Abstract
Antimicrobial resistance (AMR) and the emergence of multidrug-resistant organisms are one of the most eminent impediments of recent times. The intrinsic resistance mechanisms of the pathogenic organisms, especially the efflux pumps, play a significant part in the development of AMR and have to date resulted in the failure of a wide range of antimicrobial drugs. The efflux mechanisms exude the drugs from the cells and prevent their accumulation, thereby rendering the drug futile against the pathogenic microbe. Therefore, in order to combat with these evolving MDR microorganisms, their intrinsic resistance processes like the efflux pumps need to be targeted. In this regard, the advent of phytotherapeutic molecules isolated from specific medicinal plants has shown promising potential as efflux pump inhibitors (EPIs) with their capability of modifying the resistance mechanisms of pathogens. These botanicals show structural similarity or mimic certain peptide structure that increases their binding affinity to the efflux pump proteins. As a result, either the EPI binds to the drug binding pocket or the configuration of the efflux pumps is altered as a result of the binding, thus preventing the drug efflux. This review primarily focuses on the mode of action of the microbial efflux pumps, their contribution to enhancing microbial resistance, the emerging role of phytotherapeutics, and their potential to act as efflux pump inhibitors (EPIs).
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Antimicrobial resistance (AMR)
- Multidrug resistance
- Efflux pumps
- Efflux pump inhibitors
- Phytotherapeutics
- Medicinal plants
4.1 Introduction
One of the most significant obstacles faced by the contemporary world of research is the aspect of antimicrobial resistance (AMR). The vast range of infections caused by microorganisms such as bacteria, fungi, viruses, and parasites, no longer remain susceptible to the standardized therapeutics developed against them, primarily owing to AMR (Abushaheen et al. 2020). The evolving capacity of these microorganisms inculcates them with the power to develop resistance mechanisms, thereby allowing them to easily bypass the action of the applied drugs and thrive in their very presence. Resistant organisms therefore have the power to wreak havoc in the host system without getting affected by any of the administered drugs (Hughes and Andersson 2017). Furthermore, the rapid development of resistant mechanisms of microbes has almost rendered them imperishable, and this has turned out to be a major challenge in the times of modern research. The advent of microbial resistance to multiple drugs also known as multidrug resistance (MDR) has increased the threat by multiple folds (Piddock 2006a). The mechanisms via which the organisms develop resistance can be broadly distributed into three categories, namely, adaptive resistance, acquired resistance, and intrinsic resistance (Kabra et al. 2019). Gradual exposure to increasing doses of drugs often results in an onset of sudden inception of resistance in certain specific microorganisms that is to some extent associated with the diverseness of patterns of gene expression and might require phenotypic and epigenetic inheritance (Kabra et al. 2019). This form of AMR is known as adaptive resistance. On the contrary, some microorganisms develop acquired resistance primarily because of resistant specific horizontal gene transfer or due to some form of genetic mutation. Intrinsic resistance, however, is mainly associated with the internal mechanisms of microorganisms, which are predominantly concerned with reducing the uptake of drugs by the cell with the help of efflux pumps or with decreasing the cellular concentration of the drugs that have already been taken up by the cell (Reygaert et al. 2018). Therefore, with respect to AMR, the only probable option to deter the threat posed by the resistant superbugs is to develop therapeutic strategies that target the very core of resistant mechanisms in the microbial cells (Motta et al. 2015). Targeting the resistant developing mechanisms in the microbes would provide an opportunity to increase the susceptibility of the organisms to a wide range of drugs (Golparian et al. 2014). The aspect of targeting the AMR mechanisms mainly efflux pumps with medicinal plant extracts has come to the forefront and is being considered a prospective option of therapy (Li and Nikaido 2009; Poole 2001). This study mainly deals with the efflux pump mechanisms highlighting their association with the development of antimicrobial resistance, the advent of phytotherapeutics, and its role in efflux pump inhibition.
4.2 Efflux Pumps: Antimicrobial Resistance Mechanisms
Even after 94 years of the advent of penicillin, the aspect of microbial infections and manifested diseases still remains an unfinished battle (Poole 2001). At present, although an arsenal of treatment options is available for these infections, the microbes stand undefeated powered by their developed AMR (Prestinaci et al. 2015). One of the strongest components of the intrinsic resistance mechanisms is the efflux pumps (Poole 2007). It was the last AMR mechanism discovered and was first observed in Escherichia coli, where it provided resistance against tetracycline. According to the initial discovery of efflux pumps related to Tetracycline, four different plasmids, namely, R144, RP1, RA1, and R222, coded for tetracycline-related efflux proteins in several strains of E. coli and these efflux machineries were significantly responsible for reducing the cellular accumulation of tetracycline in the E. coli cells (Poole 2007). In the later years, a number of essential efflux mechanisms have come to the forefront, which are either chromosome or plasmid-encoded, and have been categorized into various classes on the basis of their protein family as well as on the basis of the antimicrobial resistance features shown by them (Lomovskaya and Watkins 2001). Structurally, efflux pumps can be predominantly defined as transmembrane proteins, whose presence is mostly ubiquitous in all living organisms. Functionally, they have been majorly divided into six protein families (Borges-Walmsley et al. 2003), namely, Major Facilitator Superfamily (MFS), Resistance Nodulation Division (RND) family, Small Multidrug Resistant (SMR) Superfamily, Proteobacterial Antimicrobial Compound Efflux (PACE) Superfamily, Multidrug and Toxic Compound Extrusion (MATE) Superfamily, and, finally, the ATP Binding Cassette (ABC) Superfamily (Du et al. 2018; Eswaran et al. 2004). For most of these efflux pumps, viz., MFS, RND, SMR, MATE, PACE, etc., the source of energy is either the proton motive force (PMF) or the sodium ion gradient, and therefore, they primarily function as secondary active transporters (Marquez 2005; Nikaido 2011). However, for ABC family proteins, the energy source is ATP, and therefore they act as the primary active transporter (Hollenstein et al. 2007). Each of these efflux pumps has separate mechanisms of action, thereby providing resistance against a wide range of microbial infections.
