Abstract
Cardiac development in the human embryo is characterized by the interactions of several transcription and growth factors leading the heart from a primordial linear tube into a synchronous contractile four-chamber organ. Studies on cardiogenesis showed that cell proliferation, differentiation, fate specification and morphogenesis are spatiotemporally coordinated by cell-cell interactions and intracellular signalling cross-talks. In recent years, research has focused on a class of inter- and intra-cellular modulators called non-coding RNAs (ncRNAs), transcribed from the noncoding portion of the DNA and involved in the proper formation of the heart. In this chapter, we will summarize the current state of the art on the roles of three major forms of ncRNAs [microRNAs (miRNAs), long ncRNAs (lncRNAs) and circular RNAs (circRNAs)] in orchestrating the four sequential phases of cardiac organogenesis.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Background
Successful development of the embryonic heart sees the cardiac progenitor cells proliferate and differentiate into beating cardiomyocytes (CMs). Cardiac organogenesis requires exquisite modulation of gene expression, and transcriptional dysregulation in this process underpins congenital heart diseases. Most of the literature to be discussed in this chapter will mainly focus on murine studies, as heart development has been mainly investigated using transgenic mouse models. From a clinical standpoint, a comparison between mouse and human cardiac development by means of episcopic fluorescence image capture revealed the relevance of this model, as the cellular events leading to the formation of the heart are comparable in both mammals.
The early stage embryo is a disc formed by the three sheets of ectoderm, mesoderm and endoderm known as the three germ layers. The tissues forming the heart mainly come from the mesoderm germ layer. However, some of the cells migrate from the ectoderm and form the cardiac neural crest cells. The latter will participate in the septation of the cardiac outflow tract into aorta and pulmonary artery, remodel the pharyngeal arch arteries, develop the valves, and take part in the formation of the cardiac conduction system.
The phases of cardiac development in human and mouse have different timings, as summarized in Fig. 11.1. Following gastrulation, the heart muscle cells start developing from a pool of mesodermal cardiac precursor cells found in the anterior lateral plate of the embryonic mesoderm. These progenitors will then migrate to the cranial and cranio-lateral regions of the developing embryo. The subsequent phases of cardiac development can be divided into the following key steps, which warrant the correct formation and maturation of the three-dimensional structures of the heart: cardiac crescent (CS8), linear heart tube (CS9), cardiac looping (CS10), chamber formation (CS11-19), and maturation (CS20-birth).
In order to regulate the fate of the several progenitor cells to eventually form the heart, non-coding RNAs (ncRNAs) have been recognized to play a fundamental role in cardiac development and pathologies [1, 2], thanks to recent technological advances in sequencing and computational algorithms. Additionally, the discovery of ncRNAs has also expanded the functional complexity of transcriptome, adding new molecular dimension to temporal regulation, cellular and tissue specificities and functional diversity in heart organogenesis [3, 4]. At genetic level, ncRNAs modulate gene expression patterns by interrogating transcription, chromatin modification and post-transcriptional alterations [2]. First referred to as the ‘junk DNA’, a large part of the non-coding portion of the human genome (up to 90%) has now been proven to be actively transcribed into several types of ncRNAs which hold several biological functions throughout prenatal development and post-natal life [5, 6]. Based on the molecular length and function, three main categories of ncRNAs have been identified: long ncRNAs (lncRNAs; longer than 200 nucleotides), microRNAs (miRNAs; maximum 22 nucleotides long), and circular RNAs (circRNAs; formed by 1-5 exons).
The widest subgroup of ncRNAs consists of lncRNAs [7], mostly transcribed by RNA polymerase II, which causes them to undergo capping at 5′ end and polyadenylation at 3′ end. lncRNAs have a limited to absent protein-coding potential due to the lack of open reading frames. LncRNAs appeared to be critical regulators of gene expression in both transcription and post-transcription gene regulation events, with the majority of them exhibiting developmental stage-specific regulation paralleling mRNA expression patterns [8]. They can act either in cis, in order to regulate the nearby genes, or in trans, which let them modulate the expression of the target genes by means of several mechanisms. These mechanisms include: DNA looping, recruiting chromatin modifiers and transcription factors, miRNA sponges, and influencing mRNA splicing, translation or degradation. Through genome-wide RNA sequencing, more than 100 annotated and newly described IncRNAs have been defined in the cardiac differentiation and maturation signatures [9]. Nevertheless, the exact transcriptomic profiling and roles of lncRNAs during heart development (i.e. CMs differentiation, heart wall development, cardiac chamber and outflow tract formation, and cardiac cell electricphysiology and conduction) have not yet been detailed.
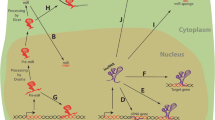
The mechanism of gene expression regulation by miRNAs, on the other hand, is at the post-transcriptional level, with silencing of genes that occurs via targeting the protein-coding and non-coding genes. Following synthesis of pri-miRNAs by RNA polymerase II and III in the nucleus, the microprocessor complex Drosha-Dgcr8 cleaves the pri-miRNAs into pre-miRNAs. These are then transported into the cytosol, where the Dicer-TRBP complex cleaves pre-miRNAs to form the mature 22 nucleotide-long miRNA molecules. Here, miRNAs will go through the RNA-induced silencing complex, formed by the Argonaute proteins, which guides the miRNAs towards the target mRNA for its degradation [10, 11]. The fundamental role played by miRNAs in cardiac development was proved by Dicer knock-down in murine ESCs which, among other effects, led to cardiac development defects [12].
circRNAs are single-stranded circular RNAs predominantly found in the cytoplasm. Thanks to the absence of 5′ and 3′ ends, they have a more stable structure making them more resistant to the exonuclease-mediated degradation to which the other ncRNAs undergo. Based on the derivation sources, circRNAs can be categorized into: (1) circRNA derived from exons (ecircRNA; the most abundant form of circRNAs), (2) circRNA derived from lariat introns (ciRNA), and (3) circRNA derived from exons wih retained introns (ElciRNA) [13,14,15]. circRNA length ranges between 100–1000 bases and, although their abundance is relatively low, some are expressed at higher levels compared to their linear transcripts. Although circRNAs are ubiquitously expressed, they accumulate in fully differentiated somatic cells while being quite diluted in proliferating cells including tumour cells. Mechanistically, it has been shown that circRNAs can act as miRNA sponges to counteract the inhibition induced by the latter. Indeed, the phenotype induced by gain and loss of function esperiments in zebrafish indicated that a specific circRNA could have functions beyond sequestering specific miRNAs [16]. As a single-stranded RNA, circRNAs can bind the trans 3’ UTRs of target mRNAs to concur in gene expression regulation. In addition, circRNAs can be involved in the regulation of RNA-binding proteins [17]. Compared to lncRNAs and miRNAs, however, the functions of thousands of described circRNAs remain limited. Interestingly, based on deep RNA sequencing analysis, the top-expressed circRNAs in the human heart were associated with cardiac- or skeletal muscle genes including TTN, RYR2 and DMD [18].
Since ncRNAs regulators have only been recently related to cardiac development and disease, a detailed understanding on the expression dynamic of these ncRNAs during each stages of the embryonic heart development is quintessential. Thus, in this chapter we will summarize the roles that ncRNAs play in the development of the heart.
