Abstract
Traditional deep learning models have limited ability to extract features from pneumonia images. This study combines convolutional attention modules with transfer learning to improve the model's feature extraction ability and simplify training.This article has discovered an improved CBAM-Xception neural network pneumonia model. This network uses Xception as the main network, and combines with the convolutional attention CBAM module to enhance the expression of lesion information and suppress irrelevant information interference; At the same time, transfer learning is introduced to prevent over fitting when the sample data amount is small. In order to evaluate the effectiveness of the optimization model, experimental simulation tests were conducted on the Mendeley Data public pneumonia dataset. The improved model achieves 94.2% accuracy in classifying four images of COVID-19, bacterial pneumonia, viral pneumonia and normal chest on the test set, and the optimization effect is significant. In order to further verify the performance of this method, the experiment divided the small sample data set to train the model, and divided the large sample data to test the generalization performance of the model. The results show that the model in this paper has good generalization ability. This model can provide an important basis for the auxiliary diagnosis and treatment of pneumonia.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Since the COVID-19 pandemic, the word pneumonia has gradually become active in public view.Pneumonia is prone to infection in children and the elderly with weak resistance, and the mortality rate is increasing. As of May 16, 2023, there were approximately 676.1 million COVID-19 [1] cases worldwide and approximately 6 million 880 thousand million deaths from COVID-19.
With the development of computer hardware performance, artificial diagnosis is no longer the only solution. Hospitals have brought computer-assisted diagnosis (CAD) [2] into clinical practice to facilitate doctors’ reference and diagnosis. There are many different solutions for computer-aided diagnosis of pneumonia at home and abroad.Yuan Maozhou [3] used the LBP(Local Binary Pattern) algorithm and gray level co-occurrence matrix to extract the lung region features of the image, and the obtained features were recognized using a support vector machine classifier for the lung image. However, the new algorithm requires manual threshold setting, and the extracted lung features are similar. Yue Lu et al. [4] applied the decision tree algorithm to the data of 200 children with pneumonia, and finally achieved an accuracy rate of 80%. Jun et al. [5] used support vector machines [6] to classify two types of interstitial pneumonia, and compared with the recognition results of artificial pneumonia, the results showed a difference of only 5% to 6%. In the implementation scheme of pneumonia recognition, traditional algorithms have excellent performance, but with the gradual increase of data sets, the performance of traditional algorithms, both in terms of robustness and recognition efficiency, is unsatisfactory.
With the development of artificial intelligence and big data, artificial intelligence has been active in the field of disease image recognition as a new solution in recent years. Deep learning [7], as a popular solution in artificial intelligence, includes branch methods such as convolutional neural networks [8], and is gradually being applied to disease recognition. Zhou Qihao et al. [9] combined deep dense aggregation structure with DenseNet-201 and proposed a deep learning based classification network DLDA-A-DenseNet, which aggregates feature information from different stages and classifies the dataset provided by the China Chest CT Image Survey and Research Association. The recognition accuracy improved by 2.24% compared to the original DenseNet-201. Guo Yi et al. [10] used GhostNet lightweight network to simplify DenseNet network parameters, and classified the open COVID-19 dataset. Under the condition of ensuring the recognition accuracy of 83%, the medical computer took 236 ms. Khanun Roisatul Ummah et al. [11] evaluated Watershed segmentation, smoothing image method of Median and Gaussian filters, and other image preprocessing methods for automatic detection of new coronal pneumonia based on CT images. In order to further improve the classification accuracy of the model, scholars often increase the number of layers of the network. Although networks with more layers have improved classification accuracy, their parameters are far more than shallow networks, increasing the difficulty of model training and posing a serious challenge to computer performance. Therefore, this paper proposes a CBAM-Xception neural network that combines the convolutional attention CBAM [12] module with transfer learning [13]. Convolutional attention enhances the feature extraction ability of the model. Transfer learning is performed on the parameters trained in ImageNet [14] data, allowing the model to quickly fit the parameters, thereby reducing the burden of computer performance.
2 Model Selection and Improvement
2.1 Transfer Learning
During the actual data collection process, it is difficult to collect medical images of pneumonia due to patient privacy issues, as well as complex features of pneumonia lesions, varying sizes of lesions, and different manifestations of different lesions at different time points, making it difficult for pneumonia data to form a standard training database. However, using a small amount of data sets to train in the Xception network can result in model overfitting. Therefore, this article uses migration learning to pre train the model on the ImageNet dataset, and then migrate parameters with stronger general feature extraction capabilities to the Xception network. On this basis, the model is trained to improve the accuracy of model recognition and reduce the difficulty of computer training.
