Abstract
Purple photosynthetic bacteria are metabolically versatile, anoxygenic phototrophs that produce bacteriochlorophylls a or b and display a wide range of metabolic lifestyles, which are reflected in their diverse range of habitats. Under oxic conditions, energy is derived from aerobic respiration, and the synthesis of photosynthetic pigments, and pigment-binding proteins, is repressed; when conditions shift to anoxic, the ultrastructure of the cytoplasmic membrane of purple bacteria changes, invaginations forming towards the inside of the cell creating intracytoplasmic membranes (ICMs). The photosynthetic machinery for the light-dependent reactions is housed in these ICMs. Light-harvesting (LH) antenna complexes absorb light energy, which is then transferred through a network of pigment–protein complexes, eventually promoting charge separation in the reaction centre, ultimately resulting in the formation of a proton motive force, used to drive ATP synthesis. ATP is then used to reduce inorganic carbon into organic compounds in the Calvin-Benson-Bassham cycle. Purple bacteria have evolved highly intricate assemblies of pigment–protein complexes to efficiently carry out this process. The assembly and organisation of these complexes differ among different species. In this chapter, we will discuss the structure, function and organisation of the photosynthetic components and the mechanisms underlying the photosynthetic process.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Antenna
- Atomic force microscopy
- ATP synthase
- Bacteriochlorophyll
- Bacteriopheophytin
- Calvin-Benson-Bassham cycle
- Carotenoid
- Core complex
- Cytochrome
- Excitation
- Intracellular membrane
- Light-harvesting
- Photosynthesis
- Pigment
- Proton motive force
- Purple bacteria
- Quinone
- Reaction centre
1 General Introduction
Photosynthesis is a globally important, fundamental process by which plants, algae and some bacteria convert sunlight into chemical energy. The process relies on a set of highly-ordered, multicomponent assemblies of pigment–protein complexes, located within invaginated membranes; the organisation and dynamics of these complexes are vital for efficient photosynthesis. Although the overall objective of photosynthesis across different organisms is similar—the production of metabolites for the cell—the multiprotein assemblies and compartments that perform this differ among different organisms.
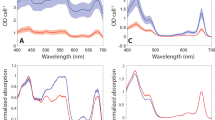
Unlike the chlorophyll-containing plants, algae and cyanobacteria that perform oxygenic photosynthesis, purple bacteria are anoxygenic phototrophs that use bacteriochlorophyll (BChl) a or b (Fig. 1), as well as carotenoid pigments that are ubiquitous in photosynthesis. The differences in structure between the bacteriochlorophyll and chlorophyll pigments result in varied absorption properties of phototrophic organisms (Fig. 1). Purple phototrophic bacteria are Proteobacteria, the most metabolically versatile of the seven phyla of photosynthetic prokaryotes: Chlorobi, Acidobacteria, Firmicutes, Cyanobacteria, Proteobacteria, Chloroflexi and Gemmatimonadetes (Gupta and Khadka 2016). Purple bacteria display a wide range of metabolic lifestyles with the capability to be photoautotrophic, photoheterotrophic, chemoautotrophic and mixotrophic, capable of fermentation and aerobic or anaerobic respiration, along with many different metabolic pathways for energy generation and carbon and sulfur metabolism (Imhoff et al. 2005). This is reflected in the diverse habitats in which they grow, including lakes, ponds, estuaries, marine environments, microbial mats, sewage and waste lagoons, as well as extreme niches including hot, cold, acidic, alkaline and hypersaline environments (Madigan 2003). Purple bacteria include both purple sulphur and purple non-sulphur bacteria, classically dependent on their ability to metabolise and assimilate sulphur compounds (Frigaard and Dahl 2008), although more contemporary studies have demonstrated that all purple bacteria are capable of sulphur metabolism to an extent (Hansen and Gemerden 1972). They can use a variety of reductants as electron donors, including H2S, other sulphur-containing compounds (such as cyst(e)ine and thiosulfate) or H2 (Truper and Fischer 1982; Brune 2004); some species can use Fe2+ iron as an electron donor (Ehrenreich and Widdel 1994). Table 1 shows the basic characteristics of various species of purple bacteria.
Structures of (bacterio)chlorophylls. Chemical structures of common (bacterio)chlorophylls. The carbon numbering system displayed on chlorophyll a is that approved by the International Union of Pure and Applied Chemistry (IUPAC). Structural differences between these pigments are highlighted in gray. Absorption spectra of these pigments in methanol are displayed (inset)
Anoxygenic photosynthesis is a simpler form of the process carried out by oxygenic organisms, which makes purple bacteria ideal model systems for dissecting the physiology, biochemistry and molecular biology of photosynthesis. Under oxic conditions, energy is derived from aerobic respiration, and the synthesis of photosynthetic pigments and pigment-binding proteins is repressed. As conditions change through microoxic to anoxic, the ultrastructure of the cytoplasmic membrane forms invaginations towards the inside of the cell called intracytoplasmic membranes (ICMs) (Hunter et al. 2008); these can take the form of vesicles, tubes or lamellae. Figure 2 shows a variety of different ICMs from different species.
Intracellular membranes of purple bacteria. (a) Schematic illustration of vesicular and lamellar ICM architectures (LaSarre et al. 2018). (b) Transmission electron micrograph of Rhodospirillum (Rsp.) rubrum cells, ICM vesicles are visible throughout the cytoplasm (Scheuring and Sturgis 2009). (c) TEM of photosynthetic membranes of Phs. molischianum cells containing stacked lamellar-type ICM continuous with the cytoplasmic membrane (Scheuring and Sturgis 2009)
Photosynthesis occurs in two stages: the light-dependent reactions, followed by the light-independent reactions. The light-dependent reactions occur in the ICM, where the photosynthetic apparatus is housed. The pigments within light-harvesting (LH) antenna complexes absorb photons of light as resonance energy. The derived excitation energy is then transferred through a network of pigments in the LH complexes towards the reaction centre (RC), where a ‘special pair’ of BChl pigments are housed. In the RC, charge separation occurs leading to the release of excited electrons, which are then passed down an electron transfer chain to RC-bound quinone electron carriers. This quinone is fully reduced to quinol after two electron transfers, at which point it is released from the RC and diffuses to cytochrome bc1 (cyt bc1). The quinol is then oxidised, releasing two protons into the periplasm of the cell; the electrons are passed to the soluble cytochrome c2 (cyt c2) and eventually back to the special pair in ‘cyclic electron flow’. This results in an electrochemical gradient between the cytoplasm and the periplasm, generating a proton motive force (pmf) utilised by ATP synthase (ATPase) in the production of ATP (Hu et al. 1998) (Fig. 3). In the light-independent stage, the energy produced in the light-dependent reactions is used to fix CO2 in the Calvin-Benison-Bassham cycle (see details below) (Tabita 2004; Blankenship 2014).
Schematic of the light-dependent reactions of purple bacterial photosynthesis. Light energy is first absorbed by B800 in LH2; this is then transferred to lower energy BChls until it reaches and excites the special pair BChls in the RC. An electron is then passed along a series of electron donors to QB, and after a second charge separation, QB is fully reduced to quinol; the electrons are then passed to cyt bc1 and cyt c2 which re-reduces the special pair. This creates an electrochemical gradient across the membrane, which is used by ATPase to create ATP. Note the exact locations of cyt bc1 and ATPase are not known. Yellow arrows, excitation transfer; red arrows, electron transfer; blue arrows, proton transfer; BChl bacteriochlorophyll; LH light harvesting; RC reaction centre; SP special pair; BPhe bacteriopheophytin; Q quinone; QH2 quinol; cyt cytochrome, ATPase ATP synthase
Aerobic anoxygenic phototrophs (AAPs) are a unique purple bacterial functional group that contain fully functional photosynthetic apparatus; however, this is assembled and operative under oxic conditions. Their photosynthetic machinery has various extensive modifications, including different peripheral antennas, and some organisms using zinc-chelated BChls in place of the more common magnesium-containing pigments (Wakao et al. 1996). AAPs have a much lower number of photosynthetic complexes per cell and a huge abundance of carotenoids (Rathgeber et al. 2004). Interestingly, although they are incapable of photoautrophy and rely on heterotrophy for 80% of their cellular metabolism, phototrophy can double organic carbon assimilatory efficiency (Kolber et al. 2001). BChl synthesis in these organisms is inhibited by sunlight (Aagaard and Sistrom 1972). Examples of these organisms can be seen in Table 1.
The majority of the genetic information needed to build the photosynthetic apparatus in purple bacteria is clustered into large (40–50 kbp) groups of genes called the photosynthesis gene cluster (PGC). The precise organisation of these genes within the cluster is highly variable. This cluster enables the precise control of expression levels and spatial proximity of the components (Alberti et al. 2004). The discovery of a phototrophic member of the Gemmatimonadetes bacteria, carrying a PGC of purple bacterial origin, demonstrates that the organisation of these genes into this superoperon permitted horizontal gene transfer between distant bacterial phyla (Zeng and Koblizek 2017).
2 Structural Components
2.1 Peripheral Antenna Complexes
The RC itself is a poor antenna, harvesting insufficient photons to support rapid growth via phototrophy. Purple bacteria have thus evolved specialised antenna complexes to increase the cross-sectional area available for light harvesting and maximise photon capture. The energy absorbed by the antenna complexes is then passed to the RC. Purple bacteria typically contain two types of antenna complexes: the ‘core’ light-harvesting complex 1 (LH1) intimately associates with the RC, encircling it to form the ‘core complex’ and RC–LH1 complex, found in all purple bacterial species. There are also peripheral antennas, known as LH2, which are separated from the RC by the LH1 ring. These antenna complexes are embedded in the ICM of purple bacteria, and bind the light-absorbing BChl and carotenoid pigments non-covalently.
2.1.1 Light-Harvesting Complex 2
The peripheral LH2 complexes funnel excitation energy to the RC via the core LH1 complexes (discussed in detail later). The LH2 complexes have been well studied across different species; all exist as octa- or nonameric quaternary structures. The crystal structures of LH2 from Rhodoblastus (Rbl.) acidophilus (Fig. 4a) (McDermott et al. 1995; Prince et al. 1997; Papiz et al. 2003) and Phaeospirillum (Phs.) molischianum (Fig. 4b) (Koepke et al. 1996) have been solved to 2.0 Å and 2.4 Å resolution, respectively. Both structures adopt the same modular principle: oligomers consisting of numerous pairs of α- and β-polypeptides, known as the αβ-heterodimer, forming double-ring structures with their associated pigments. The α- and β-polypeptides of the two species share 26% and 31% sequence identity, respectively.
Structures of different forms of light-harvesting antenna complexes in purple bacteria. (a) Structure of the LH2 complex from Rbl. acidophila 10,050. Cartoon representation of the nonameric B800-B850 LH2 complex viewed from above. The α- and β-polypeptides are represented in light-green and purple, respectively. The BChls α-B850, β-B850 and B800 are represented in red, green and blue, respectively, and the carotenoid is represented in orange (Papiz et al. 2003). (b) Crystal structure of the LH2 complex from Phs. molischianum. The octameric B800-B850 LH2 complex viewed from above with the N-termini pointing upwards. The α- and β-polypeptides are represented in orange and red, respectively. The BChl a molecules are in grey (Koepke et al. 1996). (c) Structure of the LH3 complex from Rbl. acidophilus 7050. Schematic representation of the nonameric LH3 complex from Rbl. acidophilus 7050 (McLuskey et al. 2001). (d) Electron densities obtained from Rps. palustris LH4 X-ray data, revealing an octameric structure (Hartigan et al. 2002). (e) AFM analysis of peripheral antenna complexes from Rps. palustris grown under low light; the white circles represent LH4 complexes (Scheuring et al. 2006). (f) Non-symmetrised (left) and symmetrised averages (right) of nonameric LH2 and octameric LH4 (LL-LH2) from Rps. palustris grown in low light (Scheuring et al. 2006)
The LH2 complex of Rbl. acidophilus is formed of a nonameric ring structure of 9 inner α- and 9 outer β-polypeptide chains (Fig. 4a), with ring diameters of 36 Å and 68 Å, respectively; each polypeptide crosses the membrane once, via an α-helical transmembrane domain (McDermott et al. 1995). The LH2 structure acts as a scaffold for the associated pigments, including carotenoids and two populations of BChls. One BChl population in the Rbl. acidophilus LH2 contains nine pairs of overlapping pigments held perpendicular to the plane of the membrane via histidine residues in the α- and β-polypeptides (Prince et al. 1997); the short distances between these paired BChls (~9 Å) provide strong excitonic coupling between the pigments, shifting the absorption maximum to ~850 nm; this population is known as B850 (after their corresponding absorption maxima). The complex also contains 9 B800 BChl molecules inserted between the β-polypeptides, parallel to the membrane plane. The central Mg atom of these BChls are coordinated by methionine residues in the α-polypeptide (Papiz et al. 2003); these pigments are not tightly coupled and so can be considered ‘monomeric’, thus contribute a band at ~800 nm, close to the absorption maximum of the pigment in solution. Finally, a carotenoid (rhodopin glucoside in the Rbl. acidophilus LH2) threads the space between the α- and β-pairs. Together, these pigments contribute to the structural and functional integrity of LH2 (Lang and Hunter 1994).
The overall structure of the B800–850 LH2 complex from Phs. molischianum is similar to that of Rbl. acidophilus (McDermott et al. 1995; Koepke et al. 1996), but with some key differences. The ring structure of this complex is made up of 8 αβ-pairs (Fig. 4b), thus possessing 3 BChls and 1 carotenoid fewer than that from Rbl. acidophilus, as well as reduced diameters of the inner (31 Å) and outer rings (62 Å) (Koepke et al. 1996). The 8 B800 molecules are coordinated by aspartate rather than methionine residues of α-polypeptides almost parallel to the membrane plane, with a distance of 2.45 Å. The acetyl carbonyl groups of BChl were also shown to interact with tryptophan residues in the α- and β-polypeptides; these residues were shown to be important in stabilising the BChl–LH1 interaction (Davis et al. 1997; Kehoe et al. 1998). The 8 carotenoids found in the complex are lycopene pigments stretched between the B800 and B850 molecules (Koepke et al. 1996). The crystal structure of Phs. molischianum LH2, along with mutational studies (Olsen et al. 1997), indicated that His residues also form a hydrogen bond with the carbonyl group of the coordinated BChl. The difference in oligomerisation seen between the two LH2 proteins was suggested to be due to the differences in the interaction angle between subunits: 40° for nonameric rings and 45° for octomeric rings (Janosi et al. 2006); these angles are believed to be determined by the surface interactions in the transmembrane regions (Janosi et al. 2006). The intermolecular forces underlying the ring shape of the LH2 complex were shown to influence the structural and functional integrity of the LH2 complex in Pararhodospirillum (Psp.) photometricum (previously Rhodospirillum) (Liu et al. 2011a, b).