2 From Cardiac Crescent to Looping Heart Tube: The Role of ncRNAs
In the early stages of development, the cardiac precursor cell population is found in the two symmetrical sides of the lateral plate mesoderm of the flat tri-laminar disc. At CS8 (human 15–20 days, mouse E7.5), the lateral plate gets divided by the intraembryonic coelom in two layers, i.e. the somatic and the splanchnic mesoderm. Once the two sides start merging, the splanchnic mesoderm merges cranially and forms a horseshoe-shaped field named the cardiac crescent. The cells that form the cardiac crescent are termed the first heart field (FHF) and will contribute to the left ventricle and atrioventricular canal [19]. At the medial sides of the cardiac crescent processes, a separate population of cells forms the second heart field (SHF) which will contribute to the outflow tract myocardium, right ventricle and both atria. Cells derived from the FHF will first fuse at the midline to form the linear heart tube at CS9, after which SHF cells will add to the heart tube and increase it in size. Subsequently, the heart tube loops at CS10 [20].
Several genes are expressed in the committed mesodermal cells towards cardiac lineage. The earliest genes involved in commitment of embryonic stem cells (ESCs) towards cardiac mesoderm are the transcription factor Brachyury and eomesodermin (Eomes). Both Brachyury and Eomes are critical for the primitive streak patterning and the mesendoderm specification in the early embryo. In particular, Eomes is the key transcription factor required for the formation of either endoderm or cardiovascular mesoderm according to a high or low level of Activin, respectively. Following commitment to mesoderm, Eomes will then induce the expression of Mesp1 [21], which will eventually start the cardiovascular differentiation [22].
Several ncRNAs collaborate with Eomes in the early commitment of ESCs towards cardiac mesoderm. For instance, the exon 2 of lncRNA linc1405 was shown to co-localize with Eomes in the primitive streak and played a major role in the activation of Mesp1-mediated cardiac mesoderm specification of ESCs [23]. The lncRNA Fendrr (ENSMUSG00000097336) was shown to be expressed in EOMES-positive cells at E6.5-7, with its loss resulted in embryonic lethality in mice [24]. Finally, it has been reported that other lncRNAs and circRNAs are either transcriptionally regulated (LINC00467) or co-expressed (RP3428L16.2, RP11829H16.3; circPSD3, circSLC39A8, circALMS1) with EOMES in human cardiac progenitors [25].
Mesp1-expressing cells contribute to FHF and SHF derivatives, which will eventually give rise to the three main compartments of the heart, i.e. cardiac muscle (made by CMs), vessels (endothelial cells) and epicardium [26, 27]. Downstream of Mesp1, the FHF expresses the transcription factors Nkx2.5, Hand1 and Tbx5 [28, 29], while SHF expresses Nkx2.5, Gata4/6, Hand2, Tbx1/2, Mef2c and Isl1 [30,31,32,33,34,35,36].
Upon fusion at the midline of the cell populations derived from the FHF, the heart tube forms (CS9, mouse E8, human day 21), and CMs arrest the proliferation process. SHF precursor cells simultaneously migrate from the pericardium to the heart tube at the venous and arterial poles [36]. As they are mediated by WNT/β-catenin signaling [37], they proliferate at high rates and thus contribute to the heart tube’s growth. During their addition to the heart tube, the SHF-derived CMs temporarily stop proliferating. Noncanonical WNT and Notch signaling also regulate differentiation during second heart field deployment [38, 39].
For the expression of Mesp1, the downregulation of miR-142-3p during ESC differentiation is required. Conversely, Mesp1 activates the miR-322/-503 cluster during the heart looping [40]. In mice, the lncRNA Braveheart (Bvht, AK143260) is required to induce Mesp1, and the depletion of Bvht in mouse ESCs impairs the formation of CMs. Intriguingly, to date the transcript of Bvht has not yet been identified in human. Conversely, the lncRNA Carmen was seen to be conserved from mouse to human, and its expression is induced between the mesodermal and cardiac progenitor stage. Similar to Bvht, depletion of Carmen was associated with a significant reduction in the expression of differentiation makers and cardiac transcription factors, including NKX2.5, TBX5, GATA4, MYH6, MYH7, and TNNI [41]. Moreover, the expression of the master cardiac transcription factor – Nkx2.5 was modulated by novlnc6 which influenced the expression of BMP10 (a key signaling ligand for cardiogenesis during embryonic stem cell cardiac differentiation) [42].
Mesp2 has redundant functions compensating for Mesp1 upon knock-out of the latter [43, 44]. However, Mesp1 plays a major role in the motility of progenitors required for the correct cell migration and cardiac development [45]. A group of ncRNAs were reported to be co-expressed with MESP2 in the early cardiac mesoderm (circPSD3; RP11445F12.1, RP11445F12.2, RP3428L16.2, LINC00467) [25].
The T-box family genes start being expressed in the FHF and SHF. In the FHF, Tbx5 expression is modulated by miR-218 family, with the overexpression of Tbx5 affecting heart development in both humans and mice, resulting in heart chamber abnormalities and heart-looping defects [46]. Intriguingly, the ectopic expression of Gata4 and Tbx5, combined with chromatin remodeling component Baf60c/Smarcd3, was shown to induce beating myocardium in mesoderm [47]. Conversely, in the SHF TBX1 interferes with BMP signaling cascade components and has a negative regulatory effect on Mef2c transcript and SRF protein levels [48,49,50]. The subsequent differentiation of the myocardium at the arterial pole of the heart tube is reinforced by BMP which drives the miRNA 17-92-mediated repression of Isl1 and Tbx1 [25]. Repression of Tbx1 during heart maturation is of utmost importance, as its overexpression leads to Gata4 and Mef2c downregulation with subsequent blockage of the cardiac differentiation pathway [51]. This finding is corroborated by the required upregulation of MEF2C during induction of cardiac differentiation of the human embryonic stem cells which was found to be modulated via overexpression of miRNA-499 and miRNA-1 [52].
SHF and neural crest cells involved in cardiac development are characterized by the expression of Isl1 [53, 54] although it has been shown to be transiently expressed in FHF cells as well, albeit with no related function [33, 55]. A group of ncRNAs were shown to be co-expressed (MEIS1-AS2; circ-PTPN13, circ-ENC1, circ-PPP2R3A, circ-FUT8) or transcriptionally regulated (LINC01021, AC009518.4) with ISL1 in human cardiac progenitors [25, 55]. ISL1 is targeted by miR-17-92 to promote differentiation of the myocardium at the arterial pole in the final stages of maturation [51]. Finally, the expression of HAND2 – critical for ventricular CMs expansion – is initially discovered in the cardiac crescent at E7.75 and will continue throughout the linear heart tube at E8.5. It has been recently shown that the lncRNA Uph (also named Hand2as or lncHand2), playing critical roles in the regulation of the precise expression of HAND2, together with miR-1-2 family in looping heart, to eventually lead to chamber formation [56, 57].
Myocardin (Myocd) is a master regulator of the smooth muscle cell phenotype. It is expressed in cardiac crescent and it coactivates several factors including Gata4, Tbx5, serum response factor (Srf) – which regulates BMP10 in cardiac maturation – and MEF2 [58, 59]. MYOCD is activated by lncRNAs MYOSLID [60] and SENCR, although the latter has an indirect influence on it. In mice, mm67 and mm85 have been shown to activate Myocd. In subsequent stages, Myocd is shown to be modulated by miR-1 [61], and miR-214 has been shown to indirectly regulate its expression [62, 63]. Myocardin is required for CMs survival and heart function maintenance after birth [64]. Finally, for the correct formation of heart and vessels the lncRNA ALIEN was identified in mesendodermal tissues between cardiac crescent and heart tube [3].
