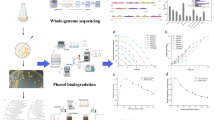
Abstract
Phenol is present as the basic structural compound of many synthetic organics. This pollutant generates from different sources such as herbicides, wood preservatives, petroleum industries, pharmaceuticals, etc. US Environmental Protection Agency marked it as the priority pollutant. The objective of this study is to isolate a potent bacterial strain from soil which is capable to remove phenol from wastewater. For this, soil sample was collected from the hospital area and enriched with 500 ppm phenol for 10 days. After serial dilution of the soil sample, colonies were developed in petri plate on nutrient agar medium. The isolated colonies were transferred to the slant and screening was done to select the most potent strain in liquid nutrient medium containing 500 ppm of phenol. The most efficient strain, P25, was able to reduce almost ~99.44% of phenol concentration in 24 h, 37 °C temperature, pH 6.8. The isolated strain was acclimatized in MSM (minimum salt medium) for 2 months and habituated to remove 700 mg/L phenol concentration. Simultaneously the strain was biochemically characterized and identified by 16S rDNA sequence analysis.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Many pollutants that are very hazardous for the nature are discharged from chemical, pharmaceuticals and oil refinery industries including many aliphatic and aromatic hydrocarbons (Taghreed and El-Naas 2014). Phenol is one of them, which has the capacity to damage the gastrointestinal tract, irritation of respiratory tracts, muscle tremors. Damage of liver, kidney, and nervous system are the adverse effects of phenol. Also phenol is very much hazardous to the aquatic ecosystems (Szczyrba et al. 2016). Therefore, it is mandatory to treat the phenol and phenolic wastes properly before disposal into the nature (Szczyrba et al. 2016; Kumar et al. 2005). The maximum permissible level of phenol in land water is 1 ppm according to the Central Pollution Control Board (CPCB) and IS:2490-1974 (Cheela et al. 2014; Lathasree et al. 2004; Saravanan et al. 2009). As per World Health Organization (WHO), the safety limit of phenol in drinking water should not exceed 1 mg/L (Bakhshi et al. 2011; Saravanan et al. 2008; Wang et al. 2010).
Several treatment methods are there to treat phenol such as adsorption, chlorination, ozonation as well as many physicochemical methods (Szczyrba et al. 2016; Tamer et al. 2010). Due to high cost effect of these methods and production of toxic intermediate compounds, involvement of biological processes is necessary in the treatment of phenol (Szczyrba et al. 2016). These biological treatments may be biosorption, biodegradation, bioaccumulation, etc., involving bacteria, algae, fungi, etc. Many such studies have been done previously as Mohanty and Jena (2017) did his experiment on biodegradation of phenol using Pseudomonus sp. NBM11 that was able to degrade up to 1000 ppm phenol completely in the temperature ranging between 30 and 32 ℃ and pH 6.8–7.2. Another study was done by Parvathy and Prabhakumari (2017) involving Pseudomonas aeruginosa, isolated from industrial soil to remove catechol.
The current study has been carried out aiming to isolate the most potent bacterial strain to degrade phenol and the identification of the strain.
2 Materials and Methods
2.1 Soil
Soil was collected from the local hospital area.
2.2 Reagents
Phenol-(analytical grade), 4-amino antipyrene, potassium ferricyanide, ammonium chloride and ammonium hydroxide solution. Our targeted concentration of phenol was prepared by mixing the properly weighed phenol to distilled water.
2.3 Bacterial Media
The nutrient agar medium with following composition was used for cultivation of bacteria.
Peptone—0.5%, beef extract—0.3%, agar—3.0%.
2.4 Isolation and Screening of Most Potent Bacterial Strain
The bacterial strain capable of removing phenol was isolated by soil enrichment and serial dilution plate count method. The soil was enriched with 500 ppm phenol for 10 days. After serial dilution and plate count isolated colonies were transferred in individual slant. A total of 29 isolated colonies were transferred to slant.
Screening was done to select most potent strain. Each of the isolated colonies was transferred to liquid medium and incubated 37 ℃ for 24 h in the presence of 500 ppm phenol. After fermentation, the fermented broth was centrifuged and the clear supernatant was used for spectrophotometric estimation of residual phenol content.
2.5 Estimation of Residual Phenol Content
Residual phenol concentration was measured in spectrophotometer at 510 nm wavelength followed by APHA method. Residual phenol content was calculated from the standard curve made with the known concentrations of phenol.
2.6 Morphological, Biochemical, and Phylogenetic Characterization of the Isolated Bacterial Strain
Morphological and biochemical characterization of the isolated bacterial strain was done as per Bergey’s Manual of Determinative Bacteriology (Holt et al. 1993). Phylogenetic assay was made by 16s rDNA method.
