Abstract
CRISPR/Cas has emerged as a game-changing technology for genome editing with widespread applications ranging from human therapeutics to engineering bacterial genomes for beneficial purposes to editing plant genomes for agricultural purposes. Successful genome editing through CRISPR/Cas relies on two components: an appropriate Cas endonuclease and a 20-base-pair (bp), single-guide RNA (sgRNA). CRISPR/Cas is currently favored as a genome editing technique due to its simple design rules and efficient editing capabilities that do not necessarily involve adding any foreign DNA at the target site. Cas endonucleases can be programmed to target any site in the genome by changing the gRNA sequence, highlighting the importance of gRNA design for increased specificity and efficiency, and reduced off-targeting in CRISPR/Cas genome editing. The rapid rise in CRISPR/Cas genome editing and associated applications has led to the development of numerous computational tools for effective sgRNA design. In this chapter, we discuss the essentials of gRNA design and provide an overview of the design process. In addition to summarizing factors which affect gRNA specificity and CRISPR cleavage efficiency, we discuss predictions of target efficiency and off-target detection algorithms. Finally, we describe the application-specific (knockout, activation, repression, base editing, and RNA editing) requirements of gRNA design and different tools to facilitate gRNA design.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- CRISPR/Cas
- Design tools
- On-targeting efficiency
- Off-targeting
- Bioinformatic tools
- Specificity of CRISPR/Cas
3.1 Introduction
CRISPR/Cas is an adaptive immune system of archaea and bacteria, providing a defense against invading plasmids and viruses (Garneau et al. 2010). Natural CRISPR/Cas systems consist of three core components:
-
An array of repeats encompassing unique sequences called spacers
-
A promoter sequence upstream of CRISPR arrays
-
An operon encoding a set of effector Cas proteins, essential for processing information coded within arrays
Native CRISPR/Cas defense systems consist of three stages: adaptation or acquisition, expression or biogenesis, and interference. During acquisition, a foreign genetic element (a “protospacer”) is cleaved and incorporated into the CRISPR locus as a new spacer. In biogenesis, these arrays are expressed as precursor CRISPR RNA (pre-crRNA) and subsequently processed into mature crRNA. Finally, in the interference stage, Cas endonucleases cleave the invading double-stranded DNA using crRNA as a guide sequence (Brazelton et al. 2015). Multiple studies have confirmed that the adoption and interference stages also require a protospacer adjacent motif (PAM) in the immediate vicinity of the protospacer (Fig. 3.1).
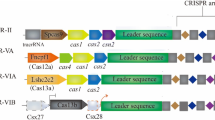
Based on effector Cas protein organization and non-coding RNA species architecture, CRISPR/Cas systems have been classified into two main classes and six types (Lino et al. 2018). Class 1 systems are defined as multi-Cas proteins acting in a cascade manner or Cas module-RAMP (repeat-associated mysterious proteins), i.e., Cmr complexes. In contrast, class 2 systems are compact and utilize a single effector Cas protein. For detailed classification of CRISPR/Cas systems, see Chap. 2. Due to their compact architecture and single effector Cas protein, class 2 systems have been adopted for genome editing applications in eukaryotes (Jinek et al. 2013; Makarova and Koonin 2015; Mali et al. 2013). Cas9 from Streptococcus pyogenes (SpCas9) requires a non-coding RNA known as transactivating crRNA (tracrRNA) in addition to crRNA. In today’s genome editing applications, these two non-coding RNAs are synthetically fused into one sgRNA (Alkhnbashi et al. 2020). So, an sgRNA in an engineered CRISPR/Cas9 system consists of a permanent part and a programmable part. The programmable part can be tailored to target Cas9 anywhere in the genome. The target site in DNA consists of a 20-nucleotide (nt)-long region complementary to sgRNA plus a PAM sequence (NGG for SpCas9 and TTTV for Cpf1) (Table 3.1). If there is no PAM adjacent to the target site, Cas endonuclease will not cleave the target site. If the sgRNA pairs with the DNA target sequence followed by PAM, it could create a double-stranded break (DSB) in the target site. The DSB will be repaired by either non-homologous end joining (NHEJ) or homology-directed repair (HDR) (Tian et al. 2017) (Fig. 3.2). The sgRNAs are not selected randomly; they must be associated with a PAM that is present in the target DNA but not included in the sgRNA. Bacteria use PAM to differentiate between self and non-self, thereby protecting their own DNA from cleavage because PAMs are only present in phage DNA (Fig. 3.3). With this simple and straightforward design, CRISPR/Cas can be programmed to any sequence in the genome. However, this simple, two-component (sgRNA and PAM) process also has disadvantages, as exactly similar or closely similar sgRNA sequences may occur at multiple locations and some of them could be tolerated by Cas endonuclease, leading to so-called off-targets (Cui et al. 2018). Cas endonuclease may also tolerate specific sequence changes in PAM. For example, while spCas9 specifically recognizes NGG (where N is any nucleotide base; G is guanine), it may also recognize NAG (where N is any nucleotide base; A is adenine; G is guanine), albeit less efficiently (Thomas et al. 2019). It is critical to reduce the number of potential off-target sites for improved CRISPR/Cas specificity, especially in human therapeutic applications, germline modifications, and genome editing for important agricultural purposes.
The rapid rise in CRISPR/Cas applications has prompted researchers to devise bioinformatic tools using different algorithms and design rules for effective sgRNA design, specific targeted modification, and low off-targets. Such tools facilitate gRNA design with maximum on-target efficiency in available genomes with user-defined PAM sequence and Cas endonuclease (Cui et al. 2018). Many design tools exist, but all have their own individual strengths and limitations. Most vary in terms of design parameters, specifications, available genomes, on-target efficiency score, off-target predictions, and so on. For example, design tools such as CRISPR-P (Li and Durbin 2009), E-CRISPR (Heigwer et al. 2014), CasOT (Xiao et al. 2014), and Cas-OFFinder (Bae et al. 2014) were mainly developed to predict off-targets in CRISPR/Cas experiments. However, in CRISPR/Cas applications such as CRISPR screening, cleavage efficiency is also important (Ma et al. 2016). Therefore, design tools such as sgRNA Designer, CRISPR-ERA (Liu et al. 2015), and Benchling predict on-targets as well as off-targets. Other genomic features such as sgRNA guanine-cytosine (GC) content, PAM flanking sequences, chromatin structure, methylation status, regulatory potential, and evolutionary conservation are also important in sgRNA design (Shi et al. 2015). Another critical factor in designing an efficient sgRNA is the application-specific (knockout (KO), knock-in (KI), CRISPR interference (CRISPRi), CRISPR activation (CRISPRa), and base editing) location of sgRNA in the genome. “WeReview: CRISPR Tools” is an online, live repository which helps researchers choose the best and latest tools for CRISPR/Cas applications (Torres-Perez et al. 2019). The current chapter aims to help researchers select the most useful tools for sgRNA design with maximum specificity and limited off-targets. This chapter also seeks to help users who are designing sgRNA with application-specific parameters in CRISPR/Cas.
3.2 Fundamentals of CRISPR/Cas Experiment and sgRNA Design
Engineered CRISPR/Cas system relies on sgRNA and PAM for genome modification in the target site of the genome. The prerequisites for designing an efficient sgRNA are:
-
1.
Target gene and target region
-
2.
Specific Cas endonuclease (e.g., Cas9, Cas9 nickase (nCas9), nuclease-dead Cas9 (dCas9), Cpf1) and an appropriate PAM for the Cas endonuclease
-
3.
Promoter selection for in vivo or in vitro expression of sgRNA
-
4.
Cloning strategy for sgRNA, e.g., sgRNA cloned in expression vector or used as template for RNA production
-
5.
For multiple gRNA, whether expressed from a single promoter or individual promoters
Also important for sgRNA design are application-specific parameters (e.g., for KO, KI, CRISPRi, CRISPRa, and base editing) coupled with the intended DSB repair system. For example, in KO applications, off-targets on other chromosomes may be cleared by backcrossing. Moreover, the sgRNA position for CRISPRi and CRISPRa applications would be different to that for KO and KI applications. In addition, two or more sgRNAs are required in some applications, such as two sgRNAs with nCas9, a pair of sgRNAs in CRISPRa, and a pair of distal sgRNAs in KI applications (Mohr et al. 2016). Here we summarize the essentials of an effective sgRNA for different CRISPR/Cas systems.
3.2.1 Good Gene Annotation: An Essential Requirement
From a genome editing perspective, good gene annotation is a prerequisite for designing an appropriate sgRNA. Online databases and tools are available to help designers view sgRNA in a relevant genome browser, as successful editing in most CRISPR/Cas applications depends upon gRNA positioning relative to specific features of the gene. For example, in CRISPRa, the sgRNA must be located within 50–500 bp of the transcription start site (TSS), but in CRISPRi, the gRNA should be near TSS. For KO applications using NHEJ, appropriate target regions may include a common coding exon, while in KI, a specific coding exon, intron, or a region coding for a protein domain could be appropriate (Gilbert et al. 2014; Shalem et al. 2014; Wang et al. 2014; Shi et al. 2015). High-quality genome databases with regularly updated gene annotations based on experimental data are available for models such as drosophila, zebrafish, mouse, rat, and Arabidopsis. These databases assist in formed design of gRNA relative to the position of gene features. However, in non-model species, the lack of genome databases with appropriate gene annotations is a limiting factor on the design of specific gRNA (Mohr et al. 2016).
3.2.2 Different Guidelines for Different Applications
With rapid development in CRISPR/Cas systems has come the development of bioinformatic tools and algorithms to predict on-target efficiency, as well as off-targets. Off-target tools mostly focus on sequence similarity with on-target sites and use a defined cut-off for possible number of mismatches that can be tolerated. However, even for off-target sites with mismatches, creating a bulge or gap sometimes leads to a valid target site for a DSB. Although several tools can predict off-targets, it is not feasible to apply those rules for every gRNA and every application. Some rules for gRNA effectiveness are not relevant to all CRISPR/Cas applications or even the same application in different species (Mohr et al. 2016). For example, a CRISPRi application in Escherichia coli showed that gRNA must target the non-template strand (also called the coding strand or sense strand) (Qi et al. 2013), but similar studies in eukaryotes showed that gRNA binding to either strand is effective. Moreover, as compared with KO applications, off-target effects will be of less concern in CRISPRi and CRISPRa applications, because binding may not be within effective range of the promoter sequence (Mohr et al. 2016). A recent study showed that sgRNA effectiveness parameters for cleavage efficiency in CRISPRi were not valid for CRISPRa applications (Doench et al. 2016). This suggests that different applications require different design principles. However, it is not yet clear to what extent general design rules are relevant to various applications or to what extent optional parameters will be required for a particular species, tissue, or cell.
3.2.3 Best Design Linked with Availability of More Data
Improvements in CRISPR/Cas design require more data to be available. When designing sgRNA, researchers must be aware of the design tool’s criteria for maximizing specificity and limiting off-targets. Researchers must also know the background of the design criteria: the study, species, delivery method, and specific applications from which a particular parameter was derived (Mohr et al. 2016). Sharing results and data from good designs and poor ones, along with species information and specific applications, will help researchers to continue improving the design and efficiency of CRISPR/Cas systems. In addition, information and data sharing will help researchers better understand the universal and application-specific factors that influence the effective design of sgRNA.
3.3 sgRNA Design Process: An Overview
The key aspect of sgRNA design is to define the target site in the genome. This can easily be done by locating the PAM sequence (NGG for spCas9 and TTTV for Cpf1) in the target region or gene. All PAM sequences recognized by different Cas endonucleases are listed in Table 3.1. Theoretically, if 5′-20 nt of the sgRNA pairs with a complementary target site in the genome, the sgRNA/Cas9 complex will create a DSB. However, several practical studies have suggested that cleavage efficiency varies significantly among different gRNAs. So, predictive models and algorithms are essential for selecting the best high-efficiency gRNA with limited off-targets. An additional challenge in CRISPR applications is off-target activity caused by both sgRNA and Cas9. Several studies have confirmed that CRISPR/Cas9 can tolerate several mismatches and cleave the DNA at sites other than the intended site of modification (target site) leading to off-target mutations. Although spCas9 systems recognize 5′-NGG-3′ as PAM, spCas9 can also recognize 5′-NAG-3′ and 5′-NGA-3′ albeit with low efficiency. Many models and computational tools are available to help researchers design an effective gRNA with high efficiency and specificity (Cui et al. 2018). In the following section, we present an overview of the design process in CRISPR/Cas applications.
3.3.1 Selection of Desired Genetic Modification
The first step in the design process is to define the desired genetic modification, e.g., KO, point mutation, transcriptional control, or KI. Because different modifications require different CRISPR/Cas reagents, a clear understanding of the desired genetic manipulation will narrow down the selection of appropriate CRISPR/Cas components (Thomas et al. 2019). However, although a broad range of CRISPR reagents and components exist, it is better to customize these components if perfect reagents do not exist for the chosen application.
