Abstract
Actinomycetes strain VSM-1 isolated from the marine sediment samples of north coastal Bay of Bengal was identified as Streptomyces nanhaiensis by conventional and molecular approaches. S. nanhaiensis recorded significant broad-spectrum activity against all human and plant pathogenic microorganisms tested. An attempt has been made to evaluate the influence of medium components on the antimicrobial metabolite production of VSM-1 and to optimize an effective fermentation medium to enhance its bioactive metabolite yield. Bioactive metabolite production by VSM-1 was initially optimized using one-factor-at-a-time (OFAT) method where the yield of metabolites was recorded to generate inhibition zones of 30, 29, 28, 27, and 29 mm against test microorganisms, i.e., Shigella flexneri, Serratia marcescens, Proteus vulgaris, Pseudomonas aeruginosa, and medically important dermatophyte Candida albicans. The optimal values obtained from OFAT were selected, and the experimental model was designed using response surface methodology. Central composite design (CCD) was employed to study the influence of the variables on the production of bioactive metabolites by VSM-1 and their effect on the responses (test microorganisms). The statistical analysis showed that the variables which have a significant effect (P < 0.001) on the metabolite production at both the interactive and individual levels were measured in terms of antimicrobial activity. Maximum yield of metabolites was recorded to generate increased zones of inhibition, i.e., 34.2, 32, 31.6, 32.3, and 33.6 mm, against test microorganisms Shigella flexneri, Serratia marcescens, Proteus vulgaris, Pseudomonas aeruginosa, and Candida albicans, respectively. Unstructured kinetic models were used to fit the results obtained from experiments, and kinetic parameters were also estimated. Further, chemometric profile of the ethyl acetate extract of the strain was performed by gas chromatography and mass spectroscopy (GC-MS). Interpretation on mass spectrum of GC-MS was carried out using NIST Mass Spectral Data Base, with NIST MS search v.2.0. The results of the present study revealed the presence of various active principles with a diverse range of positive pharmacological actions, and hence this strain could be a possible source of novel bioactive compounds.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Natural products and their derivatives have been deep rooted as precious source of therapeutic agents. The natural product structures have the traits of excessive chemical diversity, biochemical specificity, and adequate amount of chiral centers, aggravated steric complexity, and exclusive molecular properties that make them favorable lead structures for drug discovery and differentiate them from synthetic and combinatorial compounds. Natural products have been deemed to be a natural combinatorial chemical library which could provide unrivaled compounds with chemical diversity and proper pharmacological chemical profiles (Bredholdt et al. 2007). The abundance and diversity of natural products having biological activity ends in the opportunity for the invention of drugs and the drugs derived from the natural assets play a widespread position in the prevention and treatment of human illnesses (Koehn and Carter 2005). Natural products continue to be the prominent source in view to the urgent need and demand for the new drugs, especially antibiotics and anticancer drugs. Hence, natural products have been witnessed as a highly compelled source to develop promising new drugs (Appendino and Banfi 2011).
The potential sources of the commercially essential bioactive compounds are the actinomycetes which have been historically wealthy in generating the vital bioactive compounds which provide a direct evidence to be a reliable supply of novel compounds (Parthasarathi et al. 2010). The list of novel actinobacteria and products derived from poorly explored areas of the world stresses the importance of investigating new habitats. The probability of locating novel bioactive compounds relies on the number of strains screened and their degree of diversity, uniqueness, and capability to produce secondary metabolites (Donadio et al. 2002). In this context awareness on rare resources which includes marine actinomycetes has captivated unique attention. This reality is due to their potency to produce biologically active secondary metabolites with many of them as potent antibiotics and lead compounds that cannot be produced and discovered by terrestrial microbes (Xiong et al. 2013).
Marine ecosystems harbor many unique forms of actinomycetes which appear to be widely distributed through the ocean and found in intertidal zones, sea water, sponges, and ocean sediments (Ramesh and Mathivanan 2009; Sun et al. 2010). Therefore, there is a need to bio-prospect untapped geographical assets and explore new strains of actinomycetes to maximize the discovery of novel bioactive metabolites (Leal et al. 2012). The terrestrial soils have been the predominant and extensively exploited source, and investigations on marine actinobacteria are few and inconclusive, despite the fact that they are the critical assets for new bioactive compounds (Newman and Cragg 2007). In recent years, there has been a growing awareness of the potential value of deep sea sediments as sources of actinobacteria that produce useful bioactive metabolic products. Literature on the isolation of actinobacteria from marine sediments suggested that these sources may be treasured for the isolation of novel actinobacteria with the potential to yield useful bioactive products (Jensen et al. 2007).
The search for filamentous actinobacterium offering incredibly precious bioactive compounds has been remarkably successful, and about two thirds of the naturally occurring antibiotics have been purified from actinomycetes species. Among the filamentous actinomycetes species that accounts for a major fraction of the bioactive metabolites, Streptomyces is by far the most prolific genus that the pharmacy industry has isolated and screened millions of strains over several decades. It has been advised that only a fraction of the bioactive compound producers of Streptomyces strains have been discovered and figuring out the undiscovered portion will require substantial effort (Solanki et al. 2008).
Optimization of the process variables can be computed by response surface methodology (RSM) effectively, if the mathematical formulations of RSM model are close to physics (being modeled) significantly (Wang et al. 2017). The drafted article of RSM by Box and Hunter (1957) sketched the principles for the design of the model. RSM designs the experiments using the mathematical and statistical techniques; it also builds models, evaluates the effects of parameters of the system, and engineers the optimum conditions for the target responses (Fatma et al. 2013). RSM is an up-to-date approach for constructing the model approximation that optimizes a response (output variable result) which is influenced by many independent process variables. The traditional methods-based optimization follows the optimization by one-factor-at-a-time (OFAT), but optimization using RSM for any production has an edge by reduction in the number of the experimental runs, time reduction for the execution of the experiments, and analyzing the influence of the variable interactions against the response (Wang et al. 2017). In addition RSM evaluates the conjunction between the variables and responses and defines the effect of variables against responses either as a single variable or via combination in process (Zafari et al. 2013).
Fermentation process provides a controlled environment to carry out complex reactions within the cell. Careful understanding of its metabolic stoichiometry in general is supported by mathematical expression with good approximations. These developed kinetic models pave the way for better elucidation of bioactive metabolite synthesis in terms of their yield and production rates. Simulation of the chosen models allows the bioengineers to estimate the kinetic parameters (or constants), which in turn validates the kinetic models. Several researchers have used unstructured models to explain the biological systems for effective synthesis of microbial products (Rama Krishna et al. 2016). In this study, estimation of kinetic parameters in bioactive metabolite production using Streptomyces nanhaiensis VSM-1 (KU507594) was attempted.