3 Results and Discussion
3.1 Isolation and Screening of Most Potent Bacterial Strain
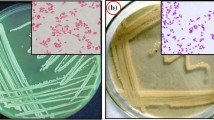
After serial dilution of the enriched soil sample and transfer of the diluted soil sample into nutrient agar medium, 29 isolated colonies were obtained. Colony characteristics of the isolated colonies are shown in Table 1.
After screening, it was found that the strain marked as P25 showed maximum phenol removal capability (~99.44%) (Table 1), so it was selected for further study.
3.2 Morphological, Biochemical, and Phylogenetic Characterization of the Isolated Bacterial Strain
Morphological characterization of the isolated bacterial strain P25 is shown in Tables 2 and 3. Gram characteristics and spore characteristics are shown in Figs. 1, 2, and 3. Physicochemical characteristics are shown in Table 4. The phylogenetic tree is shown in Fig. 4.
4 Conclusion
The isolated most potent bacterial strain was identified as Brevibacillus formosus strain NRRL NRS-863, which was able to reduce almost 99.44% of phenol. However, the mechanism of removal of phenol by the isolated strain i.e. whether the strain degrades it or accumulates it is not known till now. Our further study will reveal it.
References
Bakhshi Z, Najafpour G, Kariminezhad E et al (2011) Growth kinetic models for phenol biodegradation in a batch culture of Pseudomonusputida. Environ Technol 32(16):1835–1841
Cheela VRS, Kumar GS, Padma DV et al (2014) Biodegradation of Phenol using pure and mix culture Bacteria. e-J Sci and Technol (e-JST) 2(9):91–95. https://hypatia.lb.teiath.gr/bitstream/11400/6242/1/Cheela_35.pdf
Holt JG, Kreig NR, Sneath JT, Williams ST (1993) Bergey’s manual of determinative bacteriology, 9th edn. Wolters Kluwer
Kumar A, Kumar S, Kumar S (2005) Biochem Eng J 22:151–159
Lathasree S, Rao AN, Sivasankar B, Sadasivam V, Rengaraj K (2004) Heterogeneous photocatalytic mineralization of phenols in aqueous solutions. J Mol Catal A-Chem 223:101–105
Mohanty SS, Jena HM (2017) Biodegradation of phenol by free and immobilized cells of a novel Pseudomonus sp. NBM11. Braz J Chem Eng 34(01):75–84
Parvathy G, Prabhakumari C (2017) Isolation, characterization and optimization of catechol degrading Pseudomonas aeruginosa from Cashew Industrial soil. Int J Adv Biotechnol Res 7(1):13–23
Saravanan P, Pakshirajan K, Saha P (2009) Degradation of Phenol by Tio2-based heterogeneous photocatalysis in presence of sunlight. J Hydro-Environ Res 3(1):45–50
Saravanan P, Pakshirajan K, Saha P (2008) Growth kinetics of an indigenous mixed microbial consortium during phenol degradation in a batch reactor. Bioresour Technol 99:205–209
Szczyrba E, Szczotka A, Bartelmus G (2016) Modelling of aerobic biodegradation of phenol by Stenotrophomonas maltophilia KB2 strain. Proc ECO Pole 10(2)
Taghreed Al-Khalid T, El-Naas MH (2014) Biodegradation of phenol and 2,4 dichlorophenol: the role of glucose In biomass acclimatization. Int J Eng Res Technol 3(1):1579–1586
Tamer E, Amin MA, El Tayeb O, Mattiasson B, Guieysse B (2010) Kinetics and metabolic versatility of highly tolerant phenol degrading Alcaligenes strain TW1. J Hazard Mater 173(1–3):783–788
Wang L, Li Y, Yu P et al (2010) Biodegradation of phenol at high concentration by a novel fungal strain Paecilomyces variotii JH6. J Hazard Mater 183:366–371
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Maity, D., Kundu, P., (Nee Pramanik), S.A. (2021). Isolation of a Most Potent Bacterial Strain from Soil for Bioremediation of Phenol. In: Ramkrishna, D., Sengupta, S., Dey Bandyopadhyay, S., Ghosh, A. (eds) Advances in Bioprocess Engineering and Technology . Lecture Notes in Bioengineering. Springer, Singapore. https://doi.org/10.1007/978-981-15-7409-2_29
Download citation
DOI: https://doi.org/10.1007/978-981-15-7409-2_29
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-7408-5
Online ISBN: 978-981-15-7409-2
eBook Packages: EngineeringEngineering (R0)