [138]. However, depending on the extent of end-processing and the fidelity of end-pairing, C-NHEJ-based repair may result in mutations or chromosomal rearrangement and therefore is considered error-prone in a relative sense. For this reason, while the quiescent state of stem cells help minimize their chance of incurring replicative and oxidative DNA damage, it also subjects them to the error-prone C-NHEJ repair mechanism instead of the error-free HR repair mechanism. Genes involved in the C-NHEJ repair are by and large increased in pluripotent stem cells, such as ES cells and iPS cells [113, 114, 139]. However, at least one of the C-NHEJ factors, DNA-PKcs, was shown to be down-regulated in ES cells compared to differentiated cells [133]. Unlike pluripotent stem cells, it was reported that bulge stem cells in the hair follicle exhibit a higher efficiency in C-NHEJ repair of DNA damage compared to epidermal cells as a result of their higher nuclear expression of DNA-PKcs [140]. Similarly, it was shown that thrombopoietin can promote C-NHEJ repair in hematopoietic stem cells [141]. Those differences may reflect the mitotically active and quiescent state of pluripotent stem cells versus bulge/hematopoietic stem cells, respectively. The efficient but error-prone NHEJ repair mechanism in bulge stem cells promotes their short-term survival at the cost of their long-term genomic stability. These findings highlight the notion that stem cells in general exhibit a higher efficiency in DNA damage repair, but the preferred choice of DSB repair pathways may vary among different stem cell types.
HR
Replication-induced DSBs most commonly evoke the HR machinery and the DNA helicases/nucleases for repair [142–145]. The ssDNA exposed by the initial 5′ limited end resection recruits ssDNA-binding protein, RPA, which then assembles ATR, Rad17-Rfc2-5, and the 9-1-1 complexes to trigger G2/M arrest [69–72]. Besides replicative damage, ATR can also be activated by DSBs via an ATM-mediated pathway in a cell cycle-dependent manner [146]. The activation of ATR turns on the HR repair mechanism by recruiting RAD51. RAD51 is homologous to the bacterial RecA, and is the core HR enzyme in eukaryotes that forms the presynaptic filament by binding ssDNAs in place of RPA [147–149]. The nucleoprotein complex of ssDNA-bound RAD51 is stabilized by RAD51 paralogues, which include five members in human, that is, XRCC2, XRCC3, RAD51B/RAD51L1, RAD51C/RAD51L2, and RAD51D/RAD51L3. RAD51 serves a key role in initiating strand invasion at the homologous sequence and driving the branch migration of the Holiday junction [150]. The mechanism for RAD51 recruitment following replication stalling is not entirely clear, but may involve breast cancer 2 (BRCA2) [151], SUMOylated BLM [152], SUMOylated RPA70 [153], RAD52 [154], and X-ray repair cross-complementing proteins 2 and 3 (XRCC2 and XRCC3) [155, 156]. In mice, germ-line deletion of RAD51 results in early embryonic lethality [157]. If the end-resection process uncovers direct repeat sequences, both ssDNA ends can be annealed and repaired by a process called SSA. SSA is a RAD51-independent repair mechanism regulated by RAD52, ERCC1, and XPF. SSA repair often leads to deletions of the sequence between repeats—a completely different outcome from HR-based repair in terms of the fidelity of repair. It is therefore an undesirable choice for DSB repair in stem cells.
It is noted that HR, although accurate in its repair, operates with a very slow kinetics, which poses a challenge for fast dividing cells with a large genome size. Therefore, in fast dividing higher eukaryotic cells that primarily use HR for genome maintenance, such as in the case of mouse ES cells, the HR activity needs to be boosted so that stalled/collapsed replication forks can be restarted/repaired in a timely manner and that their genome integrity can be maintained. In many other types of higher eukaryotic cells, HR appears to play a minor role in DSB repair, as C-NHEJ repair is more efficient than HR and is active throughout the cell cycle. The exact mechanisms controlling the preferential choice of HR over C-NHEJ in mitotically active stem cells have yet to be elucidated. Generally speaking, human ES cells show higher expression levels of HR repair genes, such as RAD51 and RAD54, compared to differentiated cells [113, 133, 158]. Another way to address this issue is to find new targets that are required for stem cell self-renewal and play a role in promoting HR repair. One such candidate is a stem and cancer cell-enriched protein with a well-established function in self-renewal maintenance—nucleostemin (NS) [159–163]. NS has been shown to play indispensable roles in several fundamental biological events, including early and late embryogenesis, adult tissue regeneration, and pluripotency reprogramming [164–170]. We recently discovered its key mechanism of action in protecting proliferating cells from DNA damage during the S-phase [168, 169, 171–174], which highlights the importance of genome maintenance in self-renewal and suggests NS as a new regulatory component in the repair of replicative DNA damage [174, 175]. The role of NS in genome protection was first discovered by its ability to reduce telomeric DNA damage [171]. It was shown that NS mechanistically promotes the SUMOylation of TRF1 and the telomeric recruitment of PML-IV through interaction with SUMOylated TRF1. More recently, a role of NS in protecting against replication-induced damage on non-telomeric chromosomes was uncovered in developing stem/progenitor cells and regenerating hepatocytes [168, 169]. Our data showed that NS-knockout (NSKO)-induced DNA damage occurs independently of the p53 status or rRNA synthesis, and that NS is directly recruited to DNA damage sites and regulates the recruitment of RAD51 to stalled replication-induced DNA damage foci. Early studies suggested a link between NS and the MDM2-p53 pathway [159, 176–178]. However, it is now clear that the obligatory effect of NS loss on cell proliferation and survival occurs in the absence of p53 [165, 179–181]. Our current model states that the ability of NS to protect the integrity of telomeric and non-telomeric chromosomes during genome replication is required for the maintenance of self-renewal. It operates constitutively by the nucleoplasmic pool of NS proteins. In contrast, the MDM2-regulatory function of NS is mostly silent under normal growth conditions and becomes activated only when the nucleolar organization is disassembled under nucleolar stress conditions (Fig. 4).
NS promotes HR repair of replication-triggered DNA damage in stem cells. Our current model states that the obligatory function of NS resides in its ability to maintain the integrity of replicating genome. NS does so by promoting HR repair of DNA damage in the S-phase via a RAD51-mediated mechanism and/or a TRF1-mediated mechanism. So far, there is no evidence to indicate that localization in the nucleolus (yellow circle) is required for the essential activity of NS. When the nucleolus is dissembled under nucleolar stress conditions, most of the nucleolar contents, including NS, are released into the nucleoplasm in bulk. The massive increase of NS in the nucleoplasm triggers its interaction with MDM2 and thereby suppresses the p53 activity. The green and red arrows indicate an excitatory/increase or an inhibitory/decrease effect, respectively
Alt-NHEJ
An alternative repair pathway, Alt-NHEJ, is defined as an end-joining event that does not require Ku proteins or DNA ligase 4. This alternative repair mechanism was first observed two decades ago in C-NHEJ-deficient yeasts and mammalian cells [182–185]. It has been questioned ever since whether Alt-NHEJ is simply a by-product of persistent reactive DSB ends that are repaired by surrogates when C-NHEJ and HR are unavailable [186] or stands as an evolutionarily conserved, bona fide end-joining repair pathway [187]. A recent study showed that Alt-NHEJ can occur at approximately 10 % of the C-NHEJ efficiency in C-NHEJ-proficient as well as C-NHEJ-deficient cells [188]. Another study reported the discovery of Alt-NHEJ in E. coli, which lacks C-NHEJ components [189]. These findings indicate that Alt-NHEJ is a mechanistically distinct pathway in its own right that might have preceded C-NHEJ in evolution [190]. Biologically, this pathway may help save genetic information at the cost of introducing mutagenic events when more accurate mechanisms of repair are not available during, for example, mitosis [191, 192]. Pathologically, it is recognized as the major pathway responsible for chromosomal translocation [193–195]. Our current knowledge describes that some Alt-NHEJ may operate by annealing microhomology unmasked by limited end resection. Based on this reason, it is sometimes referred to as microhomology-mediated end-joining (MMEJ), although microhomology may not always be present at the repaired junction [196, 197]. Alt-NHEJ is active during the S and G2 phase, and has the propensity to introduce deletions and chromosomal rearrangement [192, 196]. To date, the molecular mechanism underlying Alt-NHEJ is not entirely clear but appears to require enzymes that perform limited end resection, end recognition, microhomology pairing, flap removal, gap filling, and ligation [198, 199]. The initial 5′-end resection step of Alt-NHEJ is carried out by MRE11 in yeasts [197, 200] and mammals [201–203], and is also mediated by CtIP [204]. The protein involved in sensing DSB ends for Alt-NHEJ is believed to be PARP1. PARP1 has been shown to promote Alt-NHEJ by competing with Ku proteins for free DSB ends [205–209]. In arabidopsis, PARP mutants display less Alt-NHEJ products [210]. Removal of non-homologous flaps may be performed by the RAD1/RAD10 endonuclease in yeasts [197, 211] or the ERCC1-XPF (ERCC4) complex in mammals [212]. Finally, gap filling and ligation may require Werner syndrome ATP-dependent helicase (WRN), DNA polymerase λ [Pol λ (lamda)], DNA ligase 3/XRCC1, and DNA ligase1 [205, 207, 213–219]. Other than these promoting factors, Alt-NHEJ may be suppressed by PARP2 and proteins involved in driving the repair decision toward C-NHEJ and possibly HR [207]. As Alt-NHEJ often leads to variable-sized interstitial deletions, inversion, chromosomal translocation, and telomere fusion [220, 221], its activity needs to be tightly controlled in stem cells.
