Abstract
Members of the genus Alphavirus are mostly mosquito-borne pathogens that cause disease in their vertebrate hosts. Chikungunya virus (CHIKV), which is one member of the genus Alphavirus [1], has been a major health problem in endemic areas since its re-emergence in 2006. CHIKV is transmitted to mammalian hosts by the Aedes mosquito, causing persistent debilitating symptoms in many cases. At present, there is no specific treatment or vaccine. Experiments involving live CHIKV need to be performed in BSL-3 facilities, which limits vaccine and drug research. The emergence of pseudotyped virus technology offered the potential for the development of a safe and effective evaluation method. In this chapter, we review the construction and application of pseudotyped CHIKVs, the findings from which have enhanced our understanding of CHIKV. This will, in turn, enable the exploration of promising therapeutic strategies in animal models, with the ultimate aim of developing effective treatments and vaccines against CHIKV and other related viruses.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
16.1 Biological Characteristics of Chikungunya Virus
Viruses belonging to the Alphavirus genus can infect humans and often cause symptoms such as joint pain and fever. According to the latest revision report published by the International Committee on Taxonomy of Viruses (ICTV) in 2021, the Alphavirus genus comprises 32 species, including Aura virus, Barmah Forest virus, Bebaru virus, Caaingua virus, Cabassou virus, Chikungunya virus (CHIKV), Sindbis virus, Eastern equine encephalitis virus, Western equine encephalitis virus, Mayaro virus, Semliki Forest virus, and Onyong-nyong virus.
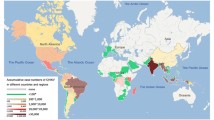
CHIKV is an arthropod-borne Alphavirus that is transmitted to humans primarily via the bite of an infected mosquito. CHIKV is becoming widespread in Africa, South Asia, and Southeast Asia, with high infection rates [2, 3]. Infection of humans by CHIKV can cause Chikungunya fever (CHIKF), an acute febrile illness associated with severe, often debilitating, polyarthralgias, fever, rash, and even death [4,5,6]. CHIKV has emerged in more than 100 countries, with approximately one million people infected each year. In 2017, the World Health Organization (WHO) reported 10 potential infectious diseases for priority research, with CHIKV on the list.
16.1.1 Molecular Structure
CHIKV is an arbovirus of the Alphavirus genus, which belongs to the Togaviridae family [7]. It is a single-stranded positive-sense RNA virus with a full-length genome of 11.8 kb, including 5′ and 3′ untranslated regions (UTRs) and two open reading frames (ORFs) (Fig. 16.1) [8]. One ORF encodes four non-structural proteins, namely, nsP1, nsP2, nsP3, and nsP4; the other ORF encodes five structural proteins, namely, capsid proteins C, E3, E2, 6 K, and E1 [9].
Capsid protein C includes two functional domains: an N-terminal RNA binding domain and a C-terminal protease domain. The former binds to genomic RNA, while the latter is involved in the assembly of structural proteins [10, 11]. E1 contains a hydrophobic fusion loop, and under acidic conditions, the CHIKV membrane protein undergoes a conformational change, exposing the fusion loop and promoting nucleocapsid release [12]. Glycoprotein E2 binds to receptors on the host cell membrane to form an endosome and enter the cytoplasm [13, 14]. E3 prevents premature fusion of the heterodimer formed by E2 and E1 with the cell membrane [15]. Studies have shown that two amino acid residues, Gly91 and His230, play a key role in the membrane fusion of E1 [16]. 6 K is the signal peptide of glycoprotein E1 and consists of only 55–60 amino acids [15, 17]. Recently, through structural analysis, Chinese scholars found that the matrix remodeling-related protein 8 (MXRA8) molecule widely distributed on the surface of chondrocytes, muscle cells, and skeletal muscle cells is the receptor of CHIKV, which confirmed the mechanism of host cell invasion by CHIKV and provided a new target for the research and development of vaccines and broad-spectrum neutralizing antibodies [18,19,20].
16.1.2 Genotypes and Variants
According to genetic analysis, CHIKVs comprise one serotype and three genotypes, namely, the West African genotype, the East-Central-Southern African (ECSA) genotype, and the Asian genotype [21].
Sequence analysis of the isolated viruses revealed that the re-emerging strains of CHIKV originating from Reunion Island belonged to the ESCA genotype [22, 23], while the strains currently circulating in Southeast Asia include both Asian and ESCA genotypes [24,25,26,27]. The strains introduced into St. Maarten and the Americas belong to the Asian genotype [24, 25, 28]. Some evidence suggests that the CHIKV strain isolated in Brazil in 2014 has ESCA genes similar to those circulating in Angola [29].
