Abstract
It is believed that Idiopathic Pulmonary Fibrosis (IPF) is an age-related chronic, progressive, and histopathologically associated fibrosing interstitial lung disorder which primarily affects the elderly. Despite tremendous progress in our knowledge of pathophysiology of diseases, we still do not know the possible causes of IPF. According to current research evidences, it is proposed that IPF may develop genotype as a result of repeated alveolar damage causing an abnormal wound-healing response. Genomic variations in epithelial integrity and host defence genes put people at risk for IPF, whereas immunosuppression and overt respiratory infection are supposed to have a high death rate. The role of infection in disease etiopathogenesis has long been suspected and its progression, or as a cause of acute aggravation, although preliminary investigations using classic culture procedures have formed inconsistent findings. Current approach of culture-independent microbiological analysis procedures to IPF patients has previously revealed various unacknowledged variations in lung microbiome and also a high microbial burden in bronchoalveolar lavage (BAL) in patients with IPF. However, connection does not always imply causation. Furthermore, lung microbiome is still incompletely defined, and more studies need to be done to explore species other than viruses and bacteria, such as fungus. The knowledge of microbiome’s role in aetiology and IPF progression might leads to its modification, allowing targeted therapeutic treatment.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Idiopathic Pulmonary Fibrosis (IPF)
- Interstitial Lung Disease (ILD)
- Microbiome
- Pathogenesis
- Acute Exacerbation
- Infection in Lung
13.1 Introduction
Microbiome refers as pathogenic and symbiotic organisms and a commensal’s ecological community which share human bodily space and also the intricate dealings of these microbes with the host. Various studies are conducted on the microbiome of gastrointestinal tract, with about 100 trillion microorganisms; yet, the lower respiratory tract’s epithelial surface is considered to be one of least inhabited areas of human body which has been supposed as sterile in past. Identifying and isolating microorganisms were challenging due to the difficulty of physically sampling lower airways and limitations of bacterial culture, leading to the incorrect idea. To better understand the respiratory tract’s microbiome, researchers switched from using culture-dependent methods to using methods independent of culture. High-throughput DNA sequencing methods use sequence similarity in extremely conserved genes like 16S ribosomal RNA gene to rapidly identify multifaceted bacterial communities (with species which cannot be grown) [1, 2]. Because of this, scientists are now studying lung microbiome in healthy volunteers and also patients with chronic respiratory diseases as cystic fibrosis, bronchiectasis, COPD and ILD. As a result of this investigation, researchers have discovered diverse communities of fungi, bacteria and viruses [3, 4].
IPF is a debilitating, severe, fibrosing and deadly fibrotic ILD that predominantly affects elderly people and eventually causing respiratory failure with the cause of chronic dyspnoea, an inevitable reduction in the functions of lung. It is a degenerative interstitial lung disease associated with ageing, via median diagnostic age of 66 years, and a serious condition via 2.5–.5 years of survival. The IPF factors are still unknown and yet, the particular or main causing factor has not been acknowledged, the disease is supposed to be induced by abnormality in wound-healing mechanisms in genetically susceptible individuals in response to unidentified environmental triggers (such as gastric micro-aspiration, viral infections, cigarette smoke, particulate dust, etc.) [5, 6].The resulting extracellular matrix deposition & development of fibroblastic foci reasons irreparable damages in lung architecture, resulting in alveolar structure loss, impeded gas exchange and eventually causing respiratory failure. Infectious agents, such as bacteria and viruses, can cause damage in alveolar epithelial cells and apoptosis, as well as alter the host’s response toward injury. Furthermore, researches involving genetic vulnerability to IPF have identified an amplified risk with genetic polymorphisms involved in characteristic host response control. A single nucleotide polymorphism in promoter region of the mucin 5B gene (MUC5B) (rs35705950), which codes for critical component of airway mucus, and single nucleotide polymorphism in the toll-interacting protein (TOLLIP) gene (rs5743890), which codes for adaptor protein that controls signaling through toll-like receptors (TLRs), are two specific examples [7, 8].