2.1.2 Light-Harvesting Complexes 3 and 4
Under stress conditions, such as low light or low temperature, absorption bands of several purple bacteria vary due to the expression of atypical peripheral light-harvesting complexes, known as LH3 and LH4 (Evans et al. 1990; Mcluskey et al. 2001; Hartigan et al. 2002; Niedzwiedzki et al. 2011). These are spectral variants of LH2, functioning to increase the size of the photosynthetic unit and broaden the spectral range of solar energy captured by the cell.
Both the Rbl. acidophilus 7050 and 7750 strains contain LH3 complexes (Fig. 4c) (Cogdell et al. 1983; Angerhofer et al. 1986). In the 7050 strain, LH3 complexes always coexist with LH2 complexes; however, low-light conditions can also induce the expression of LH3 (Cogdell et al. 1983; Angerhofer et al. 1986). LH2 was almost completely replaced with LH3 complexes in the 7750 strain under low-light conditions (Gardiner et al. 1993). The LH3 complexes absorb light at 800–820 nm, a blue shift from the 800–850 nm absorption seen in LH2. The overall structure of the LH3 complex from Rbl. acidophilus 7050 (Mcluskey et al. 2001) is analogous to LH2 from Rbl. acidophilus 10,050 (McDermott et al. 1995), with the exception of reorientation of acetyl groups at the C3 position of B820 BChl in LH3 relative to the membrane plane; this causes a change in the hydrogen bonding patterns between the protein and the coupled BChl a molecules, resulting in the change in spectral properties of the apoprotein (Mcluskey et al. 2001).
Rhodopseudomonas (Rps.) palustris predominantly contains LH4 complexes in low-light conditions (Tharia et al. 1999). This complex, also referred to as the low-light LH2 B800 complex, is composed of eight αβ-polypeptide pairs; each pair contains four BChl molecules, distinct from the three BChls in LH2 complexes (Hartigan et al. 2002); electron microscopy (EM) and atomic force microscopy (AFM) studies have confirmed its octameric structure (Scheuring et al. 2006) (Fig. 4d, e).
The majority of purple phototrophic bacteria that use BChl a employ a peripheral antenna, but there are some exceptions; Rhodospirillium (Rsp.) rubrum and AAPs within the Eryth-Citro clde lack the genes that encode LH2 (and LH3 and LH4) (Munk et al. 2011; Zheng et al. 2011). To date, peripheral antennas have not been found in any BChl b-containing phototrophs; one possible explanation being that these antennas evolved in an ancestor using BChl a, after divergence of the two pigment pathways.
2.2 The Core Complex of Purple Bacterial Photosynthesis
Once light is harvested by LH2, the excitation energy is then passed to the BChl molecules housed in the LH1 antenna proteins and subsequently to the special pair BChls in the RC, enabling charge separation. Across all species, the LH1 complex surrounds the RC forming a ring; this RC–LH1 supercomplex, central to purple bacterial photosynthesis, is known as the core complex. As mentioned above, some purple bacteria, e.g. BChl a-containing Rsp. rubrum and BChl b-containing Blastochloris (Blc.) viridis, exclusively use the RC–LH1 core complex for both light harvesting and photochemical functions. The aggregation states of the RC–LH1 core complexes amongst different species are diverse (Fig. 5), with both monomeric and dimeric cores being found in the membranes.
Diverse structures of RC–LH1 complexes from different purple bacterial species. Cartoon representations viewed perpendicular to the membrane plane (top) and from the membrane plane (bottom) of (a) RC–LH1 from Blc. viridis (PDB: 6ET5) (Qian et al. 2018), (b) RC–LH1 of Thc. tepidum (5Y5S) (Yu et al. 2018), (c) RC–LH1-PufW complex of Rps. palustris (PDB: 1PYH) (Roszak et al. 2003), (d) RC–LH1-PufX dimer of Rba. sphaeroides (PDB: 4V9G) (Qian et al. 2013). The 4Hcyt, L, M and H subunits of the RC are represented in yellow, bright green, pale green and lime green, respectively; α -, β- and γ-chains of LH1 are shown in blue, light blue and cyan, respectively. PufX and protein W are represented in black and dark grey, respectively
2.2.1 Light-Harvesting Complex 1
The LH1 complex is composed of a ring of α- and β-polypeptides forming a heterodimer together with their associated pigment molecules: BChl a or b and carotenoids. These generally short, 40–70 amino acid residue, α-helical antenna polypeptides contain three domains: polar C- and N-terminal domains lying either side of a hydrophobic transmembrane domain (Brunisholz and Zuber 1992; Zuber and Cogdell 2004). A conserved histidine residue acts as the ligand to the central magnesium atom of the BChl (Qian 2017); in Blc. viridis, the C3 acetyl groups of BChl b hydrogen bond to tryptophan residues in LH1 (Qian et al. 2018), adopting an in-plane conformation similar to coordination in the LH2 of Rbl. acidophilus.
2.2.2 The Photochemical Reaction Centre
The RC, home of the light-induced charge separation across the intracytoplasmic membrane of Blc. viridis, was the first membrane protein structure to be solved at high resolution (Deisenhofer et al. 1985; Deisenhofer and Michel 1988). The RC consists of a common, basic structure of three transmembrane polypeptides, the L, M and H subunits (Fig. 6b), and their associated pigments, similar in all species of purple photosynthetic bacteria. L and M subunits each contain five transmembrane helices, and the LM heterodimer displays pseudo twofold axis symmetry, perpendicular to the membrane plane. These subunits possess 25–30% sequence identity. L and M act as a scaffold for the arrangement of cofactors: 4 BChls, 2 bacteriopheophytin (BPhe) pigments, 1 non-haem iron, 2 quinones and 1 carotenoid. The cofactors are also arranged with a pseudo twofold symmetry and form two separate branches, A and B, within the M and L subunits, respectively (Fig. 6c). Electron transfer has been shown to occur through branch A (Kirmaier et al. 1985; Bylina and Youvan 1988; Kellogg et al. 1989). The H subunit contains only 1 transmembrane helix, anchoring the subunit to the membrane, a cytoplasmic globular domain docks to L and M. The RC structure of Blc. viridis revealed an extra periplasmic tetrahaem cytochrome subunit (4Hcyt) attached to the membrane via a covalently attached fatty acid molecule (Deisenhofer et al. 1985). The 4Hcyt houses four haem groups covalently attached to cysteine residues, arranged in a high-low-high-low redox potential pattern (Fig. 6c), functioning to re-reduce the special pair BChl (Ortega et al. 1999). The conserved structure of RC is demonstrated by the crystal structures of Rhodobacter (Rba.) sphaeroides (Fig. 6d) (Allen and Holmes 1986; Chang et al. 1986) and Thermochromatium (Thc.) tepidum (Fig. 6e) (Nogi et al. 2005) RCs, both of which are very similar to that of Blc. viridis (Qian et al. 2018); however, Rba. sphaeroides (Allen and Holmes 1986; Chang et al. 1986) lacks the 4Hcyt subunit and both contain BChl a rather than BChl b.
Structures of LH1 and RC complexes. Cartoon representation of (a) LH1 from Blc. viridis (PDB: 6ET5) (Qian et al. 2018) from the periplasmic side of the membrane and (b) RC from Blc. viridis (PDB: 6ET5) (Qian et al. 2018) from the membrane plane. The RC resides in the centre of the LH1 complex; this has been removed from the structure. (c) Arrangement of cofactors in the RC from Blc. viridis; cofactors are represented as sticks. The redox potentials of the haems are labelled. (d) Crystal structure of the RC from Rba. sphaeroides (PDB: 2RCR) (Camara-Artigas et al. 2002). (e) Crystal structure of the RC from Thc. tepidum (PDB: 5Y5S) (Yu et al. 2018). The α-, β- and γ-chains of LH1 are shown in blue, light blue and cyan, respectively. The 4Hcyt, L, M and H subunits are represented in yellow, bright green, pale green and limon, respectively
2.2.2.1 Quinones
As well as being the home of charge separation, the RC also functions to produce fully reduced quinol (QH2) to drive electron transfers through cyt bc1, ultimately resulting in the formation of an electrochemical gradient driving ATPase. The RCs of purple bacteria consist of two acceptor quinones that act in series. The primary quinone, QA, is bound tightly to the RC, and cycles between oxidised quinone and singly reduced semiquinone. The secondary quinone, QB, is reduced twice by QA to form a doubly reduced, fully protonated QH2. The electron donor for these reactions is cyt c2, so the RC can be thought of as a ‘cytochrome c2/ubiquinone’ photo-oxidoreductase enzyme. After two turnovers of the RC, QH2 is released from the QB site and replaced by oxidised quinone to complete the quinone cycle. In the majority of purple bacteria, both QA and QB are ubiquinone (UQ) molecules, but in Blc. viridis, Allochromatium vinosum and Thc. tepidum, QA is a menaquinone (MQ). The X-ray crystal structure of the RC from Rba. sphaeroides revealed hydrogen bonding to QA (UQ-10) through carbonyl oxygens, a histidine residue and an amide backbone; a similar hydrogen bonding pattern was seen in QA from Blc. viridis (MQ-9) (Gast et al. 1985; Shopes and Wraight 1985). The acceptor quinones are symmetrically positioned about a non-haem iron atom coordinated by four histidines, and a glutamate residue, QA, is bound in the L subunit and QB within the M subunit (Deisenhofer and Michel 1988).
2.2.3 Additional Core Complex Components
2.2.3.1 PufX
The core complex of many species within the Rhodobacter genus contains an extra transmembrane polypeptide, PufX (Youvan et al. 1984; Sheng and Hearst 1986; DeHoff et al. 1988; Holden-Dye et al. 2008). PufX creates a break in the LH1 ring, keeping it in an ‘open’ conformation to facilitate rapid quinone/quinol exchange between the RC and cyt bc1 (Klug et al. 1988; Farchaus et al. 1990; Lilburn et al. 1992). PufX represents an α-helical structure consisting of 34 residues (Tunnicliffe et al. 2006); its N-terminus is largely unstructured, followed by a structured helical domain positioned to ensure basic residues are at the membrane interface to fulfil a membrane-anchoring role (Parkes-Loach et al. 2000). Studies on Rba. capsulatus and Rba. sphaeroides indicated that deletion of the pufX gene led to the inability of these strains to grow phototrophically (Klug et al. 1988; Farchaus et al. 1990; Lilburn et al. 1992). For a history of PufX, see the review by Holden-Dye et al. (Holden-Dye et al. 2008).
2.2.3.2 Protein W
The X-ray crystal structure of the core complex from Rps. palustris revealed that the RC was surrounded by an incomplete elliptical ring of 15 αβ-polypeptide pairs. Electron density maps suggested that a single transmembrane helix, named protein W, prevented closure of the LH1 ring (Fig. 5c) (Roszak et al. 2003). The break in the LH1 ring created by protein W is positioned adjacent to the secondary electron acceptor quinone (QB) binding site, as is the case for PufX, which suggested a similar potential role in quinone/quinol exchange with cyt bc1. The identity of protein W remained unknown for many years, until proteomic analysis of purified core complexes from the sequenced Rps. palustris strain CGA009 identified a polypeptide (RPA4402) that was present in the preparation (Jackson et al. 2018). Biochemical and electron microscopic analysis indicated that RPA4402/protein W is only present in ~10% of the purified core complexes, and deletion of the encoding gene had no measurable effect on phototrophic growth; thus, the precise role of protein W remains enigmatic (Jackson et al. 2018).
2.2.3.3 The Gamma Subunit
An additional LH1 subunit was also identified in core complex preparations of Blc. viridis (Jay et al. 1983); unlike protein W and PufX, this γ-polypeptide was found to be in apparent equimolar ratio with the LH1 α- and β-polypeptides (Brunisholz et al. 1985). Subsequently, the structure of the core complex from this organism revealed the location of γ, packing between β-polypeptides on the outside of the LH1 ring (Qian et al. 2018). The α:β:γ ratio was found to be 17:17:16, the ‘missing’ γ-subunit creating the channel for quinone diffusion, converse to the proposed roles for PufX and protein W, whose presence at a single location around the LH1 ring is believed to create this channel. An additional role of the γ-polypeptide in increasing the packing of LH1 BChls was also proposed; the resulting shorter intra-subunit Mg-Mg distances increase excitonic coupling, making the Blc. viridis RC–LH1 complex the most red-shifted photosynthetic complexes described to date (Qian et al. 2018).
2.2.4 Architectures of Core Complexes
The RC–LH1 complexes possess different forms across different purple photosynthetic bacteria (Fig. 5). Table 2 details the RC–LH1 composition for several species of purple bacteria.
Early, low-resolution electron microscopy studies on membranes from Blc. viridis and Halorhodospira (Hlr.) halochloris, followed by single particle analysis on purified core complexes from Phs. molischianum, indicated that the common structure of RC–LH1 complexes consisted of a circular LH1 antenna surrounding the RC (Stark et al. 1984; Engelhardt et al. 1986; Boonstra et al. 1994). Later studies employing atomic force microscopy (AFM) to interrogate the topography of membranes from purple bacteria (Liu and Scheuring 2013; Liu 2016) achieved sufficient resolution to assign the number of αβ-polypeptide pairs in the LH1 antenna from Blc. viridis, Rsp. rubrum, Psp. photometricum and Phs. molischianum; all considered the RC to be enclosed by 16 αβ dimers, with those from Blc. viridis (Fig. 7a) and Rsp. rubrum interpreted to be elliptical in shape (Fotiadis et al. 2003; Scheuring et al. 2003a, 2004a; Gonçalves et al. 2005). This elliptical shape was reinforced with the 4.8 Å crystal structure of the Rps. palustris core complex, although the LH1 ring was found to contain the W polypeptide in place of one of the 16 αβ pairs (Roszak et al. 2012). The ‘gap’ in the elliptical ring created by protein W was also identified by AFM (Fig. 7d) (Scheuring et al. 2006). The contemporary, high-resolution structures of the RC–LH1 complexes from Tch. tepidum and Blc. viridis (Fig. 7a), consist of 16 and 17 αβ heterodimers, respectively; the latter also containing 16 γ-subunits, illustrating that monomeric core complexes do not have a uniform architecture.
