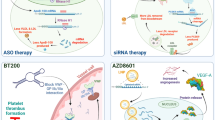
Abstract
Ribonucleic acid (RNA) is being exploited and understood in its many aspects of function and structure for development of valuable tools in the therapeutics of various diseases such as cardiovascular etc. The expanded knowledge regarding function of RNA in the genomics and inside the cell has dramatically changed the therapeutic strategies in the past few years. RNA has become a spotlight of attention for developing novel therapeutic schemes and hence variety of therapeutic strategies is being coming into the picture that includes RNA interference, use of aptamers, role of microRNA (miRNA) that can alter the complex gene expression patterns. It is due to the fact that RNA offers various advantages in disease management as it can be edited and modified in its various forms such as secondary and tertiary structures. Although scientists are in process of manufacturing RNA-targeting therapies using variety of endogenous gene silencing regulators, Small interfering RNAs (Si RNAs), aptamers and microRNA for cardiovascular diseases yet the development of a novel, risk free therapeutic strategy is a major challenge and need of the hour in cardiovascular medicine. In this regard these agents are required to overcome pleothra of barriers such as stability of drug targets, immunogenicity, adequate binding, targeted delivery etc. to become effective drugs. Recent years have witnessed the progress of RNA therapeutic strategies in cardiovascular diseases that are likely to significantly expand the cardiovascular therapeutic repertoire within the next decade. The present manuscript has been compiled to summarize various approaches of siRNA based therapies in cardiovascular diseases along with the advantages, outcomes and limitations if any in this regard. In addition, the future prospects of RNA therapeutic modalities in cardiovascular diseases are summarized.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Cardiovascular disease (CVD) is becoming a leading cause of morbidity, mortality as well as disability across the globe despite of the advancements in therapeutics and risk management strategies [1].The disease has been reported to be chronic in nature and the symptoms of the disease deteriorate as time period increases. Besides noteworthy therapeutic achievements in CVD, there are multiple undesirable risk factors associated with therapeutic modalities such as drug toxicity, complexity, resistance and many more. Hence the development of a novel, risk free therapeutic strategy is a major challenge and need of the hour in cardiovascular medicine. As per the availability of current literature, several lines of evidences are available supporting the function of RNA specifically the application of small interfering RNAs in silencing the disease causing genes in cardiovascular diseases [2]. It has been considered that RNA plays dynamic and versatile role in regulating the gene expression by acting as an intermediate molecule between DNA and proteins [3, 4].
RNA can offer various advantage in disease management as it can be edited and modified in its various forms such as secondary and tertiary structures. Moreover it may undergo tight, dynamic and various post transcriptional regulatory modifications by using plenty of RNA binding proteins [5, 6]. Hence RNA interference (RNAi) can propose major advantage over pharmacological therapy as they can target specific pathogenic genes that are associated with CVD with low toxicity and high potency. In addition to it, it is pertinent to mention that various biotechnology companies are already in process of manufacturing RNA-targeting therapies using various drug- able targets. For the same purpose, variety of endogenous gene silencing regulators, Small interfering RNAs (Si RNAs), aptamers and microRNA are being exploited to investigate the potential therapeutic agents [7].
It has been observed that novel technologies involved multiple types of small RNA that include miRNA, siRNA, snRNA, piRNA and snoRNA etc. Recent studies are evidencing the extremely broad spectrum of RNA species. Amongst these classes, the expression of mRNA target is inhibited by using siRNA whereas the miRNA are either used for inhibiting other miRNA or to mimic as some other miRNA which will be antagonizing the function of endogenous miRNA due to mimic behaviour [8]. This hypothesis explains how subclasses/varieties of RNA talk to each other and respond differently that leads to altered functional genetic information playing a vital role in various pathological conditions such as CVD etc. To date, therapies involving RNA agents are being exploited to treat various diseases such as cancer [9], infectious [10] and neurodegenerative diseases [11] as well.
The therapeutic potential of RNA based agents are being explored in multiple clinical trials in context to cardiovascular diseases. This manuscript focuses to summarize three such approaches that include siRNA, miRNA and aptamers as well along with the clinical trials conducted in this regard. Further the future directions of RNA therapeutics alongwith their outcomes and limitations in regard to cardiovascular diseases are critically summarized.
2 RNA and Its World: Traditional Concept vs. Current Concept
2.1 Traditional Concept of RNA World
As per the description of various classical studies, it has been observed that a large amount of RNA is transcribed into the cell. The structure and function of variety of RNA transcribed in the cell is becoming better understood with respect to their role in molecular biology as well as their therapeutic potential. Most of the RNA transcribed in the cell do not encode for proteins rather only a part of it (such as tRNA and rRNA) is involved in the process of translation and its regulation. As per the traditional knowledge, RNAs were considered to be transmitters of genetic information i.e. from DNA to RNA and then to the ribosome for proteins synthesis, and hence considered to be the regulators of protein synthesis/Gene expression [12]. The traditional concept of RNA and its role in gene expression was dictated in central dogma which is represented as follows in Fig. 23.1.