While comparing 130 IPF patients’ peripheral blood transcriptomes with controls, 4 genes involved in immune defence, including alpha-defensins, were found to be upregulated. These findings propose that innate immunological vulnerability can contribute a significant role in IPF aetiology, and supports hypothesis that infection, in combination with host immune system, contributes toward an abnormal fibrosis sequence of events. This review will be looking at what we currently know about the function of the respiratory microbiome in IPF, as well as extents of debate &further research objectives and priorities [9, 10].
13.2 Microbiome Development and Composition in Healthy Lungs
Initially thought to be sterile, epithelial surfaces of respiratory tract have been revealed to support dynamic microbial populations utilizing various culture-independent methods. Bacterial DNA was identified in 95.7% specimens of bronchoalveolar lavage (BAL) using high-throughput sequencing of bacterial 16s-rRNA, compared with 39.1% of BAL samples using conventional standard culture methods. Healthy lungs have bacterial communities that are quite similar to those observed in the mouth, but with a bacterial load that is two to four times lower. Previous studies have reported that there are approximately 10–100 bacterial cells per 1000 human cells in lung tissues. Interestingly despite changes in temperature, pH, & oxygen concentration, level of microbiome in healthy volunteers is quite consistent [11, 12]. Firmicutes (including genera Veillonella sp. and Streptococcus sp.), Bacteroidetes (including the species Prevotella sp.) to a slighter extent, Actinobacteria and Proteobacteria are the most commonly found phyla in normal airways.
The microbiota composition of the lungs is largely determined by three factors: microbial immigration, which is brought on by oro-nasal cavity mucosal dispersion, micro-aspiration of gastric contents and air inhalation; microbial elimination, which is caused by cough, mucociliary clearance and immunity; and the microbiological growth environment including oxygen tension, temperature, pH and nutrition availability [13,14,15].
The microbiota present in lungs reflects a stable condition among microbial inflow, outflow and reproduction level, and as a result of these three variables, with the latter being primarily impacted in the event of pathological processes of chronic diseases. The microbiome of lung is changed across every lung disease examined and compared to healthy volunteers. Many studies have found pollutants samples of upper respiratory tract during sampling due to the sensitivity of molecular technologies used, resulting in an inaccurate representation of the true microbiome. The risk of oropharyngeal contamination should be considered, as the majority of published research works have utilized BAL samples to describe the lung microbiome of healthy volunteers. Furthermore, heterogeneity of the microbial composition of lung at spatially distinct lung locations within subjects has been demonstrated in healthy participants, but this variation is smaller than inter-subject community variance [16,17,18]. Contamination has recently been shown to have a negligible impact on microbial plethora in bronchoscopy-acquired samples, supporting utility of bronchoscopy to study microbiome of lungs. Contamination can occur at any point during a microbiome study, not only during bronchoscopy [19].
While comparing data of microbiome from very identical subject specimen utilizing distinct sequencing channel and techniques, significant variance was observed. Other significant sources of contamination include agents and extraction kits and in low biomass samples they become important factor as those obtained from the respiratory system. Recall that BAL DNA sequencing provides “instantaneous” “snapshot” in time of bacterial diversity of lower airways, but does not assess chronically changing microbial communities over time. Several research works have focused on viruses and fungi in addition to the study of lung microbiota. A new study has found that commensal fungi have an effect on both the host immune system and bacteria in the gut. This has implications for the restoration of a healthy microbiome following antibiotic therapy. Because of the wide variety of viruses that can be found in the lungs, they are thought to be a catalyst for many different types of lung disease [20,21,22].
13.3 Microbiome in Idiopathic Pulmonary Fibrosis
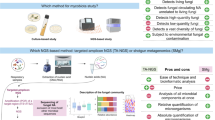
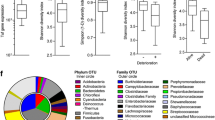
The lung microbiome: Previously considered to be sterile, the respiratory tract’s epithelial surfaces have been demonstrated to support dynamic microbial populations utilizing culture-independent methods. With biochemical sequence analysis of the factor 16s-rRNA genomic regions, bacterial species can now be recognized; in other microbiome scientific studies, groups of bacteria with similar genetic codes are categorized into operational taxonomic units (OTUs) and evaluated by comparing to the 16s rRNA data base. Higher microbial 16s-rRNA sequencing recognizes bacterial DNA in 95.7% of BAL specimens, compared to comparison to standard culture systems, that can locate bacteria in 39.1% of BAL samples. Using these genetic methods in characterizing microbial flora in respiratory tract of both sick population and healthy controls have shown correlations that imply the microbiome–host interaction that may be important in the aetiology and development of lung disease. Moreover, when severe asthma patients were compared with non-severe asthmatic patients and related controls, changes in the microbiome were observed, showing that the disease phenotype may be influenced by the microbial populations in the airway [23,24,25] (Fig. 13.1).