In the looping heart, miR-1-2 family targets NOTCH ligands, HDAC4, Hand2, MEF2 and SRF to eventually allow the progenitor cells to proliferate and differentiate. The miR-1/133a cluster is positively regulated by Myocd, which aids in the specification of immature embryonic CMs into fetal ones [61, 65]. miR-1 is polycistronically clustered on the same chromosome with miR-133, however they have different – and sometimes opposing – effects during cardiac differentiation. The deletion of miR-133a genes led to ventricular septal defects and abnormal cardiomyocytic proliferation which eventually leading to neonatal death [66]. However, its overexpression in mouse and human ESCs caused the repression of cardiac markers [67, 68].
Another miRNA involved in the looping of the heart is miR-499, encoded by Myh7b. In vitro, its overexpression was shown to speed up the beating embryoid bodies formation while its inhibition blocked cardiac differentiation [65].
During cardiac differentiation, several circRNAs were seen to be overexpressed. Circ-SLC8A1-1 is expressed from the gene NCX1 (Na+/Ca++ exchanger, also known as SLC8A1) during CMs differentiation in hESC and mouse [69]. In a study comparing human, mice and rat hearts, circSLC8A1-1 was shown to be the most abundant circRNA in the hearts [70]. Intriguingly, upregulation of circ-SLC8A1 was observed in the DCM [71]. Other reported circRNAs during cardiac differentiation include circ-TTN-90, circ-TTN-275, circ-TPM1-1, circ-HIPK3-2, circ-EXOC6B-14, circ-MB-2, circ-ALPK2-2, circ-MYBPC3-3, circ-NEBL-19 and circ-RYR2-113 in hESC differentiating towards CMs [18]. On the contrary, the circr-Foxo3 was found to interact with multiple stress- and senescence-related factors (e.g. ID-1, E2F1 and FAK), which was highly associated with heart samples from both aged patients and mice [72].
3 Chamber Formation and the Final Phases of Heart Maturation
It is perceivable that heart being a mechanical pump requires three-dimensionality (in term of chambers, valves, septation and blood vessels) to fullfil its biological functions, and cardiac function dictates its form to a large extent. Hence, the formation and maturation of the heart structure are highly associated with the contractile force and hemodynamic demands towards the systemic circulation, in addition to influences by other factors such as oxygen gradient and nutrient enviroment. In the final stages of heart formation, the major contribution in cardiac growth comes from the intracardiac myocardial cells. In particular, the ventricular and atrial myocardium arises from the outer curvature of the heart, whereas the cardiac cushion develops from the endocardium beneath the atrioventricular canals and outflow tract myocardium [73].
Many transcriptional regulation in organogenesis involve members of the ancient family of T-box transcription factors, including the specification of cardiac chambers and the conduction system [74]. Herein, the T-box activators and repressors work together for the cardiac ballooning by inducing cardiac cushions (TBX2 and TBX3) limited to the atrioventricular canal (TBX20 and TBX5). Several circRNAs have been seen to co-express with TBX5, including circ-HIPK3, circ-PLOD2_1, circ-RHOBTB3, circ-PSMB1, circ-SLC8A1_1 and circ-MYH6/7_1. Similarly, the expression of TBX2 was seen to co-express with several lncRNAs and circRNAs, including TTN-AS1, RP11-617F23.1, circ-PHKB_1, circ-HIPK3, circ-SLC8A1_1, circ-MYH6/7_1 and circ-PALM2 [25]. Cells that are originating from TBX2-expressing prognitors will contribute to right and left ventricular walls [19], and the repressive interaction of Tbx20 upstream of Tbx2 underlie the primary lineage specification to chamber and non-chamber myocardium, thereby determining heart integrity and contractile function [75]. Moreover, the chamber formation is also mediated by the expression of several key regulators, including Gata4, Nkx2.5, Tbx5, dHand, eHand, Pitx2, MEF2C, and Irx4 [76]. Intriguingly, analysis of paired human atrial and ventricular samples revealed that 17–28% of the total lncRNA transcripts were differentially regulated in the four chambers, vastly attributed to their distinctive roles in cardiac functions [77]. The lncRNA uc.457 has also been associated with ventricular septal defect in human, and was recently revealed to regulate proliferation and differentiation of CMs by inhibiting the protein expression of histone cell cycle regulation defective homolog a, cardiac muscle troponin T, natriuretic peptide A and mef2C, respectively [78].
Chamber-specific expression of miRNA signatures in human heart has also been reported recently [79]. By performing miRNA deep sequencing, Kakimoto Y et al. revealed that the miRNA-1 was the most abundant in both atrial (21%) and ventricular (26%) chambers, and the miRNA-208 family showed prominent chamber specificity in the atrial (miRNA-208b-3p and miRNA-208a-3p) and ventricle (miRNA-208-3p and miRNA-208b-5p). In zebrafish, it has been shown that the miRNA-143-adducin3 is essential for chamber morphogenesis through direct inhibition of adducin3 which encodes an F-actin capping protein. Disruption of this miRNA led to ventricular collapse and decreased contractility [80]. The miRNA-138 is another molecule that is required to establish appropriate gene expression restricted to the atrio-ventricular valve region, and its dysregulation caused abnormal ventricular formation [81]. For cardiac valvulogenesis, Kopla HJ et al. reported that the miRNA-21 was necessary for proper development of the atrio-ventricular valve by repressing the tumor suppressor programmed cell death 4 (PDCD4b) expression, since miRNA-21 expression is known to be restricted to valvular endothelium and implicated in the response to several forms of cardiac stress [82].
During perinatal transition of heart, maturation of the cardiac tissue is required to warrant functional adaptation of the changes in nutrient environment and hemodynamic load after birth. The maturation and final septation of the heart requires, together with Gata4, Nkx2-5 and Tbx5, the expression of RxRa, FOG-2, Pitx2, Sox4, NF-Atc, TEF-1, Tbx1, Hey2, CITED, and ZIC3 [76]. At the cellular level, majority of CMs undergo dramatic changes in the morphology, proliferation, gene expression and metabolism. Therefore, any abberant transcriptional pertubation occur at this stage often lead to congenital heart defects. In fact, during CM maturation many lncRNAs are strictly regulated by maturation stage-specific transcription factors. For instances, it has been reported that approximately 70% of the lncRNAs that were highly expressed at CM maturation stage could bind to NFAT – an important CM maturation regulator when coupled with calcineurin [25]. Abnormal NFAT signaling causes pathological cardiac hypertrophy and heart failure. Of all lncRNAs, 90% of them are enriched for the MEIS1 motif which has been implicated in heart development [83].
Furthermore, a recent study reported a high-resolution landscape on neonatal cardiac lncRNAs interactions with neighboring transcriptomic molecules during cardiac maturation and postnatal stress in murine [8]. Specifically, the study revealed the Ppp1r1b-lncRNA as a regulator of its partner gene Tcap which encodes the muscle protein titin and the expression ratio of Ppp1r1b-lncRNA/Tcap could be used as a molecular signature for ventricular septum defect in human infantile hearts. Impuls conduction through the heart is the fundamental phenomenon of a synchronized muscle fiber contraction, proper transcriptional regulation of muscle fiber assembly and maturation is of quintessential. In this context, the cardiac conduction regulatory RNA (lncRNA-CCRR) was found to control cardiac conduction by promoting binding of connexin43 to the interacting protein CIP85. Silencing or knockdown lncRNA-CCRR causes malformation of intercalated discs and gap junctions that slow longitudinal cardiac conduction [84]. Other examples of lncRNAs that regulates CMs proliferation, differentiation and maturation includes uc. 40, uc.167, uc.245 and TUC40 [85,86,87,88].
