Abstract
Cullin-RING ubiquitin ligases (CRLs) determine the substrate specificity of ubiquitination reactions, and substrates are recruited to the cullin core through binding to their cognate substrate receptor modules. Because a family of substrate receptors compete for the same cullin core, the assembly and activity of CRLs are dynamically regulated to fulfill the needs of the cell to adapt to the changing pool of proteins demanding ubiquitination. Cullins are modified by NEDD8, a ubiquitin-like protein. This process, referred to as neddylation, promotes the E3 activity of CRLs by inducing conformational rearrangement in the Cullin-RING catalytic core. Cand1 is a cullin-associated protein whose binding is excluded by cullin neddylation. Although early biochemical studies suggested that Cand1 inhibits CRL activity, genetic studies revealed its positive role in ubiquitination. Emerging evidence from kinetic and quantitative proteomic studies demonstrated that Cand1 stimulates assembly of new Skp1-Cul1-F-box protein (SCF) complexes by exchanging the Skp1-F-box protein substrate receptor modules. Furthermore, aided by refined experimental design as well as computational simulation, an attractive model has been developed in which substrate, neddylation cycle and Cand1-mediated “adaptive exchange” collaborate to maintain the dynamics of the cellular SCF repertoire. Here, we review and discuss recent advances that have deepened our understanding of CRL regulation.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
3.1 Introduction to NEDD8
Neural precursor cell expressed, developmentally downregulated 8 (NEDD8) is a highly conserved protein across eukaryotic species (83% identity between humans and Arabidopsis thaliana), although different copy numbers for the NEDD8 gene were found in genomes of several plant species (Rao-Naik et al. 1998; Mergner and Schwechheimer 2014). NEDD8 was first reported as a highly expressed mRNA in embryonic mouse brain (Kumar et al. 1992). Although NEDD8 mRNA exhibits a broad expression pattern in adult tissues, it is highly accumulated in the heart and skeletal muscle in humans (Kamitani et al. 1997). Its expression is increased during the early stage of mouse embryogenesis but decreased in the brain at late developmental stages (Kumar et al. 1992; Kamitani et al. 1997). In terms of subcellular localization, exogenous NEDD8 showed strong expression in the nucleus and much weaker expression in the cytoplasm (Kamitani et al. 1997).
NEDD8 is a ubiquitin-like protein. Among all of the ubiquitin-like proteins, NEDD8 possesses the greatest similarity with ubiquitin (~60% identical to human ubiquitin). It even contains the same C-terminal Leu-Arg-Gly-Gly residues, which are crucial for the conjugation of ubiquitin to its substrate proteins. Interestingly, both ubiquitin and Nedd8 are produced as precursors that need to be processed at the C-terminus to reveal the terminal glycine (Kamitani et al. 1997). Cdc53, the budding yeast Cul1, was the first protein identified to be covalently modified by NEDD8, suggesting a regulatory role of neddylation in the ubiquitin-proteasome system (Lammer et al. 1998; Liakopoulos et al. 1998). Genetic studies have demonstrated that neddylation is essential in multiple model organisms including mice (Tateishi et al. 2001), Arabidopsis (Dharmasiri et al. 2003), Drosophila (Ou et al. 2002), and fission yeast (Osaka et al. 2000), suggesting a conserved and important function of neddylation in development.
Given the high sequence similarity, it is not surprising that the overall structure of NEDD8 is similar with that of ubiquitin (Rao-Naik et al. 1998; Whitby et al. 1998). NEDD8 is comprised of a globular core and a C-terminal tail that is flexible in solution. Ubiquitin displays an asymmetric distribution of charged residues, thereby forming the acidic and basic patches (Wilkinson 1988). The globular domain of NEDD8 maintains a similar arrangement of acidic and basic patches, although charged residues making up the patches are only moderately conserved. Similarly, two surface-exposed hydrophobic patches, which are crucial for protein-protein interactions in ubiquitin (Shih et al. 2000; Sloper-Mould et al. 2001; Hu et al. 2002), are both conserved in NEDD8 (Enchev et al. 2015). However, NEDD8 and ubiquitin each have specific functions due to small differences in their structures. Seven unique residues that are conserved across NEDD8 orthologues but differ from the ones in ubiquitin are responsible for NEDD8-specific interactions. One such residue, Ala-72, was shown to play a critical role in preventing the interaction of NEDD8 with the ubiquitin E1 enzyme (Whitby et al. 1998). The other six charged residues present on the surface of NEDD8 are essential for regulating ubiquitin ligase activities by NEDD8 (Wu et al. 2002).
3.2 The NEDD8 Conjugation System
NEDD8 is initially translated as a precursor that requires proteolytic processing to expose the C-terminal Gly residue. Ubiquitin C-terminal hydrolase L3 (UCHL3), a member of the C12 family peptidases, displays a dual specificity for efficient processing of both NEDD8 and ubiquitin (Wada et al. 1998; Johnston et al. 1999; Linghu et al. 2002). In contrast, the deneddylase 1 (DEN1) from the C48 family peptidases acts exclusively on NEDD8. DEN1 can mediate the proteolytic processing of NEDD8 precursor as well as the deconjugation of NEDD8 from some protein substrates (Gan-Erdene et al. 2003; Mendoza et al. 2003; Wu et al. 2003). Of note, UCHL3 knockout in mice or DEN1 knockout in Drosophila does not result in neddylation defects (Kurihara et al. 2000; Chan et al. 2008), suggesting redundant function between both enzymes.
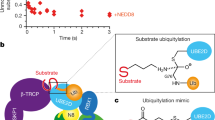
The processed NEDD8 is covalently conjugated to substrate proteins in a manner highly similar to ubiquitin conjugation, through a cascade of enzymes comprising E1 NEDD8-activating enzyme (NAE), E2 NEDD8-conjugating enzyme, and E3 NEDD8 ligase (Fig. 3.1). Initially, via the C-terminal Gly-76, NEDD8 forms a thioester bond with NAE. The human NAE complex is composed of two subunits, amyloid-β precursor protein binding protein 1 (APP-BP1) and ubiquitin-activating enzyme 3 (UBA3) (Osaka et al. 1998; Gong and Yeh 1999). Interestingly, the APP-BP1 and UBA3 are highly homologous to the amino- and carboxy-terminal regions of ubiquitin E1, respectively. In Arabidopsis, the corresponding NAE has also been identified, with AXR1 as the counterpart of APP-BP1 and Ecr1 as the orthologue of UBA3 (Pozo 1998). The Arabidopsis axr1 mutant exhibits abnormal response to the essential plant hormone auxin (Lincoln et al. 1990; Leyser et al. 1993), further demonstrating the importance of neddylation. Two active sites in the heterodimeric NAE complex, an adenylation domain in UBA3, and a catalytic Cys domain, are required for efficient activation and conjugation of NEDD8 (Walden et al. 2003a). The C-terminal ubiquitin-fold domain (UFD) in UBA3 plays an important role in mediating interactions with the E2 NEDD8-conjugating enzymes (Huang et al. 2005; Walden et al. 2003b).