Numerous studies have shown that the large-scale CHIKF outbreak in Reunion Island in 2005–2006 was caused by a CHIKV strain (ECSA genotype) transmitted via Aedes albopictus. This CHIKV epidemic strain harbors a A226V mutation on the E1 protein, which changes the transmission host of CHIKV from Aedes aegypti to A. albopictus [30,31,32,33].
In 2009, a new ECSA subtype (GenBank accession no. HM159390) was discovered in Hyderabad, India, with a K211E mutation in the E1 protein (E1-K211E) [34]. In 2010, a CHIKV strain containing two novel mutations E1-K211E and E2-V264A was found among all CHIKV isolates from New Delhi, India [35]. In 2011 and 2012, the abovementioned epidemic strains (E1-K211E and E2-V264A) were found in Tamil Nadu and Kolkata, respectively [34, 36].
16.1.3 Pathogenic Mechanisms and Biosafety Risk
CHIKV can be cultured on a variety of passage cell lines, such as C6/36, AP61, Vero, LLC- MK2, and BHK-21 cells [37]. CHIKV can also be isolated and cultured in vivo by intracranial inoculation of suckling rats aged 1–3 days.
CHIKV enters the host’s skin 24–48 hours after a mosquito bite, resulting in viremia. Viremia manifests as a systemic proinflammatory response associated with leukopenia and mononucleosis. The virus circulates through the blood and attacks fibroblasts in muscles and joints, macrophages in lymphoid tissue, and endothelial cells in the liver [12, 38, 39].
CHIKV is considered to cause a high level of individual harm but a low level of group harm. Nucleic acid and serological testing of A. albopictus-inactivated serum and frozen specimens can be performed in biosafety level (BSL)-2 laboratories, but potential and confirmed CHIKV-positive samples have to be handled in BSL-3 facilities. It is important that researchers adhere to strict safety precautions to avoid unnecessary biosafety risks.
16.2 Construction of Pseudotyped CHIKV
Currently, there are three commonly used packaging backbone vectors for pseudotyped CHIKVs; these are human immunodeficiency virus (HIV-1)-based lentiviral vectors, vesicular stomatitis virus (VSV)-based vectors, and murine leukemia virus (MLV)-based vectors. In addition, when the abovementioned packaging backbone vector cannot construct a pseudotyped virus, a recombinant pseudotyped virus can be constructed by reverse genetics technology.
16.2.1 Construction of Pseudotyped CHIKV Using Different Vectors
16.2.1.1 Lentiviral Vectors
The HIV-1-based lentiviral system is the most commonly used packaging system for the construction of pseudotyped CHIKVs. As early as 2017, our group constructed the West African genotype pseudotyped CHIKV using the HIV-1 system [40]. The full-length CHIKV West African genotype (strain 37,997, GenBank accession no. AY726732) structural protein (C-E3-E2-6K-E1) was synthesized, and after codon optimization in mammalian cells, it was cloned into the eukaryotic expression vector pcDNA3.1(+), which constituted the CHIKV membrane protein expression plasmid pCMV3.1-CHIKV. Then, 293 T cells were transfected with the appropriate ratio (1:2) of membrane protein expression plasmid pCMV3.1-CHIKV and backbone plasmid pSG3Δenv.cmv.Fluc, and the supernatant was collected after 48 hours. The final prepared CHIKV pseudotyped virus titer was 1.0 × 107 TCID50/ml. In 2019, Korean scholars constructed a pseudotyped CHIKV containing two reporter genes based on a lentivirus system [41]. In this study, a three-plasmid system was used to co-transfect 293 T cells with a full-length CHIKV ECSA genotype (strain KNIH/2009/77, GenBank accession no. MN158171.1), structural protein (C-E3-E2-6K-E1), expression plasmid pCMV2-FLAG-CHIKVst, a lentivirus-based dual vector (Luc2P-pLVX-IRES-ZsGreen1), and a packaging vector of lentivirus reporter (psPAX2). The supernatant was harvested to obtain the pseudotyped virus, CHIKVpseudo. There are two important findings from this study. First, when constructing the membrane protein expression plasmid, FLAG tag was added to aid the subsequent detection of protein expression. Second, the system contains two reporter genes, encoding luciferase and GFP. The tracer results of the two reporter genes can be compared simultaneously.
The packaging process of pseudotyped CHIKV based on lentiviral vectors is generally conducted as follows:
-
1.
Seed 293 T cells one day before transfection.
-
2.
Incubate the cells with DMEM (high glucose) containing 10% FBS for 12–18 h. The cells should be 80% confluent before transfection.
-
3.
Co-transfect CHIKV structural protein expression plasmid and lentivirus-based packaging vector plasmid/plasmids using Lipofectamine 3000 or other transfection reagents. Dilute the plasmid with Opti-MEM if necessary.
-
4.
Incubate at 37 °C for 4–6 h.
-
5.
Remove the medium, add fresh DMEM with 10% FBS, and incubate at 37 °C overnight for another 42–44 h.
