Abstract
In this work, we have developed a DNA AND logic gate based on graphene oxide (GO) absorbing single DNA and G-quadruplex interacting with N-methyl mesoporphyrin IX(NMM) for detecting breast cancer biomarker carbohydrate antigen 15-3 (CA15-3). With CA15-3 and NMM as the two inputs, the fluorescence intensity of the NMM is the output signal. A hairpin DNA probe is designed, consisting of CA15-3 aptamer and partly anti-CA15-3 aptamer sequences as a long stem and G-rich sequences as a quadruplex-forming oligomer. In the presence of CA15-3 or NMM alone, there is no significant fluorescence enhancing, and the output of the signal is “0”. While in the presence of CA15-3 protein and NMM, the fluorescence signal was dramatically increasing and the output of the signal is “1”. This biosensor platform also exhibited good reproducibility, selectivity and showed high sensitivity for CA15-3 protein in a range of 10U/mL-500U/mL with the detection limit of 10U/mL. In addition, the fluorescence DNA AND logic gate can successfully be applied to the determination of CA15-3 in spiked human serum. In summary, the proposed fluorescent DNA AND logic gate could construct a simple, fast, label-free and highly specific sensing methods for CA15-3.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
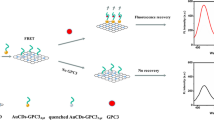
Molecular logic system is a device to realize logical processing of molecular information based on the interaction between molecular modules in physical, chemical and biological processes [1]. Through the combination of molecular logic system and all kinds of analysis and detection technology, it can construct a digital, multi-group differentiation, intelligent biochemical sensing device, which has a wide application prospect in nanotechnology, biomedicine, biochemical analysis, environmental analysis and other fields. For instance, Huo et al. [2] successfully constructed a series of colorimetric and fluorescent logic gate (AND, NAND, OR, NOR, INHIBIT and IMPLICATION) based on the interaction of TMPipPrOPP and G-Quadruplex to realize pH-sensing. Lu et al. [3] used the recovery of fluorescence intensity and no fluorescence recovery as the output signal to detect arginine and Cu2+. Moreover, Deng et al. [4] finished a pathogenic bacterial gene-induced logically reversible logic gate, which is good for implement advanced computational processing. Molecular logic gate attracted extensive attention in terms of biosensors [5], intelligent diagnosis [4] and molecular computing [6] in recent years. Although obtained many progressive results, it is challenging to construct easy-to-design, versatile and robust logic circuits integrated on a molecular scale.
However, molecular logic gate always used fluorescent dyes to realize fluorescence signal, [7, 8] this method is relatively expensive, low yield and singly labeled impurities and fluorescent labeling may have a certain effect on the affinity between the target and the aptamer. As a result, non-fluorescent labeled fluorescent aptamer biosensor technology has been developed. G-quadruplex is a kind of DNA or RNA secondary structure, which contains guanine-rich bases. Every four guanine-rich bases are connected by Hoogsteen hydrogen bond and fold into a helix formation, which is first discovered in 1962 [9]. The G-quadruplex is stabilized through K+, Na+ and Mg2+ , etc. [10, 11]. Furthermore, hemin, N-methylmesoporphyrin IX (NMM) or Thioflavine T can be inserted into the structure of G-quadruplex to construct colorimetric and fluorescent biosensors [12,13,14,15,16], which can lower the fluorescence back signal. Two-dimensional graphene oxide nanosheets have been widely used in fluorescent aptasensor because of their good properties [17,18,19,20,21]. GO can absorb single strand DNA via π-π stacking between nucleobases and GO, rather than double-stranded DNA, triple-strand DNA or aptamer–target complexes [22, 23]. Owing to simple and specific interactions between targets and aptamers, good water dispersibility and biocompatibility of GO [24], many GO-based fluorescent aptasensor have been constructed for a variety of targets detection [25,26,27,28,29,30].
Here we constructed a DNA AND logic gate based on graphene oxide (GO) absorbing single DNA and G-quadruplex interacting with NMM for detecting breast cancer biomarker carbohydrate antigen 15-3 (CA15-3). CA15-3 can specifically interact with aptamer and open the hairpin structures, leading the G-rich sequence fell off from GO and G-quadruplex stabilized through K+. And then, NMM can be bind to G-quadruplex, which caused a great fluorescence enhancement. In contrast, the hairpin structures cannot be opened without CA15-3 added into experiment, which are strongly absorbed on GO resulting no fluorescent signal. Therefore, a DNA AND logic gate can be constructed using CA15-3 and NMM as the two inputs, and the fluorescence signal as the output. Furthermore, this fluorescent DNA AND logic gate was used for the assay of CA15-3 in a biological sample and satisfactory experimental results were obtained.