4.3 Small Multidrug Resistant (SMR) Superfamily
4.3.1 Structure
The SMR efflux family is mainly driven by the energy provided by the proton motive force (PMF) (Paulsen et al. 1996). They are hydrophobic by nature and are primarily lipophilic cations that render their range of molecular recognition significantly lower, compared with other efflux pumps (Bay and Turner 2009). Structurally SMR family proteins are homotetramers, being constituted of four transmembrane segments (TMS) (Poulsen et al. 2009). SMR pump genes have been reported to be expressed in the chromosomal DNA and in certain cases on transposable elements and plasmids as well (Bay and Turner 2009). Although the substrate range of SMR efflux pumps is considerably lower than others, it confers resistance to several drugs that are mostly β-lactams and aminoglycosides in certain cases (Kornelsen and Kumar 2021). For example, the SMR pump of Escherichia coli (EmrE pump) reportedly transports drugs such as erythromycin, tetracycline, and vancomycin, thereby rendering the organism invincible against these drugs (Bay et al. 2008; Yerushalmi et al. 1996). On the contrary, Staphylococcus epidermidis has also shown resistance provided by SMR pumps that have effectively transported tetracycline, erythromycin, as well as ampicillin (Blair et al. 2014; Kumar and Schweizer 2005).
4.3.2 Mechanism
The SMR pumps comprise oligomeric functional complexes and a hydrophobic core. The pump pathway lying in the transmembrane region, through which the toxic molecules and drugs are effluxed out of the cell, comprises the polar countenance of the amphipathic membrane molecules, along with the residues of serine, tyrosine, tryptophan, as well as glutamate (Rath et al. 2006). The substrates generally bind to the transmembrane during the process of efflux with the help of the conserved glutamate residues, being specifically cationic in nature (Kolbusz et al. 2010). Moreover, as SMRs are associated with the efflux of multiple drugs and hence the name, the versatility and pliability are achieved owing to the presence of conserved sequences of proline and glycine, the active sites of which allow the conformational affability (Jack et al. 2000). Initially, the drug interacts with the transmembrane glutamate residues of the SMR efflux pumps, followed by the exchange of the drug cation with a proton. It then forms a complex with the carrier moiety via a carboxyl group and is regarded as the drug carrier complex, which crosses the hydrophobic internal region of the membrane of the cell. The translocation across the membrane occurs on the basis of several intricate reversible alterations in the protein structure. Following this step, protons acquired from the medium come in contact with the drug carrier complex and replace the drug with the help of competitive binding. Therefore, the drug is released and the efflux proteins are once again complexed with the proton, which results in the reversal of the protein’s confirmation, bringing it back to its original form (Spengler et al. 2017).
4.4 Proteobacterial Antimicrobial Compound Efflux (PACE) Superfamily
4.4.1 Structure
The PACE superfamily is so named owing to the presence of these pump proteins in a wide range of proteobacteria. These pumps have four transmembrane segments and comprise alpha helices (Zgurskaya and Introduction 2021). In PACE pumps, four amino acid residues are universally conserved. They are, namely, a residue of glutamic acid that is present within the first transmembrane segment, a residue of asparagine present in the second transmembrane segment, a residue of alanine present within the periplasmic space of the fourth transmembrane section, and, finally, a residue of aspartic acid, present in the boundary of the cytoplasmic membrane of the fourth transmembrane segment (Ahmad et al. 2018). These pumps are mainly responsible for the efflux of drugs such as chlorhexidine, benzalkonium, acriflavine, dequalinium, proflavine, and so on (Spengler et al. 2017).
4.4.2 Mechanism
Proteobacterial antimicrobial compound efflux superfamily proteins are mainly transporters of biocides such as acriflavine and chlorhexidine. The Acinetobacter chlorhexidine efflux pumps (AceI) are one of the most significant prototypes of PACE efflux pumps (Abdi et al. 2020). The AceI proteins primarily use the electrochemical proton gradient as the prominent source of energy, utilizing which they transport the substrates. AceI pumps are to some extent similar to the SMR family efflux pumps, as they are both small in size and share a similar secondary structure; however, they do not have any common functional attributes. The mechanism of transport mainly comprises the recognition of the substrates with the help of the glutamic acid residue lying in the core of the first transmembrane segment (Reddy Bolla et al. 2020). Post recognition, the substrate forms a complex with the efflux protein machinery, thereby resulting in a conformational change. Finally, translocation takes place and the drug molecules are released outside the cell (Hassan et al. 2019). The intricate details of the mechanism of action of the PACE efflux pumps other than AceI have not yet been thoroughly researched to date.
4.5 Major Facilitator Superfamily (MFS)
4.5.1 Structure
The MFS efflux pumps function on the basis of the energy derived from the transport of NA+ or H+ ions. The MFS fold can be defined as a canonical section comprising 12 transmembrane segments, which can be further subdivided into two domains, namely, the N-terminal domain and the C-terminal domain. Each of the two domains consists of six transmembrane segments, which are interconnected with the help of a cytoplasmic loop and might also sometimes bear the presence of a helix that is amphipathic by nature (Varela and Mukherjee n.d.). These pumps are mainly associated with the transport of metabolites, drugs, anions, and so on. They have the capacity to efflux out drugs such as tetracycline, macrolides, and so on, thereby rendering them inactive (Kumar et al. 2020). These pumps show a great amount of variation with respect to their substrates. For example, organisms like Acinetobacter baumannii express distinctly separate MFS efflux pumps, i.e., the SmvA pump specific for erythromycin and the CraA pump for chloramphenicol (Pasqua et al. 2019). E. coli also expresses separate MFS pumps such as MefB, QepA, and Fsr; MefB shows affinity toward macrolides, Qep toward fluoroquinolones, and Fsr toward trimethoprim (Kumar et al. 2013).
4.5.2 Mechanism
The MFS superfamily proteins are ubiquitously present and are susceptible to a wide range of molecules or substrates such as oligosaccharides, small metabolites, amino acids, ions, as well as several other antimicrobial molecules, including a variety of drugs (Kumar et al. 2013). The structure and organization of the MFS efflux pumps are more or less reserved and are termed the MFS fold. The mechanism of transport is carried out by continuous changing of the binding site of the substrate between the extracellular side of the cell membrane and the intracellular or periplasmic side of the cell membrane (Ranaweera et al. 2015). The models developed to demonstrate the mode of action of the MFS efflux pumps are termed the “clamp switch model” and the “‘rocker switch model.” According to these models, the metamorphosis between different states includes both structural modifications and rotations of fixed and rigid sections. The fifth transmembrane segment is technically responsible for the transition between the states of inwardly open to states of outwardly open, along with the reassembly of the hydrophobic centers present in the N terminal domain and the C terminal domain’s cytoplasmic loop, in order to render the cytoplasmic entrance as closed (Du et al. 2018).