In term of miRNA modulation, miRNA-27b has been reported to play critical roles in skeletal muscle development [89], and it is robustly expressed within the myocardium in the adult heart [90]. Via microarray analysis, Chinchilla A et al. found that relatively few miRNAs display discrete peak of decreasing or increasing expression profiles during ventricular maturation. In particular, the miRNA-27b (an early stage marker of ventricular chamber formation) displays an overt myocardial expression during cardiogenesis, and it regulates the cardiac myogenesis transcription factor – Mef2c without disturbing the expression of other cardiac genes [91]. This specific role of miRNA-27b on Mef2c suggests potential therapeutic for cardiac hypertrophy. Interestingly, the miRNA-27a exhibited a strongly upregulatory role on the β-MHC gene by targeting the thyroid hormone receptor β1 (TRβ1) in ventricular CMs [92]. The miRNA-143 plays an essential role in mechanotransduction pathway, in particular on circulatory adaptation and regulation between the outflow tracts and ventricles by suppressing retinoic acid signaling [93]. Besides miRNA itself, the miRNA-processing enzyme Dicer also plays a critical role in promoting cardiac outflow tract aligment and chamber septation by upregulating the morphogen Pitx2c and Sema3c. Due to impairment of miRNA processing at later-stage, cardiac-specific Dicer deficiency mice exhibited misexpression of cardiac contractile proteins and rapidly developed dilated cardiomyopathy, heart failure and postnatal lethality [94]. Moreover, the miRNA-208a is reported as a novel modulator of cardiac hypertrophy and electrical conduction. Overexpression of miRNA-208a (which is encoded within an intron of α-cardiac muscle myosin heavy chain gene (Myh6)) in mice induced muscle hypertrophy and arrhythmias, whereas sufficient level of miRNA-208a expression was required for proper cardiac conduction and the expression of cardiac genes such as GATA4 and connexin 40 [95].
The miRNAs also play important roles in cardiac extracellular matrix remodeling. For instances, the miRNA-133 and miRNA-30 were reported to directly downregulate connective tissue growth factor (CTGF), which is a key molecule in maintaining proper extracellular matrix remodeling in myocardium [96]. Overexpression of these miRNAs resulted in low CTGF level accompanied by decreased production of collagen, whereas knocking down their expression causing cardiac fibrosis. Furthermore, in CMs derived from rats at 4 weeks, the miRNA-29a was found to be differentially upregulated which inversely regulated CMs proliferation by targeting to Cyclin D2 (CCND2) [97]. This finding suggest an inhibition role of miRNAs in CMs proliferation during postnatal development. The circRNAs play a critical role in cardiac cell specification from cardiac progenitor cells to CMs. It is reported that nearly 500 and 200 circRNAs were positively (e.g. circ-SLC8A1-1, circ-TTN-275, and circ-ALPK2-1) and negatively (e.g. circ-DNMT3B-4, circ-OSBPL10 and circ-FGD4-7) correlated to the differentiation of human embryonic stem cells to CMs [13, 18]. Of interest, the circ-TTN was differentially expressed in neonatal and adult rat hearts [70] and revealed to be co-expressed with MYL4 – mutation of which leads to abberant sarcomere formation, atrial enlargement and fibrillation [98]. By circRNA profiling, circ-TTN expression was dynamically regulated in mice with dilated cardiomyopathy, and largely downregulated in mice lacking the RNA-binding motif protein 20 (RBM20), suggesting a novel mechanistic insights for diated cardiomyopathy [99]. Expression of other circRNAs, such as circ-SLC8A1, circ-CHD7, circ-ATXN10 and circ-DNAJC6 was also found to be prominent in patients with dilated cardiomyopathy [100].
4 Future Perspectives
ncRNAs have gained interest in the past decades due to their role in modulating cell fate at a post-transcriptional level. The modulation occurring in the prenatal life at the embryo level helps us shedding a light on the tuning required for the proper formation of the heart and the other organs. More importantly, it gives us the possibility to better understand how congenital heart diseases occur.
Potentially, ncRNAs could be used both for diagnostic and therapeutic purposes. In this view, the fact that circRNA concentration profiles change during cardiac development and disease independently from their host gene expression, they represent novel and more stable biological markers. Although they are still at their infancy, artificial circRNAs similarly to miRNA mimics and antagomirs could represent promising tools in regenerative medicine since they are highly stable and can regulate a wide range of cellular functions. Recent studies have highlighted the extensive network of interactions among microRNAs, lncRNAs and circRNAs, forming crucial regulatory axis participating in the modulation of cardiac differentiation [101].
In order to obtain mature CMs from induced pluripotent stem cells (iPSCs), Miyamoto has recently shown that the use of Gata4-Mef2c-Tbx5, or GMT, led to the correct formation of cardiac cells [102, 103]. Emerging literature is showing the cardiac differentiation potential of PSCs but also the limitations to generate fully mature CMs to model cardiac diseases or for drug screening porpuses. Specific ncRNAs control and promote the differentiation of PSCs and mesodermal progenitors into CMs and the use of microvesicles to transfer those peculiar ncRNAs is a fascinating possibility to better coordinate cardiogenic maturation of healthy and pathological progenitor cells. This will be critical to better understand the role of ncRNAs in the regulation of cardiovascular system development and eventually in the progression of cardiovascular disease.
World-leading laboratories are investing in gene editing, mainly in CRISPR/Cas9 technology, to edit efficiently any genomic locus with high DNA sequence specificity and possibly without undesired byproducts. However, CRISPR/Cas9 technology is still a very young gene-editing technology that can result in off-target effects with unexpected consequence and the long-term impact of genetic alteration on future generations is yet unknown. In addition, small indels generated by CRISPR/Cas9 system can alter or prevent functional modifications of ncRNAs or affect overlapping/adjacent genes in loci characterized by bidirectional promoters or sense/antisense genes (where lncRNAs are generated) [104]. Although a prudent path should be considered for CRISPR/Cas9-based in vivo applications, these novel gene editing approaches will allow us to perform more precise perturbation studies to uncover the basic principles of cardiac development and better collocate transcription factors, ncRNA networks and molecular pathways that contribute to CM maturation (Table 11.1).
Abbreviations
- circRNAs:
-
circular RNAs
- CS:
-
Carnegie Stage
- dpc:
-
days post coitum
- ESCs:
-
embryonal stem cells
- FHF:
-
first heart field
- lncRNAs:
-
long non-coding RNAs
- miRNAs:
-
microRNAs
- ncRNAs:
-
non-coding RNAs
- RNA-seq:
-
RNA sequencing
- SHF:
-
secondary heart field
References
Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009;10(3):155–9.
Mercer TR, Mattick JS. Structure and function of long noncoding RNAs in epigenetic regulation. Nat Struct Mol Biol. 2013;20(3):300–7.
Kurian L, Aguirre A, Sancho-Martinez I, Benner C, Hishida T, Nguyen TB, Reddy P, Nivet E, Krause MN, Nelles DA, Esteban CR, Campistol JM, Yeo GW, Belmonte JCI. Identification of novel long noncoding RNAs underlying vertebrate cardiovascular development. Circulation. 2015;131(14):1278–90.
Matkovich SJ, Edwards JR, Grossenheider TC, de Guzman Strong C, Dorn GW 2nd. Epigenetic coordination of embryonic heart transcription by dynamically regulated long noncoding RNAs. Proc Natl Acad Sci U S A. 2014;111(33):12264–9.
Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C, Kodzius R, Shimokawa K, Bajic VB, Brenner SE, Batalov S, Forrest AR, Zavolan M, Davis MJ, Wilming LG, Aidinis V, Allen JE, Ambesi-Impiombato A, Apweiler R, Aturaliya RN, Bailey TL, Bansal M, Baxter L, Beisel KW, Bersano T, Bono H, Chalk AM, Chiu KP, Choudhary V, Christoffels A, Clutterbuck DR, Crowe ML, Dalla E, Dalrymple BP, de Bono B, Della Gatta G, di Bernardo D, Down T, Engstrom P, Fagiolini M, Faulkner G, Fletcher CF, Fukushima T, Furuno M, Futaki S, Gariboldi M, Georgii-Hemming P, Gingeras TR, Gojobori T, Green RE, Gustincich S, Harbers M, Hayashi Y, Hensch TK, Hirokawa N, Hill D, Huminiecki L, Iacono M, Ikeo K, Iwama A, Ishikawa T, Jakt M, Kanapin A, Katoh M, Kawasawa Y, Kelso J, Kitamura H, Kitano H, Kollias G, Krishnan SP, Kruger A, Kummerfeld SK, Kurochkin IV, Lareau LF, Lazarevic D, Lipovich L, Liu J, Liuni S, McWilliam S, Madan Babu M, Madera M, Marchionni L, Matsuda H, Matsuzawa S, Miki H, Mignone F, Miyake S, Morris K, Mottagui-Tabar S, Mulder N, Nakano N, Nakauchi H, Ng P, Nilsson R, Nishiguchi S, Nishikawa S, Nori F, Ohara O, Okazaki Y, Orlando V, Pang KC, Pavan WJ, Pavesi G, Pesole G, Petrovsky N, Piazza S, Reed J, Reid JF, Ring BZ, Ringwald M, Rost B, Ruan Y, Salzberg SL, Sandelin A, Schneider C, Schonbach C, Sekiguchi K, Semple CA, Seno S, Sessa L, Sheng Y, Shibata Y, Shimada H, Shimada K, Silva D, Sinclair B, Sperling S, Stupka E, Sugiura K, Sultana R, Takenaka Y, Taki K, Tammoja K, Tan SL, Tang S, Taylor MS, Tegner J, Teichmann SA, Ueda HR, van Nimwegen E, Verardo R, Wei CL, Yagi K, Yamanishi H, Zabarovsky E, Zhu S, Zimmer A, Hide W, Bult C, Grimmond SM, Teasdale RD, Liu ET, Brusic V, Quackenbush J, Wahlestedt C, Mattick JS, Hume DA, Kai C, Sasaki D, Tomaru Y, Fukuda S, Kanamori-Katayama M, Suzuki M, Aoki J, Arakawa T, Iida J, Imamura K, Itoh M, Kato T, Kawaji H, Kawagashira N, Kawashima T, Kojima M, Kondo S, Konno H, Nakano K, Ninomiya N, Nishio T, Okada M, Plessy C, Shibata K, Shiraki T, Suzuki S, Tagami M, Waki K, Watahiki A, Okamura-Oho Y, Suzuki H, Kawai J, Hayashizaki Y, Consortium F, Group RGER, Genome Science G. The transcriptional landscape of the mammalian genome. Science. 2005;309(5740):1559–63.
Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermuller J, Hofacker IL, Bell I, Cheung E, Drenkow J, Dumais E, Patel S, Helt G, Ganesh M, Ghosh S, Piccolboni A, Sementchenko V, Tammana H, Gingeras TR. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science. 2007;316(5830):1484–8.
Scheuermann JC, Boyer LA. Getting to the heart of the matter: long non-coding RNAs in cardiac development and disease. EMBO J. 2013;32(13):1805–16.
Touma M, Kang X, Zhao Y, Cass AA, Gao F, Biniwale R, Coppola G, Xiao X, Reemtsen B, Wang Y. Decoding the long noncoding RNA during cardiac maturation: a roadmap for functional discovery. Circ Cardiovasc Genet. 2016;9(5):395–407.
Devaux Y, Zangrando J, Schroen B, Creemers EE, Pedrazzini T, Chang CP, Dorn GW 2nd, Thum T, Heymans S, Cardiolinc network. Long noncoding RNAs in cardiac development and ageing. Nat Rev Cardiol. 2015;12(7):415–25.
Rotini A, Martinez-Sarra E, Pozzo E, Sampaolesi M. Interactions between microRNAs and long non-coding RNAs in cardiac development and repair. Pharmacol Res. 2018;127:58–66.
Quattrocelli M, Sampaolesi M. The mesmiRizing complexity of microRNAs for striated muscle tissue engineering. Adv Drug Deliv Rev. 2015;88:37–52.
Zhao Y, Ransom JF, Li A, Vedantham V, von Drehle M, Muth AN, Tsuchihashi T, McManus MT, Schwartz RJ, Srivastava D. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell. 2007;129(2):303–17.
Bei Y, Yang T, Wang L, Holvoet P, Das S, Sluijter JPG, Monteiro MC, Liu Y, Zhou Q, Xiao J. Circular RNAs as potential Theranostics in the cardiovascular system. Mol Ther Nucleic Acids. 2018;13:407–18.
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, Marzluff WF, Sharpless NE. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19(2):141–57.
Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, Zhong G, Yu B, Hu W, Dai L, Zhu P, Chang Z, Wu Q, Zhao Y, Jia Y, Xu P, Liu H, Shan G. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015;22(3):256–64.
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, Loewer A, Ziebold U, Landthaler M, Kocks C, le Noble F, Rajewsky N. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495(7441):333–8.
Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, Evantal N, Memczak S, Rajewsky N, Kadener S. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell. 2014;56(1):55–66.
Tan WL, Lim BT, Anene-Nzelu CG, Ackers-Johnson M, Dashi A, See K, Tiang Z, Lee DP, Chua WW, Luu TD, Li PY, Richards AM, Foo RS. A landscape of circular RNA expression in the human heart. Cardiovasc Res. 2017;113(3):298–309.
Aanhaanen WT, Brons JF, Dominguez JN, Rana MS, Norden J, Airik R, Wakker V, de Gier-de Vries C, Brown NA, Kispert A, Moorman AF, Christoffels VM. The Tbx2+ primary myocardium of the atrioventricular canal forms the atrioventricular node and the base of the left ventricle. Circ Res. 2009;104(11):1267–74.
Gunthel M, Barnett P, Christoffels VM. Development, proliferation, and growth of the mammalian heart. MolTher. 2018;26(7):1599–609.
Costello I, Pimeisl IM, Drager S, Bikoff EK, Robertson EJ, Arnold SJ. The T-box transcription factor Eomesodermin acts upstream of Mesp1 to specify cardiac mesoderm during mouse gastrulation. Nat Cell Biol. 2011;13(9):1084–91.
van den Ameele J, Tiberi L, Bondue A, Paulissen C, Herpoel A, Iacovino M, Kyba M, Blanpain C, Vanderhaeghen P. Eomesodermin induces Mesp1 expression and cardiac differentiation from embryonic stem cells in the absence of Activin. EMBO Rep. 2012;13(4):355–62.
Guo X, Xu Y, Wang Z, Wu Y, Chen J, Wang G, Lu C, Jia W, Xi J, Zhu S, Jiapaer Z, Wan X, Liu Z, Gao S, Kang J. A Linc1405/Eomes complex promotes cardiac mesoderm specification and Cardiogenesis. Cell Stem Cell. 2018;22(6):893–908.. e896
Grote P, Wittler L, Hendrix D, Koch F, Wahrisch S, Beisaw A, Macura K, Blass G, Kellis M, Werber M, Herrmann BG. The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev Cell. 2013;24(2):206–14.
Li Y, Zhang J, Huo C, Ding N, Li J, Xiao J, Lin X, Cai B, Zhang Y, Xu J. Dynamic organization of lncRNA and circular RNA regulators collectively controlled cardiac differentiation in humans. EBioMedicine. 2017;24:137–46.