2 Materials and Methods
2.1 Reagents and Materials
The carbohydrate antigen 15-3 (CA15-3) were purchased from Shanghai Linc-Bio Science Co. Ltd. Graphene oxide was purchased from XFNANO Co. Ltd. (Nanjing, China, http://www.xfnano.com). The healthy human serum and streptavidin (SA) were obtained from solarbio (Beijing, China, http://www.solarbio.com). The lysozyme, thrombin and bovine serum albumin (BSA) were purchased from Sigma (St. Louis, MO, USA). The hpDNA (5’GAAGTGAATATGACAGATCACAACTAATGGGTAGGGCGGGTGGGAGTTGTGATCTGTCA-3’) were synthesized by Sangon Biological Engineering Technology Co. Ltd. (Shanghai, China, http://www.sangon.com) and purified using high performance liquid chromatography. The underline indicates the CA15-3 aptamer, and the bold denotes the stem region of the G-rich sequences. N-methylmesoporphyrin IX (NMM) was purchased from J&K Scientific Ltd. (Beijing, China). All of the reagents were diluted to the required concentration with working buffer (10 mM PBS, pH = 7.2) before use.
2.2 Assay Procedures
Firstly, 250 nM hpDNA and 5 mg mL−1 GO was first mixed and kept at 37 ℃ for 30 min, followed by adding 100U mL−1 of CA15-3. 30 min later, the reaction solution was added with 20 mM K+, 1 μM NMM and incubated at 37 ℃ for 25 min, then the solution was diluted to 100μL. Finally, the fluorescence of the mixture was carried out on an EnSpire ELIASA from PerkinElmer. In control experiments, the measurement process was all the same with the above except the addition of CA15-3. Unless otherwise noted, each fluorescence measurement was repeated three times, and the standard deviation was plotted as the error bar.
3 Results
3.1 Principle of Design
In the present study, the principle of AND logic gate was constructed (Fig. 1). In the absence of NMM and CA15-3 input, the result shows no prominent fluorescence band at 610 nm output 0. Operation by NMM as (Input 1) to this mixture solution, the output remained ‘0’ and with the sole addition of another input CA15-3 (input 2), the output remained ‘0’ again. In the simultaneous presence of both the chemical inputs (1 and 2), the output becomes ‘1’. Thus, the strategy of detecting CA15-3 for AND molecular logic gate is turned out. The implementation of this fluorescence AND logic platform is mainly due to the hairpin DNA probe consisting of CA15-3 aptamer and partly anti- CA15-3 aptamer sequences as a long stem and a G-rich sequences as a quadruplex-forming oligomer. In the absence of CA15-3, the GO can strongly adsorb single-stranded nucleic acids, thus, the hpDNA is adsorbed onto the surface of GO via π–π stacking between DNA bases and GO, resulting in low fluorescence signal. Upon adding the CA15-3, the aptamer sequences could be specific recognized by CA15-3 to form aptamer/CA15-3 complexes, leading to explode G-rich sequence. Then, when the K+ and NMM are added, G-rich sequence can fold into a quadruplex by K+, and then G-quadruplex interacts with NMM, which is a specific G-quadruplex binder, leading to a dramatic increase in fluorescence of NMM. The quantities of CA15-3 can be achieved by fluorescence increment.
3.2 Feasibility Analysis of the Developed Method for CA15-3 Detection
To further verify the feasibility of our strategy, Fig. 2 shows the fluorescence emission spectra under different conditions. When mixture solution containing GO, hp DNA and K+, there also was low fluorescence (black line), showing that hpDNA can strongly be absorbed on GO sheets. The fluorescence signal produced by mixture solution containing hpDNA, GO, K+ and NMM (red line) is relatively weak in the absence of CA15-3, because CA15-3 cannot specifically bind to aptamer and open the hairpin DNA. And then, when mixture solution containing hpDNA, GO, K+ and CA15-3, there was low fluorescence (blue line), since no NMM interacting with G-quadruplex. However, upon adding NMM and CA15-3, significant enhancement of the fluorescence intensity was observed (green line), as a result of the specific binding of aptamer to CA15-3, leading to the aptamer/CA15-3 complex formation, and keep the hpDNA away from GO. The G-rich sequence can fold into a quadruplex by K+, and then G-quadruplex interacts with NMM. Taking the above results together, the feasibility of the proposed aptasensor for CA15-3 detection by our design was confirmed.