-
6.
Harvest the culture supernatant. Filter using a 0.45 μM pore-size filter and store at ˗70 °C until use.
All work involving pseudotyped CHIKV was performed in a BSL-2 facility.
16.2.1.2 VSV-Based Vectors
A pseudotyped CHIKV based on the VSV vector system has also been constructed [42]. Researchers constructed a West African genotype (strain 37,997, GenBank accession no. AY726732, E3-E2-6K-E1) CHIKV pseudotyped virus based on the VSV vector containing the GFP reporter gene and the firefly luciferase reporter gene, respectively, for the subsequent study of the virus entry mechanism.
The packaging process of pseudotyped CHIKV based on VSV vectors is generally conducted as follows:
-
1.
Seed 293 T cells into a 10-cm dish one day before transfection.
-
2.
Incubate the cells with 10 ml of DMEM (high glucose) containing 10% FBS. The cells should be 80% confluent before transfection.
-
3.
Remove the medium, wash the cells once with the medium, and then add 5 ml of fresh medium.
-
4.
Transfect 5 μg of CHIKV structural protein expression plasmid using Lipofectamine 3000 or other transfection reagents. Dilute the plasmid with Opti-MEM if necessary.
-
5.
Incubate at 37 °C for 6 h.
-
6.
Remove the medium, add fresh DMEM with 10% FBS, and incubate at 37 °C overnight for another 18 h.
-
7.
Inoculate with a multiplicity of infection of 1.0 of VSVΔG/GFP*-G, VSVΔG/Luci*-G, or VSVΔG/SEAP*-G.
-
8.
Adsorb the viruses at 37 °C for 6–8 h.
-
9.
Remove the medium and wash with phosphate-buffered saline (PBS) or serum-free DMEM three times.
-
10.
Add 10 ml of the medium and incubate at 37 °C overnight (17–18 h).
-
11.
Harvest the culture supernatant. Filter using a 0.45 μM pore-size filter and store at ˗70 °C until use.
All work involving pseudotyped CHIKV was performed in a BSL-2 facility.
16.2.1.3 MLV-Based Vectors
Like HIV-1, MLV is also a retrovirus that is commonly used as a pseudotyped viral packaging vector. French researchers constructed pseudotyped CHIKV using an MLV vector. Briefly, 293 T cells were co-transfected with the pTG5349 MLV packaging plasmid, the pTG13077 plasmid encoding an MLV-based vector containing a CMV-GFP internal transcriptional unit, a plasmid encoding the Gag-pol proteins of MLV, and a plasmid encoding the CHIKV La Réunion infectious clone (strain LR2006 OPY1, GenBank accession no. KT449801.1) viral envelope glycoproteins (C-E3-E2-6K-E1) [43]. After 24 hours of co-transfection with plasmids, the packaging effect of pseudotyped virus can be determined by observing the fluorescence signal intensity of GFP.
According to published studies and our previous research, all three genotypes of CHIKV can successfully be used to construct pseudotyped viruses. For pseudotyped CHIKV, the expression of E protein can be improved by removing capsid protein C, but whether or not the full-length structural protein is selected, the E2 and E1 protein genomes are essential for the construction of pseudotyped CHIKV [44].
16.2.2 CHIKV Infectious Clones and Virus-like Particles
Compared with other viruses, alphaviruses are more suitable for the construction of infectious clones and virus-like particles. In addition to pseudotyped viruses, several CHIKV infectious clones expressing different reporter genes have been developed to study viral replication and pathogenic mechanisms. The insertion position of the reporter gene in the Alphavirus genome is critical for the stability of infectious clones and the virulence of animals [45].
A common strategy for constructing Alphavirus infectious clones is to initiate reporter gene expression via an additional subgenomic promoter upstream of the true subgenomic promoter (5’-26S) or the E1 protein (3’-DP) downstream. Compared with 3’-DP, the 5’-26S method is more stable with minimal loss of viral virulence [45, 46]. A reproducible CHIKV infectious clone containing a near-infrared fluorescent protein (iRFP) reporter gene—CHIKV-iRFP—was developed, which was used to establish in vivo imaging of small animals. Briefly, the iRFP gene was fused to the full-length CHIKV genome (Asian lineage, GenBank accession no. KC488650) between nsP4 and the capsid (C), and another subgenomic (SG) T7 promoter was introduced downstream of the iRFP gene to facilitate the replication of recombinant viruses [47].
16.3 Application of Pseudotyped CHIKV
16.3.1 Neutralizing Assay Based on Pseudotyped CHIKV
The main application of pseudotyped CHIKV is to establish a pseudotyped virus-based neutralizing antibody assay (PBNA). The process of pseudotyped CHIKV infection of cells can be blocked by neutralizing antibodies in test samples. After a test sample interacts with the pseudotyped CHIKV, detecting the expression of the reporter gene carried by the pseudotyped virus can reflect the titer of the neutralizing antibody to be tested. Compared with plaque reduction neutralization test (PRNT), this method has the advantages of high safety, simple operation, easy access, and potential high throughput. In addition, PBNA allows for a reduction in the biosafety level of CHIKV neutralizing antibody detection (from BSL-3 to BSL-2), which was comparable to the detection results obtained with the live virus method.