3.3.2 Choice of Appropriate Expression System
To achieve the desired objective in a CRISPR/Cas experiment, Cas9 and gRNA must be expressed in the target cells or organism. Factors that can affect the desired modification, off-target numbers, and efficiency include the selected expression system (transient or stable), promoter choice (constitutive or tissue specific), reagents (plasmid, mRNA or RNPs), and delivery systems (viral, non-viral, or physical) (Graham and Root 2015). Standard protocols and reagents may suffice for CRISPR/Cas applications in easy-to-transfect cell lines, e.g., HEK293 (Banan 2020).
3.3.3 Selection of Appropriate Cas Endonuclease
Of the two classes of CRISPR/Cas systems described above, Class 1 systems use multiple Cas proteins, while Class 2 use a single effector Cas protein to create DSB in the target DNA. Choosing the right Cas endonuclease is essential. Cas9 and Cpf1 (Cas12a), the two most widely used Cas endonucleases, are both Class 2 CRISPR/Cas systems. Cas9 is a type II endonuclease that recognizes NGG as PAM sequence and creates DSB with blunt ends, three bp upstream of PAM site. Multiple engineered Cas9 variants have been generated, for example, nCas9, which produces single-stranded breaks (SSB), while dCas9 is used for site-specific binding of DNA. In contrast, Cpf1 is a type V endonuclease that recognizes the TTTN PAM sequence. Cpf1 cleaves 18–23 bp away from PAM and produces staggered ends with 5′ overhangs. Because it is smaller than spCas9, it is easy to pack into viral vectors for delivery. So, selection of expression system depends upon the desired modifications (Luo 2019).
3.3.4 Selection of Gene or Genetic Element
To manipulate a gene with a particular CRISPR application, a researcher must first identify the target gene’s genomic sequence. Selection of target region (promoter, exons, or introns) in the gene will depend upon the desired genetic modification. For example, for KO applications, 5′ constitutive expressed exon is the best target. Alternatively, gRNA can be targeted to an exon that codes an essential protein domain. For HDR applications, the target sequence should be in close proximity (within 10 bp) to the desired edit site.
3.3.5 Searching of Target Site for Intended Gene Modification
Most CRISPR/Cas design tools search target regions using either a sequence-based or a genome-based approach. In sequence-based searching, the user must input the sequence to define the target site for gRNA design. The CRISPOR design tool searches on sequence and requires an input of <2000 nt for gRNA design and display. In a genome-based approach, the user must provide a gene name, ID, or similar input to display gRNA relative to the gene features. For example, the WGE (Wellcome Sanger Institute genome editing) tool requires a gene symbol in order to display sgRNA relative to the gene features (Thomas et al. 2019).
3.3.6 Sequencing of Target Site and Design of sgRNA
Once the desired manipulation, expression system, Cas endonuclease, and CRISPR reagents are decided, the next step is to confirm the site and design sgRNA. SgRNA design is a prime concern in CRISPR applications. Because features in the target DNA site affect the sgRNA efficiency, therefore, it is better to sequence the target region before designing gRNA, because variations in the target region and gRNA may occur and this can reduce cleavage efficiency. Most CRISPR/Cas applications require an efficient and specific sgRNA, but this task is quite challenging because there are many criteria to obey. So, to identify the most suitable gRNA with maximum efficiency, design criteria are very important. Various sequence features influence the efficiency of gRNA. For example, the presence of guanine (G) at 5′ end of sgRNA (GX19NGG) was crucial for expression from U6 promoter. G was also required on the first or second position adjacent to PAM, probably for loading of Cas9. The presence of cytosine (C) at this position was not favored. Thymine (T) at the fourth position closest to PAM is undesirable too, because the presence of multiple uracil (U) decreases sgRNA expression. Adenine (A) is suitable in the middle region of gRNA; G is preferred in the distal region of sgRNA. Overall, A and G make sgRNA more stable and more efficient. In addition to gRNA sequence features, novel features in PAM affect sgRNA reproducibility. For example, in the variable nucleotide N of NGG for spCas9, C is preferred, while T is not favored. Moreover, Cas9 preferences for particular sgRNA sequence features are quite different from those in a dCas9-mediated application. A 19-nucleotide sgRNA in dCas9-mediated CRISPRi and CRISPRa showed the highest efficiency compared with 20 nt or 17–18 nt truncated sgRNA for Cas9. Moreover, the seed region of sgRNA is of key importance in CRISPR/Cas9, while all sgRNA nucleotides contribute to gRNA efficiency in CRISPR/dCas9.
3.3.7 Selection of Suitable gRNA
A given target sequence or gene may have many potential gRNAs. It is important to select the most suitable gRNA with the highest efficiency for the intended modification. Suitability is assessed in terms of position relative to target site, high on-target activity, and low off-target activity. This can be achieved with tools such as WGE and CRISPOR using custom filters. Filtering for gRNAs with low off-targets will identify candidates with minimum off-targets. However, a gRNA with high on-target activity may have significantly low specificity leading to high off-targets. A gRNA with a high on-target score and high specificity would be an ideal sgRNA candidate for the desired CRISPR application (Thomas et al. 2019).
3.3.8 Design Criteria for Genome-Wide CRISPR Libraries
In contrast to individual gRNA design, CRISPR libraries are designed to screen mutations (or desired modifications) in many genes or across an entire genome. As a result, sgRNA design for genome-wide CRISPR libraries is entirely computer-based because it is impossible to evaluate each gRNA. Instead, multiple sgRNAs are designed for each gene in the genome at different locations. Users can design their own custom libraries or use libraries according to their chosen application (Thomas et al. 2019). Selected libraries and their applications are listed in Table 3.2.
3.4 Specificity in CRISPR/Cas
After selecting PAM and potential target sites, the next step is to identify the site most likely to result in efficient genome editing. In addition to choosing an sgRNA to match the target site, researchers try to select one with no additional binding sites in the genome. While the ideal sgRNA would have no homologous sites in the genome, in practice an sgRNA will have partial homology to many additional sites in the genome, i.e., off-targets (Duan et al. 2014). Off-target sites with mismatches near PAM will not be cleaved efficiently; such sgRNA would have lower off-targets effects and will be associated with the highest specificity as compared to those sgRNA in which mismatches are away from PAM in off-target sites. Off-target sites may be effectively minimized by predicting CRISPR/Cas specificity and designing a specific and optimal sgRNA. The two main approaches for predicting sgRNA specificity are based on either (1) alignment or (2) scoring. In the first method, sgRNA sequences are aligned to a given genome using conventional or specialized tools to discover all off-targets, and only frequency of the mismatches in the gRNA sequence is considered. In a scoring-based approach, sgRNA are scored and ranked after the initial alignment in order to select the most specific sgRNA for a given experiment. In this scoring-based approach, in addition to frequency of mismatches, positional weighing of each mismatch is calculated. Two scoring-based approaches are commonly used: (1) a learning-based method and (2) a hypothesis-driven method. Below we discuss alignment- and scoring-based methods in detail (Liu et al. 2020).
3.4.1 Alignment-Based Approach to Predict Specificity
Alignment-based methods for assessing sgRNA sequences involve aligning the sgRNA with a reference genome and identifying potential off-targets based on sequence homology. Bowtie (Langmead et al. 2009) and Burrows-Wheeler aligner (BWA) mapping tools are used to predict off-targets, but neither identify small PAM sequences. Because these tools allow a limited number of mismatches in the sgRNA seed region, they cannot identify all off-targets. CHOPCHOP and CCTOP design tools use Bowtie to find off-targets for a candidate sgRNA, while CRISPOR uses BWA. Alignment-based Cas-OFFinder and Cas-OT also predict off-targets (Liu et al. 2020). Cas-OFFinder is popular for finding off-targets with no mismatch limitations and can even predict off-targets with a 1-bp insertion or deletion (Thomas et al. 2019). Cas-OT can identify off-targets with 6-bp mismatches in the seed region and predict off-targets in coding exons of genes. Alignment-based CRISFlash and FlashFry use tree-based algorithms and user-defined data to optimize sgRNA. As well as off-target predictions, FlashFry provides additional information such as GC content and on-target score for sgRNA (Liu et al. 2020).
3.4.2 Specificity Prediction Through Scoring-Based Tools
3.4.2.1 Hypothesis-Driven Methods
Alignment-based methods can reliably predict off-targets. However, not all nucleotide positions with mismatches in sgRNA are equally effective in terms of off-target cleavage. In addition, alignment-based predictions for off-targets are sometimes false positives. One study found that only a few of the off-targets predicted by Cas-OFFinder and CC-Top were valid, and the tools also failed to predict some valid off-targets. So, there was a need to limit the features that contribute to the non-specific off-targets in CRISPR/Cas (Liu et al. 2020). These issues can be addressed in CRISPR/Cas systems by using the MIT specificity score (named after the institution) to evaluate off-targets (Hsu et al. 2013). Hsu et al. studied more than 700 sgRNAs and evaluated sgRNA/Cas9 sequence features such as contribution of position and numbers of mismatched nucleotide in the target site (Hsu et al. 2013). The MIT score is adopted to predict off-targets in design tools such as CHOPCHOP and CRISPOR (Haeussler et al. 2016; Labun et al. 2016). Cutting frequency determination (CFD) score is also popular for evaluating off-targets in CRISPR/Cas (Liu et al. 2020). In addition to recognizing NGG PAM, Cas9 recognizes non-canonical PAM sites such as NAG, NGA, and NCG, thus leading to off-targets. Doench et al. (2016) used PAM sequence features in their scoring matrix to predict off-targets. CFD score is considered a better performer better than MIT score and has been adopted by many design tools, such as GuideScan (Perez et al. 2017) and CRISPRscan (Moreno-Mateos et al. 2015). Other design tools use sgRNA/Cas9 structural features to predict off-targets. For example, CRISPR-OFF (Alkan et al. 2018) and uCRISPR (Zhang et al. 2019) use structural features because their off-target prediction accuracy is better than sequence features.
3.4.2.2 Learning-Based Methods
Compared to empirical algorithms, learning methods use multiple features (including PAM, GC contents, methylation state, and chromatic structure) to improve their off-target predictions. Most recent tools use machine learning with multiple features for predicting CRISPR/Cas system specificity and off-targets. For example, CRISPR target assessment (CRISTA), which uses machine learning to predict efficiency, was found to perform better than other tools (Liu et al. 2020). The computer platform DeepCRISPR, which incorporates sgRNA on-targets and off-targets into a single framework, has been found to perform better than other tools for predicting efficiency and off-targets (Chuai et al. 2018).
3.5 Factors Affecting Specificity
Numerous studies have revealed different factors that may affect CRISPR/Cas specificity. These factors can be classified into two categories: (a) an intrinsic specificity of Cas9 which recognize the importance of position of every sgRNA nucleotide to create DSB and (b) relative abundance of sgRNA/Cas9 for effective target cleavage. Factors that may contribute to CRISPR/Cas system specificity are discussed below.
3.5.1 Importance of PAM in CRISPR/Cas Specificity
To be recognized by an individual Cas9 domain, PAM must be next to the 3′ end of the genome target sequence (Wu et al. 2014b). Because PAM sequences vary across Cas endonucleases, users can select a different Cas endonuclease if a particular PAM (e.g., NGG for Cas9) does not exist in the target sequence. The most commonly used Cas endonuclease, Cas9, recognizes NGG for cleavage but can also recognize the canonical PAM sites NGA and NAG, thus increasing the number of off-targets. Some of these Cas proteins require a longer PAM sequence such as SaCas9 protein, derived from Staphylococcus aureus, which has “NNGRRT” PAM requirement. It is assumed that such Cas9 proteins which recognize a longer PAM will have less targetable sites in the genome and, therefore, will have fewer off-target sites in a given target DNA. PAM sequences with appropriate Cas endonucleases are listed in Table 3.1.
3.5.2 Seed Sequence of sgRNA
Recruiting Cas9 to the genome target site requires sgRNA. In vitro studies have shown that Cas9 can tolerate mismatches in the first seven nucleotides in the region distal to PAM. However, studies with bacteria and mammals have confirmed that mismatches in 10–12 bp PAM proximal region (also called seed region) of the gRNA will result in reduced cleavage or complete abolishment. Other studies suggest there is no clearly defined seed region, but have confirmed that mismatches in the PAM proximal region stop Cas9 cleavage of DNA (Cong et al. 2013). In contrast, genome-wide binding datasets have shown a clearly defined seed region, limited to five nucleotides proximal to PAM (Wu et al. 2014b). The differences in seed region might arise from factors such as concentration and time required for Cas9 binding and cleavage.
