Abstract
Bovine viral diarrhea (BVD) is prevalent worldwide and causes high economic losses in cattle due to a variety of disease syndromes. BVD is caused by three bovine pestiviruses, bovine viral diarrhea virus 1 (BVDV-1), BVDV-2 and HoBi-like pestivirus (HoBiPeV) with considerable genetic and antigenic heterogeneity. Bovine pestiviruses belong to the Pestivirus genus within the Flaviviridae family that also comprises the genera Flavivirus, Hepacivirus and Pegivirus. As per the latest (10th) ICTV report, pestiviruses have been classified into 11 approved species, including bovine pestiviruses, which have been classified into species, Pestivirus A, Pestivirus B and Pestivirus H. The term BVDV in this chapter commonly refers to all the three bovine pestiviruses. The pathogenesis of BVDV infection is complex, with infection pre- and post-gestation leading to different outcomes. BVDVs are highly successful to persist and spread in their host populations due to their unique ability to produce persistent infection through evasion of adaptive immune response and innate immune response. Recent advances in diagnostic methods, nucleotide sequencing, and computer-assisted phylogenetic analyses have so far identified 21 BVDV-1 subtypes (BVDV-1a to BVDV-1u), 4 BVDV-2 subtypes (BVDV-2a to BVDV-2d) and 4 HoBiPeV subtypes (HoBiPeV-a to HoBiPeV-d). Providing acquired immune protection against BVDV is challenging due to the antigenic diversity among BVDV strains and ability of BVDV to infect the fetus. Both killed and live attenuated vaccines have been reported to be effective in the field, and recent advancements in molecular studies have helped toward future development of new-generation vaccines against BVD. However, over the years, vaccination alone has not resulted in the elimination of BVDV-related clinical disease or a significant reduction in BVDV losses. All successful BVDV control programs are based on identification and removal of PI animals, movement controls, strict biosecurity and surveillance. To date, BVDV control programs without vaccination have been implemented successfully in Scandinavian countries, Austria and Switzerland, while control with vaccination has been used in Germany, Belgium, Ireland and Scotland. This chapter will focus on advances in research involving all aspects of BVDV with special emphasis on molecular biology, genetic and antigenic diversity, diagnosis, prevention and control besides discussion on future perspectives.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Pestivirus
- Bovine viral diarrhea virus
- Persistent infection
- BVDV-1
- BVDV-2
- HoBiPeV
- Epidemiology
- Diagnosis
- Control
- Cattle
1 Prologue
Bovine viral diarrhea (BVD) is one of the major economically important viral diseases of cattle and is prevalent in cattle populations worldwide. From its discovery in 1946 to the present date, without doubt BVD is one of the most complex infectious diseases encountered in veterinary medicine with regard to its pathogenesis, diagnosis, management and control. Bovine viral diarrhea virus (BVDV) causes BVD and mucosal disease (MD) and is highly complex, with respect to the heterogeneity in genetic and antigenic properties, host spectrum, host-virus interaction, virulence and immune response. Although its impact on cattle health and production remained underestimated for long time, the economic impact has recently been appreciated with control programs being implemented in several countries in Europe with an ultimate goal of BVD eradication.
Bovine viral diarrhea (BVD) was first reported in the USA in 1946 (Olafson et al. 1946) in association with epizootics of acute disease characterized by fever, leukopenia, reduced milk yield, high rates of abortion, diarrhea and erosive lesions of the digestive tract of cattle. The symptoms were similar to rinderpest (RP) with morbidity of 33–88% and mortality of 4–8%. In the 1950s, a special form of viral diarrhea called mucosal disease (MD), with hemorrhages and intestinal erosions, was reported in Iowa, USA (Ramsey and Chivers 1953). However, the relationship between the two illnesses could be established after many years, when it was clear that both BVD and MD were caused by BVDV.
Two biotypes of BVDV exist based on their effects on cell cultures, the non-cytopathic (ncp) and cytopathic (cp). First isolation of ncp BVDV from BVD clinical cases was reported by Lee and Gillespie (1957), while cp BVDV was isolated first by Underdahl et al. (1957). Although BVD could be reproduced early experimentally, it took many years until the 1980s when MD could be experimentally reproduced, hypothesis of immune tolerance was proved, and mechanisms of persistent infection (PI) and pathogenesis of MD were established (Malmquist 1968; McClurkin et al. 1984; Brownlie et al. 1984). Only ncp BVDVs establish persistent infections, while cp BVDV is generated in PI animals by mutations in ncp BVDV and causes MD.
In 1987, BVD associated with thrombocytopenia, hemorrhage and high mortality was reported in US dairy herds (Perdrizet et al. 1987) followed by other reports from the USA and Canada (Bolin and Ridpath 1992; Pellerin et al. 1994). The causative agent involved in these outbreaks was identified as BVDV-2 and the classical BVDV strains identified earlier were termed as BVDV-1 (Ridpath et al. 1994). First identified in fetal bovine serum originating from Brazil in 2004, HoBi-like pestivirus has recently been recognized as a bovine pathogen in Europe, South America and Asia that causes clinical symptoms akin to the classical BVDV-1 or BVDV-2 infections (Schirrmeier et al. 2004; Liu et al. 2009; Bauermann et al. 2013; Mishra et al. 2014).
Animal health economics highlight the importance of the disease and play a major role in decision-making process regarding selection of control strategies. Although prevalence rates vary, BVDV is prevalent in most of the cattle populations. BVDV infection causes significant economic losses in cattle production and the economic impact varies within and between countries (Richter et al. 2017). Direct losses due to BVDV occur on account of increased morbidity and mortality in adult cattle due to acute infection, reduced milk yield, respiratory disorders, extended calving interval, reproductive disorders such as repeat breeding and abortions, congenital defects, increased neonatal mortality, non-thriving and death among young stock besides the costs for treatment and prevention (Houe 2003). BVDV-induced indirect losses arise due to implementation of control programs and trade restrictions.
BVDV can have devastating effects on the economy of individual dairy farmers which has been seen in the form of severe acute BVD outbreaks in many countries in North America and Europe, specifically in the USA, Canada and Germany. Significant differences exist in the virulence of BVDV strains, and both BVDV-1 and BVDV-2 have high or low virulent strains, and hence wide differences in economic impact due to BVD have been reported in various countries (Houe 1999). A recent study on direct financial losses due to BVDV infection in 15 countries around the world over the past 30 years has shown that direct financial losses due to BVDV were in the range of 0.50–687.80 USD per animal and the average direct losses were higher per dairy cow than per beef cow (Richter et al. 2017). Another study analyzed the data of 31 published studies undertaken around the world during 1991–2015 and has shown that the economic impact of BVD ranges from 0 to 552 GBP per cow per year and is based on outcome of the disease (Yarnall and Thrusfield 2017). Despite variation in calculation methods, calculation of economic losses has been a significant motivator for considering implementation of BVD control programs and mitigation activities in many countries in Europe.
The aim of this chapter is not only to summarize the available facts on BVDV but also to critically analyze the data and highlight recent advances on BVDV taxonomy, molecular biology, epidemiology, diagnosis and control.
2 Taxonomy
Bovine pestiviruses belong to the Pestivirus genus in the family Flaviviridae that also comprises the genera Flavivirus, Hepacivirus and Pegivirus. As per the ninth report of the International Committee on Taxonomy of Viruses (ICTV), the Pestivirus genus was comprised of four approved species, Bovine viral diarrhea virus 1 (BVDV-1), Bovine viral diarrhea virus 2 (BVDV-2), classical swine fever virus (CSFV), and border disease virus (BDV), and four tentative species, namely, Giraffe-1 pestivirus; Pronghorn antelope pestivirus; atypical bovine pestivirus, also termed as BVDV-3 or HoBi-like pestivirus; and Bungowannah virus (Simmonds et al. 2012). While several other pestiviruses have been proposed as additional species including the atypical porcine pestivirus (APPV), Linda virus, bat (Rhinolophus affinis) pestivirus and Norway rat pestivirus, there was no change in the taxonomy of Pestivirus genus by ICTV during 1999–2017.
Pestiviruses have several characteristics that differentiate them from other members of the Flaviviridae family. The two proteins unique to the Pestivirus genus are the Erns envelope glycoprotein, which has RNase activity, and the nonstructural protease Npro, which releases itself autocatalytically from the polyprotein. On the basis of genetic and antigenic characteristics, the four existing Pestivirus species have been demarcated using a range of criteria including complete coding nucleotide sequences that differ by more than 25%, displaying >10-fold differences in cross-neutralization titers, and may have differing or overlapping host range (Becher et al. 2003). Based on genetic analysis, several genotypes or subtypes within Pestivirus species have been proposed, but these subdivisions have not yet been officially approved.
The taxonomy of pestiviruses was problematic for long, since the earlier Pestivirus species names had been derived from names of virus isolates, which were based on host range and disease attributes. Hence, a new uniform naming system, analogous to that used for species belonging to Pegivirus and Hepacivirus genera of Flaviviridae, has been approved by ICTV recently for Pestivirus species with the format Pestivirus X, where X represents a different capital letter for each species without change in virus isolate names (Smith et al. 2017). The four existing species have been designated as Pestivirus A, which comprises of Bovine viral diarrhea virus 1 (BVDV-1); Pestivirus B, which comprises of Bovine viral diarrhea virus 2 (BVDV-2); Pestivirus C, which comprises of Classical swine fever virus (CSFV); and Pestivirus D, comprising of Border disease virus (BDV) along with seven additional species. Atypical bovine pestivirus or Hobi-like pestivirus (HoBiPeV) or BVDV-3 belongs to the species Pestivirus H.
3 BVDV Structure
The BVDV virions are enveloped, spherical particles which are 40–60 nm diameter in size. The BVD virus particle is composed of a core region, consisting of the genomic RNA coated with structural capsid or core protein which is surrounded by a lipid envelope. The capsid is about 30 nm diameter and appears as an electron-dense inner core (Horzinek et al. 1971). Cryo-electron microscopy of purified BVDV virions has recently shown that viral particles display an electron-dense capsid surrounded by a phospholipid bilayer with no visible spikes and most BVDV particles are 50 nm diameter in size and about 2% are 65 nm in size, suggesting some size flexibility during BVDV morphogenesis (Callens et al. 2016). However, whether BVDV capsid is icosahedral or not remains to be determined in future.
The lipid bilayer envelope is made up of three virus-encoded glycoproteins, Erns, E1, and E2. The heavily glycosylated Erns glycoprotein is loosely associated with the virus particle and is secreted in soluble form by infected cells, while E1 and E2 glycoproteins are integral membrane proteins and E1-E2 heterodimer is essential for BVDV entry. BVDV particles have a higher concentration of Erns than E1 and E2. Due to the pleomorphic nature of envelope and association of BVDV particles with host cells, it is extremely difficult to achieve highly purified infectious particles by ultracentrifugation and identification by electron microscopy. The buoyant density of virion in sucrose is 1.134 gm/ml and molecular weight of the virion is estimated as 6.0 × 107 (Lindenbach et al. 2013). Similar to other enveloped viruses, BVDV is inactivated by organic solvents and detergents, but it is resistant to low pH unlike flaviviruses.
4 Genomic Organization
Similar to other pestiviruses, BVDV genome consists of a positive-sense single-stranded RNA of about 12.3 kb size. However, the size may vary up to 16.5 kb due to insertions, genomic duplications, and genomic recombination events (Becher et al. 1999). No subgenomic RNA is transcribed during BVDV replication and the plus strand genomic RNA only represents the viral mRNA and codes all viral proteins.