Diversity of core-complex architectures among purple photosynthetic bacteria species. Core complexes in their native membranes (left) and the models of individual core complexes (right) for (a) Blc. viridis (LH116-RCL,M,H-4Hcyt) (Scheuring et al. 2003a), (b) Rba. blasticus (PufX2-LH113-RCL,M,H)2 (Scheuring et al. 2005a), (c) Psp. photometricum (LH116-RCL,M,H) (Scheuring et al. 2004a), (d) Rps. palustris (W-LH115-RCL,M,H) (Scheuring et al. 2006) and (e) Rba. veldkampii (Liu et al. 2011a, b)
This lack of uniformity is compounded by the presence of dimeric core complexes in species of Rhodobacter. Cryo-EM of negatively stained membranes from an LH2 mutant of Rba. sphaeroides (the mutation causing the switch from its native vesicular to tubular membranes) revealed highly ordered core complexes in S-shaped conformations, believed to be comprised of two RCs, each surrounded by a C-shaped LH1 (Jungas et al. 1999). Sucrose density gradients of solubilised membranes from Rba. sphaeroides also revealed that monomeric and dimeric core complexes could be isolated in which PufX was detected in a 1:1 ratio with the RC, but when pufX was deleted, only monomeric RC–LH1 could be isolated, indicating that PufX promotes dimerisation of the core complex (Francia et al. 1999). AFM analysis of membranes from Rba. sphaeroides and Rba. blasticus revealed that the majority of core complexes exist in dimeric form in the native membranes of these strains (Fig. 7b) (Bahatyrova et al. 2004; Scheuring et al. 2005a). A subsequent structure of the dimeric core from Rba. sphaeroides revealed that each RC was surrounded by 14 αβ heterodimers, with each PufX adjacent to the QB site and interacting with the RC-H subunit via its N-terminal extension, and both N- and C-termini of PufX promoting dimerisation by interacting with a β polypeptide in the other half of the dimer (Qian et al. 2013). Interestingly, the core complex of Rba. veldkampii was found to contain a PufX polypeptide, but sedimentation and single particle analysis, and subsequent AFM analysis of its native membrane, demonstrated the complex exists solely as monomeric cores in this organism (Fig. 7e) (Gubellini et al. 2006; Liu et al. 2011a, b). More recently, a systematic analysis of additional Rhodobacter species demonstrated that dimeric core complexes were also found in Rba. azotoformans and Rba. changlensis, but those of Rba. capsulatus and Rba. vinaykumarii were monomeric, yet all contained PufX (Crouch and Jones 2012). This study also indicated that there was no clear motif within the amino acid sequences of PufX from the tested strains that could be identified as key for dimerisation; further work is required to clarify the biochemical basis of this morphological difference.
2.3 Cofactors and Pigments
2.3.1 Carotenoids
Carotenoids have several essential functions within photosynthetic systems:
-
1.
They are accessory pigments functioning in the collection and absorption of light energy and transferring this to BChl molecules.
-
2.
They function in the photoprotection process, rapidly quenching triplet excited states of BChl and preventing them from reacting with oxygen and forming the highly reactive and damaging excited singlet state of oxygen. If singlet oxygen is formed, carotenoids can also quench this (Blankenship 2014).
-
3.
Carotenoids also play an important role in the assembly of LH complexes; in order to reversibly dissociate LH complexes, removal of carotenoids from the complex is required (Loach and Parkes-Loach 1995).
In LH2 complexes, carotenoids extend between the subunits (McDermott et al. 1995; Koepke et al. 1996; Prince et al. 1997), stabilising the complex and interacting with the BChls.
Structures of several carotenoids present in purple bacteria can be found in Fig. 8. All of these carotenoids are extended molecules with delocalised π electron systems. A number of common characteristics exist including, but not limited to, the presence of tertiary hydroxy and methoxy groups at C-1, keto groups at C-2 and frequent double bonds in the C-3,4 position (Takaichi 2000).
2.3.2 Bacteriochlorophylls
The main structural cofactors and LH pigments of photosynthetic antennae of purple bacteria are BChls, which also play a key role in photochemistry in the RC; energy is harvested and transferred from BChls within the LH complexes to the RC for charge separation at the special pair. Many different types of BChl exist within phototrophic prokaryotes, but only two types exist in purple photosynthetic bacteria, BChl a and BChl b (Fig. 1). BChl a is the most common pigment found in purple bacteria, permitting absorption of wavelengths between 800 and 970 nm when assembled into LH complexes, shifting the Qy absorption band of the pigment in solution into the near infrared (Imanishi et al. 2019) (Table 1). BChl b is only found in a few species; it differs from BChl a by the presence of an exocyclic double bond at C-8 in ring B, known as an ethylidene substituent (Fig. 1). BChl b permits absorption between 960 and ~1050 nm, the longest wavelength absorbance band of any chlorophyll-type pigment. The use of BChls a or b allows purple bacteria to absorb wavelengths of light outside the range of the chlorophylls used by plants and cyanobacteria.
2.3.3 Bacteriopheophytins
BPhe are easier to reduce than BChls and hence function as electron acceptors within the sequence of electron carriers in places where BChls are not active (Fajer et al. 1975). They are metal-free BChls, lacking the central Mg2+ present in BChls. They are predominantly recognised as breakdown products, due to their formation during the degradation of BChl. However, small amounts of BPhe are essential components of RCs, which are produced in specific, yet unknown, pathways. In charge separation, BPhe (termed HA) function as the first electron acceptor subsequent to the excitation of the special pair of BChls (Zinth and Wachtveitl 2005; Jones 2009).
2.4 Cofactor-Cofactor and Protein-Protein Interactions
A large, overlapping array of BChl molecules is formed by dense packing of LH1 and LH2 proteins in the photosynthetic membranes. This ensures that absorbed energy and the excited state are both quickly delocalised among the array and thus can be easily transferred to other LH complexes and eventually to the RC. Oligomerisation of B820 is required for the formation of LH1 and LH2 complexes (Miller et al. 1987; Chang et al. 1990). B820 subunits readily associate with the corresponding LH apoproteins to form LH1 and LH2 complexes with dependence on the details of cofactor-protein and protein-protein interactions. The heterodimeric B820 complex is stabilised by the interaction of the N-terminal regions of α- and β-polypeptides (Parkes-Loach et al. 2004). Several studies have demonstrated that fine-tuning of the absorption maximum of the associated LH complex may be due to evolution of the α-polypeptide (Todd et al. 1998). Dissociation and reassociation experiments revealed that under in vitro conditions, B820 reassociates to form LH1 via several intermediate species, reflecting the different numbers of subunits that associate during growth of the complex and eventual formation of the ring. Once this structure is formed, some rearrangements of the protein segments occur on a slower timescale (Miller et al. 1987; Pandit et al. 2003).
2.5 Assembly of Complexes
Assembly factors LhaA and PucC are important, but not essential, in the specific assembly of LH1 and LH2 complexes in vivo, respectively. These factors form oligomers at sites of initiation of membrane invagination. LhaA makes associations with RCs, BChl synthase, the protein translocase subunit YajC and the membrane protein insertase YidC, aiding coordination of pigment delivery, co-translational insertion of the LH polypeptides and the folding and assembly of these form functioning complexes (Mothersole et al. 2016). Reconstitution experiments demonstrated that LH1 subunits possess the ability to reassociate with RCs to form functional, native-like core complexes, in the absence of LhaA (Bustamante and Loach 1994). The RC–LH1 may follow a similar pattern of assembly in vivo. As is seen in Rba. sphaeroides (Pugh et al. 1998), the RC is likely to form first, allowing the LH1 subunits to subsequently encircle it. This theory suggests that the LH1 subunits are static within the complete circle; however, these subunits may in reality be flexible (Jamieson et al. 2002; Bahatyrova et al. 2004). The forces underlying the stability of the B820 subunit may be minimal, allowing the subunit to readily dissociate; this would provide a possible pathway for quinones on release from the QB binding site (Hunter et al. 2008).
2.6 Spectroscopic Properties of Light-Harvesting Complexes
Understanding the spectroscopic properties of LH complexes is fundamental as they are the key factors in directing the flow of excitation energy towards the RC. Although many antenna proteins contain the same BChl molecules, their absorption spectra can differ. BChl a and BChl b (Fig. 1) molecules of LH1 complexes absorb at 870–890 nm and near 1000 nm, respectively, whereas the BChl a molecules of LH2 complexes absorb at 800 and 850 nm. Additionally, a number of strains have been shown to synthesise modified antenna proteins absorbing at unusual wavelengths, for instance Merichromatium purpuratum (830 nm) (Cogdell et al. 1990) and Roseococcus thiosulfatophilus (856 nm) (Gall et al. 1999). The arrangement of BChl molecules within LH complexes could modify the spectroscopic properties of LH complexes. 24–32 and 16–18 BChl molecules associate with LH1 and LH2, respectively, through van der Waals contacts, leading to the red shifting of the absorption properties of the complexes (van Grondelle et al. 1994), compared to the BChls in solution.
The absorption spectra of LH3 and LH4 complexes are shifted to be shorter than 820 nm. The primary sequence of LH proteins between complexes and species differs; changes in the amino acid sequence in the protein environment of BChls can affect the hydrogen-bonding state and the dielectric properties of the protein environment (Fowler et al. 1994, 1997; Gall et al. 1997); this explains the part of the blue shift observed. Recent experiments showed that a change in the hydrogen bonding network affected the Qy transition of BChls but also indirectly affected the role of charge-transfer states (Nottoli et al. 2018), leading to a new explanation of the blue shift seen in LH3 and LH4 complexes.
Homogenous LH proteins do not necessarily share the same electronic properties (van Mourik et al. 1992); slight differences in the conformation of the immediate environment of the BChls or dynamic fluctuations in the system, known as the static and dynamic disorder, respectively, lead to changes in the local physicochemical properties of the BChl and heterogeneity in the electronic properties. This disorder in the structure changes the properties of otherwise chemically identical BChls, which results in the localisation of excited states onto a subset of BChls within the interacting group (Hunter et al. 2008).
2.7 Cytochrome bc1
Cyt bc1 is a multisubunit membrane-bound enzyme and an essential component of the energy transduction machinery in purple photosynthetic bacteria (Fig. 9) (Berry et al. 2000, 2004; Crofts 2004). The enzyme functions to electronically connect the quinone pool to the downstream electron transport chain (ETC) and translocate protons across the membrane helping to produce the pmf (Crofts 2004). The cyt bc1 oxidises QH2 and donates the electron to the membrane-soluble electron carrier cyt c2 (Jenney and Daldal 1993). Therefore, this enzyme can be thought of as a ‘ubiquinol/cytochrome c2 oxidoreductase’, operating in the opposite direction to the RC and completes the cyclic electron flow utilised by purple bacterial photosynthesis.
Structures of cytochrome bc1 and ATP synthase. (a) Structure of RC–LH1 from Blc. viridis (Qian et al. 2018) shown for comparison, colour scheme identical to Fig. 5. (b) Structure of cyt bc1 from Rba. capsulatus (PDB: 1ZRT) (Berry et al. 2000). The cyt b, cyt c1 and Rieske subunits are represented in yellow, red and blue, respectively. (c) Structure of the ATPase from Bacillus PS3 (PDB: 6 N30) (Guo et al. 2019). The c-ring, A, B, α, β, δ, γ and ε subunits are represented in orange, blue, purple, red, yellow, light blue, pink and green, respectively. The grey lines represent the intracytoplasmic membrane
The overall structure of cyt bc1 (Fig. 9b) consists of an intertwined homodimer with two monomers organised about a twofold molecular axis; each monomer is comprised of three subunits: a high-potential [2Fe–2S] cluster-containing subunit, known as the Rieske protein (Rieske et al. 1964), an integral membrane cyt b subunit and a c-type cytochrome subunit, known as cyt c1, in which the haem cofactor is covalently attached to the polypeptide (Berry et al. 2000; Darrouzet et al. 2004). Cyt b consists of two b-type haems located on the positive (p) and negative (n) sides of the membrane. The central core of the monomer is formed by ten transmembrane helices, one from the Rieske protein, one from cyt c1 and eight from cyt b. The large protrusion on the p side of the membrane consists mainly of the hydrophilic parts of the Rieske protein and the cyt c1 subunits, whereas the n side is seemingly devoid of proteins (Fig. 9) (Berry et al. 2004). Each monomer contains two Q/QH2 binding sites referred to as Qo (QH2 oxidation, H+ output) and Qi (Q reduction, H+ input) sites; the Qo site is at the interface of the Rieske protein and cyt b on the p side of the membrane, and the Qi site is in the cyt b subunit, closer to the n side of the membrane (Sarıbaş et al. 1998; Berry et al. 2000).
2.8 ATP Synthase
The electrochemical gradient produced by the actions of the photosynthetic machinery is used by F0F1-ATPase, a membrane protein that synthesises ATP from ADP and inorganic phosphate (Pi). The enzyme contains two distinct portions of multisubunit complexes; a hydrophobic proton-translocating F0 proton channel is embedded within the membrane, consisting of three subunits, A, B and C, and a hydrophilic, enzymatic F1 portion, consisting of α, β, ε and γ subunits, protrudes from the plane of the membrane by ~100 Å (Fig. 9) (Guo et al. 2019). Three αβ heterodimers in F1 form a hexamer, with a long central cavity filled with the elongated portion of the γ-subunit comprising an N-terminal coiled-coil structure and an α-helical domain close to the C-terminus; this protrudes ~30 Å into the stalked region. A short α-helix of the γ-subunit is at a 45° incline to the coiled-coil domain at the bottom of F1. The γ-subunit makes up a coupling domain, coupling ATP hydrolysis with ion pumping (Capaldi et al. 1996; Junge et al. 1997).
Structure of cytochrome c2. (a) Crystal structure of cyt c2 from Rba. sphaeroides (PDB: 1CXC) (Axelrod et al. 1994); the haem group is represented in green. (b) Crystal structure of Rba. sphaeroides cyt c2:RC (PDB: 1L9B) (Axelrod et al. 2002). Cyt c2 is represented in purple, the L, M and H subunits of the RC are represented in magenta, cyan and orange, respectively. The haem group in cyt c2 and the special pair BChl in RC are represented in black to highlight their positioning in the complex
The αβ heterodimer contains six nucleotide-binding sites within the six clefts between adjacent subunits; only three of these participate in catalysis (Weber et al. 1994, 1995); the role of the other three clefts is not known but they are believed to play a regulatory role (Milgrom et al. 1991; Jault and Allison 1993; Jault et al. 1995; Matsui et al. 1997). The catalytic sites are mainly formed by the β subunit. Although they are all identical, at any given time, they have different conformations and affinities to nucleotides (Kayalar et al. 1977). Nucleotide binding to a particular site triggers the release of a nucleotide from another site; this is intimately linked to the rotation of the γ-subunit within the αβ heterodimer (Senior et al. 2002; Weber and Senior 2003).