Repairing without actual repairing: the immortal strand hypothesis
An appealing but still controversial theory of chromosome cosegregation was proposed nearly four decades ago (Fig. 5). This theory explains how long-term dividing cells (i.e. adult stem cells) at steady state minimize the consequence of replication errors or replicative damage by selective cosegregation of the parental templates or chromosomes (the immortal DNA strands) from the newly synthesized daughter chromosomes (the mortal DNA strands) [222]. This phenomenon has been described in somatic stem cells undergoing asymmetric cell division in the epidermis [223], small intestinal crypts [224], mouse mammary epithelium [225], and muscle satellite cells [226, 227]. It has also been shown in cultured cells engineered with an inducible p53 [228], as well as in neural stem cells [229]. A modified CO-FISH method was recently developed to differentially label sister chromatids with unidirectional probes to telomeric satellite DNAs [230]. Using this CO-FISH method, it was observed that apparent non-random segregation of sister chromatids occurs in a subset of colon crypt epithelial cells, which supports asymmetry of template DNA strand segregation [230]. By retaining the original DNA templates and passing the newly synthesized DNA strands down to their differentiated progeny, those stem cells are guaranteed to dodge the high frequency of replicative DNA damage literally in a repair-free manner. Some studies have begun to address the potential mechanisms underlying the selective cosegregation of parental chromosomes. In adult skeletal muscle, stem cells with long-term self-renewal express more Pax7 than cells undergoing myogenic commitment. It has been shown by the CO-FISH analysis that the Pax7-high subpopulation displays a high frequency of template DNA strand cosegregation, whereas Pax-low subpopulation separates their chromatids randomly. Some satellite cells display non-random segregation of template DNA strands and the Numb protein during growth in muscle fibers in vivo as well as in culture [226]. Cardiac progenitors also exhibit asymmetrical chromatid segregation, where Pim-1, which is a kinase that enhances cardiac repair, plays a role by increasing the asymmetrical chromatid segregation and promoting self-renewal of cardiac progenitors [231]. Conversely, there are studies that refute the immortal DNA strand theory. For example, one study showed that sister chromatids display random distribution between daughter cells in cultured lung fibroblasts and ES cells [230]. Another recent study reported that the accumulation rate of mutations in healthy stem cells of the colon, blood, head, and neck tissues are strikingly similar to those expected without the protection from the immortal strand mechanism [232]. Using the chromosome labeling approach, one group demonstrated that crypt base columnar stem cells in adult intestinal crypts segregate most of their chromosomes randomly both in intact and in regenerating epithelium [233]. Taken together, the published data seem to suggest that adult skeletal muscles and epithelial cells may reduce the long-term impact of replication-associated mutagenesis by retaining the original DNA strands in the quiescent stem cell population with self-renewal capacity. The ability of template DNA strands to cosegregate allows long-term stem cells to avoid transmitting erroneous genetic information to inherited daughter stem cells. This skill, however fascinating, may not be commandeered by all stem cells and may rely heavily on the cell division pattern (asymmetric versus symmetric cell division). In addition, it may be confounded by the methods used to detect the existence of immortal DNA strands. Finally, the immortal strand hypothesis cannot fix the problem of replication-unrelated damages that can be directly inflicted upon both chromosomal strands.
The immortal strand hypothesis. Adult stem cells (yellow) that undergo asymmetric cell division may display a phenomenon, where the parental template DNAs (the immortal DNA strands, marked by green) are cosegregated into one daughter cell (i.e. stem cells) and the newly synthesized DNAs (the mortal DNA strands, marked by red) are passed down to the other daughter cells (i.e. progeny). For those that divide symmetrically, the template (green) strands are randomly segregated so that each daughter cells, no matter whether they are stem cells or differentiated cells, have equal chances to inherit the errors created during genome replication. The immortal strand theory explains why some adult stem cells accumulate less chromosomal mutations than expected, despite their long lifespan and self-renewal activity. Mutations accumulated during first to third round of genome replication are symbolized by asterisks, blue diamonds, and open circles, respectively
The lesser of two evils: choosing between survival with defects and death
In response to DNA damage, stem cells, like all other types of cells, recruit an evolutionarily conserved pathway, known as DNA damage response or DDR, which induces cell cycle arrest and activates DNA damage repair mechanisms [234]. The ultimate goal of the DDR pathway is to restore the damaged DNA and maintain cell survival. Under the condition when a complete reversal of genomic damage cannot be achieved and the resulting damage cannot be tolerated, those that harbor the damaged chromosomes will be eliminated by apoptosis, become senescent, undergo differentiation, or resume cycling at the risk of oncogenic transformation or mitotic catastrophe. Even though immortal DNA theory appears to be an ideal solution for tissues to eliminate damaged chromosomes without the cost of depleting the original stem cell pool, still it remains a theory and may not be applicable to all conditions for reasons stated above. Therefore, some consequences would have to be taken under most situations. It remains an intriguing question how stem cells weigh among these various outcomes when faced with improperly repaired chromosomal damage.
Senescence is when cells stay in the G0 phase indefinitely. Senescence, along with the cell death event, ensures that the damaged chromosomes will not be passed down to other stem cells or their progenies. p53 is still the key regulator of senescence and cell death in stem cells. While disarming of the p53 guardian mechanism does allow more stem cells to survive with a defective genome, it also exposes them to higher risks of tumorigenicity [235]. Contrarily, excessive use of the p53 mechanism will eventually result in the depletion of stem cells, which may then lead to impaired tissue homeostasis, organ failure, and premature aging. This idea has been supported by studies showing that, on one hand, p53 hyperactivation is associated with bone marrow failure [236, 237], and, on the other hand, p53 deficiency promotes blood production and leukemia development at the same time [238, 239]. It has been reported that, in mice, adult hematopoietic stem cells and bulge stem cells in hair follicles are more resistant to IR exposure compared to other blood cells and epidermal cells, respectively [138, 240–242]. The relative resistance to IR in those stem cells may be explained by their higher Bcl2 expression level and shorter duration of p53 activation [138, 140]. In contrast, intestinal stem cells undergo massive cell death in response to DNA damage, which may be caused by their lower Bcl-2 expression and longer duration of p53 activation [243, 244]. Similar to intestinal stem cells, fetal hematopoietic stem cells are also more sensitive to IR exposure than their progeny cells [245]. The difference in response to DNA damage between fetal and adult hematopoietic stem cells may reflect their respective developmental natures as well as their distinctive proliferative properties (i.e. asymmetric versus symmetric cell division). Autophagy is another mechanism that promotes the survival of stem cells under genotoxic and metabolic stress conditions [246]. Autophagy is a highly conserved pathway that removes and recycles damaged organelles sequestered in autophagosomes. As we begin to understand more about the biological role of autophagy and its molecular regulation and participating molecules, now is the time to examine how this event plays into the self-renewal of various stem cells. To begin with, autophagy may contribute to the low mitochondrial mass seen in some stem cells, given its close connection to the regulation of mitochondrial activity. Readers interested in the relationship between autophagy and stem cell self-renewal and differentiation may find more information in a review article recently published [247]. Finally, differentiation is another outcome of stem cells in stress. It has been shown that oxidative stress can induce the differentiation of hematopoietic stem cells [248]. Melanocyte stem cells also undergo DNA damage-induced differentiation [249]. The differentiation of stem cells following DNA damage may be mediated by a STAT3-regulated mechanism that increases the expression of BATF [250]. Together, the current data indicate that a combination of pathways, including p53, Bcl-2 family genes, autophagy, and JAK-STAT, may cooperatively determine the survival versus apoptosis outcomes of stem cells.
Last but not least: repairing chromosomal damage at the end
Telomeres are key protectors of chromosomal integrity but prone to damage during the DNA replication process. On one hand, DNA replication shortens the telomere length. On the other hand, it may introduce breaks on the double-stranded telomere repeat region due to replication fork stalling. Therefore, maintaining telomere integrity has been a major task for all dividing cells, particularly those undergoing self-renewing proliferation. Indeed, telomere dysfunction is linked to several aging disorders and cancers [251–255]. In most cells, the telomere length is maintained primarily by the telomerase complex [256, 257]. Therefore, it should come as no surprise that male germ line and most stem cells show high levels of telomerase activity. While the discovery of the telomerase complex nicely resolves the end-replication problem, it may not represent the whole picture of telomere biology. It was noted that hematopoietic stem cells from mice overexpressing telomerase reverse transcriptase (TERT) can be serially transplanted only to the same amount of times as those isolated from wild-type mice [258]. This result indicates that the telomere length is not the sole determinant of the longevity of hematopoietic stem cells. Other factors, such as the integrity of telomeric and non-telomeric chromosomes, come into play as well. In 10–15 % of human cancers, the telomerase activity is undetectable, and the telomere length is maintained by the alternative lengthening of telomere (ALT) mechanism. Those cells, also known as ALT cells, are characterized by the hallmarks of telomere sister chromatid exchange (T-SCE), extrachromosomal telomere repeats (ECTR), heterogeneous telomere length, and ALT-associated PML bodies (APB) [259, 260]. APB is a cell biological entity defined by the overlapping of telomere foci and PML bodies. It is believed that ALT cells may use the HR mechanism for telomere elongation. While the appearance of T-SCE and ECTR may be the result of telomere HR, the biological significance of APB remains to be found. Some have postulated that APB may serve the function of sequestering low molecular weight telomeric DNAs [261]. Others suggest that APB may be actively involved in the HR event [259, 262–265]. Interestingly, it was reported that telomere lengthening is carried out by a recombination-based, ALT-like mechanism during the early cleavage cycles after fertilization, which later transitions into the telomerase-based mechanism [266]. Reciprocally, whether the ALT state can be established from TA cells and how it is done if so happens remains unclear in a general sense. For a few selective TA cell types, e.g. HCT15 cells and T cell lymphoma, ALT can be induced by TERT inhibition or telomerase RNA component (TERC) deletion, respectively [267, 268]. One study showed that the 5′ cytosine-rich overhangs at the telomere may be linked to the HR program [269]. Other studies have identified a strong correlation between the ALT state and alpha thalassemia/mental retardation syndrome X-linked (ATRX, also known as RAD54) gene mutation in pancreatic neuroendocrine tumors, pediatric glioblastoma, and TA-ALT hybrid cells [270–272], which suggests that ATRX might be an ALT repressor.
Conclusion
Stem cells are at a higher risk of incurring DNA damage than their differentiated progeny because of their longevity in life and self-renewal requirement. The amount of damage accumulated on the genome becomes a major limiting factor that restricts their proliferative lifespan. As outlined in this review, there is good evidence to support that stem cells take care of this problem by obliterating the occurrence of DNA damage at first sight, accelerating the repair process, and selecting the less detrimental outcome (Fig. 1). Failure to do so may result in grave consequences, including organ failure, premature aging, and/or cancer formation. Such is the case with Cockayne syndrome, Werner syndrome, ataxia telangiectasia, xeroderma pigmentosum, trichothiodystrophy, and Hutchinson-Gilford progeria. While stem cells in general may be equipped with a heightened defense against genomic stress, they do not always do it in the same way. Mitotically active stem cells tend to use HR for damage repair and select apoptosis as the outcome in exchange for long-term genomic stability, whereas quiescent stem cells tend to do the opposite. It is my hope that a review on this subject may have a measurable impact on our understanding of the biology underlying tissue homeostasis, premature aging, and tumor formation through stating the current state of knowledge on how different types of stem cells maintain their genome integrity while undergoing self-renewal throughout life and building a conceptual framework to catalyze future research.