2.2 CBAM Module
Various types of pneumonia lesions vary in size at different stages and tend to concentrate in a single area. Allowing the model to assign more parameter weights to the lesion area will improve the recognition accuracy of the model. Therefore, this article introduces a CBAM module that integrates spatial attention and channel attention to improve the feature extraction ability of the model, extracting diverse lesion features in both spatial and channel aspects, and allocating more weight of model parameters to focus areas. The CBAM module structure is shown in Fig. 1. After combining the CBAM module with the Xception main network, the CBAM module extracts the focus features output from the Xception main network through both spatial and channel aspects, making the model focus on the focus area. The feature map output from the Xception main network is multiplied by the feature map output from the attention module to enhance the expression ability of features in the spatial channel dimension.
2.3 CBAM Xception Transfer Learning Pneumonia Detection Model
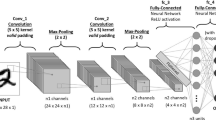
This article proposes a convolutional attention neural network CBAM-Xception for pneumonia lesion recognition by introducing the CBAM module, and simplify the model training process through transfer learning. The CBAM-Xception model consists of an Xception module, a CBAM module, and a fully connected layer module. The Xception module and CBAM module extract the features of the input pneumonia image and submit the extracted features to the classification module for classification. The model structure is shown in Fig. 2. The Xception feature extraction block contains deep separable convolutions. Deep separable convolution has a lower number of parameters than conventional convolution but shares the same feature extraction capability. The CBAM module includes channel attention modules and spatial attention modules. Better results can be achieved than an attentional mechanism that focuses only on the channel. The fully connected classification module and Softmax function classify the extracted features and output the confidence levels of four types of pneumonia labels.
3 Experiment and Analysis
3.1 Data Set
The data used in this paper is the pneumonia dataset from Mendeley Data, and the images of bacterial pneumonia, viral pneumonia, COVID-19 and other pneumonia are taken as research objects. To prevent network overfitting, four data enhancement operations are performed on the original dataset, including random rotation angle, random horizontal flip, shear change angle, and horizontal offset. The image enhancement effect is shown in Fig. 3.
3.2 Experimental Setup
Divide the pneumonia image dataset into training, validation, and testing sets in a ratio of 8:1:1. The experiment was conducted under the Windows 10 operating system, with Python version 3 6. The version of Tensorflow is 2.2.0. The model is constructed using the TensorFlow deep learning framework. The parameter optimizer is a gradient descent function, and the average cross entropy is a loss function. Train 100 samples per training session.
3.3 Model Identification Results
The four classification confusion matrix obtained from the test set classification in this model is shown in Fig. 4. The Kappa coefficient of the model is 0.917, and the overall classification accuracy (OA) is 94.20%. Most of the test data are concentrated on the diagonal of the confusion matrix, and the recognition rate is high. The recognition rate of each type of pneumonia is shown in Table 1. The recognition accuracy of both COVID-19 and normal pictures exceeded 97.00%; The recognition accuracy of bacterial pneumonia is the lowest, at 88.88%. By observing the confusion matrix, it can be found that bacterial pneumonia is easy to be identified as viral pneumonia, and viral pneumonia is also easy to be mistakenly classified as bacterial pneumonia by the model. Based on the comparison of the dataset, it can be seen that there is a small difference between bacterial pneumonia and viral pneumonia, making it prone to classification errors.
3.4 Model Performance Evaluation.
Ablation Experiment.
The ablation experiment was used to verify the effect of CBAM module and transfer learning, and compare the effects of Xception from zero training, CBAM-Xception from zero training, and CBAM-Xception transfer learning. The results of the ablation experiment are shown in Table 2. The three methods used the same experimental environment and dataset, only changed the experimental comparison section. By comparing the accuracy, loss value, and recall of the three methods, the performance of the methods is determined. The training process of different methods on the pneumonia dataset is shown in Figs. 5.
In Fig. 5, it can be seen that the recognition accuracy of our method is superior to other methods. After adding transfer learning, the model achieves higher accuracy and lower loss value, and the model can converge quickly. After adding the CBAM attention module to the network, the classification accuracy improved by 4.8% (Table 2). Therefore, transfer learning and CBAM module effectively enhance the ability of the model to extract pneumonia features. The above experiments show that CBAM-Xception transfer learning method can improve the accuracy of pneumonia classification for pneumonia dataset.
Generalization Performance Verification.
On data with the same probability distribution, partitioning a small number of datasets to train the model and testing a large amount of data on the model can evaluate the generalization performance of the model. In this experiment, the training set and test set are exchanged, and the ratio between the training set, validation set, and test set is 1:1:8. The transfer learning CBAM-Xception network model is trained with a training set with a small amount of data, and then the model is evaluated with a verification set and a test set with a large amount of data. The training process of model loss value and accuracy is shown in Fig. 6.