2.9 Cytochrome c2
The water-soluble c-type haem protein cyt c2 serves as an electron donor to the RC, rapidly reducing the oxidised special pair created during the charge separation step due to its optimisation for rapid association and dissociation (Tiede and Dutton 1993). The haem group of cyt c2 is covalently linked to the protein through thio-ether linkages; one edge of this haem group is exposed to solvent, which is important for reduction of the RC (Axelrod et al. 1994) (Fig. 10). Many positively charged Lys and Arg residues surround the solvent-exposed haem edge; these interact electrostatically with a cluster of negatively charged residues on the surface of the RC, positioning the cyt c2 in the centre of the periplasmic surface of the RC, so that the solvent-exposed haem edge is directly over the BChl dimer, the primary donor, for rapid electron transfer (Rosen et al. 1983; Caffrey et al. 1992; Drepper et al. 1997). This binding region exhibits close van der Waals contacts important for both binding and electron transfer. A crystal structure of the cyt c2:RC complex from Rba. sphaeroides has been determined, confirming this binding position (Axelrod et al. 2002). The position of the haem is similar to that found in the 4Hcyt subunit of the RC of Blc. viridis (Fig. 6b) (Deisenhofer and Michel 1988). Binding of the cyt c2 to RC leads to a slight conformational change creating a kink in the polypeptide chain of the RC, indicating a rigidly fixed-specific binding site of the cyt c2 on the RC (Axelrod et al. 2002).
3 Organisation and Assembly of Photosynthetic Membranes
Knowledge of the static structures of individual membrane proteins is insufficient to fully understand their function and physiological coordination. An appreciation of in situ assembly and distribution of the proteins within their native membrane, and analysis under near-native conditions, are fundamental. In addition, it is increasingly accepted that structural heterogeneity exists within these proteins, between proteins from different species and between distinct gene products within a species, and between individual complexes. In addition to studying the structures of individual membrane proteins, AFM has matured to be a powerful technique to visualise the organisation of different membrane proteins in the native environment, enabling visualisation of the macromolecular structures in biological membranes and the dynamics of adaptation to different conditions (Scheuring et al. 2005b, 2007). AFM imaging of photosynthetic membranes from various purple bacteria have shown a variety of different organisational patterns of the photosynthetic apparatus, ranging from highly ordered to less ordered arrangements of proteins (Liu and Scheuring 2013; Liu 2016).
The simplest purple bacterial photosynthetic membrane is that of Blc. viridis (Scheuring et al. 2003a; Qian et al. 2018), which contains simplified photosynthetic units lacking LH2 complexes. Small chromatophores were isolated from Blc. viridis and analysed using AFM (Scheuring et al. 2003a), revealing a highly organised architecture composed of hexagonally dispersed single RCs, each surrounded by a closed ellipsoid of LH1 subunits (Fig. 11b), similar to the cryo-EM structure (Qian et al. 2018). After removing the 4Hcyt subunit from the RC, AFM images revealed that the underlying RC-L and -M subunits adopt an asymmetric topography, whereby orientation of the long axis of LH1 coincides with this, representing an important constraint for energy transfer (Scheuring et al. 2003b). When the entire RC is removed, the LH1 units adopt a closed circular structure of ~100 Å diameter. This change reflects the flexibility of the LH1 heterotrimer assembly and the strong and specific interactions between the RC and LH1 components. This flexible motion could promote a ‘breathing’ motion to facilitate quine/quinol exchange, similar to that proposed for the LH1 of Thc. tepidum (Niwa et al. 2014).
Molecular resolution AFM topographs of purple photosynthetic apparatus. (a) Protein organisation in the vesicular ICMs of Rba. sphaeroides (Bahatyrova et al. 2004). (b) Protein organisation in the flat ICM of Blc. viridis (Scheuring et al. 2003b). (c) Protein organisation in the stacked ICMs of Psp. photometricum (Scheuring et al. 2004a)
The presence of LH2 complexes leads to a slightly more complex photosynthetic unit (PSU) arrangement; nevertheless, AFM images of the photosynthetic membrane of wild-type Rba. sphaeroides revealed a relatively ordered, interconnected network of LH2 and dimeric RC–LH1-PufX core complexes, coexisting with ordered LH2-only domains (Fig. 11a). The RC–LH1-PufX dimers form linear arrays, composed of pairs of elliptical structures with a protruding protein in the centre (Fig. 11a), interconnected with rows of peripheral antenna LH2 complexes (Bahatyrova et al. 2004). This physical continuity between the complexes ensures a highly efficient transfer of excitation energy from the peripheral LH2 to the LH1 complexes and onto an active RC (Bahatyrova et al. 2004; Scheuring et al. 2005a). The LH1 complexes are positioned to serve as a hub for the collection of excitons from LH2; the excitons can migrate along a series of dimers until they reach an open RC (Bahatyrova et al. 2004). LH2 was found to preferentially hexagonally pack within convex vesicular regions of the membrane (Scheuring et al. 2014). Cyt bc1 was postulated to localise in disordered areas adjacent to RC–LH1 complexes; cyt bc1 is outnumbered by RC–LH1 by a 3:1 ratio, placing them out of direct contact with the core complexes (Cartron et al. 2014; Scheuring et al. 2014). Mutants of Rba. sphaeroides lacking PufX had a completely reorganised ICM with ordered arrays of monomeric RC–LH1 complexes from which LH2 is largely excluded and segregated to the curved membrane domains (Frese et al. 2004). The interconnection of RC–LH1-PufX arrays interspersed with rows of LH2 complexes was also prevented in the mutant, suggesting that PufX plays a long-range organisational role within the membrane (Frese et al. 2004).
Despite the similarity in the structures of the RC–LH1-PufX complex in Rba. blasticus (Scheuring et al. 2005a) and Rba. sphaeroides (Jungas et al. 1999; Qian et al. 2008), AFM images revealed that the Rba. blasticuus RC–LH1-PufX complexes did not form rows that were seen in Rba. sphaeroides (Bahatyrova et al. 2004). Instead individual S-shaped dimeric core complexes are interspersed with nonameric LH2 rings (Fig. 7d) (Scheuring et al. 2005a).
Other LH2 containing species displayed much less regular arrangements of their PSUs. Psp. photometricum ICMs, consisting of stacked, flattened thylakoid-like discs, contain a mixed, disordered arrangement of monomeric RC–LH1 complexes and circular nonameric LH2 rings (Fig. 11c) (Scheuring et al. 2004a, b), and inter-complex clustering was visualised (Scheuring et al. 2004b; Scheuring and Sturgis 2005; Liu et al. 2009). The membranes are heterogeneously organised with RC–LH1-rich domains separate from hexagonally packed, almost crystalline LH2 domains. Some of the core complexes were completely surrounded by 7 LH2 complexes, whereas others made multiple contacts with other core complexes, increasing the probability that excitons will find an open RC, enhancing rapid energy trapping as is seen in the more organised Rba. sphaeroides membranes (Bahatyrova et al. 2004). This organisation provides efficient movement of excitons towards the RC whilst opening the ring of LH2 complexes to ensure all LH2 are in contact with one another. A heterogeneous variability in ring sizes of LH2 was also observed including octamers, nonamers and decamers, representing a natural variability in ring sizes, suggesting a possible strategy for broadening absorption to maximise the collection of near-IR radiation and optimisation of the packaging of photosynthetic components within the membrane (Scheuring et al. 2004b). Although cyt bc1 was demonstrated to be in the membrane preparations, it was not identified in AFM images (Scheuring et al. 2004a).
Similarly, the ICMs of Phs. molischianum also consist of thylakoid-like structures giving rise to flat membrane sheets. The LH2 and core complexes are also arranged similarly to that in Psp. photometricum; LH complexes are segregated into two structurally different domains: one consists of a mixture of core complexes and a small amount of octameric LH2 complexes, and the other consists of paracrystalline, hexagonally packed octameric LH2 rings (Gonçalves et al. 2005). The crystal structure of the LH2 complex from Phs. molischianum (Koepke et al. 1996) enabled the exact pigment distances between LH2 complexes to be deduced (Hu et al. 2002); intercomplex distances between central Mg2+ atoms are ~16.3–28.3 Å, enabling efficient energy transfer.
The membranes of Rps. palustris consist of a complex ICM structure made of infoldings of the cytoplasmic membrane forming regular bundles of stacked and flattened thylakoid-like membrane sacs. AFM images of Rps. palustris membranes were obtained under differing light conditions, reflecting the versatility of photosynthetic apparatus in chromatic adaptation (Fig. 7f) (Scheuring et al. 2006). Under low light, mixed domains were commonly observed, along with paracrystalline domains of peripheral LH2 complexes, as is seen for other purple bacteria containing lamellar ICM structures. The mixed domains are likely to arise from the proximal layers within the lamellar ICM folds, whereas the LH2-only regions arise from distal layers formed subsequently; this is supported by the assembly sequence of complexes in developing ICMs (Koblízek et al. 2005). Under high light, regions of crystalline RC–LH1 core complexes were observed, with a few LH2 complexes. This arrangement of randomly ordered RCs would be expected to facilitate efficient energy trapping under high photon fluxes. Chromatic adaptation also resulted in the modification of LH2 ring sizes and absorption spectra (Scheuring et al. 2006). AFM images also confirmed the structures of the RC–LH1 complex, consisting of LH1 rings composed of 15 heterodimers interrupted by a gap, presumed to be the location of protein W (Scheuring et al. 2006).
3.1 Common Features of the Photosynthetic Membranes
Although the organisations of photosynthetic membranes in differing species vary, there are a number of recurring features. All the species contain membranes densely clustered with photosynthetic complexes, ensuring efficient excitation energy capture and transfer between antennae and ultimately to the RC for charge separation. In addition, the organisation of photosynthetic apparatus is never entirely random; clustering of RC–LH1 and formation of LH2 domains are often seen. Although the locations of cyt bc1 and ATPase in most species remain enigmatic, the location of Rba. sphaeroides cyt bc1 has been proposed (Cartron et al. 2014; Scheuring et al. 2014), and AFM on intact chromatophores from the same organism revealed a high protrusion from the membrane surface, speculated to be ATPase (Kumar et al. 2016).
3.2 Functional Importance of Photosynthetic Membrane Organisation
The architectural variability of the photosynthetic membranes permits the systems to work efficiently in their native environments under various conditions. It is difficult to simultaneously satisfy the paradox of the organisation of the photosynthetic apparatus; these include, firstly, the expectation that the LH system should completely surround the RC for maximum efficiency and to minimise the distances for energy to travel between complexes. Next, the core complexes should be connected in order to avoid energy loss from closed RCs by allowing energy transfer between core complexes (Bahatyrova et al. 2004; Scheuring et al. 2005a). And thirdly, the cyt bc1 complex is expected to be in close proximity to the RC to close the electron circuit by quinol diffusion.
The first problem lies with the location of cyt bc1 and the release of quinone/quinol. For Rba. sphaeroides and Rps. palustris whereby the RC–LH1 complexes contain PufX and protein W, respectively, the diffusion of quinone is fairly simple due to the gap produced by these proteins creating a possible pathway (Roszak et al. 2003; Bahatyrova et al. 2004; Qian et al. 2005). In other species, however, the ring of LH1 was observed to be closed; therefore, quinone transfer across the LH1 wall would depend on the dynamics of the process by which the integrity of the ring is breached. Circular and elliptical closed forms of LH1 were observed in AFM images of Rsp. rubrum 2D crystals, suggesting the structural flexibility of LH1 that facilitates the transfer of quinone through molecular motions (Jamieson et al. 2002; Siebert et al. 2004). The missing 17th gamma subunit of Blc. viridis RC–LH1 (Qian et al. 2018) also provides a path for this diffusion, in this case through the absence of an additional subunit, rather than the presence of one.
Once the quinone is released from the RC, another problem arises in its rapid diffusion through a crowded membrane to cyt bc1 complexes. A lipid environment was found to be created around the core complexes of Psp. photometricum, due to a size mismatch of the LH2 and core complexes, providing a potential long-range pathway for quinones to travel (Liu et al. 2009). However, it was revealed that static paracrystalline LH2 complexes in the photosynthetic membrane, with significantly restricted diffusion, provided no space for quinones to diffuse between them (Scheuring et al. 2006).
4 Energy Transfer
4.1 Transfer of Excitation Energy
The knowledge of the structures and spatial arrangement of photosynthetic complexes and association with cofactors provide a basis for understanding energy transfer performed by the photosynthetic system. Energy is collected by antenna systems via the funnel concept; the pigments of the most peripheral antenna complexes, e.g. LH2, absorb shorter wavelengths than the pigments within RC. Photons of shorter wavelengths are higher in energy than longer wavelength photons and thus ensure ‘downhill’ energy flow: LH2 ➔ LH1 ➔ RC. A certain amount of energy is lost as heat during each transfer, providing some irreversibility and resulting in the funnelling of excitation energy into the RC. This funnel model is enabled by the spatial and energetic organisation of the antenna pigments. This concept also applies within LH complexes in the energy transfer from carotenoids to BChls; however, some energy transfers occur ‘uphill’, e.g. between LH1 and the RC in Rba. sphaeroides and Blc. viridis (Sumi 2002).
Higher-energy photons are first absorbed by carotenoids in LH2, or in LH1 in species lacking LH2; carotenoids absorb short wavelengths, 400–550 nm, of light and subsequently transfer the energy to both Qx and Qy states of B800 in LH2. Upon absorption of light, carotenoids are promoted from the ground state, S0 to the S2 state (transition to S1 state is symmetry-forbidden); the molecule then quickly relaxes to the S1 state in less than 300 fs. The lifetime of the S1 state varies from 300 ps to ~1 ps, depending on the conjugation length of the carotenoid (Sundström 2004). Once the excitation energy is passed from S1, or sometimes S2 (Polívka and Frank 2010), to B800, it is quickly transferred to B850 in ~1 ps by Förster resonance energy transfer (FRET). The excitation energy is subsequently delocalised over the ring of tightly coupled B850 pigments, i.e. it hops rapidly around the ring, within 100 fs (van Oijen et al. 1999). The excited state is thus equally probable to transfer from any site within the ring, implying that no fixed arrangement of LH2 and LH1 complexes is required for efficient transfer (Cogdell et al. 1999). If the energy does not reach another LH complex within 1 ns, it will decay. Energy from B850 is subsequently transferred to the B875 pigments in ~3 ps, and, again, it is delocalised over the tightly coupled B875 pigments. The relatively large distance between B875 and the RC makes the transfer between these pigments the slowest step, occurring in ~35–50 ps, which is essentially irreversible (Pullerits and Sundström 1996; Sundstrom and Grondelle 1999).
4.2 Charge Separation in the RC
The transfer of excitation energy to the RC induces the separation and stabilisation of charge across the photosynthetic membrane. A BChl dimer, known as the special pair (P), acts as the primary electron donor and is excited to P∗; this excited state then transfers an electron to a BPhe molecule via a transient radical pair state; BPhe then subsequently transfers an electron to QA. A c-type cytochrome, or the 4HCyt subunit in certain species, reduces oxidised P+, whilst QA− transfers an electron to QB. At this point, an electron has been transferred across the membrane, and the primary reactants are ready for the next electron transfer.