3.5.3 Effective Concentration of Cas9/sgRNA Complex
The effective concentration of Cas9/sgRNA influences the specificity of CRISPR/Cas systems. Studies have confirmed that cleavage becomes less specific at higher effective concentrations of Cas9/sgRNA. For example, an in vitro study found that higher concentrations of Cas9/sgRNA complex resulted in greater tolerance of mismatches, leading to cleavage of non-specific sites. Hsu and co-authors suggested that decreasing the amount of plasmid in transfected cells led to increasedCas9 specificity (Wu et al. 2014b; Hsu et al. 2013). Another study showed that a 2.6-fold increase in Cas9 concentration led to a similar increase in off-targets. When Cas9 level remained constant, the amount of sgRNA influenced off-target number (Wu et al. 2014a).
3.5.4 Importance of sgRNA Sequence
SgRNA sequence is the key to Cas9 specificity because it contributes to Cas9 loading and Cas9/sgRNA binding to the target site. Differences in sgRNA sequence influence Cas9 tolerance of mismatches at every position in 20 nucleotides. A possible underlying mechanism for this change in specificity is that different sgRNA sequences may influence effective concentration of sgRNA. For example, it has been reported that seed sequence mutations in sgRNA increase its transcription by U6 promoter. Changes in sgRNA sequence may also contribute to chromatin state, off-targets, and thermodynamic stability of sgRNA-DNA duplex (Wu et al. 2014b). We describe these effects in detail below.
3.5.4.1 Chromatin Accessibility and Epigenetic Features Affecting Binding of Cas
Chromatin state, i.e., whether packed or open, may influence Cas9’s ability to access the target site. DNase I hypersensitivity (DHS) is a strong predictor of chromatin accessibility. DHS peaks for a number of accessible seed sequences and PAM have been found to accurately predict the number of chromatin immunoprecipitation (ChIP) peaks in vivo. Wu and colleagues have suggested that chromatin accessibility does not impact significantly on-target activity of sgRNA as compared to off-target binding (Wu et al. 2014a, b).
Methylation of CpG sites (where cytosine and guanine are adjacent, with guanine closer to 3′) is an epigenetic mechanism that has been found to be linked with chromatin silencing. A study confirmed that CpG methylation of target sites may restrict Cas9 binding to the target site. Target site methylation showed strong correlation with ChIP signal, and less binding was observed in highly methylated sites (Wu et al. 2014a, b). Hsu et al. showed that Cas9 can mutate highly methylated promoters in vivo. However, an in vitro study found that CpG methylation had no significant effect on Cas9 cleavage (Hsu et al. 2013). Taking these studies together suggests that CpG methylation may affect only off-target sites.
3.5.4.2 Numbers of Seed Sequence in the Genome
Depending on sgRNA seed sequence length (5–12 nt), a mammalian genome may contain hundreds of thousands of seed match sites followed by PAM. However, nucleotide preference in the seed regions may mean that specific seed match sequences could be dramatically low. For example, for Cas9, a mouse genome contains about one million AAGGA + NGG seed sites but less than 10,000 CGTCG + NGG sites (Wu et al. 2014a, b). The relative abundance of seed sites is an important factor in designing specific sgRNA, especially in dCas9 applications.
3.5.4.3 Length of Target Sequence Influences Specificity
Length of sgRNA is important for Cas9 specificity. A 20-bp gRNA is optimal for guiding Cas9 to a target site. Although one might speculate that specificity may increase with sgRNA length, Ran et al. found that when sgRNA length was increased by extending the 5′-end, the extended sequence at the 5′-end was degraded in vivo (Ran et al. 2013). In contrast, truncated sgRNA with 17–18 nt of length increased Cas9 specificity. While the underlying mechanism is not clear, it may be that the first two nucleotides do not contribute to Cas9 stability, but instead contribute to off-targets (Fu et al. 2014).
3.5.5 sgRNA Scaffold
The impact of modifications in the sgRNA scaffold region has not been studied in detail. However, it is known that truncation or extension at the 3′ end may contribute to Cas9 stability and specificity by changing sgRNA expression, in similar fashion to 5′-end modifications in sgRNA. Increasing the length of the hairpin bound by Cas9 has been found to increase sgRNA efficiency for imaging and transcriptional regulation, probably due to efficient loading of sgRNA, but the exact mechanism remains unclear (Hsu et al. 2013; Wu et al. 2014b).
3.5.6 Repair Outcomes of DSBs
In addition to the above factors, DNA repair outcomes and sequence variations are likely to influence the selection of specific sgRNA. Several studies have identified a bias in repair outcomes for KO applications. These studies have shown that nucleotide comparison of target site adjacent to the cleavage site is important for single-nucleotide insertion or deletion in NHEJ repair pathway (Mao et al. 2013). The presence of thymine (T) adjacent to the cleavage site was associated with precise insertion of a single homologous nucleotide at the cleavage site (T to TT). However, having a dinucleotide repeat adjacent to the cleavage site led to single-nucleotide deletion with removal of homologous base (CC to C). Moreover, microhomologies in sequences flanking the cleavage site resulted in deletion of 30 nucleotides through microhomology-mediated end joining (MMEJ) repair. These findings highlight a bias in repair outcomes linked to the presence of specific sequences in target sites and the competing roles of NHEJ and MMEJ. Based on these studies, computational tools such as Favored Outcomes of Repair Events at Cas9 Targets (FORECasT) and inDelphi have been developed to predict the most likely mutational outcomes of CRISPR/Cas experiments.
3.6 Efficiency of sgRNA
Initially it was believed that CRISPR/Cas9 could target any genome sequence that was followed by PAM (NGG). As a result, most of the early bioinformatic tools were constructed based on simple methods to locate target site and PAM to design sgRNAs. Some of these tools predicted sgRNA position relative to gene features. However, several later studies demonstrated that Cas9 cleavage efficiency varies significantly between different sites, i.e., not all sites are cleaved with the same efficiency (Cong et al. 2013; Jinek et al. 2012; Mali et al. 2013; Wang et al. 2014). For example, two sgRNAs can have 100% homology with their target sites but different cleavage efficiency, indicating that cleavage efficiency may also be affected by specific nucleotides and nucleotide composition. Subsequent studies identified additional factors such as sequence features (GC contents, specific nucleotide positions, and sequence composition), genetic and epigenetic factors (methylation and chromatin arrangement), and thermodynamic properties (sgRNA secondary structure, melting temperature (Tm), and free energy) that influence on-target cleavage efficiency.
Nucleotide position and composition in the target sequence is critical for CRISPR/Cas on-target efficiency (Wilson et al. 2018; Wong et al. 2015). CRISPR/Cas-based screening in mammals has shown that G is highly preferred at positions 1 and 2 upstream to PAM, while T is not favored at position 4 in close proximity to PAM. The GC content of positions 4–13 proximal to PAM is also important for Cas9 cleavage efficiency. Using sequence features such as GC content, preferred nucleotide position, and sgRNA position relative to gene features, predictive models have been developed to design efficient sgRNA for CRISPR/Cas applications. Several laboratories have used these models to develop individual design platforms such as E-CRISP, CHOPCHOP, CRISPR-FOCUS, and CCTOP for predicting sgRNA efficiency (Table 3.3).
Genetic and epigenetic features also contribute to target-site cleavage efficiency. Studies have shown that nucleosomes (sections of chromatin) may reduce Cas9 cleavage efficiency, and DNase I hypersensitivity (DHS) and epigenetic signatures may influence on-target efficiency. Predict-SGRNA is an R package (R is a free software environment) that uses epigenetic features to predict sgRNA cleavage efficiency (Liu et al. 2020). CRISPRpred and uCRISPR predict sgRNA efficiency using the energy properties of sgRNA, DNA, and Cas9 complex and sgRNA secondary structure. Because not all sgRNAs are effective, even when using the best design tools, multiple sgRNAs are used for each target gene. Multiple sgRNAs are also required to distinguish on-target perturbation from any off-target effect of an individual sgRNA.
3.7 Off-Targeting in CRISPR/Cas
Off-targets are a major challenge for the CRISPR/Cas community because Cas9 can bind and create DSBs even when there is only partial complementarity between sgRNA and target site. Numerous studies have reported that CRISPR/Cas may produce substantial numbers of off-targets. For example, a study in human beings found that Cas9 can tolerate up to five mismatches between sgRNA and target site, leading to DNA cleavage frequencies even higher than the intended target site (Carroll 2013; Hsu et al. 2013; Xie et al. 2014). Off-targets are not random changes but are induced by the PAM and target site. Natural off-targets in a bacterial defense system may degrade hypervariable nucleic acids (i.e., those vary much more than their counterparts in other similar regions) or plasmids beneficial for archaea and bacteria. However, from a genome editing perspective, off-targets may lead to undesirable changes at random sites in the genome, thus compromising the benefits of genome modifications. Predicting and minimizing off-targets in advance is essential for safe use of CRISPR/Cas, especially in therapeutic applications and translational research. It is also important to identify all off-targets and confirm that a desired phenotype has arisen from on-target modification instead of off-targets.
Several sgRNA design tools have a special focus on limiting off-targets in CRISPR/Cas (Table 3.4). Most of these produce sgRNA with minimal off-targets and show predicted off-targets for a given sgRNA. Different tools use different scoring methods to predict off-targets. Most of these tools score off-targets either by using data from systematic mutation studies or by having user-provided input penalties such as mismatch number and positions. Others use binary criteria, e.g., defined proximal or distal region, or sites with less than a defined number of mismatches. SgRNA candidates are then ranked by off-target number or the weighted sum of all off-target scores (Wu et al. 2014b). Some tools give option of using alternate PAM site to predict off-targets, e.g., NAG or NGA for Cas9.
As with on-target prediction tools, most design tools for off-target prediction initially focused on Cas9 and predicted off-targets through alignment-based methods using seed sequence followed by NGG. However, the discovery that Cas9 also binds NAG or NGA PAM made it apparent that many off-targets were being missed. The early tools were superseded by tools that used sequence similarity or dCas9-mediated binding to confirm off-target sites, but these later approaches were biased and not comprehensive. Unbiased approaches were then developed based on high-throughput, next-generation sequencing (NGS). For example, DSBCapture used integrase-deficient lentiviral vectors (IDLV) and sequencing, while Digenome-seq, ChIP-NGS (whole genome binding), and direct in situ breaks labeling, enrichment on streptavidin, and next-generation sequencing (BLESS) were developed to detect off-targets in CRISPR/Cas applications. However, these approaches also had advantages and disadvantages. IDLV and BLESS could detect genome-wide off-targets, but they were less efficient because most off-target sites are transient. In addition, both approaches could generate false-positive off-targets because DSBs may arise from endogenous processes. Although whole genome sequencing is ideal and unbiased, it can miss perfectly repaired off-targets and binding sites without cleavage. Moreover, ChIP-NGS could be biased towards open chromatin and highly expressed genes. Guide-seq has good efficiency but does not work for DNA nicks (single-stranded cuts). Digested genome sequencing (Digenome-seq) does not consider other factors that affect cleavage. All things considered, the above approaches are all useful but need refinement because in vitro results can differ from in vivo (Peng et al. 2016).
Over the last few years, considerable effort has gone into limiting off-targets and improving specificity. Approaches have included lowering GC content, employing paired nickase enzymes, and using truncated sgRNA (17–18 bp). Lower GC content may reduce off-targets because higher GC content improves RNA/DNA duplex stability, thereby increasing the chance of tolerated mismatches. SgRNA and target site mismatches that produce bulges at the 5′ end, the 3′ end, or 7–12 nucleotides proximal to PAM must be avoided. The combined use of paired nickases and paired sgRNAs will generate two closely associated single-stranded breaks and eventually make a DSB.
3.8 Application-Specific Design of sgRNA
Although all CRISPR/Cas applications rely on sgRNA to guide Cas9 to the target sequence, DSBs are not always required. KO and KI applications always require DSB creation to delete or insert DNA at a precise location respectively. Large-scale deletions and insertions require more than one DSB. In KO applications, the NHEJ repair pathway will introduce a small indel into the coding framework, leading to a frameshift mutation and thus disruption of protein formation. However, for repair templates with suitable homology arms, DSBs will be repaired by HDR pathway, consequently leading to site-specific insertion of the repair template. Because NHEJ is the preferred pathway in cells, HDR efficiency must be improved for KI applications. In contrast to KO and KI applications, CRISPRi and CRISPRa use dCas9, which does not create a DSB, but instead recruits a transcriptional activator (VP64) or repressor (Krüppel-associated box (KRAB) domain proteins) to the promoter region of a gene (Graham and Root 2015). Similarly, sgRNA position in CRISPRi and CRISPRa varies significantly between KO and KI applications. However, despite differences in sgRNA position relative to gene features, the same basic principle underlies sgRNA design in all applications. Here we summarize application-specific sgRNA design in CRISPR/Cas.
