Abstract
Neural circuit function is determined not only by anatomical connections but also by the strength and nature of the connections, that is functional or physiological connectivity. To elucidate functional connectivity, selective stimulation of presynaptic terminals of an identified neuronal population is crucial. However, in the central nervous system, intermingled input fibers make selective electrical stimulation impossible. With optogenetics, this becomes possible, and enables the comprehensive study of functional synaptic connections between an identified population of neurons and defined postsynaptic targets to determine the functional connectome. By stimulating convergent synaptic inputs impinging on individual postsynaptic neurons, low frequency and small amplitude synaptic connections can be detected. Further, the optogenetic approach enables the measurement of cotransmission and its relative strength. Recently, optogenetic methods have been more widely used to study synaptic connectivity and revealed novel synaptic connections and revised connectivity of known projections. In this chapter, I focus on functional synaptic connectivity in the striatum, the main input structure of the basal ganglia, involved in the motivated behavior, cognition, and motor control, and its disruption in a range of neuropsychiatric disorders.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Nucleus accumbens
- Spiny projection neurons
- Cholinergic interneurons
- GABA interneurons
- Channelrhodopsin
- Synaptic responses
1 Introduction
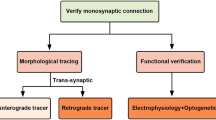
The mammalian brain contains billions of neurons and the majority of its functions are determined by their connectivity. Since the identification of synapses as the connections between neurons by Sherrington (Shepherd 1994), determining the wiring diagram of the brain has been a central interest of neuroscientists in order to understand brain function. Neuronal connectivity is evaluated in two ways: the physical wiring diagram, that is anatomical connectivity, and the strength and nature of the connection, that is physiological or functional connectivity. Anatomical connectivity is typically determined using retrograde and orthograde tracers, including viruses with reporter proteins. Synaptic connections can be identified by trans-synaptic tracers (e.g., pseudorabies virus), green fluorescent protein (GFP) reconstitution across synaptic partners (GRASP) (Feinberg et al. 2008), or observation with electron microscopy. However, the functional strength of a particular synaptic connection is not revealed by these methods. Direct measurement of synaptic responses in identified populations of neurons is crucial for determining functional (physiological) connectivity.
1.1 Functional Connectome
We introduced the concept of the functional connectome to describe comprehensive studies of functional (physiological) connectivity (Chuhma et al. 2011). The word “connectome” was used by Sporns in 2005 to describe the wiring diagram of the human brain (Sporns et al. 2005). The current definition of connectome is the comprehensive mapping of wiring in a brain (not limited to human) or a brain region (Lichtman and Sanes 2008; Seung 2009). In contrast to connectome, defined as the comprehensive study of anatomical connections of neurons, functional connectome can be defined as the comprehensive study of the physiological strength and nature (e.g., kinds of transmitters) of monosynaptic connections between two identified populations of neurons. It should be noted that the concept of functional connectivity used in the functional connectome approach is different from the functional connectivity used in brain imaging, which refers to temporally associated activity of two or more brain regions during a certain brain state (Friston et al. 1993). Functional connectivity in brain imaging is not necessarily relevant to direct synaptic connections among recorded regions, while the functional connectome is restricted to direct synaptic connections from an identified population of neurons to their defined postsynaptic target neurons.
1.2 Why the Optogenetic Approach is Suitable for Functional Connectome Studies
For a functional connectome study, selective and reliable stimulation of an identified population of neurons and recording from identified target neurons is crucial. Although identification of postsynaptic neurons is possible in vivo, intracellular recording from brain slices has advantages. In contrast to multiple ways of identifying postsynaptic neurons, selective stimulation of identified presynaptic neurons has been hard to achieve. Particularly in the basal ganglia, where cell architecture is not layered and input fibers from multiple structures are intermingled, population-selective stimulation with electrical stimulation is almost impossible. The advent of optogenetics has enabled selective activation of identified neuronal populations by genetically limited expression of photoactivatable proteins. Optogenetic activation enables convergent stimulation of inputs to a single neuron and detection of low-frequency small-amplitude responses (Eskenazi et al. 2020). The optogenetic approach also enables measurement of cotransmission. Immunohistochemistry or gene-expression studies may indicate the potential for cotransmission; however, the functional contribution of any cotransmitter is difficult to discern without physiological measurement.
For functional connectome studies, fast-on fast-off excitation current generation is necessary. Suitable photoactivatable proteins are nonselective cation channel channelrhodopsin (ChR)-2 and its mutants with faster kinetics and/or higher light sensitivity (Wietek and Prigge 2016; Yizhar et al. 2011). Because of its membrane-targeted nature, ChR2 can also be used as a genetically controlled axonal tracer. Although this makes coordinated anatomical studies feasible, this chapter focuses on excitation of genetically identified axon terminals to reveal functional connectivity. In the following section, I focus on the striatum (Str) and its intrinsic and extrinsic connectivity. The Str is one of the most suitable locations to apply the optogenetic approach to the functional connectome, because of intermingled inputs and existence of genetic drivers for conditional expression of ChR2. In this chapter, “functional connectivity” is used as physiologically confirmed direct synaptic connection from an identified cell population to a single identified neuron.