4.3 Electron Transfer in Cytochrome c2
In species that do not contain the 4Hcyt subunit, cyt c2 re-reduces P+. C-type cytochromes contain covalently attached haem groups; the haem group is located in the crevice of cyt c2, with one edge exposed to the outside of the protein (Axelrod et al. 2002). This exposed edge is surrounded by a group of positively charged lysine residues which interact with the negative charges of the reaction partner forming a tightly bound complex. Cyt c2 thus binds the RC at a specific site with the haem optimally positioned for electron transfer (Fig. 9) (Axelrod et al. 2009). The oxidation kinetics of cyt c2 are multiphasic; when bound to the RC, the oxidation occurs within ~1 μs; however when the reduced cyt c2 is unbound prior to electron transfer, the phase is slower (Mathis et al. 1994). The cytochrome needs to diffuse to the RC, and dock in place ready for oxidation; this results in the slow phase.
In species that contain the 4Hcyt subunit, the cytochrome is permanently positioned for optimal electron transfer. After flash excitation, the haem located closest to P is oxidised within a few hundred ns; electron transfer then occurs down the chain of haems within 4Hcyt re-reducing the first haem in μs. The 4Hcyt is then reduced by cyt c2 (Menin et al. 1998; Myllykallio et al. 1998). At this stage, protons have been taken up from the cytoplasmic side of the membrane, and cyt c2 has been oxidised in the periplasmic side.
4.4 Modified Q Cycle
Cyt bc1, located in the ICM (Xia et al. 1997; Zhang et al. 1998; Berry et al. 2000; Crofts 2004), mediates the completion of cyclic electron transfer in the modified Q cycle (Fig. 3) (Crofts 2004). Ubiquinone (UQ) binds the Qo site (see Sect. 2.3 for details on the structure of cyt bc1) of cyt b. UQ then transfers the electron to the Rieske subunit, leaving ubisemiquinone in the Qo site, and a proton is released to the periplasmic side of the membrane. Next, ubisemiquinone transfers a second electron to the lower redox potential haem and subsequently to the higher redox haem; a second proton is also released into the periplasmic side of the membrane. The Rieske subunit then undergoes a large amplitude motion, moving from the cyt b to the cyt c1 subunit, displacing the Fe-S centre. UQ next binds to the Qi site and is reduced to the semiquinone by the higher redox haem of cyt b. Oxidised UQ is then dissociated from the Qo site, and the Rieske Fe-S centre reduces cyt c1, which in turn reduces cyt c2. The Rieske subunit then moves back to its original position. At this stage, one quinol has been oxidised, an electron has traversed the full electron transport chain, and another has reduced the oxidised UQ to the semiquinone state for the following turnover. A new UQ binds to the Qo site and transfers its electron to the Rieske subunit, which then subsequently passes it to cyt c1 and then cyt c2 along one branch and then the two haem groups from cyt b on the other branch. The electron on the higher redox potential haem then reduces semiquinone to quinol, taking two protons from the cytoplasmic side. Finally, the reduced quinol dissociates from the complex (Crofts 2005).
Two turnovers of cyt bc1 result in the transfer of two electrons to cyt c2, two UQ oxidised to quinone form and one UQ reduced to hydroquinone quinol form and four protons transferred from the cytoplasmic to the periplasmic side of the membrane. Overall, this electron flow gives rise to pmf across the membrane which is subsequently utilised by ATPase.
4.5 Proton Translocation and ATP Synthase
ATPase exploits the proton motive force created by the photosynthetic electron transfer chain to produce ATP from ADP and Pi. Specifically, the F0 portion of the enzyme uses the proton motive force to generate a torque, which is then used by the F1 portion in the synthesis of ATP (Noji et al. 1997; Wada et al. 1999; Pänke et al. 2000; Kinosita et al. 2004; Junge et al. 2009) (Fig. 8). ATPase operates via the binding change mechanism (Boyer 1993). The major energy-requiring step is the simultaneous binding of substrates to, and release of products from, the ATP catalytic sites (Boyer et al. 1973; Kayalar et al. 1977). The rotation of subunits extending through the ATPase complex couples these affinity changes to proton transport. Three ATP catalytic sites located in the β-subunit of the complex sequentially alternate between open, loose and tight binding sites. As the γ-subunit rotates, it acts as a camshaft alternatively distorting the β-subunit, leading to the cycling between the three binding sites (Hunter et al. 2008). Rotation of the γ-subunit in the centre of the F1 portion deforms the surrounding catalytic subunits, giving rise to the three binding sites (Cross 2000). Rotation of the c subunit relative to the α subunit, however, is required for completion of the proton pathway (Vik and Antonio 1994; Duncan et al. 1995; Hatch et al. 1995; Junge et al. 1996). The energy from this conformational change is transduced into the ATP phosphoanhydride bond. Entrance and exit channels for protons are contained within the F0 subunit, but there is no connection between the two channels (Junge et al. 1997). An essential glutamic acid residue in a transmembrane helices within the entrance channel is protonated by residing protons; this conformational change ratchets the c subunit with respect to the α subunit (Rastogi and Girvin 1999; Junge et al. 2009; Pogoryelov et al. 2009). The c ring rotates; in turn each σ subunit picks up a proton and releases it through the exit channel. Therefore, during one complete cycle, a proton is transported through each c subunit, and each of the three β-subunits goes through open, loose and tight conformations, each producing and releasing a molecule of ATP.
5 Calvin-Benson-Bassham Cycle
The final stage in purple bacterial photosynthesis is the use of the energy generated in the steps discussed to drive the conversion of inorganic carbon into organic compounds. Purple photosynthetic bacteria contain the most diverse metabolism of carbon compounds. Both purple non-sulphur and sulphur bacteria assimilate CO2 into 3-phosphoglycerate (3-PG) and reduce CO2 mainly via the Calvin-Benson-Bassham (CBB) cycle (Fig. 12) (Benson 2002; Tabita 2004). The CBB cycle is a complex series of reactions, which can be broken down into three phases: carboxylation, reduction and regeneration (Heldt and Piechulla 2011).
Schematic of the Calvin-Benson-Bassham (CBB) cycle. The CBB cycle occurs in three main stages; RuBisCO is first used in carboxylation to produce six molecules of 3-PGA from the incorporation of inorganic carbon (CO2) and RuBP. ATP and NADPH are then utilised to phosphorylate and reduce 3-PGA to produce 3-PG in the reduction stage. Finally, in the regeneration stage, 5 3-PG molecules are recycled to regenerate RuBP, and one molecule of 3-PG is used to make glucose; two cycles of the cycle are required to make a molecule of glucose. Orange circle = carbon, green diamond = phosphate
Carboxylation is the first stage of the cycle and utilises ribulose-bisphosphate carboxylase/oxygenase (RuBisCO) to incorporate carbon from CO2 into an organic molecule. Ribulose-1,5-bisphosphate (RuBP) is attached to a molecule of CO2 to form a six-carbon compound; this spontaneously breaks down into two molecules of 3-phosphoglyceric acid (3-PGA). The next stage of the cycle is reduction (Heldt and Piechulla 2011); 3-PGA molecules are converted to 3-PG by phosphorylation. First ATP donates a phosphate group to each of the two 3-PGA molecules producing 1,3-bisphosphoglycerate. This is then reduced by NADPH releasing a phosphate group and producing 3-PG. The final stage is regeneration, in which some of the 3-PG molecules are used to make glucose, while others are recycled to regenerate the RuBP acceptor.
AAPs are obligate photoheterotrophs; no autotrophic carbon assimilation pathways have been reported (Yurkov and Beatty 1998; Fuchs et al. 2007; Swingley et al. 2007). AAPs and some anaerobic anoxygenic phototrophs have been reported to grow heterotrophically on acetate using the ethylmalonyl-CoA pathway and the oxidative glyoxylate cycle (Tang et al. 2011; De Meur et al. 2018).
References
Aagaard, J., & Sistrom, W. R. (1972). Control of synthesis of reaction center bacteriochlorophyll in photosynthetic bacteria. Photochemistry and Photobiology, 15(2), 209–225. https://doi.org/10.1111/j.1751-1097.1972.tb06240.x.
Alberti, M., Burke, D. H., & Hearst, J. (2004). Advances in photosynthesis. Anoxygenic photosynthetic Bacteria. Available at: https://springerlink.bibliotecabuap.elogim.com/content/pdf/10.1007%2F0-306-47954-0.pdf. Accessed 29 Aug 2019.
Allen, J. F., & Holmes, N. G. (1986). A general model for regulation of photosynthetic unit function by protein phosphorylation. FEBS Letters. https://doi.org/10.1016/0014-5793(86)80682-2.
Angerhofer, A., Cogdell, R. J., & Hipkins, M. F. (1986). A spectral characterisation of the light-harvesting pigment–protein complexes from Rhodopseudomonas acidophila. Biochimica et Biophysica Acta (BBA) – Bioenergetics, 848(3), 333–341. https://doi.org/10.1016/0005-2728(86)90208-2.
Axelrod, H. L., et al. (1994). Crystallization and X-ray structure determination of cytochrome c2 from Rhodobacter sphaeroides in three crystal forms. Acta Crystallographica Section D Biological Crystallography. International Union of Crystallography (IUCr), 50(4), 596–602. https://doi.org/10.1107/S0907444994001319.
Axelrod, H. L., et al. (2002). X-ray structure determination of the cytochrome c2: Reaction center electron transfer complex from Rhodobacter sphaeroides. Journal of Molecular Biology. Academic Press, 319(2), 501–515. https://doi.org/10.1016/S0022-2836(02)00168-7.
Axelrod, H., Miyashita, O., & Okamura, M. (2009). Structure and function of the cytochrome c 2: Reaction center complex from Rhodobacter sphaeroides. In (323–336). Dordrecht: Springer. https://doi.org/10.1007/978-1-4020-8815-5_17.
Bahatyrova, S., et al. (2004). The native architecture of a photosynthetic membrane. Nature. Nature Publishing Group, 430(7003), 1058–1062. https://doi.org/10.1038/nature02823.
Benson, A. A. (2002). P aving the P ath. Annual Review of Plant Biology. Annual Reviews 4139 El Camino Way, P.O. Box 10139, Palo Alto, CA 94303-0139, USA, 53(1), 1–25. https://doi.org/10.1146/annurev.arplant.53.091201.142547.
Berry, E. A., et al. (2000). Structure and function of cytochrome bc complexes. Available at: http://www.life.illinois.edu/crofts/pdf_files/ARB_review.pdf. Accessed 30 Aug 2019.
Berry, E. A., et al. (2004). X-Ray structure of rhodobacter capsulatus cytochrome bc 1: Comparison with its mitochondrial and chloroplast counterparts. Photosynthesis Research, 81(3), 251–275. https://doi.org/10.1023/B:PRES.0000036888.18223.0e.
Blankenship, R. E. (2014). Molecular mechanisms of photosynthesis Robert E. Blankenship (2nd ed.). St Louis: Wiley Blackwell.
Boonstra, A. F., Germeroth, L., & Boekema, E. J. (1994). Structure of the light harvesting antenna from Rhodospirillum molischianum studied by electron microscopy. Biochimica et Biophysica Acta (BBA) – Bioenergetics. Elsevier, 1184(2–3), 227–234. https://doi.org/10.1016/0005-2728(94)90227-5.
Boyer, P. D. (1993). The binding change mechanism for ATP synthase — Some probabilities and possibilities. Biochimica et Biophysica Acta (BBA) – Bioenergetics. Elsevier, 1140(3), 215–250. https://doi.org/10.1016/0005-2728(93)90063-L.
Boyer, P. D., Cross, R. L., & Momsen, W. (1973). A new concept for energy coupling in oxidative phosphorylation based on a molecular explanation of the oxygen exchange reactions. Proceedings of the National Academy of Sciences of the United States of America. National Academy of Sciences, 70(10), 2837–2839. https://doi.org/10.1073/pnas.70.10.2837.
Brune, D. (2004) Advances in photosynthesis. Anoxygenic photosynthetic Bacteria. Available at: https://springerlink.bibliotecabuap.elogim.com/content/pdf/10.1007%2F0-306-47954-0.pdf. Accessed 29 Aug 2019.
Brunisholz, R. A., & Zuber, H. (1992). Structure, function and organization of antenna polypeptides and antenna complexes from the three families of Rhodospirillaneae. Journal of Photochemistry and Photobiology, B: Biology. https://doi.org/10.1016/1011-1344(92)87010-7.
Brunisholz, R. A., et al. (1985). The light-harvesting polypeptides of Rhodopseudomonas viridis. The complete amino-acid sequences of B1015-alpha, B1015-beta and B1015-gamma. Biological Chemistry Hoppe-Seyler, 366(1), 87–98. https://doi.org/10.1515/BCHM3.1985.366.1.87.
Busselez, J., et al. (2007). Structural basis for the PufX-mediated dimerization of bacterial photosynthetic Core complexes. Structure, 15(12), 1674–1683. https://doi.org/10.1016/j.str.2007.09.026.
Bustamante, P. L., & Loach, P. A. (1994). Reconstitution of a functional photosynthetic receptor complex with isolated subunits of Core light-harvesting complex and reaction centers. Biochemistry, 33(45), 13329–13339. https://doi.org/10.1021/bi00249a020.
Bylina, E. J., & Youvan, D. C. (1988). Directed mutations affecting spectroscopic and electron transfer properties of the primary donor in the photosynthetic reaction center. Proceedings of the National Academy of Sciences of the United States of America. National Academy of Sciences, 85(19), 7226–7230. https://doi.org/10.1073/pnas.85.19.7226.
Caffrey, M., et al. (1992). Cytochrome c2 mutants of Rhodobacter capsulatus. Archives of Biochemistry and Biophysics, 292(2), 419–426. https://doi.org/10.1016/0003-9861(92)90011-k.
Camara-Artigas, A., Brune, D., & Allen, J. P. (2002). Interactions between lipids and bacterial reaction centers determined by protein crystallography. Proceedings of the National Academy of Sciences of the United States of America, 99(17), 11055–11060. https://doi.org/10.1073/pnas.162368399.
Capaldi, R. A., et al. (1996). Structural changes in the γ and ε subunits of the Escherichia coli F1F0-type ATPase during energy coupling. Journal of Bioenergetics and Biomembranes. Kluwer Academic Publishers-Plenum Publishers, 28(5), 397–401. https://doi.org/10.1007/BF02113980.
Cartron, M. L., et al. (2014). Integration of energy and electron transfer processes in the photosynthetic membrane of Rhodobacter sphaeroides. Biochimica et Biophysica Acta (BBA) – Bioenergetics, 1837(10), 1769–1780. https://doi.org/10.1016/j.bbabio.2014.02.003.
Chang, C.-H., et al. (1986). Structure of Rhodopseudomonas sphaeroides R-26 reaction center.
