Abstract
Of 504 tonsil samples from slaughtered fattening pigs, 5.6 % were positive for Listeria monocytogenes by culture after enrichment. The 28 L. monocytogenes isolates were analyzed to gain insights into genetic relationships and virulence-associated traits. Of the 28 isolates, 57 % belonged to serotype 1/2a (genetic lineage II), 25 % to serotype 4b (genetic lineage I) and 18 % to serotype 1/2b (genetic lineage I). These serotypes are commonly associated with human listeriosis cases. Multilocus sequence typing assigned the 28 isolates to 16 clonal complexes (CCs) and three singletons, including one new sequence type (ST768). Several of these CCs were also found in strains from human infections. Sequence analysis of the whole internalin A gene (inlA) showed that all but one isolate (CC6/serotype 4b) encoded full-length proteins. Clinical strains from human patients commonly harbor full-length inlA. On the other hand, genes for benzalkonium chloride tolerance were not found and all but one isolate lacked genes for the stress survival islet (SSI-1). Thus, tonsils of slaughtered fattening pigs can be colonized with L. monocytogenes of public health impact. To counteract this threat during slaughter, prevention of contamination of carcasses and the environment is of major importance, in particular adherence to good slaughter hygiene practice. With regard to pig tonsils, special attention must be given to the handling and contamination of head meat and pig tongues.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Listeria monocytogenes is an important foodborne pathogen with a significant impact on public health and economy worldwide. L. monocytogenes has the potential to cause serious and potentially life-threatening conditions (including septicemia, meningitis, meningoencephalitis and abortion) in persons with diminished immunity (Allerberger and Wagner 2010). Because of its high case fatality rate in persons at risk, listeriosis ranks among the most frequent causes of death due to foodborne illness. In the European Union, a total of 1763 confirmed human cases of listeriosis (notification rate of 0.44 cases per 100,000 population) were reported in 2013 (EFSA/ECDC 2015). L. monocytogenes are widely distributed in the environment and certain strains may become established and persist in the processing environment (Blatter et al. 2010; Carpentier and Cerf 2011; Larivière-Gauthier et al. 2014). Other reservoirs include domestic and wild animals, but their significance in view of foodborne diseases and potential transmission routes (during slaughter) remains to be elucidated.
With regard to slaughtered pigs, the animals might be asymptomatic carriers of important human pathogens such as L. monocytogenes. Several studies thereby reported that L. monocytogenes was isolated more frequently from pig tonsils than from fecal samples, but detection rates in the tonsils of slaughtered pigs varied considerably (Buncic 1991; Fredriksson-Ahomaa et al. 2009; Kanuganti et al. 2002). Pig tonsils may play a role in the contamination of pluck sets, carcasses and the slaughterhouse environment during slaughter (Fredriksson-Ahomaa et al. 2009). To evaluate the potential health risk for humans, data on the animals’ probability of carrying L. monocytogenes must be complemented by analysis of genetic relationships and virulence-associated traits of strains. The aim of the present study was therefore to determine the occurrence of L. monocytogenes in the tonsils from fattening pigs at slaughter and to further characterize isolated L. monocytogenes. Thereby, serotypes of isolated L. monocytogenes were compared with those associated with human infections, multilocus sequence typing (MLST) was performed to get information on genetic relationships (between isolates and potential human associations) and internalin A gene sequence profiling was used to detect mutations that might attenuate the virulence. Moreover, factors contributing to the persistence of L. monocytogenes in food-associated environments (benzalkonium chloride tolerance, stress survival islet) were examined.
2 Materials and methods
2.1 Abattoir and sampling
In an abattoir processing about 250 pig carcasses per hour, 504 tonsil samples were collected from 504 healthy fattening pigs. Sampled pigs originated from the north and central part of Switzerland and were about 6 months old. Sampling comprised two phases: in the first phase (March–May 2011), 250 carcasses from 126 batches were sampled; in the second phase (October 2014), 254 carcasses from 177 batches were sampled. Tonsil samples (Tonsilla veli palatini) were obtained at the end of slaughtering when the head had been cut off. Samples were excised using sterile forceps and scissors, placed into stomacher bags and transported cooled to the laboratory. Examinations in the laboratory were performed within 4 h after sampling.