2 Functional Connectome of the Striatum with Optogenetics
2.1 Basics of Striatal Circuits
The rodent Str comprises the nucleus accumbens (NAc) and dorsal or neostriatum (dStr). The NAc corresponds to the ventral striatum in primates. Although the striatal region corresponding to the primate caudate or putamen is not clear in rodent, the ventromedial dStr is regarded as roughly homologous to the caudate and dorsolateral dStr roughly homologous to the putamen (Graybiel 2008). Both the NAc and dStr share similar cytoarchitectures: about 95% GABAergic spiny projection neurons (SPNs) and about 5% interneurons (Wilson 2004). SPNs are the sole population of output neurons and their major projection sites are the globus pallidus (GP)/ventral pallidum (VP) and ventral midbrain (Wilson 2004). Striatal interneurons comprise cholinergic and GABAergic interneurons. Cholinergic interneurons (ChIs) are a single population and makeup about 1% of striatal neurons, while GABAergic interneurons comprise several different types, namely fast-spiking interneurons (FSIs), low-threshold spike interneurons (LTSIs), and neurogliaform interneurons (Kreitzer 2009; Tepper et al. 2010). A subset of GABA interneurons expresses tyrosine hydroxylase (TH), although they do not release dopamine (DA) (Tepper et al. 2018). While interneurons make up small minority populations, they exert strong control of SPN excitability and modulate DA neuron afferents axo-axonally (Exley and Cragg 2008; Nelson et al. 2014; Oldenburg and Ding 2011; Tepper et al. 2004).
Both the dStr and NAc receive inputs from the cortex (Ctx), thalamus (Thal), and ventral midbrain (Yin and Knowlton 2006). The NAc is distinguished by inputs from limbic structures: the hippocampus (Hipp) and basolateral amygdala (BLA) (Belujon and Grace 2011; Pennartz et al. 2009). Those inputs show a topographic projection pattern. Ventral midbrain DA neurons in the medial ventral tegmental area (VTA) project to the NAc medial shell, the most ventromedial subregion of the Str, while more lateral DA neurons in the substantia nigra pars compacta (SNc) project to dorsolateral subregion of the Str (Haber et al. 2000; Ikemoto 2007). These connections modulate Ctx-basal ganglia-Thal loops that run in parallel from the ventral “limbic loop” to the dorsal “sensorimotor loop” (Pennartz et al. 2009; Yin and Knowlton 2006).
2.2 Connectivity Between Str Neurons and Extra Str Regions
Functional connectome studies have proceeded from examination of connectivity between principal neurons to more subregion-specific and cell-type-specific functional connectivity. Now the input–output connectivity in each Str neuron type has been examined.
2.2.1 Spiny Projection Neurons
SPNs are divided into two populations based on their projections to the output nuclei of the basal ganglia, directly (the direct pathway), or via the GP/VP (the indirect pathway). The output nuclei comprise the VTA/SN and entopeduncular nucleus, the rodent counterpart of the internal segment of the GP in primates, which in turn projects to the Thal (Gerfen and Surmeier 2011). Direct pathway SPNs (dSPNs) express DA D1 receptors and enkephalin, while indirect pathway SPNs (iSPNs) express DA D2 receptors and substance P (Gerfen and Surmeier 2011). These two populations of SPNs are almost completely segregated in the dStr and NAc core in adult rodents (Bertran-Gonzalez et al. 2008). In the NAc shell, 17–38% of SPNs express both D1 and D2 receptors (Bertran-Gonzalez et al. 2008; Gangarossa et al. 2013).
2.2.1.1 Direct Pathway Spiny Projection Neurons
dSPNs receive glutamatergic synaptic connection from the Ctx and Thal, both in the dStr and NAc (Choi et al. 2019; Ding et al. 2012; Ellender et al. 2013; Parker et al. 2016; Zhu et al. 2016). In the medial dStr, inputs from the anterior cingulate Ctx are stronger than those from parafascicular nucleus of the Thal, which rarely fire dSPNs even after blockade of GABAA receptors (Choi et al. 2019) (Fig. 27.1a). There are direct inputs to the Str from somatostatin (SOM)-positive and parvalbumin (PV)-positive GABA neurons in the primary (M1) and secondary (M2) motor areas of the Ctx (Melzer et al. 2017). dSPNs receive strong input from M1 PV and M2 SOM GABA neurons and weaker inputs from M1 SOM GABA neurons. M2 PV GABA neurons make only weak connections to Str GABA interneurons (Fig. 27.1a). dSPNs receive direct synaptic inputs from ventral midbrain DA neurons, though the connection was by glutamate and GABA cotransmission, not DA (Chuhma et al. 2018; Kim et al. 2015; Tritsch et al. 2012) (Fig. 27.1a). Although GABA cotransmission appears to be more widely observed in dStr subregions, glutamate cotransmission to SPNs is limited to the lateral dStr (Chuhma et al. 2018). The dStr and GP make reciprocal GABAergic connections (Kita 2007). Pallidal neurons make direct synaptic connections to both dSPNs and iSPNs, primarily from neuronal PAS domain protein (Npas)1-positive neurons, and inputs to iSPNs are consistently larger than those to dSPNs (Glajch et al. 2016) (Fig. 27.1a).