4.6 Multidrug and Toxic Compound Extrusion (MATE) Superfamily
4.6.1 Structure
The MATE family efflux pumps are mostly related to the efflux of fluoroquinolones. However, some MATE pumps are also known to be associated with aminoglycosides (Piddock 2006b). Structurally, it is made up of 12 transmembrane segments or helices, along with the amino acid residues existing within a range of 400–700 (Radchenko et al. 2015). Most of the MATE pumps that are found in bacteria have been observed in Gram-negative organisms. One of the first MATE pumps was observed in the organism Vibrio parahaemolyticus, and the pump was named NorM. Other organisms such as Neisseria meningitidis and Neisseria gonorrhoeae also express the NorM MATE efflux pump (Rouquette-Loughlin et al. 2003).
4.6.2 Mechanism
MATE efflux pumps function as proton antiporters, sodium antiporters, and drug antiporters, being present in archaea, bacteria, as well as eukaryotic cells. Each MATE transporter protein comprises two domains, the N- and C-terminal domains (Sun et al. 2014). These domains are so placed inside the MATE transporters, that they form a central cavity that is shaped like a V. The lower section of this cavity ends halfway in the bilayer of the cellular membrane, therefore making the structure acquire an outwardly open stable state. When the efflux mechanism of the substrate is initiated, the substrate and the cation enter into a competition as both of them target to interact with the same groups of amino acids. In most cases, the drug-binding sites of the MATE efflux proteins are the same as the cation-binding sites; however, separate drug-affiliated pockets are also present in certain MATE efflux pumps, like NorM, where both the cation and the substrate have the capacity to bind simultaneously to the protein owing to the presence of distinct and separate binding sites. After drug binding is achieved, the proton undergoes a conformational change in order to switch between the inwardly open state and the outwardly open state, thereby resulting in the efflux of the drug. The MATE family recognizes a vast range of drugs, viz., kanamycin, cimetidine, ampicillin, ethidium, norfloxacin, triethylammonium, chloramphenicol, metformin, ciprofloxacin, and so on, thereby proving protection against the microorganisms against such potent antimicrobial agents (Du et al. 2018; Kuroda and Tsuchiya 2009).
4.7 Resistance Nodulation Division (RND) Family
4.7.1 Structure
RND efflux pumps are mainly localized in Gram-negative bacteria. They are responsible for the efflux of a variety of compounds such as detergents, heavy metals, yes, antibiotics, and so on and thus functionally diverse. Some RND pumps such as the Tet and Mef pumps are substrate-specific and are only associated with the transport of tetracycline and macrolides, respectively (Blair et al. 2015). On the contrary, RND pumps like MexAB-OprM found in Pseudomonas aeruginosa provide protection to the organism against a wide variety of substances ranging from beta-lactams, tetracycline, chloramphenicol, sulfamethoxazole, and trimethoprim to fluoroquinolones as well as (Kourtesi et al. 2013) eighth transmembrane segments.
4.7.2 Mechanism
The RND Superfamily of efflux pumps also functions as antiporters of proton and/or drugs and are found in bacteria, eukaryotes, and archaea, although RND pumps are primarily present in Gram-negative bacteria (Routh et al. 2011). With respect to AcrB, an established prototype of RND efflux pumps, the structure of RND proteins constitutes of three states that are not equivalent to each other, namely, the loose (L) state or access state, the tight (T) state or bonding state, and the open (O) state or extrusion state. In other words, the RND transporters show stable interchangeable and reversible shifts between three stages—L, T, and, O, that is loose (access), tight (bonding), and open (extrusion) states, respectively. According to this model, the drug enters the binding site or drug pocket that usually lies in the periplasmic domain, when the RND transporter molecules are in the loose or access state (Du et al. 2018). Post entry, the drug moves ahead in the drug-associated pocket, and then the transporter shifts its state from access or loose to binding or tight state, thereby allowing the drug to bind to the transporter protein and form a complex. Post binding, the drug is further exuded into the docking domain of the extrusion or open state (Shafer et al. 1998). The side chains present in the transmembrane segments of the RND transporter proteins undergo protonation, which in turn allows the change in the states to take place (Fernando and Kumar 2013).
4.8 ATP-Binding Cassette (ABC) Superfamily
4.8.1 Structure
The efflux pumps of the ABC superfamily obtain their energy from the hydrolysis of ATP. They have a wide range of substrates starting from ions, amino acids, proteins, and polysaccharides to drugs (Davidson and Chen 2004). Structurally, these pumps comprise six transmembrane segments, each made up of alpha helices. They generally are active when present in pairs, either in the homodimeric state or in the heterodimeric state (Cole and Deeley 1998). They generally show concomitance with cytoplasmic ATPases. For example, the VcaM efflux pump present in Vibrio cholerae is a noteworthy ABC family efflux pump and is responsible for the efflux of drugs like tetracycline and fluoroquinolones (Blair et al. 2015; Davidson and Chen 2004).
4.8.2 Mechanism
ABC efflux pumps of pathogenic microorganisms are one of the principal contributors of intrinsic antimicrobial resistance. The range of molecules exuded by the ABC cascade is extremely vast and huge, and these proteins have therefore been termed orphan proteins. The ABC transporters are constituted of two separate domains, one is the transmembrane domains containing binding sites for substrates, and the other is the nucleotide-binding domain, where, upon binding of ATP, it is hydrolyzed, thereby providing the energy required to initiate and complete the process of transport (Prasad and Goffeau 2012). ABC transporters show two forms, the homodimeric and heterodimeric forms. The homodimeric form falls under intrinsic resistance and possesses a separate binding site for substrate and ATP, thereby allowing ATP hydrolysis to fuel the process of drug exudation. However, the heterodimeric form is related to acquired AMR and often possesses a degenerative site that does not allow hydrolysis of ATP. With respect to the homodimeric form, it too shows the reversible changes and alterations between two states, that is, the switch between an occluded state that is inward open and the outward open state, in order to ensure the translocation of substrates across the bilayer of the cell membrane, and this process is referred to as the process of alternating access mechanism. In this regard, another crucial factor is the changes taking place in the nucleotide-binding domain. The structural and conformational changes associated with the ABC transporters are directly linked with the dissociation as well as dimerization of the nucleotide-binding domain, owing to the binding and hydrolysis of ATP. The change between these two states is also facilitated by the presence of certain enzymes like flippase that facilitate the shift of lipid precursors from the cytoplasmic leaflet associated with the inner membrane to the periplasmic leaflet (Du et al. 2018).