Research toward the exploration of marine actinobacteria has not progressed much in India. As a result, an attempt has been made to isolate potent indigenous actinobacterium strain VSM-1 with precise bioactive capability that was isolated from marine sediment samples of Bay of Bengal (Visakhapatnam) of Andhra Pradesh. Attempts have been additionally made to optimize the cultural and environmental conditions of the potent isolate that facilitate improved production of biologically active compounds. The present study additionally describes the extraction and biological evaluation of the antimicrobial compounds (by GC-MS analysis) produced by the strain.
2 Materials and Methods
2.1 Sampling and Isolation
The marine sediment samples were collected at different depths of the Bay of Bengal of coastal Andhra Pradesh, India. The collected sediment samples were transported to the laboratory in sterile bags and air-dried at room temperature for further isolation studies. The air-dried sediment sample was pretreated with calcium carbonate (10:1 w/w) and incubated at 37 °C for 4 days (Alferova et al. 1989). The treated sediment samples were suspended in sterile distilled water (1 g in 100 mL), homogenized by vortexing and 0.1 mL of serially diluted sample (10−4 dilution) that was spread over the surface of Bennet’s agar (0.1% yeast extract, 0.1% beef extract, 0.2% casein enzymic hydrolysate, 1% dextrose, and 2% agar) (pH 8) supplemented with nalidixic acid (50 μg/mL) and secnidazole (50 μg/mL). After incubation for 3 weeks at 30 °C, distinct strains were selected and maintained by subculturing on yeast extract malt extract dextrose agar (ISP-2) slants.
2.2 Identification
Characterization of the isolate was carried out as described by the International Streptomyces Project by microscopy, morphological, cultural, physiological, and biochemical characteristics. The cultural characteristics of the strain were studied on different media such as tryptone yeast extract agar (ISP-1), YMD agar (ISP-2), oatmeal agar (ISP-3), starch inorganic salts agar (ISP-4), glycerol asparagine agar (ISP-5), peptone yeast extract iron agar (ISP-6), tyrosine agar (ISP-7), starch-casein agar, Czapek-Dox agar, and nutrient agar media (Shirling and Gottlieb 1966). The micromorphology of the strain was examined under scanning electron microscopy (model JOEL-JSM 5600) (Bozzola and Russell 1999). The utilization of carbon sources by the strain was carried out in minimal medium containing different carbon sources at 1% concentration according to the method described by Isik et al. (1999). Biochemical tests which include IMVIC, H2S production (Cowan 1974), nitrate reduction (Gordon 1966), catalase (Jones 1949), and starch and gelatin hydrolysis (Waksman 1961) were also evaluated. Physiological characterization such as the effect of pH (5–9), temperature (20–60 °C), and salinity tolerance were analyzed. In addition the sensitivity of the strain to different antibiotics was determined by paper disc method (Williams 1989). Molecular identification of the strain was carried out according to the procedure of Nilsson and Strom (2002).
2.3 Optimization
2.3.1 Incubation Period
The growth pattern and bioactive metabolite production of the strain were studied at regular intervals up to 10 days. Two-week-old culture of the strain was cultivated in seed medium (ISP-2 broth) comprising of yeast extract (0.4%), malt extract (1%), dextrose (0.4%), and CaCO3 (0.2%) at room temperature for 48 h. Seed culture at a rate of 10% was inoculated into Bennet’s broth (production medium) consisting of 0.1% yeast extract, 0.1% beef extract, 0.2% casein enzymic hydrolysate, and 1% dextrose with pH 7.3. The fermentation process was carried out for 10 days under agitation at 120 rpm. At every 24 h interval, the flasks were harvested, and the biomass was separated from the culture filtrate. Biomass was determined in terms of total cell dry weight. Antimicrobial metabolite production was determined in terms of their antimicrobial spectrum (Narayana and Vijayalakshmi 2008). The culture filtrates were extracted with ethyl acetate and evaporated to dryness in a water bath at 80 °C. The solvent extracts were concentrated, and 50 μL of crude extract was tested for antimicrobial activity by employing agar well diffusion method against test organisms like Shigella flexneri (MTCC 1457), Serratia marcescens (MTCC 118), Proteus vulgaris (ATCC 6380), Pseudomonas aeruginosa (ATCC 9027), and Candida albicans (ATCC 10231).
2.3.2 Culture Conditions for the Optimum Production of Bioactive Metabolites
Bioactive metabolite production of the strain was optimized by using different parameters including pH, temperature, salt concentration, carbon and nitrogen sources, and minerals.
2.3.2.1 pH and Temperature
To determine the influence of initial pH on growth and bioactive metabolite production, Streptomyces nanhaiensis VSM-1 was cultured in the medium with different initial pH, ranging from 4 to 10, and at specific starting temperatures, from 20 to 50 °C. The biomass and bioactive metabolite production were envisioned to determine optimal pH and temperature conditions which were used in this study.
2.3.2.2 NaCl Concentration
The impact of salinity on growth and bioactive metabolite production by Streptomyces nanhaiensis VSM-1 was recorded by cultivating the strain in the fermentation medium amended with different concentrations of NaCl (0.1–9%) at optimum pH and temperature for 9 days. The salt concentration in which the strain exhibits optimum levels of bioactive metabolites was fixed for further studies.
2.3.2.3 Carbon and Nitrogen Sources
To determine the impact of carbon sources on biomass and bioactive metabolite production, different carbon sources like maltose, lactose, fructose, sucrose, glucose, starch, mannitol, arabinose, raffinose, and rhamnose each at a concentration of 1% were added separately into the production medium, maintaining all other conditions at optimum levels. The impact of varying concentrations of the best carbon source (0.5–5%) on the bioactive metabolite production was evaluated. Similarly, the influence of various nitrogen sources on antimicrobial metabolite production was evaluated by amending different nitrogen sources like soya peptone, Bio peptone, asparagine, meat extract, yeast extract, tryptone, peptone, casein, beef extract, and malt extract each at a concentration of 0.5% that were individually supplemented into the production medium containing an optimum amount of the superior carbon source. The growth and bioactive metabolite production were determined after 9 days of incubation at optimum pH, temperature, and salt concentration. Further, the impact of varying concentrations of optimized nitrogen source (0.1–2%) was studied to standardize the maximum antimicrobial metabolite production.