Simplified scheme of cullin neddylation. Following precursor processing of NEDD8 by UCH, mature NEDD8 was attached to cullins in consecutive steps including E1 (UBA3/APPBP1)-mediated activation, E2 (UBE2M/F)-mediated conjugation, and E3 (Rbx/DCN1)-mediated ligation. NEDD8 isopeptidases such as the COP9 signalosome are able to catalyze the deneddylation of cullins
Once activated by NAE, NEDD8 is transferred to a conserved cysteine on the E2 NEDD8-conjugating enzyme via a transthiolation reaction. To date, two NEDD8 E2-conjugating enzymes have been identified: the well-studied Ubc12 (also known as UBE2M) and less-characterized UBE2F (Liakopoulos et al. 1998; Osaka et al. 1998; Gong and Yeh 1999; Huang et al. 2009). Structural studies showed that the N-terminal extension and catalytic core domain of either Ubc12 or UBE2F bind to UBA3’s hydrophobic groove and UFD of NAE, respectively (Huang et al. 2004, 2005, 2007, 2009). Although they interact with NAE in similar fashions, Ubc12 and UBE2F exhibit distinct substrate specificities for neddylation. Ubc12 pairs with RING-box protein 1 (Rbx1) to regulate neddylation of Cullins 1–-4, whereas UBE2F specifically interacts with RING-box protein 2 (Rbx2) to mediate Cullin 5 neddylation (Huang et al. 2009). Interestingly, these two E2s were recently demonstrated to cross-talk with each other, wherein Ubc12 promotes UBE2F ubiquitylation and degradation under both physiological and stressed conditions (Zhou et al. 2018).
Following thioester bond formation, the E2~NEDD8 complex interacts with NEDD8 E3 enzymes for specific transfer of NEDD8 onto target proteins. The RING domain proteins, characterized by the zinc coordination sites in the RING domain, represent the first reported NEDD8 E3 ligases (Deshaies and Joazeiro 2009). The N-terminal domain of Rbx1 mediates stable assembly with cullins, whereas the C-terminal RING binds and activates E2 enzymes, including the E2~NEDD8 intermediate. Biochemical and structural studies have revealed a dual E3 mechanism—involving Rbx1 and Defective in Cullin Neddylation 1 (DCN1)—that mediates cullin neddylation (Kurz et al. 2008; Scott et al. 2010, 2011; Kim et al. 2008). Early studies demonstrated that Ubc12 stimulates cullin neddylation in an Rbx1-dependent manner (Kamura et al. 1999; Gray 2002), while DCN1 is required for optimal neddylation of cullin in C. elegans and S. cerevisiae (Kurz et al. 2005). Furthermore, in the presence of N-terminally acetylated E2, DCN proteins dramatically stimulate cullin neddylation in vitro (Monda et al. 2013; Scott et al. 2014). Thus, DCN1 is considered a co-E3, which is capable of binding both cullin and the acetylated N-terminus of Ubc12 via its potentiating neddylation (PONY) domain, restricting the flexible RBX1-bound Ubc12~NEDD8 to a catalytically competent orientation. Similarly, N-terminal acetylation in UBE2F also promotes DCN1-dependent neddylation of Cul5 that specifically employs Rbx2 as the RING protein (Monda et al. 2013). More recently, a sophisticated neddylation model was revealed, in which the NEDD8 and substrate protein synergistically regulate neddylation specificity by toggling E2/E3 conformations (Scott et al. 2014).
3.3 Activation of CRLs by Neddylation
The family of cullin-RING ligases (CRLs) is characterized by a cullin core linking an E2-binding RING and a substrate-binding receptor module. Early crystallographic studies of cullin-RING complexes revealed a highly elongated structure in which cullin serves as a rigid scaffold (Zheng et al. 2002a; Duda et al. 2008; Angers et al. 2006). The N-terminal domain (NTD) of cullin, which consists of three repeats of a novel five-helix structural motif, adopts a stalk-like structure that recruits substrate-binding adaptors. The C-terminal domain (CTD) of cullin is comprised of a four-helix bundle (4HB), an α/β and two winged-helix (WHA and WHB) subdomains. In terms of Rbx1 binding, two interaction surfaces were identified in the CTD (Zheng et al. 2002a). The Rbx1 RING domain docks into a V-shaped groove formed by the cullin α/β and WHB domains, whereas the N-terminal β-strand of Rbx1 is incorporated into the β-sheet of the cullin α/β domain. Importantly, in the structural model of a E2~ubiquitin thioester bound to Rbx1 in a CRL complex, the catalytic cysteine of E2 is positioned approximately 50 Å from the tip of the substrate receptor protein. Furthermore, the rigidity of the Cul1 scaffold that imposes this gap is required for E3 activity, as evidenced by the finding that introducing flexibility into the N-terminal repeat domain disrupts ubiquitination of a specific substrate (Zheng et al. 2002a). These observations thus beg the question of how ubiquitin is transferred to the substrate and to the continually elongating ubiquitin chain.
The cullin family members are the best-characterized neddylation substrates. Neddylation of cullin takes place on a highly conserved lysine residue, which is located in the C-terminal WHB subdomain (Wada et al. 1999). Mutagenesis studies indicate that cullin neddylation efficiently promotes the ubiquitination activity of Skp1-Cul1-F-box protein (SCF), the founding member of CRLs, resulting in rapid turnover of protein substrates (Wu et al. 2000; Read et al. 2000; Morimoto et al. 2000; Podust et al. 2000). In vitro biochemical evidence indicates that neddylation stimulates recruitment of ubiquitin charged E2 to the SCF complex (Kawakami et al. 2001). Interestingly, the proximity of the cullin neddylation site to the RING domain of Rbx1 suggests that neddylation might stimulate the activity of CRLs by regulating the interaction between NEDD8 and Rbx1-bound E2~ubiquitin (Wu et al. 2002; Zheng et al. 2002a; Read et al. 2000; Kawakami et al. 2001).
The crystal structure of neddylated Cul5ctd-Rbx1 complex revealed that neddylation induces a striking reorientation of the cullin WHB subdomain (Duda et al. 2008), which frees the RING domain of Rbx1 from the compact WHB-RING architecture in the unneddylated cullin. As a result, Rbx1 adopts flexible conformations, providing potential catalytic geometries for ubiquitin transfer. Besides the C-terminus of NEDD8, the Leu8/Ile44/His68/Val70 face is also involved in the interaction between NEDD8 and Cul5 (Duda et al. 2008). However, a prior study reported that the hydrophobic patch around Ile44 of NEDD8 binds ubiquitin E2, but not the NEDD8 E2, in nuclear magnetic resonance (NMR) spectroscopy (Sakata et al. 2007). This disagreement between the two structural studies suggests that in the presence of E2~ubiquitin under conditions of ubiquitin transfer, structural rearrangements may occur to favor contacts between NEDD8 and E2~ubiquitin (see Chap. 2 for full discussions on the structure of CRLs with and without neddylation). Detailed insights into the multiple mechanisms by which NEDD8 activates SCF emerged from a kinetic study (Saha and Deshaies 2008). First, neddylation enhances E2 recruitment to SCF as well as the rate of ubiquitin transfer to substrates. Second, neddylation improves the rate of ubiquitin chain elongation. Third, neddylation enables crosslinking of substrate to E2 and enhances amide bond formation in the E2 active site. Finally, neddylation increases the fraction of ubiquitin-charged substrate molecules as well as the average number of ubiquitin molecules attached to a modified substrate. Overall, two distinct effects of neddylation on CRL activity were proposed: (1) enhancement of ubiquitin chain initiation by bridging the gap between the SCF substrate and the E2~ubiquitin and (2) improvement of chain elongation via stimulating E2 recruitment and enhancing E2 activity. Consistent with the structural and kinetic findings, unneddylated SCF complexes containing a mutant Cul1 with WHB deletion, which were engineered to mimic neddylation by constitutively releasing the RING, exhibit increased activity in vitro (Duda et al. 2008; Yamoah et al. 2008) and in vivo (Boh et al. 2011). Thus, the unneddylated CRLs can be considered as autoinhibited or off, and neddylation activates the complex by inducing profound conformational rearrangements.