2.2 Detection and identification of L. monocytogenes
Examination for L. monocytogenes was performed in accordance with ISO 11290-1:2004 using a two-step enrichment procedure (Half-Fraser broth, 24 h at 30 °C; Fraser broth 24 h at 37 °C; Oxoid, Pratteln, Switzerland). Subcultures were streaked onto Chromogenic Listeria Agar (Oxoid) supplemented with Listeria Selective and Differential Supplement and incubated for 48 h at 37 °C. Presumptive L. monocytogenes colonies (green–blue colonies surrounded by an opaque halo on the chromogenic agar) were streaked onto sheep blood agar for appraisal of hemolysis.
2.3 Serotyping and lineage PCR
Listeria monocytogenes isolates were serotyped at the Swiss National Reference Center for Enteropathogenic Bacteria and Listeria (NENT; Institute for Food Safety and Hygiene, University of Zurich) using the commercial set of Listeria O-factor and H-factor antisera from Denka Seiken (Pharma Consulting, Burgdorf, Switzerland). The lineage-specific multiplex PCR assay was performed using the primers and protocol described by Ward et al. (2004), with minor modifications. Briefly, amplifications were performed in 20-µl volumes with 0.4 µM final primer concentrations (actA1, plcB2 and actA3-plcB3), 10 µl of GoTaq Green Master Mix (Promega, Madison, WI, USA) and 100 ng of extracted genomic DNA (DNeasy Blood and Tissue Kit; Qiagen, Hilden, Germany). The cycling conditions were: 94 °C for 3 min, followed by 30 cycles of 94 °C for 30 s, 56 °C for 30 s, 72 °C for 30 s and a final step at 72 °C for 2 min.
2.4 Multilocus sequence typing (MLST)
MLST was performed as described by Ragon et al. (2008) with the following modifications: amplifications were performed in 50-µl volumes with 0.4 µM final primer concentrations, 25 µl of HotStarTaq Master Mix (Qiagen) and 50 ng of extracted genomic DNA (Qiagen). The cycling conditions were: for the bglA, cat and ldh loci: 94 °C for 4 min, followed by 35 cycles of 94 °C for 30 s, 45 °C for 30 s and 72 °C for 2 min; and for the abcZ, dapE, dat and lhkA loci: 94 °C for 4 min, followed by 35 cycles of 94 °C for 30 s, 52 °C for 30 s and 72 °C for 2 min. Amplifications were concluded with a 10-min 72 °C step. Amplicons were purified using the GenElute™ PCR Clean-Up Kit (Sigma–Aldrich, St Louis, MO, USA). Sequencing of the amplicons (Microsynth, Balgach, Switzerland) was performed using the universal primers described by Ragon et al. (2008). Alleles and sequence types (STs) are publicly available at http://www.pasteur.fr/mlst.
2.5 Internalin A gene (inlA) sequence profiling
To cover full-length inlA, four pairs of previously described primers were used for inlA profiling (Orsi et al. 2007). Amplifications were performed in 50-μl volumes using the Phusion Green Hot Start II High-Fidelity DNA Polymerase in accordance with the manufacturers’ instructions (Thermo Fisher Scientific, Waltham, MA, USA). The cycling conditions were: 98 °C for 30 s, followed by 35 cycles of 98 °C for 5 s, 50 °C for 30 s, 72 °C for 1.5 min and a final step at 72 °C for 10 min. Amplicons were sequenced commercially by Microsynth and analyzed for mutations with CLC Main Workbench 6.9.1.
2.6 Benzalkonium chloride (BC) tolerance and stress survival islet (SSI-1)
BC tolerance was examined by testing for bcrABC genes using primers p1 and p2 (Elhanafi et al. 2010) and for the Tn6188 transposon (Müller et al. 2013). Amplifications were performed using the GoTaq Green Master Mix (Promega). In addition, the presence of SSI-1 (lmo0444-lmo0448) was examined as described by Ryan et al. (2010). The Phusion Green Hot Start II High-Fidelity DNA Polymerase was used for the amplifications in accordance with the manufacturers’ instructions (Thermo Fisher Scientific). The cycling conditions were (lmo0444-lmo0448): 98 °C for 30 s, followed by 35 cycles of 98 °C for 5 s, 59 °C for 15 s, 72 °C for 4.5 min and a final step at 72 °C for 10 min.