3.8.1 sgRNA for KO Applications
Being able to KO an individual gene is a powerful tool for functional genomics. Knockout (KO) of single and multiple genes is often studied to evaluate phenotypic changes in cells, tissue, or organisms and by subsequently characterizing those genes for their potential roles in different functions. CRISPR/Cas has become the gold standard for producing KO models for functional characterization of genes (Graham and Root 2015). The KO of a gene or genetic element may be achieved by creating a DSB that is repaired through the NHEJ pathway. Exon size and relative position are important for generating KO alleles. For example, larger exons would have multiple choices of sgRNA, making it easier to select an efficient sgRNA. However, small exons are easy to delete with two DSBs. In addition, sgRNA position relative to the gene features may affect the outcomes of KO applications. Targeting sgRNA too close to a translation initiation codon ATG may reinitiate translation at a downstream ATG, leading to N-terminal truncated protein. Similarly, targeting sgRNA close to the 3′-end of a gene may result in insufficient disruption of protein functions. With sgRNA design for KO applications, selecting an optimal target such as a functional domain, active site, or the transmembrane helical domain of a protein (Fig. 3.4) can increase the likelihood of completely disrupting protein functions (Thomas et al. 2019). Using multiple sgRNAs can help ensure that the curated phenotype in a KO experiment has resulted from disrupting the respective gene instead of off-targets. For large-scale design, multiple sgRNAs per gene are also recommended for increased screen efficiency. In addition to generating KO for a single gene, multiplex genomes using CRISPR/Cas can be used to simultaneously disrupt multiple genes in order to study their interactions and discover pathways.
3.8.2 Position of sgRNA for KI Applications
While the NHEJ pathway may lead to disruption of a gene, KI approaches using repair templates can use the HDR pathway to precisely insert a single nucleotide change or add a large template such as green fluorescent protein (GFP) (Wu et al. 2018), a tag (Chen et al. 2018), or a fluorophore. For the HDR-based repair pathway, the desired repair template must be introduced along with sgRNA and Cas9 or nCas9. The length and nature of the repair template depend on the size of the intended modification. For example, for a single-base replacement, ssDNA repair template with 50 bp homology arms on both sides of DSB could work efficiently. However, for larger insertions such as a GFP, tag, or fluorophore, a repair template with long homology arms (400–1000 bp) is desirable (Fig. 3.4). It is also advisable to exclude PAM site in the repair template. Moreover, mutating PAM site and sgRNA binding site with silent mutations would prevent subsequent binding and cleavage of target site after insertion of the repair template. These silent mutations may also assist genotyping following insertion of the desired repair template (Graham and Root 2015).
3.8.3 Designing sgRNA for CRISPRi and CRISPRa
In contrast to KO and KI applications that use Cas9 or nCas9, respectively, transcriptional regulation through CRISPR/Cas relies on dCas9, which does not create DSB but simply binds at a precise location in the genome. Binding dCas9 with an appropriate activator or repressor to a gene’s promoter region may subsequently activate or repress the gene by blocking binding of RNA polymerase or transcriptional factors. SgRNA position relative to the transcription start site (TSS) may affect the efficiency of activation or repression. Accurate TSS identification is highly desirable for transcriptional regulation through CRISPR/Cas. Generally, the target site for sgRNA design in CRISPRi should be downstream (within a 300 bp window) of TSS, while for CRISPRa it should be upstream (within a 400 bp window) (Fig. 3.4). Designing multiple sgRNAs for a target region should assist in achieving the best results (Davis et al. 2018; Noguchi et al. 2017; Thomas et al. 2019).
3.8.4 SgRNA in Epigenetic Regulation
dCas9 can be used to alter gene expression by recruiting epigenetic modifiers such as lysine-specific demethylase 1 (LSD1), ten-eleven translocation gene protein 1 (TET1), DNA methyltransferase MQ1, and histone acetyltransferase p300 to modify the methylation state of cytosine in the promoter region by inducing demethylation or histone acetylation (Brocken et al. 2017). Epigenetic modifiers sometimes work better than CRISPRi or CRISPRa.
3.8.5 Design Criteria for Base Editing
In CRISPR/Cas system, base editing was initially achieved by providing a repair template using the HDR pathway, which has low efficiency. To overcome the low efficiency, researchers developed two CRISPR-mediated base editing platforms for DNA (cytosine base editor (CBE) and adenine base editor (ABE)) and an RNA base-editing platform. CBE and ABE were developed by fusing either cytosine deaminase or adenine deaminase with an appropriate Cas protein (dCas9 or nCas9) (Liang and Huang 2019). The RNA base editor was developed by fusing type VI CRISPR/Cas effector (dCas13b) with hyper-activated adenosine deaminase 2 that acts on RNA (ADAR2) to create a programmable RNA base editor known as REPAIR (RNA editing for programmable A to I (G) replacement). In base editing, sgRNA position depends on the targeted nucleotide’s location in the protospacer region. The targeted nucleotide must be present within the active base editing window on the non-targeted strand, thus deciding the position and orientation of the sgRNA. The size of the active base window (usually four to eight nucleotides) depends on which base editor is used (Thomas et al. 2019). Base-editing efficiency can sometimes be very low at certain positions because these are inaccessible due to nucleosomes.
3.8.6 Designing sgRNA for RNA Editing
An alternative CRISPR/Cas system for regulation at transcriptional level uses CRISPR/Cas13, which specifically targets single-stranded RNA (ssRNA). CRISPR/Cas13 uses CRISPR RNA (crRNA) to recognize and cleave ssRNA (Freije et al. 2019). In bacteria, non-specific cleavage of RNA has been observed after initial cleavage with Cas13. Cas13 is used in a very sensitive diagnostic platform known as the specific high-sensitivity enzymatic reporter unlocking (SHERLOCK) assay for differentiating Zika virus strains (Kellner et al. 2019), genotyping human beings, and RNA imaging (Yang et al. 2019). SHERLOCK could also be useful for detecting SARS-CoV-2, the RNA virus that causes coronavirus disease 2019 (COVID-19) (Joung et al. 2020).
3.9 Design Tools for sgRNA
Design tools available to the CRISPR/Cas community have been developed by both academic and commercial institutes. Although the basic objective of these tools is to design and select an optimal sgRNA and provide information about the target site, each tool has its own particular features and benefits. Similarly, these tools all aim to provide sgRNA with minimal off-targets in the genome, but they employ various methods to score these off-targets. For example, off-target scoring in CHOPCHOP is based on empirical data from multiple studies, while Cas-Finder and E-CRISP evaluate off-targets using user-defined values for mismatch number and position.
Some design tools are application- or species-specific. For example, CRISPR-ERA and BE-Designer specifically design sgRNA for transcriptional regulation (CRISPRi/CRISPRa) and base editing, respectively. FlyCRISPR and CRISPR-PLANT are specialized for sgRNA design in Drosophila and plants, respectively (Liu et al. 2020). Some sgRNA design tools provide users with additional options for selecting alternative PAM sites, as well as Cas effector. Some useful sgRNA design tools are listed in Table 3.5, after which we discuss some of these potential tools.
3.9.1 CHOPCHOP
More than 200 genomes are available on the CHOPCHOP website; users can input gene name or target sequence. This tool supports gRNA design for multiple applications (KO, KI, CRISPRi, and CRISPRa); users can choose application-specific Cas effector endonuclease. CHOPCHOP ranks potential sgRNAs on position, GC contents, mismatch number, and efficiency scores (Liu et al. 2020).
3.9.2 Base Editing (BE)-Analyzer and BE-Designer
These are publicly available design tools for base editing. Both tools help researchers select sgRNA for desired region and analyze outcomes of base editing from NGS data. BE-Designer also lists all potential sgRNAs for a given DNA sequence and provides off-targets for a given sgRNA against a large number of species (Hwang et al. 2018).
3.9.3 CRISPOR
One of the best tools for designing efficient sgRNA, CRISPOR contains 19 different PAMs and 417 different genomes. It can accept genome coordinates or user-provided sequences. Each sgRNA will be ranked for off-targets, specificity, and efficiency. Outcome predictions, GC contents, and poly T will also be given for each sgRNA (Liu et al. 2020).
3.9.4 CRISFlash
Like CHOPCHOP, CRISFlash can use sequenced genome or genome sequence to design sgRNA. In addition, it accepts user-defined values to optimize sgRNA and off-targets. CRISFlash is considered a faster tool for sgRNA design and scoring off-targets (Jacquin et al. 2019).
3.10 Prospects
CRISPR/Cas technology is a revolutionary tool for functional genomic human therapeutics and agricultural advances. Because sgRNA plays an indispensable role in CRISPR/Cas-mediated genome editing, numerous tools have been developed for designing efficient and specific sgRNA with minimal off-targets. However, off-targets continue to represent a major challenge for CRISPR/Cas-mediated genome manipulation. Systematic studies show that predictive models for efficient sgRNA design are not always effective for all applications and all species. This makes it imperative that scientists know the weaknesses and strengths of each model for sgRNA design. As new knowledge about CRISPR continues to emerge, it is clear that sgRNA and PAM are not the only influences on CRISPR/Cas cleavage, with additional such factors now including GC contents and chromatin accessibility. The ongoing discovery of new and novel features that contribute to CRISPR/Cas specificity and efficiency will also help minimize off-targets. Moreover, it has become clear that CRISPR/Cas outcomes are specific rather than random. Such findings will facilitate more precise editing with CRISPR/Cas.
In summary, recent advances in our understanding of CRISPR mechanisms and factors affecting specificity and efficiency, combined with the further development of bioinformatics tools, will enable more precision in achieving desired on-target modifications without potential off-targets. Directed evolution using EvolvR may also help scientists to engineer new Cas proteins with improved specificity.