Bondue A, Tannler S, Chiapparo G, Chabab S, Ramialison M, Paulissen C, Beck B, Harvey R, Blanpain C. Defining the earliest step of cardiovascular progenitor specification during embryonic stem cell differentiation. J Cell Biol. 2011;192(5):751–65.
Lescroart F, Chabab S, Lin X, Rulands S, Paulissen C, Rodolosse A, Auer H, Achouri Y, Dubois C, Bondue A, Simons BD, Blanpain C. Early lineage restriction in temporally distinct populations of Mesp1 progenitors during mammalian heart development. Nat Cell Biol. 2014;16(9):829–40.
Devine WP, Wythe JD, George M, Koshiba-Takeuchi K, Bruneau BG. Early patterning and specification of cardiac progenitors in gastrulating mesoderm. elife. 2014;3:e03848.
Ivanovitch K, Temino S, Torres M. Live imaging of heart tube development in mouse reveals alternating phases of cardiac differentiation and morphogenesis. elife. 2017;6:e30668.
Tsuchihashi T, Maeda J, Shin CH, Ivey KN, Black BL, Olson EN, Yamagishi H, Srivastava D. Hand2 function in second heart field progenitors is essential for cardiogenesis. Dev Biol. 2011;351(1):62–9.
Greulich F, Rudat C, Kispert A. Mechanisms of T-box gene function in the developing heart. Cardiovasc Res. 2011;91(2):212–22.
Laugwitz KL, Moretti A, Caron L, Nakano A, Chien KR. Islet1 cardiovascular progenitors: a single source for heart lineages? Development. 2008;135(2):193–205.
Dodou E, Verzi MP, Anderson JP, Xu SM, Black BL. Mef2c is a direct transcriptional target of ISL1 and GATA factors in the anterior heart field during mouse embryonic development. Development. 2004;131(16):3931–42.
Molkentin JD, Lin Q, Duncan SA, Olson EN. Requirement of the transcription factor GATA4 for heart tube formation and ventral morphogenesis. Genes Dev. 1997;11(8):1061–72.
Harrelson Z, Kelly RG, Goldin SN, Gibson-Brown JJ, Bollag RJ, Silver LM, Papaioannou VE. Tbx2 is essential for patterning the atrioventricular canal and for morphogenesis of the outflow tract during heart development. Development. 2004;131(20):5041–52.
van den Berg G, Abu-Issa R, de Boer BA, Hutson MR, de Boer PA, Soufan AT, Ruijter JM, Kirby ML, van den Hoff MJ, Moorman AF. A caudal proliferating growth center contributes to both poles of the forming heart tube. Circ Res. 2009;104(2):179–88.
Kwon C, Arnold J, Hsiao EC, Taketo MM, Conklin BR, Srivastava D. Canonical Wnt signaling is a positive regulator of mammalian cardiac progenitors. Proc Natl Acad Sci U S A. 2007;104(26):10894–9.
High FA, Jain R, Stoller JZ, Antonucci NB, Lu MM, Loomes KM, Kaestner KH, Pear WS, Epstein JA. Murine Jagged1/Notch signaling in the second heart field orchestrates Fgf8 expression and tissue-tissue interactions during outflow tract development. J Clin Invest. 2009;119(7):1986–96.
Rochais F, Dandonneau M, Mesbah K, Jarry T, Mattei MG, Kelly RG. Hes1 is expressed in the second heart field and is required for outflow tract development. PLoS One. 2009;4(7):e6267.
Shen X, Soibam B, Benham A, Xu X, Chopra M, Peng X, Yu W, Bao W, Liang R, Azares A, Liu P, Gunaratne PH, Mercola M, Cooney AJ, Schwartz RJ, Liu Y. miR-322/−503 cluster is expressed in the earliest cardiac progenitor cells and drives cardiomyocyte specification. Proc Natl Acad Sci U S A. 2016;113(34):9551–6.
Ounzain S, Micheletti R, Arnan C, Plaisance I, Cecchi D, Schroen B, Reverter F, Alexanian M, Gonzales C, Ng SY, Bussotti G, Pezzuto I, Notredame C, Heymans S, Guigo R, Johnson R, Pedrazzini T. CARMEN, a human super enhancer-associated long noncoding RNA controlling cardiac specification, differentiation and homeostasis. J Mol Cell Cardiol. 2015;89(Pt A):98–112.
Ounzain S, Micheletti R, Beckmann T, Schroen B, Alexanian M, Pezzuto I, Crippa S, Nemir M, Sarre A, Johnson R, Dauvillier J, Burdet F, Ibberson M, Guigo R, Xenarios I, Heymans S, Pedrazzini T. Genome-wide profiling of the cardiac transcriptome after myocardial infarction identifies novel heart-specific long non-coding RNAs. Eur Heart J. 2015;36(6):353–368a.
Saga Y. Genetic rescue of segmentation defect in MesP2-deficient mice by MesP1 gene replacement. Mech Dev. 1998;75(1–2):53–66.
Saga Y, Miyagawa-Tomita S, Takagi A, Kitajima S, Miyazaki J, Inoue T. MesP1 is expressed in the heart precursor cells and required for the formation of a single heart tube. Development. 1999;126(15):3437–47.
Chiapparo G, Lin X, Lescroart F, Chabab S, Paulissen C, Pitisci L, Bondue A, Blanpain C. Mesp1 controls the speed, polarity, and directionality of cardiovascular progenitor migration. J Cell Biol. 2016;213(4):463–77.
Liberatore CM, Searcy-Schrick RD, Yutzey KE. Ventricular expression of tbx5 inhibits normal heart chamber development. Dev Biol. 2000;223(1):169–80.
Takeuchi JK, Bruneau BG. Directed transdifferentiation of mouse mesoderm to heart tissue by defined factors. Nature. 2009;459(7247):708–11.
Fulcoli FG, Huynh T, Scambler PJ, Baldini A. Tbx1 regulates the BMP-Smad1 pathway in a transcription independent manner. PLoS One. 2009;4(6):e6049.
Chen L, Fulcoli FG, Tang S, Baldini A. Tbx1 regulates proliferation and differentiation of multipotent heart progenitors. Circ Res. 2009;105(9):842–51.
Pane LS, Zhang Z, Ferrentino R, Huynh T, Cutillo L, Baldini A. Tbx1 is a negative modulator of Mef2c. Hum Mol Genet. 2012;21(11):2485–96.
Wang J, Greene SB, Bonilla-Claudio M, Tao Y, Zhang J, Bai Y, Huang Z, Black BL, Wang F, Martin JF. Bmp signaling regulates myocardial differentiation from cardiac progenitors through a MicroRNA-mediated mechanism. Dev Cell. 2010;19(6):903–12.
Wilson KD, Hu S, Venkatasubrahmanyam S, Fu JD, Sun N, Abilez OJ, Baugh JJ, Jia F, Ghosh Z, Li RA, Butte AJ, Wu JC. Dynamic microRNA expression programs during cardiac differentiation of human embryonic stem cells: role for miR-499. Circ Cardiovasc Genet. 2010;3(5):426–35.
Cai CL, Liang X, Shi Y, Chu PH, Pfaff SL, Chen J, Evans S. Isl1 identifies a cardiac progenitor population that proliferates prior to differentiation and contributes a majority of cells to the heart. Dev Cell. 2003;5(6):877–89.
Engleka KA, Manderfield LJ, Brust RD, Li L, Cohen A, Dymecki SM, Epstein JA. Islet1 derivatives in the heart are of both neural crest and second heart field origin. Circ Res. 2012;110(7):922–6.
