Abstract
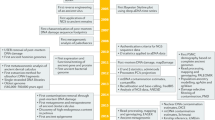
The introduction of next generation sequencing into archaeogenetics transformed the field into a powerhouse of the genetic study of human history. Although the processing of ancient DNA sequencing data follows the same basic principle for that of high-quality modern genomes, there are properties specific to ancient DNA molecules that need to be taken into account during data processing and analysis. In this chapter, I summarize the up-to-date practice of key steps of ancient DNA sequence data, such as the removal of Illumina sequencing adapter, read mapping to the reference genome, tabulation of post-mortem chemical damages, estimation of contamination level, and genotype calling. For each step, I highlight the current practices and the underlying logic for them, as well as directions of future method development.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120
Bos KI, Schuenemann VJ, Golding GB et al (2011) A draft genome of Yersinia pestis from victims of the black death. Nature 478:506–510
Bos KI, Harkins KM, Herbig A et al (2014) Pre-Columbian mycobacterial genomes reveal seals as a source of New World human tuberculosis. Nature 514:494–497
Dabney J, Meyer M, Pääbo S (2013) Ancient DNA damage. Cold Spring Harb Perspect Biol 5:a012567
Daly KG, Maisano Delser P, Mullin VE et al (2018) Ancient goat genomes reveal mosaic domestication in the Fertile Crescent. Science 361:85–88
DePristo MA, Banks E, Poplin R et al (2011) A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 43:491–498
Feldman M, Harbeck M, Keller M et al (2016) A high-coverage Yersinia pestis genome from a sixth-century Justinianic Plague victim. Mol Biol Evol 33:2911–2923
Fu Q, Mittnik A, Johnson PLF et al (2013) A revised timescale for human evolution based on ancient mitochondrial genomes. Curr Biol 23:553–559
Furtwängler A, Reiter E, Neumann GU et al (2018) Ratio of mitochondrial to nuclear DNA affects contamination estimates in ancient DNA analysis. Sci Rep 8:14075
Gaunitz C, Fages A, Hanghøj K et al (2018) Ancient genomes revisit the ancestry of domestic and Przewalski’s horses. Science 360:111–114
Green RE, Krause J, Briggs AW et al (2010) A draft sequence of the Neandertal genome. Science 328:710–722
Hofmanová Z, Kreutzer S, Hellenthal G et al (2016) Early farmers from across Europe directly descended from Neolithic Aegeans. Proc Natl Acad Sci U S A 113:6886–6891
Jeong C, Balanovsky O, Lukianova E et al (2019) The genetic history of admixture across inner Eurasia. Nat Ecol Evol 3:966–976
Jónsson H, Ginolhac A, Schubert M et al (2013) mapDamage2.0: fast approximate Bayesian estimates of ancient DNA damage parameters. Bioinformatics 29:1682–1684
Keller A, Graefen A, Ball M et al (2012) New insights into the Tyrolean Iceman’s origin and phenotype as inferred by whole-genome sequencing. Nat Commun 3:698
Kircher M, Sawyer S, Meyer M et al (2012) Double indexing overcomes inaccuracies in multiplex sequencing on the Illumina platform. Nucleic Acids Res 40:e3–e3
Korneliussen TS, Albrechtsen A, Nielsen R et al (2014) ANGSD: analysis of next generation sequencing data. BMC Bioinf 15:356
Kousathanas A, Leuenberger C, Link V et al (2017) Inferring heterozygosity from ancient and low coverage genomes. Genetics 205:317–332
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760
Link V, Kousathanas A, Veeramah K et al (2017) ATLAS: analysis tools for low-depth and ancient samples. bioRxiv 105346
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17:10–12
Meyer M, Kircher M, Gansauge MT et al (2012) A high-coverage genome sequence from an archaic Denisovan individual. Science 338:222–226
Meyer M, Arsuaga JL, de Filippo C et al (2016) Nuclear DNA sequences from the Middle Pleistocene Sima de los Huesos hominins. Nature 531:504–507
Miller W, Drautz DI, Ratan A et al (2008) Sequencing the nuclear genome of the extinct woolly mammoth. Nature 456:387–390
Orlando L, Ginolhac A, Zhang G et al (2013) Recalibrating Equus evolution using the genome sequence of an early Middle Pleistocene horse. Nature 499:74–78
Ottoni C, Van Neer W, De Cupere B et al (2017) The palaeogenetics of cat dispersal in the ancient world. Nat Ecol Evol 1:0139
Peltzer A, Jäger G, Herbig A et al (2016) EAGER: efficient ancient genome reconstruction. Genome Biol 17:60
Prüfer K (2018) snpAD: an ancient DNA genotype caller. Bioinformatics 34:4165–4171
Raghavan M, Skoglund P, Graf KE et al (2014) Upper palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505:87–91
Rasmussen M, Li Y, Lindgreen S et al (2010) Ancient human genome sequence of an extinct Palaeo-Eskimo. Nature 463:757–762
Rasmussen M, Guo X, Wang Y et al (2011) An Aboriginal Australian genome reveals separate human dispersals into Asia. Science 334:94–98
Renaud G, Stenzel U, Kelso J (2014) leeHom: adaptor trimming and merging for Illumina sequencing reads. Nucleic Acids Res 42:e141–e141
Renaud G, Slon V, Duggan AT et al (2015) Schmutzi: estimation of contamination and endogenous mitochondrial consensus calling for ancient DNA. Genome Biol 16:224
Rohland N, Harney E, Mallick S et al (2015) Partial uracil–DNA–glycosylase treatment for screening of ancient DNA. Philos Trans R Soc B 370:20130624
Sawyer S, Krause J, Guschanski K et al (2012) Temporal patterns of nucleotide misincorporations and DNA fragmentation in ancient DNA. PLoS One 7:e34131
Schubert M, Lindgreen S, Orlando L et al (2016) AdapterRemoval v2: rapid adapter trimming, identification, and read merging. BMC Res Notes 9:88
Schuenemann VJ, Peltzer A, Welte B et al (2017) Ancient Egyptian mummy genomes suggest an increase of Sub-Saharan African ancestry in post-Roman periods. Nat Commun 8:15694
Skoglund P, Ersmark E, Palkopoulou E et al (2015) Ancient wolf genome reveals an early divergence of domestic dog ancestors and admixture into high-latitude breeds. Curr Biol 25:1515–1519
Smith CI, Chamberlain AT, Riley MS et al (2003) The thermal history of human fossils and the likelihood of successful DNA amplification. J Hum Evol 45:203–217
The, 1000 Genomes Project Consortium (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491:56–65
Verdugo MP, Mullin VE, Scheu A et al (2019) Ancient cattle genomics, origins, and rapid turnover in the Fertile Crescent. Science 365:173–176
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Singapore Pte Ltd.
About this entry
Cite this entry
Jeong, C. (2021). Ancient DNA Study. In: Shin, D.H., Bianucci, R. (eds) The Handbook of Mummy Studies. Springer, Singapore. https://doi.org/10.1007/978-981-15-3354-9_11
Download citation
DOI: https://doi.org/10.1007/978-981-15-3354-9_11
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-3353-2
Online ISBN: 978-981-15-3354-9
eBook Packages: HistoryReference Module Humanities and Social SciencesReference Module Humanities