Abstract
Rice is one of the most essential staple foods for most of the countries in the world. Yield of this important crop is severely hampered worldwide by an increasing number of microbial attacks. Among these, bacterial blight (BB) caused by Xanthomonas oryzae pv. oryzae is one of the main constraints of rice production. Different management strategies are promoted to alleviate this serious problem. Conventional breeding practices are mostly utilized to develop resistant cultivars in the different parts of the world as the approaches are cost effective and environment friendly. However, the fruitful results may not be achieved due to low yield and reduced effectiveness against that pathogen. On the other hand, pathogen races are gradually changing their organization to adopt the unfavorable environment. In this present situation, the research efforts have been shifted to find out resistance genes in plants or in others, against specific pathogens. Till date around 40 genes have been detected in rice. Only few genes have been cloned successfully and tested against this devastating pathogen. However, scientists are now concerning about the durability and long-term protection by developing new molecular tools which might also help in sustainable agriculture. In this chapter, we are trying to summarize different aspects of this disease and recent researches on rice plant against this pathogen.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Rice is one of the most accepted staple crops of almost two thirds of the world and specially for developing countries like India. However, cultivation of this popular crop has been constrained due to rigorous climate change. Rice farming is still challenging due to global population expansion, extensive urbanization, and constant shifting of cultivable land to industrial park. Greater modernization promotes greater abiotic stresses to the plants like rise in temperature, prolonged drought condition, and water stress, increases harmful radiations, upholds reactive oxygen species (ROS) generation, etc. Employment of excessive amount of fertilizers ultimately reduces soil fertility. Use of harmful chemical pesticides and fungicides contaminates groundwater and also encourages pathogens to improve themselves and to develop new races. Application of non-judicious farming technology and mistreatment of high throughput up to date agricultural technologies has ultimately gifted non-cultivable lands (Delteil et al. 2010). To fulfill the rising demand of food supply throughout the world, growers have to produce more than 40% extra rice by 2030 (Khush 2005; Delteil et al. 2010). However, it will be a herculean task today due to tremendous challenging factors. In spite of other abiotic factors, bacterial and fungal pathogens cause a serious threat on rice cultivation (Delteil et al. 2010). Near about 20–30% of rice production is regularly hampered per year globally by various rice pathogens (Chattopadhyay et al. 2017). In this connection, Xanthomonas oryzae pv. oryzae, a Gram-negative bacterium and the causal organism of bacterial leaf blight of rice, causes almost more than 50% crop loss in respect to others (Ishiyama 1922; Chattopadhyay et al. 2017). Though it is mainly a seed-borne disease, all the developmental stages are susceptible to the pathogen under encouraging environmental situations (Chattopadhyay et al. 2017).
This disease outbreak occurs throughout the world and causes serious crop loss and economic suppression mainly in different parts of Western Africa and Asia. Moreover, intense wind associated with rains may amplify the epidemic of bacterial blight (Chattopadhyay et al. 2017). It basically occurs in high-yielding rice varieties grown in the monsoon season, under profound nitrogen fertilization and especially in the irrigated and rain-feed ecosystems (Laha et al. 2017). Bacterial blight epidemics were reported from different parts of India (Laha et al. 2009, 2017; Yugander et al. 2014). More than 50% crop loss was reported in different West African countries due to this major destructive disease (Basso et al. 2011; Laha et al. 2017). Singh et al. (2013) recorded the highest economic loss in the following series Pusa Basmati-1 (45%) > Haryana Shankar Dhan-1 (31%) > HKR 47 (23%) in India. Besides that, severe economic loss was also noticed in various regions of Southern China, Japan, the Philippines, Pakistan, Nepal, and South and Central American countries (Adhikari and Mew 1991; Mew et al. 1993; Khan et al. 2000; Akhtar et al. 2003; Qi 2009; Corral et al. 2013).
It was the most challenging task to control this devastating pathogen over time. The most convenient, economic, useful, and sustainable method was to generate resistant cultivars against this pathogen (Chattopadhyay et al. 2017). However, selection of wild resistant varieties was tedious, and new varieties of pathogen races were also challenging over time. Researchers sometimes utilize some abiotic and biotic inducers also, which are able to boost up the innate immunity of the plants. However, it was not so effective in this pathosystem. Nowadays, more researches move on to developing transgenic rice varieties against this pathogen. More than 40 genes have been tested till date to generate elite variety of rice (Chattopadhyay et al. 2017; Laha et al. 2017). By virtue of the advancement of molecular biology and biotechnological methods, multi-genes are incorporated within a single rice variety through gene pyramiding and marker-assisted selection techniques (Chattopadhyay et al. 2017). Consequently, whole genome editing tools like clustered regularly interspaced short palindromic repeats/CRISPR-associated protein (CRISPR/Cas9) and transcription activator-like effector nucleus (TALEN) are also utilized (Chattopadhyay et al. 2017; Laha et al. 2017).
This chapter emphasizes on the detailed outlook of the important disease and also provides information about the various strategies including molecular tools for the improvement of bacterial blight resistance of rice.