The genomic organization of BVDV is similar to all the recognized pestiviruses (Fig.14.1). The RNA genome codes for a polyprotein of about 3900 amino acids in 1 large open reading frame that is flanked by 5′- and 3′-untranslated regions (UTR) of about 400 and 200 nucleotides, respectively. The translated polyprotein is then processed by viral and cellular proteases resulting in 12–13 proteins, Npro; C (capsid); the envelope proteins Erns, E1, and E2; and nonstructural proteins p7, NS2, NS3, NS2–3, NS4A, NS4B, NS5A and NS5B (Lindenbach et al. 2013). The arrangement of proteins in the polyprotein is NH2–Npro/C/Erns/E1/E2/p7/NS2/NS3/NS4A/NS4B/NS5A/NS5B–COOH.
The 5′-UTR of BVDV has a stable stem loop structure (Ia hairpin), which is involved in both translation initiation and replication of the viral RNA (Grassmann et al. 2005). BVDV RNAs do not possess a 5′-cap and is not polyadenylated at 3′-end. The internal ribosomal entry site (IRES), located toward the end of 5′-UTR and first part of Npro, directly recruits the small ribosomal subunit, positions it at the translational start site, and promotes start of translation even in the presence of a non-AUG codon (Poole et al. 1995; Pestova et al. 2008). The 3′-UTR has a conserved stem-loop structure, and the single-stranded domain at the 3′-end of the genomic RNA is necessary for efficient RNA replication. Significant variation exists in the length of 3′-UTR sequence among pestiviruses including BVDV. Although BVDV 3′-UTR is less conserved than 5′-UTR, it has both conserved and variable parts. BVDV-1 isolates have an eight-nucleotide repeat sequence (TGTATATA) in the variable part of 3′-UTR, while BVDV-2 isolates contain TGTAAATA repeat sequence.
Polyprotein processing occurs co-translationally and begins with the N-terminal autoprotease (Npro) release. The Npro, found only in pestiviruses, is responsible for cleavage at its own carboxy-terminus, thereby releasing amino-terminus of the C protein (Stark et al. 1993). The C/Erns cleavage occurs by cellular signal peptidase (SPase) and signal peptide peptidase (SPPase), while processing at the Erns/E1, E1/E2, E2/p7 and p7/NS2 borders is carried out by SPase (Rumenapf et al. 1993). The processing of NS2 and NS3 occurs by protease located within NS2, whereas all processing downstream of NS3 is done by NS3 protease. NS4A acts as a cofactor of the NS3 protease and is essential for NS4B/NS5A and the NS5A/NS5B site processing.
BVDV strains exist as two biotypes, non-cytopathic (ncp) and cytopathic (cp). Cytopathic viruses result following changes in the NS2/3 protein coding region leading to generation of NS2 and NS3 proteins. These genomic changes include point mutations, genomic duplications, and insertion of cellular mRNA sequences (Meyers and Thiel 1996; Kummerer and Meyers 2000). The genomic changes in the Npro, capsid and NS4B have also been reported in some cp strains of BVDV.
5 Viral Structural and Nonstructural Proteins
With the exception of Npro, the first coding region of the ORF encodes the structural proteins which are integral components of the viral particle. These include the capsid or core protein and three glycoproteins, Erns, E1 and E2. BVDV capsid protein C is highly basic and binds RNA with low affinity. The Erns glycoprotein, present on the virus particle and in virus-free supernatant or in the blood of infected cells, exhibits several unusual characteristics including its exceptional membrane anchor and RNase activity (Lindenbach et al. 2013). Erns is highly glycosylated and commonly exists as disulfide-linked homodimers (Hulst and Moormann 2001). RNase activity of Erns is responsible for both single-stranded and double-stranded RNA degradation, which is considered important in limiting the host innate immune response. However, the enzyme becomes active only in endoplasmic reticulum environment and when it gets separated from the viral genome by the lipid envelope.
The E1 and E2 glycoproteins are integral membrane proteins and interact to form disulfide-linked E1-E2 heterodimers which are responsible for BVDV entry and infectivity (Ronecker et al. 2008). The structure and function of E1 is not yet known and antibodies against E1 are not found in infected animals. However, it has recently been proposed that E1 contains the fusion peptide necessary for the fusion during entry. The 53 kDa BVDV E2 glycoprotein is responsible for virus attachment, generation of neutralizing antibodies, and host tropism (Weiland et al. 1990; Liang et al. 2003). BVDV E2 in general is 373 amino acids long and contains 17 cysteine residues in the homodimer form with four highly conserved N-glycosylation sites. BVDV E2 possesses three domains, domain I and II contain neutralizing epitopes that are exposed on the viral surface, while the domain III acts as an anchor.
The first nonstructural protein, Npro, is an autoprotease and is unique to pestiviruses and comprises 168 amino acids in all the pestiviruses detected so far. In BVDV-infected cells, it inhibits interferon (IFN-1) production and thereby interferes with antiviral activity of the host cell. Although earlier studies showed that Npro is a papain-like cysteine protease with Glu22-His49-Cys69 building a catalytic triad, a subsequent study classified it into C53 protease family (Rawlings et al. 2012). BVDV Npro crystallographic study has revealed that a catalytic dyad of His49 and Cys69 is responsible for proteolytic activity of Npro (Zogg et al. 2013). The p7 nonstructural protein, existing as free p7 or E2-p7, has ion channel activity and a role in assembly of infectious progeny virus.
The NS2 and NS3 proteins are found predominantly as the unprocessed NS2/3 in cells infected with ncp viruses and primarily as NS2 and NS3 in cells infected with cp viruses, but recent studies have shown that cleavage of NS2/3 is necessary for replication of ncp viruses very early in the infection (Meyers and Thiel 1996; Lackner et al. 2004; Mishra et al. 2010). The NS2 protease is essential for efficient NS2-3 cleavage, while cellular Jiv protein acts as an essential cofactor for ncp viruses (Tautz et al. 1996; Lackner et al. 2004). Insertions of cellular sequences or duplication of BVDV genomic sequences commonly occurs in NS2-3 region. The 80 kDa NS3 protein is immunogenic and possesses two distinct enzymatic activities, the N-terminus serine protease domain, which along with cofactor NS4A is necessary for proteolytic cleavage of viral polyprotein beyond NS3 and virus viability, and the C-terminus RNA helicase and NTPase domain that participates in RNA replication.
The NS4A protein functions as a cofactor for the NS3 serine protease and has an important role in the morphogenesis of virions. NS4B is predicted as an integral membrane protein localized at intracellular membranes and is associated with RNA replication complex. NS5A co-localizes with membrane-bound NS4B and NS5B in the RNA replication complex and BVDV NS5A is tolerant to some deletions but its role in viral replication is not clear. The NS5B protein, containing GDD and NTPase functional motifs, acts as RNA-dependent RNA polymerase and functions as the major protein in genomic RNA replication, and C-terminus of BVDV NS5B is important for morphogenesis of virions.
6 BVDV Attachment and Entry
BVDV entry into bovine cells involves virion attachment to cellular receptors, internalization, and membrane fusion. Bovine CD46 has been reported to act as a cellular receptor for BVDV (Maurer et al. 2004), but it may not be sufficient and some unknown elements may be required for BVDV entry. For initiating the infection process, BVDV E1-E2 heterodimer binds the cellular receptor. BVDV entry into bovine cells occurs through clathrin-dependent endocytosis and endosomal fusion in a pH-dependent manner (Grummer et al. 2004; Krey et al. 2005), and a similar mechanism exists for entry into ovine cells (Mathapati et al. 2010).
Following uncoating and release of the genomic RNA, transcription and translation of viral proteins take place in cellular cytoplasm. However, the mechanisms of viral assembly and release from the cell following translation and maturation of viral proteins are not clear at present. BVDV virions are assembled in the endoplasmic reticulum (ER) along with final post-translational modifications. The viral proteins are maintained exclusively in the ER and Golgi bodies, hence are not displayed on the cell surface.
7 BVDV Replication
BVDV replication takes place in cellular cytoplasm. Following IRES-mediated initiation of translation, BVDV replication process begins with a positive strand replicase complex consisting of viral and cellular components formed at the 3′-terminus of the genome. The replicase complex catalyzes transcription of positive-sense RNA into full-length complementary negative-sense RNA, which then acts as template for synthesizing additional positive-sense RNA, using a semiconservative asymmetric replication model (Warrilow et al. 2000). In comparison to negative-sense RNA, a large amount of newly synthesized positive-sense RNA is generated. In this model, three virus-specific RNAs, a double-stranded replicative form (RF), a partially single-stranded and partially double-stranded replicative intermediate (RI) and a single-stranded viral RNA are involved. The same positive-sense genomic RNA acts as a template for both replication and translation. Regulation of this process is mediated by secondary structure of IRES in the 5′-UTR. The budding of BVDV takes place in endoplasmic reticulum, where the RNA-core complex is packed in envelopes (Schmeiser et al. 2014). Following assembly, virions are released from the cell through the secretory pathway.
8 Genetic and Antigenic Diversity
BVDV strains show high genetic diversity, which results from accumulation of point mutations, homologous and non-homologous RNA recombination. Variation in evolutionary rates (5.9 × 10−4 to 9.3 × 10−3 substitutions/site/year) has been reported for BVDV-1. Mutations may lead to producing a population of viruses, called quasispecies, with each possessing a small number of nucleotide differences from the population mean. Like other RNA viruses, BVDV isolates exist as quasispecies in infected animals. A recent study has shown that similar to classical BVDVs, differences exist between the swarms circulating within HoBiPeV PI animals from the same inoculum suggesting involvement of host factors in the selection of genetic variants in PI animals (Weber et al. 2015). Besides the reports of homologous RNA recombination in BVDV-1 and BVDV-2, generation of cp BVDV variants with a variety of genomic alterations following non-homologous RNA recombination has been described. BVDV antigenic diversity happens by replication in vaccinated or previously infected animals, but a recent study has shown that antigenic changes may arise in absence of an immune response.
Genetic typing of BVDV strains is important for BVD epidemiology and control. Accurate genetic typing of BVDV has been obtained from the application of advanced molecular techniques such as reverse transcription polymerase chain reaction (RT-PCR), next-generation sequencing, and phylogenetic analysis. Although sequence analysis of the highly conserved 5′-UTR is mostly used for the classification of pestivirus isolates into species level, segregation into subtypes/subgenotypes within each species is more accurate by analysis of complete Npro and E2 genes (Becher et al. 2003; Vilcek et al. 2001, 2004; Yesilbag et al. 2017). However, discrepancies in allocation of some BVDV isolates into subtypes have been reported either due to inconsistent use of different genomic regions or inconsistent use of methods for phylogenetic analysis. Additionally, antigenic similarity detected by monoclonal antibody (MAb) binding and cross-neutralization assays with homologous and heterologous antisera is used for determining antigenic diversity of BVDV strains (Paton et al. 1995; Becher et al. 2003; Dias et al. 2017).
So far, BVDV-1 has been segregated into 21 subtypes (1a–1u), while BVDV-2 has been segregated into 4 subtypes (2a–2d) and HoBiPeV has been segregated into 4 subtypes (a–d). Global distribution of BVDV subtypes is shown in Table 14.1. In India, all the three bovine pestivirus species, Pestivirus A, Pestivirus B and Pestivirus H and two genotypes/subtypes of BVDV-1 (1b, 1c), BVDV-2 (2a, 2b), and HoBiPeV (c, d) within these species have been detected so far (Mishra et al. 2011, 2014).