2.3.2.4 Minerals
Impact of minerals on the production of biomass and bioactive metabolites was studied by supplementing different minerals like MgSO4, FeSO4, ZnSO4, K2HPO4, and KH2PO4 each at a concentration of 0.05% (w/v) to the optimized medium.
2.3.2.5 Evaluation of Antimicrobial Activity
The bioactive metabolites of the strain produced under optimized conditions were tested in opposition to various strains of microorganisms, viz., Shigella flexneri (MTCC 1457), Serratia marcescens (MTCC 118), Proteus vulgaris (ATCC 6380), Pseudomonas aeruginosa (ATCC 9027), and medically important dermatophyte Candida albicans (ATCC 10231). The test microorganisms were procured from ATCC and MTCC and preserved at 4 °C.
2.3.2.6 Experimental Design by RSM
Experimental design applied for optimization of process variables (incubation time in days, pH, temperature, concentrations of mannitol and peptone) is executed using RSM. RSM is a combination of mathematical and statistical methods applied commonly for optimization of industrial, agricultural, and chemical reaction experimental designs (Sathish and Vivekanandan 2016). RSM is applied to classify the relationship among the process variables and the response (zone of inhibition against the pathogenic microorganisms by the bioactive metabolite produced by VSM-1), as well as optimize the best condition of the variables to predict model and to probe the best values of the responses (Güray et al. 2008). Central composite design (CCD) of RSM is an ideal constructive experimentation as it allows the reasonable amount of data to test lack of fit when sufficient number of experimental values exists. The five significant independent variables selected for the present study include incubation time in days, pH, temperature, and concentrations of mannitol and peptone, where A, B, C, D, and E are the values of the respective variable. The experimental ranges of the actual and the coded values of the process variables are shown in Table 38.1.
The independent variables experimental ranges were selected as low (−1), middle (0), and high (+1) based on some preliminary experimental studies (Myers et al. 2009). The experiments were designed and carried out as per central composite design (CCD) for 5 independent variables, of which a total 50 (=25 + 2 × 5 + 8) experiments consists of 32 factorial, 10 axial, and 8 center points, based on the following equation:
where N is the total number of experiments, n the number of independent variables, and nc the number of central points (replicates). ANOVA (analysis of variance) is applied for prediction of equation, and the information obtained from the experiments was fitted with the second-order polynomial equation (Wang et al. 2016):
where Y is the predicted response, β0 is the intercept coefficient, βi is the linear coefficient, βij is the interaction coefficients, βii is the quadratic coefficients, and Xi and Xj are the coded values of the five additive variables under study.
2.3.2.7 Model Adequacy Verification
Analysis of variance (ANOVA) is crucial and useful for evaluating the fitted model because the fitted mathematical model sometimes is not sufficient in explaining the experimental region of the independent variable (Marcos et al. 2008). In order to exercise the statistical discrepancy between at least three or more independent group means and their associated source of variations, Table 38.7 presents the results of ANOVA for the main effects, the quadratic terms, and the obtained data interactions. The sum of squares is the sum of the squares of the deviation from the mean. Mean square is an estimate of population variance that is obtained by dividing the associated sum of squares by the degrees of freedom. The mean square is compared with the residual mean square by the F value. The probability P value (Prob > F) is the probability of obtaining the F value or what we observed in the sample that determine the trueness of the null hypothesis (no significant difference of factor effects). Determining the coefficient determination (R2) is another common way to validate how well is the model’s goodness of fit. The R2 is measured on the scale of 1; if the R2 is close to 1, it means that the predicted values are in close agreement with the original experimental values. The 3D graphs were obtained using the Design-Expert software. The interactive effect between the process variables and their effect against the responses was plotted by 3D surface figure. Finally by solving the polynomial equation derived and the designed quadratic model and the grid search in RSM 3D plots, the optimal conditions were obtained.
2.3.2.8 Unstructured Mathematical Modeling
Basic mathematical and unstructured kinetic models which quantitatively describe the substrate utilization and growth-associated production formation kinetics in a batch system were taken from Ushakiranmayi et al. (2017). Models of logistic and Luedeking-Piret were used to simulate the cell growth and bioactive metabolite productions of Streptomyces nanhaiensis VSM-1 (KU507594). The data acquired from the logistic (L) model were used to calculate the maximum specific cell growth rate (μmax), d−1; the specific production rate of bioactive metabolite, d−1; and the maximum biomass concentration (Xm), g/L. Constants of growth-associated substrate consumption, γ, and non-growth-associated substrate consumption, η, were calculated using the logistic incorporated modified Luedeking-Piret (LIMLP) model. Similarly, growth-associated product formation constant, α, and non-growth-associated product formation constant, β, were determined from stationary phase data (where \( \frac{dX}{dt}=0 \)), using the logistic incorporated Luedeking-Piret (LILP) model.
2.3.2.9 Statistical Analysis
Results on the growth and the production of antimicrobial metabolites by S. nanhaiensis VSM-1 exposed to different cultural conditions are statistically analyzed with two-way analysis of variance (ANOVA).
2.3.2.10 Fermentation
A loopful of properly sporulated culture of S. nanhaiensis VSM-1 was inoculated into 100 mL of seed medium (YMD broth) and incubated on a rotary shaker at 30 °C for 48 h. Seed culture at the rate of 10% (v/v) was transferred to the optimized production medium consisting of 1% (w/v) mannitol, 1% (w/v) peptone, 0.05% (w/v) K2HPO4, and 3% (w/v) NaCl at pH 8. The inoculated flasks were incubated on rotary shaker at 30 °C and incubated for 9 days. The fermented broth collected at the end of 9 days was centrifuged, and the culture filtrate obtained was extracted with ethyl acetate. The solvent extracts were pooled and concentrated in vacuum to dryness, and the resultant crude extract was subjected to gas chromatography and mass spectroscopy.
2.3.2.11 Identification of Antimicrobial Metabolites by GC-MS
Identification of the chemical compounds present in the crude extract (CE) was carried out by GC-MS. Analysis was conducted on Agilent GC-MS machine (GC: 5890 series II; MSD 5972). The fused silica HP-5 capillary column (30 m × 0.25 mm, ID, film thickness of 0.25 μm) was directly coupled to the MS. The carrier gas was helium with a flow rate of 1.2 mL/min. Oven temperature was programmed (50 °C for 1 min and then 50–280 °C at a rate of 5 °C/min) and, subsequently, held isothermally for 20 min. The temperature of the injector port was maintained at 250 °C and that of detector at 280 °C (Boussada et al. 2008). The peaks of the obtained compounds in the gas chromatography were subjected to mass spectral evaluation. The spectra have been analyzed from the library data, NIST MS search (version 2.0) (included with NIST’02 mass spectral library, Agilent p/n G1033 A).