2 Geographical Distribution and Past History of Bacterial Blight
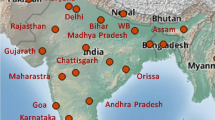
The devastating pathogen of bacterial blight disease of rice has been first identified and reported in the year of 1884 from Japan (Tagami and Mizukami 2008; Chattopadhyay et al. 2017). Later on, it was characterized properly and named as Xanthomonas oryzae pv. oryzae (Ishiyama) Swings et al. (Xoo) in the year 1922. Though almost all the rice-growing regions of the Caribbean island like Salvador, Mexico, Costa Rica, Honduras, and Panama were affected by this disease, it was first reported from the Mali region of West Africa (Buddenhagen et al. 1979). In South America, the affected regions include Colombia, Ecuador, Bolivia, and Venezuela (Lozano 1977; Sere et al. 2013). Louisiana and Texas from North America were most vulnerable for this pathogen (Jones et al. 1989). Furthermore, it was accounted from Australia and all the Asian countries (Ou 1985; Devadath 1992; Win et al. 2013). Srinivasan et al. (1959) reported the incidence of this disease from the state Maharashtra in India. However, its occurrence has not been established in Europe. The worldwide distribution of this disease is presented in Fig. 1.
World distribution of blast disease of rice (Laha et al. 2017)
3 Symptoms
It is a representative of typical vascular disease among others and has three discrete phases of symptoms. In the first phase of the disease, it shows pale-green to straw-colored striped water-soaked lesions with wavy margins on the tip of the isobilateral leaves which migrates longitudinally downward (Fig. 2). The leaf blight lesions may appear on both edges of the leaf and ultimately cover the whole leaf in the severe condition. Finally, it turns to whitish to grayish black due to the progression of saprophytic fungal growth. A small opaque yellowish drop containing bacterial eluents may be observed in the humid areas at the morning, which dries up and turns into small spherical yellowish beads (Chattopadhyay et al. 2017).
In the tropical countries, kresek or wilt phase is the harshest phase of this disease. In this situation, the leaves become yellow to grayish in color and roll completely with wilting symptoms. In the acute phases, the tillers shrivel, and the affected plants ultimately die (Laha et al. 2017).
On the other hand, in the Philippines, the symptoms appear in the cluster of leaves which turns into pale yellow or whitish, and finally the affected leaves wither and dry up (Laha et al. 2017).
4 Pathogen
Xanthomonas oryzae pv. oryzae (Ishiyama) Swings et al. is a rod-shaped, non-spore-forming, Gram-negative, and motile uniflagellate bacterium having a single polar flagellum. It belongs to Xanthomonadaceae (family), Xanthomonadales (order), Gammaproteobacteria (class), and Proteobacteria (phylum) in the domain Bacteria. It secretes xanthomonadin (a brominated aryl polyene pigment), a typical non-diffusible yellow pigment (Laha et al. 2017).
5 Disease Cycle and Epidemiology
According to the inception of different studies on this disease carried out by different scientific organizations like the International Rice Research Institute (IRRI), Manila, Philippines, All India Coordinated Rice Improvement Project (AICRIP), the main cause of the disease is seed infection (Laha et al. 2009). However, there are few such references which indicate that the disease may not occur due to planting of those seeds. Furthermore, it was observed that the infection may occur in the self-grown plants, from the pre-infected stubble straws and also from the infected wild rice varieties. The pathogen may also survive on some wild grasses like Panicum repens, Leersia hexandra, and Cyperus rotundus and contaminate irrigation water which may also act as a source of primary inoculum. Cloudy humid conditions along with moderate temperature like 28–30 °C and excess use of nitrogenous fertilizer aggravate rapid growth and spreading of the disease (Ezuka and Kaku 2000) (Fig. 3).
6 Brief Overview of Disease Management Strategies Employed till Date
Various control measures including host nutrition and physical, cultural, and chemical control are employed to restrict this devastating disease. However, the best possible outcome may not be achieved due to regular modification of the pathogenic races, though improvement of plant resistance by the introduction of new gene or by regular breeding technique may be the best solution to overcome this disease.
Use of cultural control as an essential part of integrated disease management may be useful to some extent to control this disease. Removing infected plant trashes, wild rice plants, and weeds like Leersia sp. and Cyperus sp. may be quite useful to confine pathogen progress. Restriction of unnecessary pruning of plant parts and field to field irrigation may be helpful to control this disease. Not only that, use of pathogen-free certified seeds and judicious use of manures mainly nitrogen may be beneficial (Ezuka and Kaku 2000; Laha et al. 2009, 2017).
After disease inception, the use of different chemicals may act as SOS to control this destructive disease, though chemicals which are utilized in the field may not be eco-friendly. Till date various chemicals like chloramphenicol, cellomate, Sankel, streptomycin, phenazine, etc. have been used in the field (Laha et al. 2017). However, satisfactory results may not be achieved and sometimes more than one chemical was required to control this pathogen and the remnants of which may cause serious health hazards (Srivastava 1972; Laha et al. 2017). Seed soaking with various chemicals like Agrimycin 100, streptomycin, Terramycin, chlortetracycline hydrochloride, Ceresan, etc. at variable proportions along with hot water treatment may diminish the pathogen inoculum from seeds (Laha et al. 2017). Srivastava (1972) reported that TF-130 and ATDA (2-amino-1,3,4-thiadiazole) were most efficient against this pathogen in Japan and adjoining countries. Agrimycin 100 and Fytolan (copper oxychloride) in a specific proportion (50:500) may be useful to check secondary infection (Singh et al. 1980). According to Laha et al. (2009), 12.5 kg/ha Klorocin application 10 days after transplanting may give useful results. The highest disease control by the application of chemicals was achieved by applying Streptocycline (200 mg/l), copper oxychloride (2.5 g/l), and 2,4-D ethyl ester (1.0 ml/l) in combination (Singh et al. 2012b; Laha et al. 2017).