2.2 Current Concept of RNA World
Since last decade the multiple studies have expended the role of RNA within the cell and as other catalytic RNAs [13, 14]. As per current status, it is obvious that RNA is not only the intermediary molecule to encode proteins from genes rather some of its types may act as functional end products to control the gene expression in a different but strategic manner. Moreover, recent studies are evidencing the use of new classes of small RNAs such as miRNA as well as siRNA which are generated as a product of some novel biosynthetic pathways and are helpful to mediate regulatory functions [15]. Earlier it was considered that RNA are of two types that include coding RNA (that codes for proteins) and Non coding RNA (which do not encode for proteins). The various types of RNA include rRNA, tRNA, mRNA, snRNA, siRNA and snoRNA etc. The availability of different types of RNA along with their percentage is shown in Fig. 23.2.
Although a variety of RNAs are known to the scientist still the current understanding of RNA molecules and its function is only the tip of the iceberg. The RNA research is gaining momentum on a fast pace due to the rapid development of the molecular biotechnology as well as the various classes of RNA have attracted considerable attention of the scientists to unfold their role in gene regulation and in developing novel drug discovery and development targets. Amongst the above mentioned classes, microRNAs (miRNAs) as well as small interfering RNAs (siRNAs) are being highly exploited for their therapeutic potential and has been depicted by number of scientists against various deadly diseases such as cancer and others [16, 17]. Moreover these small molecules have a potential to become non-druggable target to cure plethora of diseases without undergoing drug induced side effects [18].
3 Small Interfering RNAs
SiRNAs are small non-coding RNA; a novel class of RNA based therapeutic agent with an important role in gene expression. siRNAs have a well-defined structure usually varying in its length from 20 to 24-bp with phosphorylation on 5′ end and hydroxylation on 3′ end [19]. In addition to it, siRNAs exists with overhanging nucleotides on both the sides. In the cell itself, the Dicer enzyme is responsible for the production of siRNAs from either long dsRNAs or small hairpin RNAs. Although these molecules are only of length 20–25 base pairs yet they work to inhibit the gene expression. These are the key molecules to direct post transcriptional gene silencing process called RNA interference (RNAi) rather their existence was first defined by the participation of siRNA in RNAi. It has been demonstrated that the siRNA are derived from the longer transcripts and the processing involves an enzyme named as DICER [20]. The general properties of the siRNA are shown in Table 23.1.
4 Mechanism of Action of siRNA
siRNAs are double stranded RNAs that direct post transcriptional gene silencing process called RNAi. These dsRNAs can be introduced into the cell with the help of transfection method. With the help of complementary siRNA sequence any of the gene can be knocked down and hence this RNA class is becoming an important tool for validating the gene functions as well as drug targeting. RNAi is a normal process that took place in almost all the eukaryotic cells. In this highly conserved process, dsRNA molecules i.e. siRNAs silence the post transcriptional effect of any gene of interest [7, 21]. Now a day, synthetic siRNAs are being synthesized by various biotechnology companies that are used to silence the effect of some pre-decided target genes. The knockdown of the target genes is entirely based upon the complementarily between a siRNA and the target gene [22]. Although, synthetic siRNAs are designed to silence targeted gene, but sometime unintended genes may also get knock down, due to imperfect complementarily to non-targeted mRNAs [22].
The steps involved in the RNA interference are as follows:
-
1.
Synthesis of siRNA: The synthesis of siRNA is the main stage of RNAi. To achieve the synthesis of siRNA, long dsRNAs are transfected inside the cell followed by cleavage of the ds DNA by an endo-ribonuclease enzyme. The enzyme used for the cleavage of the dsRNA is known as Dicer. This enzyme helps to cut a long piece of dsRNA into smaller pieces of 21–25 bp dsRNAs having 2 nucleotides on the 3′ terminus alongwith phosphate groups on 5′ terminus. The smaller dsRNA produced in this process are known either as silencing RNA or short interfering RNA [23].
-
2.
Incorporation of siRNA to RISC: In this step, siRNA duplexes are inserted into the RNA-induced silencing complex (RISC). RISC complex plays a crucial role in RNAi. It is a RNA/protein nuclease complex that initially binds to the siRNA duplexes [24]. For the incorporation of siRNA to RISC, 5′- terminal phosphorylation of the siRNA is compulsory. RISC has two domains named as PAZ and PIWI that help to recognize the 5′-terminus and 3′-terminus of the actual guiding strand followed by targeting the homologous regions in the guiding strand [25].
-
3.
Gene silencing: After incorporation into RISC complex, siRNAs are identified by RNA-induced silencing complex (RISC) as well as Argonaute 2 (AGO2) followed by uncoiling of the dssiRNAs into single strands [26,27,28]. Single stranded siRNA are recognized by their complementary mRNA and after binding with perfect complementary strands target mRNA is finally degraded by exonucleases and hence mRNA cleavage is induced. After the mRNA cleavage, the same will not be recognized by the cell as it is recognized as abnormal mRNA which will in turn silence the gene due to no translation [29, 30]. The step to step process of RNA interference is shown in Fig. 23.3.
