Abstract
Plants are exposed to a plethora of microorganisms in their environment. A number of these microorganisms are plant pathogens. In order to defend themselves against pathogen attack, plants have evolved specialized sensory receptors to recognize some of the conserved molecular features (PAMPs, DAMPs, HAMPs, and NAMPs) as well as secreted effector molecules of pathogens. A cascade of signal transduction events are triggered which causes transcriptional rewiring leading to activation of defense responses. Closure of stomata, strengthening of cell wall along with accumulation of secondary metabolites, and induction of a hypersensitive response (HR) and pathogenesis-related (PR) proteins are some of the key defense strategies of the host. Interestingly, through secretion of volatile organic compounds (VOCs), plants have the ability to induce defense responses in uninfected tissues as well as surrounding plants. In this chapter, we elaborate on the mechanisms by which plants perceive pathogen attack and transduce the signal to downstream signaling molecules, culminating in the activation of defense responses.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Defense hormones
- Effector-triggered immunity
- Pathogen perception
- Pathogen-triggered immunity
- Plant defense responses
- Resistance genes
- Secondary messengers
1 Introduction
Plants are constantly exposed to a diverse array of microorganisms. Among them some are pathogenic on the host, whereas others grow in harmony with the host without causing any damage. Plants possess a proficient and dynamic sensory system to distinguish between them. In case of beneficial interactions, plants have adapted to harbor these microorganisms in specialized compartments, thus maintaining a suitable niche inside their tissue (Oldroyd 2013; Jones et al. 2007). However, in case of a negative interaction, the microorganism tries to forcefully colonize to obtain nutrients from the host plant. Plants being sessile cannot evade from such harmful interactions but possess several robust defense mechanisms to inhibit the growth of such pathogenic organisms.
The first step in mounting an immune response lies in the ability of host to perceive the pathogen attack, and this is achieved via a wide array of specialized extracellular receptors that are present on the plant cell membrane. Generally, plants recognize bacterial pathogens by conserved structural components such as flagellin, lipopolysaccharides (LPS), peptidoglycans (PG), etc. or bacterial molecules such as EfTu or RaxX that are released into the extracellular milieu (Couto and Zipfel 2016). Fungal pathogens are sensed by the recognition of chitin or fungal secreted proteins such as NLPs (NEP1 like proteins) (Kaku et al. 2006). These conserved microbe-specific molecules are known as PAMPs/MAMPs (pathogen-/microbe-associated molecular patterns). Herbivory is perceived by the presence of certain herbivore-associated molecular patterns (HAMPs) present in the oral secretion of the insect at the time of attack (Mithofer and Boland 2008). Nematodes also secrete molecules that are known to elicit plant defense responses, and these molecules are known as nematode-associated molecular patterns (NAMPs) (Mendy et al. 2017). Besides these signals, plants can also sense molecules that are released from their own cells as a consequence of pathogen attack and use them as cues to mount an immune response (Bacete et al. 2018). These molecules are known as DAMPs (damage-associated molecular patterns). Classic examples of DAMPs are degradation products that are released following the action of microbial enzymes on various components of the plant cell wall. Also, plants have cytoplasmic receptors to sense effector molecules secreted by potential pathogens to mount a robust immune response (Schreiber et al. 2016).