Crop improvement by conventional breeding methods may also help to combat against this disease. In the year 1969, two bacterial blight rice varieties named IR 20 and IR 22 were developed from IRRI, mostly on the basis of the major resistance genes like Xa4 (Khush et al. 1989; Laha et al. 2017). However, those high-yielding varieties (HYVs) showed different degrees of susceptibility in different countries against this pathogen, which may be due to the development of more virulent new pathogenic races (Laha et al. 2017). Consequently, new resistance genes like Xa5 and Xa7 have been incorporated by plant researchers (Khush et al. 1989; Laha et al. 2017). Gradually other bacterial blight resistance-related new genes were discovered from wild rice varieties like Xa21, obtained from Oryza longistaminata (Khush et al. 1990; Laha et al. 2017). Till date more than 35 genes including Xa23, Xa27, Xa30(t)/Xa38, and Xa33 have been identified from various sources and customized for marker-assisted selection breeding (Sundaram et al. 2014; Kim et al. 2015; Laha et al. 2017). Nowadays, to achieve the highest durable resistance against this destructive pathogen, amalgamation of different genes into a single rice cultivar was taking place by using the process of marker-assisted selection breeding (Laha et al. 2017). By using this technique, various rice varieties were obtained which are listed in Table 1.
Researches on host resistance against this pathogen are not so wide in Africa as in Asia. According to available reports, NERICA 4, NERICA 8, and NERICA 14 varieties showed modest level of bacterial blight resistance (Banito et al. 2012). However, few accessions of O. glaberrima from Mali showed high degree of resistance against this pathogen race A3 (Djedatin et al. 2011; Laha et al. 2017).
7 Why Modify the Employment of Bacterial Blight Resistance Genes?
It is well-known that the effective genes present in the elite clones of different rice varieties are evenly distributed all across the vast geographic areas. Sometimes they interact with the existing pathogen community of local inhabitants. Very often, long-term contact with efficient pathogen may cause an epidemic outbreak, which is mainly due to the loss of durability of the resistance genes (Dossa et al. 2015). A detailed understanding of pathogen physiology, host metabolism, and disease epidemiology and consequently knowledge of interdisciplinary approaches are very much necessary to control the pathogen dynamics. Studies on suppressive environments and effective genes through effector biological programs may employ a new arena in the field of integrated disease management. The outcome of those experiments should be properly informed to the forerunner of the cultivation.
Near about 40 genes were identified for bacterial blight of rice mostly from the wild and cultivable varieties (Khan et al. 2014; Zhang et al. 2014; Dossa et al. 2015). In Asia, the resistance genes like Xa 4, Xa 5, Xa 13, and Xa 21 are mostly used for breeding purposes (Khan et al. 2014). Interestingly, the bacterial blight pathogen uses transcription activator-like (TAL) effectors to colonize the host system. In the large deployment of disease resistance against this pathogen and in contrast to the other pathosystems, Xa genes can be further classified into sub-categories (Boch et al. 2014; Dossa et al. 2015). The roles of different Xa genes are discussed in Table 2.
8 Genes and Techniques Employed to Rice Research
Various techniques have been utilized to produce stable transgenic rice plants in the market till date. However, transformation through Agrobacterium tumefaciens has been accepted as a routine technique in most of the laboratories (Toki et al. 2006; Delteil et al. 2010). Through this technique, efficiency of different rice varieties like japonica, indica, etc. has been improved sufficiently (Delteil et al. 2010). Later on, marker-assisted selection techniques were familiarized to the breeders for crop improvement (Ballini et al. 2009; Delteil et al. 2010). White and Yang (2009) describe the specific roles of major Xa resistance genes which confer the resistance against Xanthomonas. According to Peng et al. (2015), Xa21 was the most prime gene in bacterial blight resistance which is also connected to signaling pathways. Other genes such as OsBRR1 and OsWAK1 were shown to be helpful in coding for receptor-like proteins and provide basal resistance to rice plant (Delteil et al. 2010). Moreover, genes which are utilized for the making of transgenic rice are basically related to defense signaling and transcription factors including XB15, OsPLDB1, OsDR8, SPL18, SPL11, PACK1, OsGAP1, and OsRac1 (Delteil et al. 2010). Moreover, few PR genes out of 14 found in rice have been overexpressed (Van Loon et al. 2006; Delteil et al. 2010).
In a recent study on RNA-Seq showed that rice cultivar IRBB61 containing Xa7 when infected with Xanthomonas oryzae at high temperature stress, gives protection against the pathogen and was not dependent on salicylic acid pathway but related to abscisic acid (Cohen et al. 2017).