Dorn T, Goedel A, Lam JT, Haas J, Tian Q, Herrmann F, Bundschu K, Dobreva G, Schiemann M, Dirschinger R, Guo Y, Kuhl SJ, Sinnecker D, Lipp P, Laugwitz KL, Kuhl M, Moretti A. Direct nkx2-5 transcriptional repression of isl1 controls cardiomyocyte subtype identity. Stem Cells. 2015;33(4):1113–29.
Anderson KM, Anderson DM, McAnally JR, Shelton JM, Bassel-Duby R, Olson EN. Transcription of the non-coding RNA upperhand controls Hand2 expression and heart development. Nature. 2016;539(7629):433–6.
Han X, Zhang J, Liu Y, Fan X, Ai S, Luo Y, Li X, Jin H, Luo S, Zheng H, Yue Y, Chang Z, Yang Z, Tang F, He A, Shen X. The lncRNA Hand2os1/Uph locus orchestrates heart development through regulation of precise expression of Hand2. Development. 2019;146(13):dev176198.
Huang J, Elicker J, Bowens N, Liu X, Cheng L, Cappola TP, Zhu X, Parmacek MS. Myocardin regulates BMP10 expression and is required for heart development. J Clin Invest. 2012;122(10):3678–91.
Belian E, Noseda M, Abreu Paiva MS, Leja T, Sampson R, Schneider MD. Forward programming of cardiac stem cells by homogeneous transduction with MYOCD plus TBX5. PLoS One. 2015;10(6):e0125384.
Wang K, Long B, Liu F, Wang JX, Liu CY, Zhao B, Zhou LY, Sun T, Wang M, Yu T, Gong Y, Liu J, Dong YH, Li N, Li PF. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur Heart J. 2016;37(33):2602–11.
Wystub K, Besser J, Bachmann A, Boettger T, Braun T. miR-1/133a clusters cooperatively specify the cardiomyogenic lineage by adjustment of myocardin levels during embryonic heart development. PLoS Genet. 2013;9(9):e1003793.
Sahoo S, Meijles DN, Al Ghouleh I, Tandon M, Cifuentes-Pagano E, Sembrat J, Rojas M, Goncharova E, Pagano PJ. MEF2C-MYOCD and Leiomodin1 suppression by miRNA-214 promotes smooth muscle cell phenotype switching in pulmonary arterial hypertension. PLoS One. 2016;11(5):e0153780.
Wu Y, Li Z, Yang M, Dai B, Hu F, Yang F, Zhu J, Chen T, Zhang L. MicroRNA-214 regulates smooth muscle cell differentiation from stem cells by targeting RNA-binding protein QKI. Oncotarget. 2017;8(12):19866–78.
Huang J, Min Lu M, Cheng L, Yuan LJ, Zhu X, Stout AL, Chen M, Li J, Parmacek MS. Myocardin is required for cardiomyocyte survival and maintenance of heart function. Proc Natl Acad Sci U S A. 2009;106(44):18734–9.
Sluijter JP, van Mil A, van Vliet P, Metz CH, Liu J, Doevendans PA, Goumans MJ. MicroRNA-1 and -499 regulate differentiation and proliferation in human-derived cardiomyocyte progenitor cells. Arterioscler Thromb Vasc Biol. 2010;30(4):859–68.
Liu N, Bezprozvannaya S, Williams AH, Qi X, Richardson JA, Bassel-Duby R, Olson EN. microRNA-133a regulates cardiomyocyte proliferation and suppresses smooth muscle gene expression in the heart. Genes Dev. 2008;22(23):3242–54.
Ivey KN, Muth A, Arnold J, King FW, Yeh RF, Fish JE, Hsiao EC, Schwartz RJ, Conklin BR, Bernstein HS, Srivastava D. MicroRNA regulation of cell lineages in mouse and human embryonic stem cells. Cell Stem Cell. 2008;2(3):219–29.
Takaya T, Ono K, Kawamura T, Takanabe R, Kaichi S, Morimoto T, Wada H, Kita T, Shimatsu A, Hasegawa K. MicroRNA-1 and MicroRNA-133 in spontaneous myocardial differentiation of mouse embryonic stem cells. Circ J. 2009;73(8):1492–7.
Xu T, Wu J, Han P, Zhao Z, Song X. Circular RNA expression profiles and features in human tissues: a study using RNA-seq data. BMC Genomics. 2017;18(Suppl 6):680.
Werfel S, Nothjunge S, Schwarzmayr T, Strom TM, Meitinger T, Engelhardt S. Characterization of circular RNAs in human, mouse and rat hearts. J Mol Cell Cardiol. 2016;98:103–7.
Lei W, Feng T, Fang X, Yu Y, Yang J, Zhao ZA, Liu J, Shen Z, Deng W, Hu S. Signature of circular RNAs in human induced pluripotent stem cells and derived cardiomyocytes. Stem Cell Res Ther. 2018;9(1):56.
Du WW, Yang W, Chen Y, Wu ZK, Foster FS, Yang Z, Li X, Yang BB. Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur Heart J. 2017;38(18):1402–12.
Kelly RG, Buckingham ME, Moorman AF. Heart fields and cardiac morphogenesis. Cold Spring Harb Perspect Med. 2014;4(10):a015750.
Stennard FA, Harvey RP. T-box transcription factors and their roles in regulatory hierarchies in the developing heart. Development. 2005;132(22):4897–910.
Stennard FA, Costa MW, Lai D, Biben C, Furtado MB, Solloway MJ, McCulley DJ, Leimena C, Preis JI, Dunwoodie SL, Elliott DE, Prall OW, Black BL, Fatkin D, Harvey RP. Murine T-box transcription factor Tbx20 acts as a repressor during heart development, and is essential for adult heart integrity, function and adaptation. Development. 2005;132(10):2451–62.
Chen H, VanBuren V. A provisional gene regulatory atlas for mouse heart development. PLoS One. 2014;9(1):e83364.
Johnson EK, Matkovich SJ, Nerbonne JM. Regional differences in mRNA and lncRNA expression profiles in non-failing human atria and ventricles. Sci Rep. 2018;8(1):13919.
Zhang Q, Cheng Z, Yu Z, Zhu C, Qian L. Role of lncRNA uc.457 in the differentiation and maturation of cardiomyocytes. Mol Med Rep. 2019;19(6):4927–34.
Kakimoto Y, Tanaka M, Kamiguchi H, Hayashi H, Ochiai E, Osawa M. MicroRNA deep sequencing reveals chamber-specific miR-208 family expression patterns in the human heart. Int J Cardiol. 2016;211:43–8.
Deacon DC, Nevis KR, Cashman TJ, Zhou Y, Zhao L, Washko D, Guner-Ataman B, Burns CG, Burns CE. The miR-143-adducin3 pathway is essential for cardiac chamber morphogenesis. Development. 2010;137(11):1887–96.
Morton SU, Scherz PJ, Cordes KR, Ivey KN, Stainier DY, Srivastava D. microRNA-138 modulates cardiac patterning during embryonic development. Proc Natl Acad Sci U S A. 2008;105(46):17830–5.
Kolpa HJ, Peal DS, Lynch SN, Giokas AC, Ghatak S, Misra S, Norris RA, Macrae CA, Markwald RR, Ellinor P, Bischoff J, Milan DJ. miR-21 represses Pdcd4 during cardiac valvulogenesis. Development. 2013;140(10):2172–80.
Wamstad JA, Alexander JM, Truty RM, Shrikumar A, Li F, Eilertson KE, Ding H, Wylie JN, Pico AR, Capra JA, Erwin G, Kattman SJ, Keller GM, Srivastava D, Levine SS, Pollard KS, Holloway AK, Boyer LA, Bruneau BG. Dynamic and coordinated epigenetic regulation of developmental transitions in the cardiac lineage. Cell. 2012;151(1):206–20.