Abbreviations
- ABC:
-
ATP-binding cassette
- ALT:
-
Alternative lengthening of telomere
- Alt-NHEJ:
-
Alternative non-homologous end-joining
- APB:
-
ALT-associated PML bodies
- ATM:
-
Ataxia telangiectasia mutated
- ATR:
-
Ataxia telangiectasia and Rad3 related
- ATRX:
-
Alpha thalassemia/mental retardation syndrome X-linked (also known as RAD54)
- BLM:
-
Bloom helicase
- BRCA1/2:
-
Breast cancer 1/2
- CO-FISH:
-
Chromosome orientation fluorescence in situ hybridization
- C-NHEJ:
-
Classical non-homologous end-joining
- CtBP:
-
C-terminal binding protein
- CtIP:
-
CtBP interacting protein
- DDR:
-
DNA damage response
- DNA-PKcs:
-
DNA-dependent protein kinase catalytic subunit
- DSBs:
-
Double-stranded breaks
- ECTR:
-
Extrachromosomal telomere repeats
- ERCC1:
-
Excision repair cross-complementation group 1
- ERCC4:
-
Excision repair cross-complementation group 4 (also known as XPF)
- ES:
-
Embryonic stem
- FANCD1:
-
Fanconi anemia complementation group D1
- HLTF:
-
Helicase-like transcription factor
- HR:
-
Homologous recombination
- ICL:
-
Interstrand crosslink
- iPS:
-
Induced pluripotent stem
- IR:
-
Ionizing radiation
- KD:
-
Knock-down
- KO:
-
Knock-out
- MDR:
-
Multidrug resistance
- MRN:
-
MRE11/RAD50/NBS1
- MSH2:
-
mutS homolog 2
- NS:
-
Nucleostemin
- PARP1/2:
-
Poly(ADP)ribose polymerase 1 or 2
- PCNA:
-
Proliferating cell nuclear antigen
- PML:
-
Promyelocytic leukemia protein
- RFC:
-
Replication factor C
- RPA:
-
Replication protein A
- ROS:
-
Reactive oxygen species
- SOD:
-
Superoxide dismutase
- SHPRH:
-
SNF2 histone-linker PHD ring helicase
- SSA:
-
Single strand annealing
- SSBs:
-
Single-stranded breaks
- ssDNA:
-
Single-stranded DNA
- TERC:
-
Telomerase RNA component
- TERT:
-
Telomerase reverse transcriptase
- TLS:
-
Translesion synthesis
- TopBP1:
-
Topoisomerase II binding protein 1
- TRF1:
-
Telomeric repeat factor 1
- T-SCE:
-
Telomere sister chromatid exchange
- XLF:
-
XRCC4-like factor (also known as Cernunnos)
- XRCC1:
-
X-ray repair cross-complementing group 1
- WRN:
-
Werner syndrome ATP-dependent helicase
References
Fuchs E, Segre JA (2000) Stem cells: a new lease on life. Cell 100(1):143–155
Reynolds BA, Weiss S (1992) Generation of neurons and astrocytes from isolated cells of the adult mammalian central nervous system. Science 255(5052):1707–1710
Davis AA, Temple S (1994) A self-renewing multipotential stem cell in embryonic rat cerebral cortex. Nature 372(6503):263–266
Tsai RY (2004) A molecular view of stem cell and cancer cell self-renewal. Int J Biochem Cell Biol 36(4):684–694
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674
Reya T, Morrison SJ, Clarke MF, Weissman IL (2001) Stem cells, cancer, and cancer stem cells. Nature 414(6859):105–111
Wang JC, Dick JE (2005) Cancer stem cells: lessons from leukemia. Trends Cell Biol 15(9):494–501
Lapidot T, Sirard C, Vormoor J, Murdoch B, Hoang T, Caceres-Cortes J, Minden M, Paterson B, Caligiuri MA, Dick JE (1994) A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature 367(6464):645–648
Bonnet D, Dick JE (1997) Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nat Med 3(7):730–737
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF (2003) Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci USA 100(7):3983–3988
Singh SK, Clarke ID, Terasaki M, Bonn VE, Hawkins C, Squire J, Dirks PB (2003) Identification of a cancer stem cell in human brain tumors. Cancer Res 63(18):5821–5828
Collins AT, Berry PA, Hyde C, Stower MJ, Maitland NJ (2005) Prospective identification of tumorigenic prostate cancer stem cells. Cancer Res 65(23):10946–10951
Kim CF, Jackson EL, Woolfenden AE, Lawrence S, Babar I, Vogel S, Crowley D, Bronson RT, Jacks T (2005) Identification of bronchioalveolar stem cells in normal lung and lung cancer. Cell 121(6):823–835
O’Brien CA, Pollett A, Gallinger S, Dick JE (2007) A human colon cancer cell capable of initiating tumour growth in immunodeficient mice. Nature 445(7123):106–110
Hermann PC, Huber SL, Herrler T, Aicher A, Ellwart JW, Guba M, Bruns CJ, Heeschen C (2007) Distinct populations of cancer stem cells determine tumor growth and metastatic activity in human pancreatic cancer. Cell Stem Cell 1(3):313–323
Schatton T, Murphy GF, Frank NY, Yamaura K, Waaga-Gasser AM, Gasser M, Zhan Q, Jordan S, Duncan LM, Weishaupt C, Fuhlbrigge RC, Kupper TS, Sayegh MH, Frank MH (2008) Identification of cells initiating human melanomas. Nature 451(7176):345–349
Mandal PK, Blanpain C, Rossi DJ (2011) DNA damage response in adult stem cells: pathways and consequences. Nat Rev Mol Cell Biol 12(3):198–202
Naka K, Hirao A (2011) Maintenance of genomic integrity in hematopoietic stem cells. Int J Hematol 93(4):434–439
Nagaria P, Robert C, Rassool FV (2013) DNA double-strand break response in stem cells: mechanisms to maintain genomic integrity. Biochim Biophys Acta 1830(2):2345–2353
Rocha CR, Lerner LK, Okamoto OK, Marchetto MC, Menck CF (2013) The role of DNA repair in the pluripotency and differentiation of human stem cells. Mutat Res 752(1):25–35
Cervantes RB, Stringer JR, Shao C, Tischfield JA, Stambrook PJ (2002) Embryonic stem cells and somatic cells differ in mutation frequency and type. Proc Natl Acad Sci USA 99(6):3586–3590
Tilgner K, Neganova I, Moreno-Gimeno I, Al-Aama JY, Burks D, Yung S, Singhapol C, Saretzki G, Evans J, Gorbunova V, Gennery A, Przyborski S, Stojkovic M, Armstrong L, Jeggo P, Lako M (2013) A human iPSC model of Ligase IV deficiency reveals an important role for NHEJ-mediated-DSB repair in the survival and genomic stability of induced pluripotent stem cells and emerging haematopoietic progenitors. Cell Death Differ 20(8):1089–1100
Araki R, Fujimori A, Hamatani K, Mita K, Saito T, Mori M, Fukumura R, Morimyo M, Muto M, Itoh M, Tatsumi K, Abe M (1997) Nonsense mutation at Tyr-4046 in the DNA-dependent protein kinase catalytic subunit of severe combined immune deficiency mice. Proc Natl Acad Sci USA 94(6):2438–2443
Reese JS, Liu L, Gerson SL (2003) Repopulating defect of mismatch repair-deficient hematopoietic stem cells. Blood 102(5):1626–1633
Ito K, Hirao A, Arai F, Matsuoka S, Takubo K, Hamaguchi I, Nomiyama K, Hosokawa K, Sakurada K, Nakagata N, Ikeda Y, Mak TW, Suda T (2004) Regulation of oxidative stress by ATM is required for self-renewal of haematopoietic stem cells. Nature 431(7011):997–1002
Prasher JM, Lalai AS, Heijmans-Antonissen C, Ploemacher RE, Hoeijmakers JH, Touw IP, Niedernhofer LJ (2005) Reduced hematopoietic reserves in DNA interstrand crosslink repair-deficient Ercc1−/− mice. EMBO J 24(4):861–871
Navarro S, Meza NW, Quintana-Bustamante O, Casado JA, Jacome A, McAllister K, Puerto S, Surralles J, Segovia JC, Bueren JA (2006) Hematopoietic dysfunction in a mouse model for Fanconi anemia group D1. Mol Ther 14(4):525–535
Nijnik A, Woodbine L, Marchetti C, Dawson S, Lambe T, Liu C, Rodrigues NP, Crockford TL, Cabuy E, Vindigni A, Enver T, Bell JI, Slijepcevic P, Goodnow CC, Jeggo PA, Cornall RJ (2007) DNA repair is limiting for haematopoietic stem cells during ageing. Nature 447(7145):686–690
Rossi DJ, Bryder D, Seita J, Nussenzweig A, Hoeijmakers J, Weissman IL (2007) Deficiencies in DNA damage repair limit the function of haematopoietic stem cells with age. Nature 447(7145):725–729
Takubo K, Ohmura M, Azuma M, Nagamatsu G, Yamada W, Arai F, Hirao A, Suda T (2008) Stem cell defects in ATM-deficient undifferentiated spermatogonia through DNA damage-induced cell-cycle arrest. Cell Stem Cell 2(2):170–182
Zhang QS, Marquez-Loza L, Eaton L, Duncan AW, Goldman DC, Anur P, Watanabe-Smith K, Rathbun RK, Fleming WH, Bagby GC, Grompe M (2010) Fancd2−/− mice have hematopoietic defects that can be partially corrected by resveratrol. Blood 116(24):5140–5148
Zhang S, Yajima H, Huynh H, Zheng J, Callen E, Chen HT, Wong N, Bunting S, Lin YF, Li M, Lee KJ, Story M, Gapud E, Sleckman BP, Nussenzweig A, Zhang CC, Chen DJ, Chen BP (2011) Congenital bone marrow failure in DNA-PKcs mutant mice associated with deficiencies in DNA repair. J Cell Biol 193(2):295–305
Hoeijmakers JH (2009) DNA damage, aging, and cancer. N Engl J Med 361(15):1475–1485
Lodish H, Berk A, Matsudaira P, Kaiser CA, Krieger M, Scott MP, Zipursky SL, Darnell J (2004) Molecular biology of the cell, 5th edn. Freeman, New York, p 963
Challen GA, Little MH (2006) A side order of stem cells: the SP phenotype. Stem Cells 24(1):3–12
Tercero JA, Diffley JF (2001) Regulation of DNA replication fork progression through damaged DNA by the Mec1/Rad53 checkpoint. Nature 412(6846):553–557
Katou Y, Kanoh Y, Bando M, Noguchi H, Tanaka H, Ashikari T, Sugimoto K, Shirahige K (2003) S-phase checkpoint proteins Tof1 and Mrc1 form a stable replication-pausing complex. Nature 424(6952):1078–1083
Pacek M, Tutter AV, Kubota Y, Takisawa H, Walter JC (2006) Localization of MCM2-7, Cdc45, and GINS to the site of DNA unwinding during eukaryotic DNA replication. Mol Cell 21(4):581–587
Bakker ST, Passegue E (2013) Resilient and resourceful: genome maintenance strategies in hematopoietic stem cells. Exp Hematol 41(11):915–923
Savatier P, Lapillonne H, Jirmanova L, Vitelli L, Samarut J (2002) Analysis of the cell cycle in mouse embryonic stem cells. Methods Mol Biol 185:27–33
Kapinas K, Grandy R, Ghule P, Medina R, Becker K, Pardee A, Zaidi SK, Lian J, Stein J, van Wijnen A, Stein G (2013) The abbreviated pluripotent cell cycle. J Cell Physiol 228(1):9–20