3 Results and Discussion
Taxonomic position of the strain VSM-1 was described on the basis of conventional and molecular approaches. Cultural characteristics of the strain were studied by growing the isolate on ten selective media (seven ISP media and three non-ISP media), and the results are tabulated in Table 38.2. The strain exhibited good growth on ISP-2, ISP-3, ISP-4, ISP-5, Czapek-Dox agar medium, and nutrient agar medium. The strain is aerobic actinomycetes that form grayish white aerial mycelium, while the substrate mycelium varied from pale yellow to white. No pigment was observed on any of the tested medium including melanin pigmentation on ISP-7. The biochemical and physiological characteristics of the isolate are summarized in Table 38.3. The strain is gram positive, and it has shown positive results for indole, Voges-Proskauer, citrate utilization, starch, casein hydrolysis, nitrate reduction, and catalase test but negative for gelatin hydrolysis, H2S production, and methyl red and coagulase tests. Growth of the strain occurred in the pH range of 6–10 with optimum growth at pH 7.5. The temperature range for growth was 20–45 ° C with the optimum growth at 30 °C. Tolerance of the strain to NaCl concentration additionally serves as an important character for species identification. VSM-1 exhibited salt tolerance up to 8% with optimum growth at 3% NaCl; therefore, the strain may be positioned in intermediate salt tolerance group. The salt concentration has a profound effect on the production of antibiotic from microorganisms due to its effect on the osmotic pressure to the medium (Tresner et al. 1968). The utilization of diverse carbon sources by the strain indicated its wide pattern of carbon assimilation ability (Table 38.4). The strain utilized a wide range of carbon sources such as fructose, galactose, glucose, maltose, sucrose, and xylose but did not utilize arabinose, inositol, lactose, rhamnose, and raffinose. Carbohydrate utilization plays a distinguished role in the taxonomic characterization of actinomycetes strains (Pridham and Gottlieb 1948). The strain was sensitive to the majority of the antibiotics tested but showed resistance to streptomycin, gentamycin, and ampicillin (Table 38.5).
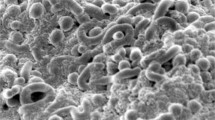
The spore chain ornamentation and spore morphology of the isolate were observed by scanning electron microscopy. The arrangement of spores was in spiral fashion, and the surface of the spore was rough (Fig. 38.1), and hence it can be placed in the spira group of the family Streptomycetaceae and the genus Streptomyces (Pridham et al. 1958). These results were confirmed by the identification of the culture by 16S rDNA sequencing. Gene sequence of 16S rRNA of VSM-1 was blasted against nucleotide database of the NCBI. The library search reported matching strains, and the sequences had been aligned with the set of published sequence on the basis of the conserved primary sequence and additionally with the aid of nucleotide blast similarity search evaluation. The 16S rDNA gene sequence of the isolate VSM-1 showed a close relation with Streptomyces nanhaiensis VSM-1 (Fig. 38.2). The rDNA sequence was deposited in the GenBank database of NCBI with an accession number KU507594.
3.1 Media Optimization
3.1.1 Incubation Period
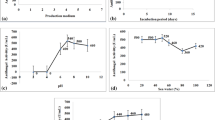
The growth pattern of S. nanhaiensis VSM-1 was studied on Bennet’s broth. The stationary phase of the strain extended from 168 to 216 h of incubation, after which it declined (Fig. 38.3a). The secondary metabolites obtained from 9-day-old culture exhibited high antimicrobial activity against the test microorganisms. The incubation period for the production of bioactive metabolites appears to differ among Streptomyces strains. Saurav and Kannabiran (2010) stated that the maximum incubation duration required for optimum growth and antibiotic yield by the isolate Streptomyces VITSVK9 sp. was 9 days which was in complete accordance with the earlier report, whereas metabolites accumulated from 10-day-old culture of S. crystallinus AZ-A151 producing hygromycin B exhibited good antimicrobial activity (Atta et al. 2012).
(a) Growth pattern and antimicrobial activity of S. nanhaiensis VSM-1. (b) Effect of pH on growth and bioactive metabolite yield of S. nanhaiensis VSM-1. (c) Effect of temperature on growth and bioactive metabolite yield of S. nanhaiensis VSM-1. (d) Effect of NaCl on growth and bioactive metabolite yield of S. nanhaiensis VSM-1. (e) Effect of different carbon sources on growth and bioactive metabolite yield of S. nanhaiensis VSM-1. (f) Effect of different concentrations of (continued) mannitol on growth and bioactive metabolite yield of S. nanhaiensis VSM-1. (g) Effect of different nitrogen sources on growth and bioactive metabolite yield of S. nanhaiensis VSM-1. (h) Effect of different concentrations of peptone on growth and bioactive metabolite yield of S. nanhaiensis VSM-1. (i) Effect of different minerals on growth and bioactive metabolite yield of S. nanhaiensis VSM-1 (Data on cell growth and bioactive metabolite yield were statistically analyzed by two-way ANOVA and found to be significant at 1%)
3.1.2 pH and Temperature
The bioactive metabolite production was found to be influenced by pH of the medium. The maximum growth as well as bioactive metabolite production of the strain was obtained at pH 8 (Fig. 38.3b). This result suggests the inclusion of this strain in the alkaliphilic actinomycetes group. Medium maintained at pH 8 was reported to support enhanced antimicrobial metabolite production by Streptomyces tanashiensis strain A2D (Singh et al. 2009). The results are also comparable with some Streptomyces spp. reported to produce antibiotics against bacteria, fungi, and yeast at alkaline pH (Basilio et al. 2003). The impact of temperature on biomass and bioactive metabolite production of the strain was studied. Highest growth in addition to antimicrobial metabolite production was obtained at 30 °C (Fig. 38.3c). In terms of its optimum temperature for growth, the organism appeared to be mesophilic in nature. This is in agreement with earlier reports for several of Streptomyces spp. (Ellaiah et al. 2004; Mustafa 2011; Atta et al. 2011).
3.1.3 NaCl Concentration
Optimum salt requirement for antimicrobial metabolite production was examined in liquid basal medium supplemented with different salt concentrations ranging from 0.1% to 9%. 3% NaCl was found to be optimum for maximum growth as well as antimicrobial compound production of the strain VSM-1 (Fig. 38.3d). Further increase in salt concentration reduced the antimicrobial agent biosynthesis. The requirement of NaCl for the production of bioactive metabolites seems to be different among actinomycetes strains. Singh et al. (2009) recorded that antibiotic production by S. tanashiensis was maximum at 2% NaCl in the medium. Culture medium with 5% NaCl was reported to support high antimicrobial metabolite production by Streptomyces VITSVK9 (Saurav and Kannabiran 2010).