Abbreviations
- ABE:
-
Adenine base editor
- BLESS:
-
Breaks labeling, enrichment on streptavidin
- BWA:
-
Burrows-Wheeler aligner
- CBE:
-
Cytosine base editor
- CFD:
-
Cutting frequency determination
- ChIP:
-
Chromatin immunoprecipitation
- Cmr:
-
Cas module-RAMP
- Cpf1:
-
CRISPR from Prevotella and Francisella 1
- CRISPR/Cas:
-
Clustered regularly interspaced short palindromic repeats/CRISPR associated protein
- CRISPRa:
-
CRISPR activation
- CRISPRi:
-
CRISPR interference
- CRISTA:
-
CRISPR target assessment
- crRNA:
-
CRISPR RNA
- DHS:
-
DNase I hypersensitivity
- DSB:
-
Double-stranded break
- GFP:
-
Green fluorescent protein
- gRNA :
-
Guide RNA
- HDR:
-
Homology-directed repair
- HEK293:
-
Human embryonic kidney 293 cells
- IDLV:
-
Integrase-deficient lentiviral vectors
- KI:
-
Knock-in
- KO:
-
Knockout
- KRAB:
-
Krüppel-associated box
- LSD1:
-
Lysine-specific demethylase 1
- MMEJ:
-
Microhomology-mediated end joining
- nCas9:
-
Cas9 nickase
- NGS:
-
Next-generation sequencing
- NHEJ:
-
Non-homologous end joining
- PAM:
-
Protospacer adjacent motif
- RAMP:
-
Repeat-associated mysterious proteins
- RNP:
-
Ribonucleoprotein
- sgRNA :
-
Single-guide RNA
- SHERLOCK:
-
Specific high-sensitivity enzymatic reporter unlocking
- TET1:
-
Ten-eleven translocation gene protein 1
- TSS:
-
Transcription start site
- WGE:
-
Wellcome Sanger Institute genome editing
References
Abudayyeh OO, Gootenberg JS, Konermann S, Joung J, Slaymaker IM, Cox DB, Severinov K (2016) C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353(6299):aaf5573
Alkan F, Wenzel A, Anthon C, Havgaard JH, Gorodkin J (2018) CRISPR-Cas9 off-targeting assessment with nucleic acid duplex energy parameters. Genome Biol 19(1):1–13
Alkhnbashi OS, Meier T, Mitrofanov A, Backofen R, Voß B (2020) CRISPR-Cas bioinformatics. Methods 172:3–11
Bae S, Park J, Kim JS (2014) Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30(10):1473–1475
Banan M (2020) Recent advances in CRISPR/Cas9-mediated knock-ins in mammalian cells. J Biotechnol 308:1–9
Bassett AR et al (2013) Mutagenesis and homologous recombination in Drosophila cell lines using CRISPR/Cas9. Biol Open 3:42. https://doi.org/10.1242/bio.20137120
Brazelton VA Jr, Zarecor S, Wright DA, Wang Y, Liu J, Chen K, Lawrence-Dill CJ (2015) A quick guide to CRISPR sgRNA design tools. GM Crops Food 6(4):266–276
Brocken DJ, Tark-Dame M, Dame RT (2017) dCas9: a versatile tool for epigenome editing. Curr Issues Mol Biol 26:15–32
Carroll D (2013) Staying on target with CRISPR-Cas. Nat Biotechnol 31(9):807–809
Chen B, Zou W, Xu H, Liang Y, Huang B (2018) Efficient labeling and imaging of protein-coding genes in living cells using CRISPR-Tag. Nat Commun 9(1):1–9
Chuai G, Ma H, Yan J, Chen M, Hong N, Xue D, Gu F (2018) DeepCRISPR: optimized CRISPR guide RNA design by deep learning. Genome Biol 19(1):80
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339(6121):819–823
Cui Y, Xu J, Cheng M, Liao X, Peng S (2018) Review of CRISPR/Cas9 sgRNA design tools. Interdiscipl Sci Comput Life Sci 10(2):455–465
Davis CA, Hitz BC, Sloan CA, Chan ET, Davidson JM, Gabdank I, Hilton JA, Jain K, Baymuradov UK, Narayanan AK, Onate KC, Graham K, Miyasato SR, Dreszer TR, Strattan JS, Jolanki O, Tanaka FY, Cherry JM (2018) The Encyclopedia of DNA elements (ENCODE): data portal update. Nucleic Acids Res 46(D1):D794–D801. https://doi.org/10.1093/nar/gkx1081
Doench JG, Hartenian E, Graham DB, Tothova Z, Hegde M, Smith I, Root DE (2014) Rational design of highly active sgRNAs for CRISPR-Cas9–mediated gene inactivation. Nat Biotechnol 32(12):1262–1267
Doench JG, Fusi N, Sullender M, Hegde M, Vaimberg EW, Donovan KF, Virgin HW (2016) Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat Biotechnol 34(2):184–191
Duan J, Lu G, Xie Z, Lou M, Luo J, Guo L, Zhang Y (2014) Genome-wide identification of CRISPR/Cas9 off-targets in human genome. Cell Res 24(8):1009–1012
Freije CA, Myhrvold C, Boehm CK, Lin AE, Welch NL, Carter A, Yozwiak NL (2019) Programmable inhibition and detection of RNA viruses using Cas13. Mol Cell 76(5):826–837
Fu Y, Sander JD, Reyon D, Cascio VM, Joung JK (2014) Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat Biotechnol 32(3):279–284
Garcia-Doval C, Jinek M (2017) Molecular architectures and mechanisms of Class 2 CRISPR-associated nucleases. Curr Opin Struct Biol 47:157–166
Garneau JE, Dupuis M-È, Villion M, Romero DA, Barrangou R, Boyaval P, Moineau S (2010) The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468(7320):67–71
Gilbert LA, Horlbeck MA, Adamson B, Villalta JE, Chen Y, Whitehead EH, Qi LS (2014) Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159(3):647–661
Graham DB, Root DE (2015) Resources for the design of CRISPR gene editing experiments. Genome Biol 16(1):260
Haeussler M, Schönig K, Eckert H, Eschstruth A, Mianné J, Renaud JB, Joly JS (2016) Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol 17(1):148
Harrington LB, Burstein D, Chen JS, Paez-Espino D, Ma E, Witte IP, Doudna JA (2018) Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 362(6416):839–842
Heigwer F, Kerr G, Boutros M (2014) E-CRISP: fast CRISPR target site identification. Nat Methods 11(2):122–123
Hirano H, Gootenberg JS, Horii T, Abudayyeh OO, Kimura M, Hsu PD, Nishimasu H (2016) Structure and engineering of Francisella novicida Cas9. Cell 164(5):950–961
Ho SM, Hartley BJ, Julia TCW, Beaumont M, Stafford K, Slesinger PA, Brennand KJ (2016) Rapid Ngn2-induction of excitatory neurons from hiPSC-derived neural progenitor cells. Methods 101:113–124
Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V, Cradick TJ (2013) DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 31(9):827–832
Hwang GH, Park J, Lim K, Kim S, Yu J, Yu E, Bae S (2018) Web-based design and analysis tools for CRISPR base editing. BMC Bioinformatics 19(1):542
Jacquin AL, Odom DT, Lukk M (2019) Crisflash: open-source software to generate CRISPR guide RNAs against genomes annotated with individual variation. Bioinformatics 35(17):3146–3147
Jeon Y, Choi YH, Jang Y, Yu J, Goo J, Lee G, Jeong C (2018) Direct observation of DNA target searching and cleavage by CRISPR-Cas12a. Nat Commun 9(1):1–11
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816–821
Jinek M, East A, Cheng A, Lin S, Ma E, Doudna J (2013) RNA-programmed genome editing in human cells. elife 2:e00471
Joung J, Ladha A, Saito M, Segel M, Bruneau R, Huang MLW, Greninger AL (2020) Point-of-care testing for COVID-19 using SHERLOCK diagnostics. MedRxiv
Kellner MJ, Koob JG, Gootenberg JS, Abudayyeh OO, Zhang F (2019) SHERLOCK: nucleic acid detection with CRISPR nucleases. Nat Protoc 14(10):2986–3012
Kleinstiver BP, Prew MS, Tsai SQ, Topkar VV, Nguyen NT, Zheng Z, Aryee MJ (2015) Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523(7561):481–485
Labun K, Montague TG, Gagnon JA, Thyme SB, Valen E (2016) CHOPCHOP v2: a web tool for the next generation of CRISPR genome engineering. Nucleic Acids Res 44(W1):W272–W276
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10(3):R25
Lee CM, Cradick TJ, Bao G (2016) The Neisseria meningitidis CRISPR-Cas9 system enables specific genome editing in mammalian cells. Mol Ther 24(3):645–654
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25(14):1754–1760
Liang P, Huang J (2019) Off-target challenge for base editor-mediated genome editing. Cell Biol Toxicol 35:185
Lino CA, Harper JC, Carney JP, Timlin JA (2018) Delivering CRISPR: a review of the challenges and approaches. Drug Deliv 25(1):1234–1257
Liu H, Wei Z, Dominguez A, Li Y, Wang X, Qi LS (2015) CRISPR-ERA: a comprehensive design tool for CRISPR-mediated gene editing, repression and activation. Bioinformatics 31(22):3676–3678
Liu L, Chen P, Wang M, Li X, Wang J, Yin M, Wang Y (2017) C2c1-sgRNA complex structure reveals RNA-guided DNA cleavage mechanism. Mol Cell 65(2):310–322
Liu G, Zhang Y, Zhang T (2020) Computational approaches for effective CRISPR guide RNA design and evaluation. Comput Struct Biotechnol J 18:35–44
Luo Y (2019) CRISPR gene editing. Springer, New York, NY
Ma J, Köster J, Qin Q, Hu S, Li W, Chen C, Xu H (2016) CRISPR-DO for genome-wide CRISPR design and optimization. Bioinformatics 32(21):3336–3338
Makarova KS, Koonin EV (2015) Annotation and classification of CRISPR-Cas systems CRISPR. Springer, New York, NY, pp 47–75
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Church GM (2013) RNA-guided human genome engineering via Cas9. Science 339(6121):823–826
Mao Y, Zhang H, Xu N, Zhang B, Gou F, Zhu JK (2013) Application of the CRISPR–Cas system for efficient genome engineering in plants. Mol Plant 6(6):2008–2011
Mohr SE, Hu Y, Ewen-Campen B, Housden BE, Viswanatha R, Perrimon N (2016) CRISPR guide RNA design for research applications. FEBS J 283(17):3232–3238
Montague TG, Cruz JM, Gagnon JA, Church GM, Valen E (2014) CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res 42(W1):W401–W407
Moon SB, Lee JM, Kang JG, Lee NE, Ha DI, Kim SH, Kim YS (2018) Highly efficient genome editing by CRISPR-Cpf1 using CRISPR RNA with a uridinylate-rich 3′-overhang. Nat Commun 9(1):1–11
Moreno-Mateos MA, Vejnar CE, Beaudoin JD, Fernandez JP, Mis EK, Khokha MK, Giraldez AJ (2015) CRISPRscan: designing highly efficient sgRNAs for CRISPR-Cas9 targeting in vivo. Nat Methods 12(10):982–988
Morgens DW, Wainberg M, Boyle EA, Ursu O, Araya CL, Tsui CK, Li A (2017) Genome-scale measurement of off-target activity using Cas9 toxicity in high-throughput screens. Nat Commun 8(1):1–8
Naito Y, Hino K, Bono H, Ui-Tei K (2015) CRISPRdirect: software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 31(7):1120–1123
Nishimasu H, Cong L, Yan WX, Ran FA, Zetsche B, Li Y, Nureki O (2015) Crystal structure of Staphylococcus aureus Cas9. Cell 162(5):1113–1126
Noguchi S, Arakawa T, Fukuda S, Furuno M, Hasegawa A, Hori F, Ishikawa-Kato S, Kaida K, Kaiho A, Kanamori-Katayama M, Kawashima T, Kojima M, Kubosaki A, Manabe RI, Murata M, Nagao-Sato S, Nakazato K, Ninomiya N, Nishiyori-Sueki H, Noma S, Saijyo E, Saka A, Sakai M, Simon C, Suzuki N, Tagami M, Watanabe S, Yoshida S, Arner P, Axton RA, Babina M, Baillie JK, Barnett TC, Beckhouse AG, Blumenthal A, Bodega B, Bonetti A, Briggs J, Brombacher F, Carlisle AJ, Clevers HC, Davis CA, Detmar M, Dohi T, Edge ASB, Edinger M, Ehrlund A, Ekwall K, Endoh M, Enomoto H, Eslami A, Fagiolini M, Fairbairn L, Farach-Carson MC, Faulkner GJ, Ferrai C, Fisher ME, Forrester LM, Fujita R, Furusawa JI, Geijtenbeek TB, Gingeras T, Goldowitz D, Guhl S, Guler R, Gustincich S, Ha TJ, Hamaguchi M, Hara M, Hasegawa Y, Herlyn M, Heutink P, Hitchens KJ, Hume DA, Ikawa T, Ishizu Y, Kai C, Kawamoto H, Kawamura YI, Kempfle JS, Kenna TJ, Kere J, Khachigian LM, Kitamura T, Klein S, Klinken SP, Knox AJ, Kojima S, Koseki H, Koyasu S, Lee W, Lennartsson A, Mackay-Sim A, Mejhert N, Mizuno Y, Morikawa H, Morimoto M, Moro K, Morris KJ, Motohashi H, Mummery CL, Nakachi Y, Nakahara F, Nakamura T, Nakamura Y, Nozaki T, Ogishima S, Ohkura N, Ohno H, Ohshima M, Okada-Hatakeyama M, Okazaki Y, Orlando V, Ovchinnikov DA, Passier R, Patrikakis M, Pombo A, Pradhan-Bhatt S, Qin XY, Rehli M, Rizzu P, Roy S, Sajantila A, Sakaguchi S, Sato H, Satoh H, Savvi S, Saxena A, Schmidl C, Schneider C, Schulze-Tanzil GG, Schwegmann A, Sheng G, Shin JW, Sugiyama D, Sugiyama T, Summers KM, Takahashi N, Takai J, Tanaka H, Tatsukawa H, Tomoiu A, Toyoda H, van de Wetering M, van den Berg LM, Verardo R, Vijayan D, Wells CA, Winteringham LN, Wolvetang E, Yamaguchi Y, Yamamoto M, Yanagi-Mizuochi C, Yoneda M, Yonekura Y, Zhang PG, Zucchelli S, Abugessaisa I, Arner E, Harshbarger J, Kondo A, Lassmann T, Lizio M, Sahin S, Sengstag T, Severin J, Shimoji H, Suzuki M, Suzuki H, Kawai J, Kondo N, Itoh M, Daub CO, Kasukawa T, Kawaji H, Carninci P, Forrest ARR, Hayashizaki Y (2017) FANTOM5 CAGE profiles of human and mouse samples. Sci Data 4:170112. https://doi.org/10.1038/sdata.2017.112
Peng R, Lin G, Li J (2016) Potential pitfalls of CRISPR/Cas9-mediated genome editing. FEBS J 283(7):1218–1231
Perez AR, Pritykin Y, Vidigal JA, Chhangawala S, Zamparo L, Leslie CS, Ventura A (2017) GuideScan software for improved single and paired CRISPR guide RNA design. Nat Biotechnol 35(4):347–349
Pliatsika V, Rigoutsos I (2015) Off-Spotter: very fast and exhaustive enumeration of genomic look alikes for designing CRISPR-Cas guide RNAs. Biol Direct 10:4
Prykhozhij SV, Rajan V, Gaston D, Berman JN (2015) CRISPR multitargeter: a web tool to find common and unique CRISPR single guide RNA targets in a set of similar sequences. PLoS One 10(3):e0119372
Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP, Lim WA (2013) Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152(5):1173–1183
Ran FA, Hsu PD, Lin CY, Gootenberg JS, Konermann S, Trevino AE, Scott DA, Inoue A, Matoba S, Zhang Y (2013) Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154:1380–1389. PubMed: 23992846
Ren B, Liu L, Li S, Kuang Y, Wang J, Zhang D, Zhou H (2019) Cas9-NG greatly expands the targeting scope of the genome-editing toolkit by recognizing NG and other atypical PAMs in rice. Mol Plant 12(7):1015–1026
Sanjana NE, Shalem O, Zhang F (2014) Improved vectors and genome-wide libraries for CRISPR screening. Nat Methods 11(8):783
Sanson KR, Hanna RE, Hegde M, Donovan KF, Strand C, Sullender ME, Doench JG (2018) Optimized libraries for CRISPR-Cas9 genetic screens with multiple modalities. Nat Commun 9(1):1–15
Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelsen TS, Zhang F (2014) Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343(6166):84–87
Shi J, Wang E, Milazzo JP, Wang Z, Kinney JB, Vakoc CR (2015) Discovery of cancer drug targets by CRISPR-Cas9 screening of protein domains. Nat Biotechnol 33(6):661–667