In addition to mediating RING domain rearrangement, neddylation also disrupts the binding interface between Cul1-Rbx1 and CAND1 (cullin-associated NEDD8-dissociated protein 1), the substrate receptor exchange factor (see below). Moreover, both WHB and NEDD8 are reoriented to contact the cullin NTD in the neddylated CRL models, evoking the hypothesis that the CTD-NTD interface might be remodeled by WHB and/or NEDD8 to favor CRL activity (Duda et al. 2008) (see Fig. 2.4 in Chap. 2 for structural details). Further studies are required to provide new insight into the architecture of neddylated full-length CRLs.
The NEDD8 modification is reversible. Once cullins are neddylated, they can be rapidly deneddylated by the COP9 signalosome (CSN) in which CSN5 serves as the catalytic subunit (see Chap. 4 for full discussions on CSN and its role in regulating CRLs). CSN specifically recognizes neddylated cullins as its substrates. Recent studies have shown that neddylated Cul1 and Cul4 have a much higher affinity for CSN than their unneddylated forms (Cavadini et al. 2016; Mosadeghi et al. 2016). A key aspect of CSN activity is that it is inhibited by substrate binding to the CRL (Fischer et al. 2011; Emberley et al. 2012; Enchev et al. 2012). This is due to a steric clash between the bound substrate and CSN (Cavadini et al. 2016; Fischer et al. 2011; Enchev et al. 2012). As a consequence, a CRL in vivo should be most efficiently deneddylated when there is little or no substrate available. The mutually exclusive binding of substrate and CSN to a CRL lies at the heart of how differential assembly states of different CRL complexes is controlled.
Based on its biochemical activity, it is not surprising that CSN inhibits the activity of SCF ubiquitin ligase in vitro (Lyapina et al. 2001; Zhou et al. 2003). However, multiple lines of genetic evidence revealed that CSN promotes degradation of substrates mediated by SCF and other CRLs in vivo, suggesting a positive role of CSN in regulating CRL activity (Schwechheimer et al. 2001; Cope et al. 2002; Pintard et al. 2003). This apparent paradox about CSN function was partially resolved by the finding that the inhibitory effects of CSN exhibited in vitro prevent the autoubiquitination and degradation of CRL substrate receptors in vivo, thereby promoting CRL activity (Zhou et al. 2003; Wee et al. 2005; He et al. 2005). More importantly, deneddylation is also required for Cand1-mediated dynamic exchange of CRL substrate receptors, a process critical for sustaining continuous cycles of CRL assembly and disassembly (see later portion of this chapter).
3.4 CAND1: An Inhibitor or Activator of CRLs?
Cand1, formerly known as TATA-binding protein (TBP)-interacting protein 120A (TIP120A), was originally identified as a TBP-interacting protein in rat liver nuclear extracts (Yogosawa et al. 1996). A subsequent study showed that Cand1 globally stimulates transcription driven by three classes of eukaryotic RNA polymerases (Makino et al. 1999). To date, the significance of these functions remains unknown. A milestone for understanding the role of Cand1 is the finding by four groups that Cand1 interacts with cullins (Zheng et al. 2002b; Liu et al. 2002; Hwang et al. 2003; Min et al. 2003; Oshikawa et al. 2003). Both the Skp1-binding N-terminus and the unneddylated C-terminal domain of cullins are required for Cand1-cullin association. Reciprocally, deletion of either N-terminal or C-terminal residues from Cand1 abolished its binding with Cul1, suggesting that Cand1-cullin interaction may involve both ends of Cand1. The crystal structure of Cand1-Cul1-Rbx1 brought a more comprehensive understanding of the Cand1-cullin complex architecture (Goldenberg et al. 2004). By virtue of 27 tandem huntingtin-elongation-A subunit-TOR (HEAT) repeats, Cand1 forms a highly sinuous superhelical structure that coils into a U-shaped belt that cradles Cul1-Rbx1. In the complex, Cand1 and Cul1 interact with each other in a head-to-tail pattern. Overall, like a clamp, Cand1 uses its N-terminal and C-terminal arches as two prongs to grip both ends of Cul1. Importantly, a β-hairpin motif protruding from the Cand1 main body interacts with the Skp1-binding helices located on the Cul1 N-terminus. Distally, two helical repeats on the N-terminus of Cand1 occupy a cleft on the Cul1 CTD where the neddylation acceptor lysine residue resides. These findings are consistent with the biochemical data that showed that either Skp1 or neddylation dissociated Cand1 from cullins (Zheng et al. 2002b; Liu et al. 2002; Hwang et al. 2003; Min et al. 2003; Oshikawa et al. 2003). In turn, the binding of Cand1 to Cul1 dissociated Skp1 from Cul1 and inhibited the ubiquitination of p27 in vitro (Zheng et al. 2002b). All these findings converge to the conclusion that Cand1 inhibits CRL activity in vitro.
Several lines of evidence, however, revealed that Cand1 is required for optimal CRL activity in vivo. Although RNAi-mediated silencing of Cand1 in human cells significantly increases Cul1-Skp1 association, the level of substrate protein also increases slightly (Zheng et al. 2002b). The Arabidopsis Cand1 loss-of-function mutants exhibit defects in CRL regulated signaling events and reduced turnover rates of CRL substrate proteins (Feng et al. 2004; Chuang et al. 2004; Cheng and Dai 2004). All these observations are reminiscent of the CSN paradox, and it was naturally hypothesized that Cand1 helps maintain the stability of CRL substrate receptors by sequestering cullin cores, thus promoting CRL activity in vivo (Zheng et al. 2002b; Cope and Deshaies 2003; Min et al. 2005). In support of this, a slight decrease in the level of Skp2 is observed in Cand1 knockdown Hela cells (Zheng et al. 2002b). However, several lines of evidence argued against the model of Cand1-mediated adaptor stabilization (Schmidt et al. 2009; Chua et al. 2011), and a few studies reported that Cand1 can promote CRL activity independently of adaptor stability (Lo and Hannink 2006; Bosu et al. 2010). Cand1 deletion does not affect the stability of at least a subset of substrate receptors in fission yeast (Schmidt et al. 2009), and Cand1 knockdown human cells exhibit a decrease in CRL3Keap1 activity despite an increase in the level of assembled CRL3Keap1 complexes (Lo and Hannink 2006). So, the question still remains: how does Cand1 promote CRL activity in vivo? An important hint came from the biochemical data demonstrating that human Skp2-Skp1 promotes the dissociation of Cand1 from Cul1. Furthermore, as mentioned in the prior section, substrate can inhibit binding and deneddylation of CRLs by CSN and thus maintaining neddylation (Cavadini et al. 2016; Fischer et al. 2011; Emberley et al. 2012; Enchev et al. 2012; Chew and Hagen 2007). Schmidt et al. suggested a refined model in which transient interaction of Cand1 with Cul1-Rbx1 scaffold drives dynamic cycles between association and displacement of different substrate receptors. This cycle would enable rapid remodeling of the CRL repertoire in the absence of substrates, and when substrates are available, they induce the neddylation of their cognate CRLs to promote their ubiquitination. Upon substrate consumption, CSN-mediated deneddylation would direct the CRLs again into the Cand1 cycle. Though this model can explain how Cand1 activates CRLs while competing against substrate receptors for binding to cullins, there was no direct biochemical evidence for the dynamic recycling of CRL complexes nor how Cand1 could promote such recycling.