Chang, M. C., et al. (1990). Spectroscopic characterization of the light-harvesting complex of Rhodospirillum rubrum and its structural subunit. Biochemistry. American Chemical Society, 29(2), 421–429. https://doi.org/10.1021/bi00454a017.
Cogdell, R. J., et al. (1983). The isolation and partial characterisation of the light-harvesting pigment–protein complement of Rhodopseudomonas acidophila. BBA – Bioenergetics, 722(3), 427–435. https://doi.org/10.1016/0005-2728(83)90058-0.
Cogdell, R. J., et al. (1990). Isolation and characterisation of an unusual antenna complex from the marine purple sulphur photosynthetic bacterium Chromatium purpuratum BN5500. Biochimica et Biophysica Acta (BBA) – Bioenergetics. Elsevier, 1019(3), 239–244. https://doi.org/10.1016/0005-2728(90)90199-E.
Cogdell, R. J., et al. (1999). How photosynthetic bacteria harvest solar energy. Journal of bacteriology. American Society for Microbiology (ASM), 181(13), 3869–3879. Available at: http://www.ncbi.nlm.nih.gov/pubmed/10383951. Accessed 6 Sept 2019.
Crofts, A. R. (2004). The cytochrome BC 1 complex: Function in the context of structure. Annual Review of Physiology, 66, 689–733. https://doi.org/10.1146/annurev.physiol.66.032102.150251.
Crofts, A. R. (2005). The Q-cycle — A personal perspective. In Discoveries in photosynthesis (pp. 479–499). Berlin/Heidelberg: Springer-Verlag. https://doi.org/10.1007/1-4020-3324-9_46.
Cross, R. L. (2000). The rotary binding change mechanism of ATP synthases. Biochimica et Biophysica Acta (BBA) – Bioenergetics. Elsevier, 1458(2–3), 270–275. https://doi.org/10.1016/S0005-2728(00)00079-7.
Crouch, L. I., & Jones, M. R. (2012). Cross-species investigation of the functions of the Rhodobacter PufX polypeptide and the composition of the RC–LH1 core complex. Biochimica et Biophysica Acta (BBA) – Bioenergetics, 1817(2), 336–352. https://doi.org/10.1016/j.bbabio.2011.10.009.
Darrouzet, E., Cooley, J. W., & Daldal, F. (2004). The cytochrome bc 1 complex and its homologue the b 6 f complex: Similarities and differences. Photosynthesis Research, 79(1), 25–44. https://doi.org/10.1023/B:PRES.0000011926.47778.4e.
Davis, C. M., et al. (1997). Evaluation of structure-function relationships in the core light- harvesting complex of photosynthetic bacteria by reconstitution with mutant polypeptides. Biochemistry. American Chemical Society, 36(12), 3671–3679. https://doi.org/10.1021/bi962386p.
De Meur, Q., et al. (2018). Genetic plasticity and ethylmalonyl coenzyme a pathway during acetate assimilation in Rhodospirillum rubrum S1H under photoheterotrophic conditions. Applied and Environmental Microbiology. American Society for Microbiology, 84(3), e02038-17. https://doi.org/10.1128/AEM.02038-17.
DeHoff, B. S., et al. (1988). In vivo analysis of puf operon expression in Rhodobacter sphaeroides after deletion of a putative intercistronic transcription terminator. Journal of Bacteriology, 170(10), 4681–4692. https://doi.org/10.1128/jb.170.10.4681-4692.1988.
Deisenhofer, J., & Michel, H. (1988). The photosynthetic reaction center from the purple bacterium rhodopseudomonas viridis. Available at: http://science.sciencemag.org/. Accessed 4 July 2019.
Deisenhofer, J. et al. (1985). Structure of the protein subunits in the photosynthetic reaction center of. Nature Available at: https://www.nature.com/articles/318618a0.pdf. Accessed 3 July 2019.
Drepper, F., Dorlet, P. A., & Mathis, P. (1997). Cross-linked electron transfer complex between cytochrome c2 and the photosynthetic reaction center of Rhodobacter sphaeroides†. American Chemical Society. https://doi.org/10.1021/BI961350U.
Duncan, T. M., et al. (1995). Rotation of subunits during catalysis by Escherichia coli F1-ATPase. Proceedings of the National Academy of Sciences of the United States of America. National Academy of Sciences, 92(24), 10964–10968. https://doi.org/10.1073/pnas.92.24.10964.
Ehrenreich, A., & Widdel, F. (1994). Anaerobic oxidation of ferrous Iron by purple bacteria, a new type of phototrophic metabolism, applied and environmental microbiology. Available at: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC202013/pdf/aem00029-0311.pdf. Accessed 29 Aug 2019.
Engelhardt, H., Engel, A., & Baumeister, W. (1986). Stoichiometric model of the photosynthetic unit of Ectothiorhodospira halochloris. Proceedings of the National Academy of Sciences, 83(23), 8972–8976. https://doi.org/10.1073/pnas.83.23.8972.
Evans, M. B., Hawthornthwaite, A. M., & Cogdell, R. J. (1990). Isolation and characterisation of the different B800–850 light-harvesting complexes from low- and high-light grown cells of Rhodopseudomonas palustris, strain 2.1.6. Biochimica et Biophysica Acta (BBA) – Bioenergetics, 1016(1), 71–76. https://doi.org/10.1016/0005-2728(90)90008-R.
Fajer, J. et al. (1975). Primary charge separation in bacterial photosynthesis: Oxidized chlorophylls and reduced pheophytin. Proceedings of the National Academy of Sciences. National Academy of Sciences, 72(12), 4956–4960. https://doi.org/10.1073/PNAS.72.12.4956.
Farchaus, J. W., Gruenberg, H., & Oesterhelt, D. (1990) Complementation of a reaction Center-deficient Rhodobacter sphaeroides pufLMX deletion strain in trans with pufBALM does not restore the photosynthesis-positive phenotype. Journal of Bacteriology. Available at: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC208526/pdf/jbacter01044-0475.pdf. Accessed 29 Aug 2019.
Fotiadis, D. et al. (2003) Structural analysis of the RC–LH1 photosynthetic core complex of Rhodospirillum rubrum using atomic force microscopy downloaded from. JBC Papers in Press. Available at: http://www.jbc.org/. Accessed 20 Feb 2019.
Fowler, G. J. S., et al. (1994). Blue shifts in bacteriochlorophyll absorbance correlate with changed hydrogen bonding patterns in light-harvesting 2 mutants of Rhodobacter sphaeroides with alterations at α-Tyr-44 and α-Tyr-45: Figure 2. Biochemical Journal, 299(3), 695–700. https://doi.org/10.1042/bj2990695.
Fowler, G. J. S., et al. (1997). The role of βArg − 10 in the B800 Bacteriochlorophyll and carotenoid pigment environment within the light-harvesting LH2 complex of Rhodobacter sphaeroides †. Biochemistry, 36(37), 11282–11291. https://doi.org/10.1021/bi9626315.
Francia, F., et al. (1999). The reaction Center−LH1 antenna complex of Rhodobacter sphaeroides contains one PufX molecule which is involved in dimerization of this complex. American Chemical Society. https://doi.org/10.1021/BI982891H.
Frese, R. N., et al. (2004). The long-range organization of a native photosynthetic membrane. Proceedings of the National Academy of Sciences of the United States of America, 101(52), 17994–17999. https://doi.org/10.1073/pnas.0407295102.
Frigaard, N. U., & Dahl, C. (2008). Sulfur metabolism in phototrophic Sulfur bacteria. Advances in Microbial Physiology. https://doi.org/10.1016/S0065-2911(08)00002-7.
Fuchs, B. M., et al. (2007). Characterization of a marine gammaproteobacterium capable of aerobic anoxygenic photosynthesis. Proceedings of the National Academy of Sciences, 104(8), 2891–2896. https://doi.org/10.1073/pnas.0608046104.
Gall, A., et al. (1997). Influence of the protein binding site on the absorption properties of the monomeric Bacteriochlorophyll in Rhodobacter sphaeroides LH2 complex†. American Chemical Society. https://doi.org/10.1021/BI9717237.
Gall, A., et al. (1999). Certain species of the Proteobacteria possess unusual bacteriochlorophyll environments in their light-harvesting proteins. Biospectroscopy, 5(6), 338–345. https://doi.org/10.1002/(SICI)1520-6343(1999)5:6<338::AID-BSPY3>3.0.CO;2-D.
Gardiner, A. T., Cogdell, R. J., & Takaichi, S. (1993). The effect of growth conditions on the light-harvesting apparatus in Rhodopseudomonas acidophila, Photosynthesis Research. Kluwer Academic Publishers. Available at: https://springerlink.bibliotecabuap.elogim.com/content/pdf/10.1007%2FBF00146415.pdf. Accessed 9 July 2019.
Gast, P., et al. (1985). Determination of the amount and the type of quinones present in single crystals from reaction center protein from the photosynthetic bacterium Rhodopseudomonas viridis. FEBS Letters. https://doi.org/10.1016/0014-5793(85)80544-5.
Gonçalves, R. P., et al. (2005). Architecture of the native photosynthetic apparatus of Phaeospirillum molischianum. Journal of Structural Biology, 152(3), 221–228. https://doi.org/10.1016/j.jsb.2005.10.002.
Gubellini, F., et al. (2006). Functional and structural analysis of the photosynthetic apparatus of Rhodobacter veldkampii †. Biochemistry, 45(35), 10512–10520. https://doi.org/10.1021/bi0610000.
Guo, H., Suzuki, T., & Rubinstein, J. L. (2019). Structure of a bacterial ATP synthase. eLife, 8. https://doi.org/10.7554/eLife.43128.
Gupta, R. S., & Khadka, B. (2016). Evidence for the presence of key chlorophyll-biosynthesis-related proteins in the genus Rubrobacter (Phylum Actinobacteria) and its implications for the evolution and origin of photosynthesis. Photosynthesis Research, 127(2), 201–218. https://doi.org/10.1007/s11120-015-0177-y.
Hansen, T. A., & Gemerden, H. (1972). Sulfide utilization by purple nonsulfur bacteria. Archiv für Mikrobiologie. Springer, 86(1), 49–56. https://doi.org/10.1007/BF00412399.
Hartigan, N., et al. (2002). The 7.5-Å electron density and spectroscopic properties of a novel low-light B800 LH2 from Rhodopseudomonas palustris. Biophysical Journal. https://doi.org/10.1016/S0006-3495(02)75456-8.
Hatch, L. P., Cox, G. B., & Howitt, S. M. (1995). The essential arginine residue at position 210 in the alpha subunit of the Escherichia coli ATP synthase can be transferred to position 252 with partial retention of activity. The Journal of biological chemistry. American Society for Biochemistry and Molecular Biology, 270(49), 29407–29412. https://doi.org/10.1074/jbc.270.49.29407.
Heldt, H.-W., & Piechulla, B. (2011). Plant biochemistry. Academic.
Holden-Dye, K., Crouch, L. I., & Jones, M. R. (2008). Structure, function and interactions of the PufX protein. Biochimica et Biophysica Acta, 1777, 613–630. https://doi.org/10.1016/j.bbabio.2008.04.015.
Hu, X., et al. (1998). Architecture and mechanism of the light-harvesting apparatus of purple bacteria. Computational Biomolecular Science, 95, 5935–5941.
Hu, X., et al. (2002). Photosynthetic apparatus of purple bacteria. Quarterly Reviews of Biophysics. Cambridge University Press, 35, 1–62. https://doi.org/10.1017/S0033583501003754.
Hunter, C., et al. (2008). The purple phototrophic bacteria. Dordrecht: Springer.
Imanishi, M., et al. (2019). A dual role for ca 2+ in expanding the spectral diversity and stability of light-harvesting 1 reaction Center Photocomplexes of purple phototrophic Bacteria. Biochemistry. American Chemical Society, 58(25), 2844–2852. https://doi.org/10.1021/acs.biochem.9b00351.
Imhoff, J. F., Hiraishi, A., & Süling, J. (2005) Anoxygenic phototrophic purple bacteria, Bergey’s manual® of systematic bacteriology. https://doi.org/10.1007/0-387-28021-9_15.
Jackson, P. J., et al. (2018). Identification of protein W, the elusive sixth subunit of the Rhodopseudomonas palustris reaction center-light harvesting 1 core complex. Biochimica et Biophysica Acta (BBA) – Bioenergetics. Elsevier, 1859(2), 119–128. https://doi.org/10.1016/j.bbabio.2017.11.001.
Jamieson, S. J., et al. (2002). Projection structure of the photosynthetic reaction centre-antenna complex of Rhodospirillum rubrum at 8.5?? resolution. EMBO Journal. https://doi.org/10.1093/emboj/cdf410.
Janosi, L., et al. (2006). Influence of subunit structure on the oligomerization state of light-harvesting complexes: A free energy calculation study. Chemical Physics. https://doi.org/10.1016/j.chemphys.2005.08.038.
Jault, J., & Allison, W. (1993). Slow binding of ATP to noncatalytic nucleotide binding sites which accelerates catalysis is responsible for apparent negative cooperativity exhibit... – PubMed – NCBI, The Journal of Biological Chemistry. Available at: https://www.ncbi.nlm.nih.gov/pubmed/8420930. Accessed 30 Aug 2019.
Jault, J.-M. et al. (1995). Complex of the Fi-ATPase from thermophilic Bacillus PS 3 containing the CX.D261N substitution fails to dissociate inhibitory MgADP from a catalytic site when ATP binds to noncatalytic sites1", Biochemistry. Available at: https://pubs.acs.org/sharingguidelines. Accessed 30 Aug 2019.
Jay, F., Lambillotte, M., & Mühlethaler, K. (1983). Localisation of Rhodopseudomonas viridis reaction centre and light harvesting proteins using ferritin-antibody labelling. European Journal of Cell Biology, 30(1), 1–8. Available at: http://www.ncbi.nlm.nih.gov/pubmed/6189715. Accessed 18 Sept 2019.
Jenney, F. E., & Daldal, F. (1993). A novel membrane-associated c-type cytochrome, cyt cy, can mediate the photosynthetic growth of Rhodobacter capsulatus and Rhodobacter sphaeroides. The EMBO Journal. European Molecular Biology Organization, 12(4), 1283–1292. https://doi.org/10.1002/J.1460-2075.1993.TB05773.X.
Jones, M. R. (2009). The petite purple photosynthetic powerpack. Biochemical Society Transactions, 37(2), 400–407. https://doi.org/10.1042/BST0370400.
Jungas, C., et al. (1999). Supramolecular organization of the photosynthetic apparatus of Rhodobacter sphaeroides. The EMBO Journal, 18(3), 534–542.
Junge, W., Sabber, D., & Engelbrecht, S. (1996). ATP-synthesis. Rotatory catalysis by F-ATPase: Real-time recording of intersubunit rotation. Berichte der Bunsengesellschaft für physikalische Chemie. John Wiley & Sons, Ltd, 100(12), 2014–2019. https://doi.org/10.1002/bbpc.19961001215.