Stemmer M, Thumberger T, del Sol KM, Wittbrodt J, Mateo JL (2015) CCTop: an intuitive, flexible and reliable CRISPR/Cas9 target prediction tool. PLoS One 10(4):e0124633
Stemmer M, Thumberger T, del Sol Keyer M, Wittbrodt J, Mateo JL (2017) Correction: CCTop: an intuitive, flexible and reliable CRISPR/Cas9 target prediction tool. PLoS One 12(4):e0176619
Thomas M, Parry-Smith D, Iyer V (2019) Best practice for CRISPR design using current tools and resources. Methods 164–165:3
Tian P, Wang J, Shen X, Rey JF, Yuan Q, Yan Y (2017) Fundamental CRISPR-Cas9 tools and current applications in microbial systems. Synth Syst Biotechnol 2(3):219–225
Torres-Perez R, Garcia-Martin JA, Montoliu L, Oliveros JC, Pazos F (2019) WeReview: CRISPR tools—live repository of computational tools for assisting CRISPR/Cas experiments. Bioengineering 6(3):63
Uniyal AP, Mansotra K, Yadav SK, Kumar V (2019) An overview of designing and selection of sgRNAs for precise genome editing by the CRISPR-Cas9 system in plants. 3 Biotech 9(6):223
Wang T, Wei JJ, Sabatini DM, Lander ES (2014) Genetic screens in human cells using the CRISPR-Cas9 system. Science 343(6166):80–84
Wang T et al (2015) Identification and characterization of essential genes in the human genome. Science 350(6264):1096
Wang T, Guan C, Guo J, Liu B, Wu Y, Xie Z, Xing XH (2018) Pooled CRISPR interference screening enables genome-scale functional genomics study in bacteria with superior performance. Nat Commun 9(1):1–15
Wheeler EC, Vu AQ, Einstein JM, DiSalvo M, Ahmed N, Van Nostrand EL, Yeo GW (2020) Pooled CRISPR screens with imaging on microraft arrays reveals stress granule-regulatory factors. Nat Methods 17:636–642
Wilson LO, O’Brien AR, Bauer DC (2018) The current state and future of CRISPR-Cas9 gRNA design tools. Front Pharmacol 9:749
Wong N, Liu W, Wang X (2015) WU-CRISPR: characteristics of functional guide RNAs for the CRISPR/Cas9 system. Genome Biol 16(1):1–8
Wu X, Scott DA, Kriz AJ, Chiu AC, Hsu PD, Dadon DB, Chen S (2014a) Genome-wide binding of the CRISPR endonuclease Cas9 in mammalian cells. Nat Biotechnol 32(7):670–676
Wu X, Kriz AJ, Sharp PA (2014b) Target specificity of the CRISPR-Cas9 system. Quant Biol 2(2):59–70
Wu Z, Zhao J, Qiu M, Mi Z, Meng M, Guo Y, Yuan Z (2018) CRISPR/Cas9 mediated GFP knock-in at the MAP1LC3B locus in 293FT cells is better for bona fide monitoring cellular autophagy. Biotechnol J 13(11):1700674
Xiao A, Cheng Z, Kong L, Zhu Z, Lin S, Gao G, Zhang B (2014) CasOT: a genome-wide Cas9/gRNA off-target searching tool. Bioinformatics 30:1180
Xie S, Shen B, Zhang C, Huang X, Zhang Y (2014) sgRNAcas9: a software package for designing CRISPR sgRNA and evaluating potential off-target cleavage sites. PLoS One 9(6):e100448
Yamano T, Nishimasu H, Zetsche B, Hirano H, Slaymaker IM, Li Y, Ishitani R (2016) Crystal structure of Cpf1 in complex with guide RNA and target DNA. Cell 165(4):949–962
Yang LZ, Wang Y, Li SQ, Yao RW, Luan PF, Wu H, Chen LL (2019) Dynamic imaging of RNA in living cells by CRISPR-Cas13 systems. Mol Cell 76(6):981–997
Yennamalli RM, Kalra S, Srivastava PA, Garlapati VK (2017) Computational tools and resources for CRISPR/Cas 9 genome editing method. MOJ Proteom Bioinform 5(4):116
Zhang D, Hurst T, Duan D, Chen SJ (2019) Unified energetics analysis unravels SpCas9 cleavage activity for optimal gRNA design. Proc Natl Acad Sci 116(18):8693–8698
Zhu LJ, Holmes BR, Aronin N, Brodsky MH (2014) CRISPRseek: a bioconductor package to identify target-specific guide RNAs for CRISPR-Cas9 genome-editing systems. PLoS One 9(9):e108424
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Appendices
Appendix 1: List of Useful Bioinformatics Tools and Databases for Gene Modification Research
Tool | Description | Link |
|---|---|---|
“AlleleID® is a comprehensive desktop tool designed to address the challenges of bacterial identification, pathogen detection or species identification” | http://premierbiosoft.com/bacterial-identification/index.html | |
It is an Oligo and cDNA Microarray Design Software. “It designs probes for SNP detection, microarray gene expression and gene expression profiling. In addition, comprehensive support for tiling arrays and resequencing arrays is available” | ||
Autoprime is a very useful software for designing Reverse Transcription Real Time PCR (Q-RT-PCR) primers that are specific to the exon-intron boundaries | ||
“Beacon Designer™ automates the design of real time primers and probes” | ||
The site harbors a number of biocomputational online tools (Cleaner, Translator, NetPlasmit, Aligner, PatSearch, etc. for nucleotide and protein sequences) and half a dozen of software | ||
“BioEdit is a biological sequence alignment editor written for Windows 95/98/NT/2000/XP/7.” One can download and then work with the molecular sequences for alignment, restriction mapping, RNA analysis, translation, graphical viewing of electropherogram, etc. | ||
Basic local alignment search tool, provided by NCBI | ||
Cas-Database | Cas-Database is a genome-wide gRNA library design tool for Cas9 nucleases from Streptococcus pyogenes (SpCas9) | |
Cas-Designer | A bulge-allowed quick guide-RNA designer for CRISPR/Cas-derived RGENs | |
CINEMA stands for Color INteractive Editor for Multiple Alignments. It is a free software for sequence alignment with color editor | ||
“Click2Drug contains a comprehensive list of computer-aided drug design (CADD) software, databases and web services. These tools are classified according to their application field, trying to cover the whole drug design pipeline” | ||
The latest form of Clustal alignment program. It is online and command-line based. The distinguishing feature of Clustal-omega is its scalability, as several thousands of medium to large-sized sequences can be aligned simultaneously. It will also make use of multiple processors, where present. In addition, the quality of alignments is superior to the previous versions. The algorithm uses seeded guide trees and HMM profile-profile progressive alignments | ||
A very popular site for pairwise and multiple sequence alignment. It runs on Windows, Linux/Unix, and Mac operating systems | ||
Latest version of ClustalX 2.0 is provided by “Plate-Forme Bio-Informatique de Strasbourge,” along with detailed instructions (help) for operating ClustalX. Besides, this site also provides online tools (viz. Actin-Related Proteins Annotation server, EMBOSS, Gene Ontology Annotation, SAGE experiment parameters, GPAT, etc,) and database (SRS, BAliBase, InPACT), Documentation (tutorials to elucidate the parameters of Clustal, GCG, EMBOSS, Bioinformatics protocols, etc.) | ||
“The COnsensus-DEgenerate Hybrid Oligonucleotide Primers (CODEHOP) program is hosted by the Fred Hutchinson Cancer Research Center in Seattle, Washington and designs PCR (Polymerase Chain Reaction) primers from protein multiple-sequence alignments” | ||
The Comparative RNA Web (CRW) Site disseminates information about RNA structure and evolution that has been determined using comparative sequence analysis | ||
The Computational Biology and Bioinformatics Group of the Biosciences Division of Oak Ridge National Laboratory provides data and bioinformatics tools for prokaryotic and some eukaryotic genome and related analysis. The tools are “Gene Channel,” “Generation Microbial Gene Prediction System,” “Microbial Gene Prediction System Internet Linked,” “Genome Analysis Pipeline,” etc. | ||
“CRDD (Computational Resources for Drug Discovery) is an important module of the in silico module of OSDD. The CRDD web portal provides computer resources related to drug discovery on a single platform. Following are major features of CRDD” | ||
The tool “compute pI/Mw is a tool which allows the computation of the theoretical pI (isoelectric point) and Mw (molecular weight) for a list of UniProt Knowledgebase (Swiss-Prot or TrEMBL) entries or for user entered sequences” | ||
COSMID | A Web-based tool for identifying and validating CRISPR/Cas Off-target sites | |
“CPHmodels 3.2 is a protein homology modeling server. The template recognition is based on profile-profile alignment guided by secondary structure and exposure predictions” | ||
CRISPR gRNA Design tool | CRISPR gRNA Design tool lets you design gRNA(s) to efficiently engineer your target and minimize off-target effects using ATUM Scoring Algorithms | |
CRISPR multitargeter | CRISPR MultiTargeter is a web-based tool for automatic searches of CRISPR guide RNA targets | |
CRISPRdb | It enables the easy detection of CRISPR in locally produced data and consultation of CRISPRs present in the data base | |
CrisprGE | CrisprGE is a central hub of CRISPR-based genome editing | |
This page is maintained by CSIR and harbors the software/tools developed for bioinformatics analysis | ||
The Database for Annotation, Visualization and Integrated Discovery (DAVID) v6.7 “provides a comprehensive set of functional annotation tools for investigators to understand biological meaning behind large list of genes” | ||
“Swiss-PdbViewer (aka DeepView) is an application that provides a user friendly interface allowing to analyze several proteins at the same time. The proteins can be superimposed in order to deduce structural alignments and compare their active sites or any other relevant parts. Amino acid mutations, H-bonds, angles and distances between atoms are easy to obtain thanks to the intuitive graphic and menu interface” | ||
One can calculate the GC content of a nucleotide sequence | ||
Dotlet is a free online software used as a tool for diagonal plotting of sequences | ||
Dot-plot(+) software is used to identify the overlapping portions of two sequences and to identify the repeats and inverted repeats of a particular sequence | http://bip.weizmann.ac.il/education/materials/gcg/dotplot.html | |
Dotter is a graphical dotplot program for detailed comparison of two sequences. It runs on MAC, Linux, Sun solaris, and Windows OS | ||
A wonderful site that harbors a number of drug designing applications for smart mobiles | ||
This webpage maintains several entries to drug designing. One can learn and make use of these software/links | ||
The European Molecular Biology Open Software Suite (EMBOSS) “is a free Open Source software analysis package specially developed for the needs of the molecular biology (e.g. EMBnet) user community.” Some of the applications are prophet (Gapped alignment for profiles), infoseq (Displays some simple information about sequences), water (Smith-Waterman local alignment), pepstats (Protein statistics), etc. | ||
“The Ensembl project produces genome databases for vertebrates and other eukaryotic species, and makes this information freely available online” | ||
“This tool takes a list of variant positions and alleles, and predicts the effects of each of these on overlapping transcripts and regulatory regions annotated in Ensembl. The tool accepts substitutions, insertions and deletions as input” | ||
RNAi construct designer | ||
“ESyPred3D is an automated homology modeling program. The method gets the benefit of the increased alignment performances of an alignment strategy that uses neural networks” | https://www.unamur.be/sciences/biologie/urbm/bioinfo/esypred/ | |
A resource portal supported by Expert Protein Analysis System and Swiss Institute of Bioinformatics for analyzing bioinformatics data | ||
It is an online tool that “allows the translation of a nucleotide (DNA/RNA) sequence to a protein sequence” | ||
“ExPASy is the SIB Bioinformatics Resource Portal which provides access to scientific databases and software tools (i.e., resources) in different areas of life sciences including proteomics, genomics, phylogeny, systems biology, population genetics, transcriptomics etc.” | ||
This server is hosted by the University of Virginia, USA. It harbors a multiple online software for sequence (nucleic acid and amino acid) comparison, local and global alignment, hydropathy plotting, and protein secondary structure prediction | ||
“FastPCR is an integrated tool for PCR primers or probe design, in silico PCR, oligonucleotide assembly and analyses, alignment and repeat searching.” This program can be downloaded and run on PCs | ||
“Galaxy is an open, web-based platform for data intensive biomedical research. Whether on the free public server or your own instance, you can perform, reproduce, and share complete analyses” | ||
GAS is UNIX or DOS-based downloadable, command-line oriented “integrated computer program designed to automate and accelerate the acquisition and analysis of genomic data” | ||
It is a commercial product. “GelCompar II consists of the Basic Software and five modules: Cluster analysis, Identification & Libraries, Comparative Quantification and Polymorphism Analysis, Dimensioning techniques & Statistics, and Database Sharing Tools” | https://www.applied-maths.com/modules-and-features-gelcompar-ii | |
For analyzing 1D Gel | ||
The “Gel-Quant” software is used to analyze one-dimensional gel images. The gel image is saved in “bitmap” format, following electrophoresis and scanning the gel | ||
“GeneFisher is an interactive web-based program for designing degenerate primers.” The underlying assumption is “assumption that genes with related function from different organisms show high sequence similarity, degenerate primers can be designed from sequences of homologues genes.” This assumption “leads to isolation of genes in a target organism using multiple alignments of related genes from different organisms” | ||
GeneCopoeia offers comprehensive tools for microRNA (miRNA) functional analysis so researchers can detect, express, validate, or knockdown microRNA of interest confidently. All known human, mouse, and rat microRNA in miRBase covered | ||
“geneid is a program to predict genes in anonymous genomic sequences designed with a hierarchical structure” | ||