3.5 CAND1: The Substrate Receptor Exchange Factor
A key breakthrough in the current understanding of Cand1 emerged from a kinetic analysis of SCF assembly and disassembly (Pierce et al. 2013). Using a fluorescence resonance energy transfer (FRET) assay, the binding dynamics between an F-box protein and the Cul1-Rbx1 scaffold were monitored in a real-time manner. With a K D in the picomolar range, Fbxw7-Skp1 forms an extremely tight complex with Cul1-Rbx1 regardless of the neddylation state. Importantly, addition of Cand1 to pre-assembled and unneddylated SCFFBXW7 complexes increases the dissociation rate of Fbxw7-Skp1 by more than one million fold, with a maximum rate of 1.3 s−1. As expected, neddylation of Cul1 abolishes the effect of Cand1 due to its ability to block binding of Cand1 to Cul1. Reciprocally, the spontaneous dissociation of Cand1 from Cul1-Rbx1 is extremely slow, and this dissociation rate is accelerated greatly by Fbxw7-Skp1. Furthermore, a Skp1 mutant lacking a loop that is predicted to clash with a β-hairpin of Cand1 (Goldenberg et al. 2004) fails to displace Cand1 from Cul1-Rbx1, and the Cand1 mutant lacking the β-hairpin also fails to displace F-box–Skp1 from Cul1-Rbx1 (Liu et al. 2018). When either the Skp1 loop or the β-hairpin of Cand1 is deleted, Skp1 and Cand1 can form a stable ternary complex with Cul1 (Goldenberg et al. 2004; Liu et al. 2018).
The biochemical and kinetic results described above can be explained by hypothesizing the existence of a transient ternary complex consisting of Cand1, substrate receptor module, and Cul1 that upon dissociation of either Cand1 or the substrate receptor module would yield a stable SCF or Cul1-Cand1 complex. Given the ability of both Cand1 and the substrate receptor module to destabilize each other’s association with Cul1-Rbx1, it was proposed that Cand1 mediates the exchange of substrate receptors on the Cul1-Rbx1 scaffold, which exactly mimics how guanine nucleotide exchange factors (GEFs) act on Ras-like GTPases (Klebe et al. 1995; Goody and Hofmann-Goody 2002; Guo et al. 2005). In support of this hypothesis, addition of Cand1 to a mixture of pre-assembled SCFβ-TrCP and free Fbxw7-Skp1 significantly increases the ubiquitylation of the SCFFbxw7 substrate, CycE (Pierce et al. 2013). In contrast to the previous in vitro finding that Cand1 inhibits the ubiquitin ligase activity of SCF, these results showed for the first time that Cand1 promotes the ubiquitination of SCF substrates through accelerating the assembly of specific substrate receptors with Cul1 recycled from the pre-existing pool of SCFs. Based on its biochemical mechanism of action, Cand1 is defined as a “substrate receptor exchange factor” (Pierce et al. 2013).
Besides the few substrate receptors tested in vitro, quantitative mass spectrometry revealed that the cellular repertoire of SCF is dramatically altered by Cand1 depletion, in both human (Pierce et al. 2013) and yeast cells (Wu et al. 2013). Furthermore, in budding yeast lacking Cand1, Skp1-Grr1 failed to associate with Cdc53 (yeast Cul1) in response to glucose, likely due to inefficient release of Cdc53 from pre-existing SCF complexes (Zemla et al. 2013). To further reveal the power of Cand1-mediated exchange of substrate receptors, metabolic pulse-labelling assays were designed to examine SCF dynamics in cells (Pierce et al. 2013; Wu et al. 2013). As expected, several newly synthesized F-box proteins exhibit reduced assembly into Cul1 in the absence of Cand1. Importantly, the level of these F-box proteins in total cell lysate is unaffected, indicating that the rate of synthesis is not responsible for the reduced incorporation. Taken together, these data indicate that Cand1 modulates the SCF repertoire by acting as an exchange factor that equilibrates Cul1-Rbx1 with the entire pool of substrate receptors. This exchange activity of Cand1 ensures that the cellular repertoire of CRLs remains dynamic, and when specific substrates arise, they can stabilize their cognate CRLs to sustain their efficient degradation.
3.6 Regulation of CRL Assembly by Both Cand1 and Neddylation
Through blocking access of Cand1 to cullins, neddylation inhibits Cand1-mediated exchange, and therefore, neddylation activity should also regulate the assembly of CRLs. Surprisingly, a couple of early studies found that when neddylation was eliminated by MLN4924 (pevonedistat), a potent inhibitor of NAE (Soucy et al. 2009), the CRL network was marginally affected (Bennett et al. 2010; Lee et al. 2011). In light of the updated knowledge about the working mechanism of Cand1, methods used to probe cellular CRL complexes were re-evaluated. In a conventional immunoprecipitation assay, the CRL complexes may have been rearranged due to post-lysis exchange of substrate receptors by Cand1. This possibility was tested using an assay based on SILAC mass spectrometry (Reitsma et al. 2017). HEK293 cells with (heavy) or without (light) a 3xFLAG tag on the endogenous Cul1 are grown in media with and without heavy-labeled amino acids as indicated. Immediately before lysis, cells are mixed and the percentage exchange of F-box proteins is determined by immunoprecipitation of 3XFLAG-tagged Cul1 followed by quantitative mass spectrometry. Strikingly, robust exchange of F-box proteins is observed in the shortest immunoprecipitation performed (10 min). Importantly, this exchange is greatly suppressed for Skp1 and almost all F-box proteins in the lysate of Cand1/2 double-knockout (DKO) cells, suggesting that Cand1/2 accounts for the fast post-lysis re-equilibration of SCF complexes. Inhibition of neddylation by pevonedistat further enhances exchange for all FBPs, apparently through exposing Cand1 binding sites on Cul1. These findings point out that to precisely capture the landscape of cellular CRLs, it is necessary to inhibit the Cand1-mediated post-lysis exchange of substrate receptors.
Inspired by the finding that both SCF and Cul1-Cand1 complexes are extremely stable (Pierce et al. 2013; Reitsma et al. 2017), it was proposed that an excess of recombinant Cul1-Rbx1 in the lysis buffer, like a molecular sponge, could suppress exchange by sequestering any free Cand1/2 and F-box proteins. As predicted, exchange of most F-box proteins is drastically diminished in the immunoprecipitates when recombinant Cul1-Rbx1 is added to the lysis buffer (Reitsma et al. 2017). By virtue of this molecular sponge, it is now possible to investigate how the cellular SCF repertoire changes in response to various perturbations. Experiments using this new immunoprecipitation system revealed that F-box proteins show huge variations in their percentage association with Cul1, suggesting a non-equilibrium pool of SCF complexes, and most F-box proteins exhibit low percentages of Cul1 association. In contrast to previous reports that only a small fraction of Cul1 stably associates with Cand1 in 293T cells (Bennett et al. 2010), the new assay revealed that nearly half of the Cul1 assembles with Cand1 in HEK2933xFLAG-Cul1 cells, underscoring the potential of Cand1 to rigorously regulate the SCF repertoire. Importantly, depletion of Cand1/2 leads to an increase in net assembly of F-box proteins, whereas pevonedistat treatment elicits the opposite effect in a Cand1/2-dependent manner. These results indicate that neddylation together with Cand1/2 activity maintain a non-equilibrium pool of SCF complexes.