3.3 Optimization of Reaction Conditions
To achieve optimal sensing performance, several reaction conditions such as the concentration of hpDNA, the concentration of GO, the CA15-3 incubated time and incubated temperature were optimized. While the F and F0 were the fluorescence intensities in the presence and absence of CA15-3, respectively. The fluorescence intensity and the value of F/F0 are selected to evaluate the effects of the reaction conditions on the sensing performance of the method. As shown in Fig. 3(a), Maximum F/F0 value is observed when the concentration of hpDNA was 250 nM, however, F/F0 value decreases obviously along with the further increasing of hpDNA concentration. Thus, the hpDNA concentration of 250 nM was confirmed as the optimized concentration.
(a) The effect of DNA Concentration on the fluorescence response of this system; (b) The effect of GO Concentration on the fluorescence response of this system; (c) The effect of CA15-3 incubation temperature on the fluorescence response of this system; (d) The effect of CA15-3 incubation time on the fluorescence response of this system.
At the same time, the concentration of GO is another important factor affecting fluorescence intensity. As depicted in Fig. 3(b), with the increase in the concentration of GO, the fluorescence intensity initially increased and then gradually decrease, and maximum F/F0 values is observed when the concentration of GO is 5 mg mL−1. Thus, 5 mg mL−1 GO are used in the subsequent experiments.
In addition, the CA15-3 incubation time and incubation temperature are another important reaction condition affecting fluorescence intensity for this sensor. Figure 3(c) shows that the incubation temperature could obviously affect the sensitivity; the F/F0 value reached a maximum when the incubate temperature was 37 ℃ and then decreased gradually. Therefore, 37 ℃was confirmed as the optimized incubate temperature. The fluorescence intensity and the value of F/F0 change is related to the CA15-3 incubation time. As shown in Fig. 3(d), maximum F/F0 value is observed when time is 30 min. Thus, 30 min of CA15-3 incubated time was selected for the rest of the experiments.
3.4 Sensitivity and Specificity
Under the optimal reaction conditions, the sensitivity of the sensor for detection of CA15-3 is investigated. Figure 4(a) shows the fluorescence emission spectra of the biosensor incubated in different concentrations of CA15-3. We found that the fluorescence dramatically enhanced with the increasing concentration of CA15-3 from 0 to 3000 U mL−1. It illustrates a highly concentration dependence of the sensor for detection of CA15-3. At the same time, Fig. 4(b) shows a good linear correlation between the fluorescence intensity and the concentration of CA15-3 in the range from 10 UmL−1 to 500 UmL−1. The calibration plot of the linear equation is given as y = 4.635x + 243.73 (R2 = 0.9848), where x is concentration of CA15-3 and y is the fluorescence intensity. Furthermore, the LOD calculated using S/N ratio of three was 10 UmL−1. In addition, for the specificity study, adding different control proteins was investigated, including thrombin (0.1 μg mL−1), lysozyme (0.1 μg mL−1), streptavidin (0.1 μg mL−1), BSA (0.1 μg mL−1). As shown in Fig. 5, in the presence of other control proteins (0.1 μg mL−1), the significant increase of fluorescence signal is observed in the presence of the CA15-3 (0.1 μg mL−1), indicating that this proposed strategy exhibited good specificity for CA15-3 detection.
(a) Fluorescence spectra of NMM towards different concentrations (0, 10, 50, 100, 250, 500, 1000, 2000 and 3000 U mL−1) of CA15-3 in PBS buffer; (b) The relationship between the fluorescence intensity of NMM and different concentrations of CA15-3 in PBS buffer. The concentrations of hpDNA, GO, K+ and NMM were 250 nM, 5 mg mL−1, 20 mM and 1 μM, respectively.
3.5 CA15-3 Assay in Real Samples
To further verify the potential applicability of this present strategy in biological sample, the detection of CA15-3 in biological sample by spiking CA15-3 to human serum diluted to 0.1% with buffer solution with the different concentration of CA15-3 were performed. As shown in Fig. 6, a significant increase in biological sample was observed, compared with no spiking biological sample. These results clearly demonstrate that this sensor can be a potential analytical method to detect CA15-3 in real samples sensitively. The recoveries for the various concentrations of spiked CA15-3 in human serum were in the range of 86.97–106.13%, with the relative standard deviations (RSDs) of 8.76%, 5.05%, 2.14% and 5.50% at 100, 250, 500 and 1000 U mL−1 of CA15-3, respectively. This indicated an acceptable precision and reproducibility of the present approach for detecting CA15-3 in real samples (n = 3) (Table 1).