There are several advantages of siRNA over conventional drugs [19, 31]. Detection of target sites is pretty easier with high flexibility since both the target siRNA and mRNA are sequences-specific. The inhibitory effect of the siRNA may be realized by targeting particular region of the mRNA [32]. In the process of gene silencing only a small amount of siRNA is required to reduce the concentration of homologous mRNA drastically within a period of 24 h. Physiological impact of cells is not altered by siRNA [33]. The transcripts of interest are destroyed selectively by high level homology of siRNA to the target region of cognate transcription. In the absence of the target sequences, the siRNA remain dormant in the cells and in the presence of the target sequence, the genes are silenced stably. siRNAs displays long term biological effects [34].
5 Chronological Representation of Discoveries in the Area of RNAi
The field of RNAi is not new to us. As far as the discovery of RNAi is concerned, geneticists Craig Mello and Andrew Fire discovered that upon injecting double-stranded RNA (dsRNA) into small worms some of the corresponding genes get switched off and hence the concept of the RNAi came into picture [35]. Finally researchers were awarded with Noble Prize for Medicine and Physiology in 2006. Some of the chronological discoveries in the area of RNAi are shown in Table 23.2.
6 RNA Interference in Cardiovascular Diseases
The technique is used to down regulate the desired gene expression using easily formulated small interfering RNA fragments. The method is quickly evolving against various diseases since its discovery and is becoming a routine application in many of the laboratories. There are number of studies that explain the success of RNA interference in treatment of human diseases.
RNAi induced by siRNA has emerged as a crucial technique for screening new therapeutic targets and searching molecular mechanisms of cardiovascular diseases [49]. siRNAs can be used to inhibit the overproduction of proteins that are associated with cardiovascular disease. Proprotein convertase subtilisin/kexin type 9 (PCSK9) has been reported to increase the levels of cholesterol in plasma; inhibition of the enzyme decreases the chances of hypercholesterolemia [50]. Studies have been carried out on the monkeys and it has been concluded that the special siRNA delivery by lipidoid nanoparticles against PCSK9 can decrease the cholesterol effectively [51]. siRNA have been found to silence the CCR2 i.e. chemokine receptor that results in enhanced recovery from myocardial infarction as it reduces the penetration of inflammatory cells into the infracted area [52].
Apart from inflammation, cardiomyocyte apoptosis is also observed during myocardial in fraction. Cardiomyocyte apoptosis results in reduced cardiac contractility as observed in mouse models with myocardial infarction (MI). Post myocardial infarction, apoptosis is mediated by tyrosine phosphatase Src homology region 2 domain-containing phosphatase 1 (SHP-1) [53]. It has been reported that RNAi against SHP-1 rescues cardiomyocytes from apoptosis among mouse MI model [54]. Addition, a hurdle in RNA therapeutics has been observed in mouse MI model but it resulted in non-specific gene silencing because of the off-target binding of siRNAs [55]. To reduce off-target binding of siRNA is also very important in RNA therapy. Tough decoy (TuD) RNA has been introduced which is meant to silence the sense strand of the siRNA duplex [56]. Recently, RNAi-based gene silencing methods are being established in humans and varieties of clinical trials are ongoing which may act as a promising therapy for the fatal diseases such as cardiovascular, neurological diseases etc.
It has been observed that upcoming cardiovascular RNA drugs are used to address multiple organ systems. It is not always mandatory to address heart or vasculature directly rather the disease can be managed by targeting liver RNA drugs. Liver targeted RNA drugs are gaining success and are developed in the field of cardiovascular diseases. Another system such as immune system involving monocytes or macrophages can be targeted for delivering RNA drug to its tissue target [57]. Various studies involving the role of RNAi specifically in cardiovascular diseases are shown in Table 23.3.
7 SiRNA Based Drugs: FDA Approval
Though the idea of the gene silencing has been revealed two decades ago, yet, the researchers are desperate and making critical steps for seeking approval for newly manufactured/to be manufactured new siRNA based medicines from US Food and Drug Administration (FDA). It has been observed that there is entry of FDA approved drugs into the market and amongst those drugs the drugs for various diseases are enlisted. The FDA approval for few of the siRNA has led the field of gene silencing to the new heights due to the fact that RNAi has the flexibility to design as many as novel targets against many different genes/diseases with high specificity. Few such drugs are enlisted in Table 23.4.