Since the cellular concentrations of F-box proteins do not correlate with their percentage association with Cul1, what determines the landscape of cellular SCFs? As mentioned previously, the antagonistic binding of substrates and CSN to Cul1 (Cavadini et al. 2016; Fischer et al. 2011; Enchev et al. 2012) suggested that substrates play a key role in sculpting the dynamic SCF repertoire. To test this idea, the assembly state of individual SCF complexes was explored upon induction of substrate. As expected, assembly of a specific SCF complex is induced when its cognate substrate becomes available. Furthermore, this process depends on both Cand1/2 activity and neddylation (Liu et al. 2018; Reitsma et al. 2017).
A perplexing phenomenon found in both Arabidopsis and human Cand1-deficient cells is that increased assembly of a specific CRL complex occurs jointly with inefficient degradation of its substrate (Chuang et al. 2004; Lo and Hannink 2006; Pierce et al. 2013; Zhang et al. 2008). By exploring Tumor Necrosis Factor alpha (TNFα)-induced degradation of IκBα via SCFβ-TrCP-dependent ubiquitination, a study using quantitative approaches provided important insights into this paradox (Liu et al. 2018). Despite a higher level of SCFβ-TrCP in unstimulated DKO cells, the degradation rate of IκBα is substantially reduced compared to WT cells. Immunoprecipitation assays revealed that phosphorylated IκBα (pIκBα, the substrate of SCFβ-TrCP) in DKO cells binds β-TrCP as normal, but the level of pIκBα-bound β-TrCP recruited to Cul1 decreases. These results suggest that substrates can bind both free and Cul1-bound F-box proteins, but substrates bound to free F-box proteins cannot gain access to Cul1 for efficient ubiquitination in cells lacking Cand1/2 activity. Consistent with this explanation, β-TrCP overexpression, which leads to an increased level of SCFβ-TrCP but no change in the percentage association of β-TrCP with Cul1, fails to increase the degradation rate of IκBα in DKO cells, simply because the fraction of pIκBα bound to free β-TrCP still cannot gain access to Cul1. In contrast, Cul1 overexpression rescued the degradation defect of IκBα in DKO cells, because almost all β-TrCP molecules were driven to form SCF complexes. Taken together, these results demonstrate that dynamic exchange of F-box proteins associated with Cul1 is required for efficient substrate degradation, and restricted access of substrate to Cul1 represents the major deficiency in DKO cells.
Interestingly, although Cand1 inhibits neddylation of cullins, it promotes the binding of DCN1, the NEDD8 E3, to Cul1-Rbx1, and Cand1 decreased the K D of DCN1-Cul1 by 36 fold (Kim et al. 2008; Monda et al. 2013; Liu et al. 2018; Keuss et al. 2016). Moreover, the DCN1-Cul1 complex could not be detected in DKO cells through co-immunoprecipitation. This finding suggests that DCN1 prefers Cand1-bound Cul1, which enables immediate neddylation of Cul1 upon the removal of Cand1 by F-box-Skp1. Indeed, in the presence of F-box-Skp1 modules, Cand1-bound Cul1 exhibited a higher neddylation rate than free Cul1 (Liu et al. 2018). In addition, neddylation biases the exchange reaction toward the formation of a stable SCF by preventing Cand1 from re-binding the Cul1 and thus acting as the energy input that drives Cul1 toward the formation of new SCF. Furthermore, this mechanism may also ensure that each new SCF is neddylated and activated coincident with its formation.
3.7 Assembly and Disassembly of SCF: The Cul1 Cycle
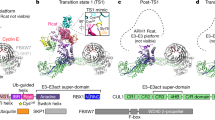
Based on all the available kinetic and quantitative studies of the components and regulators of SCFs, a mathematical model was developed to elucidate and investigate the dynamics of the SCF system (Liu et al. 2018). In this model, Cul1 constantly cycles through the exchange, neddylation and deneddylation stages, while substrate binding through an F-box protein locks Cul1 in the neddylated state, and thus the SCF is stabilized to allow substrate ubiquitination (Fig. 3.2). In addition to simulating and predicting changes in substrate degradation upon genetic and chemical alterations, the model provides novel insights into the significance of the Cand1-mediated exchange mechanism. First, the model calculated that a Cul1 molecule with no bound substrate goes through the entire cycle with an average time of ~1.5 min. This rate is consistent with the rate of Cul1 neddylation and deneddylation measured in human cells (Liu et al. 2018). Considering the molar ratio of Skp1 to Cul1, an F-box protein could gain access to Cul1 every ~4 min. This rapid cycle apparently offers a more efficient way to remodel the SCF repertoire compared to synthesis-dependent regulation. Second, the total F-box protein concentration was predicted as one of the two most sensitive parameters affecting the substrate half-life in the DKO but not the WT cells. Indeed, overexpression of a single F-box protein, including a truncated F-box protein that binds only Cul1 but no substrate protein, dramatically increases the stability of multiple proteins that are substrates of SCFs only in the DKO cells. These cells also display dramatically lower cell proliferation rates and an increased level of apoptosis marker proteins. These observations led to the hypothesis that Cand1-mediated exchange permits the SCF system to tolerate large variations in the level of individual F-box proteins. Because such large changes can naturally occur during development, loss of Cand1 could have profound effects especially on multicellular organisms. In support of this, in the absence of Cand1, Arabidopsis exhibited severe defects throughout its life cycle, whereas cultured cells and yeasts were only modestly affected. More broadly, this Cand1-mediated “adaptive exchange” mechanism could confer organism tolerance to variations in the number of expressed F-box proteins, providing a foundation for large expansion or contraction of the F-box protein gene repertoire during the evolution of different species (Liu et al. 2018).
Schematic illustration of adaptive SCF assembly. Cand1-mediated exchange of F-box proteins (FBP), together with the neddylation/deneddylation cycle, sustains the dynamicity of the SCF system. Binding of Cand1 to a pre-existing SCF complex forms an unstable ternary intermediate that expels either Cand1 or the Skp1-FBP module. After the Skp1-FBP has dissociated, the resulting Cand1-Cul1 promptly retrieves another Skp1-FBP module from the cellular pool. Disassociation of Cand1 from the newly formed ternary complexes generates a new SCF complex. Due to the preferential association of DCN1 with Cul1 that is bound by Cand1, neddylation occurs immediately after Cand1 dissociation. Binding of substrate stabilizes this neddylated SCF complex, resulting in efficient ubiquitylation of the substrate. In the absence of substrates, CSN catalyzes the deneddylation of Cul1, and the deneddylated SCF complex reenters the cycle
3.8 Closing Remarks
While the attractive picture of SCF dynamics has been increasingly revealed, important aspects of the assembly and regulation of CRLs remain to be explored. First, studies of dynamic exchange largely rely on the Cul1-based SCF system. However, Cand1 also associates with other cullins, and at steady state, Cand1 is not equally distributed across cullins, with more Cand1 associated with Cul1 and Cul5 than that with Cul2 and Cul4 (Bennett et al. 2010). Biochemical studies akin to those performed in SCF are required to answer whether Cand1 plays a similar role in regulating the dynamics of other CRLs and, if so, how important is the dynamic exchange in regulating the activity of these CRLs. Second, how is the distribution of Cand1 over different cullins determined and coordinated? Third, although the net assembly of SCF complexes was increased in the DKO cells and was decreased when neddylation was inhibited by pevonedistat, quite a few F-box proteins showed different trends in their assembly status (Reitsma et al. 2017). It is thus very interesting to study what roles Cand1 and neddylation play in the regulation of these F-box proteins. Furthermore, Cand1-mediated exchange has been hypothesized to confer tolerance to large variations in substrate receptor levels. Investigating specific effects of Cand1 at different developmental stages in multiple eukaryotic organisms will further elucidate the importance of this exchange mechanism.
