Abstract
The gut microbiome is considered as an organ that contributes to the regulation of host metabolism. Mammals possess an ‘extended genome’ of millions of microbial genomes located in the intestine: the microbiome. To date, there is rapidly booming evidence for host–microbe interaction at virtually all levels of complexity, ranging from direct cell-to-cell communication to comprehensive systemic signalling and engaging various organs and organ systems, including the central nervous system. As such, the disclosure of differential microbial composition is associated with alterations in behaviour, and cognition has consequently subsidized to establish the microbiota–gut–brain axis as an extension of the well-accepted gut–brain axis concept. Numerous exertions have been focused on demarcating a role for this axis in health and disease, ranging from stress-associated conditions such as depression, anxiety and irritable bowel syndrome (IBS) to neurodevelopmental conditions such as autism. Besides this, the gut–brain axis is also reported to influence brain disorders, e.g. Alzheimer’s disease, Parkinson’s disease and schizophrenia. There is bidirectional communication network that links the enteric and central nervous systems. This network is not merely anatomical, but it encompasses endocrine, humoral, metabolic and immune routes of intercommunication as well. The autonomic nervous system, hypothalamic–pituitary–adrenal (HPA) axis and nerves within the gastrointestinal tract all link the gut and the brain, allowing the brain to influence intestinal activities, including activity of functional immune effector cells, and the gut to influence mood and behaviour, cognition and mental and reproductive health. In this chapter, we have focused on how gut microbiomes influence physical and mental health.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction: Gut Microbiome and Brain Broadcast
1.1 Composition and Dynamics of Healthy Adult Microbiota
Earlier it was depicted that the gut microbiota is comprised of 500–1000 species of microbes (Ramakrishna and Krishnan 2007), but a large-scale study in 2007 has estimated that the collective human gut microflora is made up of more than 35,000 bacterial species (Frank et al. 2007). Additionally, if well specified from a standpoint of total bacterial genes, Human Microbiome Project and Metagenomics of the Human Intestinal Tract studies reveal that there is a presence of more than 10 million non-redundant genes in the human microbiome.
Considering the human body as an environment, human microbiota is the entire assemblage of microorganisms living at the surface and inside of our body (Dewhirst et al. 2010; Grice et al. 2006; González et al. 2014; Arumugam et al. 2011). These communities of microorganisms are vital for many more important aspects of human physiology, digestion, detoxification and immune system development. Some of the microbes that live in the gut encode proteins that are essential for the host’s health, such as enzymes that are required for the breakdown of indigestible food components and vitamin production (Flint et al. 2012; Qin et al. 2010). So we humans are having two genomes, one inherited from our parents and the other one is acquired, i.e. ‘the microbiome’. This concept is the foundation for the characterization of humans as ‘superorganisms’ (Walsh et al. 2014). The most significant difference between these two genomes is that the inherited genome remains nearly stable during our entire lifetime, but the genome acquired from microbiome is extremely dynamic and can be affected by numerous factors like age (Gajer et al. 2012), diet (David et al. 2014; Wu et al. 2011), hormonal cycles (Koren et al. 2012), travel (Yatsunenko et al. 2012), therapies, treatments (Perez-Cobas et al. 2013) and illness (Perez-Cobas et al. 2013).
1.2 Formation of Gut Microbiota During the Early Stages of Life
Infants who are fully term, vaginally delivered, breastfed, and not antibiotic-treated have the best chance of developing a healthy gut flora (Alex et al. 2013). In these newborns, facultative anaerobes like enterobacteria, staphylococci and streptococci are the most primitive microbes starting to colonize and further taking advantage of the redox potential and available oxygen in the newborn gut. These initial colonizers consume available oxygen in the gut; by this way, it creates an anaerobic ecosystem and permits the proliferation of the strict anaerobes, Clostridium, Bacteroides and bifidobacteria; after that, bifidobacteria become dominant and more numerous than all other bacterial groups and species within the first few weeks of human life. The newborn microbiota is extremely dynamic, and it is exemplified by low stability and low variety. By the end of first year of life, newborns develop a microbial profile different for each infant and attains the characteristic microbiota of an adult gut microbiome, and by age of 2.5 years, the microbiota completely resembles that of an adult in terms of composition (Lobo et al. 2014).
1.3 What Consists Gut Microbiota?
The adult microbiota has been reported to be relatively stable over time in addition to being more complex than that of the neonate (Hamady and Knight 2009). Healthy gut microbiota is mainly composed of phyla Firmicutes and Bacteroidetes. followed by phyla Actinobacteria and Verrucomicrobia. Yet this general profile remains persistent; gut microbiota displays both temporal and spatial differences in distribution at the genus level and beyond. There is a notable variation in the range and quantity of bacteria from the oesophagus distally to the rectum, ranging from 101 per gram of contents in the oesophagus and stomach to about 1012 per gram of insides in the colon and distal gut (O’Hara and Shanahan 2006). Figure 8.1 shows the time-based diversity of the gut microbiota from oesophagus distally to the colon. Streptococcus seems to be the leading genus in the distal oesophagus, duodenum and also jejunum (Pei et al. 2004; Justesen et al. 1984).
Helicobacter is the regulatory genus present in the stomach and regulates the entire microbial population of the gastric flora; that is, when Helicobacter pylori (H. pylori) populates in the stomach as a commensal, at that time, the gut attains a rich diversity with another dominant genus like Streptococcus (most dominant), Prevotella, Veillonella and Rothia (Blaser 1999; Andersson et al. 2008). This range of microbes gets disturbed when H. pylori acquires a pathogenic phenotype. The large intestine comprises more than 70% of all microbes that reside in our body. The main phyla that inhabit in the large intestine are Firmicutes and Bacteroidetes. Eventually, Firmicutes/Bacteroidetes ratio has been obtained in predisposition to disease states (Ley et al. 2006).
The remarkable variability even in healthy persons that has been noticed in the current studies makes the implication of this ratio controversial. Additionally, from Firmicutes and Bacteroidetes, the human colon is similarly having primary pathogens like Campylobacter jejuni, Salmonella enterica, Vibrio cholera, Escherichia coli (E. coli) and Bacteroides fragilis, but with very less abundance (0.1% or less of the entire gut microbiome) (Human Microbiome Project Consortium 2012; Gillespie et al. 2011). The phylum Proteobacteria is markedly low, and its deficiency along with high abundance of genera Bacteroides, Prevotella and Ruminococcus suggests a healthy gut microbiota (Hollister et al. 2014). Moreover, this longitudinal divergence, we do have axial discrepancy from the lumen to the mucosal surface of the intestine. Although Enterobacteriaceae, Enterococcus, Clostridium, Lactobacillus, Bacteroides, Bifidobacterium, Streptococcus and Ruminococcus are the predominant luminal microbial genera (can be recognized from stool analysis), solitary Clostridium, Lactobacillus, Enterococcus and Akkermansia are the principal mucosa and mucus-linked genera (which can be detected in the mucus layer and epithelial crypts of the small intestine) (Swidsinski et al. 2005). These intestinal microbiotas are known to play a key role in several metabolic, nutritional, physiological and immunological processes (O’Hara and Shanahan 2006).
Throughout human life, the healthy gut microbiota composition increases in both variety and richness (Scholtens et al. 2012) and gets maximum complexity in the human adult, with several hundred species-level phylotypes dominated by the phyla Bacteroidetes and Firmicutes (Rajilic-Stojanovic et al. 2012). Each human individual reaches a homeostatic climax composition, which likely remains relatively stable during most of a healthy adult’s life. Although the individual microbial composition has an ‘individual core’ that varies at the bacterial phylotype level and depends on the lifestyle of that individual (Zoetendal et al. 2008; Jalanka-Tuovinen et al. 2011), at the late stages of life, the microbiota composition becomes again less diverse and more dynamic, characterized by a higher Bacteroides to Firmicutes ratio, increase in Proteobacteria and decrease in Bifidobacterium (Biagi et al. 2010).
Establishment of the gut microbiota population in early life plays a key role in the microbial makeup and disease predisposition throughout the entire life span (Scholtens et al. 2012). Sometimes, a dissimilar microbiota composition is linked with chronic intestinal disorders and the severity of distress during disease and subsequent use of antibiotic (Sekirov et al. 2010). An additional important factor in microbiota composition improvement is diet. In early life, diet already has an effect on the gut microbiome. Breastfed babies has a microbiota that is more heterogeneous than that of formula-fed babies and has a better taxonomic variety (Schwiertz et al. 2010). In addition, food habits till the age of 3 years can also impact gut microbiota composition; in a malnourished child, there is lower abundance of Bacteroidetes; those are proven to be specific in breaking down of carbohydrates from energy-rich western diet foods. Briefly, human gut microbiota is having a symbiotic relationship with the gut mucosa and reveals significant nutrient metabolic, xenobiotic and drug metabolism, antimicrobial protection and immunomodulation and gut protecting jobs in the healthy person. And it obtains its beneficial nutrients from host dietary components and shed epithelial cells. As a result, it is an organ with wide metabolic competence and significant signals from the brain that can affect the motor, sensory, and secretory systems of the gut and functional smoothness.
1.4 Gut–Brain Axis
An estimated 90 percent of cells found in the human body are not belongs to the human after all but of mostly prokaryotic origin, derived from at least 40,000 bacterial strains in 1800 genera (Forsythe and Kunze 2013; Frank and Pace 2008; Luckey 1972). Though considerably smaller in size, these approximately 100 trillion cells add up to a mass of almost 1–2 kg in an adult individual (Forsythe and Kunze 2013)—approximately the weight of a full-grown human brain (1.5 kg).