8 Future Perspective
The flexibility of RNAs makes them crucial therapeutics modality for number of human diseases. Exploring new classes of RNA that possess therapeutic potential will help in its successful translation to the clinic. Understanding the mode of action of various RNAs including long-noncoding RNAs (lncRNAs), miRNA, siRNA, etc. in CVD will help in improved therapeutics among patients. lncRNAs are distinct form of short non-coding RNAs that binds the targets through complimentary base pairing. lncRNAsfold into specific tertiary structures and interact with its proteins targets and the function of lncRNAs in MI have revealed [85]. The role of lncRNAs is only partially known and recently unfolded in cardiac development and the progression of MI [86]. Hence studies relating to regulation of long-noncoding RNAs for reversal of the diseased state and triggering the endogenous regenerative process can be a promising therapeutic tool for CVD patients [87, 88]. Many diseases result from multigene dysregulation; so administration of more than one therapeutic target will be beneficial for good outcome. Combination of RNA therapeutics with protein or small molecule drugs may also become promising therapy [89]. Delivery of RNAs will improve cell-based therapies. Cell based therapies will be promising approach for rejuvenation of ischemic myocardium since adult cardiomyocytes have restricted regenerative capacity [90]. However, this therapeutic approach has some obstacles like less cell retention and poor cell survival in infracted regions. These therapeutic challenges can be improved by incorporating few RNA agents. Delivery of mRNA encoding angiogenic factors along with RNAi that is responsible for decreased inflammation and fibrosis in the host tissue may improve survival [91]. For implementation of this approach few new strategies needs to be designed for effective cell and RNAs delivery. To achieve this goal, a new strategy should be designed for effective deliver of cells and RNAs to infracted area. Role of RNA therapeutics in treating MI has been successfully demonstrated in small animals however preclinical studies on large animals and patients will further explore/establish their efficacy in future. To maximize benefits and to avoid adverse effects, there is a need to draw stringent standard operative procedures for delivery strategies, development and improvement of RNA-based MI therapeutics.
References
Benjamin EJ, Blaha MJ, Chiuve SE, Cushman M, Das SR, Deo R, de Ferranti SD, Floyd J, Fornage M, Gillespie C, Isasi CR, Jimenez MC, Jordan LC, Judd SE, Lackland D, Lichtman JH, Lisabeth L, Liu S, Longenecker CT, Mackey RH, Matsushita K, Mozaffarian D, Mussolino ME, Nasir K, Neumar RW, Palaniappan L, Pandey DK, Thiagarajan RR, Reeves MJ, Ritchey M, Rodriguez CJ, Roth GA, Rosamond WD, Sasson C, Towfighi A, Tsao CW, Turner MB, Virani SS, Voeks JH, Willey JZ, Wilkins JT, Wu JH, Alger HM, Wong SS, Muntner P, American Heart Association Statistics Committee and Stroke Statistics Subcommittee. Heart disease and stroke statistics-2017 update: a report from the American Heart Association. Circulation. 2017;135(10):e146–603.
Tang Y, Ge YZ, Yin JQ. Exploring in vitro roles of siRNA in cardiovascular disease. Acta Pharmacol Sin. 2007;28(1):1–9.
Kapranov P, Willingham AT, Gingeras TR. Genome-wide transcription and the implications for genomic organization. Nat Rev Genet. 2007;8(6):413–23.
Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009;10(3):155–9.
Stellos K. The rise of epitranscriptomic era: implications for cardiovascular disease. Cardiovasc Res. 2017;113(5):e2–3.
Stellos K, Gatsiou A, Stamatelopoulos K, Perisic Matic L, John D, Lunella FF, Jae N, Rossbach O, Amrhein C, Sigala F, Boon RA, Furtig B, Manavski Y, You X, Uchida S, Keller T, Boeckel JN, Franco-Cereceda A, Maegdefessel L, Chen W, Schwalbe H, Bindereif A, Eriksson P, Hedin U, Zeiher AM, Dimmeler S. Adenosine-to-inosine RNA editing controls cathepsin S expression in atherosclerosis by enabling HuR-mediated post-transcriptional regulation. Nat Med. 2016;22(10):1140–50.
Siomi H, Siomi MC. On the road to reading the RNA-interference code. Nature. 2009;457(7228):396–404.
Carthew RW, Sontheimer EJ. Origins and mechanisms of miRNAs and siRNAs. Cell. 2009;136(4):642–55.
Moreno PM, Pego AP. Therapeutic antisense oligonucleotides against cancer: hurdling to the clinic. Front Chem. 2014;2:87.
Schluep T, Lickliter J, Hamilton J, Lewis DL, Lai CL, Lau JY, Locarnini SA, Gish RG, Given BD. Safety, tolerability, and pharmacokinetics of ARC-520 injection, an RNA interference-based therapeutic for the treatment of chronic hepatitis B virus infection, in healthy volunteers. Clin Pharmacol Drug Dev. 2017;6(4):350–62.
Scoles DR, Meera P, Schneider MD, Paul S, Dansithong W, Figueroa KP, Hung G, Rigo F, Bennett CF, Otis TS, Pulst SM. Antisense oligonucleotide therapy for spinocerebellar ataxia type 2. Nature. 2017;544(7650):362–6.