13.4 Microbiome Effect on IPF Prognosis and Exacerbation
Exacerbations are common in the IPF progression, as they are in a variety of chronic diseases of lungs. Acute episodes and exacerbations are linked to an especially bleak prognosis. Non-survivors exhibited shorter dyspnoea durations, higher C reactive protein (CRP) values, inspiratory oxygen fraction (FiO2) ratios/lower arterial oxygen tension (PaO2), lower proportions of lymphocytes and greater proportions of neutrophils in BALF than survivors. CRP was found to be only independently associated predictor of survival among those variables, ultimately suggesting that inflammation and/or bacterial or viral infection might be one of many pathogenic mechanisms involved in causing acute episodes and aggravations [26, 27].
“An acute, clinically significant deterioration of unidentifiable cause in a patient with underlying IPF” is currently a new definition of exacerbation and it necessitates the official prohibition of infection for clinical diagnosis. The specific aetiology of acute aggravations, however, is still unidentified, and it is uncertain whether it reflects an augmented phase of increased lung damage response or an underlying fibroproliferative process to an unknown previous or coexisting infection. Respiratory tract infections carry a mortality risk in persons with IPF, and is indistinguishable with acute aggravations is one of the factors suggesting an infection involvement in aggravation. Recent investigations involving lung microbiome during aggravations of IPF and its impact on progression of disease have also cast doubt on the definition. According to these research works, an enhanced bacterial load at time of diagnosis appears to be a biomarker for a disease that progresses more quickly and has a higher mortality risk [28, 29].
Another research including 20 patients with IPF diagnosed acute aggravations and 15 matched control subjects with constant IPF condition who undergone bronchoscopy & extraction of DNA process found that patients with IPF had a four-fold greater bacterial load during aggravations. In comparison to patients with stable IPF, their BALF included a greater number of neutrophils. This suggests the idea that bacteria have a significant role in exacerbations even if active infection is present. They are supposed to use 16S rRNA gene qPCR and pyrosequencing to investigate changes in BAL microbiota in both stable and acute exacerbation groups. There was noticeable alteration in microbiota in cases of acute exacerbation, along with a substantial increase in Stenotrophomonas sp. & Campylobacter sp., and a substantial decrease in Campylobacter sp. & Veillonella sp., despite being known best as gastrointestinal pathogen, was initially demonstrated in respiratory microbiota [30,31,32]. Its occurrence in respiratory microbiota is most probable due to stomach’s gastric contents silent micro-aspiration. To conclude these findings, this pilot study shows that IPF acute exacerbation may be due to be a significant role of various bacteria. Micro-aspiration may play a role in the apparent transfer of bacteria that are normally restricted to the gastrointestinal system. Although a prospective longitudinal research work is needed to validate the findings, they give a justification for clinical trials including prophylactic antibiotics as a method to avoid acute aggravations in IPF patients [33, 34].