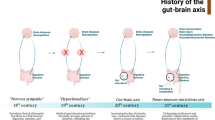
The discovery that differential microbial composition is associated with alterations in behaviour and cognition has significantly contributed to establish the ‘microbiota–gut–brain axis’ as an extension of the well-accepted ‘gut–brain axis’ concept. This concept is used to describe the bidirectional communication between the central nervous system (CNS) and intestinal organs and was first introduced in terms of ‘peripheral regulation of emotions’ by William James and Carl Lange in the 1880s and further challenged and refined by Walter Cannon in the 1920s as ‘primacy of the brain in regulating gastrointestinal function’. So gut–brain axis is a bidirectional interaction network that links both enteric and central nervous systems. This correlation is not only anatomical but also extends to incorporate endocrine, humoral, metabolic and immune paths of transmission as well. Furthermore, autonomic nervous system, hypothalamic–pituitary–adrenal (HPA) axis and nerves within the gastrointestinal (GI) tract all link the gut and the brain together, allowing the brain to influence intestinal activities and activities of functional immune effector cells; moreover, they influence mood, cognition and mental health. This host–microbe interaction is present at all levels of complexity, ranging from direct cell-to-cell communication to extensive systemic signalling and involving various organs and organ systems.
Signals coming from the brain can affect motor, sensory and secretory sensory systems of the gut, and on the other hand, visceral messages from the gut can impact brain functioning with the help of this gut–brain bidirectional transmission network (Grenham et al. 2011; Montiel-Castro et al. 2013). This correlation of brain functioning with enteric gut microbiota is less extensively studied but increasingly accepted and appreciated (Khanna and Tosh 2014). Gut microbiota predominantly consists of bacteria but also contains archaea, protozoa, fungi and viruses, all of which have co-evolved with the human host. Our colon harbours the largest numbers of microorganisms in the gut; most of these native microbes are strict anaerobes in nature (Eckburg et al. 2005). Synthesis and role of these intestinal microbiota have constantly been the subject of intense study; primarily it was analysed using culture-based microbiological methods (Grenham et al. 2011), and right now, culture-independent 16S rRNA gene sequence-based techniques are in use, and these techniques allow better understanding of microbial structure and assortment of this complex study (Arboleya et al. 2008; Qin et al. 2010). With evolving improvements in metagenomic technologies, we are able to disclose the composition of the human gut microbiota from early childhood (Palmer et al. 2007) to elderly (Claesson et al. 2012). Although lesser is known regarding the physiological impact of these microbiota on host health, comprising that of the brain, understanding the stimulus of gut microbiota on the host well-being has been portrayed as one of the most exciting areas in entire medicine (Shanahan 2012).
At the time of birth, our brain is extremely under-developed, and gut is generally interpreted as completely sterile. As described in Sect. 8.1.2, preliminary colonization is influenced by mother’s microbe environment and the environment of the hospital. This colonization plays an important role in brain development in the early post-natal period. The subsequent microbial arrangement of the newborn gut is affected by several factors including diet, use of antibiotics, mode of delivery, surrounding environment and the main maternal microbiota (Koenig et al. 2011; Marques et al. 2010; Dominguez-Bello et al. 2010).
These properties of intestinal microbiota identified in healthy full-term infants are distressed in preterm infants (Dennison 1976) that are commonly delivered via caesarean section, take antibiotics and are sometimes not fed properly (Hoy et al. 2000). Moreover, preterm infants are having functionally immature or not properly developed gut which has low levels of acidity in the stomach, because they are lacking in gastric acid secretion and they need to be fed more frequently (Hoy et al. 2000; Sondheimer and Clark 1985; Sondheimer et al. 1985), and it leads to an increase in the incidence of potentially pathogenic bacteria in the gastrointestinal (GI) tract, and preterm infants have a smaller amount of microbial variety than full-term infants (Arboleya et al. 2008; Chang et al. 2011; Jacquot et al. 2011). And these characteristics, which have been linked to the development of cerebral palsy and autism, have been the focus of research and ongoing controversy (Mangiola et al. 2016).
In the case of the elderly, when these microbiota compositions of elderly people in nursing homes are compared with those living in the community with their families, large-scale alterations were noticed. Those admitted in nursing homes have a far less varied microbiota, and this can be a result of less diverse diet (Claesson et al. 2012). It is also thinkable, sometimes, that pathological factors lead to admission into nursing homes, likewise worsening cognitive functionality and declining physical activity, might be having an important role in the reduced microbial richness and not a less diverse diet. Current studies should explain this issue, and this can be a challenge for the food industry to discover diets for the elderly to help them sustain their microbial variety. What we can justify here is that a dysregulated gut microbiota either in early childhood or in an elderly population meaningfully increases the possibility of brain dysfunction.
1.5 How Gut Microbiota Communicates with the Brain?
There are various possible direct and indirect communication routes through which the gut microbiota can communicate with the brain including neuroendocrine, neuroanatomical immune and through neurotransmitters.
1.5.1 Neuroanatomical Pathway
Human gut can interrelate with the brain with the help of two neuroanatomical pathways. One is mutual information interchange straight between the gut and the brain by autonomic nervous system (ANS) and vagus nerve (VN) in the spinal cord, and the other one is a bidirectional signalling between the gut and the brain through communication between enteric nervous system (ENS) within the gut and ANS and VN; inside the spinal cord, information from the heart, lungs, liver, pancreas, stomach and intestines is conveyed to the brain via sensory fibres in the vagus nerve (Travagli et al. 2003). Sensory vagal inputs reach the nucleus of the solitary tract (NTS) and are thence conveyed to extensive zones of the CNS and also the cerebral cortex and medulla oblongata. Preclinical studies have implicated the vagus nerve as a key route of neural communication between microbes of the gut and centrally mediated behavioural effects, as confirmed with the elimination of central Lactobacillus rhamnosus after vagotomy (Bravo et al. 2011), and those who underwent vagotomy at an early age have a reduced risk of certain neurologic disorders (Svensson et al. 2015)
1.5.2 Neuroendocrine-HPA Axis
Neuroendocrine-HPA axis provides the principal control of the stress reaction and can have a considerable impact on the brain–gut–microbiota axis (Wang and Kasper 2014; Tillisch 2014; Scott et al. 2013; Moloney et al. 2014; O’Mahony et al. 2009, 2011, 2017). It is fair enough and maybe of significance in several pathologic conditions psychological or physical stress can considerably dysregulate the HPA axis and in result the brain–gut microbiota axis, e.g. in IBS (Dinan et al. 2006). Human brain recruits these same methods to control the composition of the gut microbiota, for example, in conditions of stress. The hypothalamic–pituitary–adrenal (HPA) axis controls cortisol secretion, and cortisol can in turn impact immune cells (including cytokine secretion) locally within gut as well as systemically in body. This cortisol level can also alter gut permeability and barrier function and can in turn alter gut microbiota composition. Additionally, the gut microbiota and probiotic agents can modify the levels of circulating cytokines, and this can be effective on brain functioning.
Stress and HPA axis can also affect the formation of the gut microbiome. Initial stress and separation of the mother may possibly lead to a long-term change of HPA and had an extended effect on the microbial population (Desbonnet et al. 2008; Barouei et al. 2012). When it is evaluated with rats not separated from the mother, an assortment of 16S ribosomal RNA in adult rats, who have been through mother separation for around 3 h/day starting from day 2 to day 12 after birth, unveiled that stress extremely altered microbiome detected from faeces (O’Mahony et al. 2009). A mouse that is exposed to a long-term stress microbiome configuration was comparably different from a non-stressed mouse (Bendtsen et al. 2012). Recently, with the use of the above theories, it can be concluded that repeated social interaction and stress can diminish the number of Bacteroides in the caecum and augment the number of Clostridium. Stress can also upsurge interleukin-6 (IL 6) and monocyte chemoattractant protein 1 (MCP-1) levels in blood. MCP-1 was significantly related to the variations of three different kinds of stress-inducing bacterial strains, namely, Enterococcus faecalis, Pseudobutyrivibrio and aerogenic bacteria Dorea.
1.5.3 Immunological Pathway
The development of gut immune system is dependent on the gut microbiota (Furusawa et al. 2013; Mayer et al. 2014). Germ-free mice nearly had no immune activity, but they were able to generate immunity when fed with certain microbiota. For instance, the segmented filamentous bacterium in the gut can re-establish its full functions of gut B and T lymphocytes (Umesaki et al. 1995, 1999; Talham et al. 1999). These bacteria can communicate with the host through a variety of routes, and Toll-like receptors (TLRs) of a host cell play an important role in the broadcast between bacteria and host. Currently, ten different types of TLRs are in the human innate immune system; all of these have been identified as pattern recognition receptors (Takeuchi and Akira 2010). And they are part of the innate immune system, performs the initial step in the production of cytokine response, also widely distributed on neurons (McKernan et al. 2011). Thus, neurons likewise respond to bacterial and viral components. Thus, neurons likewise respond to bacterial and viral components. Intestinal epithelial cells are able to transfer microbial composition or metabolites in the internal environment and also with the nervous system (O’Brien et al. 2004). The equilibrium of gut microbiota may alter the regulation of inflammatory response, and this method may also engage in the control of emotion and behaviour.