Zhang Y, Sun L, Xuan L, Pan Z, Hu X, Liu H, Bai Y, Jiao L, Li Z, Cui L, Wang X, Wang S, Yu T, Feng B, Guo Y, Liu Z, Meng W, Ren H, Zhu J, Zhao X, Yang C, Zhang Y, Xu C, Wang Z, Lu Y, Shan H, Yang B. Long non-coding RNA CCRR controls cardiac conduction via regulating intercellular coupling. Nat Commun. 2018;9(1):4176.
Li H, Jiang L, Yu Z, Han S, Liu X, Li M, Zhu C, Qiao L, Huang L. The role of a novel Long noncoding RNA TUC40- in Cardiomyocyte induction and maturation in P19 cells. Am J Med Sci. 2017;354(6):608–16.
Liu H, Hu Y, Yin J, Yan X, Chen W, Wang X, Han S, Yu Z, Li M. Effects of long non-coding RNA uc.245 on cardiomyocyte-like differentiation in P19 cells via FOG2. Gene. 2019;694:83–92.
Song G, Shen Y, Ruan Z, Li X, Chen Y, Yuan W, Ding X, Zhu L, Qian L. LncRNA-uc.167 influences cell proliferation, apoptosis and differentiation of P19 cells by regulating Mef2c. Gene. 2016;590(1):97–108.
Wu R, Xue P, Wan Y, Wang S, Gu M. LncRNA-uc.40 silence promotes P19 embryonic cells differentiation to cardiomyocyte via the PBX1 gene. In Vitro Cell Dev Biol Anim. 2018;54(8):600–9.
Crist CG, Montarras D, Pallafacchina G, Rocancourt D, Cumano A, Conway SJ, Buckingham M. Muscle stem cell behavior is modified by microRNA-27 regulation of Pax3 expression. Proc Natl Acad Sci U S A. 2009;106(32):13383–7.
Hernandez-Torres F, Martinez-Fernandez S, Zuluaga S, Nebreda A, Porras A, Aranega AE, Navarro F. A role for p38alpha mitogen-activated protein kinase in embryonic cardiac differentiation. FEBS Lett. 2008;582(7):1025–31.
Chinchilla A, Lozano E, Daimi H, Esteban FJ, Crist C, Aranega AE, Franco D. MicroRNA profiling during mouse ventricular maturation: a role for miR-27 modulating Mef2c expression. Cardiovasc Res. 2011;89(1):98–108.
Nishi H, Ono K, Horie T, Nagao K, Kinoshita M, Kuwabara Y, Watanabe S, Takaya T, Tamaki Y, Takanabe-Mori R, Wada H, Hasegawa K, Iwanaga Y, Kawamura T, Kita T, Kimura T. MicroRNA-27a regulates beta cardiac myosin heavy chain gene expression by targeting thyroid hormone receptor beta1 in neonatal rat ventricular myocytes. Mol Cell Biol. 2011;31(4):744–55.
Miyasaka KY, Kida YS, Banjo T, Ueki Y, Nagayama K, Matsumoto T, Sato M, Ogura T. Heartbeat regulates cardiogenesis by suppressing retinoic acid signaling via expression of miR-143. Mech Dev. 2011;12(1–2):18–28.
Saxena A, Tabin CJ. miRNA-processing enzyme dicer is necessary for cardiac outflow tract alignment and chamber septation. Proc Natl Acad Sci U S A. 2010;107(1):87–91.
Callis TE, Pandya K, Seok HY, Tang RH, Tatsuguchi M, Huang ZP, Chen JF, Deng Z, Gunn B, Shumate J, Willis MS, Selzman CH, Wang DZ. MicroRNA-208a is a regulator of cardiac hypertrophy and conduction in mice. J Clin Invest. 2009;119(9):2772–86.
Duisters RF, Tijsen AJ, Schroen B, Leenders JJ, Lentink V, van der Made I, Herias V, van Leeuwen RE, Schellings MW, Barenbrug P, Maessen JG, Heymans S, Pinto YM, Creemers EE. miR-133 and miR-30 regulate connective tissue growth factor: implications for a role of microRNAs in myocardial matrix remodeling. Circ Res. 2009;104(2):170–8.. 176p following 178
Cao X, Wang J, Wang Z, Du J, Yuan X, Huang W, Meng J, Gu H, Nie Y, Ji B, Hu S, Zheng Z. MicroRNA profiling during rat ventricular maturation: a role for miR-29a in regulating cardiomyocyte cell cycle re-entry. FEBS Lett. 2013;587(10):1548–55.
Orr N, Arnaout R, Gula LJ, Spears DA, Leong-Sit P, Li Q, Tarhuni W, Reischauer S, Chauhan VS, Borkovich M, Uppal S, Adler A, Coughlin SR, Stainier DYR, Gollob MH. A mutation in the atrial-specific myosin light chain gene (MYL4) causes familial atrial fibrillation. Nat Commun. 2016;7:11303.
Khan MA, Reckman YJ, Aufiero S, van den Hoogenhof MM, van der Made I, Beqqali A, Koolbergen DR, Rasmussen TB, van der Velden J, Creemers EE, Pinto YM. RBM20 regulates circular RNA production from the Titin gene. Circ Res. 2016;119(9):996–1003.
Siede D, Rapti K, Gorska AA, Katus HA, Altmuller J, Boeckel JN, Meder B, Maack C, Volkers M, Muller OJ, Backs J, Dieterich C. Identification of circular RNAs with host gene-independent expression in human model systems for cardiac differentiation and disease. J Mol Cell Cardiol. 2017;109:48–56.
Aufiero S, Reckman YJ, Pinto YM, Creemers EE. Circular RNAs open a new chapter in cardiovascular biology. Nat Rev Cardiol. 2019;16(8):503–14.
Miyamoto K, Akiyama M, Tamura F, Isomi M, Yamakawa H, Sadahiro T, Muraoka N, Kojima H, Haginiwa S, Kurotsu S, Tani H, Wang L, Qian L, Inoue M, Ide Y, Kurokawa J, Yamamoto T, Seki T, Aeba R, Yamagishi H, Fukuda K, Ieda M. Direct in vivo reprogramming with Sendai virus vectors improves cardiac function after myocardial infarction. Cell Stem Cell. 2018;22(1):91–103.. e105
Sampaolesi M, Pozzo E, Duelen R. In the heart of the in vivo reprogramming. Stem Cell Investig. 2018;5:38.
Yang J, Meng X, Pan J, Jiang N, Zhou C, Wu Z, Gong Z. CRISPR/Cas9-mediated noncoding RNA editing in human cancers. RNA Biol. 2018;15(1):35–43.
Acknowledgements
Funding: Fonds Wetenschappelijk Onderzoek (#G088715N and #G0D4517N to MS, #1S90718N to EP), CARIPLO Foundation #2015_0634 and C1-KUL 3DMUSYC (#C14/17/111) grants.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Pozzo, E., Chai, Y.C., Sampaolesi, M. (2020). Comprehensive Overview of Non-coding RNAs in Cardiac Development. In: Xiao, J. (eds) Non-coding RNAs in Cardiovascular Diseases. Advances in Experimental Medicine and Biology, vol 1229. Springer, Singapore. https://doi.org/10.1007/978-981-15-1671-9_11
Download citation
DOI: https://doi.org/10.1007/978-981-15-1671-9_11
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-1670-2
Online ISBN: 978-981-15-1671-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)