9 Future Goal and Conclusion
To find out the major constraints in rice production, it has been observed that bacterial blight has huge effect on yield loss. Though various approaches are available to control this disease, nowadays, the cultivation of resistant varieties is considered as the most economical tool. Many researchers till date working with the disease to understand the detailed molecular mechanism of pathogenesis. To protect rice from this devastating pathogen, it is an urgent requirement to identify and characterize a large number of defense genes. Marker-assisted breeding and various transgenic approaches can be taken as potential tools to protect rice against this disease. Identification, characterization, and functional analysis of resistant genes can open the downstream signaling pathway associated with this disease. That may be altered at desired steps to check the efficacy of the pathogen. If we think in other way, application of various biotic and abiotic elicitors can also be taken as a tool to induce innate immunity against this disease. Application of newer genome editing tool as CRISPR against the disease can also be accepted as future area of research. Many transcriptomic, proteomic, and metabolomic approaches can be taken as tools to draw the defense signaling network.
To sum up at this point, our major goal is to protect rice plant from this pathogen. To achieve this goal and to secure our food, further research works are required for better understanding of pathogenesis and identifying probable easy tool to combat against this disease.
References
Adhikari TB, Mew TW (1991) Effect of bacterial blight on growth and yield of rice. J Inst Agric Anim Sci 12:29–40
Akhtar MA, Zakria M, Abbassi FM, Masod MA (2003) Incidence of bacterial blight of rice in Pakistan during 2002. Pak J Bot 35:993–997
Ballini E, Vergne E, Tharreau D, Nottéghem JL, Morel JB (2009) ARCHIPELAGO: towards bridging the gap between molecular and genetic information in rice blast disease resistance. In: Wang GL, Valent B (eds) Advances in genetics, genomics and control of rice blast disease. Springer, Berlin, pp 417–425
Banito A, Kadai EA, Sere Y (2012) Screening of rice varieties for resistance to bacterial leaf blight. J Appl Biosci 53:3742–3748
Bao SY, Tan MP, Lin XH (2010) Genetic mapping of a bacterial blight resistance gene Xa14 in rice. Acta Agron Sin 36:422–427
Basso A, Onasanya A, Issaka S, Sido AY, Haougui A, Adam T, Sere Y, Saadou M (2011) Bacterial leaf blight of rice in Niger: pathological diversity of isolates collected on irrigated lands. J Appl Biosci 38:2551–2563
Bhasin H, Bhatia D, Raghuvanshi S, Lore JS, Sahi GK, Kaur B, Vikal Y, Singh K (2012) New PCR-based sequence-tagged site marker for bacterial blight resistance gene Xa38 of rice. Mol Breed 30:607–611
Blair MW, Garris AJ, Iyer AS, Chapman B, Kresovich S, McCouch SR (2003) High resolution genetic mapping and candidate gene identification at the xa5 locus for bacterial blight resistance in rice (Oryza sativa L.). Theor Appl Genet 107:62–73
Boch J, Bonas U, Lahaye T (2014) TAL effectors-pathogen strategies and plant resistance engineering. New Phytol 204:823–832
Buddenhagen IW, Vuong HH, Ba DD (1979) Bacterial blight found in Africa. Int Rice Res Newsl 4:11
Chattopadhyay A, Nagaich D, Lima JM, Verma A, Tiwari KK (2017) Molecular aspects of bacterial blight resistance in rice: recent advancement. In: Shamim M, Singh KN (eds) Biotic stress management in rice molecular approaches. Apple Academic Press Inc., Waretown, pp 17–45
Cheema KK, Grewal NK, Vikal Y, Sharma R, Lore JS, Das A, Bhatia D, Mahajan R, Gupta V, Bharaj TS, Singh K (2008) A novel bacterial blight resistance gene from Oryza nivara mapped to 38 kb region on chromosome 4L and transferred to Oryza sativa L. Gen Res (Camb) 90:397–407
Chen H, Wang S, Zhang Q (2002) New gene for bacterial blight resistance in rice located on chromosome 12 identified from Minghui 63, an elite restorer line. Phytopathology 92:750–754
Chen S, Lin XH, Xu CG, Zhang Q (2000) Improvement of bacterial blight resistance of Minghui 63, an elite restorer line of hybrid rice, by molecular marker-assisted selection. Crop Sci 40:239–244