4.9 Challenges of Targeting Efflux Pumps
The aspect of targeting efflux pumps was considered a point of significant concern till a few years back, prior to the advent of phytotherapeutics, primarily due to the wide varieties in the structure and function of the efflux machineries, that make them immensely difficult drug targets. Apart from that, several key factors need to be taken into account while designing drugs against efflux pumps, the most essential one being, the efflux pump inhibitors (EPIs) which should either mimic the substrate structure or should be closely related to the substrate configuration of that respective efflux pump. Only then, it would be possible for the molecules to interfere with the functioning mechanisms of the pumps, thereby preventing drug efflux in the long run. Other processes of targeting the efflux proteins also exist, such as targeting the glutamate residues in certain efflux proteins (Dashtbani-Roozbehani and Brown 2021). This might result in effective modification of the binding site of the transporter moiety, thereby resulting in the loss of efflux function. Therefore, the specificity of the drugs is immensely necessary. However, several challenges arise when efflux pumps of the pathogen are being targeted as a potential option for therapy in order to increase the susceptibility of the microbial cells to the pre-administered drugs (Verma et al. 2021). One of the most significant hindrances can be the ubiquitous presence of efflux pumps. Efflux pump mechanisms are required in all living organisms, in order to maintain the functional stability of the cellular metabolism. Hence, efflux pumps of the same family might be present in both the host and the pathogen. Although the administration of established drugs targeting the pathogenic cells would not pose a problem, the administration of inhibitors targeting the efflux mechanisms might give rise to a major challenge (Pagès and Amaral 2009). The EPIs showing affinity toward both the host and pathogen efflux machineries will inhibit the host efflux functions as well, ultimately resulting in toxic and detrimental outcomes.
4.10 Emergence of Phytotherapeutics Against AMR: Its Potential as a Therapeutic Option
The development of MDR in microbes owing to the significant efflux of drugs from the pathogens is a cause of growing concern, and the formation of effective EPIs is of utmost importance (Pagès and Amaral 2009). Moreover, treatment with antibiotics has taken a backseat and the symptoms and manifestations of the microbial diseases have taken a turn toward fatal outcomes. Therefore, the emergence of phytotherapeutics has acted as a ray of hope for researchers in recent times. Efflux pump inhibitors (EPIs) from plant sources have reinstated the hope of vanquishing MDR organisms to a great extent (Sana et al. 2015). The primary reason why medicinal plants can act as propitious sources of efflux pump inhibitors is the abundance of secondary metabolites in these plants that are diversified both structurally and chemically and are aided with several known medicinal properties. In recent years, numerous research works have been carried out on these medicinal plants and the efficacy of their extracts to act as potent EPIs (Seukep et al. 2020). One significant obstacle is that the activity of these efflux pumps is often diverse and multiple molecular mechanisms are associated with them, because of which it becomes difficult to target these ubiquitous machineries in the microorganisms. Certain plant extracts possess molecules that have the ability to block and stop the functioning of efflux pumps of microorganisms, especially Gram-positive and Gram-negative bacteria (Fig. 4.1). This action of the plant-derived EPIs reduces the resistance of the pathogenic organisms against the proposed therapeutics and in a way re-establishes the efficiency of the initial treatment methods. Accumulation of the drugs inside the microbial cells is quite easily possible, thereby allowing the drug to exhibit its true antimicrobial effect. The plant families that have been thoroughly researched with respect to the presence of EPs or EPI-associated molecules are, namely, Cucurbitaceae, Berberidaceae, Zingiberaceae, Lamiaceae, Apocynaceae, Fabaceae, Convolvulaceae, and so on (Stavri et al. 2007). The main reason behind zoning in upon the above-mentioned plant families is because of the fact that our daily dietary elements come from plants belonging to these families. In fact, potent EPIs from plant sources were identified from simple plant products of our diet, such as tea leaves, lemon grass, pepper, grapes, pomegranates, and so on. Some of the isolated compounds from these sources are geraniol, theobromine, piperine, resveratrol, farnesol, and so on, and their potentiality has been observed in a wide variety of pathogens, including Gram-positive and Gram-negative bacteria.
The types of efflux pumps present in Gram-positive and Gram-negative bacteria. (a) Different types of efflux pumps including SMR, MATE, ABC transporters, and MFS present in Gram-positive organisms, and the drugs against which they provide resistance. (b) The types of efflux pumps including RND, ABC transporters, MFS present in Gram-negative organisms, and the drugs against which they provide resistance
One of the earliest discoveries of phytotherapeutic molecules against bacterial efflux pumps identified and analyzed a cluster of molecules that mimic the peptide structure and proved that these molecules were highly effective against certain strains of P. aeruginosa that show heightened expression of efflux pumps like MexAB (Pagès and Amaral 2009; Stavri et al. 2007). The primary compound that showed the closest structural similarity with the peptide structure was extensively studied, and its associated and derivative molecules were also taken into account, in order to ascertain the effective relationship between conformation and function. It was also shown that these compounds have the ability to act against the efflux of fluoroquinolone, the drug levofloxacin playing the role of a marker. After analyzing the group of peptide analogs, the most significant compound playing the lead role was deduced to be Phe-Arg-β-naphthylamine or PAβN (Thomas et al. 1999). The primary activity profile of PAβN was found to be the capability to inhibit the function of the efflux pumps that majorly efflux out quinolones, in this case, fluoroquinolones like levofloxacin.