This site contains description and links to various sites pertaining to Protein Secondary Structure. It is a hub for getting a quick look at several servers and metaservers that harbor databases and/or tools for prediction of protein secondary structures | ||
GeneMark is a “family of gene prediction programs developed at Georgia Institute of Technology, Atlanta, Georgia, USA” | ||
The site harbors “geneid” program which is used to “predict genes, exons, splice sites and other signals along a DNA sequence.” This site is also hyperlinked with “Gene prediction on whole genome” which is a “precomputed whole genome prediction data sets” | ||
“The GenomeTools genome analysis system is a free collection of bioinformatics tools (in the realm of genome informatics) combined into a single binary named gt. It is based on a C library named “libgenometools” which consists of several modules” | ||
“GenomePRIDE is primer design program that designs PCR primers or long oligos on an annotated sequence” | ||
GENSCAN is a freely available software used for “identification of complete gene structures in genomic DNA.” Genscan can be used “for predicting the locations and exon-intron structures of genes in genomic sequences from a variety of organisms” | ||
Glimmer (Gene Locator and Interpolated Markov ModelER) is a system for finding genes in microbial DNA, especially the genomes of bacteria, archaea, and viruses. Glimmer uses interpolated Markov models (IMMs) to identify the coding regions and distinguish them from noncoding DNA | http://www.cbcb.umd.edu/software/glimmer/glimmer2.jun01.shtml | |
The greengenes web application provides access to the current and comprehensive 16S rRNA gene sequence alignment for browsing, blasting, probing, and downloading. The data and tools presented by greengenes can assist the researcher in choosing phylogenetically specific probes, interpreting microarray results, and aligning/annotating novel sequences | https://www.ccg.unam.mx/~vinuesa/Using_the_GreenGenes_and_RDPII_servers.html | |
Homology detection and structure prediction by HMM-HMM: used for sequence database searching and structure prediction. It is fast enough and more sensitive in finding remote homologs. HHpred performs pairwise comparison of profile hidden Markov models (HMMs). It can produce pairwise query-template sequence alignments, merged query-template multiple alignments and 3D structural models calculated by the MODELLER software from HHpred alignments | ||
“HMMgene is a program for prediction of genes in anonymous DNA.” “The program predicts whole genes, so the predicted exons always splice correctly. It can predict several whole or partial genes in one sequence, so it can be used on whole cosmids or even longer sequences” | ||
To synthesize antisense oligos for a specific target sequence of interest | https://www.idtdna.com/pages/products/functional-genomics/antisense-oligos | |
I-TASSER is an online bioinformatics platform for predicting protein structure vis-à-vis function. It has been developed by Zhang Lab (University of Michigan). It has topped in the CASP ranking of structure prediction during the years 2007–2010 | ||
It is a “multiple alignment editor written in Java.” It is used in EBI Clustalw, Pfam protein domain database; however, it is “available as a general purpose alignment editor and analysis workbench” | ||
Online free tool for finding local alignment between two sequences (provided in stipulated input format, viz. plain text without header line, Swiss-Prot ID, TrEMBL ID, EMBL ID, EST ID, etc.) | ||
“LAMP Designer designs efficient primers for Loop-Mediated Isothermal Amplification assays, that amplify DNA and RNA sequences at isothermal conditions, eliminating the necessity of a PCR setup” | ||
This link enables you to download Multiple Alignment Construction and Analysis Workbench (MACAW) software. This program is used for “locating, analyzing, and editing blocks of localized sequence similarity among multiple sequences and linking them into a multiple alignment” | http://en.bio-soft.net/format/MACAW.html | |
“MAFFT is a multiple sequence alignment program for unix-like operating systems. It offers a range of multiple alignment methods, L-INS-i (accurate; for alignment of <~200 sequences), FFT-NS-2 (fast; for alignment of <~10,000 sequences)” | ||
Java platform-based online software to map the RE sites on a target sequence | ||
“MethPrimer is a program for designing bisulfite-conversion-based Methylation PCR Primers” | ||
It is a very useful site for designing primers for methylation PCR (Denatured, single-stranded DNA (ssDNA) is modified with sodium bisulfite “followed by PCR amplification using two pairs of primers, with one pair specific for methylated DNA; the other unmethylated DNA”) | ||
“mGene is a computational tool for the genome-wide prediction of protein coding genes from eukaryotic DNA sequences” | ||
The microRNA body map is a repository of RT-qPCR miRNA expression data and functional miRNA annotation in normal and diseased human tissues | ||
This website provides access to 2003 and 2005 miRNA-Target predictions for Drosophila miRNAs | ||
Predicted microRNA targets and target downregulation scores. Experimentally observed expression patterns | ||
miRNAMap 2.0 is a collection of “experimental verified microRNAs and experimental verified miRNA target genes in human, mouse, rat, and other metazoan genomes” | ||
This site maintains a number of online bioinformatics programs (assembly, database, display, hmm, phylogeny, protein, sequence, structure, etc.), workflows (alignment, db, phylogeny), and tutorial | http://www.mybiosoftware.com/mobyle-1-0-4-integration-bioinformatics-software-databanks.html | |
It is a database for “comparative protein structure models.” The pipeline used is ModPipe | ||
The homology modeling of Protein 3D structures can be done using downloadable software “MODELLER.” It can also be used for the following protein structure-based applications: databases search for amino acid sequences, sequence and structural alignments clustering, de novo modeling of structural loops, model-optimization against user-defined objective function, and so on | ||
It harbors “experimentally resolved structures of proteins, RNA, and DNA, derived from the Protein Data Bank (PDB), with value-added features such as explicit chemical graphs, computationally identified 3D domains (compact substructures) that are used to identify similar 3D structures, as well as links to literature, similar sequences, information about chemicals bound to the structures” | ||
A handy package for analyzing sequence data for pair-wise and multiple sequence alignment, phylogenetic tree (include neighbor-joining, maximum parsimony, UPGMA, maximum likelihood and minimum evolution based) construction, and estimation of evolutionary parameters | ||
Maintains links to several platforms, pipelines, libraries, software for visualization as well as software for proteomic data analysis | ||
This software is RE site mapper, hosted by New England Biolabs | ||
It is an efficient primer analysis software that can be used for determining the features of the secondary structures of the generated primer sequences | ||
Explores and utilizes several bioinformatics tools | ||
The secondary structures produced by the primer(s) can be checked, and the Gibbs free energy required to produce these structures can be calculated using online Oligo Analyzer Version 3.1 (of IDT) | ||
Calculates the melting temperature of the oligos | https://worldwide.promega.com/resources/tools/biomath/tm-calculator/ | |
“OLIGO Primer Analysis Software is the essential tool for designing and analyzing sequencing and PCR primers, synthetic genes, and various kinds of probes including siRNA and molecular beacons. Based on the most up-to-date nearest neighbor thermodynamic data, Oligo’s search algorithms find optimal primers for PCR, including TaqMan, highly multiplexed, consensus or degenerate primers. Multiple file batch processing is possible. It is also an invaluable tool for site directed mutagenesis” | ||
Calculates base-count, thermodynamic properties (ΔS and ΔH), Tm, and GC% values of a given oligo | ||
“The program helps to design primer combinations given one fixed primer” | ||
“The ORF Finder (Open Reading Frame Finder) is a graphical analysis tool which finds all open reading frames of a selectable minimum size in a user’s sequence or in a sequence already in the database” | ||
It is a very useful site to learn about the pros and cons of factors affecting PCR | ||
“The pedant genome database provides exhaustive automatic analysis of genomic sequences by a large variety of bioinformatics tools” | ||
This online tool of ExPASy “PeptideMass cleaves a protein sequence from the UniProt Knowledgebase (Swiss-Prot and TrEMBL) or a user-entered protein sequence with a chosen enzyme, and computes the masses of the generated peptides. The tool also returns theoretical isoelectric point and mass values for the protein of interest” | ||
“PHYLIP is a free package of programs for inferring phylogenies. It is distributed as source code, documentation files, and a number of different types of executables” | ||
Protein Homology/AnalogY Recognition Engine (PHYRE) is a non-commercial, very popular online protein structure prediction (homology modeling) server. The user friendly GUI is very helpful for the novice in the field of protein structure prediction | ||
The tool “searches for predicted microRNA targets in mammals” | ||
Primer Premier is one of the “most comprehensive software to design and analyze PCR primers.” Primers can be designed for standard PCR, SNP genotyping assays, multiplexing assays, along with checking the secondary structures of the designed primers | ||
It is a freely available online software for designing primers and probe from a DNA sequence. It is a very popular software due to availability of several parameters to design primers with high specificity and accuracy | ||
Extensively used for designing primer and checking the specificity of a given primer | ||
Online primer designing tool provided by IDT | ||
“Primo Degenerate 3.4 designs PCR primers based on a single peptide sequence or multiple alignments of proteins or nucleotides. For degenerate primers, the probability of binding to the target is proportional to the effective concentration of the specific primer” | ||
It is another online primer designing software. Its notable feature is that it can reduce background noise by exercising check on mispriming on non-target DNA sequence. It also “introduces a batch mode option for high throughput PCR primer design” | ||
A java-enabled online primer designing tool | ||
Promoter2.0 predicts transcription start sites of vertebrate PolII promoters in DNA sequences. It has been developed as an evolution of simulated transcription factors that interact with sequences in promoter regions. It builds on principles that are common to neural networks and genetic algorithms | ||
This site maintains links for different software and tools (viz. PromScan, SCOPE, Promoser, Arnold, WebGesTer) for scanning, predicting promoters and transcription terminators in Eukaryotes and Prokaryotes | ||
PDB is an “information portal to biological macromolecular structure.” “The PDB archive contains information about experimentally-determined structures of proteins, nucleic acids, and complex assemblies” | ||
This site contains links to several software for “calculating and displaying the 3-D structure of oligosaccharides and proteins. With the two protein analysis sites the query protein is compared with existing protein structures as revealed through homology analysis” | ||
“ProtParam is a tool which allows the computation of various physical and chemical parameters for a given protein stored in Swiss-Prot or TrEMBL or for a user entered sequence. The computed parameters include the molecular weight, theoretical pI, amino acid composition, atomic composition, extinction coefficient, estimated half-life, instability index, aliphatic index and grand average of hydropathicity (GRAVY)” | ||
“ProtScale allows you to compute and represent the profile produced by any amino acid scale on a selected protein” | ||
It is online software that applies QUARK algorithm for ab initio protein folding vis-à-vis structure prediction. It is another eminent online tool of Zhang lab that has secured esteemed ranking in CASP | ||
Another efficient protein structure prediction server that predicts the secondary and 3D protein structure. Besides, it also predicts solvent accessibility and disordered regions, and assigns the following confidence scores to indicate the quality of a predicted 3D model. It has been developed by Xu Group of Toyota Technological Institute at Chicago. RaptorX-Binding, another tool available in the homepage of RaptorX, is used for model-assisted protein binding site prediction | ||
RasMol is a molecular visualization tool for protein in 3-dimension | ||
“This site is provided for the convenience of users of RasMol and developers of open source versions of RasMol” | ||
“PCR Designer for Restriction Analysis of Sequence Mutations” | ||
Online tool for conversion of sequence format | ||
Online, freely available tool for mapping restriction endonuclease sites on a DNA sequence | ||
The RNAfold web server will predict secondary structures of single-stranded RNA or DNA sequences. Current limits are 7500 nt for partition function calculations and 10,000 nt for minimum free energy only predictions | ||
RNAhybrid is a tool for finding the minimum free energy hybridization of a long and a short RNA | ||