Bernstein C, Prasad AR, Nifonsam V, Bernstein H (2013) DNA damage, DNA repair and cancer. New Research Directions in DNA Repair, pp 1114–1116
Ito K, Hirao A, Arai F, Takubo K, Matsuoka S, Miyamoto K, Ohmura M, Naka K, Hosokawa K, Ikeda Y, Suda T (2006) Reactive oxygen species act through p38 MAPK to limit the lifespan of hematopoietic stem cells. Nat Med 12(4):446–451
Jang YY, Sharkis SJ (2007) A low level of reactive oxygen species selects for primitive hematopoietic stem cells that may reside in the low-oxygenic niche. Blood 110(8):3056–3063
Yahata T, Takanashi T, Muguruma Y, Ibrahim AA, Matsuzawa H, Uno T, Sheng Y, Onizuka M, Ito M, Kato S, Ando K (2011) Accumulation of oxidative DNA damage restricts the self-renewal capacity of human hematopoietic stem cells. Blood 118(11):2941–2950
Greer EL, Brunet A (2005) FOXO transcription factors at the interface between longevity and tumor suppression. Oncogene 24(50):7410–7425
Tothova Z, Kollipara R, Huntly BJ, Lee BH, Castrillon DH, Cullen DE, McDowell EP, Lazo-Kallanian S, Williams IR, Sears C, Armstrong SA, Passegue E, DePinho RA, Gilliland DG (2007) FoxOs are critical mediators of hematopoietic stem cell resistance to physiologic oxidative stress. Cell 128(2):325–339
Miyamoto K, Araki KY, Naka K, Arai F, Takubo K, Yamazaki S, Matsuoka S, Miyamoto T, Ito K, Ohmura M, Chen C, Hosokawa K, Nakauchi H, Nakayama K, Nakayama KI, Harada M, Motoyama N, Suda T, Hirao A (2007) Foxo3a is essential for maintenance of the hematopoietic stem cell pool. Cell Stem Cell 1(1):101–112
Yalcin S, Zhang X, Luciano JP, Mungamuri SK, Marinkovic D, Vercherat C, Sarkar A, Grisotto M, Taneja R, Ghaffari S (2008) Foxo3 is essential for the regulation of ataxia telangiectasia mutated and oxidative stress-mediated homeostasis of hematopoietic stem cells. J Biol Chem 283(37):25692–25705
Naka K, Hoshii T, Muraguchi T, Tadokoro Y, Ooshio T, Kondo Y, Nakao S, Motoyama N, Hirao A (2010) TGF-beta-FOXO signalling maintains leukaemia-initiating cells in chronic myeloid leukaemia. Nature 463(7281):676–680
Liu J, Cao L, Chen J, Song S, Lee IH, Quijano C, Liu H, Keyvanfar K, Chen H, Cao LY, Ahn BH, Kumar NG, Rovira II, Xu XL, van Lohuizen M, Motoyama N, Deng CX, Finkel T (2009) Bmi1 regulates mitochondrial function and the DNA damage response pathway. Nature 459(7245):387–392
Parmar K, Mauch P, Vergilio JA, Sackstein R, Down JD (2007) Distribution of hematopoietic stem cells in the bone marrow according to regional hypoxia. Proc Natl Acad Sci USA 104(13):5431–5436
Takubo K, Goda N, Yamada W, Iriuchishima H, Ikeda E, Kubota Y, Shima H, Johnson RS, Hirao A, Suematsu M, Suda T (2010) Regulation of the HIF-1alpha level is essential for hematopoietic stem cells. Cell Stem Cell 7(3):391–402
Yilmaz OH, Katajisto P, Lamming DW, Gultekin Y, Bauer-Rowe KE, Sengupta S, Birsoy K, Dursun A, Yilmaz VO, Selig M, Nielsen GP, Mino-Kenudson M, Zukerberg LR, Bhan AK, Deshpande V, Sabatini DM (2012) mTORC1 in the Paneth cell niche couples intestinal stem-cell function to calorie intake. Nature 486(7404):490–495
Diehn M, Cho RW, Lobo NA, Kalisky T, Dorie MJ, Kulp AN, Qian D, Lam JS, Ailles LE, Wong M, Joshua B, Kaplan MJ, Wapnir I, Dirbas FM, Somlo G, Garberoglio C, Paz B, Shen J, Lau SK, Quake SR, Brown JM, Weissman IL, Clarke MF (2009) Association of reactive oxygen species levels and radioresistance in cancer stem cells. Nature 458(7239):780–783
Studer L, Csete M, Lee SH, Kabbani N, Walikonis J, Wold B, McKay R (2000) Enhanced proliferation, survival, and dopaminergic differentiation of CNS precursors in lowered oxygen. J Neurosci 20(19):7377–7383
Chen HL, Pistollato F, Hoeppner DJ, Ni HT, McKay RD, Panchision DM (2007) Oxygen tension regulates survival and fate of mouse central nervous system precursors at multiple levels. Stem Cells 25(9):2291–2301
St John JC, Amaral A, Bowles E, Oliveira JF, Lloyd R, Freitas M, Gray HL, Navara CS, Oliveira G, Schatten GP, Spikings E, Ramalho-Santos J (2006) The analysis of mitochondria and mitochondrial DNA in human embryonic stem cells. Methods Mol Biol 331:347–374
Simsek T, Kocabas F, Zheng J, Deberardinis RJ, Mahmoud AI, Olson EN, Schneider JW, Zhang CC, Sadek HA (2010) The distinct metabolic profile of hematopoietic stem cells reflects their location in a hypoxic niche. Cell Stem Cell 7(3):380–390
Prigione A, Fauler B, Lurz R, Lehrach H, Adjaye J (2010) The senescence-related mitochondrial/oxidative stress pathway is repressed in human induced pluripotent stem cells. Stem Cells 28(4):721–733
Mandal S, Lindgren AG, Srivastava AS, Clark AT, Banerjee U (2011) Mitochondrial function controls proliferation and early differentiation potential of embryonic stem cells. Stem Cells 29(3):486–495
Le Belle JE, Orozco NM, Paucar AA, Saxe JP, Mottahedeh J, Pyle AD, Wu H, Kornblum HI (2011) Proliferative neural stem cells have high endogenous ROS levels that regulate self-renewal and neurogenesis in a PI3K/Akt-dependant manner. Cell Stem Cell 8(1):59–71
Lamb R, Bonuccelli G, Ozsvari B, Peiris-Pages M, Fiorillo M, Smith DL, Bevilacqua G, Mazzanti CM, McDonnell LA, Naccarato AG, Chiu M, Wynne L, Martinez-Outschoorn UE, Sotgia F, Lisanti MP (2015) Mitochondrial mass, a new metabolic biomarker for stem-like cancer cells: understanding WNT/FGF-driven anabolic signaling. Oncotarget 6(31):30453–30471
Farnie G, Sotgia F, Lisanti MP (2015) High mitochondrial mass identifies a sub-population of stem-like cancer cells that are chemo-resistant. Oncotarget 6(31):30472–30486
Saintigny Y, Delacote F, Vares G, Petitot F, Lambert S, Averbeck D, Lopez BS (2001) Characterization of homologous recombination induced by replication inhibition in mammalian cells. EMBO J 20(14):3861–3870
Hanada K, Budzowska M, Modesti M, Maas A, Wyman C, Essers J, Kanaar R (2006) The structure-specific endonuclease Mus81-Eme1 promotes conversion of interstrand DNA crosslinks into double-strands breaks. EMBO J 25(20):4921–4932
Rothstein R, Michel B, Gangloff S (2000) Replication fork pausing and recombination or “gimme a break”. Genes Dev 14(1):1–10
Hyrien O (2000) Mechanisms and consequences of replication fork arrest. Biochimie 82(1):5–17
Zou L, Cortez D, Elledge SJ (2002) Regulation of ATR substrate selection by Rad17-dependent loading of Rad9 complexes onto chromatin. Genes Dev 16(2):198–208
Zou L, Elledge SJ (2003) Sensing DNA damage through ATRIP recognition of RPA-ssDNA complexes. Science 300(5625):1542–1548
Zou L, Liu D, Elledge SJ (2003) Replication protein A-mediated recruitment and activation of Rad17 complexes. Proc Natl Acad Sci USA 100(24):13827–13832
Wang X, Zou L, Lu T, Bao S, Hurov KE, Hittelman WN, Elledge SJ, Li L (2006) Rad17 phosphorylation is required for claspin recruitment and Chk1 activation in response to replication stress. Mol Cell 23(3):331–341
Lee Y, Katyal S, Downing SM, Zhao J, Russell HR, McKinnon PJ (2012) Neurogenesis requires TopBP1 to prevent catastrophic replicative DNA damage in early progenitors. Nat Neurosci 15(6):819–826
Dronkert ML, Kanaar R (2001) Repair of DNA interstrand cross-links. Mutat Res 486(4):217–247
Gibbs PE, McGregor WG, Maher VM, Nisson P, Lawrence CW (1998) A human homolog of the Saccharomyces cerevisiae REV3 gene, which encodes the catalytic subunit of DNA polymerase zeta. Proc Natl Acad Sci USA 95(12):6876–6880
Gibbs PE, Wang XD, Li Z, McManus TP, McGregor WG, Lawrence CW, Maher VM (2000) The function of the human homolog of Saccharomyces cerevisiae REV1 is required for mutagenesis induced by UV light. Proc Natl Acad Sci USA 97(8):4186–4191
Yuan F, Zhang Y, Rajpal DK, Wu X, Guo D, Wang M, Taylor JS, Wang Z (2000) Specificity of DNA lesion bypass by the yeast DNA polymerase eta. J Biol Chem 275(11):8233–8239
Srivastava AK, Han C, Zhao R, Cui T, Dai Y, Mao C, Zhao W, Zhang X, Yu J, Wang QE (2015) Enhanced expression of DNA polymerase eta contributes to cisplatin resistance of ovarian cancer stem cells. Proc Natl Acad Sci USA 112(14):4411–4416
Wang SC, Nakajima Y, Yu YL, Xia W, Chen CT, Yang CC, McIntush EW, Li LY, Hawke DH, Kobayashi R, Hung MC (2006) Tyrosine phosphorylation controls PCNA function through protein stability. Nat Cell Biol 8(12):1359–1368
Bailly V, Lamb J, Sung P, Prakash S, Prakash L (1994) Specific complex formation between yeast RAD6 and RAD18 proteins: a potential mechanism for targeting RAD6 ubiquitin-conjugating activity to DNA damage sites. Genes Dev 8(7):811–820
Bailly V, Lauder S, Prakash S, Prakash L (1997) Yeast DNA repair proteins Rad6 and Rad18 form a heterodimer that has ubiquitin conjugating, DNA binding, and ATP hydrolytic activities. J Biol Chem 272(37):23360–23365
Bailly V, Prakash S, Prakash L (1997) Domains required for dimerization of yeast Rad6 ubiquitin-conjugating enzyme and Rad18 DNA binding protein. Mol Cell Biol 17(8):4536–4543
Hoege C, Pfander B, Moldovan GL, Pyrowolakis G, Jentsch S (2002) RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419(6903):135–141
Stelter P, Ulrich HD (2003) Control of spontaneous and damage-induced mutagenesis by SUMO and ubiquitin conjugation. Nature 425(6954):188–191
Watanabe K, Tateishi S, Kawasuji M, Tsurimoto T, Inoue H, Yamaizumi M (2004) Rad18 guides poleta to replication stalling sites through physical interaction and PCNA monoubiquitination. EMBO J 23(19):3886–3896