8.1 Genetic and Antigenic Diversity of BVDV-1
BVDV-1, which belongs to species Pestivirus A, is prevalent worldwide. Although initially divided into two subtypes, BVDV-1a (NADL-like) and BVDV-1b (Osloss-like), BVDV-1 isolates originating from different countries could be segregated into 11 subtypes subsequently (Vilcek et al. 2001). Further studies on genetic typing have revealed existence of at least 21 subtypes of BVDV-1 (Yesilbag et al. 2014, 2017; Giammarioli et al. 2015) and additional subtypes is likely in the future. Despite accurate distribution of BVDV subtypes in individual countries and continents is unknown, the published reports have revealed that BVDV-1b is the predominant subtype worldwide, followed by BVDV-1a and 1c. BVDV-1b is the predominant subtype in Europe, Asia and Americas, while BVDV-1c is predominantly prevalent in Australia and Mexico. Extensive genetic diversity of BVDV-1 has been reported from several countries in Europe and also in Turkey, China and Japan, but it is comparatively lesser in Americas, Africa and Australia as well as in India.
Although BVDV-1 isolates are antigenically closely related than BVDV-2 or HoBiPeV, significant antigenic differences between BVDV-1 subtypes have been found similar to that observed between BVDV-1 and BVDV-2 or HoBiPeV. Hence, significant antigenic differences should be taken into consideration while developing vaccines and designing effective control programs. Significant antigenic differences have been reported between BVDV-1a and BVDV-1b strains in Europe (Becher et al. 2003), between BVDV-1a, BVDV-1b and BVDV-1c strains in Chile and the USA (Pizarro-Lucero et al. 2006; Ridpath et al. 2010), between BVDV-1e and BVDV-1 k strains in Switzerland (Bachofen et al. 2008), and between BVDV-1n and BVDV-1o and BVDV-1a, BVDV-1b, BVDV-1c and BVDV-1j strains in Japan (Nagai et al. 2008).
8.2 Genetic and Antigenic Diversity of BVDV-2
Although distributed in all continents, BVDV-2 strains are genetically less diverse than BVDV-1 and occur less commonly than BVDV-1. Although BVDV-2 has been divided into four (2a–2d) subtypes, the strain belonging to 2d subtype is questionable due to a single report from Argentina. BVDV-2a is the most prevalent subtype in all continents, while BVDV-2b has been detected in Americas (Brazil, Argentina, Uruguay, USA), Europe (Portugal, Spain, Slovakia, Turkey) and Asia (India, China), and BVDV-2c has been detected only in Europe (Germany, Ireland) and North America (USA). BVDV-2 can occur also in sheep and goats. In sheep, BVDV-2a has been detected in the USA and Italy, while BVDV-2b has been reported from India (Mishra et al. 2008b) and Turkey (Yesilbag et al. 2008). However, only BVDV-2a subtype has so far been reported in goats (Mishra et al. 2007a) and significant antigenic variation between BVDV-2 subtypes has not yet been observed.
8.3 Genetic and Antigenic Diversity of HoBiPeV
Although less commonly found, clinical disease following natural HoBiPeV infections is similar to that caused by classical BVDV-1 and BVDV-2 infections. Till 2014, all the previously reported HoBiPeV strains, except the Thai and Bangladesh strains, were found to be closely related genetically. However, our previous work on the basis of sequence analysis of combined datasets of 5′-UTR and full Npro gene showed that HoBiPeV strains can be classified into three subtypes with two highly divergent HoBiPeV lineages co-circulating in Indian cattle (Mishra et al. 2014). Further studies have shown that HoBiPeV strains can be classified into 4 subtypes (a–d). HoBiPeV-a has been detected in South America (Brazil) and Europe (Italy) and as contaminants of FBS, while HoBiPeV-b has been reported from Bangladesh and HoBiPeV-c and HoBiPeV-d have been detected in India (Mishra et al. 2014; Giammarioli et al. 2015). Marked antigenic differences exist between BVDV-1, BVDV-2 and HoBiPeV, while minor to moderate antigenic variation among HoBiPeV-a field isolates has been reported recently in Brazil.
9 Epidemiology of BVD
A large number of variables, such as clinical, pathological, virological, serological and production measures can be used to quantify the occurrence of BVDV infections. Long presence of antibodies (often lifelong) in acutely infected animals and presence of virus in PI animals throughout their life make prevalence studies more suitable for BVD. Sensitivity and specificity of diagnostic tests play a key role in determining true prevalence of BVDV infection. BVDV antibody prevalence studies are useful mostly in unvaccinated populations. Overall, PI animals play a major role in virus transmission than transiently infected cattle, since they shed virus in large amounts in all bodily fluids throughout their life, while virus shedding is limited to only for a few days or weeks in most other viral diseases of livestock. Vertical transmission and transmission through semen also play vital roles in epidemiology and BVDV is introduced into a susceptible herd mostly by introduction of PI animals or pregnant animals carrying a PI fetus.
9.1 Abroad
BVDV infections are widespread throughout the world except in a few countries in Europe, where it has been eradicated or in final stages of eradication. Cattle of all ages are susceptible to BVDV infection and are the primary hosts. Buffaloes and domestic non-bovid species including sheep, goats, new world camelids and swine have also been reported to carry and spread BVDV. Natural infection of BDV in cattle has been reported with clinical signs similar to BVD. Despite variation in prevalence rates among surveys, BVDV infection is endemic in many populations having 1–2% of the cattle being persistently infected (PI) and > 90% of the cattle being antibody positive. Variations in BVDV prevalence rates among different countries or regions or introduction of virus into BVDV-naïve herds is often determined by cattle population density, cattle trade and pasturing practices.
BVDV prevalence is high in areas with high cattle population density and larger herds. BVDV-1 and BVDV-2 are distributed in all continents, while HoBiPeV has so far been reported in South America, Europe and Asia. Geographical patterns in distribution of BVDV-1 subtypes have been reported around the world. BVDV-1 subtypes, 1m, 1n, 1o, 1p, and 1q have been found only in some countries in Asia, while 1f, 1g, 1h, 1k, 1l, 1r, 1s and 1t have been found exclusively in Europe. Besides cattle, buffaloes, sheep and goats, a number of BVDV-1 subtypes have been identified in wild ruminants. BVDV-1a has been detected in Canadian bison; BVDV-1b in alpaca, pudu, Canadian bison and bongo; BVDV-1c in yak and deer; BVDV-1d in roe deer; BVDV-1f in mouse deer; and BVDV-1j in deer (Vilcek and Nettleton 2006; Mishra et al. 2008a).
BVDV-2, first detected in cattle of the USA and Canada in association with hemorrhagic disease with high mortalities, was found later in several other countries of South America, Europe and Asia and also in Australia (Vilček et al. 2005). BVDV-2a is the predominant BVDV-2 subtype circulating around the world. A very special case of fatal disease with high mortalities in cattle resulting from infection with BVDV-2c isolates consisting of 3 genomic variants (dup+, dup1−, dup2−) has been reported in Germany (Jenckel et al. 2014). The recent association of HoBiPeV with severe respiratory and reproductive disease and mucosal disease in cattle and respiratory disease in small ruminants has raised concerns (Bauermann et al. 2013; Weber et al. 2015). Natural HoBiPeV infection in cattle has so far been reported sporadically in Brazil, Italy, Thailand, India and Bangladesh, but in some regions like in Northeastern Brazil, HoBiPeVs have been more frequently detected in cattle than BVDV-1 and BVDV-2. Natural infection has also been reported in buffalo in Brazil and in sheep and goats in China.
9.2 India
Serological evidence of BVDV infection in cattle in India was first reported in 1981 in Orissa State (Nayak et al. 1981) followed by a report from Gujarat State in 1989. A seroepidemiological study on sera collected from 17 states then demonstrated an overall apparent BVD seroprevalence rate of 17% in cattle in most parts of the country (Sudharsana et al. 1999). Subsequently, varying rates (up to 52%) of BVD seroprevalence in cattle and buffaloes in different parts of the country involving both commercial dairies and small holder units have been reported. Besides cattle and buffaloes, serological evidence of BVDV infection has been reported in sheep and goats (Mishra et al. 2009), in yaks (Mishra et al. 2008a), and also in mithun (Singh et al. 2017). A BVDV seroprevalence study involving sheep and goats from 13 states during 2004–2008 reported a true prevalence rate of 23.4% in sheep and 16.9% in goats and provided evidence of BVDV-1 infection predominantly and BVDV-2 occasionally (Mishra et al. 2009).
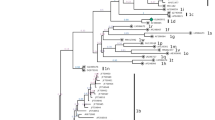
Although earlier studies indicated serological evidence of BVDV infection, the conclusive evidence of BVDV in Indian cattle was provided by virus isolation and subsequent phylogenetic analysis of BVDV isolates in 2004 (Mishra et al. 2004). The phylogenetic analysis of 13 BVDV isolates originating from cattle in eastern, northern, and western India in 5′-UTR and Npro genes revealed that they belong to BVDV-1b subtype and are closely related (Fig. 14.2). In the same year, BVDV-1 was also reported in lambs showing RVF-like symptoms (Yadav et al. 2004). Although systematic surveillance studies involving all the states are lacking, genetic typing of BVDV strains collected during later studies revealed that BVDV-1b is the predominant subtype circulating in cattle (Mishra et al. 2007b, 2014; Behera et al. 2011). Further studies have reported existence of BVDV-1b and BVDV-1c subtypes in buffaloes (Mishra et al. 2007b) and in sheep and goats (Mishra et al. 2012). Besides domestic ruminants, BVDV-1 was detected in yaks (Bos grunniens) in the Himalayan region for the first time and phylogenetic analysis revealed that they belonged to BVDV-1c subtype (Mishra et al. 2008a). It seems that BVDV has evolved well-developed strategies to become successful in replicating in different animal species.
Genetic typing of Indian cattle BVDV-1 isolates in 5′-UTR and Npro regions. The unrooted tree was based on partial sequence analysis of the 5′-UTR (245 nt) and the Npro (385 nt) and was prepared using the neighbor-joining method (Kimura 2-parameter method, transition/transversion 2.0). Sequences of BVDV isolates from India are labelled in bold and other sequences were taken from GenBank. The Npro sequences of strains 519, 721, Deer NZ1, and Deer GB1 have the following GenBank Acc. numbers: AF144464, AF144463, U80903 and U80902. Numbers over branches indicate the percentage of 1000 bootstrap replicates. (Reprinted from our work Mishra et al. 2004)
BVDV-2a was detected in cattle from Jammu & Kashmir State in 2011 and recently in bull semen from Tamil Nadu (Behera et al. 2011; Mishra et al. 2018). BVDV-2a has earlier been detected in goats from Northern India, while BVDV-2b has been reported in sheep from Western India (Mishra et al. 2007a, 2008b). Molecular epidemiology studies on BVDV-2 have provided evidence of circulation of genetically divergent BVDV-2a strains in Southern India and in Northern India (Fig. 14.3).
Genetic typing and relationship of Indian BVDV-2 strains originating from cattle, sheep and goats in the 5′-UTR and Npro regions. The phylogenetic tree was prepared based on 240 nt in the 5′-UTR and 474 nt in Npro gene using neighbor-joining method in MEGA version 6.0. Numbers in nodes indicate the percentage of 1000 bootstrap replicates that support each group. BVDV strains (cattle) labelled as filled triangles originated from Southern India, while BVDV-2 strains Ind 141353 (cattle), Ind 5197 (goat) and Ind 51966 (sheep) originated from Northern India. (Reprinted from our work Mishra et al. 2018)
Natural infection with HoBiPeV, an emerging bovine pestivirus, was identified recently in cattle in the states of Maharashtra, Punjab and Chhattisgarh while conducting systematic surveillance in cattle from 21 dairy farms across India. Molecular characterization of HoBiPeV strains revealed co-circulation of two novel and divergent lineages of HoBiPeV (c, d) in India (Fig. 14.4), highlighting the independent evolution of at least 3–4 lineages of HoBiPeV strains globally (Mishra et al. 2014). These novel findings extended the knowledge on the epidemiology and genetic diversity of HoBiPeV strains globally which is important for management and control of BVD.