Abbreviations
- 4HB:
-
Four-helix bundle
- APP-BP1:
-
Amyloid-β precursor protein binding protein 1
- CAND1:
-
Cullin-associated NEDD8-dissociated protein 1
- CAND2:
-
Cullin-associated NEDD8-dissociated protein 2
- CRLs:
-
Cullin-RING ubiquitin ligases
- CSN:
-
COP9 signalosome
- CTD:
-
C-terminal domain
- DCN1:
-
Defective in cullin neddylation 1
- DEN1:
-
Deneddylase 1
- DKO:
-
Double knockout
- FBPs:
-
F-box proteins
- FRET:
-
Fluorescence resonance energy transfer
- GEFs:
-
Guanine nucleotide exchange factors
- HEAT:
-
Huntingtin-elongation-A subunit-TOR
- NAE:
-
NEDD8-activating enzyme
- NEDD8:
-
Neural precursor cell expressed, developmentally downregulated 8
- NMR:
-
Nuclear magnetic resonance
- NTD:
-
N-terminal domain
- PONY:
-
Potentiating neddylation
- Rbx1:
-
RING-box protein 1
- Rbx2:
-
RING-box protein 2
- SCF:
-
Skp1-Cul1-F-box
- TBP:
-
TATA-binding protein
- TNFα:
-
Tumor necrosis factor alpha
- UBA3:
-
Ubiquitin-activating enzyme 3
- UCHL3:
-
Ubiquitin C-terminal hydrolase L3
- UFD:
-
Ubiquitin-fold domain
References
Angers S, Li T, Yi X et al (2006) Molecular architecture and assembly of the DDB1-CUL4A ubiquitin ligase machinery. Nature 443(7111):590–593
Bennett EJ, Rush J, Gygi SP et al (2010) Dynamics of cullin-RING ubiquitin ligase network revealed by systematic quantitative proteomics. Cell 143(6):951–965
Boh BK, Smith PG, Hagen T (2011) Neddylation-induced conformational control regulates cullin RING ligase activity in vivo. J Mol Biol 409(2):136–145
Bosu DR, Feng H, Min K et al (2010) C. elegans CAND-1 regulates cullin neddylation, cell proliferation and morphogenesis in specific tissues. Dev Biol 346(1):113–126
Cavadini S, Fischer ES, Bunker RD et al (2016) Cullin-RING ubiquitin E3 ligase regulation by the COP9 signalosome. Nature 531(7596):598–603
Chan Y, Yoon J, Wu JT et al (2008) DEN1 deneddylates non-cullin proteins in vivo. J Cell Sci 121(Pt 19):3218–3223
Cheng Y, Dai X, Zhao Y (2004) AtCAND1, a HEAT-repeat protein that participates in auxin signaling in Arabidopsis. Plant Physiol 135(2):1020–1026
Chew EH, Hagen T (2007) Substrate-mediated regulation of cullin neddylation. J Biol Chem 282(23):17032–17040
Chua YS, Boh BK, Ponyeam W et al (2011) Regulation of cullin RING E3 ubiquitin ligases by CAND1 in vivo. PLoS One 6(1):e16071
Chuang HW, Zhang W, Gray WM (2004) Arabidopsis ETA2, an apparent ortholog of the human cullin-interacting protein CAND1, is required for auxin responses mediated by the SCF(TIR1) ubiquitin ligase. Plant Cell 16(7):1883–1897
Cope GA, Deshaies RJ (2003) COP9 signalosome. Cell 114(6):663–671
Cope GA, Suh GS, Aravind L et al (2002) Role of predicted metalloprotease motif of Jab1/Csn5 in cleavage of Nedd8 from Cul1. Science 298(5593):608–611
Deshaies RJ, Joazeiro CA (2009) RING domain E3 ubiquitin ligases. Annu Rev Biochem 78:399–434
Dharmasiri S, Dharmasiri N, Hellmann H et al (2003) The RUB/Nedd8 conjugation pathway is required for early development in Arabidopsis. EMBO J 22(8):1762–1770
Duda DM, Borg LA, Scott DC et al (2008) Structural insights into NEDD8 activation of cullin-RING ligases: conformational control of conjugation. Cell 134(6):995–1006
Emberley ED, Mosadeghi R, Deshaies RJ (2012) Deconjugation of Nedd8 from Cul1 is directly regulated by Skp1-F-box and substrate, and the COP9 signalosome inhibits deneddylated SCF by a noncatalytic mechanism. J Biol Chem 287(35):29679–29689
Enchev RI, Scott DC, Da Fonseca PC et al (2012) Structural basis for a reciprocal regulation between SCF and CSN. Cell Rep 2(3):616–627
Enchev RI, Schulman BA, Peter M (2015) Protein neddylation: beyond cullin-RING ligases. Nat Rev Mol Cell Biol 16(1):30–44
Feng S, Shen Y, Sullivan JA et al (2004) Arabidopsis CAND1, an unmodified CUL1-interacting protein, is involved in multiple developmental pathways controlled by ubiquitin/proteasome-mediated protein degradation. Plant Cell 16(7):1870–1882
Fischer ES, Scrima A, Bohm K et al (2011) The molecular basis of CRL4DDB2/CSA ubiquitin ligase architecture, targeting, and activation. Cell 147(5):1024–1039
Gan-Erdene T, Nagamalleswari K, Yin L et al (2003) Identification and characterization of DEN1, a deneddylase of the ULP family. J Biol Chem 278(31):28892–28900
Goldenberg SJ, Cascio TC, Shumway SD et al (2004) Structure of the Cand1-Cul1-Roc1 complex reveals regulatory mechanisms for the assembly of the multisubunit cullin-dependent ubiquitin ligases. Cell 119(4):517–528
Gong L, Yeh ETH (1999) Identification of the activating and conjugating enzymes of the NEDD8 conjugation pathway. J Biol Chem 274(17):12036–12042
Goody RS, Hofmann-Goody W (2002) Exchange factors, effectors, GAPs and motor proteins: common thermodynamic and kinetic principles for different functions. Eur Biophys J: EBJ 31(4):268–274
Gray WM (2002) Role of the Arabidopsis RING-H2 protein RBX1 in RUB modification and SCF function. Plant Cell Online 14(9):2137–2144
Guo Z, Ahmadian MR, Goody RS (2005) Guanine nucleotide exchange factors operate by a simple allosteric competitive mechanism. Biochemistry 44(47):15423–15429
He Q, Cheng P, He Q et al (2005) The COP9 signalosome regulates the Neurospora circadian clock by controlling the stability of the SCFFWD-1 complex. Genes Dev 19(13):1518–1531
Hu M, Li P, Li M et al (2002) Crystal structure of a UBP-family deubiquitinating enzyme in isolation and in complex with ubiquitin aldehyde. Cell 111(7):1041–1054
Huang DT, Miller DW, Mathew R et al (2004) A unique E1-E2 interaction required for optimal conjugation of the ubiquitin-like protein NEDD8. Nat Struct Mol Biol 11(10):927–935
Huang DT, Paydar A, Zhuang M et al (2005) Structural basis for recruitment of Ubc12 by an E2 binding domain in NEDD8’s E1. Mol Cell 17(3):341–350
Huang DT, Hunt HW, Zhuang M et al (2007) Basis for a ubiquitin-like protein thioester switch toggling E1-E2 affinity. Nature 445(7126):394–398
Huang DT, Ayrault O, Hunt HW et al (2009) E2-RING expansion of the NEDD8 cascade confers specificity to cullin modification. Mol Cell 33(4):483–495