Solomon DA, Cardoso MC, Knudsen ES (2004) Dynamic targeting of the replication machinery to sites of DNA damage. J Cell Biol 166(4):455–463
Kannouche PL, Wing J, Lehmann AR (2004) Interaction of human DNA polymerase eta with monoubiquitinated PCNA: a possible mechanism for the polymerase switch in response to DNA damage. Mol Cell 14(4):491–500
Bienko M, Green CM, Crosetto N, Rudolf F, Zapart G, Coull B, Kannouche P, Wider G, Peter M, Lehmann AR, Hofmann K, Dikic I (2005) Ubiquitin-binding domains in Y-family polymerases regulate translesion synthesis. Science 310(5755):1821–1824
Ghosal G, Leung JW, Nair BC, Fong KW, Chen J (2012) Proliferating cell nuclear antigen (PCNA)-binding protein C1orf124 is a regulator of translesion synthesis. J Biol Chem 287(41):34225–34233
Niimi A, Chambers AL, Downs JA, Lehmann AR (2012) A role for chromatin remodellers in replication of damaged DNA. Nucleic Acids Res 40(15):7393–7403
Lorenz A, Osman F, Folkyte V, Sofueva S, Whitby MC (2009) Fbh1 limits Rad51-dependent recombination at blocked replication forks. Mol Cell Biol 29(17):4742–4756
Fugger K, Chu WK, Haahr P, Kousholt AN, Beck H, Payne MJ, Hanada K, Hickson ID, Sorensen CS (2013) FBH1 co-operates with MUS81 in inducing DNA double-strand breaks and cell death following replication stress. Nat Commun 4:1423
Jeong YT, Rossi M, Cermak L, Saraf A, Florens L, Washburn MP, Sung P, Schildkraut CL, Pagano M (2013) FBH1 promotes DNA double-strand breakage and apoptosis in response to DNA replication stress. J Cell Biol 200(2):141–149
Bacquin A, Pouvelle C, Siaud N, Perderiset M, Salome-Desnoulez S, Tellier-Lebegue C, Lopez B, Charbonnier JB, Kannouche PL (2013) The helicase FBH1 is tightly regulated by PCNA via CRL4(Cdt2)-mediated proteolysis in human cells. Nucleic Acids Res 41(13):6501–6513
Pfander B, Moldovan GL, Sacher M, Hoege C, Jentsch S (2005) SUMO-modified PCNA recruits Srs2 to prevent recombination during S phase. Nature 436(7049):428–433
Moldovan GL, Pfander B, Jentsch S (2006) PCNA controls establishment of sister chromatid cohesion during S phase. Mol Cell 23(5):723–732
Gali H, Juhasz S, Morocz M, Hajdu I, Fatyol K, Szukacsov V, Burkovics P, Haracska L (2012) Role of SUMO modification of human PCNA at stalled replication fork. Nucleic Acids Res 40(13):6049–6059
Hofmann RM, Pickart CM (1999) Noncanonical MMS2-encoded ubiquitin-conjugating enzyme functions in assembly of novel polyubiquitin chains for DNA repair. Cell 96(5):645–653
Ulrich HD (2003) Protein-protein interactions within an E2-RING finger complex. Implications for ubiquitin-dependent DNA damage repair. J Biol Chem 278(9):7051–7058
Motegi A, Sood R, Moinova H, Markowitz SD, Liu PP, Myung K (2006) Human SHPRH suppresses genomic instability through proliferating cell nuclear antigen polyubiquitination. J Cell Biol 175(5):703–708
Unk I, Hajdu I, Fatyol K, Szakal B, Blastyak A, Bermudez V, Hurwitz J, Prakash L, Prakash S, Haracska L (2006) Human SHPRH is a ubiquitin ligase for Mms2-Ubc13-dependent polyubiquitylation of proliferating cell nuclear antigen. Proc Natl Acad Sci USA 103(48):18107–18112
Sood R, Makalowska I, Galdzicki M, Hu P, Eddings E, Robbins CM, Moses T, Namkoong J, Chen S, Trent JM (2003) Cloning and characterization of a novel gene, SHPRH, encoding a conserved putative protein with SNF2/helicase and PHD-finger domains from the 6q24 region. Genomics 82(2):153–161
Motegi A, Liaw HJ, Lee KY, Roest HP, Maas A, Wu X, Moinova H, Markowitz SD, Ding H, Hoeijmakers JH, Myung K (2008) Polyubiquitination of proliferating cell nuclear antigen by HLTF and SHPRH prevents genomic instability from stalled replication forks. Proc Natl Acad Sci USA 105(34):12411–12416
Unk I, Hajdu I, Fatyol K, Hurwitz J, Yoon JH, Prakash L, Prakash S, Haracska L (2008) Human HLTF functions as a ubiquitin ligase for proliferating cell nuclear antigen polyubiquitination. Proc Natl Acad Sci USA 105(10):3768–3773
Motegi A, Kuntz K, Majeed A, Smith S, Myung K (2006) Regulation of gross chromosomal rearrangements by ubiquitin and SUMO ligases in Saccharomyces cerevisiae. Mol Cell Biol 26(4):1424–1433
Haracska L, Torres-Ramos CA, Johnson RE, Prakash S, Prakash L (2004) Opposing effects of ubiquitin conjugation and SUMO modification of PCNA on replicational bypass of DNA lesions in Saccharomyces cerevisiae. Mol Cell Biol 24(10):4267–4274
Johnson RD, Jasin M (2000) Sister chromatid gene conversion is a prominent double-strand break repair pathway in mammalian cells. EMBO J 19(13):3398–3407
Lundin C, Erixon K, Arnaudeau C, Schultz N, Jenssen D, Meuth M, Helleday T (2002) Different roles for nonhomologous end joining and homologous recombination following replication arrest in mammalian cells. Mol Cell Biol 22(16):5869–5878
Sonoda E, Sasaki MS, Buerstedde JM, Bezzubova O, Shinohara A, Ogawa H, Takata M, Yamaguchi-Iwai Y, Takeda S (1998) Rad51-deficient vertebrate cells accumulate chromosomal breaks prior to cell death. EMBO J 17(2):598–608
Bolderson E, Scorah J, Helleday T, Smythe C, Meuth M (2004) ATM is required for the cellular response to thymidine induced replication fork stress. Hum Mol Genet 13(23):2937–2945
Saleh-Gohari N, Bryant HE, Schultz N, Parker KM, Cassel TN, Helleday T (2005) Spontaneous homologous recombination is induced by collapsed replication forks that are caused by endogenous DNA single-strand breaks. Mol Cell Biol 25(16):7158–7169
Wilson DM 3rd, Bohr VA (2007) The mechanics of base excision repair, and its relationship to aging and disease. DNA Repair (Amst) 6(4):544–559
Maynard S, Swistowska AM, Lee JW, Liu Y, Liu ST, Da Cruz AB, Rao M, de Souza-Pinto NC, Zeng X, Bohr VA (2008) Human embryonic stem cells have enhanced repair of multiple forms of DNA damage. Stem Cells 26(9):2266–2274
Momcilovic O, Knobloch L, Fornsaglio J, Varum S, Easley C, Schatten G (2010) DNA damage responses in human induced pluripotent stem cells and embryonic stem cells. PLoS One 5(10):e13410
Hildrestrand GA, Diep DB, Kunke D, Bolstad N, Bjoras M, Krauss S, Luna L (2007) The capacity to remove 8-oxoG is enhanced in newborn neural stem/progenitor cells and decreases in juvenile mice and upon cell differentiation. DNA Repair (Amst) 6(6):723–732
Narciso L, Fortini P, Pajalunga D, Franchitto A, Liu P, Degan P, Frechet M, Demple B, Crescenzi M, Dogliotti E (2007) Terminally differentiated muscle cells are defective in base excision DNA repair and hypersensitive to oxygen injury. Proc Natl Acad Sci USA 104(43):17010–17015
Modrich P (2006) Mechanisms in eukaryotic mismatch repair. J Biol Chem 281(41):30305–30309
Casorelli I, Pelosi E, Biffoni M, Cerio AM, Peschle C, Testa U, Bignami M (2007) Methylation damage response in hematopoietic progenitor cells. DNA Repair (Amst) 6(8):1170–1178
Lin B, Gupta D, Heinen CD (2014) Human pluripotent stem cells have a novel mismatch repair-dependent damage response. J Biol Chem 289(35):24314–24324
Shuck SC, Short EA, Turchi JJ (2008) Eukaryotic nucleotide excision repair: from understanding mechanisms to influencing biology. Cell Res 18(1):64–72
Araujo SJ, Tirode F, Coin F, Pospiech H, Syvaoja JE, Stucki M, Hubscher U, Egly JM, Wood RD (2000) Nucleotide excision repair of DNA with recombinant human proteins: definition of the minimal set of factors, active forms of TFIIH, and modulation by CAK. Genes Dev 14(3):349–359
Volker M, Mone MJ, Karmakar P, van Hoffen A, Schul W, Vermeulen W, Hoeijmakers JH, van Driel R, van Zeeland AA, Mullenders LH (2001) Sequential assembly of the nucleotide excision repair factors in vivo. Mol Cell 8(1):213–224
Mocquet V, Laine JP, Riedl T, Yajin Z, Lee MY, Egly JM (2008) Sequential recruitment of the repair factors during NER: the role of XPG in initiating the resynthesis step. EMBO J 27(1):155–167
Nouspikel T, Hanawalt PC (2000) Terminally differentiated human neurons repair transcribed genes but display attenuated global DNA repair and modulation of repair gene expression. Mol Cell Biol 20(5):1562–1570
Nouspikel T, Hanawalt PC (2006) Impaired nucleotide excision repair upon macrophage differentiation is corrected by E1 ubiquitin-activating enzyme. Proc Natl Acad Sci USA 103(44):16188–16193
Naim V, Rosselli F (2009) The FANC pathway and mitosis: a replication legacy. Cell Cycle 8(18):2907–2911
Soulier J (2011) Fanconi anemia. Hematol Am Soc Hematol Educ Program 2011:492–497
Walden H, Deans AJ (2014) The Fanconi anemia DNA repair pathway: structural and functional insights into a complex disorder. Annu Rev Biophys 43:257–278
Lu WT, Lemonidis K, Drayton RM, Nouspikel T (2011) The Fanconi anemia pathway is downregulated upon macrophage differentiation through two distinct mechanisms. Cell Cycle 10(19):3300–3310
Huang Y, Li L (2013) DNA crosslinking damage and cancer—a tale of friend and foe. Transl Cancer Res 2(3):144–154
Essers J, Hendriks RW, Swagemakers SM, Troelstra C, de Wit J, Bootsma D, Hoeijmakers JH, Kanaar R (1997) Disruption of mouse RAD54 reduces ionizing radiation resistance and homologous recombination. Cell 89(2):195–204
Essers J, van Steeg H, de Wit J, Swagemakers SM, Vermeij M, Hoeijmakers JH, Kanaar R (2000) Homologous and non-homologous recombination differentially affect DNA damage repair in mice. EMBO J 19(7):1703–1710