Phylogenetic tree and genetic relationship of Hobi-like pestivirus (HoBiPeV) strains from Indian cattle with globally circulating HoBiPeV strains. The tree was based on the combined datasets of 5′-UTR (239 bp) and Npro (504 bp) sequences and the maximum likelihood tree was generated using concatenated datasets of 5′-UTRand Npro under the GTR + gamma substitution model in RAXML. Numbers indicate the percentage of 1000 bootstrap replicates that support each phylogenetic branch. The HoBiPeV isolates from India are labelled red, and previously reported Indian isolates of BVDV-1 and BVDV-2 are labelled blue. (Reprinted from our work Mishra et al. 2014)
10 Risk Factors
Several risk factors have been associated with BVDV infection which may vary between different geographical regions and cattle rearing practices. The presence of PI animals in the vicinity of susceptible animals is considered as the highest proven risk of BVDV infection and spread. Purchase of animals without BVDV testing is a high-risk factor in high prevalence regions than low prevalence regions. Some of the other risk factors include cattle on common pasture, sheep in pasture with cattle, over pasture fence contact, mixing of herds in pasture, wild animals in pasture, exchange of calves, large herd size, high cattle density, movement of animals pregnant with PI calves (Trojan animals), infection in contiguous farms, veterinarian reusing needles between farms, use of artificial insemination without testing semen for BVDV, use of contaminated live attenuated vaccine and livestock trade such as import of live cattle.
11 Host Range
BVDV has a wide host range and infects a variety of animals, both domesticated and wild. Among the domesticated ruminants, it infects cattle, buffalo, sheep, goats, and yaks, while among wild ruminants it infects buffalo, eland, Canadian bison, alpaca, pudu, bongo, deer, roe deer, mousedeer, reindeer, giraffe, European bison, chamois, pronghorn antelope and mithun. BVDV has been isolated in over 40 species of ruminants, and serological evidence indicates susceptibility of most free-ranging ruminants to BVDV infection (Vilcek and Nettleton 2006). Serological evidence of BVDV infection has also been reported in European rabbits. Although natural infection with BVDV occurs mainly in cattle, sheep and goats, it occurs also in pigs. Natural infection with BDV occurs mostly in sheep, but has also been reported in cattle and goats in many countries, whereas natural infection of CSFV and Bungowannah virus found in pigs has not been reported in cattle. However, natural or experimental BVDV persistent infection has been reported in mountain goats and domestic goats, domestic sheep, swine, alpaca, eland, mule deer, white-tailed deer and mouse deer and PI animals pose greatest risk of BVDV transmission.
12 Transmission
BVDV transmission occurs through several modes. BVDV spreads horizontally within a herd while vertical transmission occurs from cow to calf. PI animal results following infection of fetus with ncp BVDV in the first or second trimester (45–125 days) of pregnancy, before maturation of its immune system. The most common method of producing PI calves is through primary acute BVDV infections of pregnant cows. However, PI cows also invariably give birth to PI calves. Capability to cause fetal infections is exclusively a biotype-specific property of ncp BVDV. Horizontal transmission occurs not only by viremic transiently infected (TI) animals but also by PI animals that shed virus lifelong in all secretions, such as nasal and ocular discharges, milk/colostrum, semen, urine and feces (Van Campen and Frolich 2001). PI animals are the principal reservoirs of BVDV transmission, while TI animals transmit BVDV transiently and occasionally.
Semen from transiently infected bulls can transmit BVDV infection and virus can be detected up to 28 days in such bulls (Kirkland et al. 1994). In contrast, in both raw and extended semen of PI bulls, BVDV concentration remains high and semen from PI bulls infects susceptible animals consistently. Persistent testicular infection (PTI) occurs following acute BVDV infection in bulls. BVDV persists in semen or testicular tissue of these non-viremic and seropositive bulls, and seronegative cows may be infected via artificial insemination. Environmental conditions favoring crowding and aerosol transmission enhance the chances of BVDV transmission from acutely infected calves having respiratory form of BVDV infection. Transient shedding of vaccine strains in animals vaccinated with modified live BVD vaccine has been reported with probable consequences of secondary transmission to pregnant animals in contact with vaccinated animals (Fulton et al. 2003).
BVDV can be transmitted indirectly from contaminated pens, rectal examination gloves, hypodermic needles, nose tongs, BVDV-contaminated live vaccines, and ambient air (Niskanen and Lindberg 2003). Sufficient evidence exists regarding the spread of BVDV from domestic ruminants to wild ruminants. But there is no conclusive evidence that BVDV spreads from wild ruminants to domestic ruminants. However, serologic data from camels and roe deer strongly suggest circulation of BVDV in these animals independent of cattle, sheep and goats.
The four main factors which affect BVDV transmission are infectiousness (virulence) of the virus strain, the number of adequate contacts per time period between infectious and susceptible animals, the prevalence of infectious animals in a herd and the presence of truly susceptible animals.
13 Immunopathobiology
13.1 Pathogenesis
BVDV replicates in epithelial cells and lymphoid tissues of the oropharynx following infection via oronasal route. The phagocytic cells carry BVDV and/or BVDV-infected cells to peripheral lymphoid tissues leading to viremia that occurs 2–4 days after exposure and spreads BVDV to internal organs. The pathogenesis of BVDV is a complex interaction between the agent, host and environmental factors, and hence clinical signs are highly variable (Baker 1995). The virulence of BVDV strains and biotype of BVDV (ncp and cp) are the major determinants from the agent’s side, while immune status, immune competence and stage of pregnancy are the major determinants from host’s side.
In cultured cells infected by cp BVDV strains, there is rounding up and detachment of cells and cells die due to apoptosis, whereas in cells infected by ncp BVDV, no microscopically detectable alterations are seen. But both cp and ncp BVDV isolates induce apoptosis of T and B cells in vivo. Moreover, ncp viruses are preponderant in nature and pathogenic to the host and establish persistent infection upon fetal infection leading to lifelong virus shedding by PI animals. The host interactions with BVDV are highly variable, ranging from lack of immune response to a purifying immune response and from lack of clinical signs to highly lethal infection. Hence, disease is not obligatory for BVDV replication in PI animals or for viral transmission. Reproductive failure or immunosuppression are major consequences of BVDV infections because of its affinity for the fetus and for cells of lymphatic organs. Strong affinity of BVDV for lymphoreticular tissues causes necrosis in lymph nodes and spleen and destruction of Peyer’s patches. The four major syndromes associated with BVDV pathogenesis are acute infection, transplacental infection, persistent infection and mucosal disease.
13.2 Acute Infection
Acute BVDV infections in cattle develop when seronegative and immunocompetent cattle are infected with BVDV and the disease may be subclinical, severe acute, or chronic. The majority of postnatal BVDV infections are inapparent and virulence of BVDV strains is the key determinant of outcome of acute infection. Incidences of field cases of acute BVDV infection associated with thrombocytopenia, severe clinical signs and mortality in all age groups have been reported with death of about 40,000 animals, due to high virulent strains of BVDV-2 (Perdrizet et al. 1987; Carman et al. 1998). However, subsequent studies showed that BVDV-1 is able to induce hemorrhagic disease and all the three bovine pestiviruses, BVDV-1, BVDV-2 and HoBiPeV, encompass strains of high, moderate, and low virulence (Ridpath et al. 2000; Decaro et al. 2012; Mishra et al. 2014). Moreover, virulence is not correlated with the biotype, since all the three bovine pestiviruses have both ncp and cp strains. The differences in replication have a great impact on the virulence of BVDV strains, since strains that produce highest degree of viremia result in the most severe clinical symptoms (Walz et al. 2001). Acute BVDV infection also causes immunosuppression by depletion of both B and T lymphocytes thereby leading to suppression of immune functions in the infected animal. BVDV-induced immunosuppression not only directly causes enteritis but also predisposes calves to development of bovine respiratory disease (BRD) and secondary bacterial infections and enhances severity of bovine rota viral enteritis. Chronic viral shedding may also occur following acute infection, which has been reported in some bulls, where BVDV was found in semen for up to 7 months.
13.3 Transplacental Infection
The major economic impact of BVDV is due to its ability to cause intrauterine and transplacental infection in cattle, which may result from acute infection during or immediately before pregnancy and contaminated semen through artificial insemination or natural service. The outcome of BVDV fetal infection depends on the stage of development of the fetal immune system at the time of infection. Fetal infections during the first two trimesters of gestation may have severe reproductive consequences and may result in persistent infections, fetal death and abortion, or congenital anomalies (Brownlie et al. 1998). Dual infection of the fetus with both BVDV-1 and BVDV-2 has also been reported. Although abortions due to BVDV occur mostly in the early stages of gestation (< 125 days), abortions in the late phase of gestation do occur. Necrotizing inflammatory reaction with mononuclear cell infiltration in several tissues of fetus has been reported. The intrauterine infection of an immune-competent fetus in late gestation is similar to that of acute BVD. BVDV-induced congenital anomalies mostly occur during mid-gestation (80–150 days) and may involve the nervous system, eye, immune system, integumentary system, musculoskeletal system, or respiratory system.
13.4 Persistent Infection
Pestiviruses including BVDV use novel strategies to persist and spread in the host population through persistent infection. PI animals develop following infection of fetus with ncp BVDV strains during the first trimester of gestation, before the development of lymphoid tissues and functional immune responses and the immunotolerance is specific to the persisting BVDV strain (McClurkin et al. 1984). Primary acute BVDV infection of pregnant cows is the predominant cause of producing PI calves, although PI cows most often give birth to PI calves, whereas Trojan cow, a non-PI cow carrying a PI fetus, is immune to BVDV and possesses significantly higher antibody titers during mid-late pregnancy. The ability to cause persistent infection is exclusively a biotype-specific feature of ncp BVDV. Immunotolerance develops through selective evasion of innate immunity in the fetus by inhibition of IFN-I synthesis following ncp BVDV infection in addition to complete avoidance of the adaptive immune system. However, immune tolerance is BVDV strain specific and immune response is induced in PI animals following infections with other BVDV strains (Fulton et al. 2003). BVDV remains widely distributed in organs and secretions of PI animals and these animals may live for several years without any immune response to the persisting BVDV strain. Besides cattle, persistent infection occurs also in buffaloes, sheep, goats, alpacas, mouse deer, mountain goats and white-tailed deer.
13.5 Mucosal Disease
Mucosal disease (MD) is a sporadic disease of cattle, in which mostly 6–24-month-old animals succumb, but it may arise in adult animals. MD is highly fatal and is associated with the presence of closely related ncp and cp biotypes of BVDV in these animals. The first hypothesis on the mechanism of MD was that it develops only in PI animals. The second hypothesis was that animals suffering from MD always harbor a cp virus along with the persisting ncp strain (Bolin et al. 1985; McClurkin et al. 1985). Then it was established that close antigenic relationship between ncp and cp strains from the same animal is obligatory for development of MD (Meyers et al. 1991). Mucosal disease arises in PI animals by complex mutations of the ncp biotype to a cp biotype, or by superinfection of PI animals with a cp BVDV. In acute MD, genome analyses showed cp strain-specific genomic alterations which are mainly due to recombination events resulting in insertion of cellular sequences or duplications and deletions of viral sequences. The genomic alterations enhance NS3 protein production, which provides an apoptotic signal for the infected cells. Cp strains have been reported earlier for BVDV-1 and BVDV-2 and recently for HoBiPeV. Insertion of ubiquitin coding sequences upstream of the NS3 gene and insertion of host-cell origin Jiv sequences within NS2 upstream of the NS2/3 cleavage site are the most commonly observed genomic changes (Tautz et al. 1993; Becher and Tautz 2011).