16.3.1.1 Correlation of PBNA and PRNT
Chung et al. constructed a pseudotyped CHIKV based on a lentiviral vector and established a stable PBNA using the pseudotyped virus [41]. The activity of CHIKV E2 antibody was detected using this method, as well as using classical PRNT. The results showed that the use of PBNA yielded a mean Log IC50 value of 5.85 ± 0.09 pg/ml, while the use of PRNT yielded a Log IC50 value of 5.68 ± 0.06 pg/ml. The neutralization assay of CHIKVpseudo showed a CV value of 1.09%, similar to the PRNT CV value of 0.98%. In conclusion, PBNA using CHIKVpseudo is a safe, rapid, and reliable neutralization assay that can be used to evaluate the neutralizing activity of serum or monoclonal antibody against CHIKV.
16.3.1.2 Factors Closely Related to CHIKV PBNA
Undoubtedly, a high titer of pseudotyped virus reliably provides PBNA results. However, other factors can also affect the normal operation of PBNA.
-
(a)
Pseudotyped virus-infected cells: In general, irrespective of the vector used to construct pseudotyped CHIKV, cells susceptible to pseudotyped virus should be selected as target cells for PBNA. Pseudotyped CHIKV has a wide range of cell tropism [48], and 293 T cells [40, 48] and HeLa cells [41] can be used as infective cells in PBNA. In addition, the source of cells should be clear and the number of passages should be controlled. For experiments with longer cell culture times (more than 48 hours), the evaporative effect in edge wells should be considered. When establishing this method, the cell seeding concentration range should be experimentally validated and optimized.
-
(b)
Negative and positive controls: A perfect PBNA should have negative and positive controls to ensure the accuracy of the test results. If international or domestic standards cannot be obtained, the positive serum obtained after animal immunization can be used as a positive control.
-
(c)
Amount of CHIKV pseudotyped virus: Adding too much virus can result in false negatives or can cause the inhibition rate to reverse the concentration gradient, which will affect the experimental results.
-
(d)
Incubation time for CHIKV pseudotyped virus: The co-culture time of cells and pseudotyped CHIKV should be selected according to the specific characteristics of the pseudotyped virus. For example, the co-culture time for PBNA-based VSV vector is generally 24 h, while for the PBNA-based HIV backbone, the co-culture time is generally 48–72 h.
-
(e)
Cut-off value: To determine the critical values for the negative and positive controls (cut-off value), the serum of the negative group with a similar level of background should be selected as the source of the negative sample for establishing the cut-off value. Generally, the test sample size should not be less than 100. The experiment should be repeated at least three times, and appropriate methods should be used to determine the cut-off value, such as using the geometric mean ± 2SD or 3SD as the final cut-off value.
16.3.2 Establishment of an in Vivo Imaging Model of Small Animals
The pseudotyped virus can be used to establish a visual dynamic monitoring model in mice, to study the tissue distribution after virus infection, and to evaluate the in vivo efficacy of monoclonal antibodies, small molecule inhibitors, and vaccines. Using the accurate quantitative performance and imaging characteristics of the system, the infection site of pseudotyped virus in animals can be dynamically observed, the degree of infection can be judged according to the luminescence intensity and location, and the effect of monoclonal antibodies, drugs, and vaccines can be evaluated [49,50,51].
Our group infected mice with a HIV-1 vector-based pseudotyped CHIKV, and after optimizing variables, such as the mouse strain, infection route, and detection time, finally established a visual CHIKV pseudotyped virus infection model. Subsequently, the passive immunization effect of antiserum and the active immunization effect of the DNA vaccine were evaluated in vivo using the established mouse infection model [40].
16.3.3 Use of a Capture Antigen in CHIKV IgM Detection
The reference method for the detection of CHIKV-specific IgM is the IgM capture enzyme-linked immunosorbent assay (MAC-ELISA). In the MAC-ELISA method, CHIKV viral lysate is usually used as the antigen bound to the specific IgM to be detected. However, obtaining viral lysates is time-consuming and requires BSL-3 facilities. Chemical treatments such as beta-propiolactone must be used to inactivate the virus before use. However, beta-propiolactone treatment may alter the overall structure of enveloped viral proteins, which may modulate the affinity of antibodies against these proteins, thereby reducing the sensitivity of detection [52, 53].