Immune signalling from the gut to the brain facilitated by cytokine molecules is an additional documented route of communication (El Aidy et al. 2014). Cytokines produced at the level of the gut can penetrate bloodstream to the brain. Under normal physiologic conditions, it is unlikely that they cross the blood–brain barrier (BBB), but growing evidence implies a capacity to signal across the BBB and to affect brain areas like hypothalamus, where the BBB is lacking. It is through the latter mechanism the cytokines interleukin (IL)-1 and IL-6 activate the hypothalamic–pituitary–adrenal (HPA) axis, bringing about the release of cortisol. This is the most potent activator of the stress system.
1.5.4 Neurotransmitters Regulating Gut–Brain Axis
Gut microbiota likewise regulates important central neurotransmitters, such as serotonin, with varying levels of precursors; for example, Bifidobacterium infantis has shown to raise plasma tryptophan levels, and so it influences central serotonin (5HT) transmission (O’Brien et al. 2004). Interestingly, some bacteria associated in the synthesis and release of neurotransmitters have been already reported. Lactobacillus and Bifidobacterium spp. can synthesize g-aminobutyric acid (GABA); Escherichia, Bacillus and Saccharomyces spp. are able to produce noradrenaline; Candida, Streptococcus, Escherichia and Enterococcus spp. have been synthesizing serotonin; Bacillus can produce dopamine; likewise Lactobacillus can generate acetylcholine (Lyte 2013, 2014). These neurotransmitters of microbial origin are able to penetrate into the mucosal layer of the intestine, even though it is extremely improbable that these bacterial species can directly affect brain function. Even if they enter into the bloodstream, which is by no means sure, they will be capable of crossing the blood–brain barrier (BBB). That is why their effect on brain function is almost indirect, by acting on the enteric nervous system (ENS). SCFAs (short-chain fatty acids), which include butyrate, propionate and acetate, are indispensable metabolic end products of gut microbial activity and may apply central effects through G-protein–coupled receptors, even though such receptors are sparsely concentrated in the brain. It is more obvious that they act as epigenetic modulators through histone deacetylases (Stilling et al. 2014) SCFAs are also engaged in energy balance and metabolism and able to regulate adipose tissue, liver tissue and skeletal muscle and function (Canfora et al. 2015). Therefore, a lot of essential neurotransmitters in the body are formed by the gut microbiota, employing impact on the human body including the brain. Therefore, a lot of essential neurotransmitters in the body are formed by the gut microbiota, employing impact on the human body including the brain from which several neurotransmitters produced by gut microbiota are defined as critical molecules.
2 Gut–Microbiome–Brain Implications on Physical Health
Generally, the intestinal microbiota composition of healthy individuals is comparatively stable; however, alterations in the microbiota community may lead to a permanent imbalance known as dysbiosis (Lynch and Pedersen 2016). Numerous factors such as antibiotics, diet (comprising specific probiotic and prebiotic intake), the host immune system and acidic environment have been seen to influence the microbiota composition of the gut. Perturbation to the gut microbiota ecosystem resulting in dysbiosis can lead to gastrointestinal diseases. With current research advising dysbiosis of gut microbiota is having potential implication not only in IBS, but also in other disorders such as obesity (Turnbaugh and Gordon 2009), diabetes (Qin et al. 2012), metabolic syndrome (D’Aversa et al. 2013), cardiovascular disease and IBD as well as on reproductive health.
2.1 Irritable Bowel Syndrome (IBS)
Irritable bowel syndrome (IBS) is a common gastrointestinal (GI) disorder categorized by persistent abdominal pain allied with alterations in bowel habits. Aspects associated to IBS symptom development comprise history of enteric infection, deviations in the gut microbiota, immunomodulation, alterations in brain–gut processing and vagaries in visceral sensation and motility (Ford et al. 2017). IBS can be clinically subtyped into IBS with constipation (IBS-C), IBS with diarrhoea (IBS-D) and mixed IBS (IBS-M). In addition, IBS patients seemed to have a higher degree of psychosocial stress, a poorer quality of life and inferior levels of work productivity. Alterations in the normal gut microbiota have been proposed as etiologic factors in the development of functional gastrointestinal disorders such as IBS and functional dyspepsia and shared GI disorders of unknown aetiology (Upadhyay et al. 2018). The pathogenesis and pathophysiology of IBS are incompletely unstated, but abnormal GI motility, visceral hypersensitivity, altered brain–gut function, low-grade inflammation and psychosocial factors are considered to subsidize. IBS has been significantly associated with small intestinal bacterial overgrowth (SIBO) (4–78%) (Ghoshal and Ghoshal 2017) and prior GI infection (5–32%), suggesting that enteric dysbiosis (i.e. disrupted microbial homeostasis) is a potential pathogenic mechanism of IBS. In recent years, many research groups have engrossed on recognizing the gut microbiota composition of the large intestine of IBS patients, using modern culture-independent techniques. Next-generation sequencing has revealed that IBS patients, compared with healthy controls, show significantly lower abundance in enteric Lactobacillus, Bifidobacterium and Faecalibacterium prausnitzii (O’Mahony et al. 2005). During periods of dysbiosis, the gut microbiome influences inflammation metabolism inside the GI tract, primarily through the production of cytokines (such as interleukin [IL]-10 and IL-4) and other cellular communication mediators, such as interferon-gamma. In irritable bowel syndrome (IBS), atypical microbiota populations stimulate mucosal innate immune responses, which increases gut epithelial permeability, triggers gut pain sensory pathways and dysregulates the enteric nervous system (Mayer et al. 2014); both brain–gut and gut–brain dysfunctions arise, the prior being dominant. Obstruction in the gut–brain axis affects intestinal motility and secretion, confers to visceral hypersensitivity and leads to cellular alterations of the entero-endocrine and immune systems (Kennedy et al. 2014).
2.2 Metabolic Diseases
The human gut microbiota has been studied for more than a century. Examination that the gut microbiota, as an environmental factor, donates to adiposity and has further increased curiosity in the field. The human microbiota can be altered by diet, and macronutrients work as substrates for various microbial metabolites, such as short-chain fatty acids (SCFA) and bile acids, and are able to modulate host metabolism. Obesity predisposes towards type 2 diabetes and cardiovascular disease. The gut microbiota shows a significant role in the regulation of the host’s metabolism and the extraction of energy from ingested food. Gut microbiotas have not only beneficial roles for the host but also have pathophysiological relations with the host, particularly in the case of obesity and related metabolic disorders. Recent studies have revealed that changes in the gut microbiota may be associated in the pathogenesis of obesity and diabetes. Obesity is the outcome of a long-term positive imbalance between energy intake and expenditure, which is controlled by multiple pathways comprising metabolites, hormones and neuropeptides (Upadhyay et al. 2018) Gut hormones seem to interconnect information from the gastrointestinal tract to the regulatory appetite centres within the central nervous system (CNS) via the so-called gut–brain axis. Such messages may be transferred to the CNS either via vagal or non-vagal afferent nerve signalling or directly via blood circulation (Bueter et al. 2009). Complex neural networks, distributed throughout the forebrain and brainstem, are in control of feeding and energy homeostasis (Schwartz et al. 2000). Novel research shows that the gut microbiota is involved in obesity and metabolic disorders, revealing that obese animal and human subjects have alterations in the composition of the gut microbiota compared to their lean counterparts. Moreover, transplantation of the microbiota of either obese or lean mice influences body weight in the germ-free recipient mice, suggesting that the gut ecosystem is a significant target for weight management (Harakeh et al. 2016). Native gut microbes may regulate body weight by inducing the host’s metabolic, neuroendocrine and immune functions. The intestinal microbiota, as a whole, offers supplementary metabolic functions and regulates the host’s gene expression, improving the ability to extract and store energy from the diet and contributing to body-weight gain (Ley et al. 2005). Inequalities in the gut microbiota and increasing plasma lipopolysaccharide can also act as inflammatory factors linked to the growth of atherosclerosis, insulin resistance and weight gain.
Onset of diabetes has increased rapidly and became a major public health concern worldwide. Type 1 diabetes (T1D) is an autoimmune disease characterized by insufficient insulin production because of T-cell-mediated destruction of insulin-secreting pancreatic beta cells, while type 2 diabetes (T2D) is a condition in which the body does not produce or use insulin well. Various factors are associated with the development of diabetes, such as diet, genome and intestinal microbiota. Changes in the gut microbiota can influence the levels of gut hormones involved in the regulation of satiety and glycaemic control, such as glucagon-like peptide-1 (GLP-1), which stimulates insulin secretion from the pancreas (Baggio and Drucker 2007; Tolhurst et al. 2012)
In obese individuals and patients with metabolic syndrome, an increase in insulin sensitivity is noted after 6 weeks of allogeneic or autologous faecal microbiota transplantation from normal individuals (Vrieze et al. 2012). Same results were observed earlier in mice as well (Bäckhed et al. 2004), which has become the trigger point for the researchers to study gut microbiota in diabetes (Gravitz 2012).
Carbohydrates are an essential nutritional factor for all mammals and their gut microbiota. These bacteria greatly influence glycaemic control. Undigested polysaccharides and partially digested carbohydrates reach the gut microbiota in the distal gut, where they are metabolized by bacterial enzymes (Musso et al. 2011). It has been investigated that the genera Ruminococcus, Fusobacterium and Blautia are positively associated with T2D, whereas the genera Bifidobacterium, Bacteroides, Faecalibacterium, Akkermansia and Roseburia are negatively associated with T2D (Mangiola et al. 2016).