13.5 A Gut–Lung Axis and Regulation of Host Defence in Chronic Lung Disease Aggravations: Evidence and Implications
The exact mechanism by which bacteria influence the initial immunity which present during the birth in healthy and sick is still being researched, and a few is revealed like microbiota modulates and regulates immunity of lung or the formation of lymphoid tissue associated with bronchial related. The importance of the gut commensal microbiota as a modulator of the innate immune system is being more recognized. The intestinal microbiota in healthy individuals is dominated by three phyla: Ruminococcus, Prevotella and Bacteroides. Evidences reveal that the formation of the intestinal microbiome is vital for control of an adequate immune response in lungs during a critical early period of life. Alteration in the composition of the microbiota of intestine impacts the progression and vulnerability of chronic lung diseases including asthma and cystic fibrosis. Moreover, the host is more vulnerable to lung infections, such as Klebsiella pneumoniae, Listeria monocytogenes and viruses, in the absence of normal intestinal biota. This offers the intriguing hypothesis that chronic lung disease exacerbations are caused by decreased adaptive and innate immune system as a result of changes in the intestinal microbiota of host [35,36,37]. As previously stated, individuals with progressive IPF have an enormous burden of Staphylococcus and Streptococcus species in their lungs, and earlier researches have shown that neutrophils from microbiota depleted mice had a decreased capacity to kill S. aureus and S. pneumoniae. Recently, it has been found that microbial stimulation of gut nod-like receptor sites causes an increase in the producing of free radicals in phagocytic cells—a lung’s sentinel innate immune cell. This suggests that circumstances related to loss of intestinal bacterial homeostasis (such as antibiotic use) may lead to weakened lung immunity. In COPD, viral infections can exacerbate symptoms, and the pathophysiology that follows could be linked to dysbiosis, which alters the microbiota of the airways and causes excessive inflammation [38, 39]. Despite the fact that damage of gastrointestinal commensal signaling might be responsible for impairing innate immunity of lung in this condition, cigarette smoke also makes a significant contribution to impeded lung innate immunity either directly or indirectly by altering innate immune cell phagocytosis, mucus, ciliary function and directly enhancing intestinal microbiota (e.g., enhanced formation of biofilm). These alterations may have an influence on respiratory infections’ propensity to aggravate COPD. Providing viruses’ proclivity for causing exacerbations in lung disease, it is worth considering the influence of respiratory viral infection on gut microbiota. According to Wang and associates, influenza infection can cause abnormalities in the gut microbiota, including an increase in Enterobacteriaceae and decrease in Lactococcus & Lactobacillus. As stated earlier, this might result in a reduction of beneficial bacteria, which could contribute to smoking-related illness. According to the scientists, these changes in gut microbiome were not caused by lytic influenza intestinal infection [40, 41]. Th17 cells were involved in the damage, and neutralization of IL-17 reduced the severity of the injury. In addition, reduction of the intestinal flora caused by antibiotics resulted in less intestinal damage. The relevance of effector T cell developed in the lung after infection and subsequently moved to small intestine, produced IFN-g and modified the gut microbiota, was also highlighted in the research work by scientists. Finally, Th17 response was aided via triggered epithelial-derived IL-15 due to changes in the gut microbiota. It is conceivable that the responses of IL-17 in intestine contribute to the progression of lung diseases. Certain microorganisms are eradicated by IL-17, which has been linked to the pathophysiology of sarcoidosis, asthma, cystic fibrosis, necrotizing bronchial asthma and bone marrow transplant-related pneumonitis [42, 43] (Fig. 13.2).
IL-17 might potentially have an impact in the dynamic changes that occur in pulmonary microbiome in COPD patients. In emphysema animal model, Yadava & their colleagues studied the effects of experimental changes on the lung microbiome. LPS/elastase was given to pathogen-free & axenic mice for 4 weeks. Through an excess of Lactobacillus, Pseudomonas and a reduction of Prevotella, microbiota diversity & abundance were reduced in LPS/elastase model. The loss of bacterial load was linked to a reduction in IL-17 production. Axenic mice were given microbiota-enriched fluid intranasally, which increased IL-17 production. In mice with microbiota, inhibition of IL-17 resulted in decreased inflammation and disease load. IL-17 has been linked to hepatic fibrosis in several investigations, and several experimental models of pulmonary fibrosis are IL-17A–dependent [44, 45]. In addition, research works looking into the onset of intestinal fibrosis have found a link between changes in the microbiome and Th17 responses. Through the adhesion of segmented filamentous bacteria on intestinal epithelial cells, gut is a recognized source of Th17 cells. In lungs, the situation may be identical. In animal models, Gauguet and his colleagues revealed that intestine segmented filamentous bacteria can enhance pulmonary innate immunity by inducing IL-17, resulting in resistance to S. aureus pneumonia. This adds to the growing body of data that suggests the gut–lung microbiome axis is important in regulating the lung’s innate immune response [46, 47].
