Abstract
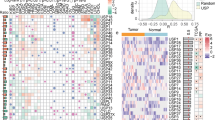
Ubiquitin-specific proteases (USPs) play a major role in the progression of cancers. In this chapter, the presence and relative abundance of paralogous genes in the USP family were studied along with their respective interaction patterns. The effect of the gene duplication events on the overall gene expression in various cancer types was also investigated. Functional divergence of the paralogous genes was evaluated and their corresponding evolution rates were also determined. It was found that the USP family contains an abundance of paralogous genes, among which most paralogous gene pairs display diverging functionalities. Furthermore, higher proportions of genes that promote cancer exist and are thus, implicated in a larger number of protein–protein interaction networks. Also, gene duplication events that lead to the production of noncancerous paralogs were observed to be preferred in this family. Lastly, it was observed that the group of paralogs which display different roles in cancer has the highest rate of evolution among all other paralogs. This evolutionary investigation would provide a further insight into the interconnection of the USP family with the JAK-STAT pathway as well as would inhibit tumorigenesis.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Clague MJ, Urbé S, Komander D (2019) Breaking the chains: deubiquitylating enzyme specificity begets function. Nat Rev Mol Cell Biol 20:338–352. https://doi.org/10.1038/s41580-019-0099-1
Mevissen TET, Komander D (2017) Mechanisms of deubiquitinase specificity and regulation. Annu Rev Biochem 86:159–192. https://doi.org/10.1146/annurev-biochem-061516-044916
Scheffner M, Nuber U, Huibregtse JM (1995) Protein ubiquitination involving an E1–E2–E3 enzyme ubiquitin thioester cascade (1995). https://doi.org/10.1038/373081a0
Aggarwal K, Massagué J (2012) Ubiquitin removal in the TGF-β pathway. Nat Cell Biol 14:656–657. https://doi.org/10.1038/ncb2534
Jin J, Liu J, Chen C, Liu Z, Jiang C, Chu H, Pan W, Wang X, Zhang L, Li B, Jiang C, Ge X, Xie X, Wang P (2016) The deubiquitinase USP21 maintains the stemness of mouse embryonic stem cells via stabilization of Nanog. Nat Commun 7:1–15. https://doi.org/10.1038/ncomms13594
Zeng H, Qu J, Jin N, Xu J, Lin C, Chen Y, Yang X, Cell XH-C (2016) Undefined: Feedback activation of leukemia inhibitory factor receptor limits response to histone deacetylase inhibitors in breast cancer. Elsevier
Cheng YC, Shieh SY (2018) Deubiquitinating enzyme USP3 controls CHK1 chromatin association and activation. Proc Natl Acad Sci U S A 115:5546–5551. https://doi.org/10.1073/pnas.1719856115
Wang YC, Wu YS, Hung CY, Wang SA, Young MJ, Hsu TI, Hung JJ (2018) USP24 induces IL-6 in tumor-associated microenvironment by stabilizing p300 and β-TrCP and promotes cancer malignancy. Nat Commun 9 (2018). https://doi.org/10.1038/s41467-018-06178-1
Zhang C, Lu J, Zhang QW, Zhao W, Guo JH, Liu SL, Wu YL, Jiang B, Gao FH (2016) USP7 promotes cell proliferation through the stabilization of Ki-67 protein in non-small cell lung cancer cells. Int J Biochem Cell Biol 79:209–221. https://doi.org/10.1016/j.biocel.2016.08.025
Liu Q, Aminu B, Roscow O, Zhang W (2021) Targeting the ubiquitin signaling cascade in tumor microenvironment for cancer therapy. Int J Mol Sci 22:1–26. https://doi.org/10.3390/ijms22020791
Hu Y, Hong Y, Xu Y, Liu P, Guo DH, Chen Y (2014) Inhibition of the JAK/STAT pathway with ruxolitinib overcomes cisplatin resistance in non-small-cell lung cancer NSCLC. Apoptosis 19:1627–1636. https://doi.org/10.1007/S10495-014-1030-Z
Yeh C-T, Huang W-C, Rao YK, Ye M, Lee W-H, Wang L-S, Tzeng DTW, Wu C-H, Shieh Y-S, Huang C-YF, Chen Y-J, Hsiao M, Wu ATH, Yang Z, Tzeng Y-M (2013) A sesquiterpene lactone antrocin from Antrodia camphorata negatively modulates JAK2/STAT3 signaling via microRNA let-7c and induces apoptosis in lung cancer cells. Carcinogenesis 34, 2918–2928 (2013). https://doi.org/10.1093/CARCIN/BGT255