The one fact that made the discovery of PAβN all the more interesting was that not only did it structurally mimic the substrate of the MexB efflux pump but was also found to affect the activity of efflux mechanisms of Klebsiella pneumoniae, E. coli, Salmonella enterica, Enterobacter aerogenes, and so on (Lomovskaya et al. 2001). The mechanism of action of PAβN indicates the fact that it shows prominent competition with the antibiotic substrates with respect to binding with the efflux proteins. As a result, PAβN binds with the efflux proteins of pumps like MexB, thereby allowing the drug to be retained within the cell and consequently resulting in an increase of cellular drug concentration, ultimately reaching the state required for the drug to exhibit its action on its respective target within the cell. Apart from this, PAβN also shows the capacity to either reduce the resistance of the microorganism toward that particular drug or permanently reverse the resistance mechanism of the microbe, thereby indicating the emergence of a long-term solution against AMR (Murakami et al. 2006). However, the functional activity of PAβN has certain limitations as well, giving rise to specific drawbacks. Although PAβN shows structural similarity with some specific antibiotics, its range leaves out several important ones, and as a result of this, it shows partial inhibition with respect to certain efflux pumps. Thus, the relationship between PAβN and antibiotics can be regarded as a disparate one, the eminent regulatory factors being the presence of the substrate or drug binding domain with affinity toward PaβN and the structural characteristics of the efflux pumps. Another disadvantage regarding PAβN might be its lack of affinity toward the antibiotic-specific site of the drug-binding domain of the efflux pumps. If PAβN binds to any site apart from the drug binding site, it might not be able to show the inhibitory actions as mentioned before. Although alteration of the structural and functional properties of the efflux pumps can be initiated in this regard, complete inhibition cannot be guaranteed. In order to overcome these limitations, several strategic schemes were considered and the synthesis of more potent derivatives of PAβN turned out to be the most effective option. The synthesis of derivative molecules of PAβN therefore led to the stability of the EPI functions and also contributed to establishing phytotherapeutics as a promising solution. The derived molecule was named MC-04,124, which showed heightened stability and features exhibiting a reduced range of toxicity and side effects. Moreover, compared with PAβN, MC-04,124 also showed greater stability in the biological system fluids (Watkins et al. 2003). To date, several derivatives of PAβN have been synthesized, and all of them have shown gradual refinement in their pharmacokinetic behavior as efflux pump inhibitors.
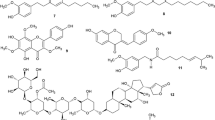
Although EPIs from natural sources working on the majority of the microbial population have been identified, the number of these molecules effectively showing their inhibitory activities against Gram-negative bacteria is relatively on the lower scale. The intricate and complex arrangement of the efflux pumps in Gram-negative organisms that is the tripartite pump system can be considered responsible for the lower range of availability of EPIs against them. Despite the daunting tripartite structure of these pumps in Gram-negative organisms, several molecules have been discovered from plant sources that effectively target these pumps (Prasch and Bucar 2015). For example, karavilagenin isolated from Momordica balsamnia and gallotannin isolated from Terminalia chebula have been reported to show their inhibitory action against the ArcAB-TolC tripartite efflux pump present in E. coli strains. Furthermore, MexAB-OprM tripartite pumps in P. aeruginosa have been reported to be inhibited by compounds such as falcarindiol, palmatine, berberine, conessine, catharanthine, osthol, curcumin, and resveratrol isolated from Levisticum officinale, Berberis vulgaris, Holarrhena antidysenterica, Catharanthus roseus, Cnidii monnieri, Curcuma longa, and Nauclea pobeguinii, respectively (Fig. 4.2).
Phytotherapeutic botanicals isolated from various plant sources inhibiting drug efflux by Eps such as TetK, MdeA, MsrA, NorA, LmrS, Bmr MDR, and so on
On the contrary, theobromine isolated from Theobroma cacao works equally effectively against both ArcAB-TolC as well as MexAB-OprM tripartite efflux pumps in Gram-negative organisms (Seukep et al. 2020) (Table 4.1).
4.11 Strategies to Overcome Intrinsic Resistance of Efflux Pumps Using Phytotherapeutics
The rapid surge in the number of multiple drug-resistant microbes urgently calls for the initiation of an effective solution. Emergence of the phytological molecules and their efflux pump inhibitory capabilities have opened up a new field of research, wherein these molecules can be considered potential drug candidates. A prime example of plant-derived EPI is 5-MHC or 5′-methoxyhydnocarpin isolated from plants such as Hydnocarpus wightianus and Berberis sp.; this chemical compound has reinstated the function of the antimicrobial berberine. In other words, 5-MHC promotes the accumulation of berberine in the pathogens, thereby allowing it to demonstrate its antimicrobial properties after reaching the desired concentration. This clearly demonstrates that berberine can effectively reverse the berberine resistance developed by pathogenic microorganisms. 5-MHC, however, does not exhibit any form of antimicrobial property by itself (Stavri et al. 2007). Therefore, in order to achieve reversal of resistance toward eminent drugs amongst the pathogenic microflora, strategies need to be developed, wherein, with the use of phytotherapeutic molecules, the intrinsic resistance mechanisms like efflux pumps of microbes can be targeted.
In general, the efflux mechanisms in pathogens can be inhibited via the following strategies:
-
1.
Inducing suppression of the genes that are responsible for the structural and functional expression of the efflux pumps
-
2.
Preventing appropriate assembly of the efflux proteins in the pumps, thereby restricting the development of efflux pump structure, making it functionally inactive
-
3.
Blocking the outer and/or inner membrane channel openings, in order to prevent the efflux of drugs
-
4.
Using antiporter directly or indirectly to prevent the pumps from using the proton motive force, thereby resulting in their collapse by cutting off the energy source
-
5.
Introducing certain chemical molecules that can exhibit either competitive or non-competitive inhibition with respect to the administered drug
-
6.
Altering the structural design of the drugs used for treatment, so as to reduce its binding affinity with the efflux pump proteins (Pagès and Amaral 2009)
Of the above-mentioned strategies, plant-derived EPIs can be used to effectively demonstrate at least two of them (Fig. 4.3). Most EPIs generally mimic the drug or substrate structure, because of which their affinity toward the efflux proteins is high. As a result, they can show significant competition with the drug, therefore, getting exuded itself, in place of the administered drug and allowing it to get accumulated within the cell. Once the drug is allowed to reach the desired concentration within the pathogen, it shows its antimicrobial activity. Thus, plant-derived EPIs can execute competitive inhibition, thereby reducing the drug-resistant capacity of pathogenic organisms (Pulingam et al. 2022). On the contrary, EPIs can also show non-competitive binding. In this case, the EPI binds to a site other than the drug binding site, and as a result of this binding, the EP protein undergoes a significant conformational change, thereby rendering the efflux pumps incapable of transporting the drugs outside the cell. Another strategy that the plant-derived chemical compounds can execute is the aspect of blocking the entry and exit channels of the efflux pumps, thus preventing the drug from getting exuded out of the cells.