Chen S, Liu X, Zeng L, Ouyang D, Yang J, Zhu X (2011) Genetic analysis and molecular mapping of a novel recessive gene xa34(t) for resistance against Xanthomonas oryzae pv. oryzae. Theor Appl Genet 7:1331–1338
Chu Z, Yuan M, Yao J, Ge X, Yuan B, Xu C, Li X, Fu B, Li Z, Bennetzen JL, Zhang Q, Wang S (2006) Promoter mutations of an essential gene for pollen development result in disease resistance in rice. Genes Dev 20:1250–1255
Cohen SP, Liu H, Argueso CT, Pereira A, Vera Cruz C, Verdier V et al (2017) RNA-Seq analysis reveals insight into enhanced rice Xa7-mediated bacterial blight resistance at high temperature. PLoS One 12(11):e0187625. https://doi.org/10.1371/journal.pone.0187625
Corral R, Leach JE, Verdier V, Vera Cruz CM (2013) Recovery plan for Xanthomonas oryzae causing bacterial blight and bacterial leaf streak of rice. Bulletin, NPDRS, 22
Delteil A, Zhang J, Lessard P, Morel J-B (2010) Potential candidate genes for improving rice disease resistance. Rice 3:56–71. https://doi.org/10.1007/s12284-009-9035-x
Devadath S (1992) Bacterial blight of paddy. In: Singh US, Mukhopadhaya AN, Kumar J, Chaube HS (eds) Plant diseases of international importance: diseases of cereals and pulses. Prentice Hall, Englewood Clifffs, pp 158–185
Djedatin G, Ndjiondjop MN, Mathieu T, Vera Cruz CM, Sanni A, Ghesquière A, Verdier V (2011) Evaluation of African cultivated rice Oryza glaberrima for resistance to bacterial blight. Plant Dis 95:441–447
Dossa GS, Sparks A, Cruz CV, Oliva R (2015) Decision tools for bacterial blight resistance gene deployment in rice-based agricultural ecosystems. Front Plant Sci 6:305. https://doi.org/10.3389/fpls.2015.00305
Ezuka A, Kaku H (2000) A historical review of bacterial blight of rice. Bulletin of the National Institute of Agrobiological Resources (Japan), No. 15 (March), 207 pp
Gao DY, Liu AM, Zhou YH, Cheng YJ, Xiang YH, Sun LH, Zhai WX (2005) Molecular mapping of a bacterial blight resistance gene Xa-25 in rice. Acta Genet Sin 32:183–188
Gao DY, Xu ZG, Chen ZY, Sun LH, Sun QM, Lu F, Hu BS, Liu YF, Tang LH (2001) Identification of a new gene for resistance to bacterial blight in a somaclonal mutant HX-3 (indica). Rice Genet Newslett 18:66
Gnanamanickam SS, Brindha PV, Narayanan NN, Vasudevan P (1999) Kavitha, S. An overview of bacterial blight disease of rice and strategies for its management. Curr Sci 77:1435–1443
Gonzalez C, Szurek B, Manceau C, Mathieu T, Sere Y, Verdier V (2007) Molecular and pathotypic characterization of new Xanthomonas oryzae strains from West Africa. Mol Plant Microbe Interact 20:534–546
Goto T, Matsumoto T, Furuya N, Tsuchiya K, Yoshimura A (2009) Mapping of bacterial blight resistance gene Xa11 on rice chromosome 3. Jpn Agric Res Q 43:221–225
Gu K, Sangha JS, Li Y, Yin Z (2008) High-resolution genetic mapping of bacterial blight resistance gene Xa10. Theor Appl Genet 116:155–163
Gu K, Tian D, Yang F, Wu L, Sreekala C, Wang D, Wang GL, Yin Z (2004) High-resolution genetic mapping of Xa27(t), a new bacterial blight resistance gene in rice, Oryza sativa L. Theor Appl Genet 108:800–807
Guo S, Zhang D, Lin X (2010) Identification and mapping of a novel bacterial blight resistance gene Xa35(t) originated from Oryza minuta. Sci Agric Sin 43:2611–2618
Guvvala LD, Koradi P, Shenoy V, Marella LS (2013) Making an Indian traditional rice variety Mahsuri, bacterial blight resistant using marker-assisted selection. J Crop Sci Biotechnol 6:111–121
He Q, Li D, Zhu Y, Tan M, Zhang D, Lin X (2006) Fine mapping of Xa2, a bacterial blight resistance gene in rice. Mol Breed 17:1–6
Hur YJ, Jeung JU, Kim YS, Park HS, Cho JH, Lee JY, Sohn YB, Song YC, Park DS, Lee CW, Sohn JG, Nam MH, Le JH (2013) Functional markers for bacterial blight resistance gene Xa3 in rice. Mol Breed 31:981–985
Ishiyama S (1922) Studies on bacterial blight of rice. Rep Agric Exp Stat Tokyo 45:233–261
Iyer AS, McCouch SR (2004) The rice bacterial blight resistance gene xa5 encodes a novel form of disease resistance. Mol Plant Microbe Interact 17:1348–1354
Jin X, Wang C, Yang Q, Jiang Q, Fan Y, Liu G, Zhao K (2007) Breeding of near-isogenic line CBB30 and molecular mapping of Xa30(t), a new resistance gene to bacterial blight in rice. Sci Agric Sin 40:1094–1100
Jones RK, Barnes LW, Gonzalez CF, Leach JE, Alvarez AM, Benedict AA (1989) Identification of low virulence strains of Xanthomonas campestris pv. oryzae from rice in the United States. Phytopathology 79:984–990