3.1.4 Carbon and Nitrogen Sources
Impact of several carbon sources on biomass and bioactive metabolite yield was shown in Fig. 38.3e. The strain was able to grow in all the tested carbon sources. The strain produced highest biomass as well as antimicrobial compound in mannitol-supplemented medium. Addition of other carbon sources to the medium separately also favored growth, but the metabolite production was less when compared with mannitol. As mannitol is the preferred carbon source for bioactive metabolite production by the strain, different concentrations of mannitol (0.5–5%) were examined to determine its optimal concentration on metabolite production. Medium containing 1% mannitol supported high levels of biomass and bioactive metabolite production by the strain (Fig. 38.3f). Mannitol at the concentration of 1.5% as the best carbon source for high rates of antibiotic production by Streptomyces spp. 201 was reported by Thakur et al. (2009).
Of all the examined nitrogen sources, yeast extract and tryptone have been favored as good nitrogen sources for biomass production, whereas peptone was found to be the excellent nitrogen source for antimicrobial agent production, of the strain VSM-1 (Fig. 38.3g). Influence of different concentrations of peptone on the production of bioactive metabolites is represented in Fig. 38.3h. Peptone at the concentration of 1% exhibited optimal production of biomass and bioactive metabolite production. This is in conformity with the findings of Chattopadhyay and Sen (1997) and Han et al. (2004) for which peptone was proved to be the best nitrogen source for antibiotic production by Streptomyces rochei G 164 and Streptomyces scabiei PK-A41, respectively.
In the present study, the optimal culture conditions for the production of bioactive metabolites by strain VSM-1 were reported. The yield of metabolites by the strain was high in production medium supplemented with 1% mannitol and 1% peptone with initial pH 8 at temperature 30 °C for 9 days of incubation.
3.1.5 Minerals
The impact of minerals on growth and bioactive metabolite production of the strain is represented in Fig. 38.3i. K2HPO4 at the concentration of 0.05% enhanced the production of biomass and bioactive metabolites. In contrast the metabolite production was low with other minerals tested. Similar results were obtained with Streptomyces fradiae in neomycin production as mentioned by Majumdar and Majumdar (1965).
3.2 RSM Modeling and Optimization of Bioactive Metabolite Production by Streptomyces nanhaiensis VSM-1
Response surface methodology has been used for model designing and to optimize the independent process variables in various fields such as food industry, dye stuff industry, biotechnology, and also wastewater treatment (Hong et al. 2017). CCD design for the production of the bioactive metabolite produced by Streptomyces nanhaiensis VSM-1 and its effect against the five responses (zone of inhibition measured in mm) is given in Appendix Table 38.6 along with the experimental and predicted responses. The experimental results were verified using statistical analysis and modified quadratic models. The regression model equation for the bioactive metabolite production was expressed as follows (Payam et al. 2017).
where Y1 is Shigella flexneri, Y2 Serratia marcescens, Y3 Proteus vulgaris, Y4 Pseudomonas aeruginosa, and Y5 Candida albicans . A is the incubation time (days), B pH, and C temperature, and D and E are the concentrations of mannitol and peptone, respectively.
Predicted values calculated for Eq. (38.2) are in good agreement with the experimental values (Appendix Table 38.6). Hence, the designed quadratic model is well suited for the present experimental setup. Model adequacy was analyzed using ANOVA that determines the significance and the goodness of fit of the model (Almeida et al. 2017). As shown from Table 38.7, the F value of all the five responses S. flexneri, S. marcescens, P. vulgaris, P. aeruginosa, and C. albicans simply that the model is highly significant that there is only 0.01 chance that this large value (Model F value) could occur due to noise (Almeida et al. 2017). Based on the F values, the optimum values of all the five variables for the bioactive metabolite production was found to be incubation time (9 days), pH (8), temperature (30°C), and concentrations of mannitol and peptone (1% and 1%) and their effect against the five responses (effect of the bioactive metabolite produced by Streptomyces nanhaiensis VSM-1 and its effect is measured as zone of inhibition in mm) S. flexneri (34.2 mm), S. marcescens (32 mm), P. vulgaris (31.6), P. aeruginosa (32.3), and C. albicans (33.6).
The model terms are said to be significant if the probability values Prob > F less than 0.05 and values exceeding greater than 0.1 indicate that model terms are not significant. The P values of all the five variables (Table 38.8) are significant since the P value is 0.0001 (Ka et al. 2017). The lack-of-fit value of all the five responses is not significant relative to the pure error. Non-significant lack of fit is desired as it is mandatory to make the goodness of the model fit (Zhang et al. 2017). The lack-of-fit value of the five responses is given in Table 38.8. The predicted response values estimated by the designed model are shown by the determination of the coefficient (R2). The difference between the R2 and the adjusted R2 should be between 0 and 0.2 to determine the adequacy of the model. The R2 and the adjusted R2 are in reasonable agreement (Table 38.7) that indicate that the model is significant. Adequate precision determines the signal-to-noise ratio. A ratio greater than 4 is desirable. The adequate precision of the five responses is given in Table 38.7.
3.3 3D Plots
The 3D response plots for the bioactive metabolite production by Streptomyces nanhaiensis VSM-1 and its effect against the five responses obtained from the above model were given in Figs. 38.4, 38.5, 38.6, 38.7, and 38.8. The 3D plots represent the main and the interactive effects of the variables against the response. Two of the variables are tested while keeping the third variable at zero level. Mutual interaction between each of the two factors all showed elliptical shapes indicating that these factors interacted with each other and produced a combined effect for the production of the bioactive metabolites and its effect against the five responses (Ying et al. 2014).
3.4 Kinetic Modeling
In this study, L, LILP, and LIMLP models were fitted for experimental data using non-linear regression of least-square method with the help of Microsoft Excel Solver 2010. Estimated kinetic parameters from model fitting were tabulated in Table 38.9. High values of regression coefficient (R2) above 0.95 indicated that the chosen models have good precision. A high value (2.68–3.86 fold) of α than β (LILP model parameters) indicates that the bioactive metabolite produced from Streptomyces nanhaiensis VSM-1 (KU507594) is more growth associated than non-growth associated in shake flask. LIMLP model parameters γ and η are also having very good agreement with experimental data. Table 38.10 compares the values of zones of inhibition from agar diffusion studies of experimental versus model predicted.