Plants possess a two-tiered detection system against pathogens (Zipfel 2014). The first tier comprises of receptors present on the surface of a cell called PRRs (pattern recognition receptors) that recognizes PAMPs, DAMPs, HAMPs or NAMPs. The PRRs can broadly be classified under two types, receptor-like kinases (RLKs; comprising of a ligand-binding ectodomain, a transmembrane domain, and a cytoplasmic kinase domain) and receptor-like proteins (RLPs; comprising of a ligand-binding ectodomain and a transmembrane domain). The immune responses that are mounted upon recognition of the pathogen by PRRs are referred to as pathogen-triggered immunity (PTI). Moreover, the immune responses that are induced following recognition of DAMPs are known as DAMP-triggered immunity (DTI). The second tier of the pathogen recognition system comprises of intracellular immune receptors that can sense secreted pathogenic effectors either directly or indirectly. The immune responses that are mounted upon recognition of these effectors are referred to as effector-triggered immunity (ETI). The receptors are classified into two types: nucleotide-binding site leucine-rich repeat (NBS-LRR) proteins and Toll-like receptor (TLR) proteins. The major difference in the signaling events during PTI and ETI is the duration and amplitude of the defense response, which is more in ETI as compared to PTI. In this chapter, we elaborate on how plants recognize various phytopathogens (bacteria and fungi) as well as herbivores and nematodes. We have described various players involved in the signal transduction events associated with pathogen perception and how the perceived signal is transduced to regulate host defense response pathways in host.
2 Perception of Pathogen Attack
Perception of danger is a key step in the activation of immune responses. However, induction of immune responses is an energy-consuming process that involves activation/deactivation of many molecular pathways, synthesis of new molecules, and alterations in basic metabolic processes (Andolfo and Ercolano 2015; Duan et al. 2013). Hence, it is crucial for plants to distinguish between a potential pathogen/pest and a random visitor to mount an appropriate immune response (Table 20.1).
2.1 Recognition of Bacterial Pathogens
Plants can recognize various structural components of bacteria or their secreted compounds to mount an immune response. Flagellin-Sensing 2 (FLS2), a LRR repeat domain-containing receptor-like kinase in Arabidopsis can recognize a 22-amino acid long peptide named flg22 derived from the flagellin of Pseudomonas syringae (Gómez-Gómez and Boller 2000). The flg22 peptide binds to the extracellular N-terminal domain of FLS2 and acts as a molecular glue between FLS2 and its co-receptor somatic embryogenesis receptor kinase 3 (SERK3) [also called as BAK1 (BRI1-associated receptor kinase 1)] (Meindl 2000; Sun et al. 2013). This complex phosphorylates downstream interacting partners and activates the immune response (Couto and Zipfel 2016). Interestingly, different plant species have evolved diverse receptors to recognize different epitopes on flagellin. Solanaceous plants such as pepper, potato and tomato recognize flgII-28 (the flagellin peptide derived from Pseudomonas syringae) by another type of LRR receptor, FLS3 (Hind et al. 2016).
Plants can also recognize peptidoglycan (PG) and lipopolysaccharide (LPS) that are major components of either bacterial cell wall or the outer membrane, respectively. Exogenous treatment with either LPS or PG activates plant immune responses (Erbs et al. 2010; Gust et al. 2007). PG is a polymer of N-acetylglucosamine and N-acetylmuramic acid linked by oligopeptides (Gust et al. 2007). Plants possess LysM domain (lysin motif)-containing proteins that can recognize glycans in N-acetylglucosamine (Gust et al. 2012). In Arabidopsis, PG is recognized by receptor-like proteins AtLYM1 and AtLYM3 where chitin receptor AtCERK1 serves as a key component in PG recognition (Willmann et al. 2011). In rice, OsLYP4 and OsLYP6 are known to interact with both chitin oligomers as well as peptidoglycan (Liu et al. 2012). Additionally, OsCERK1 appears to be a key receptor/co-receptor for LPS perception in rice (Desaki et al. 2017). In Arabidopsis, bulb-type (B-type) lectin S-domain (SD)-1 containing RLK protein LORE (lipooligosaccharide-specific reduced elicitation) is thought to be the putative LPS receptor (Ranf et al. 2015). However, the physical interaction between LPS and putative receptors are yet to be established.