Optogenetically confirmed connections between striatal spiny projection neurons (SPNs) and extra-striatal neurons, in the dStr (a) and NAc (b). Direct pathway SPNs (dSPNs; left) and indirect pathway SPNs (iSPNs; right) are shown separately. Soma (circles) colors indicate the neurons’ transmitter. The color of axon terminals (small ovals) indicates the released transmitter from the terminal. Connection strength is indicated by the thickness of the axons (lines)
In the NAc, the Hipp and BLA make glutamatergic connections in addition to the Ctx and Thal (Belujon and Grace 2011; Pennartz et al. 2009; Sesack and Grace 2010). Optical stimulation of ventral Hipp (vHipp) generates monosynaptic glutamate currents in dSPN and iSPNs in the NAc medial shell, while most strong inputs are on FSIs (Scudder et al. 2018) (Fig. 27.1b). The BLA also makes glutamatergic inputs on NAc SPNs with the same strength in dSPNs and iSPNs (Zhu et al. 2016) (Fig. 27.1b). Among these glutamate inputs, the strongest are from the vHipp, and inputs from the BLA and PFC are the same strength but significantly weaker than those from the vHipp (Britt et al. 2012). Both glutamate and GABA cotransmission of DA neurons is observed in NAc SPNs, although the subtypes of SPNs were not specified (Chuhma et al. 2014; Mingote et al. 2015; Stuber et al. 2010; Tecuapetla et al. 2010; Tritsch et al. 2014) (Fig. 27.1b).
Retrograde tracer injections show that dSPNs send axons to both DA neurons in the SNc and GABA neurons in the SN pars reticulata (SNr) (Fujiyama et al. 2011; Gerfen 1985). Early studies of output to the ventral midbrain with transgenic expression of ChR2 in the dStr (Chuhma et al. 2011) or viral injection in the NAc (Xia et al. 2011) showed GABAA responses in only non-DA neurons, but not DA neurons. In subsequent studies using viral transfection, SPN GABAergic connections to DA neurons were seen in the VTA (Bocklisch et al. 2013) and SNc (Lerner et al. 2015); however, the GABAergic input to GABA neurons was much stronger (Fig. 27.1a). The dStr is comprised of two chemically distinct compartments; striosome with prominent μ-opioid receptor expression and matrix with choline acetyltransferase (ChAT) expression (Crittenden and Graybiel 2011). A recent study revealed that SPNs innervating DA neurons were in striosome (Mcgregor et al. 2019). NAc dSPNs make direct connection to VTA neurons, preferentially to non-DA neurons (Xia et al. 2011) (Fig. 27.1b).
Most dSPNs also send collaterals to the GP (Fujiyama et al. 2011), which comprise about one-third of Str inputs to the region (Kita 2007). In vivo activation of dSPN collaterals inhibits GP neuron firing (Cazorla et al. 2014). Although the inhibition was likely mediated by direct collateral connections activating GABAA receptors, direct synaptic connections have not been confirmed.
2.2.1.2 Indirect Pathway Spiny Projection Neurons
iSPNs receive inputs from the same populations as dSPNs, however, strengths differ in some inputs. Weaker connections to iSPNs are from M2 SOM and M1 PV GABA neurons (Melzer et al. 2017), the vHipp (Scudder et al. 2018), and glutamate cotransmission from DA neurons in the lateral dStr (Chuhma et al. 2018), while stronger connections to iSPNs are from Npas1-positive pallidal GABAergic neurons (Glajch et al. 2016) (Fig. 27.1a, b).
iSPNs project to the GP/VP. Selective activation of striatal inputs, with expression of ChR2 in SPNs, evoked GABAA response in GP neurons (Chuhma et al. 2011). More than 98% of GP neurons are GABAergic projection neurons. Pallidal neurons can be classified by expression of Npas1, PV, Lim homeobox 6 (Lhx6), and transcription factor Fox P2 (Abrahao and Lovinger 2018). We observed two types of electrophysiologically distinguishable populations; slower firing high-input impedance neurons and faster firing low-input impedance neurons (Chuhma et al. 2011). The former population shares the nature of Cooper–Stanford type A neurons (Cooper and Stanford 2000) and Lhx6/Npas1 positive neurons (Abrahao and Lovinger 2018; Mastro et al. 2014); the latter shares the nature of Cooper–Stanford type B and C neurons (Cooper and Stanford 2000) and parvalbumin (PV)-positive neurons (Abrahao and Lovinger 2018, Mastro et al. 2014). Str GABAA input is stronger in type B/C-like neurons and very weak in type A-like neurons (Chuhma et al. 2011). Although the target neuron type was not specified, NAc core iSPNs make GABAergic connections to VP neurons as well (Dobbs Lauren et al. 2016).
2.2.2 Cholinergic Interneurons
Cholinergic interneurons receive direct synaptic inputs from the Ctx, Thal, GP, and ventral midbrain (Guo et al. 2015). Although all studies report strong glutamatergic inputs from the Thal and relatively weak inputs from the Ctx to ChIs (Assous et al. 2019), the difference between these two inputs varies. This was likely due to differences in origins of inputs in the Ctx or Thal, recording protocols, recording locations, and short-term plasticity. A recent study showed that glutamate synaptic currents evoked by single-pulse photostimulation were not different between inputs from M1 and parafascicular nucleus of the Thal, while effects on ChI firing were different (Mamaligas et al. 2019) (Fig. 27.2). ChIs receive GABA inputs from the motor Ctx as well (Melzer et al. 2017). Inputs are primarily from M1 SOM GABA neurons, with weaker inputs from M1 PV and M2 SOM GABA neurons. ChIs receive direct synaptic connection from ventral midbrain DA neurons (Fig. 27.2a). In addition to fast glutamate and GABA cotransmission, ChIs show D2-mediated IPSCs, D1/5-mediated delayed EPSCs, and mGluR1-mediated delayed EPSCs (Chuhma et al. 2014, 2018; Straub et al. 2014) (Fig. 27.2). The DA neuron synaptic responses showed regionally different blends of transmitters, e.g., dominant fast glutamate responses in the NAc dorsal medial shell, strong D2 and GABA responses in the medial dStr, and weak D2 responses and strong D1/5 and mGluR1 responses in the lateral dStr (Chuhma et al. 2014, 2018). The VTA contains GABA neurons projecting to the NAc (Van Bockstaele and Pickel 1995). The VTA GABA neurons generate large amplitude responses in all recorded ChIs (Brown et al. 2013) (Fig. 27.2b), and modest responses in SPNs. Recently, direct glutamatergic inputs from the pedunculopontine nucleus (PPN) to ChIs were reported (Assous et al. 2019) (Fig. 27.2a).