To overcome these challenges, researchers used pseudotyped CHIKV as a substitute for CHIKV viral lysate [54]. The purified and lyophilized pseudotyped CHIKV was directly applied to paper-based analytical devices as the coating antigen, and the CHIKV IgM antibody detection results could be obtained within 8–10 min. The results showed that the sensitivity of this method to detection using patient serum was 70.6% and the specificity was ~98%. While these results are promising, they are only preliminary, and more extensive validation with a greater number of samples is needed before pseudotyped viruses can replace traditional antigens in serological tests.
16.3.4 Drug Screening
Until now, treatment for CHIKF has been mainly symptomatic because of the lack of an FDA-approved vaccine or effective antiviral drugs. One of the reasons for the delay in the development of anti-CHIKV drugs is the need for BSL-3 facilities in which to conduct the research, which limits the potential for high-throughput operations. Using pseudoviruses to screen small-molecule compounds that have already been approved allows for rapid drug repositioning. In addition, pseudotyped viruses can also be used to screen traditional Chinese medicines with antiviral activity, all of which have the advantages of high throughput and low cost [55, 56].
German scholars determined the anti-CHIKV activity of curcumin or Boswellia serrata gum resin extract (AKBA) using pseudotyped CHIKV based on a lentiviral vector [57]. Both compounds blocked the entry of pseudotyped CHIKV and inhibited pseudotyped CHIKV infection in vitro. The IC50 values were 6.75 μM for AKBA and 10.79 μM for curcumin.
16.3.5 The Mechanism of Viral Infection
An increasing number of reports have shown that pseudotyped viruses play an important role in the study of virus invasion mechanisms. Pseudotyped viruses have the advantages of being safe, easier to manipulate, easier to construct or discover mutations in coat genes, and easier to quantify the process of entry. In addition, the viral envelope protein itself can be studied using pseudotyped viruses, and the entry process of viral replication can be studied separately from other steps. Pseudotyped viruses are mainly used to study virus infection tropism, cell tropism, receptor recognition, and the virus entry mechanism in vitro [58, 59].
There is evidence that CHIKV enters host cells via a clathrin-dependent endocytic pathway [60]. Endosomal acidification inhibitors attenuate CHIKV infection. Therefore, it is believed that CHIKV particles bind to cell surface receptors and are internalized into the host cell endosome, and then endosomal acidification promotes fusion of the virus with the endosomal membrane through the CHIKV envelope (E) protein. Researchers used MLV vector-based pseudotyped CHIKV to infect 293 T and TE671 cells and found that virus particles containing the CHIKV E protein entered 293 T cells by endocytosis but were internalized into TE671 cell vesicles by macrophage phagocytosis [44]. Through vesicle acidification, the conformation of the E protein is altered, thereby activating its ability to promote membrane fusion. In addition, it was found that digestion of E protein by endosomal cathepsin B more efficiently promoted membrane fusion activity and enhanced infection.
16.4 Conclusion
The ongoing advances in the pseudotyped virus construction system have further confirmed it as an effective virological research strategy. Although some limitations of this system still need to be taken into account, for example, pseudotyped viruses cannot fully simulate the pathogenesis of live virus infection [61], and the innovative virus packaging system is needed for application.
Abbreviations
- BSL:
-
Biosafety level
- CHIKF:
-
Chikungunya fever
- CHIKV:
-
Chikungunya virus
- ECSA:
-
East-Central-Southern African
- HIV:
-
Human immunodeficiency virus
- MLV:
-
Murine leukemia virus
- MXRA8:
-
Matrix remodeling-related protein 8
- ORF:
-
Open reading frame
- PBNA:
-
Pseudotyped virus-based neutralizing antibody assay
- PRNT:
-
Plaque reduction neutralization test
- UTR:
-
Untranslated region
- VSV:
-
Vesicular stomatitis virus
References
Chen, R., et al.: ICTV Virus Taxonomy Profile: Togaviridae. J. Gen. Virol. 99, 761–762 (2018). https://doi.org/10.1099/jgv.0.001072