Plants can also sense various bacterial secreted proteins/peptides. Elongation factor Tu (Ef-Tu) is an abundant bacterial protein that is released upon cell lysis. Ef-Tu acts as an elicitor of immune responses in various plant species (Kunze 2004). Members of the Brassicaceae family recognize a conserved 18-aa long peptide (elf18) present at the N-terminal of EF-Tu by the LRR-RLK Ef-Tu receptor (EFR) (Zipfel et al. 2006). Rice recognizes EFa50, comprising of a 50aa long peptide sequence from the middle of Ef-Tu amino acid sequence (Furukawa et al. 2014). Another secreted peptide recognized by plants is RexX21-sY, a sulfated peptide secreted by Xanthomonas oryzae pv. oryzae (Xoo) type 1 secretion system (Pruitt et al. 2015). This is recognized by rice LRR-RLK receptor Xa21. Here it is worth mentioning that Xa21 has been widely used to breed rice for bacterial blight resistance (Williams et al. 1996).
2.2 Recognition of Fungal/Oomycete Pathogens
Chitin, a polymer of N-acetyl-D-glucosamine (GlcNAc), is a major component of fungal cell walls. Plants can identify chitin oligomers by 40aa long globular LysM motif-containing receptor proteins (Kaku et al. 2006; Miya et al. 2007; Wan et al. 2008). In Arabidopsis, AtCERK1/AtLYK1 (chitin elicitor receptor kinase) recognizes chitin oligomers and mounts defense responses. Binding of 7–8-residue long chitin oligomer with AtCERK1 causes receptor homodimerization and transphosphorylation that lead to activation of defense signaling cascade (Liu et al. 2012). Rice recognizes chitin by a GPI-anchored RLP protein, OsCEBiP (chitin elicitor binding protein), that contains three extracellular LysM domains but lacks an intracellular kinase domain (Kaku et al. 2006; Kouzai et al. 2014). Ligand (GlcNAc)8 binding causes homodimerization of OsCEBiP and OsCERK1 leading to the formation of a GlcNAc8-2CEBiP-2CERK1 complex which in turn activates immune responses (Hayafune et al. 2014). Other plant receptors such as AtLYK4 (RLK), OsLYP4 and OsLYP6 (both RLP) can also recognize chitin (Liu et al. 2012; Petutschnig et al. 2010; Wan et al. 2012).
Some plants can sense presence of fungal xylanases to mount immune responses. A fungal protein ethylene-inducing xylanase (EIX) was found to activate plant immune responses in various host species (Bailey et al. 1990, 1993; Fuchs et al. 1989; Ron et al. 2000). In tomato, LeEIX is recognized by LRR-RLP LeEix2 leading to activation of immune responses (Bar and Avni 2009; Bar et al. 2009). Similarly, in Arabidopsis, LRR-RLP receptor AtRLP42 recognizes fungal endo-polygalacturonases (PGs) and activates its immune responses (Zhang et al. 2014).
2.3 Recognition of Herbivores
The plants are exposed to different insects, some of which feed upon plant parts by a process known as insect herbivory. Herbivorous insects can activate plant defense mechanisms either through mechanical wounding caused during the process of chewing or by their oral secretions. Mechanical wounding caused during herbivory induces either the activation of defense mechanisms or secretion of plant volatiles. Production of chemical factors or relaying of electrical signals across distal parts of the host tissues are some of the early plant responses generated immediately after wounding (Maffei et al. 2007).
The herbivore-associated molecular patterns (HAMPs) that are present in the oral secretions of insects are recognized by plants (Mithofer and Boland 2008). Some orally secreted compounds like fatty acid amino conjugates (FACs) act as elicitors in priming of plant defense responses (Bonaventure et al. 2011). Perception of FACs induces a MAPK signaling cascade including SIPK (salicylic acid-induced protein kinase) and WIPK (wound-induced protein kinase) along with activation of NPR1 signaling (Wu et al. 2007; Seo et al. 2007; Bonaventure and Baldwin 2010), culminating in the activation of defense responses.