13.6 Limitation
Han and colleagues were limited to naming the progression-related bacteria Staphylococcus OTU 1348 & Streptococcus OTU 1345, as 16S rRNA sequencing can never be utilized for genetic markers. More research is required to fully characterize these bacteria, either in form of microbe-specific sequencing or sequencing specific to a particular culture. Despite the fact that the cohort contained multiple Streptococcus and Staphylococcus species, only two OTUs were linked to progression of diseases. In any disease related to lung, there are certain general limits to microbiome research. Particularly in the context of the molecular technologies used, infection of samples from upper respiratory tract during taking specimen is an apparent problem in many research, yielding a misleading depiction of the real microbiome [48, 49]. Kits including reagents and extraction agents can potentially be a source of contamination, which is especially critical when working with small biomass samples like those from respiratory system. Contamination can occur at any point during a microbiome study, not particularly during the bronchoscopy stage. While corelating the data of microbiome from same patient samples using various sequencing techniques and platforms, significant variance was observed. There are a number of biases in primer design that can favour or penalize certain bacteria, resulting in the exclusion of entire genera [50, 51].
While studies on the IPF microbiome have used high-throughput molecular technologies to identify bacterial species and loads, they have not yet demonstrated a causal, mechanistic link to the disease process and advancement. In the investigational studies of IPF, it is not clear if changes to the lungs’ microbiome reflect the disease’s aetiology or are due to a lack of underpinning immune defences in this patient population. The information gleaned from this research is probable to be less significant and more irrelevant because it does not reveal how the various bacterial colonies interact with one another [52,53,54].
This study provides a “snapshot” of the lower respiratory microbiome by DNA sequencing from a BAL sample, but it does not look at longitudinal modifications. Because it is unrealistic to perform bronchoscopies on a regular basis, other approaches of tracking the lower airway microbiome over time should be explored. BAL taken from one lobe of lung may not be representative of microbiota in the other lobes, especially because histological hallmark of IPF and UIP shows spatial variability through fibrosis alongside typical parenchyma. This is the case for IPF and UIP. Ex-planted lung tissue sections via cystic fibrosis patient were sequenced using 16s rDNA to uncover variances in microbial communities within lungs [55, 56]. As per our consideration of IPF microbiome improves, sample & sequencing techniques improve and composition of patient’s microbiome might serve as a biomarker to help with prognosis & therapy stratification. A key question for future IPF research is whether or not prophylactic antibiotics should be used to target specific microbiome “signatures” in patients in order to improve survival, based on the results of a trial testing co-trimoxazole in patients with IPF [57, 58].
13.7 Conclusion
As per various studies and researches, alteration in microbiome load, composition and diversity have been linked to aetiology of disease, acute exacerbation, progression and death in idiopathic pulmonary fibrosis. Lung microbiome dysbiosis will be linked to IPF development and progression, according to the study’s findings. When it comes to IPF, microbiome manipulation could soon be a treatment modality to restore a “healthy” microbiome culture. However, a comprehensive method to account for various factors driving development of disease, advancement, & episodic exacerbating is more probable. It is unclear if antibiotics, probiotics (extrinsic microorganisms given for health purposes) or prebiotics (molecules that encourage growth and development of specific bacteria) should be used to control the lung microbiome. However, modification of microbiome should focus on pathogenic microbes while leaving the rest of the microbial population intact but that would be considerably more difficult to achieve. All of these studies suggest that anti-biotherapy may have a significant role in IPF patients, and they establish a justification for long-term anti-biotherapy related clinical trials, that acts as a modulator of immunity and anti-bioprophyl axis to avoid acute exacerbations. Future research on lung microbiome dynamics could aid in the selection of suitable, targeted & more customized anti-biotherapy over the course of disease, particularly in cases of IPF aggravations. These studies require more advanced metagenomic techniques to determine functional relevance of particular microbial species & populations in the development of IPF.