Josseran, L., et al.: Chikungunya disease outbreak, Reunion Island. Emerg. Infect. Dis. 12, 1994–1995 (2006). https://doi.org/10.3201/eid1212.060710
Kalantri, S.P., Joshi, R., Riley, L.W.: Chikungunya epidemic: an Indian perspective. Natl. Med. J. India. 19, 315–322 (2006)
Guerrero-Arguero, I., et al.: Alphaviruses: host pathogenesis, immune response, and vaccine & treatment updates. J. Gen. Virol. 102, 1 (2021). https://doi.org/10.1099/jgv.0.001644
Miner, J.J., et al.: Chikungunya viral arthritis in the United States: a mimic of seronegative rheumatoid arthritis. Arthritis Rheumatol. 67, 1214–1220 (2015). https://doi.org/10.1002/art.39027
Schilte, C., et al.: Chikungunya virus-associated long-term arthralgia: a 36-month prospective longitudinal study. PLoS Negl. Trop. Dis. 7, e2137 (2013). https://doi.org/10.1371/journal.pntd.0002137
Burt, F.J., Rolph, M.S., Rulli, N.E., Mahalingam, S., Heise, M.T.: Chikungunya: a re-emerging virus. Lancet. 379, 662–671 (2012). https://doi.org/10.1016/S0140-6736(11)60281-X
Narwal, M., et al.: Crystal structure of chikungunya virus nsP2 cysteine protease reveals a putative flexible loop blocking its active site. Int. J. Biol. Macromol. 116, 451–462 (2018). https://doi.org/10.1016/j.ijbiomac.2018.05.007
Khan, A.H., et al.: Complete nucleotide sequence of chikungunya virus and evidence for an internal polyadenylation site. J. Gen. Virol. 83, 3075–3084 (2002). https://doi.org/10.1099/0022-1317-83-12-3075
Hong, E.M., Perera, R., Kuhn, R.J.: Alphavirus capsid protein helix I controls a checkpoint in nucleocapsid core assembly. J. Virol. 80, 8848–8855 (2006). https://doi.org/10.1128/JVI.00619-06
Perera, R., Owen, K.E., Tellinghuisen, T.L., Gorbalenya, A.E., Kuhn, R.J.: Alphavirus nucleocapsid protein contains a putative coiled coil alpha-helix important for core assembly. J. Virol. 75, 1–10 (2001). https://doi.org/10.1128/JVI.75.1.1-10.2001
Schwartz, O., Albert, M.L.: Biology and pathogenesis of chikungunya virus. Nat. Rev. Microbiol. 8, 491–500 (2010). https://doi.org/10.1038/nrmicro2368
Ashbrook, A.W., et al.: Residue 82 of the chikungunya virus E2 attachment protein modulates viral dissemination and arthritis in mice. J. Virol. 88, 12180–12192 (2014). https://doi.org/10.1128/JVI.01672-14
Smith, T.J., et al.: Putative receptor binding sites on alphaviruses as visualized by cryoelectron microscopy. Proc. Natl. Acad. Sci. U. S. A. 92, 10648–10652 (1995)
Li, L., Jose, J., Xiang, Y., Kuhn, R.J., Rossmann, M.G.: Structural changes of envelope proteins during alphavirus fusion. Nature. 468, 705–708 (2010). https://doi.org/10.1038/nature09546
Kuo, S.C., et al.: Cell-based analysis of chikungunya virus E1 protein in membrane fusion. J. Biomed. Sci. 19, 44 (2012). https://doi.org/10.1186/1423-0127-19-44
Snyder, J.E., et al.: Functional characterization of the alphavirus TF protein. J. Virol. 87, 8511–8523 (2013). https://doi.org/10.1128/JVI.00449-13
Basore, K., et al.: Cryo-EM structure of chikungunya virus in complex with the Mxra8 receptor. Cell. 177, 1725–1737 e1716 (2019). https://doi.org/10.1016/j.cell.2019.04.006
Verma, J., Subbarao, N.: In silico identification of small molecule protein-protein interaction inhibitors: targeting hotspot regions at the interface of MXRA8 and CHIKV envelope protein. J. Biomol. Struct. Dyn. 1, 19 (2022). https://doi.org/10.1080/07391102.2022.2048080
Yin, P., Kielian, M.: BHK-21 cell clones differ in chikungunya virus infection and MXRA8 receptor expression. Viruses. 13 (2021). https://doi.org/10.3390/v13060949
Schuffenecker, I., et al.: Genome microevolution of chikungunya viruses causing the Indian Ocean outbreak. PLoS Med. 3, e263 (2006). https://doi.org/10.1371/journal.pmed.0030263
Arankalle, V.A., et al.: Genetic divergence of chikungunya viruses in India (1963-2006) with special reference to the 2005-2006 explosive epidemic. J. Gen. Virol. 88, 1967–1976 (2007). https://doi.org/10.1099/vir.0.82714-0
Pialoux, G., Gauzere, B.A., Jaureguiberry, S., Strobel, M.: Chikungunya, an epidemic arbovirosis. Lancet Infect. Dis. 7, 319–327 (2007). https://doi.org/10.1016/S1473-3099(07)70107-X
Lanciotti, R.S., Valadere, A.M.: Transcontinental movement of Asian genotype chikungunya virus. Emerg. Infect. Dis. 20, 1400–1402 (2014). https://doi.org/10.3201/eid2008.140268