2.4 Recognition of Nematodes
Plants are continuously exposed to a plethora of microorganisms surrounding their rhizosphere. The different varieties of root exudates secreted by the plants may either attract or deter away these microorganisms. Plants secrete flavonoid compounds that can attract symbiotic microbes like Rhizobia in case of beneficial interactions, phytoalexins to deter pathogen growth or allelopathic phenolic compounds to alter the growth of other plants (Hirsch et al. 2003). However, plant parasitic nematodes like root-knot nematode and potato cyst nematode can sense these host-derived signals. Following penetration inside the host tissue, the nematode migrates to its feeding site inside the root, wherein it feeds upon the host nutrients resulting in altered root architecture and reduced crop yield. Since long it had been speculated that plants could also mount a PTI response against nematodes, however not much was known about the compounds which elicit plant defense response. Recently, a nematode pheromone, ascaroside has been identified that is perceived by host plants as a NAMP to mount a PTI response including activation of MAP kinase cascade, upregulation of plant defense hormones such as salicylic acid and jasmonic acid, and induction of defense responses (Manosalva et al. 2015; Holbein et al. 2016; Choi et al. 2016). Moreover, a nematode immune receptor NILR1 (nematode-induced LRR-RLK 1) belonging to the LRR-RLK has been identified in Arabidopsis that perceives NAMP and mounts PTI responses (Mendy et al. 2017).
2.5 Recognition of DAMPs
The plant cell wall serves as a formidable barrier against pathogens. Pathogen secretes various proteins to degrade different components of the plant cell (Jha et al. 2005). Moreover, plants have evolved the ability to sense this damage by recognition of the cell wall degradation products. Treatment of Arabidopsis with cellulose degradation products such as cellobiose, cellotriose, etc. or cellulose synthesis inhibitors (Engelsdorf et al. 2017) activates the host immune responses (Souza et al. 2017). Similarly, the treatment of plant tissue with pectin degradation products such as oligogalacturonides (OG) can activate the host immune responses (Ferrari 2013). In Arabidopsis, wall-associated kinases (AtWAK1 and AtWAK2) can perceive pectin and pectin degradation products (OG) (Brutus et al. 2010; Decreux and Messiaen 2005; Decreux et al. 2006). The activation of immune responses by WAKs has also been reported in other plant species such as rice and maize (Delteil et al. 2016; Zuo et al. 2015; Hu et al. 2017).
In response to pathogen/damage perception, plants secrete various peptides and nucleotides in their apoplast to amplify the immune response and trigger an elaborate defense mechanism in their neighboring cells (Boutrot and Zipfel 2017). Release of plant elicitor peptides (Peps, also known as danger peptides) derived from PROPEPs (precursor proteins) has been reported in Arabidopsis upon pathogen attack (Bartels et al. 2013; Klauser et al. 2015). Arabidopsis secretes 23aa long endogenous elicitor peptides known as AtPep1, which are recognized via LRR-RLK PEP receptor (PEPR) (Krol et al. 2010). Moreover, it has been reported that extracellular ATP (eATP) can act as a DAMP in Arabidopsis (Weerasinghe et al. 2009; Wu et al. 2008). The eATP is recognized by a lectin receptor kinase-I.9 (LecRK-I.9) named DORN1 (Does not Respond to Nucleotides 1) in Arabidopsis which activates downstream defense-responsive genes (Choi et al. 2014).