3 Results and discussion
3.1 Detection, serotypes and genetic lineages of L. monocytogenes
Listeria monocytogenes were detected by culture after enrichment in 28 (5.6 %) of the 504 tonsil samples obtained from 504 slaughtered fattening pigs. With regard to the two sampling phases (March–May 2011; October 2014), 3.2 % (8/250) of the animals and 6.3 % (8/126) of the batches tested positive in the first sampling phase and 7.9 % (20/254) of the animals and 11.3 % (20/177) of the batches in the second sampling phase. The results from the two sampling phases differed significantly at the animal level (Chi-square test, P < 0.05), but the cause for this difference could not be assigned. In other European studies examining pig tonsils, the prevalence of L. monocytogenes ranged from 14 to 45 % (Autio et al. 2004; Buncic 1991; Fredriksson-Ahomaa et al. 2009).
Of the 28 L. monocytogenes strains isolated from pig tonsils in the present study (Table 1), 16 (57 %) belonged to serotype 1/2a (genetic lineage II), seven (25 %) to serotype 4b (genetic lineage I) and five (18 %) to serotype 1/2b (genetic lineage I). Serotype 1/2a strains are frequently found in food products (Gianfranceschi et al. 2009; Martín et al. 2014; Parisi et al. 2010). The majority of human listeriosis cases are associated with serotype 1/2a, 1/2b and 4b strains and infections due to serotype 1/2a strains have increased in recent years (Althaus et al. 2014; Gianfranceschi et al. 2009; Lopez-Valladares et al. 2014). In a current Swiss study characterizing 93 L. monocytogenes strains isolated during 2011–2013 from human infections (Althaus et al. 2014), the majority of the strains belonged to serotype 1/2a (62.4 %), followed by serotypes 4b (30.1 %), 1/2b and 1/2c.
3.2 Multilocus sequence typing
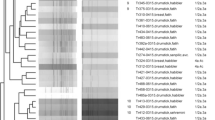
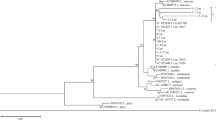
MLST grouped the 28 L. monocytogenes isolates into 19 STs that were assigned to 16 clonal complexes (CC1, CC4, CC5, CC6, CC8, CC20, CC21, CC26, CC29, CC59, CC177, CC224, CC403, CC412, CC415, CC451) and three singletons (ST200, ST226, ST768). ST768 was thereby designated for the first time and the respective isolate belonged to serotype 1/2a. Table 1 shows the distribution of CCs among serotype 1/2a, 1/2b and 4b isolates of the two sampling phases. CC1 (serotype 4b isolates), CC6 (serotype 4b isolates), CC20 (serotype 1/2a isolates) and CC224 (serotype 1/2b isolates) were thereby identified among strains from both sampling phases. Overall, the results show the clonal heterogeneity of L. monocytogenes from pig tonsils. Great genetic diversity has also been described in human- and food-derived L. monocytogenes strains (Althaus et al. 2014; Haase et al. 2014; Parisi et al. 2010). Twelve of the CCs identified among the isolates from pig tonsils in the present study (CC1, CC4, CC5, CC6, CC8, CC21, CC26, CC29, CC59, CC224, CC403, CC412) were also found in L. monocytogenes strains from human patients in Switzerland (Althaus et al. 2014).
3.3 Internalin A
Internalin A is a cell wall anchored protein, which facilitates the invasion of intestinal cells through interaction with E-cadherin receptors (Seveau et al. 2007). Multiple inlA mutations leading to premature stop codons (PMSCs) have been reported and these mutations might attenuate the virulence (Seveau et al. 2007; Van Stelten et al. 2010). Clinical L. monocytogenes strains from human patients commonly harbor full-length inlA (Althaus et al. 2014; Van Stelten et al. 2010). In the present study, sequence analysis of the inlA gene showed that all but one isolate encoded full-length proteins. The respective CC6/serotype 4b isolate showed an inlA gene PMSC mutation caused by a three-codon deletion in amino acid position 738–740.