2.2.3 GABAergic Interneurons
2.2.3.1 Fast-Spiking Interneurons
FSIs receive inputs from both the Ctx and Thal. Optogenetic stimulation of the Ctx and Thal evokes about the same strength of glutamate responses in FSIs (Sciamanna et al. 2015) (Fig. 27.2). GP neurons, likely to be Npas1-positive, make direct synaptic connections to FSIs (Glajch et al. 2016) (Fig. 27.2a). Although it is not as prominent as in Npas1-positive neurons, PV-positive GP neurons make connections to FSIs as well (Glajch et al. 2016; Saunders et al. 2016). Glutamate inputs from PPN make direct connections to FSIs, which are as strong as to ChIs (Assous et al. 2019) (Fig. 27.2a).
In the NAc, FSIs receive glutamatergic input from the vHipp that is substantially stronger than input to SPNs (Scudder et al. 2018) (Fig. 27.2b). VTA glutamate neurons, including DA-glutamate neurons, make direct synaptic connection to FSIs in the NAc medial shell (Chuhma et al. 2014, Zhang et al. 2016: 725–733) (Fig. 27.2b).
2.2.3.2 Low-Threshold Spike Interneurons
Although LTSIs receive strong glutamatergic inputs from the Ctx, they appear not to receive inputs from the parafascicular nucleus of the Thal, which sends projections to other types of Str neurons (Assous et al. 2017). PV-positive GP neurons make connections to LTSIs, but they are weaker than those to FSIs (Saunders et al. 2016). In the NAc, the vHipp makes weak inputs to LTSIs (Scudder et al. 2018) (Fig. 27.2).
2.2.3.3 Other GABA Interneurons
In addition to FSIs and LTSIs, several classes of GABA interneurons have been described: neurogliaform interneurons (NGFs), TH-positive interneurons (THINs), calretinin-expressing interneurons (CRs), fast-adapting interneurons (FAIs), and spontaneously active bursting interneurons (SABIs) (Tepper et al. 2018). NGFs received strong inputs from the Thal and moderate subthreshold inputs from the Ctx (Assous et al. 2017). THINs received strong glutamatergic inputs from the Ctx and Thal (Assous et al. 2017). However, input sources to CRs, FAIs, and SABIs have not been clearly identified.
2.3 Intra Striatum Connectivity
Str neurons make direct synaptic connections within the Str. Since Str neurons are GABAergic except for ChIs and ChIs exert inhibitory effects through muscarinic acetylcholine (ACh) receptors, the intrinsic connections are predominantly inhibitory. Possible excitatory transmission in local Str connections is limited to nicotinic ACh receptor-mediated responses and VGLUT3-mediated ChI glutamate cotransmission. Because of physical proximity, paired recordings are possible between identified Str neurons. However, paired recordings will miss weak or lower probability connections. Optogenetic stimulation of an identified neuronal population activates convergent inputs and is more likely to detect low probability synaptic connections.
2.3.1 Interneuron to Projection Neuron Connections
SPNs are under strong control of interneurons, and received direct synaptic inputs from GABAergic interneurons, besides SABIs (Assous and Tepper 2019; Tepper et al. 2018). Both dSPNs and iSPNs receive strong inputs from FSIs and LTSIs in the dStr (Straub et al. 2016; Szydlowski et al. 2013) and NAc medial shell (Scudder et al. 2018) (Fig. 27.3). Although the strength of inputs does not differ in dSPNs and iSPNs, inputs from FSIs are significantly stronger than those from LTSIs (Straub et al. 2016). NGFs also made direct inputs to SPNs in the dStr with very high incidence even in slices (Assous et al. 2017; English et al. 2012) (Fig. 27.3). ChIs make fast glutamate responses to SPNs through VGLUT3-mediated cotransmission (Higley et al. 2011) (Fig. 27.3).
2.3.2 Projection Neuron to Projection Neuron Connections
Most studies of collateral connections among SPNs have been done with simultaneous whole-cell recordings. An early study with transgenic expression of ChR2 in ~10% of SPNs in the dStr revealed a higher incidence of GABAergic connection among SPNs (Chuhma et al. 2011). In the NAc core, iSPN to dSPN collateral GABAergic connections are reliably elicited by ChR2 activation expressed in iSPNs (Dobbs Lauren et al. 2016) (Fig. 27.3).