Tichy ED, Pillai R, Deng L, Liang L, Tischfield J, Schwemberger SJ, Babcock GF, Stambrook PJ (2010) Mouse embryonic stem cells, but not somatic cells, predominantly use homologous recombination to repair double-strand DNA breaks. Stem Cells Dev 19(11):1699–1711
Chapman JR, Barral P, Vannier JB, Borel V, Steger M, Tomas-Loba A, Sartori AA, Adams IR, Batista FD, Boulton SJ (2013) RIF1 is essential for 53BP1-dependent nonhomologous end joining and suppression of DNA double-strand break resection. Mol Cell 49(5):858–871
Zimmermann M, Lottersberger F, Buonomo SB, Sfeir A, de Lange T (2013) 53BP1 regulates DSB repair using Rif1 to control 5′ end resection. Science 339(6120):700–704
Escribano-Diaz C, Orthwein A, Fradet-Turcotte A, Xing M, Young JT, Tkac J, Cook MA, Rosebrock AP, Munro M, Canny MD, Xu D, Durocher D (2013) A cell cycle-dependent regulatory circuit composed of 53BP1-RIF1 and BRCA1-CtIP controls DNA repair pathway choice. Mol Cell 49(5):872–883
Di Virgilio M, Callen E, Yamane A, Zhang W, Jankovic M, Gitlin AD, Feldhahn N, Resch W, Oliveira TY, Chait BT, Nussenzweig A, Casellas R, Robbiani DF, Nussenzweig MC (2013) Rif1 prevents resection of DNA breaks and promotes immunoglobulin class switching. Science 339(6120):711–715
Mohrin M, Bourke E, Alexander D, Warr MR, Barry-Holson K, Le Beau MM, Morrison CG, Passegue E (2010) Hematopoietic stem cell quiescence promotes error-prone DNA repair and mutagenesis. Cell Stem Cell 7(2):174–185
Fan J, Robert C, Jang YY, Liu H, Sharkis S, Baylin SB, Rassool FV (2011) Human induced pluripotent cells resemble embryonic stem cells demonstrating enhanced levels of DNA repair and efficacy of nonhomologous end-joining. Mutat Res 713(1–2):8–17
Sotiropoulou PA, Candi A, Mascre G, De Clercq S, Youssef KK, Lapouge G, Dahl E, Semeraro C, Denecker G, Marine JC, Blanpain C (2010) Bcl-2 and accelerated DNA repair mediates resistance of hair follicle bulge stem cells to DNA-damage-induced cell death. Nat Cell Biol 12(6):572–582
de Laval B, Pawlikowska P, Petit-Cocault L, Bilhou-Nabera C, Aubin-Houzelstein G, Souyri M, Pouzoulet F, Gaudry M, Porteu F (2013) Thrombopoietin-increased DNA-PK-dependent DNA repair limits hematopoietic stem and progenitor cell mutagenesis in response to DNA damage. Cell Stem Cell 12(1):37–48
Helleday T (2008) Amplifying tumour-specific replication lesions by DNA repair inhibitors—a new era in targeted cancer therapy. Eur J Cancer 44(7):921–927
Bachrati CZ, Hickson ID (2008) RecQ helicases: guardian angels of the DNA replication fork. Chromosoma 117(3):219–233
Errico A, Costanzo V (2010) Differences in the DNA replication of unicellular eukaryotes and metazoans: known unknowns. EMBO Rep 11(4):270–278
Petermann E, Helleday T (2010) Pathways of mammalian replication fork restart. Nat Rev Mol Cell Biol 11(10):683–687
Jazayeri A, Falck J, Lukas C, Bartek J, Smith GC, Lukas J, Jackson SP (2006) ATM- and cell cycle-dependent regulation of ATR in response to DNA double-strand breaks. Nat Cell Biol 8(1):37–45
Bianco PR, Tracy RB, Kowalczykowski SC (1998) DNA strand exchange proteins: a biochemical and physical comparison. Front Biosci 3:D570–D603
West SC (2003) Molecular views of recombination proteins and their control. Nat Rev Mol Cell Biol 4(6):435–445
Sung P, Krejci L, Van Komen S, Sehorn MG (2003) Rad51 recombinase and recombination mediators. J Biol Chem 278(44):42729–42732
Rossi MJ, Mazina OM, Bugreev DV, Mazin AV (2011) The RecA/RAD51 protein drives migration of Holliday junctions via polymerization on DNA. Proc Natl Acad Sci USA 108(16):6432–6437
Tarsounas M, Davies D, West SC (2003) BRCA2-dependent and independent formation of RAD51 nuclear foci. Oncogene 22(8):1115–1123
Ouyang KJ, Woo LL, Zhu J, Huo D, Matunis MJ, Ellis NA (2009) SUMO modification regulates BLM and RAD51 interaction at damaged replication forks. PLoS Biol 7(12):e1000252
Dou H, Huang C, Singh M, Carpenter PB, Yeh ET (2010) Regulation of DNA repair through deSUMOylation and SUMOylation of replication protein A complex. Mol Cell 39(3):333–345
Li X, Heyer WD (2008) Homologous recombination in DNA repair and DNA damage tolerance. Cell Res 18(1):99–113
Liu N, Lamerdin JE, Tebbs RS, Schild D, Tucker JD, Shen MR, Brookman KW, Siciliano MJ, Walter CA, Fan W, Narayana LS, Zhou ZQ, Adamson AW, Sorensen KJ, Chen DJ, Jones NJ, Thompson LH (1998) XRCC2 and XRCC3, new human Rad51-family members, promote chromosome stability and protect against DNA cross-links and other damages. Mol Cell 1(6):783–793
Sonoda E, Zhao GY, Kohzaki M, Dhar PK, Kikuchi K, Redon C, Pilch DR, Bonner WM, Nakano A, Watanabe M, Nakayama T, Takeda S, Takami Y (2007) Collaborative roles of gammaH2AX and the Rad51 paralog Xrcc3 in homologous recombinational repair. DNA Repair (Amst) 6(3):280–292
Tsuzuki T, Fujii Y, Sakumi K, Tominaga Y, Nakao K, Sekiguchi M, Matsushiro A, Yoshimura Y, Morita T (1996) Targeted disruption of the Rad51 gene leads to lethality in embryonic mice. Proc Natl Acad Sci USA 93(13):6236–6240
Adams BR, Golding SE, Rao RR, Valerie K (2010) Dynamic dependence on ATR and ATM for double-strand break repair in human embryonic stem cells and neural descendants. PLoS One 5(4):e10001
Tsai RY, McKay RD (2002) A nucleolar mechanism controlling cell proliferation in stem cells and cancer cells. Genes Dev 16(23):2991–3003
Baddoo M, Hill K, Wilkinson R, Gaupp D, Hughes C, Kopen GC, Phinney DG (2003) Characterization of mesenchymal stem cells isolated from murine bone marrow by negative selection. J Cell Biochem 89(6):1235–1249
Ohmura M, Naka K, Hoshii T, Muraguchi T, Shugo H, Tamase A, Uema N, Ooshio T, Arai F, Takubo K, Nagamatsu G, Hamaguchi I, Takagi M, Ishihara M, Sakurada K, Miyaji H, Suda T, Hirao A (2008) Identification of stem cells during prepubertal spermatogenesis via monitoring of nucleostemin promoter activity. Stem Cells 26(12):3237–3246
Tamase A, Muraguchi T, Naka K, Tanaka S, Kinoshita M, Hoshii T, Ohmura M, Shugo H, Ooshio T, Nakada M, Sawamoto K, Onodera M, Matsumoto K, Oshima M, Asano M, Saya H, Okano H, Suda T, Hamada J, Hirao A (2009) Identification of tumor-initiating cells in a highly aggressive brain tumor using promoter activity of nucleostemin. Proc Natl Acad Sci USA 106(40):17163–17168
Lin T, Meng L, Li Y, Tsai RY (2010) Tumor-initiating function of nucleostemin-enriched mammary tumor cells. Cancer Res 70(22):9444–9452
Zhu Q, Yasumoto H, Tsai RY (2006) Nucleostemin delays cellular senescence and negatively regulates TRF1 protein stability. Mol Cell Biol 26(24):9279–9290
Beekman C, Nichane M, De Clercq S, Maetens M, Floss T, Wurst W, Bellefroid E, Marine JC (2006) Evolutionarily conserved role of nucleostemin: controlling proliferation of stem/progenitor cells during early vertebrate development. Mol Cell Biol 26(24):9291–9301
Tsai RY (2011) New frontiers in nucleolar research: nucleostemin and related proteins. Nucleolus (Protein Reviews) 15:301–320
Qu J, Bishop JM (2012) Nucleostemin maintains self-renewal of embryonic stem cells and promotes reprogramming of somatic cells to pluripotency. J Cell Biol 197(6):731–745
Meng L, Lin T, Peng G, Hsu JK, Lee S, Lin S-Y, Tsai RY (2013) Nucleostemin deletion reveals an essential mechanism that maintains the genomic stability of stem and progenitor cells. Proc Natl Acad Sci USA 110(28):11415–11420
Lin T, Ibrahim W, Peng C-Y, Finegold MJ, Tsai RY (2013) A novel role of nucleostemin in maintaining the genome integrity of dividing hepatocytes during mouse liver development and regeneration. Hepatology 58(6):2176–2187
Tsai RY (2015) Pluripotency versus self-renewal of ES cells: two sides of the same coin or more? Stem Cells 33(7):2358–2359
Hsu JK, Lin T, Tsai RY (2012) Nucleostemin prevents telomere damage by promoting PML-IV recruitment to SUMOylated TRF1. J Cell Biol 197(5):613–624
Yamashita M, Nitta E, Nagamatsu G, Ikushima YM, Hosokawa K, Arai F, Suda T (2013) Nucleostemin is indispensable for the maintenance and genetic stability of hematopoietic stem cells. Biochem Biophys Res Commun 441(1):196–201
Lin T, Meng L, Wu LJ, Pederson T, Tsai RY (2014) Nucleostemin and GNL3L exercise distinct functions in genome protection and ribosome synthesis, respectively. J Cell Sci 127(10):2302–2312
Tsai RY (2014) Turning a new page on nucleostemin and self-renewal. J Cell Sci 127(18):3885–3891
Tsai RY, Meng L (2009) Nucleostemin: a latecomer with new tricks. Int J Biochem Cell Biol 41(11):2122–2124
Ma H, Pederson T (2007) Depletion of the nucleolar protein nucleostemin causes G1 cell cycle arrest via the p53 pathway. Mol Biol Cell 18(7):2630–2635
Meng L, Lin T, Tsai RY (2008) Nucleoplasmic mobilization of nucleostemin stabilizes MDM2 and promotes G2-M progression and cell survival. J Cell Sci 121(24):4037–4046
Dai MS, Sun XX, Lu H (2008) Aberrant expression of nucleostemin activates p53 and induces cell cycle arrest via inhibition of MDM2. Mol Cell Biol 28(13):4365–4376
Jafarnejad SM, Mowla SJ, Matin MM (2008) Knocking-down the expression of nucleostemin significantly decreases rate of proliferation of rat bone marrow stromal stem cells in an apparently p53-independent manner. Cell Prolif 41(1):28–35
Nikpour P, Mowla SJ, Jafarnejad SM, Fischer U, Schulz WA (2009) Differential effects of Nucleostemin suppression on cell cycle arrest and apoptosis in the bladder cancer cell lines 5637 and SW1710. Cell Prolif 42(6):762–769