Efflux pump inhibition strategies. (a) Competitive inhibition: the mode of action of efflux pumps, exuding the drug from the cells, thereby enhancing the antimicrobial resistance properties of the pathogenic microbes. (b) Competitive binding of efflux pump inhibitors (EPIs) and exclusion of EPIs from the cell in place of the drugs, resulting in the retention and accumulation of the drug within the cell. (c) Blocking the EP channels: Drug is retained within the pathogenic cell by blocking the entry and exit channels of the efflux pumps by the EPIs
Therefore, keeping these strategies in mind, chemical compounds from plant sources can be used to design accessory drugs, that would reverse, remove, or reduce the efflux pump-induced intrinsic resistance mechanisms of the pathogenic microflora and improve the efficacy of the administered drugs (Pulingam et al. 2022; Annunziato 2019).
4.12 Conclusion
Efflux pump inhibitors are therefore one of the easily accessible options via which the action of a wide range of antimicrobials can be revived. The application of EPIs along with these antimicrobials as accessory drugs might be the solution that the research world is waiting for in order to overcome hurdles such as MDR and antimicrobial resistance. Moreover, the use of botanicals as EPIs has several advantages. Being derived from medicinal plant sources, these phytotherapeutic molecules rarely show any adverse harmful side effects on hosts and do not have any long-term aftereffects as well (Prasch and Bucar 2015). These molecules have shown a very high percentage of efficiency in this field, and further investigation would definitely enlighten us further in this regard. To conclude, the inception of phytotherapeutics as EPIs might mark the end of the reign of antimicrobial resistance to pathogenic microorganisms.
Abbreviations
- ABC:
-
ATP-binding cassette
- AceI:
-
Acinetobacter chlorhexidine efflux I
- AMR:
-
Antimicrobial resistance
- ATP:
-
Adenosine triphosphate
- EmrE:
-
Efflux multidrug resistance E
- EP:
-
Efflux pump
- EPI:
-
Efflux pump inhibitor
- MATE:
-
Multidrug and toxic compound extrusion
- MDR:
-
Multidrug resistant
- MFS:
-
Major facilitator superfamily
- MHC:
-
Methoxyhydnocarpin
- MsrA:
-
Methionine sulfoxide reductase A
- PACE:
-
Proteobacterial antimicrobial compound efflux
- PAβN:
-
Phe-Arg-β-naphthylamine
- PMF:
-
Proton motive force
- RND:
-
Resistance nodulation division
- SMR:
-
Small multidrug resistant
- TMS:
-
Transmembrane segments
References
Abdi SN et al (2020) Acinetobacter baumannii efflux pumps and antibiotic resistance. Infect Drug Resist 13:423–434. https://doi.org/10.2147/IDR.S228089
Abushaheen MA et al (2020) Antimicrobial resistance, mechanisms and its clinical significance. Dis Mon 66:100971
Ahmad I et al (2018) Bacterial multidrug efflux proteins: a major mechanism of antimicrobial resistance. Curr Drug Targets 20:16–28
Annunziato G (2019) Strategies to overcome antimicrobial resistance (AMR) making use of non-essential target inhibitors: a review. Int J Mol Sci 20:5844
Bay DC, Rommens KL, Turner RJ (2008) Small multidrug resistance proteins: a multidrug transporter family that continues to grow. Biochim Biophys Acta Biomem 1778:1814–1838. https://doi.org/10.1016/j.bbamem.2007.08.015
Bay DC, Turner RJ (2009) Diversity and evolution of the small multidrug resistance protein family. BMC Evol Biol 9:140
Blair JMA, Richmond GE, Piddock LJV (2014) Multidrug efflux pumps in Gram-negative bacteria and their role in antibiotic resistance. Future Microbiol 9:1165–1177. https://doi.org/10.2217/FMB.14.66
Blair JMA, Webber MA, Baylay AJ, Ogbolu DO, Piddock LJV (2015) Molecular mechanisms of antibiotic resistance. Nat Rev Microbiol 13:42–51. https://doi.org/10.1038/nrmicro3380
Borges-Walmsley I, Mckeegan KS, Walmsley AR (2003) Structure and function of efflux pumps that confer resistance to drugs. Biochem J 376:313
Cole SPC, Deeley RG (1998) Multidrug resistance mediated by the ATP-binding cassette transporter protein MRP. BioEssays 20:931
Dashtbani-Roozbehani A, Brown MH (2021) Efflux pump mediated antimicrobial resistance by staphylococci in health-related environments: challenges and the quest for inhibition. Antibiotics 10:1502. https://doi.org/10.3390/antibiotics10121502
Davidson AL, Chen J (2004) ATP-binding cassette transporters in bacteria. Ann Rev Biochem 73:241–268. https://doi.org/10.1146/annurev.biochem.73.011303.073626
Du D et al (2018) Multidrug efflux pumps: structure, function and regulation. Nat Rev Microbiol 16:523–539. https://doi.org/10.1038/s41579-018-0048-6
Dwivedi GR et al (2018) Antibiotics potentiating potential of catharanthine against superbug Pseudomonas aeruginosa. J Biomol Struct Dyn 36:4270–4284
Eswaran J, Koronakis E, Higgins MK, Hughes C, Koronakis V (2004) Three’s company: component structures bring a closer view of tripartite drug efflux pumps. Curr Opin Struct Biol 14:741–747. https://doi.org/10.1016/j.sbi.2004.10.003
Fernando DM, Kumar A (2013) Resistance-nodulation-division multidrug efflux pumps in gram-negative bacteria: role in virulence. Antibiotics 2:163–181. https://doi.org/10.3390/antibiotics2010163
Fiamegos YC et al (2011) Antimicrobial and efflux pump inhibitory activity of caffeoylquinic acids from Artemisia absinthium against gram-positive pathogenic bacteria. PLoS One 6:e18127
Garvey MI, Rahman MM, Gibbons S, Piddock LJV (2011) Medicinal plant extracts with efflux inhibitory activity against gram-negative bacteria. Int J Antimicrob Agents 37:145–151
Golparian D, Shafer WM, Ohnishi M, Unemo M (2014) Importance of multidrug efflux pumps in the antimicrobial resistance property of clinical multidrug-resistant isolates of neisseria gonorrhoeae. Antimicrob Agents Chemother 58:3556–3559