2.3.3 Interneuron to Interneuron Connections
SPNs have been regarded to be the sole targets of FSI synaptic connections, and this was confirmed using optogenetics showing only negligible connections between FSIs and other interneurons (Straub et al. 2016, Szydlowski et al. 2013) (Fig. 27.3). Other GABAergic interneurons made connections to interneurons: LTSIs to ChIs (Straub et al. 2016), NGFs to ChIs (English et al. 2012), and THINs to LTSIs (Assous et al. 2017). ChIs excited NGFs and THINs through nicotinic ACh receptors (Assous et al. 2017; Assous and Tepper 2019; English et al. 2012) (Fig. 27.3).
3 Future Directions
Optogenetics enables selective stimulation of identified populations of neurons with intermingled inputs in the basal ganglia, making functional connectome studies possible. Functional connectome studies add information to anatomical connectome studies regarding both the strength of synaptic connections and the transmitters used. The optogenetic approach is also a powerful tool for studies of cotransmission, not only proving existence and function, but also showing relative strength of the cotransmission. The combination of genetic fluorescent markers in postsynaptic neurons and ChR2 expression in presynaptic identified neuronal populations allows the study of cell-type-specific connectivities, particularly connection to and from interneurons.
New optogenetic tools will enable refined functional connectome studies. ChR2 can be expressed with “boolean” logic by combining two types of recombinases (e.g., Cre-loxP and Flp-FRT). This system is termed INTRSECT, for “INTRonic Recombinase Sites Enabling Combinatorial Targeting” (Fenno et al. 2014). INTRSECT can be used to target specific subpopulations of neurons; e.g., those capable of cotransmission (Mingote et al. 2019; Poulin et al. 2018). Synaptic connectivity can be determined using optical recording. Postsynaptic recording can be done either genetic Ca2+ indicators (Farhi et al. 2019; Mollinedo-Gajate et al. 2019; Shen et al. 2020) or genetic voltage indicators (Bando et al. 2019; Mollinedo-Gajate et al. 2019; Shen et al. 2020; Yang and St-Pierre 2016). Genetically encoded indicators allow recording from identified populations of neurons without electrophysiological identification (Mollinedo-Gajate et al. 2019).
There are certain limitations in functional connectome studies with optogenetics. ChR2 expression depends on the strength of cell-specific promoters. If the promoter is not strong enough, expression of ChR2 may not be enough to activate presynaptic axons limiting detection of synaptic connections. Because of expression efficacy and limits in penetration of blue light through tissue (Aravanis et al. 2007; Yizhar et al. 2011), all synaptic inputs may not be activated with photostimulation. Light penetration issues can be minimized by using longer wavelength photostimulation, which penetrates deeper into tissue, with longer wavelength sensitive ChR2 variants (Yizhar et al. 2011). Another potential confound is in the interpretation of results of optogenetics experiments. Since ChR2 is activated by blue laser or high-power LED, it may activate terminals more synchronously than happens under normal physiological conditions in vivo (Kravitz and Bonci 2013: 169). Therefore, what is observed in optogenetic functional connectome studies may not be physiological; it should be interpreted as the maximum input strength of the synapses. Another potential problem is the distortion of membrane properties by expression of high levels of ectopic proteins (Zimmermann et al. 2008). ChR2 has very small single-channel conductance (Feldbauer et al. 2009; Wietek and Prigge 2016), and high expression is required for fast depolarization. A recent comparison between conditional heterozygous and homozygous expression of ChR2 in cochlea revealed only minor effects on intrinsic membrane properties (Meng et al. 2019), minimizing this concern.
Although the anatomical connectome is fundamental, anatomical information alone does not elucidate functional connectivity. It is essential to evaluate the relative strength and physiological nature of connections, for a full description of neural connectivity. Gaining information about connectivity of identified populations of neurons is crucial for understanding how neuronal wiring diagrams correlate with function. Functional connectome is one of the ways to explore how our brain works, with optogenetics shining the light to show the direction.