The disrupted GDM (gestational diabetes gut microbiota) is very similar to gut microbiota in individual patients with type 2 diabetes and associated intermediary metabolic dysfunctions. Eight months postpartum, previous GDM women have different gut microbiota than the woman with normal pregnancy. This microbial dysbiosis may increase the risk of T2D, which needs to be investigated. Feeding of probiotic dahi containing Lactobacillus acidophilus NCDC14 and L. casei NCDC19 has been tested to substantially reduce STZ-induced oxidative damage in pancreatic tissues. Thus, the modulation of the intestinal microbiota by probiotics may be effective towards prevention and management of T1D and T2D. Supplements of prebiotics improve Bifidobacterium abundance, which alters microbial dysbiosis and improves glucose tolerance in mice (Cani et al. 2007).
There is much reasonable curiosity in the interplay of drugs and intestinal microbiota. It is well known that anti-diabetic drugs can modulate microbiota and improve diabetes. Improvements in fasting blood glucose, glucose tolerance and insulin resistance were observed with the combined therapy of a prebiotic and metformin in diabetic mice (Zheng et al. 2018). Multivariate research found that there are significant discrepancies in gut composition between T2DM (Type 2 Diabetes Mellitus) and in metformin-untreated participants significant increases was observed in Escherichia species and decreases in Intestinibacter following metformin therapy (Harsch and Konturek 2018). But there is still some uncertainty in this emerging field. Whether microbiota causes diabetes or diabetes affects intestinal microbiota is not yet quite simple. Investigation of altered gut microbiota can help in the early detection of diabetes even before serological tests (Nair et al. 2018).
2.3 Reproductive Health
As of today, researchers understand that residents in human gut form a symbiotic relationship with the host and offer several benefits to the host. For example, commensal microbes consistently provide a set of services to the host such as modulation of the immune system, inhibition of pathogen colonization and releasing nutrients from food (Kim et al. 2020). Reportedly, dysbiosis of gut microbiota has been implicated in many disease states, including diabetes, obesity and cardiovascular disease (Razavi et al. 2019). Recently, a novel theory of ‘microgenderome’ associated to the potential bidirectional interaction roles between the sex hormones and gut microbiota has emerged (Aguilera et al. 2020). It has been reported that the composition of commensal microbes of male and female animals deviated at the time of puberty, which imbedded that sex hormone levels put forth particular influences on the composition of the microbiota. Abstraction of gut microbiota increased the testosterone concentration in female mice but decreased the concentration in male mice. Thus, the commensal gut microbiota also had effects on the production of male sex hormone (Yuan et al. 2020).
Despite the advances in assisted reproductive technology (ART) in women as well as in men, approximately 8–12% of the global population willing to conceive is unable to do so. Available evidence advises that vaginal and uterine microbiota have a close relationship with female infertility (Moreno et al. 2016). In fact, microbiota analysis using the 16S rRNA amplicon sequencing of cervical swabs revealed significant differences regarding the relative read count of the genus Gardnerella between females diagnosed with infectious infertility and fertile controls (Benner et al. 2018). Several mechanisms have been proposed to suggest that dysbiosis of gut microbiota can be involved in the development of polycystic ovary syndrome (PCOS). However, the data obtained from cross-sectional studies are insufficient to reveal the causality of the relationship (Zhao et al. 2020; Yurtdaş and Akdevelioğlu 2020).
3 Gut–Microbiome–Brain Implications on Mental Health
The recently emerged concept of the bidirectional communication of the gut–microbiota–brain emphasizes the relevance to study associations between neurodegenerative diseases and the gut microbiota. There exists growing evidence that gut microbiota may affect the central nervous system through communication via the vagus nerve, signalling mediators of the immune system, enteric hormones and gut microbiota-derived products (Sherwin et al. 2016). Gut bacteria produce neuroactive compounds and can modulate neuronal function, plasticity and behaviour. Furthermore, intestinal microorganisms impact the host’s metabolism and immune status which in turn affect neuronal pathways in the enteric and central nervous systems. Communication pathways between gut microbiota and the central nervous system could include autonomic, neuroendocrine, enteric and immune systems, with pathology resulting in disruption to neurotransmitter balance, increases in chronic inflammation or exacerbated hypothalamic–pituitary–adrenal axis activity.
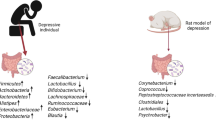
3.1 Stress/Depression
Depression is a major form of mood disorder characterized by depressed mood and/or recurrent thoughts of death and/or loss of interest or pleasure in life activities present over a period of at least 2 weeks. It results from neuro-psychiatric disturbance, immunological deregulation, genetic factors and environmental influences; nevertheless, a correlation with gut microbiota is emerging (Mangiola et al. 2016). Growing evidence links gut microbiome to the development and maturation of the central nervous system, which are regulated by microbiota potentially through stress response, neurotransmitter, neuroimmune, and endocrine pathways. The dysfunction of such microbiota–gut–brain axis is implicated in neuropsychiatric disorders, depression and other stress-related conditions (Kuo and Chung 2019). Bipolar disorder and major depression are associated with substantial disability, morbidity and reduced life expectancy. People with mood maladies have shown higher ratios of unhealthy lifestyle choices, including poor diet quality and suboptimal nutrition (Balanza-Martinez et al. 2020). Coello et al. (2019) found that gut microbiota community association differed between patients with newly diagnosed bipolar disorder and healthy individuals. Having a newly diagnosed bipolar illness was related with the prevalence of Flavonifractor, even after controlling for age, gender, physical activity, and waist size, and was mitigated by smoking status. The presence of Flavonifractor may possibly influence oxidative stress and inflammation in its host and could possibly link gut microbiota with illness pathology of bipolar disorder (Coello et al. 2019).
Sudo et al. (2004) demonstrated that the presence of gut microbiota modulated the long-range hypothalamic–pituitary–adrenal reaction to stress. These experiments showed that germ-free mice (mice raised in a sterile condition and lacking gut bacteria) exhibited a higher stress response as measured by an increased adrenocorticotropic hormone and corticosterone release compared to control mice with gut microbiota. This exaggerated hypothalamic–pituitary–adrenal response was reversed by the introduction of Bifidobacterium infantis and was somewhat reversed with stool from orthodoxly raised mice. Germ-free mice also exhibit reduced anxiety-like behaviour in addition to altered levels of brain-derived neurotrophic factors and other neurotransmitters. In 2017, Meson et al. investigated that certain gut bacteria were connected to mood symptoms in a clinical cohort of major depressive disorder patients. In this study, species richness, or the total number of detected gut bacteria, was predictive of insomnia and depression, while abundance of Enterobacteriaceae was predictive of anxiety. In the same investigation, Lactobacillus and Enterococcus abundance was also positively related to psychomotor agitation. In 2015, Luna and Foster suggested particular administration of Lactobacillus sp., Bifidobacterium sp., L. helveticus, B. longum, L. rhamnosus and Lactobacillus farciminis in murine sample led to an improvement of depression and anxiety symptoms.
3.2 Autism
Autism spectrum disorder (ASD) is a prevalent neurodevelopmental condition with no known aetiology or cure. Several possible contributing factors, both genetic and environmental, are being actively investigated. Amongst these, maternal immune dysregulation has been identified as potentially involved in promoting ASD in the offspring. An important role of gut microbiota in the maintenance of physiological state into the gastrointestinal system is supported by several studies that have shown a qualitative and quantitative alteration of the intestinal flora in a number of gastrointestinal and extra-gastrointestinal diseases. Approximately 30–50% of children and adults with autism spectrum disorders have chronic gastrointestinal symptoms, typically constipation, diarrhoea and alternating constipation and diarrhoea (Adams et al. 2019); many of them also show abnormal behavioural patterns such as aggression, anxiety and tendency to self-injure (Afroz and Alvina 2019). It has been demonstrated that a large amount of species under the genus Clostridium (ten times more) characterized the qualitative composition of faecal samples of autistic children. The composition of microbiota has been characterized, showing an imbalance of the phyla Bacteroidetes and Firmicutes (Mangiola et al. 2016). Some of the microbial products, e.g. various metabolites of aromatic amino acids, have the potential to be neuroactive and affect the functions of the enteric and central nervous systems.
Moreover, ASD patients have significantly higher intestinal permeability which causes leakage of lymphocytes and pro-inflammatory cytokines into the circulatory system. Those inflammatory molecules eventually reach the brain and cause immune activation there (Alexeev et al. 2018; Ashwood et al. 2011). As gut dysbiosis is responsible for the increased permeability of the intestinal epithelial cells, this evidence supports the idea that there is an important effect of gut dysbiosis on immune dysregulation and possibly on ASD (Afroz and Alvina 2019; Quigley 2016). Averina et al. (2020) using a whole metagenome sequencing approach found that significant differences with decreases in average abundance in the microbiota of ASD children were found for the genera Barnesiella and Parabacteroides and species Alistipes putredinis, B. caccae, Bacteroides intestinihominis, Eubacterium rectale, Parabacteroides distasonis and Ruminococcus lactaris. They also noted decreases in the abundance of genes linked to production of GABA, melatonin and butyric acid in the ASD metagenomes. In a recent research with a mouse model of autism, Sutterella correlated with a low performance in social and obsessive-compulsive disorder (marble burying) tests and TNF-α levels (Coretti et al. 2017).