Hassan KA et al (2019) Short-chain diamines are the physiological substrates of PACE family efflux pumps. Proc Natl Acad Sci U S A 116:18015–18020
Hollenstein K, Dawson RJ, Locher KP (2007) Structure and mechanism of ABC transporter proteins. Curr Opin Struct Biol 17:412–418. https://doi.org/10.1016/j.sbi.2007.07.003
Hughes D, Andersson DI (2017) Environmental and genetic modulation of the phenotypic expression of antibiotic resistance. FEMS Microbiol Rev 41:374–391. https://doi.org/10.1093/femsre/fux004
Jack DL, Storms ML, Tchieu JH, Paulsen IT, Saier MH (2000) A broad-specificity multidrug efflux pump requiring a pair of homologous SMR-Type proteins. J Bacteriol 182:2311. https://journals.asm.org/journal/jb
Joshi P et al (2014) Osthol and curcumin as inhibitors of human Pgp and multidrug efflux pumps of Staphylococcus aureus: reversing the resistance against frontline antibacterial drugs. Medchemcomm 5:1540–1547
Kabra R, Chauhan N, Kumar A, Ingale P, Singh S (2019) Efflux pumps and antimicrobial resistance: Paradoxical components in systems genomics. Prog Biophysics Mol Biol 141:15–24. https://doi.org/10.1016/j.pbiomolbio.2018.07.008
Kakarla P et al (2017) Inhibition of the multidrug efflux pump LmrS from Staphylococcus aureus by cumin spice Cuminum cyminum. Arch Microbiol 199:465–474
Kolbusz MA, ter Horst R, Slotboom DJ, Lolkema JS (2010) Orientation of small multidrug resistance transporter subunits in the membrane: correlation with the positive-inside rule. J Mol Biol 402:127–138
Kornelsen V and Kumar A (2021) Update on multidrug resistance efflux pumps in Acinetobacter spp. https://journals.asm.org/journal/aac
Kourtesi C et al (2013) Send orders of reprints at reprints@benthamscience.net microbial efflux systems and inhibitors: approaches to drug discovery and the challenge of clinical implementation. Open Microbiol J 7:34
Kumar S, Mukherjee MM, Varela MF (2013) Modulation of bacterial multidrug resistance efflux pumps of the major facilitator superfamily. Int J Bacteriol 2013:1–15
Kumar A, Schweizer HP (2005) Bacterial resistance to antibiotics: active efflux and reduced uptake. Adv Drug Deliv Rev 57:1486–1513. https://doi.org/10.1016/j.addr.2005.04.004
Kumar S et al (2013) Modulation of bacterial multidrug resistance efflux pumps of the major facilitator superfamily. Int J Bacteriol 2013:204141. https://doi.org/10.1155/2013/20414
Kumar S et al (2020) Functional and structural roles of the major facilitator superfamily bacterial multidrug efflux pumps. Microorganisms 8:266. https://doi.org/10.3390/microorganisms8020266
Kuroda T, Tsuchiya T (2009) Multidrug efflux transporters in the MATE family. Biochim Biophys Acta Proteins and Proteom 1794:763–768. https://doi.org/10.1016/j.bbapap.2008.11.012
Li XZ, Nikaido H (2009) Efflux-mediated drug resistance in bacteria: an update. Drugs 69:1555–1623. https://doi.org/10.2165/11317030-000000000-00000
Lomovskaya O, Watkins W (2001) Inhibition of efflux pumps as a novel approach to combat drug resistance in bacteria JMMB symposium. J Mol Microbiol Biotechnol 650:225. www.caister.com/bacteria-plant
Lomovskaya O et al (2001) Identification and characterization of inhibitors of multidrug resistance efflux pumps in Pseudomonas aeruginosa: novel agents for combination therapy. Antimicrob Agents Chemother 45:105–116
Mahamoud A, Chevalier J, Alibert-Franco S, Kern WV, Pagès JM (2007) Antibiotic efflux pumps in Gram-negative bacteria: the inhibitor response strategy. J Antimicrob Chemother 59:1223–1229. https://doi.org/10.1093/jac/dkl493
Marquez B (2005) Bacterial efflux systems and efflux pumps inhibitors. Biochimie 87:1137–1147. https://doi.org/10.1016/j.biochi.2005.04.012
Motta SS, Cluzel P, Aldana M (2015) Adaptive resistance in bacteria requires epigenetic inheritance, genetic noise, and cost of efflux pumps. PLoS One 10:e0118464
Murakami S, Nakashima R, Yamashita E, Matsumoto T, Yamaguchi A (2006) Crystal structures of a multidrug transporter reveal a functionally rotating mechanism. Nature 443:173–179
Negi N, Prakash P, Gupta ML, Mohapatra TM (2014) Possible role of curcumin as an efflux pump inhibitor in multi drug resistant clinical isolates of Pseudomonas aeruginosa. J Clin Diagn Res 8:DC04–DC07
Nikaido H (2011) Structure and mechanism of Rnd-type multidrug efflux pumps. Adv Enzymol Relat Areas Mol Biol 77:1–60
Oluwatuyi M, Kaatz GW, Gibbons S (2004) Antibacterial and resistance modifying activity of Rosmarinus officinalis. Phytochemistry 65:3249–3254
Pagès JM, Amaral L (2009) Mechanisms of drug efflux and strategies to combat them: challenging the efflux pump of gram-negative bacteria. Biochim Biophys Acta Proteins and Proteom 1794:826–833. https://doi.org/10.1016/j.bbapap.2008.12.011
Pasqua M et al (2019) The varied role of efflux pumps of the mfs family in the interplay of bacteria with animal and plant cells. Microorganisms 7:285
Paulsen FT et al (1996) The SMR family: a novel family of multidrug efflux proteins involved with the efflux of lipophilic drugs. Mol Microbiol 19:1167
Piddock LJV (2006a) OPINION Multidrug-resistance efflux pumps-not just for resistance. www.nature.com/reviews/micro
Piddock LJV (2006b) Clinically relevant chromosomally encoded multidrug resistance efflux pumps in bacteria. Clin Microbiol Revi 19:382–402. https://doi.org/10.1128/CMR.19.2.382-402.2006
Poole K (2001) Multidrug efflux pumps and antimicrobial resistance in Pseudomonas aeruginosa and related organisms JMMB Symposium. J Mol Microbiol Biotechnol 613:www.caister.com/bacteria-plant
Poole K (2007) Efflux pumps as antimicrobial resistance mechanisms. Ann Med 39:162–176. https://doi.org/10.1080/07853890701195262