Huang G, Meng L, Tsai RY (2015) p53 configures the G2/M arrest response of nucleostemin-deficient cells. Cell Death Discov 1:e15060
Boulton SJ, Jackson SP (1996) Identification of a Saccharomyces cerevisiae Ku80 homologue: roles in DNA double strand break rejoining and in telomeric maintenance. Nucleic Acids Res 24(23):4639–4648
Liang F, Jasin M (1996) Ku80-deficient cells exhibit excess degradation of extrachromosomal DNA. J Biol Chem 271(24):14405–14411
Wilson TE, Grawunder U, Lieber MR (1997) Yeast DNA ligase IV mediates non-homologous DNA end joining. Nature 388(6641):495–498
Kabotyanski EB, Gomelsky L, Han JO, Stamato TD, Roth DB (1998) Double-strand break repair in Ku86- and XRCC4-deficient cells. Nucleic Acids Res 26(23):5333–5342
Lieber MR (2010) The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem 79:181–211
Nussenzweig A, Nussenzweig MC (2007) A backup DNA repair pathway moves to the forefront. Cell 131(2):223–225
Corneo B, Wendland RL, Deriano L, Cui X, Klein IA, Wong SY, Arnal S, Holub AJ, Weller GR, Pancake BA, Shah S, Brandt VL, Meek K, Roth DB (2007) Rag mutations reveal robust alternative end joining. Nature 449(7161):483–486
Chayot R, Montagne B, Mazel D, Ricchetti M (2010) An end-joining repair mechanism in Escherichia coli. Proc Natl Acad Sci USA 107(5):2141–2146
Bennardo N, Cheng A, Huang N, Stark JM (2008) Alternative-NHEJ is a mechanistically distinct pathway of mammalian chromosome break repair. PLoS Genet 4(6):e1000110
Perrault R, Wang H, Wang M, Rosidi B, Iliakis G (2004) Backup pathways of NHEJ are suppressed by DNA-PK. J Cell Biochem 92(4):781–794
Peterson SE, Li Y, Chait BT, Gottesman ME, Baer R, Gautier J (2011) Cdk1 uncouples CtIP-dependent resection and Rad51 filament formation during M-phase double-strand break repair. J Cell Biol 194(5):705–720
Yan CT, Boboila C, Souza EK, Franco S, Hickernell TR, Murphy M, Gumaste S, Geyer M, Zarrin AA, Manis JP, Rajewsky K, Alt FW (2007) IgH class switching and translocations use a robust non-classical end-joining pathway. Nature 449(7161):478–482
Simsek D, Jasin M (2010) Alternative end-joining is suppressed by the canonical NHEJ component Xrcc4-ligase IV during chromosomal translocation formation. Nat Struct Mol Biol 17(4):410–416
Boboila C, Alt FW, Schwer B (2012) Classical and alternative end-joining pathways for repair of lymphocyte-specific and general DNA double-strand breaks. Adv Immunol 116:1–49
Boulton SJ, Jackson SP (1996) Saccharomyces cerevisiae Ku70 potentiates illegitimate DNA double-strand break repair and serves as a barrier to error-prone DNA repair pathways. EMBO J 15(18):5093–5103
Ma JL, Kim EM, Haber JE, Lee SE (2003) Yeast Mre11 and Rad1 proteins define a Ku-independent mechanism to repair double-strand breaks lacking overlapping end sequences. Mol Cell Biol 23(23):8820–8828
Deriano L, Roth DB (2013) Modernizing the nonhomologous end-joining repertoire: alternative and classical NHEJ share the stage. Annu Rev Genet 47:433–455
Frit P, Barboule N, Yuan Y, Gomez D, Calsou P (2014) Alternative end-joining pathway(s): bricolage at DNA breaks. DNA Repair (Amst) 17:81–97
Lee K, Lee SE (2007) Saccharomyces cerevisiae Sae2- and Tel1-dependent single-strand DNA formation at DNA break promotes microhomology-mediated end joining. Genetics 176(4):2003–2014
Dinkelmann M, Spehalski E, Stoneham T, Buis J, Wu Y, Sekiguchi JM, Ferguson DO (2009) Multiple functions of MRN in end-joining pathways during isotype class switching. Nat Struct Mol Biol 16(8):808–813
Rass E, Grabarz A, Plo I, Gautier J, Bertrand P, Lopez BS (2009) Role of Mre11 in chromosomal nonhomologous end joining in mammalian cells. Nat Struct Mol Biol 16(8):819–824
Xie A, Kwok A, Scully R (2009) Role of mammalian Mre11 in classical and alternative nonhomologous end joining. Nat Struct Mol Biol 16(8):814–818
Zhang Y, Jasin M (2011) An essential role for CtIP in chromosomal translocation formation through an alternative end-joining pathway. Nat Struct Mol Biol 18(1):80–84
Wang M, Wu W, Wu W, Rosidi B, Zhang L, Wang H, Iliakis G (2006) PARP-1 and Ku compete for repair of DNA double strand breaks by distinct NHEJ pathways. Nucleic Acids Res 34(21):6170–6182
Haince JF, McDonald D, Rodrigue A, Dery U, Masson JY, Hendzel MJ, Poirier GG (2008) PARP1-dependent kinetics of recruitment of MRE11 and NBS1 proteins to multiple DNA damage sites. J Biol Chem 283(2):1197–1208
Robert I, Dantzer F, Reina-San-Martin B (2009) Parp1 facilitates alternative NHEJ, whereas Parp2 suppresses IgH/c-myc translocations during immunoglobulin class switch recombination. J Exp Med 206(5):1047–1056
Cheng Q, Barboule N, Frit P, Gomez D, Bombarde O, Couderc B, Ren GS, Salles B, Calsou P (2011) Ku counteracts mobilization of PARP1 and MRN in chromatin damaged with DNA double-strand breaks. Nucleic Acids Res 39(22):9605–9619
Sfeir A, de Lange T (2012) Removal of shelterin reveals the telomere end-protection problem. Science 336(6081):593–597
Jia Q, den Dulk-Ras A, Shen H, Hooykaas PJ, de Pater S (2013) Poly(ADP-ribose)polymerases are involved in microhomology mediated back-up non-homologous end joining in Arabidopsis thaliana. Plant Mol Biol 82(4–5):339–351
Villarreal DD, Lee K, Deem A, Shim EY, Malkova A, Lee SE (2012) Microhomology directs diverse DNA break repair pathways and chromosomal translocations. PLoS Genet 8(11):e1003026
Ahmad A, Robinson AR, Duensing A, van Drunen E, Beverloo HB, Weisberg DB, Hasty P, Hoeijmakers JH, Niedernhofer LJ (2008) ERCC1-XPF endonuclease facilitates DNA double-strand break repair. Mol Cell Biol 28(16):5082–5092
Wang H, Rosidi B, Perrault R, Wang M, Zhang L, Windhofer F, Iliakis G (2005) DNA ligase III as a candidate component of backup pathways of nonhomologous end joining. Cancer Res 65(10):4020–4030
Liang L, Deng L, Nguyen SC, Zhao X, Maulion CD, Shao C, Tischfield JA (2008) Human DNA ligases I and III, but not ligase IV, are required for microhomology-mediated end joining of DNA double-strand breaks. Nucleic Acids Res 36(10):3297–3310
Sallmyr A, Tomkinson AE, Rassool FV (2008) Up-regulation of WRN and DNA ligase IIIα in chronic myeloid leukemia: consequences for the repair of DNA double-strand breaks. Blood 112(4):1413–1423
Zha S, Boboila C, Alt FW (2009) Mre11: roles in DNA repair beyond homologous recombination. Nat Struct Mol Biol 16(8):798–800
Lee-Theilen M, Matthews AJ, Kelly D, Zheng S, Chaudhuri J (2011) CtIP promotes microhomology-mediated alternative end joining during class-switch recombination. Nat Struct Mol Biol 18(1):75–79
Simsek D, Brunet E, Wong SY, Katyal S, Gao Y, McKinnon PJ, Lou J, Zhang L, Li J, Rebar EJ, Gregory PD, Holmes MC, Jasin M (2011) DNA ligase III promotes alternative nonhomologous end-joining during chromosomal translocation formation. PLoS Genet 7(6):e1002080
Paul K, Wang M, Mladenov E, Bencsik-Theilen A, Bednar T, Wu W, Arakawa H, Iliakis G (2013) DNA ligases I and III cooperate in alternative non-homologous end-joining in vertebrates. PLoS One 8(3):e59505
McVey M, Lee SE (2008) MMEJ repair of double-strand breaks (director’s cut): deleted sequences and alternative endings. Trends Genet 24(11):529–538
Truong LN, Li Y, Shi LZ, Hwang PY, He J, Wang H, Razavian N, Berns MW, Wu X (2013) Microhomology-mediated End Joining and Homologous Recombination share the initial end resection step to repair DNA double-strand breaks in mammalian cells. Proc Natl Acad Sci USA 110(19):7720–7725
Cairns J (1975) Mutation selection and the natural history of cancer. Nature 255(5505):197–200
Potten CS, Hume WJ, Reid P, Cairns J (1978) The segregation of DNA in epithelial stem cells. Cell 15(3):899–906
Potten CS, Owen G, Booth D (2002) Intestinal stem cells protect their genome by selective segregation of template DNA strands. J Cell Sci 115(Pt 11):2381–2388
Smith GH (2005) Label-retaining epithelial cells in mouse mammary gland divide asymmetrically and retain their template DNA strands. Development 132(4):681–687
Shinin V, Gayraud-Morel B, Gomes D, Tajbakhsh S (2006) Asymmetric division and cosegregation of template DNA strands in adult muscle satellite cells. Nat Cell Biol 8(7):677–687
Conboy MJ, Karasov AO, Rando TA (2007) High incidence of non-random template strand segregation and asymmetric fate determination in dividing stem cells and their progeny. PLoS Biol 5(5):e102
Merok JR, Lansita JA, Tunstead JR, Sherley JL (2002) Cosegregation of chromosomes containing immortal DNA strands in cells that cycle with asymmetric stem cell kinetics. Cancer Res 62(23):6791–6795
Karpowicz P, Morshead C, Kam A, Jervis E, Ramunas J, Cheng V, van der Kooy D (2005) Support for the immortal strand hypothesis: neural stem cells partition DNA asymmetrically in vitro. J Cell Biol 170(5):721–732
Falconer E, Chavez E, Henderson A, Lansdorp PM (2010) Chromosome orientation fluorescence in situ hybridization to study sister chromatid segregation in vivo. Nat Protoc 5(7):1362–1377
Sundararaman B, Avitabile D, Konstandin MH, Cottage CT, Gude N, Sussman MA (2012) Asymmetric chromatid segregation in cardiac progenitor cells is enhanced by Pim-1 kinase. Circ Res 110(9):1169–1173
Tomasetti C, Bozic I (2015) The (not so) immortal strand hypothesis. Stem Cell Res 14(2):238–241
Escobar M, Nicolas P, Sangar F, Laurent-Chabalier S, Clair P, Joubert D, Jay P, Legraverend C (2011) Intestinal epithelial stem cells do not protect their genome by asymmetric chromosome segregation. Nat Commun 2:e258