3.3 Parkinson’s Disease
Although Parkinson’s disease (PD) has been the most intensively studied, the microbiome is of interest across a range of neurodegenerative disorders. PD presently is conceptualized as a protein aggregation disease in which pathology involves both the enteric and the central nervous system, possibly spreading from one to another via the vagus nerves. PD may be of particular relevance, given the high prevalence of gastrointestinal disturbances that often precede the more well-recognized motor symptoms. An overstimulation of the innate immune system due to gut dysbiosis and/or small intestinal bacterial overgrowth, together with higher intestinal barrier permeability, may provoke local and systemic inflammation as well as enteric neuroglial activation, ultimately triggering the development of alpha-synuclein pathology. The gut microbiota and its relevant metabolites interact with the host via a series of biochemical and functional inputs, thereby affecting host homeostasis and health. Indeed, a dysregulated microbiota–gut–brain axis in PD might lie at the basis of gastrointestinal dysfunctions (Caputi and Giron 2018).
Although findings have been varied, there are some clear trends evident in the microbiome composition of patients with PD. Several studies showed an increase of Lactobacillus, Bifidobacterium, Akkermansia and Verrucomicrobiaceae in PD, while Faecalibacterium, Coprococcus, Blautia and Prevotella appear to be underrepresented (Quigley 2017; Butler et al. 2019). Similarly, Scheperjans et al. (2015) and Unger et al. (2016) found that PD patients showed a different gut microbiota than healthy controls, which was characterized by lower abundance of Prevotellaceae, Lactobacillaceae and the butyrate producer Faecalibacterium prausnitzii, whereas Enterobacteriaceae and Bifidobacterium spp. were more abundant.
Although most of the differences were associated with disease duration, lower abundance in Lachnospiraceae was the only difference between de novo PD patient and healthy control (remaining lower across almost all PD duration strata). Decreased Lachnospiraceae and increased Lactobacillaceae and Christensenellaceae were associated with a worse clinical profile, including higher frequencies of cognitive impairment, gait disturbances and postural instability. Gut microbiota may be an environmental modulator of the pathogenesis of PD and may contribute to the interindividual variability of clinical features (Barichella et al. 2019). Trace amines and their primary receptor, trace amine-associated receptor-1 (TAAR1), are widely studied for their involvement in the pathogenesis of neuropsychiatric disorders despite being found in the gastrointestinal tract at physiological levels. A therapeutic benefit of TAAR1 compounds in clinical trials is thoughtful manipulation of the brain–gut–microbiome axis to modulate symptoms of neuropsychiatric disease (Bugda Gwilt et al. 2020).
3.4 Alzheimer’s Disease
Alzheimer’s disease (AD) is the most common form of dementia and one of the major causes of disability and dependency in older people. The diversity of the gut microbiota declines in the elderly and in patients with Alzheimer’s disease (AD). Restoring the diversity with probiotic treatment alleviates the psychiatric and histopathological findings. The three different linkages between the present gut microbiome hypothesis and the other major theories for the pathogenesis of AD are as follows: bacterial metabolites and amyloids can trigger central nervous system inflammation and cerebrovascular degeneration; impaired gut microbiome flora inhibits the autophagy-mediated protein clearance process; and gut microbiomes can change the neurotransmitter levels in the brain through the vagal afferent fibres (Bostanciklioglu 2019).
Moreover, impaired memory and learning involve the dysfunction neurotransmission of glutamate, the agonist of the N-methyl-D-aspartate receptor and a major excitatory neurotransmitter in the brain. Gut microbiota including Bacteroides vulgatus and Campylobacter jejuni affect glutamate metabolism and decrease the glutamate metabolite 2-keto-glutaramic acid. Meanwhile, gut bacteria with glutamate racemase including Corynebacterium glutamicum, Brevibacterium lactofermentum and Brevibacterium avium can convert L-glutamate to D-glutamate. N-methyl-D-aspartate receptor (NMDAR)-enhancing agents have been found to potentially improve cognition in AD or Parkinson’s disease patients. These findings suggest that D-glutamate (D-form glutamate) metabolized by the gut bacteria may influence the glutamate NMDAR and cognitive function in dementia patients (Chang et al. 2020). Through metabolic activity of non-pathological microorganisms and secretion of functional by-products that increase the permeability of the intestinal mucosa, the gut microbiota influences both the production and absorption of neurotransmitters (e.g. serotonin and GABA), increasing their bioavailability to the CNS. It has been further shown some components of the gut microbiota—predominantly bacteria—synthesize and release amyloid peptides and lipopolysaccharides, which in turn activate inflammatory signalling through the release of cytokines, with potential effects on the pathophysiological cascade of Alzheimer’s disease (Vanessa et al. 2018). Depleting intestinal microbiota in AD animal models reduces amyloid-beta (Abeta) plaque deposition. Age-related changes in the microbiota contribute to immunologic and physiologic decline. Translationally relevant dietary manipulations may be an effective approach to slow microbiota changes during aging.
4 Conclusion
Due to the rapid pace of microbial science discovery, many additional functions of the microbiome are likely to be discovered. Researchers are increasingly aware that the gut and the brain communicate and are looking to leverage actions of healthy gut microbiota to treat psychological conditions. Diversity in the gut microbiota is vital not only for gut health but also for normal physiologic functioning in other organs, especially the brain. Sometimes, an altered gut microbiota in the form of dysbiosis at the extremes of life, both in the neonate and in the elderly, can have a profound impact on brain functioning. The brain is reliant on gut microbes for essential metabolic outcomes; it is not surprising that a dysbiosis can have serious negative consequences for brain function both from neurologic and mental health perspectives. However, the microbiome is a complex and dynamic ecosystem, and understanding its role in host illness and its potential for the treatment of neurological disorder will ultimately require more study.
References
Adams JB, Borody TJ, Dae-Wook K, Alexander K, Rosa K, Michael JS (2019) Microbiota transplant therapy and autism: lessons for the clinic. Expert Rev Gastroenterol Hepatol 13(11):1033–1037
Afroz KF, Alvina K (2019) Maternal elevated salt consumption and the development of autism spectrum disorder in the offspring. J Neuroinflammation 16(1):265
Aguilera M, Gálvez-Ontiveros Y, Rivas A (2020) Endobolome, a new concept for determining the influence of microbiota disrupting chemicals (MDC) in relation to specific endocrine pathogenesis. Front Microbiol 11:578007
Alex S, Lange K, Amolo T, Grinstead JS, Haakonsson AK, Szalowska E, Koppen A, Mudde K, Haenen D, Lahham S, Roelofsen H, Houtman R, Burg B, Mandrup BA, Kalkhoven E, Müller M, Hooiveld G, Kersten S (2013) Short chain fatty acids stimulate angiopoietin-like 4 synthesis in human colon adenocarcinoma cells by activating peroxisome proliferator-activated receptor gamma. Mol Cell Biol 33:1303–1316
Alexeev EE, Lanis JM, Kao DJ, Campbell EL, Kelly CJ, Battista KD, Gerich ME, Jenkins BR, Walk ST, Kominsky DJ, Colgan SP (2018) Microbiota-derived indole metabolites promote human and murine intestinal homeostasis through regulation of interleukin-10 receptor. Am J Pathol 188(5):1183–1194
Andersson AF, Lindberg M, Jakobsson H, Bäckhed F, Nyrén P (2008) Comparative analysis of human gut microbiota by barcoded pyrosequencing. PLoS One 3:e2836