Hwang J-W, Min K-W, Tamura T-A et al (2003) TIP120A associates with unneddylated cullin 1 and regulates its neddylation. FEBS Lett 541(1–3):102–108
Johnston SC, Riddle SM, Cohen RE et al (1999) Structural basis for the specificity of ubiquitin C-terminal hydrolases. EMBO J 18(14):3877–3887
Kamitani T, Kito K, Nguyen HP et al (1997) Characterization of NEDD8, a developmentally down-regulated ubiquitin-like protein. J Biol Chem 272(45):28557–28562
Kamura T, Conrad MN, Yan Q et al (1999) The Rbx1 subunit of SCF and VHL E3 ubiquitin ligase activates Rub1 modification of cullins Cdc53 and Cul2. Genes Dev 13(22):2928–2933
Kawakami T, Chiba T, Suzuki T et al (2001) NEDD8 recruits E2-ubiquitin to SCF E3 ligase. EMBO J 20(15):4003–4012
Keuss MJ, Thomas Y, McArthur R et al (2016) Characterization of the mammalian family of DCN-type NEDD8 E3 ligases. J Cell Sci 129(7):1441–1454
Kim AY, Bommelje CC, Lee BE et al (2008) SCCRO (DCUN1D1) is an essential component of the E3 complex for neddylation. J Biol Chem 283(48):33211–33220
Klebe C, Prinz H, Wittinghofer A et al (1995) The kinetic mechanism of ran-nucleotide exchange catalyzed by RCC1. Biochemistry 34(39):12543–12552
Kumar S, Tomooka Y, Noda M (1992) Identification of a set of genes with developmentally down-regulated expression in the mouse brain. Biochem Biophys Res Commun 185(3):1155–1161
Kurihara LJ, Semenova E, Levorse JM et al (2000) Expression and functional analysis of Uch-L3 during mouse development. Mol Cell Biol 20(7):2498–2504
Kurz T, Ozlu N, Rudolf F et al (2005) The conserved protein DCN-1/Dcn1p is required for cullin neddylation in C. elegans and S. cerevisiae. Nature 435(7046):1257–1261
Kurz T, Chou YC, Willems AR et al (2008) Dcn1 functions as a scaffold-type E3 ligase for cullin neddylation. Mol Cell 29(1):23–35
Lammer D, Mathias N, Laplaza JM et al (1998) Modification of yeast Cdc53p by the ubiquitin-related protein rub1p affects function of the SCFCdc4 complex. Genes Dev 12(7):914–926
Lee JE, Sweredoski MJ, Graham RL et al (2011) The steady-state repertoire of human SCF ubiquitin ligase complexes does not require ongoing Nedd8 conjugation. Mol Cell Proteomics: MCP 10(5):M110.006460
Leyser HM, Lincoln CA, Timpte C et al (1993) Arabidopsis auxin-resistance gene AXR1 encodes a protein related to ubiquitin-activating enzyme E1. Nature 364(6433):161–164
Liakopoulos D, Doenges G, Matuschewski K et al (1998) A novel protein modification pathway related to the ubiquitin system. EMBO J 17(8):2208–2214
Lincoln C, Britton JH, Estelle M (1990) Growth and development of the axr1 mutants of Arabidopsis. Plant Cell 2(11):1071–1080
Linghu B, Callis J, Goebl MG (2002) Rub1p processing by Yuh1p is required for wild-type levels of Rub1p conjugation to Cdc53p. Eukaryot Cell 1(3):491–494
Liu J, Furukawa M, Matsumoto T et al (2002) NEDD8 modification of CUL1 dissociates p120CAND1, an inhibitor of CUL1-SKP1 binding and SCF ligases. Mol Cell 10(6):1511–1518
Liu X, Reitsma JM, Mamrosh JL et al (2018) Cand1-mediated adaptive exchange mechanism enables variation in F-box protein expression. Mol Cell 69(5):773–86 e6
Lo SC, Hannink M (2006) CAND1-mediated substrate adaptor recycling is required for efficient repression of Nrf2 by Keap1. Mol Cell Biol 26(4):1235–1244
Lyapina S, Cope G, Shevchenko A et al (2001) Promotion of NEDD-CUL1 conjugate cleavage by COP9 signalosome. Science 292(5520):1382–1385
Makino Y, Yogosawa S, Kayukawa K et al (1999) TATA-binding protein-interacting protein 120, TIP120, stimulates three classes of eukaryotic transcription via a unique mechanism. Mol Cell Biol 19(12):7951–7960
Mendoza HM, Shen L-N, Botting C et al (2003) NEDP1, a highly conserved cysteine protease that deNEDDylates cullins. J Biol Chem 278(28):25637–25643
Mergner J, Schwechheimer C (2014) The NEDD8 modification pathway in plants. Front Plant Sci 5:103
Min KW, Hwang JW, Lee JS et al (2003) TIP120A associates with cullins and modulates ubiquitin ligase activity. J Biol Chem 278(18):15905–15910
Min KW, Kwon MJ, Park HS et al (2005) CAND1 enhances deneddylation of CUL1 by COP9 signalosome. Biochem Biophys Res Commun 334(3):867–874
Monda JK, Scott DC, Miller DJ et al (2013) Structural conservation of distinctive N-terminal acetylation-dependent interactions across a family of mammalian NEDD8 ligation enzymes. Structure 21(1):42–53
Morimoto M, Nishida T, Honda R et al (2000) Modification of cullin-1 by ubiquitin-like protein Nedd8 enhances the activity of SCF(skp2) toward p27(kip1). Biochem Biophys Res Commun 270(3):1093–1096
Mosadeghi R, Reichermeier KM, Winkler M et al (2016) Structural and kinetic analysis of the COP9-signalosome activation and the cullin-RING ubiquitin ligase deneddylation cycle. Elife 5:e12102
Osaka F, Kawasaki H, Aida N et al (1998) A new NEDD8-ligating system for cullin-4A. Genes Dev 12(15):2263–2268
Osaka F, Saeki M, Katayama S et al (2000) Covalent modifier NEDD8 is essential for SCF ubiquitin-ligase in fission yeast. EMBO J 19(13):3475–3484
Oshikawa K, Matsumoto M, Yada M et al (2003) Preferential interaction of TIP120A with Cul1 that is not modified by NEDD8 and not associated with Skp1. Biochem Biophys Res Commun 303(4):1209–1216
Ou CY, Lin YF, Chen YJ et al (2002) Distinct protein degradation mechanisms mediated by Cul1 and Cul3 controlling Ci stability in Drosophila eye development. Genes Dev 16(18):2403–2414
Pierce NW, Lee JE, Liu X et al (2013) Cand1 promotes assembly of new SCF complexes through dynamic exchange of F box proteins. Cell 153(1):206–215
Pintard L, Kurz T, Glaser S et al (2003) Neddylation and deneddylation of CUL-3 is required to target MEI-1/Katanin for degradation at the meiosis-to-mitosis transition in C. elegans. Curr Biol 13(11):911–921
Podust VN, Brownell JE, Gladysheva TB et al (2000) A Nedd8 conjugation pathway is essential for proteolytic targeting of p27Kip1 by ubiquitination. Proc Natl Acad Sci U S A 97(9):4579–4584