Khan MAI, Bhuiyan MR, Hossain MS, Sen PP, Ara A, Siddique MA, Ali MA (2014) Neck blast disease influences grain yield and quality traits of aromatic rice. C R Biol 337:635–641
Khan TZ, Gill MA, Khan MG (2000) Screening of rice varieties/lines for resistance to bacterial leaf blight. Pak J Phytopathol 12:71–72
Khush GS (2005) What it will take to feed 5.0 billion rice consumers in 2030. Plant Mol Biol 59:1–6
Khush GS, Angeles ER (1999) A new gene for resistance to race 6 of bacterial blight in rice Oryza sativa. Rice Genet Newslett 16:92–93
Khush GS, Bacalangco E, Ogawa T (1990) A new gene for resistance to bacterial blight from O. longistaminata. Rice Genet Newslett 7:121–122
Khush GS, Mackill DJ, Sidhu GS (1989) Breeding rice for resistance to bacterial leaf blight. In: IRRI (ed) Bacterial blight of rice. IRRI, Manila, pp 207–217
Kim SM, Suh JP, Qin Y, Noh TH, Reinke RF, Jena KK (2015) Identification and fine-mapping of a new resistance gene, Xa40, conferring resistance to bacterial blight races in rice (Oryza sativa L.). Theor Appl Genet 128:1933–1943
Kumar PN, Sujatha K, Laha GS, Rao KS, Mishra B, Viraktamath BC, Hari Y, Reddy CS, Balachandran SM, Ram T, Madhav MS, Rani NS, Neeraja CN, Reddy GA, Shaik H, Sundaram RM (2012) Identification and fine-mapping of Xa33, a novel gene for resistance to Xanthomonas oryzae pv. oryzae. Phytopathology 102:222–228
Laha GS, Reddy CS, Krishnaveni D, Sundaram RM, Srinivas Prasad M, Ram T, Muralidharan K, Viraktamath BC (2009) Bacterial blight of rice and its management. Technical Bulletin No. 41, Directorate of Rice Research (ICAR), Rajendranagar, Hyderabad, 37 pp
Laha GS, Singh R, Ladhalakshmi D, Sunder S, Prasad MS, Dagar CS, Babu VR (2017) Importance and management of rice diseases: a global perspective. In: Chauhan BS et al (eds) Rice production worldwide. Springer, pp 303–360. https://doi.org/10.1007/978-3-319-47516-5_13
Lee KS, Rasabandith S, Angeles ER, Khush GS (2003) Inheritance of resistance to bacterial blight in 21 cultivars of rice. Phytopathology 93:147–152
Lin XH, Zhang DP, Xie YF, Gao HP, Zhang Q (1996) Identifying and mapping a new gene for bacterial blight resistance in rice based on RFLP markers. Phytopathology 86:1156–1159
Liu Q, Yuan M, Zhou Y, Li X, Xiao J, Wang S (2011) A paralog of the MtN3/saliva family recessively confers race-specific resistance to Xanthomonas oryzae in rice. Plant Cell Environ 34:1958–1969
Lozano JC (1977) Identification of bacterial leaf blight in rice caused by Xanthomonas oryzae in America. Plant Dis Rep 61:644–648
Mew TW, Alvarez AM, Leach JE, Swings J (1993) Focus on bacterial blight of rice. Plant Dis 77:5–12
Miao L, Wang C, Zheng C, Che J, Gao Y, Wen Y, Li G, Zhao K (2010) Molecular mapping of a new gene for resistance to rice bacterial blight. China Agric Sci 43:3051–3058
Mir GN, Khush GS (1990) Genetics of resistance to bacterial blight in rice cultivar DV86. Crop Res 3:194–198
Noda T, Ohuchi A (1989) A new pathogenic race of Xanthomonas campestris pv. oryzae and its inheritance of differential rice variety Tetep to it. Ann Phytopathol Soc Japan 55:201–207
Noda T, Yamamoto T, Kaku H, Horino O (1996) Geographical distribution of pathogenic races of Xanthomonas oryzae pv. oryzae in Japan in 1991 and 1993. Annu Phytopathol Soc Japan 62:549–553
Ogawa T, Kaku H, Yamamoto T (1989) Resistance gene of rice cultivar, Asaminori to bacterial blight of rice. Jpn J Breed 39:196–197
Oryzabase (2011) Integrated rice science database. Available at: http://www.shigen.nig.ac.jp/rice/oryzabase
Ou SH (1985) Rice diseases, 2nd edn. Commonwealth Mycological Institute, Kew, Surrey, 380 pp
Peng H, Chen Z, Fang Z, Zhou J, Xia Z, Gao L, Chen L, Li L, Li T, Zhai W, Zhang W (2015) Rice Xa21 primed genes and pathways that are critical for combating bacterial blight infection. Sci Rep 5:12165. https://doi.org/10.1038/srep12165
Pinta W, Toojinda T, Thummabenjapone P, Sanitchon J (2013) Pyramiding of blast and bacterial leaf blight resistance genes into rice cultivar RD6 using marker assisted selection. Afr J Biotechnol 12:4432–4438
Porter BW, Chittoor JM, Yano M, Sasaki T, White FF (2003) Development and mapping of markers linked to the rice bacterial blight resistance gene Xa7. Crop Sci 43:1484–1492