Arboleya S, Binetti A, Salazar N, Fernández N, Solís G, Hernandez-Barranco A, Armougom F, Raoult D (2008) Use of pyrosequencing and DNA barcodes to monitor variations in Firmicutes and Bacteroidetes communities in the gut microbiota of obese humans. BMC Genomics 9:576
Arumugam M, Raes J, Pelletier E, Arumugam M, Raes J, Pelletier E, Paslier D, Takuji Y, Daniel RM, Gabriel RF, Julien T, Thomas B, Jean MB, Marcelo B, Natalia B, Francesc C, Leyden F, Laurent G, Torben H, Masahira H, Tetsuya H, Michiel K, Ken K, Marion L, Florence L, Chaysavanh M, Nielsen H, Trine N, Nicolas P, Julie P, Junjie Q, Thomas S, Sebastian T, David T, Edgardo U, Erwin GZ, Junwang FG, Oluf P, Willem M, Joel D, Metahit C, Jean W, Ehrlich SD (2011) Enterotypes of the human gut microbiome. Nature 473:174–180
Ashwood P, Krakowiak P, Hertz-Picciotto I, Hansen R, Pessah I, Van de Water J (2011) Elevated plasma cytokines in autism spectrum disorders provide evidence of immune dysfunction and are associated with impaired behavioral outcome. Brain Behav Immun 25(1):40–45
Averina OV, Kovtun AS, Polyakova SI, Savilova AM, Rebrikov DV, Danilenko VN (2020) The bacterial neurometabolic signature of the gut microbiota of young children with autism spectrum disorders. J Med Microbiol 69(4):558–571
Bäckhed F, Ding H, Wang T, Hooper LV, Koh GY, Nagy A, Clay FS, Gordon JI (2004) The gut microbiota as an environmental factor that regulates fat storage. Proc Natl Acad Sci 101(44):15718–15723
Baggio LL, Drucker DJ (2007) Biology of incretins: GLP-1 and GIP. Gastroenterology 132(6):2131–2157
Balanza-Martinez V, Shansis FM, Tatay-Manteiga A, López-García P (2020) Diet and neurocognition in mood disorders - an overview of the overlooked. Curr Pharm Des 26(20):2353–2362
Barichella M, Severgnini M, Roberto C, Erica C, Carlotta B, Serena C, Valentina F, Raffaella C, Camilla C, Samanta F, Giovanna P, Gianluca B, Luigi Z, Emanuele C, Clarissa C, Gianni P (2019) Unraveling gut microbiota in Parkinson’s disease and atypical parkinsonism. Mov Disord 34(3):396–405
Barouei J, Moussavi M, Hodgson DM (2012) Effect of maternal probiotic intervention on HPA axis, immunity and gut microbiota in a rat model of irritable bowel syndrome. PLoS One 7(10):e46051
Bendtsen KMB, Krych L, Sørensen DB, Pang W, Nielsen DS, Josefsen K, Lars HH, Søren JS, Hansen AK (2012) Gut microbiota composition is correlated to grid floor induced stress and behavior in the BALB/c mouse. PLoS One 7(10):e46231
Benner M, Ferwerda G, Joosten I, Van der Molen RG (2018) How uterine microbiota might be responsible for a receptive, fertile endometrium. Hum Reprod Update 24(4):393–415
Biagi E, Nylund L, Candela M, Ostan R, Bucci L, Pini E, Nikkïla J, Monti D, Satokari R, Franceschi C, Brigidi P (2010) Through ageing, and beyond: gut microbiota and inflammatory status in seniors and centenarians. PLoS One 5(5):e10667
Blaser MJ (1999) Hypothesis: the changing relationships of Helicobacter pylori and humans: implications for health and disease. J Infect Dis 179:1523–1530
Bostanciklioglu M (2019) The role of gut microbiota in pathogenesis of Alzheimer’s disease. J Appl Microbiol 127(4):954–967
Bravo JA, Forsythe P, Chew MV, Emily E, Hélène M, Timothy GD, John B, John FC (2011) Ingestion of Lactobacillus strain regulates emotional behavior and central GABA receptor expression in a mouse via the vagus nerve. Proc Natl Acad Sci U S A 108(38):16050–16055
Bueter M, Ashrafian H, le Roux CW (2009) Mechanisms of weight loss after gastric bypass and gastric banding. Obes Facts 2(5):325–331
Bugda Gwilt K, Gonzalez DP, Olliffe N, Oller H, Hoffing R, Puzan M, El Aidy S, Miller GM (2020) Actions of trace amines in the brain-gut-microbiome axis via trace amine-associated receptor-1 (TAAR1). Cell Mol Neurobiol 40(2):191–201
Butler MI, Sabrina M, Kiran VS, John FC, Timothy GD (2019) The gut microbiome and mental health: what should we tell our patients. Can J Psychiatr 64(11):747–760
Canfora EE, Jocken JW, Blaak EE (2015) Short-chain fatty acids in control of body weight and insulin sensitivity. Nat Rev Endocrinol 11(10):577
Cani PD, Amar J, Iglesias MA, Poggi M, Knauf C, Bastelica D, Burcelin R, Audrey MN, Francesca F, Kieran MT, Chantal C, Aurélie W, Evelyne D, Béatrice C, Thierry S, Bernard C, Jean F, Jean-François T, Glenn RG, Louis C, Nathalie MD, Marie CA, Rémy B (2007) Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes 56(7):1761–1772
Caputi V, Giron MC (2018) Microbiome-gut-brain axis and toll-like receptors in Parkinson’s disease. Int J Mol Sci 19:6
Chang JY, Shin SM, Chun J, Lee JH, Seo JK (2011) Pyrosequencing-based molecular monitoring of the intestinal bacterial colonization in preterm infants. J Pediatr Gastroenterol Nutr 53(5):512–519
Chang CH, Lin CH, Lane HY (2020) d-glutamate and gut microbiota in Alzheimer’s disease. Int J Mol Sci 21:8
Claesson MJ, Jeffery IB, Conde S, Power SE, O’Connor EM, Cusack S, Susan EP, Eibhlís MO, Siobhán C, Hugh MBH, Mairead C, Bhuvaneswari L, Orla O, Gerald FF, Jennifer D, Michael O, Norma H, Kieran O, Denis O, Douwe S, Martina W, Lorraine B, Catherine S, Julian RM, Anthony PF (2012) Gut microbiota composition correlates with diet and health in the elderly. Nature 488:178
Coello K, Hansen TH, Sørensen N, Munkholm K, Kessing LV, Pedersen O, Vinberg M (2019) Gut microbiota composition in patients with newly diagnosed bipolar disorder and their unaffected first-degree relatives. Brain Behav Immun 75:112–118
Coretti L, Cristiano C, Florio E, Scala G, Lama A, Keller S, Cuomo M, Russo R, Pero R, Paciello O, Raso GM, Meli R, Cocozza S, Calignano A, Chiariotti L, Lembo F (2017) Sex-related alterations of gut microbiota composition in the BTBR mouse model of autism spectrum disorder. Sci Rep 7:45356
D’Aversa F, Tortora A, Ianiro G, Ponziani FR, Annicchiarico BE, Gasbarrini A (2013) Gut microbiota and metabolic syndrome. Intern Emerg Med 8(1):11–15
David LA, Maurice CF, Carmody RN, David BG, Julie EB, Benjamin EW, Alisha VL, Sloan D, Yug V, Michael AF, Sudha BB, Rachel JD, Peter JT (2014) Diet rapidly and reproducibly alters the human gut microbiome. Nature 505:559–563
Dennison B (1976) Definition of preterm delivery. Br Med J 2(6049):1449
Desbonnet L, Garrett L, Clarke G, Bienenstock J, Dinan TG (2008) The probiotic Bifidobacteria infantis: an assessment of potential antidepressant properties in the rat. J Psychiatr Res 43(2):164–174
Dewhirst FE, Chen T, Izard J, Bruce J, Anne CRT, Wen-Han Y, Abirami L, William GW (2010) The human oral microbiome. J Bacteriol 192:5002–5017
Dinan TG, Quigley EM, Ahmed SM, Scully P, O’Brien S, O’Mahony L, Siobhan OM, Fergus S, Keeling P (2006) Hypothalamic-pituitary-gut axis dysregulation in irritable bowel syndrome: plasma cytokines as a potential biomarker. Gastroenterology 130(2):304–311
Dominguez-Bello MG, Costello EK, Contreras M, Magris M, Hidalgo G, Fierer N, Rob K (2010) Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci U S A 107:11971
Eckburg PB, Bik EM, Bernstein CN, Purdom E, Dethlefsen L, Sargent M, Steven RG, Keran EN, David AR (2005) Diversity of the human intestinal microbial flora. Science 308:1635
El Aidy S, Dinan TG, Cryan JF (2014) Immune modulation of the brain-gut-microbe axis. Front Microbiol 5:146
Flint HJ, Scott KP, Duncan SH, Louis P, Forano E (2012) Microbial degradation of complex carbohydrates in the gut. Gut Microbes 3:289–306
Ford AC, Luthra P, Tack J, Boeckxstaens GE, Moayyedi P, Talley NJ (2017) Efficacy of psychotropic drugs in functional dyspepsia: systematic review and meta-analysis. Gut 66(3):411–420
Forsythe P, Kunze WA (2013) Voices from within: gut microbes and the CNS. Cell Mol Life Sci 70(1):55–69
Frank DN, Pace NR (2008) Gastrointestinal microbiology enters the metagenomics era. Curr Opin Gastroenterol 24(1):4–10
Frank DN, St Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR (2007) Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci U S A 104:13780–13785
Furusawa Y, Obata Y, Fukuda S, Endo TA, Nakato G, Takahashi D, Yumiko N, Chikako U, Keiko K, Tamotsu K, Masumi T, Noriko NF, Shinnosuke M, Eiji M, Shingo H, Koji A, Satoshi O, Yumiko F, Trevor L, Julie MC, David LT, Masaru T, Shohei H, Osamu O, Tatsuya M, Haruhiko K, Jun K, Kenya H, Koji H, Ohno H (2013) Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504(7480):446–450