Figure 38.9 shows the comparison of model predicted and experimental profiles of S. nanhaiensis VSM-1 growth-limiting substrate utilization. Comparison of zones of inhibition from produced bioactive metabolite on media inoculated with Shigella flexneri, Serratia marcescens, Proteus vulgaris, Pseudomonas aeruginosa, and Candida albicans strains over the time was shown in Fig. 38.10. Both Figs. 38.9 and 38.10 show very good fit of data for model predicted and experimental results. Thus, the selected unstructured models provided a better approximation of kinetic profiles, and these models can be used as kinetic approximations for bioactive metabolite production (in terms of zones of inhibition studies) by S. nanhaiensis VSM-1.
3.5 Identification of Potential Bioactive Constituents by GC-MS
GC-MS chromatogram of the ethyl acetate extracts of Streptomyces nanhaiensis VSM-1 showed a total of 30 peaks, indicating the presence of 30 compounds (Fig. 38.11). The active principles with their retention time (RT), molecular formula, molecular weight, and concentration (peak area) are reported in Table 38.11. The peak area is directly proportional to the quantity of the compounds present in the ethyl acetate fraction (EAF). When compared with NIST database, the nearest compound hits for those peaks were found. According to the available library data, NIST MS search (ver. 2) (included with NIST’02 mass spectral library, Agilent p/n G1033 A), the compounds were identified as undecane (5.065); 2-propanol, 1-(2-ethoxypropoxy)- (10.643); ethanone, 1-(2,6-dihydroxy-4-methoxyphenyl) (12.141); phenol, 2,4-bis(1,1-dimethylethyl)- (18.673); 1-dodecanol, 3,7,11-trimethyl- (18.826); cis-11,14-eicosadienoic acid, methyl ester (21.056); pentadecanoic acid (23.689); 2-propenoic acid, 2-methyl-, dodecyl ester (23.885); 1-octadecene, (E)- (25.691); methyl 2-hydroxydodecanoate (25.919); N-[4-cyclooctylaminobutyl]aziridine (27.214); hexadecanoic acid methyl ester (29.715); dibutyl phthalate (30.289); benzene acetic acid, 2-phenyl ethyl ester (30.481); 2,6-dibutyl 1–4-methyl piperidine (30.747); butyric acid, pentadecyl ester (31.891); 1-nonadecene (35.358); octadecane, 6-methyl (35.953); 9-eicosene, (E)- (39.252); n-tricosane (40.178); heptacosane, 1-chloro- (49.731); sulfurous acid, hexyl pentadecyl ester (49.948); erythro-9,10-dibromopentacosane (50.27); 2-methyl octacosane (50.922); oxalic acid, 2-ethylhexyl octadecyl ester (52.091); docosane, 11-decyl- (53.21); triacontane (53.717); tetratriacontane (54.475); 10-methyldotriacontane (55.362); and 13-n-undecylpentacosane (60.827). NCBI PubChem BioAssay Database (https://pubchem.ncbi.nlm.nih.gov) and Dr. Duke’s Phytochemical and Ethnobotanical Databases (http://www.ars-gringov/duke) revealed that among the 30 peaks, 14 compounds indicate the presence of active principles which have been previously reported for their antimicrobial, antioxidant, anti-inflammatory, and cytotoxic properties (Table 38.12).
GC-MS analysis of the strain VSM-1 indicated the presence of at least 14 bioactive compounds which makes this particular strain very potential for further study. Among the active principles, phenol, 2,4-bis(1,1-dimethylethyl)-; dibutyl phthalate; 9-eicosene, (E)-; and tetratriacontane have both antibacterial and antifungal properties (Salini et al. 2014; Rangel-Sánchez et al. 2014; Roy et al. 2006; Hsouna et al. 2011; Mahmood et al. 2009), while 1-octadecene, (E)-, and n-tricosane act only on bacteria (Mishra and Sree 2007; Mihailovi et al. 2011). The compounds like 2-methyl octacosane and 1-nonadecene were previously reported for their antifungal properties (Yu et al. 2009). Several studies have attributed the antioxidant effect to phenol, 2,4-bis(1,1-dimethylethyl)-; cis-11,14-eicosadienoic acid, methyl ester; pentadecanoic acid; heptacosane, 1-chloro-; 1-octadecene, (E)-; and hexadecanoic acid, methyl ester (Ajayi et al. 2011; Mishra and Sree 2007; Hema et al. 2011). Cis-11,14-eicosadienoic acid, methyl ester, and hexadecanoic acid, methyl ester, are also reported for their anti-inflammatory active principles (Hema et al. 2011), while triacontane was reported for its antibacterial, anti-diabetic, and anti-tumor properties (Khare 2004; Mammen et al. 2010).
4 Conclusion
In the pursuit of fruitful bioactive metabolites, the study was designed to investigate the unexplored marine sediment samples of the Bay of Bengal of coastal Andhra Pradesh, India, for novel actinomycetes and their antimicrobial properties. An actinobacterium isolate VSM-1 having broad-spectrum activity was isolated from the marine sediment sample using CaCO3 pretreatment technique on Bennet’s agar media. The isolate was identified as Streptomyces nanhaiensis based on the polyphasic taxonomy. Bioactive metabolite production was initially optimized using one-factor-at-a-time (OFAT) method to select the variables. CCD of RSM was employed for further optimization that enables the evaluation of the effects of the individual factors, and their interactive influence on the responses and metabolite yield was enhanced which was represented in the form of zones of inhibition. The experimental values are in good agreement with the predicted values. Very good agreement between experimental results and model fitted data also confirmed the use of unstructured kinetic models could be better approximations for further enhancement in productivities. Lastly, the existence of various bioactive principles identified through GC-MS which is ascertained to have broad spectrum of pharmacological properties also supported that the strain VSM-1 was the most active strain. However, further studies will need to be undertaken to isolate and characterize the individual bioactive chemotypes will definitely give promising results.
