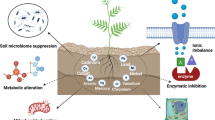
Abstract
The unprecedented augment in the concentration of diverse contaminants in the environment has grim impacts on the ecological balance of our ecosystem. Soil, being a major sink, holds up the maximum load of environmental contaminants. Heavy metals and petroleum hydrocarbons are the most common pollutants present in the soil. Plant growth promoting rhizobacteria (PGPR)-assisted phytoremediation is one of the competent methods for removal of pollutants, which has proven its efficiency in reclamation of contaminated soils. PGPR are bacteria that reside in close association with plant roots and facilitate growth and development of plants by influencing their physiological and metabolic activity. Rhizobacteria are known to amplify the effectiveness of phytoremediation by modulating contaminants transportability and accessibility to the plant via acidification, chelating agents, solubilization of phosphate, and redox changes. This chapter aims to explore the role of rhizomicrobiome in the phytoremediation of heavy metal- and petroleum-contaminated soils, the successful commercialization of PGPR, and the insights into the recent advances in PGPR research.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
4.1 Introduction
The persistence of heavy metals and petroleum hydrocarbons is one of the serious environmental concerns demanding attention of researchers throughout the world. The plants which can accumulate metal or tolerate metals stress have been used for phytoremediation in metal-polluted soils. Phytoremediation is the use of green plants for removal of both inorganic and organic pollutants from air, water, and soil (McCutcheon and Jorgensen 2008). The name phytoremediation is obtained from the Greek word phyto, i.e. plant, and the Latin word remedium, i.e. cure. Its usage in scientific literature for the first time has been traced to paper written by Cunningham and Berti in 1993 (Novo et al. 2018). Speight (2020) rightly states that the description of phytoremediation can be redefined to include the utilization of green plants and the allied microorganisms, along with appropriate soil amendments and agronomic methods to either contain, eliminate, or render lethal environmental pollutants undamaging.
Many phytoremediation projects have been carried out worldwide to mitigate contaminants like pesticides, metals, crude oil, and explosives. The phytoremediation practice utilizes specific plants with roots that can absorb contaminants over time. Many plants such as hemp, mustard, pigweed, and alpine pennycress have the ability to hyper-accumulate contaminants from polluted sites (Speight 2020). Phytoremediation has immense potential as a natural, low cost, in situ approach driven by solar energy to moderately treat polluted sites spreading over large areas. However, the plants have to be cautiously selected depending on the type contaminants (Schwitzguébel 2017). The added advantage of phytoremediation over other technologies is that various kinds of nutrients, organic materials, and oxygen are supplemented to the soil through metabolic processes of plant and microbes. This enhances the value and consistency of remediated sites, stabilizes soil, and checks wind and water erosion as well (Schwitzguébel 2017). It can be concluded that this is an esthetically pleasing technology which helps in reducing erosion, increasing biodiversity, and fixing atmospheric carbon dioxide (Cunningham and Berti 1993). Phytoremediation techniques should, however, avoid the use of food crops for cleanup. The use of ornamental plants reduces the possibility of metals passing into the food chain along with the extra benefit of improving the environment’s esthetics and producing extra earnings, together with added job opportunities from cut-flower trading and/or travel industry (Nakbanpote et al. 2016).
A survey of recent literature brings to light the numerous advantages of plant growth–promoting rhizobacteria (PGPR) to environment, agriculture, landscaping, etc. PGPRs are also known to have the potential to enhance phytoremediation processes (Jing et al. 2007). The application of PGPR for improving metal tolerance of plants is increasingly being utilized these days. Plant roots have restricted capacity to absorb metals from soil, chiefly because metals are not very soluble in the soil solution. Phytoremediation of metal-contaminated soil depends on the rate of uptake of the metal by plants. Also the phytoavailability of metal depends on soil properties and the associated PGPRs. In this chapter we attempt to outline the benefits of PGPR in phytoremediation by enhancing mitigation of petroleum and heavy metal pollutants in the environment. It also focuses on the recent developments that have taken place in decoding the genetics and genomics of PGPR.
4.2 Rhizomicrobiome
The evolution and colonization of terrestrial plants has apparently been antedated by microbiome relationships (Berg et al. 2014). A land plant does not exist individually in nature; rather a consortium of bacteria is generally associated with plants and constitutes a phytomicrobiome (Smith et al. 2017). The phytomicrobiome is of crucial importance in determining the existence of plants, or rather the holobiont. Certain plant–microbe associations (e.g., Cycas, Azolla) are so inseparable that they are known to be symbiotic ubiquitously. Members of phytomicrobiome even ascertain the survival efficiency of plants under conditions of stress. Though microbes are associated with all the major plant structures, microbes associated with rhizosphere constituting a rhizomicrobiome are most elaborated and populous (Backer et al. 2018). Venturi and Keel (2016) define the rhizosphere as a complex zone around the roots of plants with a large population of microorganisms including bacteria, protists, invertebrates, nematodes, and fungi. Bacterial communities in the rhizosphere are called the rhizobacteria. Rhizosphere has much greater amount of bacteria than the bulk soil due to the presence of root exudates such as amino acids and sugars, called rhizodeposition, which provides energy and nutrients for development (Novo et al. 2018). Rhizodeposition may comprise nearly 15% of plant total nitrogen and 10% of photosynthetically fixed carbon (Venturi and Keel 2016).
The complex composition and well-guarded regulation of rhizomicrobiome has assisted land plants against various stresses in due course of evolution (Lundberg et al. 2012; Smith et al. 2015; Zhang et al. 2017). Plant roots secrete exudates of various compositions and signal compounds in order to recruit preferential microbes (Chaparro et al. 2012; Smith et al. 2017). Apart from the considerably controlled regulation by plants, microbes do exhibit facets of self regulation (by virtue of quorum sensing in favorable conditions) depending upon the ecological conditions, which are reciprocated by plants as a mechanism for further regulation (Leach et al. 2017; Ortiz-Castro et al. 2009). This degree of regulation is directly dependent upon the affinity between roots and microbes, i.e., it is much higher for endophytes and rhizospheric bacteria (Backer et al. 2018). The co-evolution of plants and microbes has facilitated certain free-living bacteria such as PGPRs to become endophytes (Bulgarelli et al. 2013). Members of the rhizomicrobiome play pivotal roles in enhancement of plant growth by aiding in nutrient acquisition and assimilation, improving soil texture and modulating the secretion of extracellular molecules that also influence plant stress responses (Backer et al. 2018).
4.2.1 Plant Growth Promoting Rhizobacteria
Rhizobacteria can be characterized as neutral, beneficial, and harmful depending on their outcome on plant growth and development (Huang et al. 2014). Many of them have been aptly called Plant Growth Promoting Rhizobacteria (PGPRs) (Kloepper and Schroth 1978). A bacterium is called a PGPR when it can induce a positive impact on the plant after inoculation. Around, 2–5% of rhizosphere bacteria qualify as PGPR (Goswami et al. 2016). Most PGPRs belong to genera Acinetobacter, Agrobacterium, Arthobacter, Azospirillum, Azotobacter, Bacillus, Bradyrhizobium, Burkholderia, Frankia, Pseudomonads, Rhizobium, Serratia, and Thiobacillus (Glick 1995; Vessey 2003). They are known to aid plant growth by assisting in acquisition of minerals such as nitrogen and phosphorus. They also promote plant growth and development by controlling the plant hormonal balance, eliciting immune responses, mobilizing nutrients, and protecting the plant against pathogens (Glick 2012). The plant-beneficial rhizobacteria can also help to lessen the reliance on harmful fertilizers (Ahemad and Kibret 2014).
Somers et al. (2004) have categorized PGPR according to their activities as:
-
1.
Biofertilizers: ones expanding the accessibility of nutrients to the plant.
-
2.
Phytostimulators: ones that promote plant growth and development by releasing plant growth regulators.
-
3.
Rhizoremediators: ones that break down organic contaminants.
-
4.
Biopesticides: ones that control diseases by producing antibiotics and antifungal metabolites.