Gajer P, Brotman RM, Bai G, Joyce S, Urcel MES, Xue Z, Sara SKK, Li F, Zhanshan M, Xia Z, Zaid A, Larry JF, Jacques R (2012) Temporal dynamics of the human vaginal microbiota. Sci Transl Med 4:132–152
Ghoshal UC, Ghoshal U (2017) Small intestinal bacterial overgrowth and other intestinal disorders. Gastroenterol Clin 46(1):103–120
Gillespie JJ, Wattam AR, Cammer SA, Gabbard JL, Shukla MP, Dalay O, Driscoll T, Hix D, Mane SP, Mao C, Nordberg EK, Scott M, Schulman JR, Snyder EE, Sullivan DE, Wang C, Warren A, Williams KP, Xue T, Yoo HS, Zhang C, Zhang Y, Will R, Kenyon RW, Sobral BW (2011) PATRIC: the comprehensive bacterial bioinformatics resource with a focus on human pathogenic species. Infect Immun 79:4286–4298
González A, Vázquez-Baeza Y, Knight R (2014) SnapShot: the human microbiome. Cell 158:690–690
Gravitz L (2012) Microbiome: the critters within. Nature 485(7398):S12–S13
Grenham S, Clarke G, Cryan JF, Dinan TG (2011) Brain-gut-microbe communication in health and disease. Front Physiol 2:94
Grice EA, Kong HH, Conlan S, Clayton BD, Joie D, Alice CY, Gerard GB, Robert WB, Patrick RM, Eric DG, Maria LT, Julia S (2006) Topographical and temporal diversity of the human skin microbiome. Science 324:1190–1192
Hamady M, Knight R (2009) Microbial community profiling for human microbiome projects: tools, techniques, and challenges. Genome Res 19(7):1141–1152
Harakeh SM, Khan I, Kumosani T, Barbour E, Almasaudi SB, Bahijri SM, Azhar EI (2016) Gut microbiota: a contributing factor to obesity. Front Cell Infect Microbiol 6:95
Harsch IA, Konturek PC (2018) The role of gut microbiota in obesity and type 2 and type 1 diabetes mellitus: new insights into “old” diseases. Med Sci 6(2):32
Hollister EB, Gao C, Versalovic J (2014) Compositional and functional features of the gastrointestinal microbiome and their effects on human health. Gastroenterology 146:1449–1458
Hoy CM, Wood CM, Hawkey PM, Puntis JW (2000) Duodenal microflora in very-low-birth-weight neonates and relation to necrotizing enterocolitis. J Clin Microbiol 38(12):4539–4547
Human Microbiome Project Consortium (2012) Structure, function and diversity of the healthy human microbiome. Nature 486:22699609
Jacquot A, Neveu D, Aujoulat F, Mercier G, Marchandin H, Jumas-Bilak E, Picaud JC (2011) Dynamics and clinical evolution of bacterial gut microflora in extremely premature patients. J Pediatr 158(3):390–396
Jalanka-Tuovinen J, Salonen A, Nikkila J, Immonen O, Kekkonen R, Lahti L, Palva A, DeVos WM (2011) Intestinal microbiota in healthy adults: temporal analysis reveals individual and common core and relation to intestinal symptoms. PLoS ONE 6:e23035. https://doi.org/10.1371/journal.pone.0023035
Justesen T, Nielsen OH, Jacobsen IE, Lave J, Rasmussen SN (1984) The normal cultivable microflora in upper jejunal fluid in healthy adults. Scand J Gastroenterol 19:279–282
Kennedy PJ, Cryan JF, Dinan TG, Clarke G (2014) Irritable bowel syndrome: a microbiome-gut-brain axis disorder. World J Gastroenterol 20(39):14105
Khanna S, Tosh PK (2014) A clinician’s primer on the role of the microbiome in human health and disease. Mayo Clin Proc 89:107–114
Kim YS, Unno T, Kim BY, Park MS (2020) Sex differences in gut microbiota. World J Men’s Health 38(1):48
Koenig JE, Spor A, Scalfone N, Fricker AD, Stombaugh J, Knight R, Largus T (2011) Succession of microbial consortia in the developing infant gut microbiome. Proc Natl Acad Sci U S A 108(Suppl. 1):4578–4585
Koren O, Goodrich JK, Cullender TC, Spor A, Laitinen K, HeleneKling B, Gonzalez A, Jeffrey JW, Largus TA, Rob K, Fredrik B, Erika I, Seppo S, Ruth EL (2012) Host remodeling of the gut microbiome and metabolic changes during pregnancy. Cell 150:470–480
Kuo PH, Chung YE (2019) Moody microbiome: challenges and chances. J Formos Med Assoc 118(Suppl 1):S42–S54
Ley RE, Bäckhed F, Turnbaugh P, Lozupone CA, Knight RD, Gordon JI (2005) Obesity alters gut microbial ecology. Proc Natl Acad Sci 102(31):11070–11075
Ley RE, Turnbaugh PJ, Klein S, Gordon JI (2006) Microbial ecology: human gut microbes associated with obesity. Nature 444:1022–1023
Lobo C, Moreno-Ventas X, Tapia-Paniagua S, Rodriguez C, Morinigo MA, De La Banda IG (2014) Dietary probiotic supplementation (Shewanella putrefaciens Pdp11) modulates gut microbiota and promotes growth and condition in Senegalese sole larviculture. Fish Physiol Biochem 40:295–309
Luckey TD (1972) Introduction to intestinal microecology. Am J Clin Nutr 25:12
Lynch SV, Pedersen O (2016) The human intestinal microbiome in health and disease. N Engl J Med 375(24):2369–2379
Lyte M (2013) Microbial endocrinology in the microbiome-gut-brain axis: how bacterial production and utilization of neurochemicals influence behavior. PLoS Pathog 9(11):e1003726
Lyte M (2014) Microbial endocrinology and the microbiota-gut-brain axis. Microb Endocrinol 2014:3–24
Mangiola F, Ianiro G, Franceschi F, Fagiuoli S, Gasbarrini G, Gasbarrini A (2016) Gut microbiota in autism and mood disorders. World J Gastroenterol 22(1):361
Marques TM, Wall R, Ross RP, Fitzgerald GF, Ryan CA, Stanton C (2010) Programming infant gut microbiota: influence of dietary and environmental factors. Curr Opin Biotechnol 21:149–156
Mayer EA, Knight R, Mazmanian SK, Cryan JF, Tillisch K (2014) Gut microbes and the brain: paradigm shift in neuroscience. J Neurosci 34(46):15490–15496
McKernan DP, Dennison U, Gaszner G, Cryan JF, Dinan TG (2011) Enhanced peripheral toll-like receptor responses in psychosis: further evidence of a pro-inflammatory phenotype. Transl Psychiatry 1(8):e36
Moloney RD, Desbonnet L, Clarke G, Dinan TG, Cryan JF (2014) The microbiome: stress, health and disease. Mamm Genome 25(1):49–74
Montiel-Castro AJ, Gonzalez-Cervantes RM, Bravo-Ruiseco G, Pacheco-Lopez G (2013) The microbiota-gut-brain axis: neurobehavioral correlates, health and sociality. Front Integr Neurosci 7:70
Moreno I, Codoñer FM, Vilella F, Valbuena D, Martinez-Blanch JF, Jimenez-Almazán J, Simon C (2016) Evidence that the endometrial microbiota has an effect on implantation success or failure. Am J Obstet Gynecol 215(6):684–703
Musso G, Gambino R, Cassader M, Pagano G (2011) Meta-analysis: natural history of non-alcoholic fatty liver disease (NAFLD) and diagnostic accuracy of non-invasive tests for liver disease severity. Ann Med 43(8):617–649
Nair AT, Ramachandran V, Joghee NM, Antony S, Ramalingam G (2018) Gut microbiota dysfunction as reliable non-invasive early diagnostic biomarkers in the pathophysiology of Parkinson’s disease: a critical review. J Neurogastroenterol Motil 24(1):30
O’Brien SM, Scott LV, Dinan TG (2004) Cytokines: abnormalities in major depression and implications for pharmacological treatment. Hum Psychopharmacol 19(6):397–403
O’Hara AM, Shanahan F (2006) The gut flora as a forgotten organ. EMBO Rep 7:688–693
O’Mahony L, McCarthy J, Kelly P, Hurley G, Luo F, Chen K, Quigley EM (2005) Lactobacillus and bifidobacterium in irritable bowel syndrome: symptom responses and relationship to cytokine profiles. Gastroenterology 128(3):541–551
O’Mahony SM, Marchesi JR, Scully P, Codling C, Ceolho AM, Quigley EM, John FC, Dinan TG (2009) Early life stress alters behavior, immunity, and microbiota in rats: implications for irritable bowel syndrome and psychiatric illnesses. Biol Psychiatry 65(3):263–267
O’Mahony SM, Hyland NP, Dinan TG, Cryan JF (2011) Maternal separation as a model of brain–gut axis dysfunction. Psychopharmacology 214(1):71–88
O’Mahony SM, Clarke G, Dinan TG, Cryan JF (2017) Early-life adversity and brain development: is the microbiome a missing piece of the puzzle? Neuroscience 342:37–54
Palmer C, Bik EM, Digiulio DB, Relman DA, Brown PO (2007) Development of the human infant intestinal microbiota. PLoS Biol 5:e177
Pei Z, Bini EJ, Yang L, Zhou M, Francois F, Blaser MJ (2004) Bacterial biota in the human distal esophagus. Proc Natl Acad Sci U S A 101:4250–4255
Perez-Cobas AE, Gosalbes MJ, Friedrichs A, Henrik K, Alenjandro A, Kathleen E, Wolfgng O, David R, Rafael B, Martin VB, Sven CN, Carolin D, Femke AH, Amparo L, Coral B, Jana S, MDS V, Stephan JO, Manuel F, Andres M (2013) Gut microbiota disturbance during antibiotic therapy: a multi-omic approach. Gut 62:1591–1601