Abbreviations
- ACh:
-
Acetylcholine
- BLA:
-
Basolateral amygdala
- ChAT:
-
Choline acetyltransferase
- ChI:
-
Cholinergic interneuron
- ChR2:
-
Channelrhodopsin 2
- CR:
-
Calretinin-expressing interneurons
- Ctx:
-
Cortex
- DA:
-
Dopamine
- dSPN:
-
Direct-pathway spiny projection neuron
- dStr:
-
Dorsal striatum
- FAI:
-
Fast-adapting interneurons
- FSI:
-
Fast-spiking interneuron
- GP:
-
Globus pallidus
- Hipp:
-
Hippocampus
- iSPN:
-
Indirect-pathway spiny projection neuron
- LTSI:
-
Low-threshold spike interneuron
- M1:
-
Primary motor area of the cortex
- M2:
-
Secondary motor area of the cortex
- NAc:
-
Nucleus accumbens
- NGF:
-
Neurogliaform interneuron
- PFC:
-
Prefrontal cortex
- PPN:
-
Pedunculopontine nucleus
- PV:
-
Parvalbumin
- SABI:
-
Spontaneously active bursting interneurons
- SN:
-
Substantia nigra
- SNc:
-
Substantia nigra pars compacta
- SNr:
-
Substantia nigra pars reticulata
- SOM:
-
Somatostatin
- Str:
-
Striatum
- TH:
-
Tyrosine hydroxylase
- Thal:
-
Thalamus
- THIN:
-
Tyrosine hydroxylase-positive interneuron
- VGLUT:
-
Vesicular glutamate transporter
- vHipp:
-
Ventral hippocampus
- VP:
-
Ventral pallidum
- VTA:
-
Ventral tegmental area
References
Abrahao KP, Lovinger DM (2018) Classification of GABAergic neuron subtypes from the globus pallidus using wild-type and transgenic mice. J Physiol 596:4219–4235
Aravanis AM, Wang L-P, Zhang F et al (2007) An optical neural interface: in vivo control of rodent motor cortex with integrated fiberoptic and optogenetic technology. J Neural Eng 4:S143–S156
Assous M, Tepper JM (2019) Excitatory extrinsic afferents to striatal interneurons and interactions with striatal microcircuitry. Eur J Neurosci 49:593–603
Assous M, Kaminer J, Shah F et al (2017) Differential processing of thalamic information via distinct striatal interneuron circuits. Nat Commun 8:15860
Assous M, Dautan D, Tepper JM et al (2019) Pedunculopontine glutamatergic neurons provide a novel source of feedforward inhibition in the striatum by selectively targeting interneurons. J Neurosci 39:4727–4473
Bando Y, Grimm C, Cornejo VH et al (2019) Genetic voltage indicators. BMC Biol 17:71
Belujon P, Grace AA (2011) Hippocampus, amygdala, and stress: interacting systems that affect susceptibility to addiction. Ann N Y Acad Sci 1216:114–121
Bertran-Gonzalez J, Bosch C, Maroteaux M et al (2008) Opposing patterns of signaling activation in dopamine D1 and D2 receptor-expressing striatal neurons in response to cocaine and haloperidol. J Neurosci 28:5671–5685
Bocklisch C, Pascoli V, Wong JCY et al (2013) Cocaine disinhibits dopamine neurons by potentiation of GABA transmission in the ventral tegmental area. Science (New York, NY) 341:1521–1525
Britt JP, Benaliouad F, Mcdevitt RA et al (2012) Synaptic and behavioral profile of multiple glutamatergic inputs to the nucleus accumbens. Neuron 76:790–803
Brown MTC, Tan KR, O’connor EC et al (2013) Ventral tegmental area GABA projections pause accumbal cholinergic interneurons to enhance associative learning. Nature 492:452–456
Cazorla M, De Carvalho FD, Chohan MO et al (2014) Dopamine D2 receptors regulate the anatomical and functional balance of basal ganglia circuitry. Neuron 81:153–164
Choi K, Holly EN, Davatolhagh MF et al (2019) Integrated anatomical and physiological mapping of striatal afferent projections. Eur J Neurosci 49:623–636
Chuhma N, Tanaka KF, Hen R et al (2011) Functional connectome of the striatal medium spiny neuron. J Neurosci 31:1183–1192
Chuhma N, Mingote S, Moore H et al (2014) Dopamine neurons control striatal cholinergic neurons via regionally heterogeneous dopamine and glutamate signaling. Neuron 81:901–912
Chuhma N, Mingote S, Yetnikoff L et al (2018) Dopamine neuron glutamate cotransmission evokes a delayed excitation in lateral dorsal striatal cholinergic interneurons. Elife 7:e39786
Cooper AJ, Stanford IM (2000) Electrophysiological and morphological characteristics of three subtypes of rat globus pallidus neurone in vitro. J Physiol 527(Pt 2):291–304
Crittenden JR, Graybiel AM (2011) Basal ganglia disorders associated with imbalances in the striatal striosome and matrix compartments. Front Neuroanat 5:59
Ding JB, Oh W-J, Sabatini BL et al (2012) Semaphorin 3E-Plexin-D1 signaling controls pathway-specific synapse formation in the striatum. Nat Neurosci 15:215–223
Dobbs Lauren K, Kaplan Alanna R, Lemos Julia C et al (2016) Dopamine regulation of lateral inhibition between striatal neurons gates the stimulant actions of cocaine. Neuron 90:1100–1113
Ellender TJ, Harwood J, Kosillo P et al (2013) Heterogeneous properties of central lateral and parafascicular thalamic synapses in the striatum. J Physiol 591:257–272
English DF, Ibanez-Sandoval O, Stark E et al (2012) GABAergic circuits mediate the reinforcement-related signals of striatal cholinergic interneurons. Nat Neurosci 15:123–130
Eskenazi D, Chuhma N, Mingote S et al (2020) Functional connectome mapping. In: Wilson GS, Michael AC (eds) Compendium of in-vivo monitoring in real-time molecular neuroscience. World Scientific, Singapore