Thus we can say that PGPR can directly and indirectly influence plant growth. The direct mechanism involves the synthesis and modulation of phytohormones or the acceleration of resource accumulation including nitrogen fixation, phosphorus bioavailability, and iron sequestration. Indirect mechanism includes biocontrol which is the reduction of the effects of phytopathogens by production of antibiotics and antifungal compounds (Glick 2012; Novo et al. 2018).
4.3 Role of PGPRs in Phytoremediation
The use of plants and allied microbes for elimination of metal pollutants and soil reclamation has both ecological and economic benefits. In general, plant-associated microbes utilize one of these mechanisms to alleviate metal stress to plants (a) bioaccumulation, (b) bioavailability by transformation of metals into soluble form, (c) production of extracellular polymeric substances (EPSs) for binding, and (d) production of iron siderophores. Therefore, a strategy utilizing the combinatorial effect of metal-tolerant plant species along with metal-resistant plant–associated microorganisms will be more efficient. Such a novel in situ approach for bioremediation is known as rhizoremediation and the microorganisms are called heavy metal-tolerant-plant growth promoting (HMT-PGP) microbes (Mishra et al. 2017). It exploits the combined capacities of the roots of plants and allied microbial communities of rhizosphere to tackle heavy metal contamination in soils (Ullah et al. 2015). The knowledge that plants assisted microbes could prove more beneficial in enhancing metal tolerance in plants has opened up new avenues. PGPRs provide metal tolerance by either using one or multiple mechanisms such as bioaccumulation, bioavailability, and production of binding and chelating compounds, and also they enhance plant growth in such conditions by protecting against various types of abiotic and biotic stress. For instance, PGPRs produce a phytohormone indole acetic acid (IAA) which can augment the uptake of metals in the roots of plant (Khan et al. 2009; Tak et al. 2013). These microorganisms are found in abundance in plant rhizosphere, they help to diminish metal buildup in plant tissues and in addition assist in reducing metal bioavailability in soil through a variety of mechanisms. PGPRs release siderophores, organic acids, and plant growth regulators which increase the rate of phytoremediation (Tak et al. 2013). Kloepper et al. (1980) reported that some strains of the Pseudomonas fluorescens-putida act as PGPR by producing extracellular siderophores which are microbial iron transport agents. They probably deprive native microflora of iron and make it less available to them and significantly improve the yield of potato, radish, and sugar beet.
4.3.1 Role of Rhizobacteria in Phytoremediation of Contaminated Soil
4.3.1.1 Metal-Contaminated Soil
The mining and extraction of mineral resources is important for development but frequently causes great harm to neighboring ecosystems (Novo et al. 2018). Industrialization and modern life style have led to drastic pollution of biosphere. Different types of inorganic (heavy metals) and organic (hydrocarbons, volatile organic compounds, and solvents) contaminants are being incessantly released into the environment by mining and industrial activities (Manoj et al. 2020). Soils from mining areas are nutrient deficient and have reduced organic matter, pH, and cohesion and elevated concentration of metals (Novo et al. 2018). Mining waste leaches into aquifers and contaminates agricultural lands and accumulates in plants planted for food or livestock feed; hence can enter food chain (Mendez and Maier 2008). It causes damage to water and soil flora and has lethal impacts on human health because of the mutagenicity, cytotoxicity, and carcinogenicity. Release of untreated industrial waste containing precarious heavy metals such as mercury, arsenic, and cadmium into the water bodies and soil is another source of heavy metal pollution of surrounding soil and water (Nordberg et al. 2009). Aluminum (Al), Cadmium (Cd), Chromium (Cr), Copper (Cu), Lead (Pb), Mercury (Hg), and Zinc (Zn) are the most widespread polluting toxic heavy metals. These heavy metals have been declared as “priority pollutants” by United States Environmental Protection Agency because of their mutagenic and carcinogenic nature. Presence of heavy metals in soil is noxious to most plants as heavy metals ions are absorbed by roots and translocated to shoot; leading to reduction in metabolism and growth of plants (Jing et al. 2007). Soils contaminated with high metal concentration led to reduction in activity of soil microbes and thereby soil fertility (Jing et al. 2007). For example, Cd is recurrently accumulated by chief agricultural crops and its high concentration affects nutrient uptake and inhibits root and shoot growth. Such crop plants with Cd affect the health of animals and humans and negatively affect biodiversity. Cd pollution affects the activity of soil microbial communities, and the contaminated soil may eventually become unusable for crop production (Ma et al. 2011a). Therefore, research on remediation of heavy metals contaminated soils is now the topmost priority for scientists.
Few methods like thermal treatment, acid leaching, excavation and land fill, and electro-reclamation were explored for cleanup of contaminants, but they are time- and cost-consuming (Zubair et al. 2016). Currently, phytoremediation is being accepted as in situ eco-friendly technology (Abou-Shanab et al. 2019). Phytoremediation depends on the fact that numerous plant species have the capability to accrue large quantities of metals in their vegetative as well as reproductive organs. Depending upon the metal accumulating skills and tolerance, plants can be metal sensitive (excluders), having poor metal uptake and transport (indicators) or those with higher uptake efficiency (hyperaccumulators) (Khan et al. 2009). Hyperaccumulator plants have the capacity to endure high level of noxious heavy metal concentration (Ma et al. 2011a). The potential for use of a particular plant for phytoremediation depends on the BCF i.e. bioconcentration factor and the TF i.e. translocation factor. The BCF specify the capability of a plant to take up the contaminant and its accumulation in its tissues. The TF specify the capacity of the plant to transfer contaminants from the root to its aboveground parts (Novo et al. 2018). The efficiency of such plants to uptake and accumulate heavy metals also depends on edaphic factors like soil, temperature, redox potential, cation exchange capacity of the soil particles (CEC), metal bioavailability, pH, aeration, and amount of organic matter and water (Eliana Andrea et al. 2019). Also, plants chosen for the purpose of phytoremediation ought to be fast growing with high biomass production, widespread root system, ability to accumulate the pollutants, and preferably hardy, native species (Manoj et al. 2020). However, high levels of pollutants are also toxic to the plants used for reclamation of affected soil, and phytoremediation by plants alone is a very slow process. Still, this process can be accelerated by the synergistic action of plant and microbes. Plant-Microbe association improves plant development by enabling the sequestration of noxious heavy metals especially by phytostabilization and phytoextraction (Ma et al. 2011a). Rhizobacteria improve adaptation of host plants to altering environment by altering plant cell metabolism, so they can withstand exposure to high concentrations of metals (Welbaum et al. 2010).
The synergistic interaction between rhizobacteria and plants is now being investigated by many workers because of its potential to enhance plant growth, metal uptake, and tolerance during environmental stresses (Ma et al. 2011a). Rhizobacteria not only improve fertility of polluted soil but also enhance the growth and development of plants by exuding plant growth hormones (Zubair et al. 2016). Many plants like Alyssum lesbiacum and Arabidopsis halleri have been documented for phytoremediation as hyperaccumulators of Nickel (Ni) and Zn (Cluis 2004; McNair et al. 2000). Ni and Zn are known as most accumulated metals by different hyperaccumulator species (Pandey and Bajpai 2019). Khan and Bano (2018) have also emphasized the role of PGPRs in remediation of heavy metals by affecting heavy metal portability and accessibility to the plant through acidification, phosphate solubilization, and release of chelating agents. PGPRs used for phytoremediation of metal-contaminated soil in various laboratory and field experiments have been compiled in Table 4.1.