Sy, A.K., et al.: Molecular characterization of chikungunya virus, Philippines, 2011-2013. Emerg. Infect. Dis. 22, 887–890 (2016). https://doi.org/10.3201/eid2205.151268
Powers, A.M., Brault, A.C., Tesh, R.B., Weaver, S.C.: Re-emergence of chikungunya and O'nyong-nyong viruses: evidence for distinct geographical lineages and distant evolutionary relationships. J. Gen. Virol. 81, 471–479 (2000). https://doi.org/10.1099/0022-1317-81-2-471
Weaver, S.C., Forrester, N.L.: Chikungunya: evolutionary history and recent epidemic spread. Antivir. Res. 120, 32–39 (2015). https://doi.org/10.1016/j.antiviral.2015.04.016
Leparc-Goffart, I., Nougairede, A., Cassadou, S., Prat, C., de Lamballerie, X.: Chikungunya in the Americas. Lancet. 383, 514 (2014). https://doi.org/10.1016/S0140-6736(14)60185-9
Nunes, M.R., et al.: Emergence and potential for spread of chikungunya virus in Brazil. BMC Med. 13, 102 (2015). https://doi.org/10.1186/s12916-015-0348-x
Tsetsarkin, K., et al.: Infectious clones of chikungunya virus (La Reunion isolate) for vector competence studies. Vector Borne Zoonotic Dis. 6, 325–337 (2006). https://doi.org/10.1089/vbz.2006.6.325
Delatte, H., et al.: Aedes albopictus, vector of chikungunya and dengue viruses in Reunion Island: biology and control. Parasite. 15, 3–13 (2008). https://doi.org/10.1051/parasite/2008151003
Vazeille, M., et al.: Two chikungunya isolates from the outbreak of La Reunion (Indian Ocean) exhibit different patterns of infection in the mosquito, Aedes albopictus. PLoS One. 2, e1168 (2007). https://doi.org/10.1371/journal.pone.0001168
Bagny, L., Delatte, H., Quilici, S., Fontenille, D.: Progressive decrease in Aedes aegypti distribution in Reunion Island since the 1900s. J. Med. Entomol. 46, 1541–1545 (2009). https://doi.org/10.1603/033.046.0644
Naresh Kumar, C.V., Sivaprasad, Y., Sai Gopal, D.V.: Genetic diversity of 2006-2009 chikungunya virus outbreaks in Andhra Pradesh, India, reveals complete absence of E1:A226V mutation. Acta Virol. 60, 114–117 (2016)
Shrinet, J., et al.: Genetic characterization of chikungunya virus from New Delhi reveal emergence of a new molecular signature in Indian isolates. Virol. J. 9, 100 (2012). https://doi.org/10.1186/1743-422X-9-100
Taraphdar, D., Chatterjee, S.: Molecular characterization of chikungunya virus circulating in urban and rural areas of West Bengal, India after its re-emergence in 2006. Trans. R. Soc. Trop. Med. Hyg. 109, 197–202 (2015). https://doi.org/10.1093/trstmh/tru166
Sudeep, A.B., Vyas, P.B., Parashar, D., Shil, P.: Differential susceptibility & replication potential of Vero E6, BHK-21, RD, A-549, C6/36 cells & Aedes aegypti mosquitoes to three strains of chikungunya virus. Indian J. Med. Res. 149, 771–777 (2019). https://doi.org/10.4103/ijmr.IJMR_453_17
Amin, P., et al.: Chikungunya: report from the task force on tropical diseases by the world Federation of Societies of intensive and critical care medicine. J. Crit. Care. (2018). https://doi.org/10.1016/j.jcrc.2018.04.004
Akahata, W., et al.: A virus-like particle vaccine for epidemic chikungunya virus protects nonhuman primates against infection. Nat. Med. 16, 334–338 (2010). https://doi.org/10.1038/nm.2105
Wu, J., Zhao, C., Liu, Q., Huang, W., Wang, Y.: Development and application of a bioluminescent imaging mouse model for chikungunya virus based on pseudovirus system. Vaccine. 35, 6387–6394 (2017). https://doi.org/10.1016/j.vaccine.2017.10.007
Chung, W.C., Hwang, K.Y., Kang, S.J., Kim, J.O., Song, M.J.: Development of a neutralization assay based on the pseudotyped chikungunya virus of a Korean isolate. J. Microbiol. 58, 46–53 (2020). https://doi.org/10.1007/s12275-020-9384-0
Tong, W., Yin, X.X., Lee, B.J., Li, Y.G.: Preparation of vesicular stomatitis virus pseudotype with chikungunya virus envelope protein. Acta Virol. 59, 189–193 (2015). https://doi.org/10.4149/av_2015_02_189
Theillet, G., et al.: Comparative study of chikungunya virus-like particles and Pseudotyped-particles used for serological detection of specific immunoglobulin M. Virology. 529, 195–204 (2019). https://doi.org/10.1016/j.virol.2019.01.027
Izumida, M., Hayashi, H., Tanaka, A., Kubo, Y.: Cathepsin B protease facilitates chikungunya virus envelope protein-mediated infection via endocytosis or macropinocytosis. Viruses. 12, 1 (2020). https://doi.org/10.3390/v12070722