References
Anand S, Mande SS (2018) Diet, microbiota and gut-lung connection. Front Microbiol 9:2147
Bacci G, Taccetti G, Dolce D, Armanini F, Segata N, Di Cesare F et al (2020) Untargeted metagenomic investigation of the airway microbiome of cystic fibrosis patients with moderate-severe lung disease. Microorganisms 8(7):1003
Beck JM, Young VB, Huffnagle GB (2012) The microbiome of the lung. Transl Res 160(4):258–266
Bhagirath AY, Li Y, Somayajula D, Dadashi M, Badr S, Duan K (2016) Cystic fibrosis lung environment and Pseudomonas aeruginosa infection. BMC Pulm Med 16(1):174
Blanchard AC, Waters VJ (2019) Microbiology of cystic fibrosis airway disease. Semin Respir Crit Care Med 40(6):727–736
Budden KF, Shukla SD, Rehman SF, Bowerman KL, Keely S, Hugenholtz P et al (2019) Functional effects of the microbiota in chronic respiratory disease. Lancet Respir Med 7(10):907–920
Carney SM, Clemente JC, Cox MJ, Dickson RP, Huang YJ, Kitsios GD et al (2020) Methods in lung microbiome research. Am J Respir Cell Mol Biol 62(3):283–299
Caverly LJ, Zhao J, LiPuma JJ (2015) Cystic fibrosis lung microbiome: opportunities to reconsider management of airway infection. Pediatr Pulmonol 50(Suppl 40):S31–S38
Chmiel JF, Aksamit TR, Chotirmall SH, Dasenbrook EC, Elborn JS, LiPuma JJ et al (2014) Antibiotic management of lung infections in cystic fibrosis. I. the microbiome, methicillin-resistant Staphylococcus aureus, gram-negative bacteria, and multiple infections. Ann Am Thorac Soc 11(7):1120–1129
Chunxi L, Haiyue L, Yanxia L, Jianbing P, Jin S (2020) The gut microbiota and respiratory diseases: new evidence. J Immunol Res 2020:2340670
Cribbs SK, Beck JM (2017) Microbiome in the pathogenesis of cystic fibrosis and lung transplant-related disease. Transl Res 179:84–96
Cuthbertson L, Walker AW, Oliver AE, Rogers GB, Rivett DW, Hampton TH et al (2020) Lung function and microbiota diversity in cystic fibrosis. Microbiome. 8(1):45
de Almeida OGG, Capizzani C, Tonani L, Grizante Barião PH, da Cunha AF, De Martinis ECP et al (2020) The lung microbiome of three Young Brazilian patients with cystic fibrosis colonized by fungi. Front Cell Infect Microbiol 10:598938
Dickson RP (2016) The microbiome and critical illness. Lancet Respir Med 4(1):59–72
Dmitrijeva M, Kahlert CR, Feigelman R, Kleiner RL, Nolte O, Albrich WC et al (2021) Strain-resolved dynamics of the lung microbiome in patients with cystic fibrosis. MBio 12(2):e02863–e02820
Drakopanagiotakis F, Wujak L, Wygrecka M, Markart P (2018) Biomarkers in idiopathic pulmonary fibrosis. Matrix Biol 68-69:404–421
Fastrès A, Felice F, Roels E, Moermans C, Corhay JL, Bureau F et al (2017) The lung microbiome in idiopathic pulmonary fibrosis: a promising approach for targeted therapies. Int J Mol Sci 18(12):2735
Flume PA, Chalmers JD, Olivier KN (2018) Advances in bronchiectasis: endotyping, genetics, microbiome, and disease heterogeneity. Lancet 392(10150):880–890
Françoise A, Héry-Arnaud G (2020) The microbiome in cystic fibrosis pulmonary disease. Genes 11(5):536
Han MK, Huang YJ, Lipuma JJ, Boushey HA, Boucher RC, Cookson WO et al (2012) Significance of the microbiome in obstructive lung disease. Thorax 67(5):456–463
Han MK, Zhou Y, Murray S, Tayob N, Noth I, Lama VN et al (2014) Lung microbiome and disease progression in idiopathic pulmonary fibrosis: an analysis of the COMET study. Lancet Respir Med 2(7):548–556