4.3.1.2 Petroleum-Contaminated Soil
Petroleum hydrocarbons symbolize the biggest group of organic pollutants (Hawrot-Paw and Nowak 2012). Phytoremediation of petroleum is proving to be a low-cost and sustainable approach for sustainable waste management technology. This strategy can be useful in petroleum-contaminated soils where other techniques have been unsuccessful. Attempts of remediation of soils polluted with petroleum products with various herbaceous plants including Cynodon dactylon, Digitaria sanguinalis, Cyperus rotundus, Chloris babata, and Pasparlum vaginatum have also been reported (Borowik et al. 2019). However, PGPR-associated plants are much better in withstanding the pressure of growing in the crude oil contaminated soils as compared to the plants without allied PGPR (Gurska et al. 2009). Afzal et al. (2012) also stated the importance of plants and associated microorganisms to remediate petroleum hydrocarbons-contaminated soils. Gurska et al. (2009) have reported the successful establishment of a system using PGPR to enhance phytoremediation of soil contaminated with total petroleum hydrocarbons. Addition of PGPRs to soil supports an active rhizosphere, minimizes plant stress in contaminated soils, causes an increase in root biomass, and promotes degradation of oil contaminants by the plants. They also noted a noteworthy boost in fresh weight and length of shoots in experimental PGPR-associated plants.
Afzal et al. (2012) reported that not only the strains used for inoculum purpose but also the inoculation strategy (seed imbibement and soil inoculation) employed determines bacterial colonization, plant growth advancement, and degradation of hydrocarbon. When the soil contaminated with diesel, where Italian ryegrass was planted, was inoculated with amalgamation of three alkane-degrading strains namely Pantoea species ITSI10, Pantoea species BTRH79, and Pseudomonas species MixRI75, maximum hydrocarbon degradation was achieved as compared to soil where single strain was used. Also soil inoculation method gave better results than seed imbibement method. PGPRs utilized for phytoremediation of petroleum-contaminated soil in various laboratory and field experiments have been compiled in Table 4.2.
4.4 From the Lab to the Field and Commercialization
Certain PGPR mechanisms i.e., nitrogen fixation, phytohormone synthesis, 1-aminocyclopropane-1-carboxylate (ACC) deaminase activity, siderophore production, antibiosis, and phosphate solubilization are the basis of laboratory screening assays designed to develop new PGPR inocula. Under laboratory conditions, these mechanisms are difficult to screen for because of complexity of the mechanisms along with gaps in understanding. Henceforth, results obtained in laboratory conditions are not always replicated under field conditions and vice versa. Consequently, promising strains are rejected due to underperformance at classical laboratory screening scale (Cardinale et al. 2015).
PGPR formulations propound to green alternatives over conventional agrochemicals as they promote plant growth, aid in soil fertility, and suppress phytopathogens without contaminating the environment (Arora et al. 2016). The development of bioformulations includes (Backer et al. 2018):
-
(a)
Isolation of PGPR from rhizospheric soil.
-
(b)
Laboratory screening of traits.
-
(c)
Field trial under different conditions (crop varieties, soil types, seasons, locations) and management practices (use of agrochemicals, etc.)
-
(d)
Assessment of synergistic effects of possible PGPR combinations.
-
(e)
Confirmation of safety against ecotoxicology.
-
(f)
Refining PGPR formulation and knowledge about its texture and storage conditions.
-
(g)
Product registration and approval by regulatory agency of country.
-
(h)
Commercialization.
All these steps are time-consuming, laborious, and costly to perform. To facilitate this process, collaborations among industries, research institutes, and government organizations can play a vital role. Development of bioformulations is a business of intellectual property. Though living creatures and natural products can no longer be patented, formulations and their applications are patentable (Matthews and Cuchiara 2014).
Patenting holds a prominent place between discovery and commercialization of promising PGPR in the field of environmental management. Variovorax paradoxus JHP31 strain (EP2578675A1) was patented by Koga and Masuda (2015) for assisting Cd phytostabilization in the plants of Brassicaceae, Chenopodiaceae, Compositae, Gramineae, Leguminosae, Liliaceae, Polygonaceae, and Solanaceae families. PGPRs such as Achromobacter piechaudii, Agrobacterium tumefaciens, Delftia acidovorans, and Stenotrophomonas maltophilia were patented (Banerjee and Yesmin 2011) for their ability to oxidize elemental sulfur and in turn enhancing plant growth.
Some other PGPRs that have been patented are Microbacterium arabinogalactanolyticum, Microbacterium liquefaciens, and Sphingomonas macrogoltabidus for assisting phytoextraction ability of Alyssum murale (US7214516B2) (Angle et al. 2007).
Highly potent microbes possessing long shelf-life and good colonization rates present a major challenge to commercialization. Colonization rates are largely affected by inoculation and field conditions. PGPR inoculated without a suitable carrier or in amount not enough to compete with native soil microbes are the major challenges to successful rhizosphere colonization (Backer et al. 2018). Additionally, fumigation of soils with broad-spectrum biocidal fumigants during cultivation of high-value crops alters the soil ecology by affecting microflora and their interactions with plants in aiding nutrient acquisition and mobilization (Dangi et al. 2017). Many underlying issues should be tended to for substantial commercialization of PGPR strains such as the following:
-
(a)
Identification of PGPR responsively suitable for particular soil conditions and overcome environmental constraints.
-
(b)
Choosing ideal rhizoinoculation techniques depending upon cultivation conditions (i.e., greenhouse vs. field) and training farmers to apply them efficiently.
-
(c)
Selection of desirable traits-possessing strains.
-
(d)
Uniformity among regulatory agencies of different countries regarding safety and use of PGPB strains.
-
(e)
Knowledge of potential interactions among PGPR and native microflora (other bacteria, algae, and fungi) and the advantages of using them over others.
With view of all above points, PGPR strains that have been commercialized have been enlisted in Table 4.3 (Glick 2012). Progressing from research lab and greenhouse analysis to field assessments and commercialization involves development of new processes to inoculate, formulate, store, and ship these strains. Necessary instructions will need to be imparted to the users of these formulations.
4.5 Genetics and Genomics of Heavy-Metal Resistance in PGPRs
Many plant-associated microorganisms mainly bacteria and fungi are well-known to display plant-growth advancing qualities under heavy metal stress by means of various direct and indirect mechanisms e.g. Pseudomonas sp., Bacillus sp., Arthrobacter, Streptromyces, Methylobacterium, and filamentous fungi such as Trichoderma, Aspergillus, and Fusarium. There have been many genetic studies to evaluate if the heavy metal-resistance and plant growth promoter-producing bacteria found in soils would support phytoremediation. Much of the studies were done in symbiotic rhizobia and has been reviewed by Fagorzi et al. (2018). In Sinorhizobium meliloti CCNWSX0020 genetic mechanisms responsible for Cu resistance were elucidated through transposon mutagenesis combined with RTPCR (Li et al. 2014). The transcriptional analysis of Rhizobium etli revealed the increase in the levels of defense-related genes namely PvWRKY33, PvERF6, and PvPAL2 as well as ABA-synthesis-related gene PvAAO3 following infection with the pathogen (Díaz-Valle et al. 2019). Genetic screening of a cosmid genomic library of Mesorhizobium metallidurans for Cd or Zn endurance revealed the presence of a gene encoding PIB-type ATPase homologous to CadA (Maynaud et al. 2014). The mechanism of arsenite [As(III)] resistance via methylation and successive volatization was characterized by Qin et al. (2006) and the enzyme for this function was encoded by the As(III) S-adenosylmethionine methyltransferase (arsM) genes. Rhizobium leguminosarum bv trifolii, which lacks an endogenous arsM gene, was genetically engineered by using an algal As(III) methyltransferase gene (CrarsM) for arsenic bioremediation and it was able to successfully methylate arsenic reducing toxicity. In Mesorhizobium amorphae genetic mechanism of Cu resistance was investigated by transposon mutagenesis, and CopA was found to be the major determinant (Hao et al. 2015). Sinorhizobium melilotinia was shown express a P1B-5-ATPase in the nodule and its expression is activated by the presence of Ni2+and Fe2+ions (Zielazinski et al. 2013). Genomic analysis of the role of the plant-beneficial function contributing genes (PBFC genes) was probed utilizing the genomes of 25 PGPR species, and it showed favored associations among certain genes engaged in phytobeneficial qualities (Bruto et al. 2014).