Dandage R, Landry CR (2019) Paralog dependency indirectly affects the robustness of human cells. Mol Syst Biol 15:1–18 (2019). https://doi.org/10.15252/msb.20198871
Hsiao TL, Vitkup D (2008) Role of duplicate genes in robustness against deleterious human mutations. PLoS Genet 4. https://doi.org/10.1371/journal.pgen.1000014
Chen WH, Zhao XM, van Noort V, Bork P (2013) Human monogenic disease genes have frequently functionally redundant paralogs. PLoS Comput Biol 9. https://doi.org/10.1371/journal.pcbi.1003073
Bateman A (2019) UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res 47:D506–D515. https://doi.org/10.1093/nar/gky1049
Yates AD et al (2020) Ensembl 2020. Nucleic Acids Res 48:D682–D688. https://doi.org/10.1093/nar/gkz966
Pruitt KD et al (2009) The consensus coding sequence (CCDS) project: identifying a common protein-coding gene set for the human and mouse genomes. Genome Res 19:1316–1323. https://doi.org/10.1101/gr.080531.108
Keshava Prasad TS et al (2009) Human protein reference database—2009 update. Nucleic Acids Res 37:767–772. https://doi.org/10.1093/nar/gkn892
Kuo HC, Lin PY, Chung TC, Chao CM, Lai LC, Tsai MH, Chuang EY (2011) DBCAT: database of CpG islands and analytical tools for identifying comprehensive methylation profiles in cancer cells. J Comput Biol 18:1013–1017. https://doi.org/10.1089/cmb.2010.0038
Chen D, Ning Z, Chen H, Lu C, Liu X, Xia T, Qi H, Wang W, Ling T, Guo X, Tekcham DS, Liu X, Liu J, Wang A, Yan Q, Liu JW, Tan G, Piao HI (2020) An integrative pan-cancer analysis of biological and clinical impacts underlying ubiquitin-specific-processing proteases. Oncogene 39:587–602. https://doi.org/10.1038/s41388-019-1002-4
Wei R, Liu X, Yu W, Yang T, Cai W, Liu J, Huang X, Xu Gt, Zhao S, Yang J, Liu S (2015) Deubiquitinases in cancer. Oncotarget 6:12872–12889. https://doi.org/10.18632/oncotarget.3671
Young MJ, Hsu KC, Lin TE, Chang WC, Hung JJ (2019) The role of ubiquitin-specific peptidases in cancer progression. J Biomed Sci 26:1–14. https://doi.org/10.1186/s12929-019-0522-0
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45:W98–W102. https://doi.org/10.1093/nar/gkx247
Dimmer EC, Huntley RP, Barrell DG, Binns D, Draghici S, Camon EB, Hubank M, Talmud PJ, Apweiler R, Lovering RC (2008) The gene ontology—providing a functional role in proteomic studies. Proteomics 8. https://doi.org/10.1002/pmic.200800002
Baudot A, Jacq B, Brun C (2004) A scale of functional divergence for yeast duplicated genes revealed from analysis of the protein-protein interaction network. Genome Biol 5. https://doi.org/10.1186/gb-2004-5-10-r76
Goh KIl, Cusick ME, Valle D, Childs B, Vidal M, Barabási AL (2007) The human disease network. Proc Natl Acad Sci U S A 104:8685–8690. https://doi.org/10.1073/pnas.0701361104
He X, Zhang J (2006) Why do hubs tend to be essential in protein networks? PLoS Genet 2:0826–0834. https://doi.org/10.1371/journal.pgen.0020088
Acknowledgements
Authors are thankful to the Ms. Alankar Roy (Enrollment no: A91704120067) from Amity Institute of Biotechnology, Amity University, Kolkata for the support. Authors are also grateful to Department of Biochemistry and Biophysics, University of Kalyani, West Bengal for the cooperation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Ethics declarations
On behalf of all authors, the corresponding states that there is no conflict of interest.
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Ray, S., Banerjee, A. (2023). Exploring the Human USP Gene Family and Its Association with Cancer: An In Silico Study. In: Chakraborty, B., Biswas, A., Chakrabarti, A. (eds) Advances in Data Science and Computing Technologies. ADSC 2022. Lecture Notes in Electrical Engineering, vol 1056. Springer, Singapore. https://doi.org/10.1007/978-981-99-3656-4_70
Download citation
DOI: https://doi.org/10.1007/978-981-99-3656-4_70
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-99-3655-7
Online ISBN: 978-981-99-3656-4
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)