In animals suffering MD, strikingly higher numbers of cp BVDV-infected cells have been found compared to ncp-infected cells in PIs before outbreak of MD. In chronic MD, the antigenic properties of the persisting ncp and the superinfecting exogenous cp viruses are more divergent, and such animals suffer clinically for a prolonged period. In some cases, MD can develop several weeks after superinfection and is known as late-onset MD, and cp viruses in them are recombinants between cp and ncp viruses, where structural genes are shared by ncp viruses and nonstructural genes are shared by cp viruses. Direct cell damage by cp BVDV is the major mechanism of disease in PI animals suffering from MD, while cp BVDV is mostly attenuated in acutely infected animals.
14 Clinical Signs
The manifestations of BVDV infections are complex and remain a challenge for the practitioners and researchers and BVD usually refers to acute infection in seronegative immunocompetent cattle. Most of the BVDV infections in adults are subclinical or mild. However, some strains produce severe disease with mortality. After 5–7 days of incubation period, there is fever, depression, inappetence, oculonasal discharge, and occasionally oral lesions such as erosions and shallow ulcerations (Baker 1987). In calves, respiratory and gastrointestinal symptoms occur, with occasional fatal enteritis. Growth is retarded and PI calves mostly die before weaning. BVDV produces venereal infections in bulls and semen from PI bulls is infective and failure of conception in cows occurs due to fertilization failure. Repeat breeding and more number of services per conception are common in BVDV-infected herds. Semen quality is reduced because of low motility and abnormal morphology of sperm cells. The prominent characteristics of mucosal disease are bloody diarrhea along with fever, anorexia, ataxia and general debility. Mortality is often 100% and within 15 days after onset of clinical signs (Baker 1987). During postmortem, extensive ulcerative lesions in the gastrointestinal tract, affecting especially the GALT in the mucosa, are observed.
15 Immunity and Immunosuppression
Immune responses to BVDV develop following vaccination, infection, exposure to cross-reactive pestiviruses, or by passively through colostrum. Antibody response to BVDV is detectable 2–3 weeks post-infection and may plateau after about 10–12 weeks and may persist for long. Passive antibodies protect neonatal calf from BVDV infection but interfere with vaccination. Structural proteins, E2 and Erns and nonstructural protein NS3 (P80) are immunodominant proteins and induce significant antibody responses following BVDV infection. The E2 glycoprotein elicits neutralizing antibodies and is the major determinant of protective immunity, whereas Erns and NS3 elicit non-neutralizing antibodies (Donis et al. 1988). BVDV-1 E2 protein has one immunodominant epitope, while BVDV 2 has three and virus neutralizing test (VNT) is used to correlate protective immunity. Cattle vaccinated with inactivated vaccines develop a weak NS3 antibody response, while a strong NS3 antibody response is elicited following natural infection or vaccination with modified live vaccine.
BVDV causes general inhibition of cellular immune responses in cattle. Mild (10–20% decrease) or severe lymphopenia (50–60% decrease) is found depending upon the virulence of the BVDV strain and cytotoxic T-lymphocytes (CD8+) are affected more than helper T-lymphocytes (CD4+) cells (Brodersen and Kelling 1999). BVDV affects bovine monocytes, dendritic cells and macrophages and may alter function of TLRs, expression of cytokines and costimulatory molecules in bovine monocytes and macrophages resulting in an adverse effect on their ability to stimulate Th cells.
Interactions of BVDV and immune system are complicated and variable. The ncp BVDV elicits humoral immune response faster and traffics to more immune organs of mucosal immunity. Besides, BVDV antigen from ncp strains persists longer in immune tissues than the cp strains. BVDV cp strains elicit higher CMI response, while ncp strains avoid production of CMI response. Elimination of the adaptive immune response via infection before self-non-self-discrimination and inhibition of innate immune response are perfect strategies adopted by ncp BVDV for generation of PI animals. To inhibit innate immune response, ncp BVDV employs several strategies including preventing IFN-1 induction through Npro and Erns and strictly controls their RNA replication. BVDV Erns and Npro are crucial in establishment and maintenance of persistent infections and inhibit the innate immune response (Meyers et al. 2007).
16 Diagnosis
The control of BVD is highly dependent on confirmed laboratory diagnosis that defines exposure of an individual animal or population to BVDV. The clinician should have a clear intention about the diagnosis approach and communicate it to the laboratory. Laboratory diagnosis of BVD is either aimed at detecting BVDV, viral antigen, viral RNA, or antibodies against BVDV. Dramatic improvements have been made in laboratory methods for diagnosing BVDV infections during the last 20 years. In spite of recent advances in BVDV diagnosis, virus isolation and identification still remain the gold standard technique. Like other diseases, diagnosis of BVDV is an art that involves accumulation of data (laboratory test results) and reasoning from the data (interpretation).
BVDV-free animals upon testing are negative for antibody, antigen and virus, while acutely infected animals or immunocompetent fetuses are antibody positive and generally antigen or virus negative. PI animals in contrast are positive for antigen or virus and negative for antibody. Since acutely infected animals become positive for BVDV antibodies within 2–3 weeks post-infection, testing for antibody 4–8 weeks after initial testing can distinguish between acute and persistent infection in animals with positive antigen ELISA or RT-PCR results. Confirmatory diagnosis of mucosal disease relies on confirmation of PI status followed by isolation of both cp and ncp BVDV from the affected animal. However, it can also be confirmed based on identification of PI and characteristic pathological lesions. It is of utmost importance that only well-validated diagnostic tests should be employed for providing confirmatory diagnosis of BVD.
16.1 Conventional Methods
16.1.1 Virus Isolation
Virus isolation (VI) is the gold standard test for BVDV diagnosis and is the OIE recommended test for certifying individual freedom of infection during international trade. VI relies on the growth of BVDV in specific cell lines, such as MDBK or BT (bovine turbinate) followed by detection using specific antibodies or molecular methods. The cells and serum used for VI should be BVDV free and BVDV antibody free. As most of the BVDV isolates are of ncp biotype, detection by immunostaining or immunofluorescence using Erns- or NS3-specific monoclonal antibodies is recommended. In case of cp (cytopathic) strains, a characteristic cytopathic effect is observed. VI detects viremia in individual animals and is used to confirm the PI status of animals with positive ear notches, serum, or whole blood buffy coats. Microplate immuno-peroxidase assay in 96-well plates is commonly used for PI animal detection. However, these methods suffer certain limitations such as varying sensitivity and slower test results and colostral antibodies can mask BVDV in PI animals and interfere their detection.
In case a pestivirus is isolated during virus isolation, it has to be characterized antigenically further for identification of the agent, using a panel of monoclonal antibodies specific for BVDV-1, BVDV-2, or BDV in a microplate immunoperoxidase method (Paton et al. 1995). The interpretation of results needs caution due to serological cross-reactivity among pestiviruses.
16.1.2 Antigen ELISA
Direct detection of viral antigen in leukocytes, serum, or ear notch samples can be done by pestivirus antigen capture ELISA (PACE), and several commercial BVDV antigen ELISA kits are available which can be used in most of the veterinary diagnostic laboratories. PACE is usually recommended for PI screening, but it also detects some transiently infected animals, so a follow-up sample 4–6 weeks later is tested to confirm PI status. The test is based on a sandwich principle using monoclonal antibodies and usually detects BVDV NS2-3 or Erns antigen in peripheral blood leukocytes, plasma, or serum. Skin biopsies or ear notches have become a popular sample for PI testing by PACE, since PI animals are consistently detected by PACE tests regardless of antibody status and the site of the biopsy. Although the test is easy to perform and rapid, it suffers from low sensitivity and specificity. Besides, false positives are not unusual. Erns mAb-based PACE is preferable since it can detect all the three species of bovine pestivirus, BVDV-1, BVDV-2 and HoBiPeV (Mishra et al. 2014). Moreover, antigen capture ELISAs based on the detection of the p80 (NS3) protein of BVDV have diagnostic gaps prior to the age of 90 days both for serum and ear notches, instability of the p80/NS3 protein and the stronger inhibitory effect of colostral antibodies.
16.1.3 Immunohistochemistry (IHC)
Immunohistochemistry using ear notch tissue samples detects PI animals with 100% sensitivity and hence is one of the popular methods of BVDV Ag detection. While IHC is considered robust, it has many disadvantages in that it is subjective and restricted to tissue samples and is labor intensive, requiring experienced staff and its unreliability for use on samples stored in formalin for >15 days.
16.1.4 BVDV-Specific Antibody Detection
Serological tests for BVDV can be used to determine previous exposure of animals to BVDV, colostral antibodies in calves and immune response in vaccinated animals and in confirmation of acute infection. However, antibody-negative animals should be further tested for BVDV or Ag to rule out PI status. High prevalence of antibodies is indicative of current infection at a herd or region level. Although several antibody detection methods such as AGID, dot ELISA, and microsphere-based immunoassay have been reported, the virus neutralization test (VNT) and Ab ELISA are most commonly used.
16.1.5 Virus Neutralization Test (VNT)
VNT is the gold standard test to detect anti-BVDV antibodies, although antibody ELISA can alternatively be used (OIE 2017). For demonstration of seroconversion, both acute and convalescent sera should simultaneously be tested. The test is based on determination of 50% neutralizing end point and both cytopathic and non-cytopathic strains of BVDV can be used. Most preferably local isolates of BVDV-1, BVDV-2 and HoBiPeV must be used as low levels of antibody to BVDV-1 may not be detectable by a VNT that uses only BVDV-2 and vice versa. A differential neutralization test against a BDV strain should also be carried out simultaneously for serological differential diagnosis of BVD as BDV can also infect cattle naturally and serological cross-reactivity occurs.
16.1.6 BVDV Antibody ELISA
BVD antibody ELISA is useful for screening large number of cattle herds for serological diagnosis of BVD in unvaccinated animals. Both mAb-based competition ELISA and indirect ELISA are available commercially and used for detection of BVDV antibodies in serum, milk and bulk milk. Being rapid and cost effective, Ab ELISA is an efficient and economical alternative to VNT. But in BVDV-vaccinated animals, the usefulness of ELISA tests is fairly limited, while in unvaccinated populations and in eradication phase, it has more utility. Moreover, VNT detects a rise in Abs following vaccination or infection, while Ab ELISAs fail in this regard. Caution should be taken in result interpretation, since some commercial BVDV antibody ELISA kits have been reported to yield false-negative results when serum samples of calves harboring HoBiPeV antibodies were tested.
16.2 Modern Methods
Newer technologies are constantly being developed and evaluated for their use in BVDV diagnostic testing through genome detection and/or amplification. Besides, nucleotide sequencing and sequence analysis have dramatically improved the molecular epidemiology of BVDV in detecting new and divergent strains and in tracing the origin of outbreaks.
16.2.1 RT-PCR
In BVDV-infected cattle, viral RNA is detectable early and for a longer duration than virus isolation. RT-PCR employing pooled serum and milk samples has been found useful in identifying PI animals during BVDV surveillance. RT-PCR assays are able to detect acutely infected animals, PI animals and animals vaccinated with modified live vaccines, but follow-up testing is necessary to define the status of positive animals. Several RT-PCR protocols have been developed and evaluated and are being used for BVDV diagnosis. A range of samples, including blood, milk, follicular fluid, saliva, and tissue samples, can be tested successfully by RT-PCR (Dubovi 2013). The most widely used protocol utilizes primers 324/326, which is targeted at the highly conserved 5′-UTR and is pestivirus specific, but it fails to detect the highly divergent HoBiPeV strains. However, HoBiPeV-specific RT-PCR has recently been reported. Differentiation between BVDV-1, BVDV-2 and BDV is also possible by nested RT-PCR. However, extreme precautions should be taken during nucleic acid-based tests due to false-positive cases arising from cross-contamination, and a positive RT-PCR does not define the clinical status of an animal in a single animal test or in a pooled sample. Although several modifications of nucleic acid detection methods, such as RT-PCR ELISA, microarray, and LAMP tests, have been reported, real-time RT-PCR and RT-PCR are most commonly used for BVDV diagnosis.