Junge, W., Lill, H., & Engelbrecht, S. (1997). ATP synthase: An electrochemical ransducer with rotatory mechanics Trends in Biochemical Sciences. Elsevier Current Trends, 22(11), 420–423. https://doi.org/10.1016/S0968-0004(97)01129-8.
Junge, W., Sielaff, H., & Engelbrecht, S. (2009). Torque generation and elastic power transmission in the rotary FOF1-ATPase. Nature, 459(7245), 364–370. https://doi.org/10.1038/nature08145.
Kayalar, C., et al. (1977). An alternating site sequence for oxidative phosphorylation suggested by measurement of substrate binding patterns and exchange reaction inhibitions∗. The Journal of Biological Chemistry. Available at: http://www.jbc.org/. Accessed 30 Aug 2019.
Kehoe, J. W., et al. (1998). Reconstitution of Core light-harvesting complexes of photosynthetic Bacteria using chemically synthesized polypeptides. 2. Determination of structural features that stabilize complex formation and their implications for the structure of the subunit complex †. Biochemistry, 37(10), 3418–3428. https://doi.org/10.1021/bi9722709.
Kellogg, E. C., et al. (1989). Measurement of the extent of electron transfer to the bacteriopheophytin in the M-subunit in reaction centers of Rhodopseudomonas viridis. Photosynthesis Research. Kluwer Academic Publishers, 22(1), 47–59. https://doi.org/10.1007/BF00114766.
Kinosita, K., Adachi, K., & Itoh, H. (2004). Rotation of F 1 -ATPase: How an ATP-driven molecular machine may work. Annual Review of Biophysics and Biomolecular Structure, 33(1), 245–268. https://doi.org/10.1146/annurev.biophys.33.110502.132716.
Kirmaier, C., Holten, D., & Parson, W. W. (1985). Picosecond-photodichroism studies of the transient states in Rhodopseudomonas sphaeroides reaction centers at 5 K. effects of electron transfer on the six bacteriochlorin pigments. Biochimica et Biophysica Acta (BBA) – Bioenergetics. Elsevier, 810(1), 49–61. https://doi.org/10.1016/0005-2728(85)90205-1.
Klug, G. et al. (1988). Pleiotropic effects of localized Rhodobacter capsulatus puf operon deletions on production of light-absorbing pigment–protein complexes. Journal of Bacteriology. Available at: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC211687/pdf/jbacter00190-0422.pdf. Accessed 29 Aug 2019.
Koblízek, M., et al. (2005). Sequential assembly of photosynthetic units in Rhodobacter sphaeroides as revealed by fast repetition rate analysis of variable bacteriochlorophyll a fluorescence. Biochimica et Biophysica Acta (BBA) – Bioenergetics, 1706(3), 220–231. https://doi.org/10.1016/j.bbabio.2004.11.004.
Koepke, J., et al. (1996). The crystal structure of the light-harvesting complex II (B800-850) from Rhodospirillum molischianum. Structure, 4(5), 581–597. https://doi.org/10.1016/S0969-2126(96)00063-9.
Kolber, Z. S. et al. (2001). Contribution of aerobic photoheterotrophic bacteria to the carbon cycle in the ocean. Science (New York, N.Y.). American Association for the Advancement of Science, 292(5526), 2492–2495. https://doi.org/10.1126/science.1059707.
Kumar, S., et al. (2016). Direct imaging of protein organization in an intact bacterial organelle using high-resolution atomic force microscopy. https://doi.org/10.1021/acsnano.6b05647.
Lang, H. P., & Hunter, C. N. (1994). The relationship between carotenoid biosynthesis and the assembly of the light-harvesting LH2 complex in Rhodobacter sphaeroides. Biochemical Journal, 298(1), 197–205. https://doi.org/10.1042/bj2980197.
LaSarre, B., et al. (2018). Restricted localization of photosynthetic intracytoplasmic membranes (ICMs) in multiple genera of purple nonsulfur bacteria. mBio, 9(4), 780–798. https://doi.org/10.1128/mBio.00780-18.
Lilburn, T. G., et al. (1992). Pleiotropic effects of pufX gene deletion on the structure and function of the photosynthetic apparatus of Rhodobacter capsulatus. Biochimica et Biophysica Acta (BBA) – Bioenergetics, 1100(2), 160–170. https://doi.org/10.1016/0005-2728(92)90077-F.
Liu, L.-N. (2016). Distribution and dynamics of electron transport complexes in cyanobacterial thylakoid membranes. Biochimica et Biophysica Acta. Elsevier, 1857(3), 256–265. https://doi.org/10.1016/j.bbabio.2015.11.010.
Liu, L.-N., & Scheuring, S. (2013). Investigation of photosynthetic membrane structure using atomic force microscopy. Trends in Plant Science, 18, 277–286. https://doi.org/10.1016/j.tplants.2013.03.001.
Liu, L.-N., et al. (2009). Quinone pathways in entire photosynthetic Chromatophores of Rhodospirillum photometricum. Journal of Molecular Biology, 393, 27–35. https://doi.org/10.1016/j.jmb.2009.07.044.
Liu, L.-N., et al. (2011a). Forces guiding assembly of light-harvesting complex 2 in native membranes. Proceedings of the National Academy of Sciences, 108(23), 9455–9459. https://doi.org/10.1073/pnas.1004205108.
Liu, L. N., Sturgis, J. N., & Scheuring, S. (2011b). Native architecture of the photosynthetic membrane from Rhodobacter veldkampii. Journal of Structural Biology. https://doi.org/10.1016/j.jsb.2010.08.010.
Loach, P. A., & Parkes-Loach, P. S. (1995). Structure-function relationships in Core Light-Harvesting Complexes (LHI) as determined by characterization of the structural subunit and by reconstitution experiments. In Anoxygenic photosynthetic bacteria (pp. 437–471). Dordrecht: Kluwer Academic Publishers. https://doi.org/10.1007/0-306-47954-0_21.
Madigan, M. T. (2003). Anoxygenic phototrophic bacteria from extreme environments, Photosynthesis Research. Available at: http://www.life.illinois.edu/govindjee/Part2/14_Madigan.pdf. Accessed 29 Aug 2019.
Mathis, P., Ortega, J. M., & Venturoli, G. (1994). Interaction between cytochrome c and the photosynthetic reaction center of purple bacteria: Behaviour at low temperature. Biochimie, 76(6), 569–579. https://doi.org/10.1016/0300-9084(94)90181-3.
Matsui, T., et al. (1997) Catalytic activity of the 3 3 complex of F 1-ATPase without Noncatalytic Nucleotide Binding Site*. Available at: http://www-jbc.stanford.edu/jbc/. Accessed 30 Aug 2019.
McDermott, G., et al. (1995). Crystal structure of an integral membrane light-harvesting complex from photosynthetic bacteria. Nature. https://doi.org/10.1038/374517a0.
Mcluskey, K., et al. (2001). The crystallographic structure of the B800-820 LH3 light-harvesting complex from the purple bacteria Rhodopseudomonas acidophila strain 7050 †. https://doi.org/10.1021/bi010309a.
Menin, L., et al. (1998). Role of HiPIP as electron donor to the RC-bound cytochrome in photosynthetic purple bacteria. Photosynthesis Research. Kluwer Academic Publishers, 55(2/3), 343–348. https://doi.org/10.1023/A:1005989900756.
Milgrom, Y. M., Ehler, L. L., & Boyer, P. D. (1991). The characteristics and effect on catalysis of nucleotide binding to noncatalytic sites of chloroplast F1-ATPase. The Journal of Biological Chemistry, 266(18), 11551–11558. Available at: http://www.ncbi.nlm.nih.gov/pubmed/1828802. Accessed 30 Aug 2019.
Miller, J. F., et al. (1987). Isolation and characterization of a subunit form of the light-harvesting complex of Rhodospirillum rubrum. Biochemistry. American Chemical Society, 26(16), 5055–5062. https://doi.org/10.1021/bi00390a026.
Mothersole, D. J., et al. (2016). PucC and LhaA direct efficient assembly of the light-harvesting complexes in Rhodobacter sphaeroides. Molecular Microbiology. https://doi.org/10.1111/mmi.13235.
Munk, A. C., et al. (2011). Complete genome sequence of Rhodospirillum rubrum type strain (S1T). Standards in Genomic Sciences, 4(3), 293–302. https://doi.org/10.4056/sigs.1804360.
Myllykallio, H., et al. (1998). Membrane-anchored cytochrome c y mediated microsecond time range Electron transfer from the cytochrome bc 1 complex to the reaction Center in Rhodobacter capsulatus †. Biochemistry, 37(16), 5501–5510. https://doi.org/10.1021/bi973123d.
Niedzwiedzki, D. M., et al. (2011). Energy transfer in an LH4-like light harvesting complex from the aerobic purple photosynthetic bacterium Roseobacter denitrificans. Biochimica et Biophysica Acta (BBA) – Bioenergetics, 1807(5), 518–528. https://doi.org/10.1016/j.bbabio.2011.03.004.
Niwa, S., et al. (2014). Structure of the LH1–RC complex from Thermochromatium tepidum at 3.0 Å. Nature, 508(7495), 228–232. https://doi.org/10.1038/nature13197.
Nogi, T., Hirano, Y., & Miki, K. (2005). Structural and functional studies on the tetraheme cytochrome subunit and its electron donor proteins: The possible docking mechanisms during the electron transfer reaction. Photosynthesis Research. https://doi.org/10.1007/s11120-004-2416-5.
Noji, H., et al. (1997). Direct observation of the rotation of F1-ATPase. Nature, 386(6622), 299–302. https://doi.org/10.1038/386299a0.
Nottoli, M., et al. (2018). The role of charge-transfer states in the spectral tuning of antenna complexes of purple bacteria. Photosynthesis Research. Springer Netherlands, 137(2), 215–226. https://doi.org/10.1007/s11120-018-0492-1.
Olsen, J. D., et al. (1997). Site-directed modification of the ligands to the bacteriochlorophylls of the light-harvesting LH1 and LH2 complexes of Rhodobacter sphaeroides†. Biochemistry. American Chemical Society, 36, 12625–12632. https://doi.org/10.1021/BI9710481.
Ortega, J. M., Drepper, F., & Mathis, P. (1999). Electron transfer between cytochrome c(2) and the tetraheme cytochrome c in Rhodopseudomonas viridis. Photosynthesis Research, 59(2–3), 147–157. https://doi.org/10.1023/A:1006149621029.
Pandit, A. et al. (2003). Investigations of intermediates appearing in the reassociation of the light-harvesting 1 complex of Rhodospirillum rubrum. Photosynthesis Research, 3(75), 235–248. Available at: https://www.ncbi.nlm.nih.gov/pubmed/16228604. Accessed 6 Sept 2019.
Pänke, O., et al. (2000). F-ATPase: Specific observation of the rotating c subunit oligomer of EF(o)EF(1). FEBS Letters, 472(1), 34–38. https://doi.org/10.1016/s0014-5793(00)01436-8.
Papiz, M. Z., et al. (2003). The structure and thermal motion of the B800-850 LH2 complex from Rps. acidophila at 2.0 Å resolution and 100 K: New structural features and functionally relevant motions. Journal of Molecular Biology. https://doi.org/10.1016/S0022-2836(03)00024-X.
Parkes-Loach, P. S., et al. (2000). Articles role of the Core region of the PufX protein in inhibition of reconstitution of the Core light-harvesting complexes of Rhodobacter sphaeroides and Rhodobacter capsulatus †. https://doi.org/10.1021/bi002580i.
Parkes-Loach, P. S., et al. (2004). Interactions stabilizing the structure of the Core light-harvesting complex (LH1) of photosynthetic Bacteria and its subunit (B820) †. Biochemistry, 43(22), 7003–7016. https://doi.org/10.1021/bi049798f.
Pogoryelov, D., et al. (2009). High-resolution structure of the rotor ring of a proton-dependent ATP synthase. Nature Structural & Molecular Biology, 16(10), 1068–1073. https://doi.org/10.1038/nsmb.1678.
Polívka, T., & Frank, H. A. (2010). Molecular factors controlling photosynthetic light harvesting by carotenoids. Accounts of Chemical Research. American Chemical Society, 43(8), 1125–1134. https://doi.org/10.1021/ar100030m.
Prince, S. M., et al. (1997). Apoprotein structure in the LH2 complex from Rhodopseudomonas acidophila strain 10050: Modular assembly and protein pigment interactions. Journal of Molecular Biology. https://doi.org/10.1006/jmbi.1997.0966.
Pugh, R. J. et al. (1998). The LH1–RC core complex of Rhodobacter sphaeroides: Interaction between components, time-dependent assembly, and topology of the PufX protein. Biochimica et Biophysica Acta (BBA) – Bioenergetics. Elsevier, 1366(3), 301–316. https://doi.org/10.1016/S0005-2728(98)00131-5.
Pullerits, T., & Sundström, V. (1996). Photosynthetic light-harvesting pigment−protein complexes: Toward understanding how and why. American Chemical Society. https://doi.org/10.1021/AR950110O.
Qian, P. (2017). Structure and function of the reaction centre – Light harvesting 1 Core complexes from purple photosynthetic Bacteria. In Photosynthesis: Structures, mechanisms, and applications (pp. 11–31). Cham: Springer International Publishing. https://doi.org/10.1007/978-3-319-48873-8_2.
Qian, P., Hunter, C. N., & Bullough, P. A. (2005). The 8.5 Å projection structure of the core RC–LH1-PufX dimer of Rhodobacter sphaeroides. Journal of Molecular Biology. https://doi.org/10.1016/j.jmb.2005.04.032.
Qian, P., Bullough, P. A., & Hunter, C. N. (2008). Three-dimensional reconstruction of a membrane-bending complex: The RC–LH1-PufX core dimer of rhodobacter sphaeroides. Journal of Biological Chemistry. https://doi.org/10.1074/jbc.M800625200.
Qian, P., et al. (2013). Three-dimensional structure of the Rhodobacter sphaeroides RC–LH1-PufX complex: Dimerization and quinone channels promoted by PufX. Biochemistry, 52(43), 7575–7585. https://doi.org/10.1021/bi4011946.
Qian, P., et al. (2018). Cryo-EM structure of the Blastochloris viridis LH1-RC complex at 2.9 Å. Nature, 556(7700), 203–208. https://doi.org/10.1038/s41586-018-0014-5.
Rastogi, V. K., & Girvin, M. E. (1999). Structural changes linked to proton translocation by subunit c of the ATP synthase. Nature, 402(6759), 263–268. https://doi.org/10.1038/46224.
Rathgeber, C., Beatty, J. T., & Yurkov, V. (2004). Aerobic phototrophic bacteria: New evidence for the diversity, ecological importance and applied potential of this previously overlooked group. Photosynthesis Research. https://doi.org/10.1023/B:PRES.0000035036.49977.bc.