Qin J, Li R, Raes J et al (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464:59–65
Qin J, Li Y, Cai Z, Li S, Zhu J, Zhang F, Suisha L, Wenwei Z, Yuanlin G, Dongqian S, Yangqing P, Dongya Z, Zhuye J, Wenxian W, Youwen Q, Wenbin X, Junhua L, Lingchuan H, Donghui L, Peixian W, Yali D, Xiaojuan S, Zesong L, Aifa T, Shilong Z, Xiaoping L, Weineng C, Ran X, Mingbang W, Qiang F, Meihua G, Jing Y, Yanyan Z, Ming Z, Torben H, Gaston S, Jeroen R, Gwen F, Shujiro O, Mathieu A, Emmanuelle L, Pierre R, Nicolas P, Jean-Michel B, Zhaoxi Z, Hua C, Ruifu Y, Weimou Z, Songgang L, Huanming Y, Jian W, Dusko E, Rasmus N, Oluf P, Karsten K, Wang J (2012) A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490(7418):55–60
Quigley EM (2016) Leaky gut - concept or clinical entity? Curr Opin Gastroenterol 32(2):74–79
Quigley EMM (2017) Microbiota-brain-gut axis and neurodegenerative diseases. Curr Neurol Neurosci Rep 17(12):94
Rajilic-Stojanovic M, Shanahan F, Guarner F, deVos WM (2012) Phylogenetic analysis of dysbiosis in ulcerative colitis in relapse and remission. Gut 19(3):481–488
Ramakrishna B, Krishnan S (2007) The normal bacterial flora of the human intestine and its regulation. J Clin Gastroenterol 41:S2–S6
Razavi AC, Potts KS, Kelly TN, Bazzano LA (2019) Sex, gut microbiome, and cardiovascular disease risk. Biol Sex Differ 10(1):1–14
Scheperjans F, Aho V, Pereira PA, Koskinen K, Paulin L, Pekkonen E, Haapaniemi E, Kaakkola S, Eerola-Rautio J, Pohja M, Esko K, Kari M, Petri A (2015) Gut microbiota are related to Parkinson’s disease and clinical phenotype. Mov Disord 30:350–358
Scholtens PA, Oozeer R, Martin R, Amor KB, Knol J (2012) The early settlers: intestinal micro-biology in early life. Annu Rev Food Sci Technol 3:425–447
Schwartz MW, Woods SC, Porte D, Seeley RJ, Baskin DG (2000) Central nervous system control of food intake. Nature 404(6778):661–671
Schwiertz A, Taras D, Schäfer K, Beijer S, Bos NA, Donus C, Hardt PD (2010) Microbiota and SCFA in lean and overweight healthy subjects. Obesity 18(1):190–195
Scott LV, Clarke G, Dinan TG (2013) The brain-gut axis: a target for treating stress-related disorders. Inf Psychiatr 28:90–99
Sekirov I, Russell SL, Antunes LCM, Finlay BB (2010) Gut microbiota in health and disease. Physiol Rev 90(3):859–904
Shanahan F (2012) The gut microbiota-a clinical perspective on lessons learned. Nat Rev Gastroenterol Hepatol 9:609–614
Sherwin E, Sandhu KV, Dinan TG, Cryan JF (2016) May the force be with you: the light and dark sides of the microbiota-gut-brain axis in neuropsychiatry. CNS Drugs 30(11):1019–1041
Sondheimer JM, Clark DA (1985) Gastric pH in healthy preterm infants-effect of age and feeding type. In: Gastroenterology. WB Saunders Co., Philadelphia
Sondheimer JM, Clark DA, Gervaise EP (1985) Continuous gastric pH measurement in young and older healthy preterm infants receiving formula and clear liquid feedings. J Pediatr Gastroenterol Nutr 4(3):352–355
Stilling RM, Dinan TG, Cryan JF (2014) Microbial genes, brain & behaviour–epigenetic regulation of the gut–brain axis. Genes Brain Behav 13(1):69–86
Sudo N, Chida Y, Aiba Y, Sonoda J, Oyama N, Yu XN et al (2004) Postnatal microbial colonization programs the hypothalamic–pituitary–adrenal system for stress response in mice. J Physiol 558(1):263–275
Svensson E, Horváth-Puhó E, Thomsen RW, Djurhuus JC, Pedersen L, Borghammer P, Sørensen HT (2015) Vagotomy and subsequent risk of Parkinson’s disease. Ann Neurol 78(4):522–529
Swidsinski A, Loening-Baucke V, Lochs H, Hale LP (2005) Spatial organization of bacterial flora in normal and inflamed intestine: a fluorescence in situ hybridization study in mice. World J Gastroenterol 11:1131–1140
Takeuchi O, Akira S (2010) Pattern recognition receptors and inflammation. Cell 140(6):805–820
Talham GL, Jiang HQ, Bos NA, Cebra JJ (1999) Segmented filamentous bacteria are potent stimuli of a physiologically normal state of the murine gut mucosal immune system. Infect Immun 67(4):1992–2000
Tillisch K (2014) The effects of gut microbiota on CNS function in humans. Gut Microbes 5(3):404–410
Tolhurst G, Heffron H, Lam YS, Parker HE, Habib AM, Diakogiannaki E, Jennifer C, Johannes G, Frank R, Gribble FM (2012) Short-chain fatty acids stimulate glucagon-like peptide-1 secretion via the G-protein–coupled receptor FFAR2. Diabetes 61(2):364–371
Travagli RA, Hermann GE, Browning KN, Rogers RC (2003) Musings on the wanderer: what’s new in our understanding of vago-vagal reflexes: III. Activity-dependent plasticity in vago-vagal reflexes controlling the stomach. Am J Physiol 284(2):G180
Turnbaugh PJ, Gordon JI (2009) The core gut microbiome, energy balance and obesity. J Physiol 587(17):4153–4158
Umesaki Y, Okada Y, Matsumoto S, Imaoka A, Setoyama H (1995) Segmented filamentous bacteria are indigenous intestinal bacteria that activate intraepithelial lymphocytes and induce MHC class II molecules and fucosyl asialo GM1 glycolipids on the small intestinal epithelial cells in the ex-germ-free mouse. Microbiol Immunol 39(8):555–562
Umesaki Y, Setoyama H, Matsumoto S, Imaoka A, Itoh K (1999) Differential roles of segmented filamentous bacteria and clostridia in development of the intestinal immune system. Infect Immun 67(7):3504–3511
Unger MM, Spiegel J, Dillmann K-U, Grundmann D, Philippeit H, Bürmann J, Klaus F, Andreas S, Karl-Herbert S (2016) Short chain fatty acids and gut microbiota differ between patients with Parkinson’s disease and age-matched controls. Parkinsonism Relat Disord 32:66–72
Upadhyay J, Farr O, Perakakis N, Ghaly W, Mantzoros C (2018) Obesity as a disease. Med Clin 102(1):13–33
Vanessa JRD, Paula V, Forlenza AS (2018) Relevance of gut microbiota in cognition, behaviour and Alzheimer’s disease. Pharmacol Res 136:29–34
Vrieze A, Van Nood E, Holleman F, Salojärvi J, Kootte RS, Bartelsman JF, Geesje MD, Mariette TA, Mireille JS, Raish O, Muriel D, Anne D, Johan ET, Van HV, Vincent WB, Albert KG, Hans GH, Erwin GZ, Erik SS, Willem M, Joost BLH, Max N, Nieuwdorp M (2012) Transfer of intestinal microbiota from lean donors increases insulin sensitivity in individuals with metabolic syndrome. Gastroenterology 143(4):913–916
Walsh CJ, Guinane CM, O’Toole PW, Cotter PD (2014) Beneficial modulation of the gut microbiota. FEBS Lett 588(22):4120–4130
Wang Y, Kasper LH (2014) The role of microbiome in central nervous system disorders. Brain Behav Immun 38:1–12
Wu GD, Chen J, Hoffmann C, Bittinger K, Chen YY, Keilbaugh SA, Bewtra M, Knights D, Walters WA, Knight R, Sinha R (2011) Linking long-term dietary patterns with gut microbial enterotypes. Science 334(6052):105–108
Yatsunenko T, Rey FE, Manary MJ, Indi T, Maria GD, Monica C, Magda M, Glida H, Robert NB, Andrey PA, Andrew C, Barbara W, Jens R, Justin KJ, Gregory C, Catherine AL, Christian L, Jose CC, Dan K, Rob K, Jeffrey IG (2012) Human gut microbiome viewed across age and geography. Nature 486:222–227
Yuan X, Chen R, Zhang Y, Lin X, Yang X (2020) Sexual dimorphism of gut microbiota at different pubertal status. Microb Cell Factories 19(1):1–16
Yurtdaş G, Akdevelioğlu Y (2020) A new approach to polycystic ovary syndrome: the gut microbiota. J Am Coll Nutr 39(4):371–382
Zhao X, Jiang Y, Xi H, Chen L, Feng X (2020) Exploration of the relationship between gut microbiota and polycystic ovary syndrome (PCOS): a review. Geburtshilfe Frauenheilkd 80(2):161
Zheng J, Li H, Zhang X, Jiang M, Luo C, Lu Z, Xu Z, Shi J (2018) Prebiotic mannan-oligosaccharides augment the hypoglycemic effects of metformin in correlation with modulating gut microbiota. J Agric Food Chem 66(23):5821–5831
Zoetendal EG, Rajilic-Stojanovic M, De Vos WM (2008) High-throughput diversity and functionality analysis of the gastrointestinal tract microbiota. Gut 57:1605–1615
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Poojara, L., Acharya, D.K., Patel, J., Rawal, R.M. (2022). Gut–Brain Axis: Role of the Gut Microbiome on Human Health. In: Sayyed, R.Z., Khan, M. (eds) Microbiome-Gut-Brain Axis. Springer, Singapore. https://doi.org/10.1007/978-981-16-1626-6_8
Download citation
DOI: https://doi.org/10.1007/978-981-16-1626-6_8
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-1625-9
Online ISBN: 978-981-16-1626-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)