16.2.2 Real-Time RT-PCR
As real-time RT-PCR assay provides simultaneous quantitation and genotyping of BVDV, it is used more commonly now not only during BVD outbreaks but also for routine BVDV diagnosis, due to its rapidity in obtaining the results. Several real-time PCR assays, in uniplex, duplex, or multiplex formats, are available commercially for diagnosis and genetic typing of BVDV and differentiation from other pestiviruses using primers and probes targeted at 5′-UTR (Hoffmann et al. 2006; Baxi et al. 2006; Willoughby et al. 2006; Liu et al. 2008).
16.2.3 Sequencing and Next-Generation Sequencing
Genetic typing of BVDV provides useful information during BVD epidemiology and control. The more accurate genetic typing of BVDV strains is achieved from nucleotide sequencing or next-generation sequencing data followed by phylogenetic analysis. Although sequence analysis of 5′-UTR can be used for pestivirus species assignment, sequence analysis of complete Npro and E2 genes or combined datasets can classify them more accurately into genotypes/subtypes (Becher et al. 2003; Vilcek et al. 2001; Mishra et al. 2014). Recently, analyses of whole genome sequencing data of BVDV strains are being more frequently used.
17 Prevention and Control
Vaccination against BVDV has been commonly used to prevent BVDV infection by enhancing immunity in cattle populations with an aim to prevent or reduce clinical disease and prevent the spread of infection within a herd by reducing BVDV viremia and preventing fetal infection and generation of PI calf. At present, both modified live vaccine (MLV) and killed vaccines are used to control BVD. Vaccination may reduce the incidence of acute and persistent infections but may not prevent all infections in individual animals and vaccination failure is likely due to failure in broad protection arising from existence of three bovine pestivirus species (BVDV-1, BVDV-2 and HoBiPeV) and multiple subtypes. Currently most of the commercially available BVD vaccines contain antigens of BVDV-1a, BVDV-1b, BVDV-2a, or bivalent vaccines of different combinations, but no vaccine is available against HoBiPeV. Vaccination is only successful when a minimum coverage of the population is achieved with maximum number of non-susceptible animals. Although vaccination has been effective in field conditions, when used as a lone measure, it has not resulted in the elimination of BVDV-induced clinical disease or a significant reduction in BVDV losses (Ridpath 2013).
Although initially it was thought that control of BVDV infection is not possible, a range of well-planned control strategies have been designed over the years, with successful implementation of BVDV control programs in many European countries. The common strategies for control programs are identification and removal of PI animals, movement control of infected animals, strict farm biosecurity and surveillance. BVD control programs without use of vaccination have been used successfully in Scandinavian countries, Austria and Switzerland, whereas control with vaccination has been implemented in Germany, Belgium, Ireland, and Scotland (Moennig and Becher 2018). While compulsory and systematic control programs have been found most successful than voluntary control programs, there is a lack of official guidelines on BVD control in most of the countries.
18 Vaccines
18.1 Inactivated Vaccines
Since protection against homologous strains is better than heterologous viral strains, multivalent vaccines containing both BVDV-1 and BVDV-2 are better than monovalent vaccines. Development and use of BVD vaccine based on the predominant subtypes of BVDV circulating in a country has been advocated as the most viable option. BVDV-inactivated vaccine elicits primarily a humoral response targeted mainly at E2 glycoprotein with minimal cell-mediated response. As inactivated vaccines contain viral antigen(s) incapable of replication, the risk of adverse effects in vaccinates and the fetus in pregnant animals is minimal. Hence, it is safe and can be administered at any stage of gestation. But protective immunity is shorter, and booster doses are required for its efficacy, and neonatal pancytopenia, associated with the use of inactivated vaccines with powerful adjuvants, has been reported in several European countries. Fetal protection varies from incomplete to satisfactory. Despite inactivated vaccines are predominantly used, several countries allow the use of inactivated BVD vaccine before breeding, while some countries implement use of inactivated vaccine first followed by modified live BVD vaccine (two-step vaccination). Use of inactivated BVDV vaccine first and then vaccination with a live attenuated vaccine after 4 weeks have shown long-lasting immunity in vaccinates and in prevention of fetal infection following BVDV-1 and BVDV-2 challenge (Moennig et al. 2005).
18.2 Live Attenuated Vaccines
Live attenuated or modified live vaccine (MLV) against BVD was initially developed in the early 1960s, and more MLVs were produced and used subsequently. Besides generating antibody responses against E2, MLV vaccines are better inducers of CD4+ and CD8+ T-cells immune responses and provide a solid fetal protection (Reber et al. 2006). Although quite efficacious, the MLV vaccines have safety concerns due to possibility of reversion of virulence of attenuated virus, its ability to cause in utero infections, ovarian lesions leading to infertility in cows and mucosal disease, risk of contamination with adventitious viruses, and immunosuppressive effects. Hence, the MLVs are not recommended in the first 6 months of unvaccinated pregnant animals especially with ncp BVDV which may cross the placenta and infect the developing fetus. Most of the modern MLVs are prepared using cp BVDV vaccine strains. Vaccination with MLVs is not advised in calves younger than 6 weeks. To improve the safety of MLVs, a mutant ncp BVDV virus strain was developed with the deletion of Npro gene and inactivation of endoribonuclease activity of Erns and found to elicit immune response without crossing the placenta of pregnant cattle (Platt et al. 2017).
18.3 Recombinant and Vectored Vaccines
The protective efficacy of BVDV E2 antigen has been shown using various delivery platforms like viral vectors and DNA immunizations or as a recombinant protein produced in various expression systems. However, these vaccines are not available commercially either due to prohibitive cost or low protective efficacy. An adenovirus vectored subunit vaccine against BVDV, consisting of recombinant adenoviruses expressing three novel mosaic polypeptide chimeras targeting Npro, E2, and NS2-3 antigens has been reported recently and is promising (Lokhandwala et al. 2017). The prototype vaccine has been shown to induce higher BVDV-1-specific neutralizing antibody titers and lower clinical scores in calves following BVDV-2 challenge, compared to higher BVDV-2-specific neutralizing antibody titers found after MLV vaccination.
18.4 DIVA Vaccines
Although marker vaccines which allow differentiation of infected from vaccinated animals (DIVA) by serological tests have been successfully used in many other animal viral diseases, in the case of pestiviruses including BVDV, the benefit of a marker vaccine is questionable, because the PI animals, which are the main sources of infection, do not produce antibodies against the homologous strain. However, serological DIVA test may be useful in countries or regions where BVDV has been eradicated, and search for novel strategies should continue to find a solution.
19 Antivirals
There is no specific treatment for BVDV-infected animals currently, but several strategies have been tried in vitro and in vivo to identify the antivirals. Treatment of PI animal with DB772 (2-(2-benzimidazolyl)-5-[4-(2-imidazolino) phenyl] furan dihydrochloride) has shown decrease in the viral load of infected calves but it caused rapid selection of drug-resistant mutants (Newcomer et al. 2013). Similarly, iminosugar N-butyldeoxynojirimycin (NB-DNJ), an endoplasmic reticulum α-glucosidase inhibitor, has an antiviral effect against BVDV. Antiviral activity of bovine bovIFN-α and boIFN-τ against BVDV was tried but was unsuccessful, and boIFN-τ reduced the BVDV level in serum transiently when injected into PI cattle, but virus titer returned to the pre-administration level at the end of the treatment course. Essential oil of Ocimum basilicum (basil) and monoterpenes were reported to inhibit BVDV in an in vitro experiment.
20 Other Measures
20.1 Identification and Removal of PI Animals
PI animals act as BVDV reservoirs and permanently shed large amounts of infectious virus. Hence, identification and removal of PI animals are the hallmark of BVDV control. Virological screening using cost-effective BVDV antigen ELISAs and molecular tests have been found useful in identification and elimination of PI animals in successful control programs. All Scandinavian programs were successful following this method without vaccination, and the countries became mostly free from BVDV after a few years. Following success in Scandinavian countries, this approach was then implemented in Austria and Switzerland with promising outcomes.
20.2 Biosecurity
Strict farm biosecurity should be taken into account as a part of any BVD control program due to the vulnerability of susceptible herds to reinfection by PI animals. Purchase or trade with untested cattle is the predominant factor of BVDV introduction. Reports of high prevalence of BVDV-1 and BVDV-2 in sheep and BVDV transmission from sheep to cattle pose hindrances to the success of BVD control and eradication programs. Similarly wild ruminants also present a potential threat during grazing of cattle in certain geographical areas.
21 Conclusions
Molecular analyses during the last two decades have deciphered several astonishing features of BVDV and have added significantly on virus/host interplay, but many aspects of BVDV biology including the genetic basis of attenuation are still obscure and await further work at the molecular level. Similarly, problems associated with classification and nomenclature of pestiviruses including BVDV should be resolved soon and so also the subtype assignment criteria and consistency in phylogenetic analyses methods and target genes. Due to the recognition of severe acute BVDV infections earlier and with involvement of novel BVDV-2c strains recently, monitoring the role of newly emerging strains of BVDV on disease severity and on acute and persistent infections should be continued in the future, and acute BVDV infections with mucosal lesions should not be ignored. The genetic and antigenic heterogeneity of bovine pestiviruses, diverse host range, and clinical outcomes pose challenges for both laboratory diagnosis and clinical diagnosis. With regard to diagnosis, virus isolation should be carefully undertaken by eliminating cross-contamination from laboratory handling and identification of BVDV in unusual hosts should be reported with caution. Moreover, no uniform approach exists in selection of correct BVDV strains for use in serological studies, selection of antigen ELISA kits and the use of correct cells for virus isolation. Inconsistencies in selection of primers and probes to detect existing and new BVDV strains and other pestiviruses and development of new tests for detection and differentiation of all the three species of bovine pestiviruses have to be resolved in the near future. As currently available BVD vaccines, consisting of BVDV-1 or BVDV-2 strains, provide only limited protection against HoBiPeV strains, future strategies should aim at development of efficient vaccines having ability of broad protection and complete protection from fetal infection. Since contact of cattle with other domestic and wild ruminants can favor BVDV transmission, and there is a risk of introduction of HoBiPeV, surveillance strategies may be reviewed to ensure optimal performance of laboratory diagnostic tests for identification of PI animals.