Heirali AA, Acosta N, Storey DG, Workentine ML, Somayaji R, Laforest-Lapointe I et al (2019) The effects of cycled inhaled aztreonam on the cystic fibrosis (CF) lung microbiome. J Cyst Fibros 18(6):829–837
Héry-Arnaud G, Boutin S, Cuthbertson L, Elborn SJ, Tunney MM (2019) The lung and gut microbiome: what has to be taken into consideration for cystic fibrosis? J Cyst Fibros 18(1):13–21
Huang YJ, LiPuma JJ (2016) The microbiome in cystic fibrosis. Clin Chest Med 37(1):59–67
Invernizzi R, Wu BG, Barnett J, Ghai P, Kingston S, Hewitt RJ et al (2021) The respiratory microbiome in chronic hypersensitivity pneumonitis is distinct from that of idiopathic pulmonary fibrosis. Am J Respir Crit Care Med 203(3):339–347
Kanda T, Goto T, Hirotsu Y, Masuzaki R, Moriyama M, Omata M (2020) Molecular mechanisms: connections between nonalcoholic fatty liver disease, steatohepatitis and hepatocellular carcinoma. Int J Mol Sci 21(4):1525
Kitsios GD, Rojas M, Kass DJ, Fitch A, Sembrat JC, Qin S et al (2018) Microbiome in lung explants of idiopathic pulmonary fibrosis: a case-control study in patients with end-stage fibrosis. Thorax 73(5):481–484
Lipinski JH, Moore BB, O’Dwyer DN (2020) The evolving role of the lung microbiome in pulmonary fibrosis. Am J Physiol Lung Cell Mol Physiol 319(4):L675–Ll82
Lira-Lucio JA, Falfán-Valencia R, Ramírez-Venegas A, Buendía-Roldán I, Rojas-Serrano J, Mejía M et al (2020) Lung microbiome participation in local immune response regulation in respiratory diseases. Microorganisms. 8(7):1059
Lynch SV (2016) The lung microbiome and airway disease. Ann Am Thorac Soc 13 Suppl 2(Suppl 5):S462–S4s5
Mammen MJ, Scannapieco FA, Sethi S (2020) Oral-lung microbiome interactions in lung diseases. Periodontology 2000 83(1):234–241
Mendez R, Banerjee S, Bhattacharya SK, Banerjee S (2019) Lung inflammation and disease: a perspective on microbial homeostasis and metabolism. IUBMB Life 71(2):152–165
Metwally AA, Ascoli C, Turturice B, Rani A, Ranjan R, Chen Y et al (2020) Pediatric lung transplantation: dynamics of the microbiome and bronchiolitis obliterans in cystic fibrosis. J Heart Lung Transplant 39(8):824–834
Moffatt MF, Cookson WO (2017) The lung microbiome in health and disease. Clin Med (Lond) 17(6):525–529
Monsó E (2020) Look at the wood and not at the tree: the microbiome in chronic obstructive lung disease and cystic fibrosis. Arch Bronconeumol 56(1):5–6
Morris A, Gibson K, Collman RG (2014) The lung microbiome in idiopathic pulmonary fibrosis What does it mean and what should we do about it? Am J Respir Crit Care Med 190(8):850–852
Morton JT, Aksenov AA, Nothias LF, Foulds JR, Quinn RA, Badri MH et al (2019) Learning representations of microbe-metabolite interactions. Nat Methods 16(12):1306–1314
Muhlebach MS, Zorn BT, Esther CR, Hatch JE, Murray CP, Turkovic L et al (2018) Initial acquisition and succession of the cystic fibrosis lung microbiome is associated with disease progression in infants and preschool children. PLoS Pathog 14(1):e1006798
Mur LA, Huws SA, Cameron SJ, Lewis PD, Lewis KE (2018) Lung cancer: a new frontier for microbiome research and clinical translation. Ecancermedicalscience 12:866
O’Dwyer DN, Ashley SL, Gurczynski SJ, Xia M, Wilke C, Falkowski NR et al (2019) Lung microbiota contribute to pulmonary inflammation and disease progression in pulmonary fibrosis. Am J Respir Crit Care Med 199(9):1127–1138