Currently, use of a novel phytobacterial strategy that uses genetically engineered plant growth promoting bacteria along with plants seems to be promising approach to mitigate heavy metal stress in plants (Gupta and Singh 2017; Ullah et al. 2015; Ashraf et al. 2017, Tiwari and Lata 2018). Many genes belonging to metal uptake and its regulation, metabolic enzymes, metal chelators, and metal homeostasis can be used as potential target genes for such manipulation. Undoubtedly, these genetically modified microorganisms have better remediation prospective, yet their effect on biomes needs to be studied in detail. Pseudomonas aeruginosa strain Psew-MT, which was genetically modified by expressing metallothioneins to capture Cd2+, showed tolerance along with plant growth-advancing properties (Huang et al. 2016). Similarly, enhanced Cd, Hg, and Silver (Ag) resistance and accumulation were shown by genetically modified Pseudomonas putida KT244 (Yong et al. 2014).
Rhizobium–legume associations have been studied for various reasons in the past and they provide an excellent strategy which can be exploited in reclamation of heavy metal-polluted soils (Pajuelo et al. 2011; Ahemad 2012). For example, a genetically engineered Ensifer medicae MA11 strain having copAB gene from Pseudomonas fluorescens was analyzed for enhanced Cu resistance and reducing toxic effect of Cu in Medicago truncatula (Perez-Palacios et al. 2017; Delgadillos et al. 2015). Four different reports are available for the use of transgenic Meshorhizobium huakuii subsp. rengei strain B3 for Cd bioremediation in association with different plants (Ike et al. 2007, 2008; Sriprang et al. 2002, 2003). Wu et al. (2006) reported the use of genetically engineered Pseudomonas putida strain 06909 for enhanced Cd tolerance in alliance with the host plant Helianthus annuus. In another study, Weyens et al. (2013) analyzed the phytoremediation prospective of willow and its genetically engineered allied bacteria in Cd- and toluene-contaminated soils. In an independent study, the impact of genetically engineered Burkholderia pyrrocinia JK-SH007E1 on microbial communities of soil in the poplar rhizosphere during long-term use as biological control was analyzed (He et al. 2018). Whole genome sequence analysis has also been used to characterize the genetic basis of the PGPR and plant interactions. P. fluorescens Pf-5 is a remarkable organism widely recognized for its use in PGPR for its rhizosphere competence and production of broad range of secondary metabolites and antibiotics. The genome of Pseudomonas fluorescens Pf-5 was sequenced to identify the genetic features and molecular determinants responsible for biocontrol (Paulsen et al. 2005). The genome sequence Pseudomonas psychrotolerans CS51 was determined to understand the plant growth-promoting characteristics under multiple heavy metal stress (Cd, Cu, and Zn), and the existence of genes accountable for cobalt-Cd-Zn resistance, transportation of Ni, and Cu homeostasis was confirmed in the P. psychrotolerans CS51 genome. Genomes of other PGPR strains including the Serratia fonticola strain AU-P3, and Bacillus sp. strain JS, Sinorhizobium meliloti CCNWSX0020 have been sequenced, which is serving to comprehend the correlation among genes and PGPR activities (Devi et al. 2013; Song et al. 2012; Li et al. 2012).
There are few studies that have focused on the role of bacterial consortium in PGPR-mediated beneficial effects (Zolla et al. 2013). A synthetic microbial consortium containing seven 2,4-DNT-degrading microbes affiliated to Bacillus, Burkholderia, Pseudomonas, Ralstonia, and Variovorax species was found to augment root length of Arabidopsis under 2,4-DNT stress (Thijs et al. 2014). In another study, phytoremediation potential of Lupinus luteus was improved when it was inoculated with a PGPR conglomerate inclusive of Bradyrhizobium sp. and two metal resistant bacteria including Ochrobactrum cytisi and Pseudomonas sp. (Dary et al. 2010).
4.5.1 Genetically Engineered PGPRs
There have been numerous attempts to understand the molecular features that define PGPR. But, it has remained largely unsuccessful due the ability of PGPR to occupy different habits, to display alternative/selective ecological niches. Moreover, the genes that are implicated in plant-beneficial functions are also involved in the essential primary metabolism like phosphate solubilization, nif (nitrogen fixation), and phl (phloroglucinol synthesis) or in the secondary metabolic functions like pqq (pyrroloquinoline quinone synthesis). So the role of PGPR in producing plant beneficial properties needs to be experimentally verified under controlled conditions and that too in isolation for each species. That has made the process of identification of plant-beneficial traits and their corresponding genes in PGPR a relatively difficult task.
In the last few decades, there have been various reports on introduction of specific genes accountable for the expression of certain enzymes from microbial species lineally into crop plants, but very few studies have been reported on genetic manipulation in the PGPR for enhancing plant productivity under environmental stress or metal stress (Ullah et al. 2015; Saxena et al. 2019). The transgenic techniques are used to either overexpress or knock down genes playing a crucial function in metal detoxification and tolerance to metal stress like genes encoding metal binding, transport, and chelation (Dhankher et al. 2011; Ullah et al. 2015; Sarwar et al. 2017; Saxena et al. 2019).
The enzyme ACC deaminase, encoded by AcdS gene, is common in bacterial and fungal species in soil. It breaks down ACC, precursor of the plant hormone ethylene, to α-ketobutyrate and ammonium. ACC deaminase enzyme has been recognized in soil bacteria and has been anticipated to play an important function in microbe–plant association by decreasing the harmful impacts of biotic and abiotic stress to plants. The activity of ACC deaminase is one of the most widespread qualities among PGPRs (Glick 2014). ACC deaminase microbes aid allied plants in phytoremediation by biotransformation of poisonous components, rhizodegradation facilitated by root exudates, as well as detoxification of heavy metals that let host plants to sustain under unfavorable conditions. The bacterial AcdS gene has been utilized to generate transgenic plants to improve their tolerance to abiotic and biotic stress. Many genetically modified plants with foreign AcdS gene have been generated to lessen the harmful ethylene levels in plants as reviewed by Saleem et al. (2007). When Mesorhizobium ciceri was exogenously transformed with acdS gene, it showed improved plant performance under salinity stress by enhancing nodulation suggesting the significant function of ACC deaminase in assisting symbiotic interaction under salinity stress (Conforte et al. 2010; Nascimento et al. 2012; Brıgido et al. 2013). However, there is limited information on performance of these transgenic plants under farm conditions due to the environmental risks associated with them.
Thus biotechnological interventions can prove to be promising alternatives for enhancing agricultural productivity through PGPR-mediated beneficial effects. However, since the genetically engineered plant-associated microbes are mainly distributed in the rhizosphere, a detailed experimental validation is required for their use in field conditions. Currently, efforts are underway to understand the beneficial effects of root microbiome on plant productivity and stress endurance. Many differentially regulated genes were recognized in PGPR-treated roots of rice plants through microarray technique (Agarwal et al. 2019). Transgenic plants overexpressing OsASR6 (ABA STRESS RIPENING 6) showed a significant result on the growth of plant and root architecture, which could be the main reason for the positive impact of PGPRs in rice.
4.6 Conclusion
PGPRs play significant roles in assisting plant growth on soils polluted with diverse contaminants and in detoxification of soils. Microbial diversity and their interactions play an essential role in facilitating plant-based degradation of toxins. Therefore, microbiome analysis or a detailed rhiozbiome analysis in PGPR–plant interactions may provide more useful insights. The potential of “omics” technologies (such as genomics, transcriptomics, proteomics, metabolomics, and metagenomics) has to be utilized to get a clear holistic view of the role of various genetic, molecular, and regulatory mechanisms in microbe-assisted phytoremediation. For effectual utilization of genetically engineered PGPR for phytoremediation; well-structured, cost-efficient, and time-efficient tools for a trustworthy forecast of their effectiveness on contaminated sites and their repercussion on biomes required to be discoursed prior to commercialization. Also, assurance needs to be provided regarding safety upon large-scale release of strains, since public acceptance also comes into count.