3.4 Benzalkonium chloride tolerance and stress survival islet
Increased BC tolerance and genes within the SSI-1 contribute to the persistence of L. monocytogenes in food-associated environments (Mullapudi et al. 2008; Ryan et al. 2010). Of the 28 L. monocytogenes isolates from pig tonsils, none harbored bcrABC genes or the Tn6188 transposon. SSI-1 genes were only detected in one (CC224/serotype 1/2b) of the 28 isolates. Thus, these mechanisms promoting survival and growth of L. monocytogenes under food-associated stress conditions seem not to be widespread among isolates from tonsils of slaughtered fattening pigs.
4 Conclusions
Listeria monocytogenes were found in the tonsils of slaughtered fattening pigs in Switzerland (detection rate of 5.6 % at the animal level), but the prevalence was lower compared to the results from other European countries. Further characterization of the L. monocytogenes isolates from pig tonsils showed that the identified serotypes are commonly associated with human listeriosis cases, several of the clonal complexes were also found in strains from human patients and full-length internalin A genes predominated among them (as in clinical strains). Thus, tonsils of slaughtered fattening pigs pose a threat for the contamination of carcasses and the slaughterhouse environment with L. monocytogenes representing a potential health risk. To counteract this threat, prevention of contamination during slaughter is of major importance, in particular adherence to good slaughter hygiene practices and application of effective cleaning and disinfection procedures. With regard to pig tonsils colonized with L. monocytogenes (and other important bacterial pathogens as e.g. Yersinia enterocolitica), special attention must be given to the handling and the potential contamination of the head meat, pig cheeks and pig tongues.
References
Allerberger F, Wagner M (2010) Listeriosis: a resurgent foodborne infection. Clin Microbiol Infect 16:16–23. doi:10.1111/j.1469-0691.2009.03109.x
Althaus D, Lehner A, Brisse S, Maury M, Tasara T, Stephan R (2014) Characterization of Listeria monocytogenes strains isolated during 2011–2013 from human infections in Switzerland. Foodborne Pathog Dis 11:753–758. doi:10.1089/fpd.2014.1747
Autio T, Markkula A, Hellström S, Niskanen T, Lundén J, Korkeala H (2004) Prevalence and genetic diversity of Listeria monocytogenes in the tonsils of pigs. J Food Prot 67:805–808
Blatter S, Giezendanner N, Stephan R, Zweifel C (2010) Phenotypic and molecular typing of Listeria monocytogenes isolated from the processing environment and products of a sandwich-producing plant. Food Control 21:1519–1523. doi:10.1016/j.foodcont.2010.04.025
Buncic S (1991) The incidence of Listeria monocytogenes in slaughtered animals, in meat, and in meat products in Yugoslavia. Int J Food Microbiol 12:173–180. doi:10.1016/0168-1605(91)90067-Y
Carpentier B, Cerf O (2011) Persistence of Listeria monocytogenes in food industry equipment and premises. Int J Food Microbiol 145:1–8. doi:10.1016/j.ijfoodmicro.2011.01.005
EFSA/ECDC (2015) The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2013. EFSA J 13(1):3991. doi:10.2903/j.efsa.2015.3991
Elhanafi D, Dutta V, Kathariou S (2010) Genetic characterization of plasmid-associated benzalkonium chloride resistance determinants in a Listeria monocytogenes strain from the 1998–1999 outbreak. Appl Environ Microbiol 76:8231–8238. doi:10.1128/AEM.02056-10
Fredriksson-Ahomaa M, Gerhardt M, Stolle A (2009) High bacterial contamination of pig tonsils at slaughter. Meat Sci 83:334–336. doi:10.1016/j.meatsci.2009.06.004
Gianfranceschi MV, D’Ottavio MC, Gattuso A, Bella A, Aureli P (2009) Distribution of serotypes and pulsotypes of Listeria monocytogenes from human, food and environmental isolates (Italy 2002–2005). Food Microbiol 26:520–526. doi:10.1016/j.fm.2009.03.003