2.6 Recognition of Effectors
PTI and DTI form the first layer of plant immune responses. Pathogens can suppress these immune responses by secreting effector molecules directly into plant cells via the type-III-secretion system (Alfano and Collmer 2004). However, plants have evolved R gene-encoded proteins to recognize effector proteins to activate effector-triggered defense (ETD) response (Dodds and Rathjen 2010). Plants can either directly recognize effector molecules via NB-LRR (or NLR) domain-containing receptor proteins or can indirectly sense their presence by monitoring their activity (Kourelis and van der Hoorn 2018). In both cases, plants mount a robust immune response that usually culminates in a hypersensitive response and localized death of plant tissue to limit spread of the pathogen. The NLR receptor proteins are usually comprised of either coiled-coil (CC) domain or toll/interleukin-1 receptor (TIR) domains at their N-terminal (Cui et al. 2015; Schreiber et al. 2016). However, there are exceptions, wherein certain effector proteins are not directly recognized by receptor proteins, instead are recognized when bound to an accessory protein (guardee). The guard model has been proposed to explain this phenomenon (Dangl and Jones 2001). Further, a modification of this hypothesis has been proposed as a decoy model, wherein certain effector targets have evolved to function as decoys (co-receptor) which bind to the effectors and cause activation of the defense response (van der Hoorn and Kamoun 2008). Due to a few limitations in the decoy model, an improved bait-and-switch model was proposed. In this model, a two-step recognition has been proposed wherein the accessory protein (bait) associated with the receptor protein interacts with the effector protein to mount a defense response (Collier and Moffett 2009). The current hypothesis states that the receptor protein instead of recognizing the accessory protein directly recognizes the effector protein only when it is bound with its accessory protein (Dodds and Rathjen 2010). We will now provide an outline of the different effector molecules that are secreted in different pathosystems and how plants are able to recognize them.
2.6.1 Bacterial Effector Recognition
The AvrPto and AvrPtoB (also known as HopAB2) effectors secreted by pathogenic strains of P. syringae (Abramovitch et al. 2003; Ronald et al. 1992) are recognized by plants to mount immune responses. AvrPto and AvrPtoB bind to various PTI receptors and suppress immune responses. For example, AvrPto binds to various PTI receptors like FLS2 and EFR while AvrPtoB binds to FLS2, BAK1 and LysM receptor kinases and suppress immune responses (Cheng et al. 2011; Gimenez-Ibanez et al. 2009; Göhre et al. 2008; Shan et al. 2008; Xiang et al. 2008; Zeng et al. 2012). Prf/Pto protein complex recognizes the presence of both of these effector molecules, whereas Pto has binding sites for both AvrPto and AvrPtoB as well as Prf. Prf acts as a positive regulator of ETI. In the native state, Pto binds to Prf along with some other kinases to form a large macromolecular complex that keeps Prf in its inactive state (Ntoukakis et al. 2013). In presence of cognate effectors, Pto binds to the effector, gets released from Prf/Pto complex and in turn activates ETI (Abramovitch et al. 2003; Dong et al. 2009; Mathieu et al. 2014).
Rin4 (RPM1 interacting protein 4) is a membrane-localized protein that lacks any functional domain but is a part of many PRR complexes (Selote and Kachroo 2010). Rin4 can activate as well as suppress PTI depending on the phosphorylation status of the protein (Chung et al. 2014). Pathogens have evolved effector molecules such as AvrB, AvrRpt2, AvrRpm1, and HopF2 to directly or indirectly target Rin4 to suppress PTI (Lee et al. 2015; Russell et al. 2015; Wang et al. 2010; Wilton et al. 2010). In response, plants have also evolved R genes such as RPS2 (resistance to P. syringae) and RPM1 (resistance to P. syringae pv. maculicola) to sense the activity of effectors on Rin4 and mount defense responses (Chung et al. 2014; Coaker et al. 2005; Kim et al. 2005).
2.6.2 Fungal Effector Recognition
Fungal pathogens are also known to produce effector molecules which can either be secreted into the host cytoplasm or localized into the apoplastic space (Giraldo et al. 2013; Stotz et al. 2014). The recognition of apoplastic effectors is mediated by integral membrane proteins (RLPs) containing an extracellular leucine-rich repeat (eLRR) (Stergiopoulos and de Wit 2009). Induction of RLPs has been reported in tomato, apple, and oilseed rape against fungal pathogens like Cladosporium fulvum, Venturia inaequalis, and Leptosphaeria maculans, respectively (Rouxel and Balesdent 2013; Belfanti et al. 2004). However cytoplasmic effectors secreted by pathogens like Blumeria graminis, Bremia lactucae, Puccinia striiformis, Magnaporthe grisea, and Phytophthora infestans are recognized by NBS-LRR receptors that are present in the cytoplasm of respective host species (Bozkurt et al. 2010; Bai et al. 2012; Bonardi et al. 2012; Larkan et al. 2013; Rooney et al. 2005).