References
Abou-Shanab RAI, El-Sheekh MM, Sadowsky MJ (2019) Role of rhizobacteria in phytoremediation of metal-impacted sites. In: Bharagava R, Chowdhary P (eds) Emerging and eco-friendly approaches for waste management. Springer, Singapore, pp 299–328. https://doi.org/10.1007/978-981-10-8669-4_14
Afzal M, Yousaf S, Reichenauer TG, Sessitsch A (2012) The inoculation method affects colonization and performance of bacterial inoculant strains in the phytoremediation of soil contaminated with diesel oil. Int J Phytoremediation 14(1):35–47
Agarwal P, Singh PC, Chaudhry V, Shirke PA, Chakrabarty D, Farooqui A et al (2019) PGPR-induced OsASR6 improves plant growth and yield by altering root auxin sensitivity and the xylem structure in transgenic Arabidopsis thaliana. J Plant Physiol 240:153010
Ahemad M (2012) Implication of bacterial resistance against heavy metals in bioremediation: a review. J Inst Integr Omics Appl Biotechnol 3(3):39–46
Ahemad M, Kibret M (2014) Mechanisms and applications of plant growth promoting rhizobacteria: current perspective. J King Saud Univ Sci 26(1):1–20
Ajuzieogu C, Ibiene AA, Herbert OS (2015) Laboratory study on influence of plant growth promoting rhizobacteria (PGPR) on growth response and tolerance of Zea mays to petroleum hydrocarbon. Afr J Biotechnol 14(43):2949–2956
Allamin IA, Halmi M, Yasid NA, Ahmad SA, Abdullah S, Shukor Y (2020) Rhizodegradation of petroleum oily sludge-contaminated soil using Cajanus cajan increases the diversity of soil microbial community. Sci Rep 10(1):4094
Anand R, Grayston S, Chanway C (2013) N2-fixation and seedling growth promotion of lodgepole pine by endophytic Paenibacillus polymyxa. Microb Ecol 66(2):369–374
Angle J, Chaney R, Abou-shanab R, Berkum PV (2007) Bacterial effect on metal accumulation by plant. Patent no. US 7,214,516 B2
Arora NK, Mehnaz S, Balestrini R (2016) Bioformulations: for sustainable agriculture. Springer, Berlin. https://doi.org/10.1016/j.mib.2017.03.011
Ashraf MA, Hussain I, Rasheed R, Iqbal M, Riaz M, Arif MS (2017) Advances in microbe-assisted reclamation of heavy metal contaminated soils over the last decade: a review. J Environ Manag 198:132–143. https://doi.org/10.1016/j.jenvman.2017.04.060
Backer R, Rokem JS, Ilangumaran G, Lamont J, Praslickova D, Ricci E et al (2018) Plant growth-promoting rhizobacteria: context, mechanisms of action, and roadmap to commercialization of biostimulants for sustainable agriculture. Front Plant Sci 9:1473. https://doi.org/10.3389/fpls.2018.01473
Banerjee MR, Yesmin L (2002) Sulfur oxidizing rhizobacteria: an innovative environment friendly soil biotechnological tool for better canola production. In: Proceedings of agroenviron, October 26–29, Cairo, Egypt, pp 1–7
Banerjee MR, Yesmin L (2011) Sulfur-oxidizing plant growth promoting rhizobacteria for enhanced canola performance. Patent no. US 7,517,687 B2
Bashan Y, de Bashan LE (2005) Bacteria/plant growth-promotion. In: Hillel D (ed) Encyclopedia of soils in the environment. Elsevier, Oxford, pp 103–115
Berg G, Grube M, Schloter M, Smalla K (2014) Unraveling the plant microbiome: looking back and future perspectives. Front Microbiol 5:148. https://doi.org/10.3389/fmicb.2014.00148
Bhattacharyya PN, Jha DK (2012) Plant growth-promoting rhizobacteria (PGPR): emergence in agriculture. World J Microbiol Biotechnol 28:1327–1350. https://doi.org/10.1007/s11274-011-0979-9
Bhattacharyya PN, Goswami MP, Bhattacharyya LH (2016) Perspective of beneficial microbes in agriculture under changing climatic scenario: a review. J Phytol 8:26–41. https://doi.org/10.19071/jp.2016.v8.3022
Bianucci E, Godoy A, Furlan A, Peralta JM, Hernández LE, Carpena-Ruiz RO et al (2018) Arsenic toxicity in soybean alleviated by a symbiotic species of Bradyrhizobium. Symbiosis 74:167–176
Borowik A, Wyszkowska J, Kucharski M, Kucharski J (2019) Implications of soil pollution with diesel oil and BP petroleum with ACTIVE technology for soil health. Int J Environ Res Public Health 16(14):2474. https://doi.org/10.3390/ijerph16142474
Brıgido C, Nascimento FX, Duan J, Glick BR, Oliveira S (2013) Expression of an exogenous 1-aminocyclopropane-1-carboxylatedeaminase gene in Mesorhizobium spp. reduces the negative effects of salt stress in chickpea. FEMS Microbiol Lett 349:46–53
Bruto M, Prigent-Combaret C, Muller D, Moënne-Loccoz Y (2014) Analysis of genes contributing to plant-beneficial functions in plant growth-promoting rhizobacteria and related Proteobacteria. Sci Rep 4:6261
Bulgarelli D, Schlaeppi K, Spaepen S, Ver Loren Van Themaat E, Schulze-Lefert P (2013) Structure and functions of the bacterial microbiota of plants. Annu Rev Plant Biol 64:807–838. https://doi.org/10.1146/annurev-arplant-050312-120106
Cardinale M, Ratering S, Suarez C, Zapata Montoya AM, Geissler-Plaum R, Schnell S (2015) Paradox of plant growth promotion potential of rhizobacteria and their actual promotion effect on growth of barley (Hordeum vulgare L.) under salt stress. Microbiol Res 181:22–32. https://doi.org/10.1016/j.micres.2015.08.002
Chaparro JM, Sheflin AM, Manter DK, Vivanco JM (2012) Manipulating the soil microbiome to increase soil health and plant fertility. Biol Fertil Soils 48:489–499. https://doi.org/10.1007/s00374-012-0691-4
Chauhan H, Bagyaraj DJ (2015) Inoculation with selected microbial consortia not only enhances growth and yield of French bean but also reduces fertilizer application under field condition. Sci Hort 197:441–446
Cluis C (2004) Junk-greedy greens: phytoremediation as a new option for soil decontamination. BioTeach J 2:61–67
Conforte VP, Echeverria M, Sanchez C, Ugalde RA, Menendez AB, Lepek VC (2010) Engineered ACC deaminase-expressing free-living cells of Mesorhizobium loti show increased nodulation efficiency and competitiveness on Lotus spp. J Gen Appl Microbiol 56:331–338
Cunningham SD, Berti WR (1993) Phytoremediation of contaminated soils: progress and promise. In: Symposium on bioremediation and bioprocessing—ACS meeting 205, Denver, Colorado, American Chemical Society, pp 265–268
Dąbrowska G, Hrynkiewicz K, Trejgell A, Baum C (2017) The effect of plant growth-promoting rhizobacteria on the phytoextraction of Cd and Zn by Brassica napus L. Int J Phytoremediation 19:597–604. https://doi.org/10.1080/15226514.2016.1244157
Dangi S, Tirado-Corbalá R, Gerik J, Hanson B (2017) Effect of long term continuous fumigation on soil microbial communities. Agronomy 7:37. https://doi.org/10.3390/agronomy7020037
Dary M, Chamber-Pérez MA, Palomares AJ, Pajuelo E (2010) “In situ” phytostabilisation of heavy metal polluted soils using Lupinus luteus inoculated with metal resistant plant-growth promoting rhizobacteria. J Hazard Mater 177:323–330