O’Dwyer DN, Garantziotis S (2021) The lung microbiome in health, hypersensitivity pneumonitis, and idiopathic pulmonary fibrosis: a heavy bacterial burden to bear. Am J Respir Crit Care Med 203(3):281–283
Paudel KR, Dharwal V, Patel VK, Galvao I, Wadhwa R, Malyla V et al (2020) Role of lung microbiome in innate immune response associated with chronic lung diseases. Front Med 7:554
Permall DL, Pasha AB, Chen XQ, Lu HY (2019) The lung microbiome in neonates. Turk J Pediatr 61(6):821–830
Quinn RA, Adem S, Mills RH, Comstock W, DeRight GL, Humphrey G et al (2019) Neutrophilic proteolysis in the cystic fibrosis lung correlates with a pathogenic microbiome. Microbiome 7(1):23
Ramsey KA, Schultz A, Stick SM (2015) Biomarkers in paediatric cystic fibrosis lung disease. Paediatr Respir Rev 16(4):213–218
Rogers GB, Bruce KD, Hoffman LR (2017) How can the cystic fibrosis respiratory microbiome influence our clinical decision-making? Curr Opin Pulm Med 23(6):536–543
Saint-Criq V, Lugo-Villarino G, Thomas M (2021) Dysbiosis, malnutrition and enhanced gut-lung axis contribute to age-related respiratory diseases. Ageing Res Rev 66:101235
Salisbury ML, Han MK, Dickson RP, Molyneaux PL (2017) Microbiome in interstitial lung disease: from pathogenesis to treatment target. Curr Opin Pulm Med 23(5):404–410
Scialo F, Amato F, Cernera G, Gelzo M, Zarrilli F, Comegna M et al (2021) Lung microbiome in cystic fibrosis. Life. 11(2):94
Shukla SD, Budden KF, Neal R, Hansbro PM (2017) Microbiome effects on immunity, health and disease in the lung. Clin Transl Immunol 6(3):e133
Spagnolo P, Molyneaux PL, Bernardinello N, Cocconcelli E, Biondini D, Fracasso F et al (2019) The role of the Lung’s microbiome in the pathogenesis and progression of idiopathic pulmonary fibrosis. Int J Mol Sci 20(22):5618
Takahashi Y, Saito A, Chiba H, Kuronuma K, Ikeda K, Kobayashi T et al (2018) Impaired diversity of the lung microbiome predicts progression of idiopathic pulmonary fibrosis. Respir Res 19(1):34
Tan JY, Tang YC, Huang J (2020) Gut microbiota and lung injury. Adv Exp Med Biol 1238:55–72
Tojo R, Suárez A, Clemente MG, de los Reyes-Gavilán CG, Margolles A, Gueimonde M et al (2014) Intestinal microbiota in health and disease: role of bifidobacteria in gut homeostasis. World J Gastroenterol 20(41):15163–15176
Tong X, Su F, Xu X, Xu H, Yang T, Xu Q et al (2019) Alterations to the lung microbiome in idiopathic pulmonary fibrosis patients. Front Cell Infect Microbiol 9:149
Torrisi SE, Kahn N, Vancheri C, Kreuter M (2020) Evolution and treatment of idiopathic pulmonary fibrosis. Presse Med. 49(2):104025
Tracy M, Cogen J, Hoffman LR (2015) The pediatric microbiome and the lung. Curr Opin Pediatr 27(3):348–355
Trivedi R, Barve K (2020) Gut microbiome a promising target for management of respiratory diseases. Biochem J 477(14):2679–2696
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive licence to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Pathak, S. et al. (2022). Microbiome in Idiopathic Pulmonary Fibrosis. In: Gupta, G., Oliver, B.G., Dua, K., Singh, A., MacLoughlin, R. (eds) Microbiome in Inflammatory Lung Diseases. Springer, Singapore. https://doi.org/10.1007/978-981-16-8957-4_13
Download citation
DOI: https://doi.org/10.1007/978-981-16-8957-4_13
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-8956-7
Online ISBN: 978-981-16-8957-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)