References
Bachofen C, Stalder H, Braun U, Hilbe M, Ehrensperger F, Peterhans E (2008) Co-existence of genetically and antigenically diverse bovine viral diarrhoea viruses in an endemic situation. Vet Microbiol 131:93–102
Baker JC (1987) Bovine viral diarrhea virus: a review. J Am Vet Med Assoc 190:1449–1458
Baker JC (1995) The clinical manifestation of bovine viral doarrhoea infection. Vet Clin N Am Food Anim Pract 11:425–445
Bauermann FV, Ridpath JF, Weiblen R, Flores EF (2013) HoBi-like viruses: an emerging group of pestiviruses. J Vet Diagn Investig 25:6–15
Baxi M, McRae D, Baxi S, Greiser-Wilke I, Vilcek S, Amoako K, Deregt D (2006) A one-step multiplex real time RT-PCR for detection and typing of bovine viral diarrhoea viruses. Vet Microbiol 116:37–44
Becher P, Tautz N (2011) RNA recombination in pestiviruses: cellular RNA sequences in viral genomes highlight the role of host factors for viral persistence and lethal disease. RNA Biol 8:216–224
Becher P, Orlich M, Kosmidon A, Konig M, Baroth M, Thiel HJ (1999) Genetic diversity of pestiviruses: identification of novel groups and implication for classification. Virology 262:64–71
Becher P, Avalos Ramirez R, Orlich M, Cedillo Rosales S, Konig M, Schweizer M, Stalder H, Schirrmeier H, Thiel HJ (2003) Genetic and antigenic characterization of novel pestivirus genotypes: implications for classification. Virology 311:96–104
Behera SP, Mishra N, Vilcek S, Rajukumar K, Nema RK, Prakash A, Kalaiyarasu S, Dubey SC (2011) Genetic and antigenic characterization of bovine viral diarrhoea virus type 2 isolated from cattle in India. Comp Immunol Microbiol Infect Dis 34:189–196
Bolin SR, Ridpath JF (1992) Differences in virulence between two noncytopathic bovine viral diarrhea viruses in calves. Am J Vet Res 53:2157–2163
Bolin SR, McClurkin AW, Cutlip RC, Coria MF (1985) Severe clinical disease induced in cattle persistently infected with noncytopathogenic bovine viral diarrhea virus by superinfection with cytopathogenic bovine viral diarrhea virus. Am J Vet Res 46:573–576
Brodersen BW, Kelling CL (1999) Alteration of leukocyte populations in calves concurrently infected with bovine respiratory syncytial virus and bovine viral diarrhea virus. Viral Immunol 12:323–334
Brownlie J, Clarke MC, Howard CJ (1984) Experimental production of fatal mucosal disease in cattle. Vet Rec 114:535–536
Brownlie J, Hooper LB, Thompson I, Collins ME (1998) Maternal recognition of foetal infection with bovine viral diarrhea virus (BVDV) – the bovine pestivirus. Clin Diag Virol 10:141–150
Callens N, Brügger B, Bonnafous P, Drobecq H, Gerl MJ, Krey T, Roman-Sosa G, Rümenapf T, Lambert O, Dubuisson J, Rouillé Y (2016) Morphology and molecular composition of purified bovine viral diarrhea virus envelope. PLoS Pathog 12:e1005476. https://doi.org/10.1371/journal.ppat.1005476
Carman S, van Dreumel T, Ridpath J, Hazlett M, Alves D, Dubovi E, Tremblay R, Bolin S, Godkin A, Anderson N (1998) Severe acute bovine viral diarrhea in Ontario, 1993–1995. J Vet Diagn Investig 10:27–35
Decaro N, Mari V, Pinto P, Lucente MS, Sciarretta R, Cirone F (2012) Hobi-like pestivirus: both biotypes isolated from a diseased animal. J Gen Virol 93:1976–1983
Dias RK, Cargnelutti JF, Weber MN, Canal CW, Bauermann FV, Ridpath JF, Weiblen R, Flores EF (2017) Antigenic diversity of Brazilian isolates of HoBi-like pestiviruses. Vet Microbiol 203:221–228
Donis RO, Corapi W, Dubovi EJ (1988) Neutralizing monoclonal antibodies to bovine viral diarrhoea virus bind to the 56K to 58K glycoprotein. J Gen Virol 69:77–86
Dubovi EJ (2013) Laboratory diagnosis of bovine viral diarrhea virus. Biologicals 41:8–13
Fulton RW, Ridpath JF, Confer AW, Saliki JT, Burge LJ, Payton ME (2003) Bovine viral diarrhoea virus antigenic diversity: impact on disease and vaccination programmes. Biologicals 31:89–95
Giammarioli M, Ridpath JF, Rossi E, Bazzucchi M, Casciari C, De Mia GM (2015) Genetic detection and characterization of emerging HoBi-like viruses in archival foetal bovine serum batches. Biologicals 43:220–224
Grassmann CW, Yu H, Isken O, Behrens SE (2005) Hepatitis C virus and the related bovine viral diarrhea virus considerably differ in the functional organization of the 5′ non-translated region: implications for the viral life cycle. Virology 333:349–366
Grummer B, Grotha S, Greiser-Wilke I (2004) Bovine viral diarrhoea virus is internalized by clathrin-dependent receptor-mediated endocytosis. J Veterinary Med Ser B 51:427–432
Hoffmann B, Depner K, Schirrmeier H, Beer M (2006) A universal heterologous internal control system for duplex real-time RT-PCR assays used in a detection system for pestiviruses. J Virol Methods 136:200–209
Horzinek M, Maess J, Laufs R (1971) Studies on the substructure of togaviruses. Arch Gesamte Virusforsch 33:306–318
Houe H (1999) Epidemiological features and economical importance of bovine virus diarrhoea virus (BVDV) infections. Vet Microbiol 64:89–107
Houe H (2003) Economic impact of BVDV infection in dairies. Biologicals 31:137–143
Hulst MM, Moormann RJ (2001) Erns protein of pestiviruses. Methods Enzymol 342:431–440
Jenckel M, Hoper D, Schirrmeier H, Reimann I, Goller KV, Hoffmann B, Beer M (2014) Mixed triple: allied viruses in unique recent isolates of highly virulent type 2 bovine viral diarrhea virus detected by deep sequencing. J Virol 88:6983–6992
Kirkland PD, MacIntosh SG, Moyle A (1994) The outcome of widespread use of semen from a bull persistently infected with pestivirus. Vet Rec 135:527–529
Krey T, Thiel HJ, Rumenapf T (2005) Acid-resistant bovine pestivirus requires activation for pH-triggered fusion during entry. J Virol 79:4191–4200
Kummerer BM, Meyers G (2000) Correlation between point mutations in NS2 and the viability and cytopathogenicity of bovine viral diarrhea virus strain Oregon analyzed with an infectious cDNA clone. J Virol 74:390–400
Lackner T, Muller A, Pankraz A, Becher P, Thiel HJ, Gorbalenya AE (2004) Temporal modulation of an autoprotease is crucial for replication and pathogenicity of an RNA virus. J Virol 8:10765–10775
Lee KM, Gillespie JH (1957) Propagation of virus diarrhea virus of cattle in tissue culture. Am J Vet Res 18:952–953
Liang D, Sainz IF, Ansari IH, Gill LH, Vassilev V, Donis RO (2003) The envelope glycoprotein E2 is a determinant of cell culture tropism in ruminant pestiviruses. J Gen Virol 84:1269–1274
Lindenbach BD, Murray CL, Thiel HJ, Rice CM (2013) Flaviviridae. In: Knipe DM, Howley PM (eds) Fields virology, vol 6. Lippincott Williams & Wilkins, Philadelphia, pp 712–746
Liu L, Xia H, Belak S, Baule C (2008) A TaqMan real-time RT-PCR assay for selective detection of atypical bovine pestiviruses in clinical samples and biological products. J Virol Methods 154:82–85
Liu L, Xia H, Wahlberg N, Belák S, Baule C (2009) Phylogeny, classification and evolutionary insights into pestiviruses. Virology 385:351–357
Lokhandwala S, Fang X, Waghela SD, Bray J, Njongmeta LM, Herring A (2017) Priming cross-protective bovine viral diarrhea virus-specific immunity using live-vectored mosaic antigens. PLoS One 12(1):e0170425
Malmquist WA (1968) Bovine viral diarrhea-mucosal disease: etiology, pathogenesis and applied immunity. J Am Vet Med Assoc 152:763–768
Mathapati BS, Mishra N, Rajukumar K, Nema RK, Behera SP, Dubey SC (2010) Entry of bovine viral diarrhoea virus into ovine cells occurs through clathrin-dependent endocytosis and low pH-dependent fusion. In Vitro Cell Dev Biol Anim 46:403–407
Maurer K, Krey T, Moennig V, Thiel HJ, Rumenapf T (2004) CD46 is a cellular receptor for bovine viral diarrhea virus. J Virol 78:1792–1799
McClurkin AW, Littledike ET, Cutlip RC, Frank GH, Coria MF, Bolin SR (1984) Production of cattle immunotolerant to bovine viral diarrhea virus. Can J Comp Med 48:156–161
McClurkin AW, Bolin SR, Coria MF (1985) Isolation of cytopathic and noncytopathic bovine viral diarrhea virus from the spleen of cattle acutely and chronically affected with bovine viral diarrhea. J Am Vet Med Assoc 186:568–569
Meyers G, Thiel HJ (1996) Molecular characterization of pestiviruses. Adv Virus Res 47:53–118
Meyers G, Tautz N, Dubovi EJ, Thiel HJ (1991) Viral cytopathogenicity correlated with integration of ubiquitin-coding sequences. Virology 180:602–616
Meyers G, Ege A, Fetzer C, von Freyburg M, Elbers K, Carr V, Prentice H, Charleston B, Schurmann EM (2007) Bovine viral diarrhoea virus: prevention of persistent foetal infection by a combination of two mutations affecting the Erns RNase and the Npro protease. J Virol 81:3327–3338
Mishra N, Pattnaik B, Vilcek S, Patil SS, Jain P, Swamy N, Bhatia S, Pradhan HK (2004) Genetic typing of bovine viral diarrhoea virus isolates from India. Vet Microbiol 104:207–212
Mishra N, Dubey R, Rajukumar K, Tosh C, Tiwari A, Pitale SS, Pradhan HK (2007a) Genetic and antigenic characterization of bovine viral diarrhea virus type 2 isolated from Indian goats (Capra hircus). Vet Microbiol 124:340–347
Mishra N, Dubey R, Galav V, Tosh C, Rajukumar K, Pitale SS, Pradhan HK (2007b) Identification of bovine viral diarrhea virus 1 in Indian buffaloes and their genetic relationship with cattle strains in 5’ UTR. Curr Sci 93:97–100
Mishra N, Vilcek S, Rajukumar K, Dubey R, Tiwari A, Galav V, Pradhan HK (2008a) Identification of bovine viral diarrhea virus type 1 in yaks (Bos poepaghus grunniens) in Himalayan region. Res Vet Sci 84:507–510
Mishra N, Rajukumar K, Vilcek S, Tiwari A, Satav JS, Dubey SC (2008b) Molecular characterization of bovine viral diarrhea virus type 2 isolate originating from a native Indian sheep (Ovies aries). Vet Microbiol 130:88–98
Mishra N, Rajukumar K, Tiwari A, Nema RK, Behera SP, Satav JS, Dubey SC (2009) Prevalence of bovine viral diarrhoea virus antibodies among sheep and goats in India. Trop Anim Health Prod 41:1231–1239
Mishra N, Mathapati BS, Rajukumar K, Nema RK, Behera SP, Dubey SC (2010) Molecular characterization of RNA and protein synthesis during a one-step growth curve of bovine viral diarrhoea virus in ovine (SFT-R) cells. Res Vet Sci 89:130–132
Mishra N, Rajukumar K, Kalaiyarasu S, Dubey SC (2011) Pestivirus infection, an emerging threat to ruminants in India: a review. Indian J Anim Sci 81:545–551
Mishra N, Pitale SS, Rajukumar K, Prakash A, Behera SP, Nema RK, Dubey SC (2012) Genetic variety of bovine viral diarrhoea virus 1 strains isolated from sheep and goats in India. Acta Virol 56:209–215