Das AJ, Kumar R (2016) Bioremediation of petroleum contaminated soil to combat toxicity on Withania somnifera through seed priming with biosurfactant producing plant growth promoting rhizobacteria. J Environ Manag 174:79–86. https://doi.org/10.1016/j.jenvman.2016.01.031
Delgadillos J, Lafuente A, Doukkali B, Redondo-Gómez S, Mateos-Naranjo E, Caviedes MA et al (2015) Improving legume nodulation and Cu rhizostabilization using a genetically modified rhizobia. Environ Technol 36(10):1237–1245
Devi U, Khatri I, Kumar N, Kumar L, Sharma D, Subramanian S et al (2013) Draft genome sequence of a plant growth-promoting rhizobacterium, Serratia fonticola strain AU-P3(3). Genome Announc 1(6):e00946–e00913. https://doi.org/10.1128/genomeA.00946-13
Dhankher OP, Pilon-Smits EAH, Meagher RB, Doty S (2011) Biotechnological approaches for phytoremediation. Academic, Oxford, pp 309–328
Díaz-Valle A, López-Calleja AC, Alvarez-Venegas R (2019) Enhancement of pathogen resistance in common bean plants by inoculation with Rhizobium etli. Front Plant Sci 10:1317
Eliana Andrea MM, Ana Carolina TE, Tito José CB, José Luis MN, Luis Carlos GM (2019) Evaluation of contaminants in agricultural soils in an irrigation district in Colombia. Heliyon 5(8):e02217. https://doi.org/10.1016/j.heliyon.2019.e02217
Fagorzi C, Checcucci A, diCenzo GC, Debiec-Andrzejewska K, Dziewit L, Pini F et al (2018) Harnessing rhizobia to improve heavy-metal phytoremediation by legumes. Genes (Basel) 9(11):542. https://doi.org/10.3390/genes9110542
Fukami J, Cerezini P, Hungria M (2018) Azospirillum: benefits that go far beyond biological nitrogen fixation. AMB Expr 8:73. https://doi.org/10.1186/s13568-018-0608-1
Ganeshan G, Kumar AM (2005) Pseudomonas fluorescens, a potential bacterial antagonist to control plant diseases. J Plant Interact 1(3):123–134. https://doi.org/10.1080/17429140600907043
Ghnaya T, Mnassri M, Ghabriche R, Wali M, Poschenrieder C, Lutts S et al (2015) Nodulation by Sinorhizobium meliloti originated from a mining soil alleviates Cd toxicity and increases Cd-phytoextraction in Medicago sativa L. Front Plant Sci 6:1–10
Glick BR (1995) The enhancement of plant growth by free-living bacteria. Can J Microbiol 41:109–117
Glick BR (2012) Plant growth-promoting bacteria: mechanisms and applications. Scientifica Article ID 963401, 15 pages. https://doi.org/10.6064/2012/963401
Glick BR (2014) Bacteria with ACC deaminase can promote plant growth and help to feed the world. Microbiol Res 169(1):30–39
Goswami D, Thakker JN, Dhandhukia PC (2016) Portraying mechanics of plant growth promoting rhizobacteria (PGPR): a review. Cogent Food Agric 2:1–19
Gupta S, Singh D (2017) Role of genetically modified microorganisms in heavy metal bioremediation. In: Kumar R, Sharma A, Ahluwalia S (eds) Advances in environmental biotechnology. Springer, Singapore, pp 197–214
Gurska J, Wang W, Gerhardt KE, Khalid AM, Isherwood DM, Huang XD et al (2009) Three year field test of a plant growth promoting rhizobacteria enhanced phytoremediation system at a land farm for treatment of hydrocarbon waste. Environ Sci Technol 43(12):4472–4479
Hao X, Xie P, Zhu YG, Taghavi S, Wei G, Rensing C (2015) Copper tolerance mechanisms of Mesorhizobium amorphae and its role in aiding phytostabilization by Robinia pseudoacacia in copper contaminated soil. Environ Sci Technol 49:2328–2340
Hassan TU, Bano A, Naz I (2017) Alleviation of heavy metals toxicity by the application of plant growth promoting rhizobacteria and effects on wheat grown in saline sodic field. Int J Phytoremediation 19:522–529. https://doi.org/10.1080/15226514.2016.1267696
Hawrot-Paw M, Nowak A (2012) An attempt at mathematical modelling of the process of microbiological biodegradation of diesel oil. Environ Prot Eng 38:23–29
He L, Ye J, Wu B, Huang L, Ren J, Wu X (2018) Effects of genetically modified Burkholderia pyrrocinia JK-SH007E1 on soil microbial community in poplar rhizosphere. For Pathol 48:e12430
Hou J, Liu W, Wang B, Wang Q, Luo Y, Franks AE (2015) PGPR enhanced phytoremediation of petroleum contaminated soil and rhizosphere microbial community response. Chemosphere 138:592–598
Huang XF, Chaparro JM, Reardon KF, Zhang R, Shen Q, Vivanco JM (2014) Rhizosphere interactions: root exudates, microbes, and microbial communities. Botany 92(4):267–275
Huang J, Liu Z, Li S, Xu B, Gong Y, Yang Y et al (2016) Isolation and engineering of plant growth promoting rhizobacteria Pseudomonas aeruginosa for enhanced cadmium bioremediation. J Gen Appl Microbiol 62:258–265
Ike A, Sriprang R, Ono H, Murooka Y, Yamashita M (2007) Bioremediation of cadmium contaminated soil using symbiosis between leguminous plant and recombinant rhizobia with the MTL4 and the PCS genes. Chemosphere 66:1670–1676
Ike A, Sriprang R, Ono H, Murooka Y, Yamashita M (2008) Promotion of metal accumulation in nodule of Astragalus sinicus by the expression of the iron-regulated transporter gene in Mesorhizobium huakuii subsp. rengei B3. J Biosci Bioeng 105:642–648
Jing YD, He ZL, Yang XE (2007) Role of soil rhizobacteria in phytoremediation of heavy metal contaminated soils. J Zhejiang Univ Sci B 8(3):192–207. https://doi.org/10.1631/jzus.2007.B0192
Kamran MA, Syed JH, Eqani SAMAS, Munis MFH, Chaudhary HJ (2015) Effect of plant growth-promoting rhizobacteria inoculation on cadmium (Cd) uptake by Eruca sativa. Environ Sci Pollut Res 22(12):9275–9283
Khan N, Bano A (2018) Role of PGPR in the phytoremediation of heavy metals and crop growth under municipal wastewater irrigation. In: Ansari AA, Gill SS, Gill R, Lanza GR, Newman L (eds) Phytoremediation. Springer, Cham, pp 135–149. https://doi.org/10.1007/978-3-319-99651-6_5
Khan MS, Zaidi A, Wani PA, Oves M (2009) Role of plant growth promoting rhizobacteria in the remediation of metal contaminated soils. Environ Chem Lett 7:1–19
Khanna K, Jamwal VL, Gandhi SG, Ohri P, Bhardwaj R (2019) Metal resistant PGPR lowered Cd uptake and expression of metal transporter genes with improved growth and photosynthetic pigments in Lycopersicon esculentum under metal toxicity. Sci Rep 9:5855
Kloepper JW, Schroth MN (1978) Plant growth-promoting rhizobacteria on radishes. In: Proceedings of the 4th international conference on plant pathogenic bacteria, vol 2, pp 879–882. INRA, Angers
Kloepper J, Leong J, Teintze M, Schroth MN (1980) Enhanced plant growth by siderophores produced by plant growth-promoting rhizobacteria. Nature 286:885–886
Koga K, Masuda S (2015) Bacteria that reduce the content of heavy metals in a plant. Patent no: EP2578675B1
Kong Z, Glick BR, Duan J, Ding S, Tian J, McConkey BJ et al (2015) Effects of 1-aminocyclopropane-1-carboxylate (ACC) deaminase-overproducing Sinorhizobium meliloti on plant growth and copper tolerance of Medicago lupulina. Plant Soil 70:5891–5897
Leach JE, Triplett LR, Argueso CT, Trivedi P (2017) Communication in the phytobiome. Cell 169:587–596. https://doi.org/10.1016/j.cell.2017.04.025