Roly ZY, Backhouse B, Cutting A, Tan TY, Sinclair AH, Ayers KL, Major AT, Smith CA. The cell biology and molecular genetics of Mullerian duct development. Wiley Interdiscip Rev Dev Biol. 2018;7(3):e310.
Warf MB, Berglund JA. Role of RNA structure in regulating pre-mRNA splicing. Trends Biochem Sci. 2010;35(3):169–78.
Lippman Z, Martienssen R. The role of RNA interference in heterochromatic silencing. Nature. 2004;431(7006):364–70.
Jacquier A. The complex eukaryotic transcriptome: unexpected pervasive transcription and novel small RNAs. Nat Rev Genet. 2009;10(12):833–44.
Davidson BL, McCray PB Jr. Current prospects for RNA interference-based therapies. Nat Rev Genet. 2011;12(5):329–40.
Burnett JC, Rossi JJ. RNA-based therapeutics: current progress and future prospects. Chem Biol. 2012;19(1):60–71.
Lares MR, Rossi JJ, Ouellet DL. RNAi and small interfering RNAs in human disease therapeutic applications. Trends Biotechnol. 2010;28(11):570–9.
Wittrup A, Lieberman J. Knocking down disease: a progress report on siRNA therapeutics. Nat Rev Genet. 2015;16(9):543–52.
Liu X, Jiang F, Kalidas S, Smith D, Liu Q. Dicer-2 and R2D2 coordinately bind siRNA to promote assembly of the siRISC complexes. RNA. 2006;12(8):1514–20.
Donze O, Picard D. RNA interference in mammalian cells using siRNAs synthesized with T7 RNA polymerase. Nucleic Acids Res. 2002;30(10):e46.
Jackson AL, Linsley PS. Recognizing and avoiding siRNA off-target effects for target identification and therapeutic application. Nat Rev Drug Discov. 2010;9(1):57–67.
Carthew RW. Synthesis of siRNA for RNAi in Drosophila. CSH Protoc. 2006;2006(3) https://doi.org/10.1101/pdb.prot4512.
Matranga C, Tomari Y, Shin C, Bartel DP, Zamore PD. Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes. Cell. 2005;123(4):607–20.
Ameres SL, Martinez J, Schroeder R. Molecular basis for target RNA recognition and cleavage by human RISC. Cell. 2007;130(1):101–12.
Sledz CA, Holko M, de Veer MJ, Silverman RH, Williams BR. Activation of the interferon system by short-interfering RNAs. Nat Cell Biol. 2003;5(9):834–9.
Liu J, Carmell MA, Rivas FV, Marsden CG, Thomson JM, Song JJ, Hammond SM, Joshua-Tor L, Hannon GJ. Argonaute2 is the catalytic engine of mammalian RNAi. Science. 2004;305(5689):1437–41.
Meister G. Argonaute proteins: functional insights and emerging roles. Nat Rev Genet. 2013;14(7):447–59.
Hammond SM, Bernstein E, Beach D, Hannon GJ. An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature. 2000;404(6775):293–6.
Cerutti H. RNA interference: traveling in the cell and gaining functions? Trends Genet. 2003;19(1):39–46.
Sioud M. Therapeutic siRNAs. Trends Pharmacol Sci. 2004;25(1):22–8.
Tan FL, Yin JQ. RNAi, a new therapeutic strategy against viral infection. Cell Res. 2004;14(6):460–6.
McManus MT, Sharp PA. Gene silencing in mammals by small interfering RNAs. Nat Rev Genet. 2002;3(10):737–47.
Agrawal N, Dasaradhi PV, Mohmmed A, Malhotra P, Bhatnagar RK, Mukherjee SK. RNA interference: biology, mechanism, and applications. Microbiol Mol Biol Rev. 2003;67(4):657–85.
Parrish S, Fleenor J, Xu S, Mello C, Fire A. Functional anatomy of a dsRNA trigger: differential requirement for the two trigger strands in RNA interference. Mol Cell. 2000;6(5):1077–87.
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature. 1998;391(6669):806–11.
Mello CC, Conte D Jr. Revealing the world of RNA interference. Nature. 2004;431(7006):338–42.
Gehrs KM, Anderson DH, Johnson LV, Hageman GS. Age-related macular degeneration – emerging pathogenetic and therapeutic concepts. Ann Med. 2006;38(7):450–71.
Paddison PJ, Silva JM, Conklin DS, Schlabach M, Li M, Aruleba S, Balija V, O'Shaughnessy A, Gnoj L, Scobie K, Chang K, Westbrook T, Cleary M, Sachidanandam R, McCombie WR, Elledge SJ, Hannon GJ. A resource for large-scale RNA-interference-based screens in mammals. Nature. 2004;428(6981):427–31.
Wadman M. Swooping for biotech. Nature. 2005;437(7058):475.
Bernards R. The Nobel Prize in Physiology or Medicine for 2006 for the discovery of RNA interference. Ned Tijdschr Geneeskd. 2006;150(52):2849–53.