References
Ajayi GO, Olagunju JA, Ademuyiwa O (2011) Gas chromatography-mass spectrometry analysis and phytochemical screening of ethanolic root extract of Plumbago zeylanica, Linn. Med Plant Res 5:1756–1761
Alferova IV, Terekhova LP, Prauzer KH (1989) Selective medium with nalidixic acid for isolating antibiotic producing actinomycetes. Antibiot Khimioter 34:344–348
Almeida DG, Soares da Silva R d CF, Luna JM, Rufino RD, Santos VA, Sarubbo LA (2017) Response surface methodology for optimizing the production of biosurfactant by Candida tropicalis on industrial waste substrates. Front Microbiol 8:157
Appendino G, Banfi L (2011) Molecular diversity and natural products. Mol Divers 15:291–292
Atta HM, Bahobail AS, El-Sehrawi MH (2011) Studies on isolation, classification and phylogenetic characterization of antifungal substances produced by Streptomycesalbidoflavus-143. New York Sci J 4:40–53
Atta HM, Afifi MM, Elshanawany AA (2012) Production of Hygromycin-B antibiotic from Streptomyces crystallinus. AZ-A151: II. Parameters controlling of antibiotic production. Acad Arena 4:37–52
Basilio A, Gonzalez I, Vicente MF (2003) Patterns of antimicrobial activities from soil actinomycetes isolated under different conditions of pH and salinity. J Appl Microbiol 95:814–823
Boussada O, Ammar A, Saidana D (2008) Chemical composition and antimicrobial activity of volatile components from capitula and aerial parts of Rhaponticum acaule DC growing wild in Tunisia. Microbiol Res 163:87–95
Box GEP, Hunter JS (1957) Multi-factor experimental designs for exploring response surfaces. Ann Math Stat 28:1957) 195–1957) 241
Bozzola JJ, Russell LD (1999) Electron microscopy principles and techniques for biologists. Jones and Bartlett Publishers, Sudbury
Bredholdt H, Galatenko OA, Engelhardt K (2007) Rare actinomycete bacteria from the shallow water sediments of the Trondheim fjord, Norway: isolation, diversity, biological activity. Environ Microbiol 9:2756–2764
Chattopadhyay D, Sen SK (1997) Optimization of cultural conditions for antifungal antibiotic accumulated by Streptomyces rochei G 164. Hindustan Antibiot Bull 39:64–71
Cowan ST (1974) Cowan and Steel’s manual for the identification of medical bacteria. Cambridge University Press, Cambridge
Donadio S, Sosio M, Lancini G (2002) Impact of the first Streptomyces genome sequence on the discovery and production of bioactive substances. Appl Microbiol Biotechnol 60:377–380
Ellaiah P, Srinivasulu B, Adinarayana K (2004) Optimization studies on neomycin production by a mutant strain of Streptomyces marinensis in solid state fermentation. Process Biochem 39:529–534
Fatma GSB, Isil IG, Osman O (2013) Analysis of the polymer composite bipolar plate properties on the performance of PEMFC (polymer electrolyte membrane fuel cells) by RSM (response surface methodology). Energy 55:1067–1075
Gordon RE (1966) Some criteria for the recognition of Nocardia madura (Vincent) Blanchord. J Gen Microbiol 45:355–364
Güray G, Altunay P, Abdurrahman T (2008) Electrochemical treatment of deproteinated whey waste water and optimization of treatment conditions with response surface methodology. J Hazard Mater 157:69–78
Han WC, Lee JY, Park DH (2004) Isolation and antifungal and antioomycete activity of Streptomyces scabiei strain PK-A41, the causal agent of common scab disease. Plant Pathol J 20:115–126
Hema R, Kumaravel S, Alagusundaram K (2011) GC/MS determination of bioactive components of Murraya koenigii. J Am Sci 7:80–83
Hong L, Yi-min Z, Jing H (2017) Optimization of vanadium (IV) extraction from stone coal leaching solution by emulsion liquid membrane using response surface methodology. Chem Eng Res Des 123:111–119
Hsouna AB, Trigui M, Mansour RB (2011) Chemical composition, cytotoxicity effect and antimicrobial activity of Ceratonia siliqua essential oil with preservative effects against Listeria inoculated in minced beef meat. Int J Food Microbiol 148:66–72
Isik K, Chun J, Hah YC, Goodfellow M (1999) Nocardia salmonicida nom. rev., a fish pathogen. Int J Syst Bacteriol 49:833–837
Jensen PR, Williams PG, Oh DC (2007) Species specific secondary metabolite production in marine actinomycetes of the genus Salinispora. Appl Environ Microbiol 73:1146–1152
Jones K (1949) Fresh isolates of actinomycetes in which the presence of sporogenous aerial mycelia is a fluctuating characteristics. J Bacteriol 57:141–145
Ka HW, George Q, Kong M (2017) Optimisation of Pueraria isoflavonoids by response surface methodology using ultrasonic-assisted extraction. Food Chem 231:231–237
Khare CP (2004) Indian herbal remedies: rational western therapy, ayurvedic and other traditional usage, botany. Springer, Berlin
Koehn FE, Carter GT (2005) The evolving role of natural products in drug discovery. Nat Rev Drug Discov 4:206–220
Leal MC, Puga J, Serodio J (2012) Trends in the discovery of new marine natural products from invertebrates over the last two decades—where and what are we bioprospecting? PLoS One 7:1–15
Mahmood A, Ahmed R, Kosar S (2009) Phytochemical screening and biological activities of the oil components of Prunus domestica Linn. J Saudi Chem Soc 13:273–277
Majumdar MK, Majumdar SK (1965) Effect of minerals on neomycin production by Streptomyces fradiae. Appl Microbiol 13:190–193
Mammen D, Daniel M, Sane RT (2010) Seasonal and geographical variations in chemical constituents of Leptadenia reticulata. Int J Pharm Sci Rev Res 4:111–116
Marcos AB, Ricardo ES, Eliane PO (2008) Response surface methodology (RSM) as a tool for optimization in analytical chemistry. Talanta 76:965–977
Mihailovi V, Vukovic N, Niciforovic N (2011) Studies on the antimicrobial activity and chemical composition of the essential oils and alcoholic extracts of Gentiana asclepiadea L. J Med Plant Res 5:1164–1174
Mishra PM, Sree A (2007) Antibacterial activity and GC-MS analysis of the extract of leaves of Finlaysonia obovata (a Mangrove Plant). Asian J Plant Sci 6:168–172
Mustafa O (2011) Effects of some environmental conditions on biomass and antimicrobial metabolite production by Streptomyces spp. KGG32. Int J Agric Biol 3:317–324
Myers RH, Montgomery DC, Anderson-Cook CM (2009) Response surface methodology: process and product optimization using designed experiments. Wiley, Hoboken, NJ