4 Discussion
This study successfully developed a DNA AND logic gate based on graphene oxide (GO) absorbing single DNA and G-quadruplex interacting with N-methyl mesoporphyrin IX(NMM) for detecting breast cancer biomarker carbohydrate antigen 15-3 (CA15- 3). This system responded to CA15-3 linearly in the concentration ranging from 10 to 500U mL−1, and the LOD of 10 U mL−1, which was lower than the level of CA15-3 in healthy people blood. Under laboratory conditions, the method has the advantages of high sensitivity, good specificity, fewer steps and low cost. Furthermore, the system also applied to diluted healthy human serum and trustworthy for detection and measurement of CA15-3 is confirmed. But this method has some shortcomings, for instance, this method needs further experiments to lower the detection limit of CA15-3. It is expected that this system has great potential application of CA15-3 clinical diagnostics in vitro.
References
de Silva, A.P., McClenaghan, N.D.: Molecular-scale logic gates. Chem. (Weinheim an der Bergstrasse, Germany) 10, 574–586 (2004)
Huo, Y.-F., Zhu, L.-N., Li, X.-Y., Han, G.-M., Kong, D.-M.: Water soluble cationic porphyrin showing pH-dependent optical responses to G-quadruplexes: applications in pH- sensing and DNA logic gate. Sens. Actuator B-Chem. 237, 179–189 (2016)
Wenjing, L., Yifang, G., Yuan, J., Shaomin, S., Chenzhong, L., Chuan, D.: Carbon nano-dots as a fluorescent and colorimetric dual-readout probe for the detection of arginine and Cu2+ and its logic gate operation. Nanoscale 9(32), 11545–11552 (2017)
Jiankang, D., et al.: A target-induced logically reversible logic gate for intelligent and rapid detection of pathogenic bacterial genes. Chem. Commun. 54(25), 1–10 (2018). Cambridge, England
Yingying, Z., Luhui, W., Yafei, D.: A Label-free and Universal Platform for the Construction of Various Logic Circuits Based on Graphene Oxide and G-Quadruplex Structure. Anal. Sci. 35(2), 181–187 (2019)
de Silva, A.P., Uchiyama, S.: Molecular logic and computing. Nat. Nanotechnol. 2(2), 399–410 (2007)
Li, W., et al.: Graphene-based aptamer logic gates and their application to multiplex detection. ACS Nano. 6, 6659–6666 (2012)
Jing, Y., et al.: Fluorescent nanoparticle beacon for logic gate operation regulated by strand displacement. ACS Appl. Mater. Interfaces 5, 5393–5396 (2013)
Gellert, M., Lipsett, M.N., Davies, D.R.: Helix formation by guanylic acid. Proc. Nat. Acad. Sci. India A. 48, 2013–2018 (1962)
Jiande, G., Leszczynski, J.: Origin of Na+/K+ selectivity of the guanine tetraplexes in water: the theoretical rationale. J. Phys. Chem. A 106, 529–532 (2002)
Balaratnam, S., Basu, S.: Divalent cation-aided identification of physico-chemical properties of metal ions that stabilize RNA G-quadruplexes. Biopolymers 103, 376–386 (2015)
Li, R., Liu, Q., Jin, Y., Li, B.: G-triplex/hemin DNAzyme: an ideal signal generator for isothermal exponential amplification reaction-based biosensing platform. Anal. Chim. Acta 1079, 139–145 (2019)
Zhang, Y., Wang, L., Wang, Y., Dong, Y.: Label-free optical biosensor for target detection based on simulation-assisted catalyzed hairpin assembly. Comput. Biol. Chem. 78, 448–454 (2019)
Wei, Y., Wang, L., Zhang, Y., Dong, Y.: An enzyme- and label-free fluorescence aptasensor for detection of thrombin based on graphene oxide and G-quadruplex. Sensors 19, 3–11 (2019)
Khusbu, F.Y., Zhou, X., Chen, H., Ma, C., Wang, K.: Thioflavin T as a fluorescence probe for biosensing applications. Trac-Trends Anal. Chem. 109, 1–18 (2018)