Qi Z (2009) Genetics and improvement of bacterial blight resistance of hybrid rice in China. Rice Sci 16(2):83–92
Rajpurohit D, Kumar R, Kumar M, Paul P, Awasthi A, Basha OP, Puri A, Jhang T, Singh K, Dhaliwal HS (2011) Pyramiding of two bacterial blight resistance and a semi-dwarfing gene in type 3 basmati using marker-assisted selection. Euphytica 178:111–126
Rao Y, Li Y, Qian Q (2014) Recent progress on molecular breeding of rice in China. Plant Cell Rep 33:551–564
Sanchez AC, Ilag LL, Yang D, Brar DS, Ausubel F, Khush GS, Yano M, Sasaki T, Li Z, Huang N (1999) Genetic and physical mapping of xa13, a recessive bacterial blight resistance gene in rice. Theor Appl Genet 98:1022–1028
Sere Y, Fargette D, Abo ME, Wydra K, Bimerew M, Onasanya A, Akator SK (2013) Managing the major diseases of rice in Africa. In: Wopereis MCS, Johnson DE, Ahmadi N, Tollens E, Jalloh A (eds) Realizing Africa’s rice promise. Commonwealth Agricultural Bureau International, pp 213–228
Singh A, Singh VK, Singh SP, Pandian RTP, Ellur RK, Singh D, Bhowmick PK, Gopala Krishnan S, Nagarajan M, Vinod KK, Singh UD, Prabhu KV, Sharma TR, Mohapatra T, Singh AK (2012a) Molecular breeding for the development of multiple disease resistance in basmati rice. AoB Plants:pls029. https://doi.org/10.1093/aobpla/pls029
Singh R, Sunder S, Dodan DS (2012b) Sources of resistance, effectiveness of Xa/xa genes and evaluation of botanicals and non-conventional chemicals against bacterial blight of rice. Plant Dis Res 27:200–208
Singh R, Sunder S, Dodan DS (2013) Estimation of losses in grain yield due to bacterial blight and neck blast of rice in Haryana. Indian Phytopathol 66:249–251
Singh RA, Das B, Ahmed KM, Pal V (1980) Chemical control of bacterial leaf blight of rice. Trop Pest Manag 26:21–25
Singh S, Sidhu JS, Huang N, Vikal Y, Li Z, Brar DS, Dhaliwal HS, Khush GS (2001) Pyramiding three bacterial blight resistance genes (xa5, xa13 and Xa21) using marker-assisted selection into indica rice cultivar PR106. Theor Appl Genet 102:1011–1015
Song WY, Wang GL, Chen LL, Kim HS, Pi LY, Holsten T, Gardner J, Wang B, Zhai WX, Zhu LH, Fauquet C, Ronald P (1995) A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science 270:1804–1806
Srinivasan MC, Thirumalachar MJ, Patel MK (1959) Bacterial blight disease of rice. Curr Sci 28:469–470
Srivastava DN (1972) Bacterial blight of rice. Indian Phytopathol 26:1–16
Sun X, Cao Y, Yang Z, Xu C, Li X, Wang S, Zhang Q (2004) Xa26, a gene conferring resistance to Xanthomonas oryzae pv. oryzae in rice, encodes an LRR receptor kinase-like protein. Plant J 37:517–527
Sundaram RM, Chatterjee S, Oliva R, Laha GS, Cruz LJE, Sonti RV (2014) Update on bacterial blight of rice: fourth international conference on bacterial blight. Rice 7:12
Tagami Y, Mizukami T (2008) Historical review of the researches on bacterial leaf blight of rice caused by Xanthomonas. Special report on plant disease and insect pest forecasting service, vol 10. Ministry of Agriculture, Japan, pp 1–112
Tan GX, Ren X, Weng QM, Shi ZY, Zhu LL, He GC (2004) Mapping of a new resistance gene to bacterial blight in rice line introgressed from Oryza officinalis. Yi Chuan Xue Bao 31:724–729
Taura S, Ogawa T, Yoshimura A, Ikeda R, Iwata N (1992a) Identification of a recessive resistance gene to rice bacterial blight of mutant line XM6, Oryza sativa L. Jpn J Breed 42:7–13
Taura S, Ogawa T, Yoshimura A, Ikeda R, Omura T (1991) Identification of a recessive resistance gene in induced mutant line XM5 of rice to rice bacterial blight. Jpn J Breed 41:427–432
Taura S, Tabien RE, Khush GS, Yoshimura A, Omura T (1992b) Resistance gene of rice cultivar Taichung native 1 to Philippine races of bacterial blight pathogens. Jpn J Breed 42:195–201
Toki S, Hara N, Ono K, Onodera H, Tagiri A, Oka S et al (2006) Earl infection of scutellum tissue with Agrobacterium allows high speed transformation of rice. Plant J 47(6):969–976
van Loon LC, Rep M, Pieterse CMJ (2006) Significance of inducible defense-related proteins in infected plants. Annu Rev Phytopathol 44(1):135–162
Verdier V, Vera Cruz C, Leach JE (2012) Controlling rice bacterial blight in Africa: needs and prospects. J Biotechnol 159:320–328