Mack GS. MicroRNA gets down to business. Nat Biotechnol. 2007;25(6):631–8.
Haussecker D. The business of RNAi therapeutics. Hum Gene Ther. 2008;19(5):451–62.
Palanki MS, Akiyama H, Campochiaro P, Cao J, Chow CP, Dellamary L, Doukas J, Fine R, Gritzen C, Hood JD, Hu S, Kachi S, Kang X, Klebansky B, Kousba A, Lohse D, Mak CC, Martin M, McPherson A, Pathak VP, Renick J, Soll R, Umeda N, Yee S, Yokoi K, Zeng B, Zhu H, Noronha G. Development of prodrug 4-chloro-3-(5-methyl-3-{[4-(2-pyrrolidin-1-ylethoxy)phenyl]amino}-1,2,4-benzotria zin-7-yl)phenyl benzoate (TG100801): a topically administered therapeutic candidate in clinical trials for the treatment of age-related macular degeneration. J Med Chem. 2008;51(6):1546–59.
Haussecker D. The business of RNAi therapeutics in 2012. Mol Ther Nucleic Acids. 2012;1:e8.
Garber K. Alnylam terminates revusiran program, stock plunges. Nat Biotechnol. 2016;34(12):1213–4.
Rizk M, Tuzmen S. Update on the clinical utility of an RNA interference-based treatment: focus on Patisiran. Pharmgenomics Pers Med. 2017;10:267–78.
Chakraborty C, Sharma AR, Sharma G, Doss CGP, Lee SS. Therapeutic miRNA and siRNA: moving from bench to clinic as next generation medicine. Mol Ther Nucleic Acids. 2017;8:132–43.
Bartz F, Kern L, Erz D, Zhu M, Gilbert D, Meinhof T, Wirkner U, Erfle H, Muckenthaler M, Pepperkok R, Runz H. Identification of cholesterol-regulating genes by targeted RNAi screening. Cell Metab. 2009;10(1):63–75.
Tibolla G, Norata GD, Artali R, Meneghetti F, Catapano AL. Proprotein convertase subtilisin/kexin type 9 (PCSK9): from structure-function relation to therapeutic inhibition. Nutr Metab Cardiovas Dis. 2011;21(11):835–43.
Frank-Kamenetsky M, Grefhorst A, Anderson NN, Racie TS, Bramlage B, Akinc A, Butler D, Charisse K, Dorkin R, Fan Y, Gamba-Vitalo C, Hadwiger P, Jayaraman M, John M, Jayaprakash KN, Maier M, Nechev L, Rajeev KG, Read T, Rohl I, Soutschek J, Tan P, Wong J, Wang G, Zimmermann T, de Fougerolles A, Vornlocher HP, Langer R, Anderson DG, Manoharan M, Koteliansky V, Horton JD, Fitzgerald K. Therapeutic RNAi targeting PCSK9 acutely lowers plasma cholesterol in rodents and LDL cholesterol in nonhuman primates. Proc Natl Acad Sci U S A. 2008;105(33):11915–20.
Majmudar MD, Keliher EJ, Heidt T, Leuschner F, Truelove J, Sena BF, Gorbatov R, Iwamoto Y, Dutta P, Wojtkiewicz G, Courties G, Sebas M, Borodovsky A, Fitzgerald K, Nolte MW, Dickneite G, Chen JW, Anderson DG, Swirski FK, Weissleder R, Nahrendorf M. Monocyte-directed RNAi targeting CCR2 improves infarct healing in atherosclerosis-prone mice. Circulation. 2013;127(20):2038–46.
Sugano M, Tsuchida K, Hata T, Makino N. RNA interference targeting SHP-1 attenuates myocardial infarction in rats. FASEB J. 2005;19(14):2054–6.
Kim D, Hong J, Moon HH, Nam HY, Mok H, Jeong JH, Kim SW, Choi D, Kim SH. Anti-apoptotic cardioprotective effects of SHP-1 gene silencing against ischemia-reperfusion injury: use of deoxycholic acid-modified low molecular weight polyethyleneimine as a cardiac siRNA-carrier. J Control Release. 2013;168(2):125–34.
Burchard J, Jackson AL, Malkov V, Needham RH, Tan Y, Bartz SR, Dai H, Sachs AB, Linsley PS. MicroRNA-like off-target transcript regulation by siRNAs is species specific. RNA. 2009;15(2):308–15.
Mockenhaupt S, Grosse S, Rupp D, Bartenschlager R, Grimm D. Alleviation of off-target effects from vector-encoded shRNAs via codelivered RNA decoys. Proc Natl Acad Sci U S A. 2015;30:E4007–16.
Xu F, Jin L, Jin Y, Nie Z, Zheng H. Long noncoding RNAs in autoimmune diseases. J Biomed Mater Res A. 2019;107(2):468–75.
Hur K, Kim SH, Kim JM. Potential implications of long noncoding RNAs in autoimmune diseases. Immune Netw. 2019;19(1):e4.