2.6.3 Nematode Effector Recognition
Plants utilize NB-LRR immune receptors to recognize effectors secreted from root or cyst nematodes to activate host defense responses. Some common examples of immune receptors against nematodes are Gpa2, Gro1-4 and Hero (Goverse and Smant 2014). It has been observed that root-knot nematodes secrete a diffusible compound called NemF that is very similar to NF (nodulation factor) secreted by symbiotic bacteria. The NemF signal is perceived by the plant through primary receptor kinases NFR1 and NFR5 along with secondary receptor kinase SYMRK. Signal perception leads to root hair branching and waviness which in turn facilitate nematode penetration (Weerasinghe et al. 2005). Plants also encode R genes to recognize effector proteins secreted by herbivores (Hogenhout and Bos 2011). Examples of R genes which confer resistance against herbivores are Mi-1.2 (Meloidogyne 1.2), Vat (Virus aphid transmission resistance) and Bph14 (Brown planthopper 14).
2.6.4 Miscellaneous
Necrosis and ethylene-inducing peptide 1-like proteins (NLPs) are plant immunogenic proteins with cytotoxic activity produced by a vast variety of bacterial, fungal, and oomycete species (Oome et al. 2014). Plants belonging to Brassicaceae family can recognize a conserved 20aa long fragment of NLP called nlp20 to activate their immune responses (Böhm et al. 2014; Oome et al. 2014; Oome and Van den Ackerveken 2014). In Arabidopsis, the LRR-RLP AtRLP23 recognizes nlp20 and activates immune responses by making a tripartite complex with two LRR-RLK, BAK1 (brassinosteroid insensitive 1 (BRI1)-associated kinase) and SOBIR1 (Albert et al. 2015).
3 Players Involved in Transduction of a Perceived Signal
The PRR proteins present on the plant cell surface can recognize pathogen attack and mount a defense response against the pathogen. However, induction of defense responses involves an intricate signaling network that transduces the signal to downstream molecular players to trigger immune responses. These signaling molecules include protein kinases (CDPKs, MAPKs), Ca2+ burst, ROS burst, NO, lipids, 14-3-3 proteins and various phytohormones (such as SA, JA and ethylene) (Bigeard et al. 2015).
3.1 Phosphorylation Events
Phosphorylation and dephosphorylation of proteins by kinases and phosphatases play an important role in the signal transduction process. After ligand binding, conformational changes in protein/binding with co-receptors lead to phosphorylation of the receptor. Somatic embryogenesis receptor kinase (SERK) family usually works as a co-receptor for many receptor kinases such as FLS2, EFR, BRI1, Xa21, PEPR, PSKR, etc. (Ma et al. 2016). In Arabidopsis, SERK3 [also called bri1-associated receptor kinase 1 (BAK1)] is a key co-receptor for many receptor kinases and is required for proper induction of immune responses (Ma et al. 2016). In rice, OsSERK2 interacts with Xa21, Xa3 and FLS2 (Chen et al. 2014) and is required for receptor-mediated resistance against Xoo. SERKs are also involved in RLP-mediated activation of immune responses such as nlp20-triggered immunity in Arabidopsis, csp22-triggered immunity in Nicotiana, Avr4- and Avr9-induced HR in tomato (Albert et al. 2015; Postma et al. 2016; Saur et al. 2016).