Poulsen BE, Rath A, Deber CM (2009) The assembly motif of a bacterial small multi drug resistance protein. J Biol Chem 284:9870–9875
Prasad R, Goffeau A (2012) Yeast ATP-binding cassette transporters conferring multidrug resistance. Annu Rev Microbiol 66:39–63
Prasch S, Bucar F (2015) Plant derived inhibitors of bacterial efflux pumps: an update. Phytochem Rev 14:961–974. https://doi.org/10.1007/s11101-015-9436-y
Prestinaci F, Pezzotti P, Pantosti A (2015) Antimicrobial resistance: a global multifaceted phenomenon. Pathog Glob Health 109:309–318. https://doi.org/10.1179/2047773215Y.0000000030
Pulingam T et al (2022) Antimicrobial resistance: prevalence, economic burden, mechanisms of resistance and strategies to overcome. Eur J Pharm Sci 170:106103. https://doi.org/10.1016/j.ejps.2021.106103
Radchenko M, Symersky J, Nie R, Lu M (2015) Structural basis for the blockade of MATE multidrug efflux pumps. Nat Commun 6:7995
Ranaweera I et al (2015) Structural comparison of bacterial multidrug efflux pumps of the major facilitator superfamily HHS public access. Trends Cell Mol Biol 10:131
Rath A, Melnyk RA, Deber CM (2006) Evidence for assembly of small multidrug resistance proteins by a ‘two-faced’ transmembrane helix. J Biol Chem 281:15546–15553
Reddy Bolla J, Howes AC, Fiorentino F, Robinson CV (2020) Assembly and regulation of the chlorhexidine-specific efflux pump AceI. Proc Natl Acad Sci U S A 117:17011. https://doi.org/10.1073/pnas.2003271117/-/DCSupplemental
Reygaert CW (2018) An overview of the antimicrobial resistance mechanisms of bacteria. AIMS Microbiol 4:482–501
Rouquette-Loughlin C, Dunham SA, Kuhn M, Balthazar JT, Shafer WM (2003) The NorM efflux pump of Neisseria gonorrhoeae and Neisseria meningitidis recognizes antimicrobial cationic compounds. J Bacteriol 185:1101–1106
Routh MD et al (2011) Efflux pumps of the resistance-nodulation-division family: a perspective of their structure, function and regulation in gram-negative bacteria. Adv Enzymol Relat Areas Mol Biol 77:109
Roy SK, Pahwa S, Nandanwar H, Jachak SM (2012) Phenylpropanoids of Alpinia galanga as efflux pump inhibitors in mycobacterium smegmatis mc2 155. Fitoterapia 83:1248–1255
Roy SK et al (2013) NorA efflux pump inhibitory activity of coumarins from Mesua ferrea. Fitoterapia 90:140–150
Sajjad Aghayan S et al (2017) The effects of Berberine and Palmatine on efflux pumps inhibition with different gene patterns in Pseudomonas aeruginosa isolated from burn infections. Avicenna J Med Biotechnol 9:2
Sana M, Jameel H, Rahman M (2015) Miracle remedy: inhibition of bacterial efflux pumps by natural products. J Biotechnology Biochem 4:1
Seukep AJ, Kuete V, Nahar L, Sarker SD, Guo M (2020) Plant-derived secondary metabolites as the main source of efflux pump inhibitors and methods for identification. J Pharm Anal 10:277–290. https://doi.org/10.1016/j.jpha.2019.11.002
Shafer WM, Waring AJ, Lehrer RI (1998) Modulation of Neisseria gonorrhoeae susceptibility to vertebrate antibacterial peptides due to a member of the resistance nodulationdivision efflux pump family. Proc Natl Acad Sci U S A 95:1829
Siriyong T et al (2017) Conessine as a novel inhibitor of multidrug efflux pump systems in Pseudomonas aeruginosa. BMC Complement Altern Med 17:405
Spengler G, Kincses A, Gajdács M, Amaral L (2017) New roads leading to old destinations: efflux pumps as targets to reverse multidrug resistance in bacteria. Molecules 22:468. https://doi.org/10.3390/molecules22030468
Stavri M, Piddock LJV, Gibbons S (2007) Bacterial efflux pump inhibitors from natural sources. J Antimicrob Chemother 59:1247–1260. https://doi.org/10.1093/jac/dkl460
Stermitz FR, Scriven LN, Tegos G and Lewis K (n.d.) Heruntergeladen von: UC Santa Barbara
Sun J, Deng Z, Yan A (2014) Bacterial multidrug efflux pumps: mechanisms, physiology and pharmacological exploitations. Biochem Biophys Res Commun 453:254–267. https://doi.org/10.1016/j.bbrc.2014.05.090
Thomas E, Renau R et al (1999) Inhibitors of efflux pumps in Pseudomonas aeruginosa potentiate the activity of the fluoroquinolone antibacterial levofloxacin. J Med Chem 49:5597
Verma P, Tiwari M, Tiwari V (2021) Efflux pumps in multidrug-resistant Acinetobacter baumannii: current status and challenges in the discovery of efflux pumps inhibitors. Microb Pathog 152:104766
Watkins WJ et al (2003) The relationship between physicochemical properties, in vitro activity and pharmacokinetic profiles of analogues of diamine-containing efflux pump inhibitors. Bioorg Med Chem Lett 13:4241–4244
Yerushalmi H, Lebendiker M, Schuldiner S (1996) Negative dominance studies demonstrate the oligomeric structure of EmrE, a multidrug antiporter from Escherichia coli. J Biol Chem 271:31044–31048
Zgurskaya H, Introduction I (2021) Transporters, porins, and efflux pumps. Chem Rev 121:5095–5097. https://doi.org/10.1021/acs.chemrev.1c00010
Acknowledgments
TB is supported by the Early Career Research (ECR) Award, Science & Engineering Research Board (SERB) JRF fellowship. MA is supported by the UGC junior research fellowship. The authors would like to thank ECR-SERB (File No. ECR/2018/001015), Government of India, for funding.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2024 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Biswas, T., Ahmed, M., Mondal, S. (2024). Rejuvenating the Potential of Antimicrobials Via Targeted Therapy of Efflux Pumps: The Advent of Phytotherapeutics. In: Kumar, V., Shriram, V., Dey, A. (eds) Medicinal Plants and Antimicrobial Therapies. Springer, Singapore. https://doi.org/10.1007/978-981-99-7261-6_4
Download citation
DOI: https://doi.org/10.1007/978-981-99-7261-6_4
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-99-7260-9
Online ISBN: 978-981-99-7261-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)