Pozo JC (1998) The ubiquitin-related protein RUB1 and auxin response in Arabidopsis. Science 280(5370):1760–1763
Rao-Naik C, Delacruz W, Laplaza JM et al (1998) The rub family of ubiquitin-like proteins. J Biol Chem 273(52):34976–34982
Read MA, Brownell JE, Gladysheva TB et al (2000) Nedd8 modification of Cul-1 activates SCFbeta TrCP-dependent ubiquitination of Ikappa Balpha. Mol Cell Biol 20(7):2326–2333
Reitsma JM, Liu X, Reichermeier KM et al (2017) Composition and regulation of the cellular repertoire of SCF ubiquitin ligases. Cell 171(6):1326–39 e14
Saha A, Deshaies RJ (2008) Multimodal activation of the ubiquitin ligase SCF by Nedd8 conjugation. Mol Cell 32(1):21–31
Sakata E, Yamaguchi Y, Miyauchi Y et al (2007) Direct interactions between NEDD8 and ubiquitin E2 conjugating enzymes upregulate cullin-based E3 ligase activity. Nat Struct Mol Biol 14(2):167–168
Schmidt MW, Mcquary PR, Wee S et al (2009) F-box-directed CRL complex assembly and regulation by the CSN and CAND1. Mol Cell 35(5):586–597
Schwechheimer C, Serino G, Callis J et al (2001) Interactions of the COP9 signalosome with the E3 ubiquitin ligase SCFTIRI in mediating auxin response. Science 292(5520):1379–1382
Scott DC, Monda JK, Grace CR et al (2010) A dual E3 mechanism for Rub1 ligation to Cdc53. Mol Cell 39(5):784–796
Scott DC, Monda JK, Bennett EJ et al (2011) N-terminal acetylation acts as an avidity enhancer within an interconnected multiprotein complex. Science 334(6056):674–678
Scott DC, Sviderskiy VO, Monda JK et al (2014) Structure of a RING E3 trapped in action reveals ligation mechanism for the ubiquitin-like protein NEDD8. Cell 157(7):1671–1684
Shih SC, Sloper-Mould KE, Hicke L (2000) Monoubiquitin carries a novel internalization signal that is appended to activated receptors. EMBO J 19(2):187–198
Sloper-Mould KE, Jemc JC, Pickart CM et al (2001) Distinct functional surface regions on ubiquitin. J Biol Chem 276(32):30483–30489
Soucy TA, Smith PG, Milhollen MA et al (2009) An inhibitor of NEDD8-activating enzyme as a new approach to treat cancer. Nature 458(7239):732–736
Tateishi K, Omata M, Tanaka K et al (2001) The NEDD8 system is essential for cell cycle progression and morphogenetic pathway in mice. J Cell Biol 155(4):571–579
Wada H, Kito K, Caskey LS et al (1998) Cleavage of the C-terminus of NEDD8 by UCH-L3. Biochem Biophys Res Commun 251(3):688–692
Wada H, Yeh ET, Kamitani T (1999) Identification of NEDD8-conjugation site in human cullin-2. Biochem Biophys Res Commun 257(1):100–105
Walden H, Podgorski MS, Huang DT et al (2003a) The structure of the APPBP1-UBA3-NEDD8-ATP complex reveals the basis for selective ubiquitin-like protein activation by an E1. Mol Cell 12(6):1427–1437
Walden H, Podgorski MS, Schulman BA (2003b) Insights into the ubiquitin transfer cascade from the structure of the activating enzyme for NEDD8. Nature 422(6929):330–334
Wee S, Geyer RK, Toda T et al (2005) CSN facilitates cullin-RING ubiquitin ligase function by counteracting autocatalytic adapter instability. Nat Cell Biol 7(4):387–391
Whitby FG, Xia G, Pickart CM et al (1998) Crystal structure of the human ubiquitin-like protein NEDD8 and interactions with ubiquitin pathway enzymes. J Biol Chem 273(52):34983–34991
Wilkinson KD (1988) Purification and structural properties of ubiquitin. Ubiquitin. Springer, pp 5–38
Wu K, Chen A, Pan ZQ (2000) Conjugation of Nedd8 to CUL1 enhances the ability of the ROC1-CUL1 complex to promote ubiquitin polymerization. J Biol Chem 275(41):32317–32324
Wu K, Chen A, Tan P et al (2002) The Nedd8-conjugated ROC1-CUL1 core ubiquitin ligase utilizes Nedd8 charged surface residues for efficient polyubiquitin chain assembly catalyzed by Cdc34. J Biol Chem 277(1):516–527
Wu K, Yamoah K, Dolios G et al (2003) DEN1 is a dual function protease capable of processing the C terminus of Nedd8 and deconjugating hyper-neddylated CUL1. J Biol Chem 278(31):28882–28891
Wu S, Zhu W, Nhan T et al (2013) CAND1 controls in vivo dynamics of the cullin 1-RING ubiquitin ligase repertoire. Nat Commun 4:1642
Yamoah K, Oashi T, Sarikas A et al (2008) Autoinhibitory regulation of SCF-mediated ubiquitination by human cullin 1’s C-terminal tail. Proc Natl Acad Sci U S A 105(34):12230–12235
Yogosawa S, Makino Y, Yoshida T et al (1996) Molecular cloning of a novel 120-kDa TBP-interacting protein. Biochem Biophys Res Commun 229(2):612–617
Zemla A, Thomas Y, Kedziora S et al (2013) CSN- and CAND1-dependent remodelling of the budding yeast SCF complex. Nat Commun 4:1641
Zhang W, Ito H, Quint M et al (2008) Genetic analysis of CAND1-CUL1 interactions in Arabidopsis supports a role for CAND1-mediated cycling of the SCFTIR1 complex. Proc Natl Acad Sci U S A 105(24):8470–8475
Zheng N, Schulman BA, Song L et al (2002a) Structure of the Cul1-Rbx1-Skp1-F boxSkp2 SCF ubiquitin ligase complex. Nature 416(6882):703–709
Zheng J, Yang X, Harrell JM et al (2002b) CAND1 binds to unneddylated CUL1 and regulates the formation of SCF ubiquitin E3 ligase complex. Mol Cell 10(6):1519–1526
Zhou C, Wee S, Rhee E et al (2003) Fission yeast COP9/signalosome suppresses cullin activity through recruitment of the deubiquitylating enzyme Ubp12p. Mol Cell 11(4):927–938
Zhou W, Xu J, Tan M et al (2018) UBE2M is a stress-inducible dual E2 for neddylation and ubiquitylation that promotes targeted degradation of UBE2F. Mol Cell 70(6):1008–24 e6
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Wang, K., Deshaies, R.J., Liu, X. (2020). Assembly and Regulation of CRL Ubiquitin Ligases. In: Sun, Y., Wei, W., Jin, J. (eds) Cullin-RING Ligases and Protein Neddylation. Advances in Experimental Medicine and Biology, vol 1217. Springer, Singapore. https://doi.org/10.1007/978-981-15-1025-0_3
Download citation
DOI: https://doi.org/10.1007/978-981-15-1025-0_3
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-1024-3
Online ISBN: 978-981-15-1025-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)