Exley R, Cragg SJ (2008) Presynaptic nicotinic receptors: a dynamic and diverse cholinergic filter of striatal dopamine neurotransmission. Br J Pharmacol 153:S283–S297
Farhi SL, Parot VJ, Grama A et al (2019) Wide-area all-optical neurophysiology in acute brain slices. J Neurosci 39:4889–4908
Feinberg EH, Vanhoven MK, Bendesky A et al (2008) GFP reconstitution across synaptic partners (GRASP) defines cell contacts and synapses in living nervous systems. Neuron 57:353–363
Feldbauer K, Zimmermann D, Pintschovius V et al (2009) Channelrhodopsin-2 is a leaky proton pump. Proc Natl Acad Sci U S A 106:12317–12322
Fenno LE, Mattis J, Ramakrishnan C et al (2014) Targeting cells with single vectors using multiple-feature Boolean logic. Nat Methods 11:763–772
Friston KJ, Frith CD, Liddle PF et al (1993) Functional connectivity: the principal-component analysis of large (PET) data sets. J Cereb Blood Flow Metab 13:5–14
Fujiyama F, Sohn J, Nakano T et al (2011) Exclusive and common targets of neostriatofugal projections of rat striosome neurons: a single neuron-tracing study using a viral vector. Eur J Neurosci 33:668–677
Gangarossa G, Espallergues J, De Kerchove D’e A et al (2013) Distribution and compartmental organization of GABAergic medium-sized spiny neurons in the mouse nucleus accumbens. Front Neural Circuit 7:22
Gerfen CR (1985) The neostriatal mosaic. I Compartmental organization of projections from the striatum to the substantia nigra in the rat. J Comp Neurol 236:454–476
Gerfen CR, Surmeier DJ (2011) Modulation of striatal projection systems by dopamine. Annu Rev Neurosci 34:441–466
Glajch KE, Kelver DA, Hegeman DJ et al (2016) Npas1+ pallidal neurons target striatal projection neurons. J Neurosci 36:5472–5488
Graybiel AM (2008) Habits, rituals, and the evaluative brain. Annu Rev Neurosci 31:359–387
Guo Q, Wang D, He X et al (2015) Whole-brain mapping of inputs to projection neurons and cholinergic interneurons in the dorsal striatum. PLoS One 10:e0123381
Haber SN, Fudge JL, Mcfarland NR (2000) Striatonigrostriatal pathways in primates form an ascending spiral from the shell to the dorsolateral striatum. J Neurosci 20:2369–2382
Higley MJ, Gittis AH, Oldenburg IA et al (2011) Cholinergic interneurons mediate fast VGluT3-dependent glutamatergic transmission in the striatum. PLoS One 6:e19155
Ikemoto S (2007) Dopamine reward circuitry: two projection systems from the ventral midbrain to the nucleus accumbens-olfactory tubercle complex. Brain Res Rev 56:27–78
Kim J-I, Ganesan S, Luo SX et al (2015) Aldehyde dehydrogenase 1a1 mediates a GABA synthesis pathway in midbrain dopaminergic neurons. Science (New York, NY) 350:102–106
Kita H (2007) Globus pallidus external segment. Prog Brain Res 160:111–133
Kravitz AV, Bonci A (2013) Optogenetics, physiology, and emotions. Front Behav Neurosci 7:169
Kreitzer AC (2009) Physiology and pharmacology of striatal neurons. Annu Rev Neurosci 32:127–147
Lerner TN, Shilyansky C, Davidson TJ et al (2015) Intact-brain analyses reveal distinct information carried by SNc dopamine subcircuits. Cell 162:635–647
Lichtman JW, Sanes JR (2008) Ome sweet ome: what can the genome tell us about the connectome? Curr Opin Neurobiol 18:346–353
Mamaligas AA, Barcomb K, Ford CP (2019) Cholinergic transmission at muscarinic synapses in the striatum is driven equally by cortical and thalamic inputs. Cell Rep 28:1003–1014.e3
Mastro KJ, Bouchard RS, Holt HAK et al (2014) Transgenic mouse lines subdivide external segment of the globus pallidus (GPe) neurons and reveal distinct GPe output pathways. J Neurosci 34:2087–2099
Mcgregor MM, Mckinsey GL, Girasole AE et al (2019) Functionally distinct connectivity of developmentally targeted striosome neurons. Cell Rep 29:1419–1428.e5
Melzer S, Gil M, Koser DE et al (2017) Distinct corticostriatal GABAergic neurons modulate striatal output neurons and motor activity. Cell Rep 19:1045–1055
Meng X, Murali S, Cheng Y-F et al (2019) Increasing the expression level of ChR2 enhances the optogenetic excitability of cochlear neurons. J Neurophysiol 122:1962–1974
Mingote S, Chuhma N, Kusnoor SV et al (2015) Functional connectome analysis of dopamine neuron glutamatergic connections in forebrain regions. J Neurosci 35:16259–16271
Mingote S, Amsellem A, Kempf A et al (2019) Dopamine-glutamate neuron projections to the nucleus accumbens medial shell and behavioral switching. Neurochem Int 129:104482
Mollinedo-Gajate I, Song C, Knöpfel T (2019) Genetically encoded fluorescent calcium and voltage indicators. In: Barrett JE, Page CP, Michel MC (eds) Concepts and principles of pharmacology: 100 years of the handbook of experimental pharmacology. Springer International Publishing, Cham
Nelson AB, Bussert TG, Kreitzer AC et al (2014) Striatal cholinergic neurotransmission requires VGLUT3. J Neurosci 34:8772–8777
Oldenburg IA, Ding JB (2011) Cholinergic modulation of synaptic integration and dendritic excitability in the striatum. Curr Opin Neurobiol 21:425–432
Parker PRL, Lalive AL, Kreitzer AC (2016) Pathway-specific remodeling of thalamostriatal synapses in parkinsonian mice. Neuron 89:734–740