RNAiAtlas provides a siRNA oligonucleotide data from different sources and companies like Dharmacon (ThermoFisher), Qiagen, and Ambion, esiRNA for humans, and visualizes interactions between siRNA oligo and predicted off-target | https://www.hsls.pitt.edu/obrc/index.php?page=rna_interference | |
A designing tool for siRNA | ||
Robetta (Beta Version) of Baker Lab, Washington, USA, is a full-chain protein structure prediction tool. It can be used both for ab initio and comparative approaches for protein structure prediction | ||
An array of online tools (dPORE-miRNA, TcoF, PROMEX, etc.) are available which are maintained by South African National Bioinformatics Institute | ||
“The Biology WorkBench is a web-based tool for biologists. The WorkBench allows biologists to search many popular protein and nucleic acid sequence databases. Database searching is integrated with access to a wide variety of analysis and modeling tools, all within a point and click interface that eliminates file format compatibility problems” | ||
“These are a collection of protein secondary structure analysis and information sites” | ||
A suite available for almost all possible manipulation work that can be done with a given DNA or amino acid sequence, viz. Format change, Sequence splitting, CpG island detection, ORF finding, Pair-wise alignment, RE-Digestion, in silico mutation, etc. | ||
sgRNA Designer | This tool ranks and picks candidate CRISPRko sgRNA sequences for the targets provided, while attempting to maximize on-target activity and minimizing off-target activity | http://www.broadinstitute.org/rnai/public/analysis-tools/sgrna-design |
sgRNAcas9 | The BiooTools (Biological Online Tools) website is devoted to provide services to assist researchers design specific and efficient CRISPR sgRNA, primer pairs for detecting small ncRNA expression, such as miRNA, piRNA, and siRNA | |
“SIDDbase-WS is a SOAP based Web Service” that “provides interoperable access to the SIDD software, and access to the repository of stored results from calculations previously performed on complete bacterial genomes” | ||
The siDESIGN Center is an advanced, user-friendly siRNA design tool, which significantly improves the likelihood of identifying functional siRNA. One-of-a-kind options are available to enhance target specificity and adapt siRNA designs for more sophisticated experimental design | http://www.thermofisher.com/order/genome-database/browse/sirna/keyword/siDESIGN+center | |
A stand-alone program designed to run on Unix-based system. It is used for aligning an expressed DNA sequence with a genomic sequence, allowing for introns | ||
An online tool to predict secondary structure of protein | ||
It is “an exceptional tool for drawing publication and vector catalog quality plasmid maps, carrying out restriction analysis and designing cloning experiments” | ||
A short introduction to siRNA Designing Steps | ||
Online siRNA designing tool from Invitrogen | ||
InvivoGen’s siRNA Wizard™ is a software designed to help you select siRNA/shRNA sequences targeting your gene(s) of interest. This program selects siRNA/shRNA sequences that match criteria suggested by studies of RNA interference and which will have the best expression rate in psiRNA vectors | ||
It is an online protein Secondary structure prediction tool | ||
A method to identify potential splice sites in (plant) pre-mRNA by sequence inspection using Bayesian statistical models | ||
It performs several statistical analysis of the physiochemical properties and other features of the protein sequence, viz. compositional analysis, charge distributional analysis, distribution of other amino acid types, repetitive structures, multiplets, periodicity analysis | ||
This is the structural bioinformatics-related page maintained by Imperial College London. This site can be used for several purposes, viz. “analysis of protein structure and function with the aim of deriving evolutionary insights, modelling and comparison of biology networks to provide insights into Systems Biology, modelling of the activity and toxicity of small molecules as an aid to the design of novel drugs” | ||
Harbors links to several software for docking | ||
“The SIB Swiss Institute of Bioinformatics is an academic, non-profit foundation recognised of public utility.” SIB “provides high quality bioinformatics services to the national and international research community” | ||
Tree-based Consistency Objective Function For alignment Evaluation (T-Coffee) is another popular multiple sequence alignment program, developed by Cedric Notredame, CRG Centro de Regulacio Genomica (Barcelona). It allows combining results obtained from several alignment methods. The URL is http://www.ebi.ac.uk/Tools/msa/tcoffee/. The default output format is Clustal, while it accepts sequences in PIR and FASTA format | ||
It is an online primer designing software, hosted by UCSC, that allows users to design primers specific to various types of templates, viz. overlapping amplicons on a template, primers around SNP (in a GenBank), primers flanking exons and cDNA | ||
It is a Java-based free online software, to translate a given input DNA sequences and display one (at a time) of the six possible reading frames according to the selection made by the user. It also displays the graphical output for all the six reading frames together | ||
It is an interactive genome browser dedicated to human genome sequence | ||
The likelihood of secondary structure formation by the single-stranded target is checked by UnaFold software of IDT (freely available online) | ||
It is a website for universal primer designing | ||
This page harbors several “links and notes regarding bioinformatics.” This is a very useful link since a user can get link to almost all aspects of bioinformatics resources | ||
This site of Virginia Bioinformatics Institute maintains several tools for bioinformatics analysis, viz. “Analysis of Dynamic Algebraic Models,” “Complex Pathway Simulator,” “Genome Reverse Compiler,” etc. | ||
This page maintains a “list of in silico drug design online services, standalone and related databases. It is maintained by Dr. B. Villoutreix, research director at the French National Medical Research Institute (Inserm)” | ||
A simple tool for primer designing for PCR or sequencing | ||
Another RE site detection software (online, free) for linear and circular DNA | ||
This site maintains several online “tools for prediction and analysis of protein-coding gene structure” | ||
WGE | A website that provides tools to aid with genome editing of human and mouse genomes | |
What If “is a versatile molecular modelling package that is specialized on working with proteins and the molecules in their environment like water, ligands, nucleic acids, etc.” The web interface provides a number of tools, viz. Structure validation, Residue analysis, Protein analysis, 2-D graphics, 3-D graphics, Hydrogen (bonds), Rotamer related, Docking, Crystal symmetry, mutation prediction, etc. | ||
Yet Another Scientific Artificial Reality Application (YASARA) is used for predicting the rotamers (protein side chain conformations) starting with single point mutations to complete homology models of proteins |
Appendix 2: List of Commercial and Non-profit Sources of CRISPR/Cas Reagents
Resource | Description | Link |
|---|---|---|
Addgene CRISPR plasmids | A collection of CRISPR plasmids and reagents | |
Beam Therapeutics: Upleveling CRISPR’s Precision by Targeting Specific Bases | Beam Therapeutics, a company co-founded recently by leading CRISPR researchers Feng Zhang, David Liu, and J. Keith Young, is developing more precise versions of the CRISPR technology which can effectively swap one base for another in the genome without cutting the DNA or RNA | |
Caribou Biosciences: Using CRISPR to Impact Several Industries | Caribou Biosciences (@CaribouBio) is one of the companies using CRISPR technology developing tools that provide transformative capabilities to therapeutics, biological research, agricultural biotechnology, and industrial biotechnology | |
CRISPR Kits | Synthego’s CRISPR kits offer economical access to fully synthetic RNA for high fidelity editing and increased precision in genome engineering | |
CRISPRflydesign (Bullock Lab) | Offers Cas9 transgenic stocks | |
Editas Medicine: Using CRISPR to Target Point Mutations in Serious Genetic Disorders | Editas Medicine (@editasmed) is targeting mutations that cause serious genetic diseases and hopes to modify and fix these gene mutations using CRISPR | |
eGenesis: Using CRISPR to Improve Organ Transplants | eGenesis (@eGenesisBio) is pioneering an especially interesting application of CRISPR-Cas9 technology in the field of human therapeutics. This company is reviving the idea of xenotransplantation, i.e., animal-to-human organ transplants | |
FlyCas9 (Ueda Lab) | Provides reagents, protocols, and online tools for genome engineering by the designer nuclease Cas9 in Drosophila | |
flyCRISPR (O’Connor-Giles Lab, Wildonger Lab, and Harrison Lab) | Fly CRISPR resources | |
Goldstein Lab CRISPR | A genome engineering resource for the Caenorhabditis elegans research community | |
Inari Agriculture: Using CRISPR to Develop “Customized Seeds” | Inari Agriculture is an agro-biotechnology company that is revolutionizing the agricultural industry through transformational plant breeding technology. Inari uses CRISPR technology to develop seeds with traits optimized to grow best in local conditions | |
Inscripta: Increasing CRISPR’s Reach | Inscripta (@InscriptaInc) is a Colorado-based CRISPR biotech company that is revolutionizing commercially available CRISPR-associated nucleases. Inscripta’s next-generation CRISPR nucleases include natural and synthesized versions of “MADzymes,” a nomenclature inspired by the biodiversity found on the island of Madagascar | |
Intellia Therapeutics: Using Genome Editing for Personalized Disease Treatment | Intellia Therapeutics (@intelliatweets) aims to produce a new class of therapeutic products using a simplified manufacturing process. The company develops CRISPR-based solutions for personalized and curative treatments, and its current in vivo studies are focused on the use of Lipid Nanoparticles (LNPs) for delivery of the CRISPR/Cas9 complex to the liver | |
Joung Lab CRISPR | A genome engineering resource for zebrafish research community | |
Ligandal (@ligandal), one of the companies using CRISPR based in San Francisco, has developed new technology which streamlines the in vivo delivery mechanisms for CRISPR, RNA, and other genetic tools. Ligandal has developed next-generation, non-viral protein-based biomaterials to effectively deliver gene therapy materials | ||
Mammoth Biosciences: Using CRISPR to Advance Clinical Diagnostic | Mammoth Biosciences (@mammothbiosci) has capitalized on CRISPR’s unique ability to accurately find and bind to specific sequences of DNA. This company has created the first CRISPR-mediated platform for human disease detection. Their innovative point of care test allows for easy and affordable multiplexed detection of RNA/DNA sequences associated with disease | |
NTrans: Helping CRISPR Edit All Cell Types | NTrans Technologies (@NtransTech), a CRISPR technology company based in the Netherlands, is working to ensure genome engineering can be performed in all cell types. NTrans pioneered a cellular uptake mechanism which circumvents the problems with delivery of CRISPR components for therapeutic purposes | |
OxfCRISPR (Liu Lab) | Oxford Fly CRISPR Resources | |
Pairwise Plants: Using CRISPR to Grow New Varieties of Crops | Pairwise Plants (@PairwisePL) intends to create new crops and modify existing ones using gene editing technology such as CRISPR. The goal is to also assist farmers by providing them with new varieties of crops that require less resources to grow | |
Plantedit: Increasing the Worldwide Food Supply using CRISPR | Plantedit (@plantedit) is an Ireland-based CRISPR startup company aiming to produce “DNA-free” non-transgenic sustainable plant products in an attempt to introduce genome editing to food supply enhancement in a regulatory-free manner. The company focuses on creating modified plants that do not contain any foreign genetic material with a goal to meet the ever-increasing demand for “designer” crops while circumventing both the general aversion to ingesting non-plant-based DNA or RNA and the regulatory fences around traditional “GMO.” | |
Synthetic Genomics: Harnessing CRISPR to Create Sustainable Energy | Synthetic Genomics (@SynGenomeInc) manipulates microalgae for sustainable oil production. Partnering with Exxon Mobil, Synthetic Genomics identified 20 transcription factors thought to be negative regulators of lipid production in microalgae. The company then applied CRISPR-Cas9 to insert loss of function mutations in 18 of the 20 genes. They report a 200% increase in oil production from one of the modified microalgae species with little effect on growth, marking a key advancement in renewable energy biofuels | |
transOMIC | transOMIC offers reagents for CRISPR Cas9 gene editing, shRNA constructs, and cDNA and ORF clones | |
Zhang Lab Genome Engineering | CRISPR genome engineering resources website |
Rights and permissions
Copyright information
© 2022 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Ahmad, A. et al. (2022). Bioinformatic Tools in CRISPR/Cas Platform. In: Ahmad, A., Khan, S.H., Khan, Z. (eds) The CRISPR/Cas Tool Kit for Genome Editing. Springer, Singapore. https://doi.org/10.1007/978-981-16-6305-5_3
Download citation
DOI: https://doi.org/10.1007/978-981-16-6305-5_3
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-6304-8
Online ISBN: 978-981-16-6305-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)