Li Z, Ma Z, Hao X, Wei G (2012) Draft genome sequence of Sinorhizo biummeliloti CCNWSX0020, a nitrogen-fixing symbiont with copper tolerance capability isolated from lead-zinc mine tailings. J Bacteriol 194(5):1267–1268
Li Z, Ma Z, Hao X, Rensing C, Wei G (2014) Genes conferring copper resistance in Sinorhizobium meliloti CCNWSX0020 also promote the growth of Medicago lupulina in copper-contaminated soil. Appl Environ Microbiol 80(6):1961–1971
Lim JH, Kim SD (2013) Induction of drought stress resistance by multi-functional PGPR Bacillus licheniformis K11 in pepper. Plant Pathol J 29(2):201–208. https://doi.org/10.5423/PPJ.SI.02.2013.0021
Liu W, Hou J, Wang Q, Ding L, Luo Y (2014) Isolation and characterization of plant growth-promoting rhizobacteria and their effects on phytoremediation of petroleum-contaminated saline-alkali soil. Chemosphere 117:303–308
Lundberg DS, Lebeis SL, Paredes SH, Yourstone S, Gehring J, Malfatti S et al (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488:86–90. https://doi.org/10.1038/nature11237
Ma Y, Prasad MN, Rajkumar M, Freitas H (2011a) Plant growth promoting rhizobacteria and endophytes accelerate phytoremediation of metalliferous soils. Biotechnol Adv 29(2):248–258
Ma Y, Rajkumar M, Luo Y, Freitas H (2011b) Inoculation of endophytic bacteria on host and non-host plants-effects on plant growth and Ni uptake. J Hazard Mater 195:230–237
Mahdi I, Fahsi N, Hafidi M, Allaoui A, Biskri L (2020) Plant growth enhancement using rhizospheric halotolerant phosphate solubilizing bacterium Bacillus licheniformis QA1 and Enterobacter asburiae QF11 isolated from Chenopodium quinoa Willd. Microorganisms 8(6):948. https://doi.org/10.3390/microorganisms8060948
Manoj SR, Karthik C, Kadirvelu K, Arulselvi PI, Shanmugasundaram T, Bruno B et al (2020) Understanding the molecular mechanisms for the enhanced phytoremediation of heavy metals through plant growth promoting rhizobacteria: a review. J Environ Manag 254:109779
Matthews KR, Cuchiara ML (2014) Gene patents, patenting life and the impact of court rulings on US stem cell patents and research. Regen Med 9:191–200. https://doi.org/10.2217/rme.13.93
Maynaud G, Brunel B, Yashiro E, Mergeay M, Cleyet-Marel JC, Quéré AL (2014) CadA of Mesorhizobium metallidurans isolated from a zinc-rich mining soil is a PIB-2-type ATPase involved in cadmium and zinc resistance. Res Microbiol 165:175–189
McCutcheon SC, Jorgensen SE (2008) Phytoremediation. In: Jorgensen SE, Fath B (eds) Encyclopedia of ecology. Elsevier Science BV, Amsterdam, pp 2751–2766
McNair MR, Tilstone GH, Smith SS (2000) The genetics of metal tolerance and accumulation in higher plants. In: Terry N, Bañuelos G (eds) Phytoremediation of contaminated soil and water. Lewis Publishers, Boca Raton, pp 235–250
Mendez MO, Maier RM (2008) Phytostabilization of mine tailings in arid and semiarid environments—an emerging remediation technology. Environ Health Perspect 116(3):278–283
Mendis CH, Thomas VP, Schwientek P, Salamzade R, Chien JT, Waidyarathne P et al (2018) Strain-specific quantification of root colonization by plant growth promoting rhizobacteria Bacillus firmus I-1582 and Bacillus amyloliquefaciens QST713 in non-sterile soil and field conditions. PLoS One 13(2):e0193119. https://doi.org/10.1371/journal.pone.0193119
Mishra J, Singh R, Arora NK (2017) Alleviation of heavy metal stress in plants and remediation of soil by rhizosphere microorganisms. Front Microbiol 8:1706
Mitra S, Pramanik K, Sarkar A, Ghosh PK, Soren T, Maiti TK (2018) Bioaccumulation of cadmium by Enterobacter sp. and enhancement of rice seedling growth under cadmium stress. Ecotoxicol Environ Saf 156:183–196
Muratova AY, Turkovskaya OV, Antonyuk LP, Makarov OE, Pozdnyakova LI, Ignatov VV (2005) Oil-oxidizing potential of associative rhizobacteria of the genus Azospirillum. Microbiology 74(2):210–215
Nakbanpote W, Meesungnoen O, Prasad MNV (2016) Potential of ornamental plants for phytoremediation of heavy metals and income generation. In: Prasad MNV (ed) Bioremediation and bioeconomy. Elsevier, Amsterdam, pp 179–217
Nakkeeran S, Fernando WGD, Siddiqui ZA (2005) Plant growth promoting rhizobacteria formulations and its scope in commercialization for the management of pests and diseases. In: Siddiqui ZA (ed) PGPR: biocontrol and biofertilization. Springer, Dordrecht, pp 257–296
Nascimento FX, Brıgido C, Glick BR, Oliveira S, Alho L (2012) Mesorhizobium ciceri LMS-1 expressing an exogenous 1-aminocyclopropane-1-carboxylate (ACC) deaminase increases its nodulation abilities and chickpea plant resistance to soil constraints. Lett Appl Microbiol 55:15–21
Nordberg G, Fowler BA, Nordberg M, Friberg L (eds) (2009) Handbook on the toxicology of metals, 3rd edn. Academic Press, London
Novo LAB, Castro PML, Alvarenga P, da Silva EF (2018) Plant growth–promoting rhizobacteria-assisted phytoremediation of mine soils. In: Bio-geotechnologies for mine site rehabilitation. Elsevier, Amsterdam, pp 281–295
Ortiz-Castro R, Contreras-Cornejo HA, Macias-Rodriguez L, Lopez-Bucio J (2009) The role of microbial signals in plant growth and development. Plant Signal Behav 4:701–712. https://doi.org/10.4161/psb.4.8.9047
Pajuelo E, Rodŕıguez-Llorente ID, Lafuente A, Caviedes MA (2011) Legume–rhizobium symbioses as a tool for bioremediation of heavy metal polluted soils. Bioman Metal Contam Soils 20:95–123
Pandey VC, Bajpai O (2019) Phytoremediation: from theory toward practice. In: Pandey VC, Bauddh K (eds) Phytomanagement of polluted sites. Elsevier, Amsterdam, pp 1–49
Paulsen IT, Press CM, Ravel J, Kobayashi DY, Myers GS, Mavrodi DV et al (2005) Complete genome sequence of the plant commensal Pseudomonas fluorescens Pf-5. Nat Biotechnol 23(7):873–878
Perez-Palacios P, Romero-Aguilar A, Martinez JD, Doukkali B, Caviedes MA, Rodriguez-Llorente ID et al (2017) Double genetically modified symbiotic system for improved Cu phytostabilization in legume roots. Environ Sci Pollut Res 24(17):1–14
Prapagdee B, Khonsue N (2015) Bacterial-assisted cadmium phytoremediation by Ocimum gratissimum L. in polluted agricultural soil: a field trial experiment. Int J Environ Sci Technol 12(12):3843–3852
Qin J, Rosen BP, Zhang Y, Wang G, Franke S, Rensing C (2006) Arsenic detoxification and evolution of trimethylarsine gas by a microbial arsenite S-adenosylmethionine methyltransferase. PNAS 103(7):2075–2080
Radwan SS, Dashti N, El-Nemr I, Khanafer M (2007) Hydrocarbon utilization by nodule bacteria and plant growth-promoting rhizobacteria. Int J Phytoremediation 9:475–486. https://doi.org/10.1080/15226510701709580
Roy AS, Yenn R, Singh AK, Boruah HPD, Saikia N, Deka M (2013) Bioremediation of crude oil contaminated tea plantation soil using two Pseudomonas aeruginosa strains AS 03 and NA 108. Afr J Biotechnol 12(19):2600–2610