Haase JK, Didelot X, Lecuit M, Korkeala H, L. monocytogenes MLST Study Group, Achtman M (2014) The ubiquitous nature of Listeria monocytogenes clones: a large-scale multi-locus sequence typing study. Environ Microbiol 16:405–416. doi:10.1111/1462-2920.12342
Kanuganti SR, Wesley IV, Reddy PG, McKean J, Hurd HS (2002) Detection of Listeria monocytogenes in pigs and pork. J Food Prot 65:1470–1474
Larivière-Gauthier G, Letellier A, Kérouanton A, Bekal S, Quessy S, Fournaise S, Fravalo P (2014) Analysis of Listeria monocytogenes strain distribution in a pork slaughter and cutting plant in the province of Quebec. J Food Prot 77:2121–2128. doi:10.4315/0362-028X.JFP-14-192
Lopez-Valladares G, Tham W, Parihar VS, Helmersson S, Andersson B, Ivarsson S, Johansson C, Ringberg H, Tjernberg I, Henriques-Normark B, Danielsson-Tham M-L (2014) Human isolates of Listeria monocytogenes in Sweden during half a century (1958–2010). Epidemiol Infect 142:2251–2260. doi:10.1017/S0950268813003385
Martín B, Perich A, Gómez D, Yangüela J, Rodríguez A, Garriga M, Aymerich T (2014) Diversity and distribution of Listeria monocytogenes in meat processing plants. Food Microbiol 44:119–127. doi:10.1016/j.fm.2014.05.014
Mullapudi S, Siletzky RM, Kathariou S (2008) Heavy-metal and benzalkonium chloride resistance of Listeria monocytogenes isolates from the environment of turkey-processing plants. Appl Environ Microbiol 74:1464–1468. doi:10.1128/AEM.02426-07
Müller A, Rychli K, Muhterem-Uyar M, Zaiser A, Stessl B, Guinane CM, Cotter PD, Wagner M, Schmitz-Esser S (2013) Tn6188—a novel transposon in Listeria monocytogenes responsible for tolerance to benzalkonium chloride. PLoS One 8(10):e76835. doi:10.1371/journal.pone.0076835
Orsi RH, Ripoll DR, Yeung M, Nightingale KK, Wiedmann M (2007) Recombination and positive selection contribute to evolution of Listeria monocytogenes inlA. Microbiology 153:2666–2678. doi:10.1099/mic.0.2007/007310-0
Parisi A, Latorre L, Normanno G, Miccolupo A, Fraccalvieri R, Lorusso V, Santagada G (2010) Amplified fragment length polymorphism and multi-locus sequence typing for high-resolution genotyping of Listeria monocytogenes from foods and the environment. Food Microbiol 27:101–108. doi:10.1016/j.fm.2009.09.001
Ragon M, Wirth T, Hollandt F, Lavenir R, Lecuit M, Le Monnier A, Brisse S (2008) A new perspective on Listeria monocytogenes evolution. PLoS Pathog 4(9):e1000146. doi:10.1371/journal.ppat.1000146
Ryan S, Begley M, Hill C, Gahan CG (2010) A five-gene stress survival islet (SSI-1) that contributes to the growth of Listeria monocytogenes in suboptimal conditions. J Appl Microbiol 109:984–995. doi:10.1111/j.1365-2672.2010.04726.x
Seveau S, Pizarro-Cerda J, Cossart P (2007) Molecular mechanisms exploited by Listeria monocytogenes during host cell invasion. Microbes Infect 9:1167–1175. doi:10.1016/j.micinf.2007.05.004
Van Stelten A, Simpson JM, Ward TJ, Nightingale KK (2010) Revelation by single-nucleotide polymorphism genotyping that mutations leading to a premature stop codon in inlA are common among Listeria monocytogenes isolates from ready-to-eat foods but not human listeriosis cases. Appl Environ Microbiol 76:2783–2790. doi:10.1128/AEM.02651-09
Ward TJ, Gorski L, Borucki MK, Mandrell RE, Hutchins J, Pupedis K (2004) Intraspecific phylogeny and lineage group identification based on the prfA virulence gene cluster of Listeria monocytogenes. J Bacteriol 186:4994–5002. doi:10.1128/JB.186.15.4994-5002.2004
Acknowledgments
The authors thank the staff of the slaughterhouse for facilitating access to the operations and for assistance with the collection of data. Grethe Sägesser is kindly acknowledged for her support with serotyping of the strains.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sarno, E., Fierz, L., Zweifel, C. et al. Characteristics of Listeria monocytogenes isolated from tonsils of slaughtered fattening pigs in Switzerland. J. Verbr. Lebensm. 11, 19–23 (2016). https://doi.org/10.1007/s00003-015-0974-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00003-015-0974-4