Ciccia A, Elledge SJ (2010) The DNA damage response: making it safe to play with knives. Mol Cell 40(2):179–204
Marion RM, Strati K, Li H, Murga M, Blanco R, Ortega S, Fernandez-Capetillo O, Serrano M, Blasco MA (2009) A p53-mediated DNA damage response limits reprogramming to ensure iPS cell genomic integrity. Nature 460(7259):1149–1153
Barlow JL, Drynan LF, Hewett DR, Holmes LR, Lorenzo-Abalde S, Lane AL, Jolin HE, Pannell R, Middleton AJ, Wong SH, Warren AJ, Wainscoat JS, Boultwood J, McKenzie AN (2010) A p53-dependent mechanism underlies macrocytic anemia in a mouse model of human 5q-syndrome. Nat Med 16(1):59–66
Ceccaldi R, Parmar K, Mouly E, Delord M, Kim JM, Regairaz M, Pla M, Vasquez N, Zhang QS, Pondarre C, Peffault de Latour R, Gluckman E, Cavazzana-Calvo M, Leblanc T, Larghero J, Grompe M, Socie G, D’Andrea AD, Soulier J (2012) Bone marrow failure in Fanconi anemia is triggered by an exacerbated p53/p21 DNA damage response that impairs hematopoietic stem and progenitor cells. Cell Stem Cell 11(1):36–49
Zenz T, Eichhorst B, Busch R, Denzel T, Habe S, Winkler D, Buhler A, Edelmann J, Bergmann M, Hopfinger G, Hensel M, Hallek M, Dohner H, Stilgenbauer S (2010) TP53 mutation and survival in chronic lymphocytic leukemia. J Clin Oncol 28(29):4473–4479
Zhao Z, Zuber J, Diaz-Flores E, Lintault L, Kogan SC, Shannon K, Lowe SW (2010) p53 loss promotes acute myeloid leukemia by enabling aberrant self-renewal. Genes Dev 24(13):1389–1402
Meijne EI, van der Winden-van Groenewegen RJ, Ploemacher RE, Vos O, David JA, Huiskamp R (1991) The effects of x-irradiation on hematopoietic stem cell compartments in the mouse. Exp Hematol 19(7):617–623
Down JD, Boudewijn A, van Os R, Thames HD, Ploemacher RE (1995) Variations in radiation sensitivity and repair among different hematopoietic stem cell subsets following fractionated irradiation. Blood 86(1):122–127
Song S, Lambert PF (1999) Different responses of epidermal and hair follicular cells to radiation correlate with distinct patterns of p53 and p21 induction. Am J Pathol 155(4):1121–1127
Merritt AJ, Potten CS, Kemp CJ, Hickman JA, Balmain A, Lane DP, Hall PA (1994) The role of p53 in spontaneous and radiation-induced apoptosis in the gastrointestinal tract of normal and p53-deficient mice. Cancer Res 54(3):614–617
Merritt AJ, Potten CS, Watson AJ, Loh DY, Nakayama K, Nakayama K, Hickman JA (1995) Differential expression of bcl-2 in intestinal epithelia. Correlation with attenuation of apoptosis in colonic crypts and the incidence of colonic neoplasia. J Cell Sci 108(Pt 6):2261–2271
Milyavsky M, Gan OI, Trottier M, Komosa M, Tabach O, Notta F, Lechman E, Hermans KG, Eppert K, Konovalova Z, Ornatsky O, Domany E, Meyn MS, Dick JE (2010) A distinctive DNA damage response in human hematopoietic stem cells reveals an apoptosis-independent role for p53 in self-renewal. Cell Stem Cell 7(2):186–197
Warr MR, Binnewies M, Flach J, Reynaud D, Garg T, Malhotra R, Debnath J, Passegue E (2013) FOXO3A directs a protective autophagy program in haematopoietic stem cells. Nature 494(7437):323–327
Rodolfo C, Di Bartolomeo S, Cecconi F (2015) Autophagy in stem and progenitor cells. Cell Mol Life Sci (Epub ahead of print)
Owusu-Ansah E, Banerjee U (2009) Reactive oxygen species prime Drosophila haematopoietic progenitors for differentiation. Nature 461(7263):537–541
Inomata K, Aoto T, Binh NT, Okamoto N, Tanimura S, Wakayama T, Iseki S, Hara E, Masunaga T, Shimizu H, Nishimura EK (2009) Genotoxic stress abrogates renewal of melanocyte stem cells by triggering their differentiation. Cell 137(6):1088–1099
Wang J, Sun Q, Morita Y, Jiang H, Gross A, Lechel A, Hildner K, Guachalla LM, Gompf A, Hartmann D, Schambach A, Wuestefeld T, Dauch D, Schrezenmeier H, Hofmann WK, Nakauchi H, Ju Z, Kestler HA, Zender L, Rudolph KL (2012) A differentiation checkpoint limits hematopoietic stem cell self-renewal in response to DNA damage. Cell 148(5):1001–1014
Granger MP, Wright WE, Shay JW (2002) Telomerase in cancer and aging. Crit Rev Oncol Hematol 41(1):29–40
Kim Sh SH, Kaminker P, Campisi J (2002) Telomeres, aging and cancer: in search of a happy ending. Oncogene 21(4):503–511
Wong JM, Collins K (2003) Telomere maintenance and disease. Lancet 362(9388):983–988
Feldser DM, Hackett JA, Greider CW (2003) Telomere dysfunction and the initiation of genome instability. Nat Rev Cancer 3(8):623–627
Blasco MA (2005) Telomeres and human disease: ageing, cancer and beyond. Nat Rev Genet 6(8):611–622
Greider CW, Blackburn EH (1985) Identification of a specific telomere terminal transferase activity in Tetrahymena extracts. Cell 43(2 Pt 1):405–413
Shay JW, Zou Y, Hiyama E, Wright WE (2001) Telomerase and cancer. Hum Mol Genet 10(7):677–685
Allsopp RC, Morin GB, DePinho R, Harley CB, Weissman IL (2003) Telomerase is required to slow telomere shortening and extend replicative lifespan of HSCs during serial transplantation. Blood 102(2):517–520
Yeager TR, Neumann AA, Englezou A, Huschtscha LI, Noble JR, Reddel RR (1999) Telomerase-negative immortalized human cells contain a novel type of promyelocytic leukemia (PML) body. Cancer Res 59(17):4175–4179
Dunham MA, Neumann AA, Fasching CL, Reddel RR (2000) Telomere maintenance by recombination in human cells. Nat Genet 26(4):447–450
Fasching CL, Neumann AA, Muntoni A, Yeager TR, Reddel RR (2007) DNA damage induces alternative lengthening of telomeres (ALT) associated promyelocytic leukemia bodies that preferentially associate with linear telomeric DNA. Cancer Res 67(15):7072–7077
Grobelny JV, Godwin AK, Broccoli D (2000) ALT-associated PML bodies are present in viable cells and are enriched in cells in the G(2)/M phase of the cell cycle. J Cell Sci 113(Pt 24):4577–4585
Wu G, Lee WH, Chen PL (2000) NBS1 and TRF1 colocalize at promyelocytic leukemia bodies during late S/G2 phases in immortalized telomerase-negative cells. Implication of NBS1 in alternative lengthening of telomeres. J Biol Chem 275(39):30618–30622
Wu G, Jiang X, Lee WH, Chen PL (2003) Assembly of functional ALT-associated promyelocytic leukemia bodies requires Nijmegen Breakage Syndrome 1. Cancer Res 63(10):2589–2595
Nabetani A, Yokoyama O, Ishikawa F (2004) Localization of hRad9, hHus1, hRad1, and hRad17 and caffeine-sensitive DNA replication at the alternative lengthening of telomeres-associated promyelocytic leukemia body. J Biol Chem 279(24):25849–25857
Liu L, Bailey SM, Okuka M, Munoz P, Li C, Zhou L, Wu C, Czerwiec E, Sandler L, Seyfang A, Blasco MA, Keefe DL (2007) Telomere lengthening early in development. Nat Cell Biol 9(12):1436–1441
Bechter OE, Zou Y, Walker W, Wright WE, Shay JW (2004) Telomeric recombination in mismatch repair deficient human colon cancer cells after telomerase inhibition. Cancer Res 64(10):3444–3451
Hu J, Hwang SS, Liesa M, Gan B, Sahin E, Jaskelioff M, Ding Z, Ying H, Boutin AT, Zhang H, Johnson S, Ivanova E, Kost-Alimova M, Protopopov A, Wang YA, Shirihai OS, Chin L, Depinho RA (2012) Antitelomerase therapy provokes ALT and mitochondrial adaptive mechanisms in cancer. Cell 148(4):651–663
Oganesian L, Karlseder J (2011) Mammalian 5′ C-rich telomeric overhangs are a mark of recombination-dependent telomere maintenance. Mol Cell 42(2):224–236
Heaphy CM, de Wilde RF, Jiao Y, Klein AP, Edil BH, Shi C, Bettegowda C, Rodriguez FJ, Eberhart CG, Hebbar S, Offerhaus GJ, McLendon R, Rasheed BA, He Y, Yan H, Bigner DD, Oba-Shinjo SM, Marie SK, Riggins GJ, Kinzler KW, Vogelstein B, Hruban RH, Maitra A, Papadopoulos N, Meeker AK (2011) Altered telomeres in tumors with ATRX and DAXX mutations. Science 333(6041):425
Schwartzentruber J, Korshunov A, Liu XY, Jones DT, Pfaff E, Jacob K, Sturm D, Fontebasso AM, Quang DA, Tonjes M, Hovestadt V, Albrecht S, Kool M, Nantel A, Konermann C, Lindroth A, Jager N, Rausch T, Ryzhova M, Korbel JO, Hielscher T, Hauser P, Garami M, Klekner A, Bognar L, Ebinger M, Schuhmann MU, Scheurlen W, Pekrun A, Fruhwald MC, Roggendorf W, Kramm C, Durken M, Atkinson J, Lepage P, Montpetit A, Zakrzewska M, Zakrzewski K, Liberski PP, Dong Z, Siegel P, Kulozik AE, Zapatka M, Guha A, Malkin D, Felsberg J, Reifenberger G, von Deimling A, Ichimura K, Collins VP, Witt H, Milde T, Witt O, Zhang C, Castelo-Branco P, Lichter P, Faury D, Tabori U, Plass C, Majewski J, Pfister SM, Jabado N (2012) Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482(7384):226–231
Bower K, Napier CE, Cole SL, Dagg RA, Lau LM, Duncan EL, Moy EL, Reddel RR (2012) Loss of wild-type ATRX expression in somatic cell hybrids segregates with activation of alternative lengthening of telomeres. PLoS One 7(11):e50062
Acknowledgments
Due to the scope of the reviewed subject, all studies as relevant and deserving as those cited may not be exhaustively enlisted in this article. R.Y.T. is in part supported by NCI-PHS Grant R03 CA201988.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares no conflict of interest.
Rights and permissions
About this article
Cite this article
Tsai, R.Y.L. Balancing self-renewal against genome preservation in stem cells: How do they manage to have the cake and eat it too?. Cell. Mol. Life Sci. 73, 1803–1823 (2016). https://doi.org/10.1007/s00018-016-2152-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-016-2152-y