Qie Y, Sheng Y, Xu H, Jin Y, Ma F, Li L, Li X, An D. Identification of a new powdery mildew resistance gene pmDHT at or closely linked to the Pm5 locus in the Chinese wheat Landrace Dahongtou. Plant Dis. 2019;103:2645–51.
Rapanelli M, Tan T, Wang W, Wang X, Wang ZJ, Zhong P, Frick L, Qin L, Ma K, Qu J, Yan Z. Behavioral, circuitry, and molecular aberrations by region-specific deficiency of the high-risk autism gene Cul3. Mol Psychiatry. 2019; https://doi.org/10.1038/s41380-019-0498-x.
Nissen PH, Rejmark L. Expanding the spectrum of genetic variants in the calcium sensing receptor (CASR) gene in hypercalcemic individuals. Clin Endocrinol (Oxf). 2019;91(5):683–90.
Moreno AM, Palmer N, Aleman F, Chen G, Pla A, Jiang N, Chew WL, Law M, Mali P. Author correction: immune-orthogonal orthologues of AAV capsids and of Cas9 circumvent the immune response to the administration of gene therapy. Nat Biomed Eng. 2019;3(10):842.
Meng Q, Sun S, Luo Z, Shi B, Shan A, Cheng B. Maternal dietary resveratrol alleviates weaning-associated diarrhea and intestinal inflammation in pig offspring by changing intestinal gene expression and microbiota. Food Funct. 2019;10(9):5626–43.
Manor E, Gonen R, Sarussi B, Keidar-Friedman D, Kumar J, Tang HT, Tassone F. The role of AGG interruptions in the FMR1 gene stability: A survey in ethnic groups with low and high rate of consanguinity. Mol Genet Genomic Med. 2019;7(10):e00946.
Liu Z, Chen X. A novel missense mutation in human Receptor Roundabout-1 (ROBO1) gene associated with pituitary stalk interruption syndrome. J Clin Res Pediatr Endocrinol. 2019; https://doi.org/10.4274/jcrpe.galenos.2019.2018.0309.
Liu C, Kanazawa T, Tian Y, Mohamed Saini S, Mancuso S, Mostaid MS, Takahashi A, Zhang D, Zhang F, Yu H, Doo Shin H, Sub Cheong H, Ikeda M, Kubo M, Iwata N, Woo SI, Yue W, Kamatani Y, Shi Y, Li Z, Everall I, Pantelis C, Bousman C. The schizophrenia genetics knowledgebase: a comprehensive update of findings from candidate gene studies. Transl Psychiatry. 2019;9(1):205.
Li XZ, Yan Y, Zhang JF, Sun JF, Sun B, Yan CG, Choi SH, Johnson BJ, Kim JK, Smith SB. Oleic acid in the absence of a PPARgamma agonist increases adipogenic gene expression in bovine muscle satellite cells. J Anim Sci. 2019;97(10):4114–23.
Asghar O, Alam U, Hayat SA, Aghamohammadzadeh R, Heagerty AM, Malik RA. Obesity, diabetes and atrial fibrillation; epidemiology, mechanisms and interventions. Curr Cardiol Rev. 2012;8(4):253–64.
Assis R. Out of the testis, into the ovary: biased outcomes of gene duplication and deletion in Drosophila. Evolution. 2019;73(9):1850–62.
Attaran S, Saleh HZ, Shaw M, Ward A, Pullan M, Fabri BM. Does the outcome improve after radiofrequency ablation for atrial fibrillation in patients undergoing cardiac surgery? A propensity-matched comparison. Eur J Cardiothorac Surg. 2012;41(4):806–10.. discussion 810-801
Ahmad K, Spens AE. Separate Polycomb Response Elements control chromatin state and activation of the vestigial gene. PLoS Genet. 2019;15(8):e1007877.
Ahuja V, Powers-Lee SG. Human carbamoyl-phosphate synthetase: insight into N-acetylglutamate interaction and the functional effects of a common single nucleotide polymorphism. J Inherit Metab Dis. 2008;31(4):481–91.
Ai L, Liu J, Jiang Y, Guo W, Wei P, Bai L. Specific PCR method for detection of species origin in biochemical drugs via primers for the ATPase 8 gene by electrophoresis. Mikrochim Acta. 2019;186(9):634.
Akhbari M, Khalili M, Shahrabi-Farahani M, Biglari A, Bandarian F. Expression level of circulating cell free miR-155 gene in serum of patients with diabetic nephropathy. Clin Lab. 2019;65(8) https://doi.org/10.7754/Clin.Lab.2019.190209.
Yang J. Patisiran for the treatment of hereditary transthyretin-mediated amyloidosis. Expert Rev Clin Pharmacol. 2019;12(2):95–9.
Sardh E, Harper P, Balwani M, Stein P, Rees D, Bissell DM, Desnick R, Parker C, Phillips J, Bonkovsky HL, Vassiliou D, Penz C, Chan-Daniels A, He Q, Querbes W, Fitzgerald K, Kim JB, Garg P, Vaishnaw A, Simon AR, Anderson KE. Phase 1 trial of an RNA interference therapy for acute intermittent porphyria. N Engl J Med. 2019;380(6):549–58.