Kummerer, B.M., Grywna, K., Glasker, S., Wieseler, J., Drosten, C.: Construction of an infectious chikungunya virus cDNA clone and stable insertion of mCherry reporter genes at two different sites. J. Gen. Virol. 93, 1991–1995 (2012). https://doi.org/10.1099/vir.0.043752-0
Sun, C., Gardner, C.L., Watson, A.M., Ryman, K.D., Klimstra, W.B.: Stable, high-level expression of reporter proteins from improved alphavirus expression vectors to track replication and dissemination during encephalitic and arthritogenic disease. J. Virol. 88, 2035–2046 (2014). https://doi.org/10.1128/JVI.02990-13
Zhang, H.L., et al.: Visualization of chikungunya virus infection in vitro and in vivo. Emerg Microbes Infect. 8, 1574–1583 (2019). https://doi.org/10.1080/22221751.2019.1682948
Hu, D., et al.: Chikungunya virus glycoproteins pseudotype with lentiviral vectors and reveal a broad spectrum of cellular tropism. PLoS One. 9, e110893 (2014). https://doi.org/10.1371/journal.pone.0110893
Tian, Y., et al.: Development of in vitro and in vivo neutralization assays based on the pseudotyped H7N9 virus. Sci. Rep. 8, 8484 (2018). https://doi.org/10.1038/s41598-018-26822-6
Kong, Y., Cirillo, J.D.: Fluorescence imaging of mycobacterial infection in live mice using fluorescent protein-expressing strains. Methods Mol. Biol. 1790, 75–85 (2018). https://doi.org/10.1007/978-1-4939-7860-1_6
Dhadve, A., Thakur, B., Ray, P.: Construction of dual modality optical reporter gene constructs for bioluminescent and fluorescent imaging. Methods Mol. Biol. 1790, 13–27 (2018). https://doi.org/10.1007/978-1-4939-7860-1_2
Fan, C., et al.: Beta-propiolactone inactivation of coxsackievirus A16 induces structural alteration and surface modification of viral capsids. J. Virol. 91, 1 (2017). https://doi.org/10.1128/JVI.00038-17
Bonnafous, P., et al.: Treatment of influenza virus with beta-propiolactone alters viral membrane fusion. Biochim. Biophys. Acta. 1838, 355–363 (2014). https://doi.org/10.1016/j.bbamem.2013.09.021
Theillet, G., et al.: Detection of chikungunya virus-specific IgM on laser-cut paper-based device using pseudo-particles as capture antigen. J. Med. Virol. 91, 899–910 (2019). https://doi.org/10.1002/jmv.25420
Madrid, P.B., et al.: A systematic screen of FDA-approved drugs for inhibitors of biological threat agents. PLoS One. 8, e60579 (2013). https://doi.org/10.1371/journal.pone.0060579
Zhang, X., et al.: Characterization of the inhibitory effect of an extract of Prunella vulgaris on Ebola virus glycoprotein (GP)-mediated virus entry and infection. Antivir. Res. 127, 20–31 (2016). https://doi.org/10.1016/j.antiviral.2016.01.001
von Rhein, C., et al.: Curcumin and Boswellia serrata gum resin extract inhibit chikungunya and vesicular stomatitis virus infections in vitro. Antivir. Res. 125, 51–57 (2016). https://doi.org/10.1016/j.antiviral.2015.11.007
Sanders, D.A.: No false start for novel pseudotyped vectors. Curr. Opin. Biotechnol. 13, 437–442 (2002). https://doi.org/10.1016/s0958-1669(02)00374-9
Steffen, I., Simmons, G.: Pseudotyping viral vectors with emerging virus envelope proteins. Curr. Gene Ther. 16, 47–55 (2016)
Bernard, E., et al.: Endocytosis of chikungunya virus into mammalian cells: role of clathrin and early endosomal compartments. PLoS One. 5, e11479 (2010). https://doi.org/10.1371/journal.pone.0011479
Li, Q., Liu, Q., Huang, W., Li, X., Wang, Y.: Current status on the development of pseudoviruses for enveloped viruses. Rev. Med. Virol. 28, 1 (2018). https://doi.org/10.1002/rmv.1963
Acknowledgments
We thank Liwen Bianji (Edanz) (https://www.liwenbianji.cn) for editing the language of a draft of this book chapter.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Wu, J., Huang, W., Wang, Y. (2023). Pseudotyped Viruses for the Alphavirus Chikungunya Virus. In: Wang, Y. (eds) Pseudotyped Viruses. Advances in Experimental Medicine and Biology, vol 1407. Springer, Singapore. https://doi.org/10.1007/978-981-99-0113-5_16
Download citation
DOI: https://doi.org/10.1007/978-981-99-0113-5_16
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-99-0112-8
Online ISBN: 978-981-99-0113-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)