Mishra N, Rajukumar K, Pateriya A, Kumar M, Dubey P, Behera SP, Verma A, Bhardwaj P, Kulkarni DD, Vijaykrishna D, Reddy ND (2014) Identification and molecular characterization of novel and divergent HoBi-like pestiviruses from naturally infected cattle in India. Vet Microbiol 174:239–246
Mishra N, Kalaiyarasu S, Mallinath KC, Rajukumar K, Khetan RK, Gautam S, Venkatesha MD, Byregowda SM (2018) Identification of bovine viral diarrhoea virus type 2 (BVDV-2) in cattle bull semen from southern India and its genetic characterization. Curr Sci 114:666–670
Moennig V, Becher P (2018) Control of bovine viral diarrhea. Pathogens 7:29
Moennig V, Houe H, Lindberg A (2005) BVD control in Europe: current status and perspectives. Anim Health Res Rev 6:63–74
Nagai M, Hayashi M, Itou M, Fukutomi T, Akashi H, Kida H, Sakoda Y (2008) Identification of new genetic subtypes of bovine viral diarrhea virus genotype 1 isolated in Japan. Virus Genes 36:135–139
Nayak BC, Panda SN, Misra DB, Kar BC, Das BC (1981) Note on serological evidence of viral abortion in cattle in Orissa. Indian J Anim Sci 52:102–103
Newcomer BW, Neill JD, Marley MS, Ridpath JF, Givens MD (2013) Mutations induced in the NS5B gene of bovine viral diarrhea virus by antiviral treatment convey resistance to the compound. Virus Res 174:95–100
Niskanen R, Lindberg A (2003) Transmission of bovine viral diarrhoea virus by unhygienic vaccination procedures, ambient air, and from contaminated pens. Vet J 165:125–130
OIE (2017) Manual of diagnostic tests and vaccines for terrestrial animals. Chapter 2.4.7, Bovine viral diarrhea. OIE, Paris, pp 1–22
Olafson P, McCallum AD, Fox FH (1946) An apparently new transmissible disease of cattle. Cornell Vet 36:205–213
Paton DJ, Sands JJ, Lowings JP, Smith JE, Ibata G, Edwards S (1995) A proposed division of the pestivirus genus using monoclonal antibodies, supported by cross-neutralization assays and genetic sequencing. Vet Res 26:92–109
Pellerin C, Van den Hurk J, Lecomte J, Tijssen P (1994) Identification of a new group of bovine viral diarrhea virus strains associated with severe outbreaks and high mortality. Virology 203:260–268
Perdrizet JA, Rebhun WC, Dubovi EJ, Donis RO (1987) Bovine virus diarrhea-clinical syndromes in dairy herds. Cornell Vet 77:46–74
Pestova TV, de Breyne S, Pisarev AV, Abaeva IS, Hellen CU (2008) eIF2-dependent and eIF2-independent modes of initiation on the CSFV IRES: a common role of domain II. EMBO J 27:1060–1072
Pizarro-Lucero J, Celedon MO, Aguilera M, de Calisto A (2006) Molecular characterization of pestiviruses isolated from bovines in Chile. Vet Microbiol 115:208–217
Platt R, Kesl L, Guidarini C, Wang C, Roth JA (2017) Comparison of humoral and T-cell-mediated immune responses to a single dose of Bovela live double deleted BVDV vaccine or to a field BVDV strain. Vet Immunol Immunopathol 187:20–27
Poole TL, Wang CY, Popp RA, Potgieter LND, Siddiqui A, Collett MS (1995) Pestivirus translation initiation occurs by internal ribosome entry. Virology 206:750–754
Ramsey FK, Chivers WH (1953) Mucosal disease of cattle. North Am Vet 34:629–633
Rawlings ND, Barrett AJ, Bateman A (2012) MEROPS: The database of proteolytic enzymes, their substrates and inhibitors. Nucleic Acids Res 40:D343–D350
Reber AJ, Tanner M, Okinaga T, Woolums AR, Williams S, Ensley DT, Hurley DJ (2006) Evaluation of multiple immune parameters after vaccination with modified live or killed bovine viral diarrhea virus vaccines. Comp Immunol Microbiol Infect Dis 29:61–77
Richter V, Lebi K, Baumgartner W, Obritzhauser W, Käsbohrer A, Pinior B (2017) A systematic worldwide review of the direct monetary losses in cattle due to bovine viral diarrhoea virus infection. Vet J 220:80–87
Ridpath JF (2013) Immunology of BVDV vaccines. Biologicals 41:14–19
Ridpath J, Bolin SR, Dubovi EJ (1994) Segregation of bovine viral diarrhea virus into genotypes. Virology 205:66–74
Ridpath JF, Neill JD, Frey M, Landgraf JG (2000) Phylogenetic, antigenic and clinical characterization of type 2 BVDV from North America. Vet Microbiol 77:145–155
Ridpath JF, Fulton RW, Kirkland PD, Neill JD (2010) Prevalence and antigenic differences observed between bovine viral diarrhea virus subgenotypes isolated from cattle in Australia and feedlots in the southwestern United States. J Vet Diagn Investig 22:184–191
Ronecker S, Zimmer G, Herrler G, Greiser-Wilke I, Grummer B (2008) Formation of bovine viral diarrhea virus E1-E2 heterodimers is essential for virus entry and depends on charged residues in the transmembrane domains. J Gen Virol 89:2114–2121
Rumenapf T, Unger G, Strauss JH, Thiel HJ (1993) Processing of the envelope glycoproteins of pestiviruses. J Virol 67:3288–3294
Schirrmeier H, Strebelow G, Depner K, Hoffmann B, Beer M (2004) Genetic and antigenic characterization of an atypical pestivirus isolate, a putative member of a novel pestivirus species. J Gen Virol 85:3647–3652
Schmeiser S, Mast J, Thiel HJ, Konig M (2014) Morphogenesis of pestiviruses: new insights from ultrastructural studies of strain Giraffe-1. J Virol 88:2717–2724
Simmonds P, Becher P, Collett MS, Gould EA, Heinz FX, Meyers G (2012) Flaviviridae In: King AMQ, Lefkowitz E, Adams MJ, Carstens EB, Fauquet CM (eds) Ninth report of the international committee on taxonomy of viruses. Academic Press, San Diego, pp 1003–1020
Singh V, Mishra N, Kalaiyarasu S, Khetan RK, Hemadri D, Singh RK, Rajukumar K, Chamuah J, Suresh KP, Patil SS, Singh VP (2017) First report on serological evidence of bovine viral diarrhea virus (BVDV) infection in farmed and free ranging mithuns (Bos frontalis). Trop Anim Health Prod 49:1149–1156
Smith DB, Meyers G, Bukh J, Gould EA, Monath T, Muerhoff AS, Pletnev A, Rico-Hesse R, Stapleton JT, Simmonds P, Becher P (2017) Proposed revision to the taxonomy of the genus Pestivirus, family Flaviviridae. J Gen Virol 98:2106–2112
Stark R, Meyers G, Rumenapf T, Thiel HJ (1993) Processing of pestivirus polyprotein: cleavage site between autoprotease and nucleocapsid protein of classical swine fever virus. Virol 67:7088–7095
Sudharsana KJ, Suresh KB, Rajasekhar M (1999) Prevalence of bovine viral diarrhea virus antibodies in India. Rev Sci Tech 18:667–671
Tautz N, Meyers G, Thiel HJ (1993) Processing of poly-ubiquitin in the polyprotein of an RNA virus. Virology 197:74–85
Tautz N, Meyers G, Stark R, Dubovi EJ, Thiel HJ (1996) Cytopathogenicity of a pestivirus correlated with a 27 nucleotide insertion. J Virol 70:7851–7858
Underdahl NR, Grace OD, Hoerlein AB (1957) Cultivation in tissue-culture of cytopathogenic agent from bovine mucosal disease. Proc Soc Exp Biol Med 94:795–797
Van Campen H, Frolich K (2001) Pestivirus infections In: Williams ES, Barker IK (eds) Infectious diseases of wild mammals. Iowa State University Press, Iowa City, pp 232–244
Vilcek S, Nettleton PF (2006) Pestiviruses in wild animals. Vet Microbiol 116:1–12
Vilcek S, Paton DJ, Durkovic B, Strojny L, Ibata G, Moussa A, Loitsch A, Rossmanith W, Vega S, Scicluna MT, Palfi V (2001) Bovine viral diarrhea virus genotype 1 can be separated into at least eleven genetic groups. Arch Virol 146:99–115
Vilcek S, Durkovic B, Kolesarova M, Greiser-Wilke I, Paton DJ (2004) Genetic diversity of international bovine viral diarrhoea virus (BVDV) isolates: identification of a new BVDV-1 genetic group. Vet Res 35:609–615
Vilček Š, Ridpath JF, Van Campen H, Cavender JL, Warg J (2005) Characterization of a novel pestivirus originating from a pronghorn antelope. Virus Res 108:187–193
Walz PH, Bell TG, Wells JL, Grooms DL, Kaiser L, Maes RK, Baker JC (2001) Relationship between degree of viremia and disease manifestation in calves with experimentally induced bovine viral diarrhea virus infection. Am J Vet Res 62:1095–1103
Warrilow D, Lott WB, Greive S, Gowans EJ (2000) Properties of the bovine viral diarrhoea virus replicase in extracts of infected MDBK cells. Arch Virol 145:2163–2171
Weber MN, Streck AF, Silveira S, Mósena AC, Silva MS, Canal CW (2015) Homologous recombination in pestiviruses: identification of three putative novel events between different subtypes/genogroups. Infect Genet Evol 30:219–224
Weiland E, Stark R, Haas B, Rumenapf T, Meyers G, Thiel HJ (1990) Pestivirus glycoprotein which induces neutralizing antibodies forms part of a disulfide linked heterodimer. J Virol 64:3563–3569
Willoughby K, Valdazo-Gonzalez B, Maley M, Gilray J, Nettleton PF (2006) Development of a real time RT-PCR to detect and type ovine pestiviruses. J Virol Methods 132:187–194
Yadav P, Barde PV, Jadi R, Gokhale MD, Basu A, Joshi MV, Mehla R, Kumar SR, Athavale SS, Mourya DT (2004) Isolation of bovine viral diarrhea virus 1, a pestivirus from autopsied lamb specimen from Tamil Nadu, India. Acta Virol 48:223–227
Yarnall MJ, Thrusfield MV (2017) Engaging veterinarians and farmers in eradicating bovine viral diarrhoea: a systematic review of economic impact. Vet Rec 181:347. https://doi.org/10.1136/vr.104370
Yesilbag K, Forster C, Bank-Wolf B, Yilmaz Z, Alkan F, Ozkul FA, Burgu I, Rosales SC, Theil HJ, Konig M (2008) Genetic heterogeneity of bovine viral diarrhoea virus (BVDV) isolates from Turkey: identification of a new subgroup in BVDV-1. Vet Microbiol 130:258–267
Yesilbag K, Forster C, Ozyigit MO, Alpay G, Tuncer P, Theil HJ, Konig M (2014) Characterisation of bovine viral diarrhoea virus (BVDV) isolates from an outbreak with haemorrhagic enteritis and severe pneumonia. Vet Microbiol 169:42–49
Yesilbag K, Alpay G, Becher P (2017) Variability and global distribution of subgenotypes of bovine viral diarrhea virus. Viruses 9:128. https://doi.org/10.3390/v9060128
Zogg T, Sponring M, Schindler S, Koll M, Schneider R, Brandstetter H, Auer B (2013) Crystal structures of the viral protease Npro imply distinct roles for the catalytic water in catalysis. Structure 21:929–938
Acknowledgments
All the authors of the manuscript thank and acknowledge their respective institutes.
Conflict of Interest
There is no conflict of interest.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Mishra, N., Kalaiyarasu, S. (2019). Bovine Viral Diarrhea Virus. In: Malik, Y., Singh, R., Yadav, M. (eds) Recent Advances in Animal Virology. Springer, Singapore. https://doi.org/10.1007/978-981-13-9073-9_14
Download citation
DOI: https://doi.org/10.1007/978-981-13-9073-9_14
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-9072-2
Online ISBN: 978-981-13-9073-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)