Saadani O, Fatnassi IC, Chiboub M, Abdelkrim S, Barhoumi F, Jebara M et al (2016) In situ phytostabilisation capacity of three legumes and their associated plant growth promoting bacteria (PGPBs) in mine tailings of northern Tunisia. Ecotoxicol Environ Saf 130:263–269
Saleem M, Arshad M, Hussain S, Bhatti AS (2007) Perspective of plant growth promoting rhizobacteria (PGPR) containing ACC deaminase in stress agriculture. J Ind Microbiol Biotechnol 34(10):635–648. https://doi.org/10.1007/s10295-007-0240-6
Sarwar N, Imran M, Shaheen MR, Ishaq W, Kamran A, Matloob A et al (2017) Phytoremediation strategies for soils contaminated with heavy metals: modifications and future perspectives. Chemosphere 171:710–721
Saxena G, Purchase D, Mulla SI, Saratale GD, Bharagava RN (2019) Phytoremediation of heavy metal-contaminated sites: eco-environmental concerns, field studies, sustainability issues and future prospects. Rev Environ Contam Toxicol 249:71–131. https://doi.org/10.1007/398_2019_24
Schwitzguébel J (2017) Phytoremediation of soils contaminated by organic compounds: hype, hope and facts. J Soils Sediments 17:1492–1502
Smith DL, Praslickova D, Ilangumaran G (2015) Inter-organismal signaling and management of the phytomicrobiome. Front Plant Sci 6:722. https://doi.org/10.3389/fpls.2015.00722
Smith DL, Gravel V, Yergeau E (2017) Editorial: signaling in the phytomicrobiome. Front Plant Sci 8:611. https://doi.org/10.3389/fpls.2017.00611
Somers E, Vanderleyden J, Srinivasan M (2004) Rhizosphere bacterial signalling: a love parade beneath our feet. Crit Rev Microbiol 30(4):205–240
Song JY, Kim HA, Kim JS, Kim SY, Jeong H, Kang SG et al (2012) Genome sequence of the plant growth-promoting rhizobacterium Bacillus sp. Strain JS. J Bacteriol 194(14):3760–3761. https://doi.org/10.1128/JB.00676-12
Speight JG (2020) Remediation technologies. In: Speight JG (ed) Natural water remediation. Butterworth-Heinemann, Oxford, pp 263–304
Sriprang R, Hayashi M, Yamashita M, Ono H, Saeki K, Murooka Y (2002) A novel bioremediation system for heavy metals using the symbiosis between leguminous plant and genetically engineered rhizobia. J Biotechnol 99:279–293
Sriprang R, Hayashi M, Ono H, Takagi M, Hirata K, Murooka Y (2003) Enhanced accumulation of Cd2+ by a Mesorhizobium sp. transformed with a gene from Arabidopsis thaliana coding for phytochelatin synthase. Appl Environ Microbiol 69:1791–1796
Tabassum B, Khan A, Tariq RM, Ramzan M, Khan MSI, Khan I et al (2017) Bottlenecks in commercialisation and future prospects of PGPR. Appl Soil Ecol 121:102–117
Tak HI, Ahmad F, Babalola OO (2013) Advances in the application of plant growth-promoting rhizobacteria in phytoremediation of heavy metals. Rev Environ Contam Toxicol 223:33–52
Thijs S, Weyens N, Sillen W, Gkorezis P, Carleer R, Vangronsveld J (2014) Plant growth promotion by DNT-degrading bacteria. Microb Biotechnol 7:294–306
Tiwari S, Lata C (2018) Heavy metal stress, signaling, and tolerance due to plant-associated microbes: an overview. Front Plant Sci 9:452
Treesubsuntorn C, Dhurakit P, Khaksar G, Thiravetyan P (2018) Effect of microorganisms on reducing cadmium uptake and toxicity in rice (Oryza sativa L.). Environ Sci Pollut Res 25(26):25690–25701
Tripathi P, Singh PC, Mishra A, Srivastava S, Chauhan R, Awasthi S et al (2017) Arsenic tolerant Trichoderma sp. reduces arsenic induced stress in chickpea (Cicer arietinum). Environ Pollut 223:137–145
Ullah A, Heng S, Munis MFH, Fahad S, Yang X (2015) Phytoremediation of heavy metals assisted by plant growth promoting (PGP) bacteria: a review. Environ Exp Bot 117:28–40
Vejan P, Abdullah R, Khadiran T, Ismail S, Boyce AN (2016) Role of plant growth promoting rhizobacteria in agricultural sustainability—a review. Molecules 21(5):573. https://doi.org/10.3390/molecules21050573
Venturi V, Keel C (2016) Signaling in the rhizosphere. Trends Plant Sci 21(3):187–198
Vessey JK (2003) Plant growth promoting rhizobacteria as biofertilizers. Plant Soil 255:571–586
Wani SA, Chand S, Ali T (2013) Potential use of Azotobacter chroococcum in crop production—an overview. Curr Agric Res J 1:35–38
Welbaum GE, Sturz AV, Dong Z, Nowak J (2010) Managing soil microorganisms to improve productivity of agro-ecosystems. Crit Rev Plant Sci 23(2):175–193. https://doi.org/10.1080/07352680490433295
Weyens N, Schellingen K, Beckers B, Janssen J, Ceulemans R, van der Lelie D et al (2013) Potential of willow and its genetically engineered associated bacteria to remediate mixed Cd and toluene contamination. J Soils Sediments 13:176–188
Wu S, Jia S, Sun D, Chen M, Chen X, Zhong J et al (2005) Purification and characterization of two novel antimicrobial peptides Subpeptin JM4-A and Subpeptin JM4-B produced by Bacillus subtilis JM4. Curr Microbiol 51:292–296
Wu CH, Wood TK, Mulchandani A, Chen W (2006) Engineering plant-microbe symbiosis for rhizoremediation of heavy metals. Appl Environ Microbiol 72:1129–1134
Yong X, Chen Y, Liu W, Xu L, Zhou J, Wang S et al (2014) Enhanced cadmium resistance and accumulation in Pseudomonas putida KT2440 expressing the phytochelatin synthase gene of Schizosaccharomyces pombe. Lett Appl Microbiol 58:255–261
Zhang R, Vivanco JM, Shen Q (2017) The unseen rhizosphere root soil-microbe interactions for crop production. Curr Opin Microbiol 37:8–14. https://doi.org/10.1016/j.mib.2017.03.008
Zhao K, Penttinen P, Zhang X, Ao X, Liu M, Yu X et al (2014) Maize rhizosphere in Sichuan, China, hosts plant growth promoting Burkholderia cepacia with phosphate solubilizing and antifungal abilities. Microbiol Res 169(1):76–82
Zielazinski EL, González-Guerrero M, Subramanian P, Stemmler TL, Argüello JM, Rosenzweig AC (2013) Sinorhizobium meliloti Nia is a P1B-5-ATPase expressed in the nodule during plant symbiosis and is involved in Ni and Fe transport. Metallomics 5:1614–1623
Zolla G, Badri DV, Bakker MG, Manter DK, Vivanco JM (2013) Soil microbiomes vary in their ability to confer drought tolerance to Arabidopsis. Appl Soil Ecol 68:1–9. https://doi.org/10.1016/j.apsoil.2013.03.007
Zubair M, Shakir M, Ali Q, Rani N, Fatima N, Farooq S et al (2016) Rhizobacteria and phytoremediation of heavy metals. Environ Technol Rev 5(1):1
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Malik, G., Chugh, S., Hooda, S., Chaturvedi, R. (2022). Plant Growth Promoting Rhizobacteria (PGPR)-Assisted Phytoremediation of Contaminated Soils. In: Singh, A.K., Tripathi, V., Shukla, A.K., Kumar, P. (eds) Bacterial Endophytes for Sustainable Agriculture and Environmental Management. Springer, Singapore. https://doi.org/10.1007/978-981-16-4497-9_4
Download citation
DOI: https://doi.org/10.1007/978-981-16-4497-9_4
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-4496-2
Online ISBN: 978-981-16-4497-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)