In Fig. 6, overfitting is normal because the number of training sets is too small. The accuracy rate has gradually converged, and the classification accuracy rate has reached about 70%. In the iteration interval from 0 to 20, the loss value also increases gradually due to overfitting of the model. The classification results of this model on the test set are shown in Table 3.
In Table 3, it is found that the model has a strong generalization ability in COVID-19, reaching an accuracy rate of 96.7%, and surpasses the Xception model in the classification and recognition of bacterial pneumonia and viral pneumonia. The accuracy of the test set is 3% less than that of the Xception model, which indicates that the model in this paper has excellent generalization.
4 Conclusion
This paper proposes a scheme combining convolutional attention CBAM module and transfer learning. The accuracy of 94.2% was achieved on the Mendeley Data public pneumonia dataset, and the improved model has higher recognition rate and excellent generalization compared to the original Xception model, providing a certain reference for the auxiliary diagnosis and treatment of pneumonia. In future work, the number of training images for bacterial pneumonia and viral pneumonia should be increased to further improve the classification performance of the model. In summary, this article has clinical significance and practical value, providing a good research method and approach for the auxiliary diagnosis of pneumonia.
References
Johns Hopkins university. COVID-19 Dashboard by the Center for Systems Science and Engineering at Johns Hopkins University. (16 May 2022).https://coronavirus.jhu.edu/map.html
Shiraishi, J., Li, Q., Appelbaum, D., et al.: Computer-aided diagnosis and artificial intelligence in clinical omaging. Seminars Nuclear Med. 41(6) (2011)
Maozhou, Y.: Research on feature extraction and classification of X-ray lung images. Dalian University of Technology (2019). DOI:https://doi.org/10.26991/d.cnki.gdllu.2019.003229
Yue, L., Ma, L., Wei, B.: Research on children Pneumonia Clinical Syndrome Classification model based on decision tree algorithm. Electronic Test 05, 243–244 (2013)
Jun, S., Park, B., Seo, J.B., Lee, S., Kim, N.: Development of a computer-aided differential diagnosis system to distinguish between usual interstitial pneumonia and non-specific interstitial pneumonia using texture- and shape-based hierarchical classifiers on HRCT images. J. Digital Imaging, 31(2) (2018)
Hearst, M.A., Dumais, S.T.: Support vector machines. IEEE Intell. Syst. Appli. 13(4) (1998)
Sun, Z., Xue, L., Xu, Y., et al.: Overview of deep learning. Appli. Res. Comput. 29(08), 2806–2810 (2012)
Zhou, F., Jin, L., Dong, J.: A review of convolutional neural networks. Chin. J. Comput. 40(06), 1229–1251 (2017)
Zhou, Q., Zhang, J., Pu, Z., et al.: COVID-19 CT image classification method combined with deep layer dense aggregation. Appli. Res. Comput., 1–8 (2023).https://doi.org/10.19734/j.issn.1001-3695.2022.08.0502
Guo, K., Du, Q.C., Wu, A., et al.: COVID-19 recognition technology based on lightweight neural network. Chin. J. Med. Phys. 39(10), 1263–1269 (2022)
Ummah, K.R., et al.: Effect of image pre-processing method on convolutional neural network classification of COVID-19 CT scan images. Inter. J. Innov. Comput. Inform. Control 18(6), 1895–1912 (2022) https://doi.org/10.24507/ijicic.18.06.1895
Zhang, N., Wu, H., Han, X., et al.: Tomato disease recognition method based on multi-scale and attention mechanism. J. Zhejiang Agricul. 33(07), 1329–1338 (2021)
Lin, C., Zhang, G., Yang, J., et al.: Ttransfer learning based recognition for forestry business images. J. Nanjing Forestry Univ. (Nat. Sci.), 44(04), 215–221 (2020)
Li, F.-F., Jia, D., Kai, L.: ImageNet: Constructing a large-scale image database. J. Vis. 9(8) (2010)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2024 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Zhang, J., Zhang, B. (2024). Auxiliary Diagnosis of Pneumonia Based on Convolutional Attention and Parameter Migration. In: Sun, Y., Lu, T., Wang, T., Fan, H., Liu, D., Du, B. (eds) Computer Supported Cooperative Work and Social Computing. ChineseCSCW 2023. Communications in Computer and Information Science, vol 2012. Springer, Singapore. https://doi.org/10.1007/978-981-99-9637-7_37
Download citation
DOI: https://doi.org/10.1007/978-981-99-9637-7_37
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-99-9636-0
Online ISBN: 978-981-99-9637-7
eBook Packages: Computer ScienceComputer Science (R0)