Vikal Y, Chawla H, Rajiv S, Lore JS, Singh K (2014) Mapping of bacterial blight resistance gene xa8 in rice (Oryza sativa L.). Indian J Genet Plant Breed 74:589–595
Vikal Y, Das A, Patra B, Goel RK, Sidhu JS, Singh K (2007) Identification of new sources of bacterial blight (Xanthomonas oryzae pv. oryzae) resistance in wild Oryza species and O. glaberrima. Plant Genet Res 5:108–112
Wang C, Fan Y, Zheng C, Qin T, Zhang X, Zhao K (2014) High-resolution genetic mapping of rice bacterial blight resistance gene Xa23. Mol Gen Genomics 289:745–753
Wang C, Tan M, Xu X, Wen G, Zhang D, Lin X (2003) Localizing the bacterial blight resistance gene, Xa22(t), to a 100-kilobase bacterial artificial chromosome. Phytopathology 93:1258–1262
Wang C, Wen G, Lin X, Liu X, Zhang D (2009) Identification and fine mapping of the new bacterial blight resistance gene, Xa31(t), in rice. Eur J Plant Pathol 123:235–240
Wang GL, Song WY, Wu RL, Sideris S, Ronald PC (1996) The cloned gene, Xa27, confers resistance to multiple Xanthomonas oryzae pv. oryzae isolates in transgenic plants. Mol Plant Microbe Interact 9:850–855
White FF, Yang B (2009) Host and pathogen factors controlling the rice—Xanthomonas oryzae interaction. Plant Physiol 150(4):1677–1686
Win KM, Korinsak S, Sirithunya P, Lanceras-Siangliw J, Jamboonsri W, Da T, Patarapuwadol S, Toojinda T (2013) Marker assisted introgression of multiple genes for bacterial blight resistance into aromatic Myanmar rice MK-75. Field Crop Res 154:164–171
Wu X, Li X, Zu C, Wang S (2008) Fine genetic mapping of xa24, a recessive gene for resistance against Xanthomonas oryzae pv. oryzae in rice. Theor Appl Genet 118:185–191
Xiang Y, Cao Y, Xu C, Li X, Wang S (2006) Xa3, conferring resistance for rice bacterial blight and encoding a receptor kinase-like protein, is the same as Xa26. Theor Appl Genet 113:1347–1355
Yang Z, Sun X, Wang S, Zhang Q (2003) Genetic and physical mapping of a new gene for bacterial blight resistance in rice. Theor Appl Genet 106:1467–1472
Yoshimura A, Mew T, Khush G, Omura T (1983) Inheritance of resistance to bacterial blight in rice cultivar Cas 209. Phytopathology 73:1409–1412
Yoshimura S, Yamanouchi U, Katayose Y, Toki S, Wang ZX, Kono I, Kurata N, Yano M, Iwata N, Sasaki T (1998) Expression of Xa1, a bacterial blight-resistance gene in rice, is induced by bacterial inoculation. Proc Natl Acad Sci U S A 95:1663–1668
Yoshimura S, Yoshimura A, Iwata N, McCouch SR, Abenes ML, Baraoidan MR, Mew TW, Neson RJ (1995) Tagging and combining bacterial blight resistance genes in rice using RAPD and RFLP markers. Mol Breed 1:375–378
Yuan BS, Pu TM, Hua LX (2010) Genetic mapping of a bacterial blight resistance gene Xa14 in rice. Acta Agron Sin 36:422–427
Yugander A, Sundaram RM, Ladhalakshmi D, Shaik H, Sheshu Madhav M, Srinivas Prasad M, Viraktamath BC, Laha GS (2014) Pathogenic and genetic profile of Xanthomonas oryzae pv. oryzae isolates from Andhra Pradesh. Indian J Plant Prot 42:149–155
Zhang F, Zhuo DL, Zhang F, Huang LY, Wang WS, Xu JL et al (2014) Xa39, a novel dominant gene conferring broad-spectrum resistance to Xanthomonas oryzae pv. oryzae in rice. Plant Pathol. https://doi.org/10.1111/ppa.12283
Zhang F, Zhuo DL, Zhang F, Huang LY, Wang WS, Xu JL, Cruz CV, Li ZK, Zhou YL (2015) Xa39, a novel dominant gene conferring broad-spectrum resistance to Xanthomonas oryzae pv. oryzae in rice. Plant Pathol 64:568–575
Zhang J, Li X, Jiang G, Xuz Y, He Y (2006) Pyramiding of Xa7 and Xa21 for the improvement of disease resistance to bacterial blight in hybrid rice. Plant Breed 125:600–605
Zheng CK, Wang CL, Yu YJ, Zhao KJ (2009) Identification and molecular mapping of Xa32(t), a novel resistance gene for bacterial blight (Xanthomonas oryzae pv. oryzae) in rice. Acta Agron Sin 35:1173–1180
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Chakraborty, N., Sarkar, A., Acharya, K. (2020). Transgenic Rice Live Against Bacterial Blight. In: Roychoudhury, A. (eds) Rice Research for Quality Improvement: Genomics and Genetic Engineering. Springer, Singapore. https://doi.org/10.1007/978-981-15-5337-0_3
Download citation
DOI: https://doi.org/10.1007/978-981-15-5337-0_3
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-5336-3
Online ISBN: 978-981-15-5337-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)