Pennartz CMA, Berke JD, Graybiel AM et al (2009) Corticostriatal interactions during learning, memory processing, and decision making. J Neurosci 29:12831–12838
Poulin J-F, Caronia G, Hofer C et al (2018) Mapping projections of molecularly defined dopamine neuron subtypes using intersectional genetic approaches. Nat Neurosci 21:1260–1271
Qi J, Zhang S, Wang H-L et al (2016) VTA glutamatergic inputs to nucleus accumbens drive aversion by acting on GABAergic interneurons. Nat Neurosci 19:725–733
Saunders A, Huang KW, Sabatini BL (2016) Globus pallidus externus neurons expressing parvalbumin interconnect the subthalamic nucleus and striatal interneurons. PLoS One 11:e0149798
Sciamanna G, Ponterio G, Mandolesi G et al (2015) Optogenetic stimulation reveals distinct modulatory properties of thalamostriatal vs corticostriatal glutamatergic inputs to fast-spiking interneurons. Sci Rep 5:16742
Scudder SL, Baimel C, Macdonald EE et al (2018) Hippocampal-evoked feedforward inhibition in the nucleus accumbens. J Neurosci 38:9091–9104
Sesack SR, Grace AA (2010) Cortico-Basal Ganglia reward network: microcircuitry. Neuropsychopharmacology 35:27–47
Seung HS (2009) Reading the book of memory: sparse sampling versus dense mapping of connectomes. Neuron 62:17–29
Shen Y, Nasu Y, Shkolnikov I et al (2020) Engineering genetically encoded fluorescent indicators for imaging of neuronal activity: progress and prospects. Neurosci Res 152:3–14
Shepherd G (1994) Neurobiology. Oxford University Press, Oxford
Sporns O, Tononi G, Kötter R (2005) The human connectome: a structural description of the human brain. PLoS Comput Biol 1:e42
Straub C, Tritsch NX, Hagan NA et al (2014) Multiphasic modulation of cholinergic interneurons by nigrostriatal afferents. J Neurosci 34:8557–8569
Straub C, Saulnier JL, Bègue A et al (2016) Principles of synaptic organization of GABAergic interneurons in the striatum. Neuron 92:84–92
Stuber GD, Hnasko TS, Britt JP et al (2010) Dopaminergic terminals in the nucleus accumbens but not the dorsal striatum corelease glutamate. J Neurosci 30:8229–8233
Szydlowski SN, Pollak Dorocic I, Planert H et al (2013) Target selectivity of feedforward inhibition by striatal fast-spiking interneurons. J Neurosci 33:1678–1683
Tecuapetla F, Patel JC, Xenias H et al (2010) Glutamatergic signaling by mesolimbic dopamine neurons in the nucleus accumbens. J Neurosci 30:7105–7110
Tepper JM, Koós T, Wilson CJ (2004) GABAergic microcircuits in the neostriatum. Trends Neurosci 27:662–669
Tepper JM, Tecuapetla F, Koos T et al (2010) Heterogeneity and diversity of striatal GABAergic interneurons. Front Neuroanat 4:150
Tepper JM, Koós T, Ibanez-Sandoval O et al (2018) Heterogeneity and diversity of striatal GABAergic interneurons: update 2018. Front Neuroanat 12:91
Tritsch NX, Ding JB, Sabatini BL (2012) Dopaminergic neurons inhibit striatal output through non-canonical release of GABA. Nature 490:262–266
Tritsch NX, Oh WJ, Gu C et al (2014) Midbrain dopamine neurons sustain inhibitory transmission using plasma membrane uptake of GABA, not synthesis. Elife 3:e01936
Van Bockstaele EJ, Pickel VM (1995) GABA-containing neurons in the ventral tegmental area project to the nucleus accumbens in rat brain. Brain Res 682:215–221
Wietek J, Prigge M (2016) Enhancing channelrhodopsins: an overview. In: Kianianmomeni A (ed) Optogenetics. Humana Press, New York, NY
Wilson CJ (2004) The basal ganglia. In: Shepherd G (ed) The synaptic organization of the brain, 5th edn. Oxford University Press, Oxford
Xia Y, Driscoll JR, Wilbrecht L et al (2011) Nucleus accumbens medium spiny neurons target non-dopaminergic neurons in the ventral tegmental area. J Neurosci 31:7811–7816
Yang HH, St-Pierre F (2016) Genetically encoded voltage indicators: opportunities and challenges. J Neurosci 36:9977–9989
Yin HH, Knowlton BJ (2006) The role of the basal ganglia in habit formation. Nat Rev Neurosci 7:464–476
Yizhar O, Fenno LE, Davidson TJ et al (2011) Optogenetics in neural systems. Neuron 71:9–34
Zhu Y, Wienecke CFR, Nachtrab G et al (2016) A thalamic input to the nucleus accumbens mediates opiate dependence. Nature 530:219–222
Zimmermann D, Zhou A, Kiesel M et al (2008) Effects on capacitance by overexpression of membrane proteins. Biochem Biophys Res Commun 369:1022–1026
Acknowledgments
The author thanks Mihran Bakalian and Stephen Rayport for helpful comments and discussion. Writing of this chapter was supported by NIH grant MH117128 and DA038966.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Chuhma, N. (2021). Functional Connectome Analysis of the Striatum with Optogenetics. In: Yawo, H., Kandori, H., Koizumi, A., Kageyama, R. (eds) Optogenetics. Advances in Experimental Medicine and Biology, vol 1293. Springer, Singapore. https://doi.org/10.1007/978-981-15-8763-4_27
Download citation
DOI: https://doi.org/10.1007/978-981-15-8763-4_27
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-8762-7
Online ISBN: 978-981-15-8763-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)