Li, Y., Wang, Y., Zhang, B., He, Y., Wang, J., Wang, S.: A rapid fluorometric method for determination of aflatoxin B1 in plant-derived food by using a thioflavin T-based aptasensor. Microchim. Acta 186, 1–7 (2019)
Rao, C.N.R., Sood, A.K., Subrahmanyam, K.S., Govindaraj, A.: Graphene: the new two-dimensional nanomaterial. Angew. Chem.-Int. Edit. 48, 7752–7777 (2009)
Zhou, Q., et al.: Ultra- sensitive enzyme-free fluorescent detection of VEGF (165) based on target-triggered hybridization chain reaction amplification. RSC Adv. 8, 25955–25960 (2018)
Ning, Y., Hu, J., Wei, K., He, G., Wu, T., Lu, F.: Fluorometric determination of mercury (II) via a graphene oxide-based assay using exonuclease III-assisted signal amplification and thymidine-Hg (II)-thymidine interaction. Microchim. Acta. 186, 3–8 (2019)
Huang, Z., Luo, Z., Chen, J., Xu, Y., Duan, Y.: A facile, label-free, and universal biosensor platform based on target-induced graphene oxide constrained DNA dissociation coupling with improved strand displacement. ACS Sens. 3, 2423–2431 (2018)
Zhou, J., Meng, L., Ye, W., Wang, Q., Geng, S., Sun, C.: A sensitive detection assay based on signal amplification technology for Alzheimer’s disease’s early biomarker in exosome. Anal. Chim. Acta 1022, 124–130 (2018)
Lu, C.-H., Yang, H.-H., Zhu, C.-L., Chen, X., Chen, G.-N.: A graphene platform for sensing biomolecules. Angew. Chem.-Int. Edit. 48, 4785–4787 (2009)
Park, J.S., Goo, N.-I., Kim, D.-E.: Mechanism of DNA adsorption and desorption on graphene oxide. Langmuir 30, 12587–12595 (2014)
Lee, J., Kim, J., Kim, S., Min, D.-H.: Biosensors based on graphene oxide and its biomedical application. Adv. Drug Deliv. Rev. 105, 275–287 (2016)
Wang, Y., Wei, Z., Luo, X., Wan, Q., Qiu, R., Wang, S.: An ultrasensitive homogeneous aptasensor for carcinoembryonic antigen based on up conversion fluorescence resonance energy transfer. Talanta 195, 33–39 (2019)
Li, X., Ding, X., Fan, J.: Nicking endonuclease-assisted signal amplification of a split molecular aptamer beacon for biomolecule detection using graphene oxide as a sensing platform. Analyst 140, 7918–7925 (2015)
He, Y., Lin, Y., Tang, H., Pang, D.: A graphene oxide-based fluorescent aptasensor for the turn-on detection of epithelial tumor marker mucin 1. Nanoscale 4, 2054–2059 (2012)
Wang, H., Chen, H., Huang, Z., Li, T., Deng, A., Kong, J.: DNase I enzyme-aided fluorescence signal amplification based on graphene oxide-DNA aptamer interactions for colorectal cancer exosome detection. Talanta 184, 219–226 (2018)
Dolati, S., Ramezani, M., Nabavinia, M.S., Soheili, V., Abnous, K., Taghdisi, S.M.: Selection of specific aptamer against enrofloxacin and fabrication of graphene oxide based label-free fluorescent assay. Anal. Biochem. 549, 124–129 (2018)
Xu, J., et al.: A fluorescent aptasensor based on single oligonucleotide-mediated isothermal quadratic amplification and graphene oxide fluorescence quenching for ultrasensitive protein detection. Analyst 143, 3918–3925 (2018)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Hu, W., Wang, L., Wang, Y., Qian, M., Dong, Y. (2021). Label-Free Fluorescence Detection of Carbohydrate Antigen 15-3 via DNA AND Logic Gate Based on Graphene Oxide. In: Pan, L., Pang, S., Song, T., Gong, F. (eds) Bio-Inspired Computing: Theories and Applications. BIC-TA 2020. Communications in Computer and Information Science, vol 1363. Springer, Singapore. https://doi.org/10.1007/978-981-16-1354-8_32
Download citation
DOI: https://doi.org/10.1007/978-981-16-1354-8_32
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-1353-1
Online ISBN: 978-981-16-1354-8
eBook Packages: Computer ScienceComputer Science (R0)