Narayana KJP, Vijayalakshmi M (2008) Optimization of antimicrobial metabolites production by Streptomyces albidoflavus. Res J Pharmacol 2:4–7
Newman DJ, Cragg GM (2007) Natural products as sources of new drugs over the last 25 years. J Nat Prod 70:461–477
Nilsson WB, Strom MS (2002) Detection and identification of bacterial pathogens of fish in kidney tissue using terminal restriction fragment length polymorphism (T-RFLP) analysis of 16S rRNA genes. Dis Aquat Org 48:175–185
Parthasarathi S, Kim CJ, Kim PK, Sathya S, Manikandan M, Manikandan T, Balakrishnan K (2010) Taxonomic characterization and UV/VIS analysis of antagonistic marine actinomycete isolated from South West Coast of South Korea. Int J Med Res 1:99–105
Payam R, Shohreh M, Saeid V (2017) Modeling and optimization of gelatin-chitosan micro-carriers preparation for soft tissue engineering: using response surface methodology. Mater Sci Eng C 75:545–553
Pridham TG, Gottlieb D (1948) The utilization of carbon compounds by some Actinomycetales as an aid for species determination. J Bacteriol 56:107–114
Pridham TG, Hesseltine CW, Benedict RG (1958) A guide for the classification of Streptomyces according to selected groups. Appl Microbiol 6:52–79
Rama Krishna VSG, Ushakiranmayi M, Vijayalakshmi M, Sudhakar P (2016) Model-based kinetic parameters estimation in batch Pullulan fermentation using Jaggery as substrate. J Chem Pharm Res 8:217–224
Ramesh S, Mathivanan N (2009) Screening of marine actinomycetes isolated from the Bay of Bengal, India for antimicrobial activity and industrial enzymes. World J Microbiol Biotechnol 25:2103–2111
Rangel-Sánchez G, Castro-Mercado E, García-Pineda E (2014) Avocado roots treated with salicylic acid produce phenol-2,4-bis (1,1-dimethylethyl), a compound with antifungal activity. J Plant Physiol 17:189–198
Roy RRN, Laskar S, Sen SK (2006) Dibutyl phthalate, the bioactive compound produced by Streptomyces albidoflavus 321.2. Microbiol Res 161:121–126
Salini TS, Divakaran D, Shabanamol S (2014) Antimicrobial and Immuno modulatory potential of endophytic fungi Fusarium solani isolated from Withania somnifera. World J Pharm Res 3:879–890
Sathish S, Vivekanandan S (2016) Parametric optimization for floating drum anaerobic bio-digester using response surface methodology and artificial neural network. Alexandria Eng J 55:3297–3307
Saurav K, Kannabiran K (2010) Diversity and optimization of process parameters for the growth of Streptomyces VITSVK 9 spp. isolated from Bay of Bengal, India. J Nat Environ Sci 1:56–65
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Singh LS, Majumdar S, Bora TC (2009) Optimization of process parameters for growth and bioactive metabolite production by a salt tolerant and alkaliphilic actinomycete, Streptomyces tanashiensis strain A2D. J Mycol Med 19:223–225
Solanki R, Khanna M, Cal R (2008) Bioactive compounds from marine actinomycetes. Curr Sci 48:410–431
Sun W, Dai S, Jiang S (2010) Culture dependent and culture independent diversity of actinobacteria associated with the marine sponge Hymeniacidon perleve from the South China Sea. Antonie Van Leeuwenhoek 98:65–75
Thakur D, Bora TC, Bordoloi GN (2009) Influence of nutrition and culturing conditions for optimum growth and antimicrobial metabolite production by Streptomyces spp. 201. J Mycol Med 19:161–167
Tresner HD, Hayes JA, Backus EJ (1968) Differential tolerance of Streptomyces to sodium chloride as a taxonomic aid. Appl Microbiol 16:1134–1136
Ushakiranmayi M, Vijayalakshmi M, Sudhakar P, Rama Krishna Ganduri VS, Satish Babu R (2017) Optimization of culture conditions by response surface methodology and unstructured kinetic modeling for L-Asparaginase production by Pseudonocardia endophytica VUK-10. J Appl Pharm Sci 7:42–50
Waksman SA (1961) The actinomycetes, classification, identification and descriptions of genera and species. The Williams and Wilkins Co., Baltimore, MD
Wang G, Deng Y, Xu X (2016) Optimization of air jet impingement drying of okara using response surface methodology. Food Control 59:743–749
Wang Y, Qin X, Huang S, Feng J (2017) Structural-borne acoustics analysis and multi-objective optimization by using panel acoustic participation and response surface methodology. Appl Acoust 116:139–151
Williams ST (1989) In: Williams ST (ed) Bergey’s manual of systematic bacteriology, vol 4. Williams & Wilkins, Baltimore, MD
Xiong ZQ, Wang JF, Hao YY, Wang Y (2013) Recent advances in the discovery and development of marine microbial natural products. Mar Drugs 11:700–717
Ying Z, Chengyou C, Mei P (2014) Diversity of nitrogen-fixing, ammonia-oxidizing, and denitrifying bacteria in biological soil crusts of a revegetation area in Horqin Sandy Land, Northeast China. Ecol Eng 71:71–79
Yu H, Zhao X, Yue P (2009) Chemical communication in mammal population: urinary olfactory chemosignals in lactating female root voles (Microtus oeconomus pallas). Polish J Ecol 58:153–165
Zafari A, Kianmehr MH, Abdolahzadeh R (2013) Modeling the effect of extrusion parameters on density of biomass pellet using artificial neural network. Int J Recycl Org Waste Agric 2(1):9
Zhang B, Han X, Pengju G, ShuqiFang JB, Zhang B, Han X, Gu P, Fang S, Bai J (2017) Response surface methodology approach for optimization of ciprofloxacin adsorption using activated carbon derived from the residue of desilicated rice husk. J Mol Liq 238:316–325
Acknowledgments
The first author (UKM) is grateful to the University Grants Commission (UGC), New Delhi, Government of India, for providing financial assistance in the form of fellowship to carry out this work.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
1 Appendix
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Managamuri, U., Vijayalakshmi, M., Poda, S., Rama Krishna Ganduri, V.S., Satish Babu, R. (2020). Bioactive Metabolites from Streptomyces nanhaiensis VSM-1: Polyphasic Taxonomy, Optimization, and Evaluation of Antimicrobial Metabolites by GC-MS Analysis. In: Khasim, S.M., Long, C., Thammasiri, K., Lutken, H. (eds) Medicinal Plants: Biodiversity, Sustainable Utilization and Conservation. Springer, Singapore. https://doi.org/10.1007/978-981-15-1636-8_38
Download citation
DOI: https://doi.org/10.1007/978-981-15-1636-8_38
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-1635-1
Online ISBN: 978-981-15-1636-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)