Rieske, J. S., MacLennan, D. H., & Coleman, R. (1964). Isolation and properties of an iron-protein from the (reduced coenzyme Q)-cytochrome C reductase complex of the respiratory chain. Biochemical and Biophysical Research Communications. https://doi.org/10.1016/0006-291X(64)90171-8.
Rosen, D., et al. (1983). Interaction of cytochrome c with reaction centers of Rhodopseudomonas sphaeroides R-26: Localization of the binding site by chemical crosslinking and immunochemical studies. Biochemistry, 22(2), 335–341. https://doi.org/10.1021/bi00271a016.
Roszak, A. W. et al. (2003). Crystal structure of the RC–LH1 core complex from Rhodopseudomonas palustris. Science, 302. Available at: http://science.sciencemag.org/. Accessed 22 July 2019.
Roszak, A. W., et al. (2012). New insights into the structure of the reaction Centre from Blastochloris viridis: Evolution in the laboratory. Biochemical Journal, 442, 27–37. https://doi.org/10.1042/BJ20111540.
Sarıbaş, S., et al. (1998). Interactions between the cytochrome b, cytochrome c1, and Fe−S protein subunits at the Ubihydroquinone oxidation site of the bc1 complex of Rhodobacter capsulatus†. American Chemical Society. https://doi.org/10.1021/BI973146S.
Scheuring, S., & Sturgis, J. N. (2005). Chromatic adaptation of photosynthetic membranes. Science, 309(5733), 484–487. https://doi.org/10.1126/science.1110879.
Scheuring, S., & Sturgis, J. N. (2009). Atomic force microscopy of the bacterial photosynthetic apparatus: Plain pictures of an elaborate machinery. Photosynthesis Research, 102(2), 197–211. https://doi.org/10.1007/s11120-009-9413-7.
Scheuring, S., Seguin, J, et al. (2003a). Nanodissection and high-resolution imaging of the Rhodopseudomonas viridis photosynthetic core complex in native membranes by AFM. Atomic force microscopy. Proceedings of the National Academy of Sciences of the United States of America. National Academy of Sciences, 100(4), 1690–1693. https://doi.org/10.1073/pnas.0437992100.
Scheuring, S., Seguin, J., et al. (2003b). AFM characterization of tilt and intrinsic flexibility of Rhodobacter sphaeroides light harvesting complex 2 (LH2). Journal of Molecular Biology, 325(3), 569–580. https://doi.org/10.1016/s0022-2836(02)01241-x.
Scheuring, S., et al. (2004a). Watching the photosynthetic apparatus in native membranes. Sciences-New York, 101(31), 11293–112967.
Scheuring, S., Rigaud, J. L., & Sturgis, J. N. (2004b). Variable LH2 stoichiometry and core clustering in native membranes of Rhodospirillum photometricum. EMBO Journal, 23(21), 4127–4133. https://doi.org/10.1038/sj.emboj.7600429.
Scheuring, S., Busselez, J., & Lévy, D. (2005a). Structure of the dimeric PufX-containing Core complex of Rhodobacter blasticus by in Situ atomic force microscopy. Journal of Biological Chemistry, 280(2), 1426–1431. https://doi.org/10.1074/jbc.M411334200.
Scheuring, S., Lévy, D., & Rigaud, J.-L. (2005b). Watching the components of photosynthetic bacterial membranes and their in situ organisation by atomic force microscopy. Biochimica et Biophysica Acta (BBA) – Biomembranes. Elsevier, 1712(2), 109–127. https://doi.org/10.1016/J.BBAMEM.2005.04.005.
Scheuring, S., et al. (2006). The photosynthetic apparatus of Rhodopseudomonas palustris: Structures and organization. Journal of Molecular Biology, 358(1), 83–96. https://doi.org/10.1016/j.jmb.2006.01.085.
Scheuring, S., Boudier, T., & Sturgis, J. N. (2007). From high-resolution AFM topographs to atomic models of supramolecular assemblies. Journal of Structural Biology, 159(2 SPEC. ISS), 268–276. https://doi.org/10.1016/j.jsb.2007.01.021.
Scheuring, S., et al. (2014). The architecture of Rhodobacter sphaeroides chromatophores. BBA – Bioenergetics, 1837, 1263–1270. https://doi.org/10.1016/j.bbabio.2014.03.011.
Senior, A. E., Nadanaciva, S., & Weber, J. (2002). The molecular mechanism of ATP synthesis by F1F0-ATP synthase. Biochimica et Biophysica Acta (BBA) – Bioenergetics. Elsevier, 1553(3), 188–211. https://doi.org/10.1016/S0005-2728(02)00185-8.
Sheng, Y. U., & Hearst, J. E. (1986). Regulation of expression of genes for light-harvesting antenna proteins LH-I and LH-II; reaction center polypeptides RC-L, RC-M, and RC-H; and enzymes of bacteriochlorophyll and carotenoid biosynthesis in Rhodobacter capsulatus* by light and oxygen (photosynthetic bacteria/Rhodopseudomonas capsulata/puf operon/polycistronic mRNA/photooxidadve damage), Proceedings of the National Academy of Sciences of the United States of America. Available at: https://www.pnas.org/content/pnas/83/20/7613.full.pdf. Accessed 29 Aug 2019.
Shopes, R. J., & Wraight, C. A. (1985). The acceptor quinone complex of Rhodopseudomonas viridis reaction centers. BBA – Bioenergetics. https://doi.org/10.1016/0005-2728(85)90242-7.
Siebert, C. A., et al. (2004). Molecular architecture of photosynthetic membranes in Rhodobacter sphaeroides: The role of PufX. The EMBO Journal, 23, 690–700. https://doi.org/10.1038/sj.emboj.7600092.
Stark, W., et al. (1984). The structure of the photoreceptor unit of Rhodopseudomonas viridis; The structure of the photoreceptor unit of Rhodopseudomonas viridis. The EMBO Journal, 3(4), 777–783. https://doi.org/10.1002/j.1460-2075.1984.tb01884.x.
Sumi, H. (2002). Uphill energy trapping by reaction center in bacterial photosynthesis: Charge separation unistep from antenna excitation, virtually mediated by special-pair excitation. Journal of Physical Chemistry B. American Chemical Society, 106(51), 13370–13383. https://doi.org/10.1021/JP021716E.
Sundström, V. (2004). Ultrafast dynamics of carotenoid excited states−from solution to natural and artificial systems. American Chemical Society. https://doi.org/10.1021/CR020674N.
Sundstrom, V., & Van Grondelle, R. (1999). Photosynthetic light-harvesting: Reconciling dynamics and structure of purple bacterial LH2 reveals function of photosynthetic unit. Journal of Physical Chemistry B, 103, 2327–2346. https://doi.org/10.1021/jp983722+.
Swingley, W. D., et al. (2007). The complete genome sequence of Roseobacter denitrificans reveals a Mixotrophic rather than photosynthetic metabolism. Journal of Bacteriology, 189(3), 683–690. https://doi.org/10.1128/JB.01390-06.
Tabita, R. (2004). Anoxygenic photosynthetic bacteria. Available at: https://springerlink.bibliotecabuap.elogim.com/content/pdf/10.1007%2F0-306-47954-0.pdf. Accessed 29 Aug 2019.
Takaichi, S. (2000). Characterization of carotenes in a combination of a C18 HPLC column with isocratic elution and absorption spectra with a photodiode-array detector. Photosynthesis Research. Kluwer Academic Publishers, 65(1), 93–99. https://doi.org/10.1023/A:1006445503030.
Tang, K.-H., Tang, Y. J., & Blankenship, R. E. (2011). Carbon metabolic pathways in phototrophic Bacteria and their broader evolutionary implications. Frontiers in Microbiology. Frontiers, 2, 165. https://doi.org/10.3389/fmicb.2011.00165.
Tharia, H. A. et al. (1999). Characterisation of hydrophobic peptides by RP-HPLC from different spectral forms of LH2 isolated from Rps. palustris, Photosynthesis Research. Available at: https://springerlink.bibliotecabuap.elogim.com/content/pdf/10.1023%2FA%3A1006281532327.pdf. Accessed 17 July 2019.
Tiede, D. M., & Dutton, P. L. (1993). Electron transfer between bacterial reaction centers and mobile c-type cytochromes. In Photosynthetic reaction center (pp. 257–288). Academic Press. https://doi.org/10.1016/B978-0-12-208661-8.50014-0.
Todd, J. B., et al. (1998). In vitro reconstitution of the Core and peripheral light-harvesting complexes of Rhodospirillum m olischianum from separately isolated components †. Biochemistry, 37(50), 17458–17468. https://doi.org/10.1021/bi981114e.
Truper, H. G. T., & Fischer, U. (1982). Anaerobic oxidation of Sulphur compounds as electron donors for bacterial photosynthesis (R. Soc. Land. B, Trans.). Available at: https://royalsocietypublishing.org/doi/pdf/10.1098/rstb.1982.0095. Accessed 29 Aug 2019.
Tunnicliffe, R. B., et al. (2006). The solution structure of the PufX polypeptide from Rhodobacter sphaeroides. FEBS Letters, 580(30), 6967–6971. https://doi.org/10.1016/j.febslet.2006.11.065.
van Grondelle, R. et al. (1994). Energy transfer and trapping in photosynthesis. Biochimica et Biophysica Acta (BBA) – Bioenergetics. Elsevier, 1187(1), 1–65. https://doi.org/10.1016/0005-2728(94)90166-X.
van Mourik, F., Visschers, R. W., & van Grondelle, R. (1992). Energy transfer and aggregate size effects in the inhomogeneously broadened core light-harvesting complex of Rhodobacter sphaeroides. Chemical Physics Letters. North-Holland, 193(1–3), 1–7. https://doi.org/10.1016/0009-2614(92)85674-Y.
van Oijen, A. M. et al. (1999). Unraveling the electronic structure of individual photosynthetic pigment–protein complexes. Science (New York, N.Y.). American Association for the Advancement of Science, 285(5426), 400–402. https://doi.org/10.1126/science.285.5426.400.
Vik, S. B., & Antonio, B. J. (1994). A mechanism of proton translocation by F1F0 ATP synthases suggested by double mutants of the a subunit. The Journal of Biological Chemistry, 269(48), 30364–30369. Available at: http://www.ncbi.nlm.nih.gov/pubmed/7982950. Accessed 7 Sept 2019.
Wada, T., et al. (1999). A novel labeling approach supports the five-transmembrane model of subunit a of the Escherichia coli ATP synthase. Journal of Biological Chemistry, 274(24), 17353–17357. https://doi.org/10.1074/jbc.274.24.17353.
Wakao, N. et al. (1996). Discovery of natural photosynthesis using Zn-containing bacteriochlorophyll in an aerobic bacterium Acidiphilium rubrum. Plant and Cell Physiology. Narnia, 37(6), 889–893. https://doi.org/10.1093/oxfordjournals.pcp.a029029.
Weber, J., & Senior, A. E. (2003). ATP synthesis driven by proton transport in F1 F0 -ATP synthase. FEBS Letters. John Wiley & Sons, Ltd, 545(1), 61–70. https://doi.org/10.1016/S0014-5793(03)00394-6.
Weber, J., Wilke-Mounts, S., & Senior, A. E. (1994). Cooperativity and stoichiometry of substrate binding to the catalytic sites of Escherichia coli Fl-ATPase EFFECTS OF MAGNESIUM, INHIBITORS, AND MUTATION*, THE JOURNAL OF BIOIQGICAL CHEMISTRY. Available at: http://www.jbc.org/content/269/32/20462.full.pdf. Accessed 30 Aug 2019.
Weber, J. et al. (1995). α-aspartate 261 is a key residue in noncatalytic sites of Escherichia coli F 1 -ATPase. Journal of Biological Chemistry. American Society for Biochemistry and Molecular Biology, 270(36), 21045–21049. https://doi.org/10.1074/jbc.270.36.21045.
Xia, D., et al. (1997). Crystal structure of the cytochrome bc 1 complex from bovine heart mitochondria. Science, 277(5322), 60–66. https://doi.org/10.1126/science.277.5322.60.
Youvan, D. C. et al. (1984) Reaction center and light-harvesting I genes from Rhodopseudomonas capsulata (photosynthesis/DNA sequence/enhanced fluorescence mutants/R-prime plasmid/genetic map), Proceedings of the National Academy of Sciences of the United States of America. Available at: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC344636/pdf/pnas00602-0203.pdf. Accessed 29 Aug 2019.
Yu, L. J., et al. (2018). Structure of photosynthetic LH1-RC supercomplex at 1.9 Å resolution. Nature, 556(7700), 209–213. https://doi.org/10.1038/s41586-018-0002-9.
Yurkov, V. V, & Beatty, J. T. (1998). Aerobic anoxygenic phototrophic bacteria. Microbiology and Molecular Biology Reviews: MMBR. American Society for Microbiology (ASM), 62(3), 695–724. Available at: http://www.ncbi.nlm.nih.gov/pubmed/9729607. Accessed 29 Aug 2019.
Zeng, Y., & Koblizek, M. (2017). Phototrophic gemmatimonadetes: A new “purple” branch on the bacterial tree of life. In Modern topics in the phototrophic prokaryotes: Environmental and applied aspects (pp. 1–492). https://doi.org/10.1007/978-3-319-46261-5.
Zhang, Z., et al. (1998). Electron transfer by domain movement in cytochrome bc1. Nature, 392(6677), 677–684. https://doi.org/10.1038/33612.
Zheng, Q. et al. (2011) Diverse arrangement of photosynthetic gene clusters in aerobic Anoxygenic phototrophic Bacteria. PLoS One. Edited by S. Bereswill. Public Library of Science, 6(9), e25050. https://doi.org/10.1371/journal.pone.0025050.
Zinth, W., & Wachtveitl, J. (2005). The first picoseconds in bacterial photosynthesis? Ultrafast electron transfer for the efficient conversion of light energy. ChemPhysChem. John Wiley & Sons, Ltd, 6(5), 871–880. https://doi.org/10.1002/cphc.200400458.
Zuber, H., & Cogdell, R. (2004) Advances in photosynthesis. Anoxygenic photosynthetic bacteria. Available at: https://springerlink.bibliotecabuap.elogim.com/content/pdf/10.1007%2F0-306-47954-0.pdf. Accessed 29 Aug 2019.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Miller, L.C., Martin, D.S., Liu, LN., Canniffe, D.P. (2020). Composition, Organisation and Function of Purple Photosynthetic Machinery. In: Wang, Q. (eds) Microbial Photosynthesis. Springer, Singapore. https://doi.org/10.1007/978-981-15-3110-1_4
Download citation
DOI: https://doi.org/10.1007/978-981-15-3110-1_4
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-3109-5
Online ISBN: 978-981-15-3110-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)