Pasikowska M, Walsby E, Apollonio B, Cuthill K, Phillips E, Coulter E, Longhi MS, Ma Y, Yallop D, Barber LD, Patten P, Fegan C, Ramsay AG, Pepper C, Devereux S, Buggins AG. Phenotype and immune function of lymph node and peripheral blood CLL cells are linked to transendothelial migration. Blood. 2016;128(4):563–73.
Ray KK, Landmesser U, Leiter LA, Kallend D, Dufour R, Karakas M, Hall T, Troquay RP, Turner T, Visseren FL, Wijngaard P, Wright RS, Kastelein JJ. Inclisiran in patients at high cardiovascular risk with elevated LDL cholesterol. N Engl J Med. 2017;376(15):1430–40.
Kitagawa H, Ohbuchi K, Munekage M, Fujisawa K, Kawanishi Y, Namikawa T, Kushida H, Matsumoto T, Shimobori C, Nishi A, Sadakane C, Watanabe J, Yamamoto M, Hanazaki K. Phenotyping analysis of the Japanese Kampo medicine maoto in healthy human subjects using wide-targeted plasma metabolomics. J Pharm Biomed Anal. 2019;164:119–27.
Vita G, Vita GL, Stancanelli C, Gentile L, Russo M, Mazzeo A. Genetic neuromuscular disorders: living the era of a therapeutic revolution. Part 1: Peripheral neuropathies. Neurol Sci. 2019;40(4):661–9.
Tarantini G, Masiero G, Fovino LN, Mojoli M, Varricchio A, Loi B, Gistri R, Misuraca L, Gabrielli G, Cortese B, Pisano F, Moretti L, Tumminello G, Olivari Z, Mazzarotto P, Colombo A, Calabro P, Nicolino A, Tellaroli P, Corrado D, Durante A, Steffenino G, RAIi a. “Full-plastic jacket” with everolimus-eluting Absorb bioresorbable vascular scaffolds: Clinical outcomes in the multicenter prospective RAI registry (ClinicalTrials.gov Identifier: NCT02298413). Int J Cardiol. 2018;266:67–74.
Weng Y, Xiao H, Zhang J, Liang XJ, Huang Y. RNAi therapeutic and its innovative biotechnological evolution. Biotechnol Adv. 2019;37(5):801–25.
Turner L. ClinicalTrials.gov, stem cells and ‘pay-to-participate’ clinical studies. Regen Med. 2017;12(6):705–19.
Yang A, Baxi S, Korenstein D. ClinicalTrials.gov for facilitating rapid understanding of potential harms of new drugs: the case of checkpoint inhibitors. J Oncol Pract. 2018;14(2):72–6.
Mercer TR, Mattick JS. Structure and function of long noncoding RNAs in epigenetic regulation. Nat Struct Mol Biol. 2013;20(3):300–7.
Devaux Y, Zangrando J, Schroen B, Creemers EE, Pedrazzini T, Chang CP, Dorn GW 2nd, Thum T, Heymans S, Cardiolinc Network. Long noncoding RNAs in cardiac development and ageing. Nat Rev Cardiol. 2015;12(7):415–25.
Vausort M, Wagner DR, Devaux Y. Long noncoding RNAs in patients with acute myocardial infarction. Circ Res. 2014;115(7):668–77.
Ounzain S, Pedrazzini T. Long non-coding RNAs in heart failure: a promising future with much to learn. Ann Transl Med. 2016;4(15):298.
Jones SK, Merkel OM. Tackling breast cancer chemoresistance with nano-formulated siRNA. Gene Ther. 2016;23(12):821–8.
Baumann V, Winkler J. miRNA-based therapies: strategies and delivery platforms for oligonucleotide and non-oligonucleotide agents. Future Med Chem. 2014;6(17):1967–84.
Zangi L, Lui KO, von Gise A, Ma Q, Ebina W, Ptaszek LM, Spater D, Xu H, Tabebordbar M, Gorbatov R, Sena B, Nahrendorf M, Briscoe DM, Li RA, Wagers AJ, Rossi DJ, Pu WT, Chien KR. Modified mRNA directs the fate of heart progenitor cells and induces vascular regeneration after myocardial infarction. Nat Biotechnol. 2013;31(10):898–907.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Bansal, P., Arora, M. (2020). Small Interfering RNAs and RNA Therapeutics in Cardiovascular Diseases. In: Xiao, J. (eds) Non-coding RNAs in Cardiovascular Diseases. Advances in Experimental Medicine and Biology, vol 1229. Springer, Singapore. https://doi.org/10.1007/978-981-15-1671-9_23
Download citation
DOI: https://doi.org/10.1007/978-981-15-1671-9_23
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-1670-2
Online ISBN: 978-981-15-1671-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)