3.1.1 MAP Kinases
Mitogen-activated protein kinases (MAPKs) form signaling modules, which translate extracellular stimuli of pathogen attack into appropriate defense responses. MAPK cascade typically contains three sequential kinases (Rasmussen et al. 2012):
-
MAP kinase kinase kinase (MAPKKK or MEKK)
-
MAP kinase kinase (MAPKK or MKK)
-
MAP kinase (MAPK or MPK)
Usually receptor/co-receptor phosphorylates MAPKKK that phosphorylates MAPKK which phosphorylates MAPK. MAPK then phosphorylates downstream signaling components such as transcription factors and modulates defense responses (Meng and Zhang 2013). In a recent study, it has been shown that phosphorylation of OsMKK3-OsMPK7-OsWRKY30 leads to transcriptional activation of defense responses against X. oryzae in rice (Jalmi and Sinha 2016). Interestingly, in order to suppress PTI response, pathogens have evolved effector molecules that majorly target MAPK modules due to their primary role in defense signaling of plants (Feng et al. 2012).
3.1.2 CDPKs
Calcium-dependent protein kinases (CDPKs) have a serine/threonine protein kinase domain at their N-terminal and CaM-like domain with EF-hand calcium-binding sites at their C-terminal (Boudsocq and Sheen 2013). They act as Ca2+ sensors and decode the signal to generate a swift response to the external stimulus (Seybold et al. 2014, 2017). CDPK response was found to be associated with changes in host physiology such as transcriptional reprogramming, ROS accumulation, and alteration of phytohormone levels. CDPKs together with MAPKs have been found to orchestrate the transcriptional regulation of defense genes under pathogen attack (Boudsocq et al. 2010). Another group of kinases called AGC kinases, comprising of cAMP-dependent protein kinase 1 (PKA) and cGMP-dependent protein kinase (PKG) along with protein kinase C (PKC), have been shown to regulate MAPK signaling cascade upon pathogen attack (Garcia et al. 2012).
3.1.3 14-3-3
14-3-3 proteins act as phosphosensors which bind to phosphorylated proteins and regulate their functions. 14-3-3 proteins aid in phosphorylation of proteins thereby activating them (Chevalier et al. 2009). They play a crucial role in strengthening plant defense mechanisms by interacting with MAPKK proteins involved in the defense signal transduction pathway (Oh et al. 2010; Oh and Martin 2011). Induction of 14-3-3 proteins was found primarily in the penetration stage and upper epidermis of barley infected with Blumeria graminis suggesting its involvement in early signaling events (Lozano-Durán et al. 2015). 14-3-3 proteins have been found to interact with plant immune-responsive proteins such as receptor kinase BAK1 and WRKY transcription factor along with few R genes (Chang et al. 2009). 14-3-3 proteins have also been reported to regulate phytohormone levels in infected plants culminating in enhanced immune responses. (Chang et al. 2009; Camoni et al. 2018).
3.1.4 Heterotrimeric G proteins
G proteins have been found to play a critical role in defense signaling in animals. However plants lack the canonical G protein structure as observed in animals (Urano and Jones 2014). G proteins are known to activate plant defense signaling responses mediated by the action of multiple RLKs (Liu et al. 2013a, b; Maruta et al. 2015). The signals received from RLKs by G proteins are transduced downstream to different MAPKs and ROS signaling genes (Nitta et al. 2015; Cheng et al. 2015). Studies have revealed direct physical association between the Gα, Gγ1, and Gγ2 subunits and RD-type kinases CERK1, BAK1, and BIR1 to activate the plant defense network (Aranda-Sicilia et al. 2015).
3.2 Regulation of Immune Responses
Plant immune responses are metabolically costly affair; plants regulate the processes in a tight manner to avoid non-specific activation and dampen the responses when they are no longer required. This is usually achieved by dephosphorylation or degradation of receptors. After activation of immune responses, protein phosphatases (PP) such as PP2C and PP2A dephosphorylate the receptor and other intermediate kinases to negatively regulate immune responses (Durian et al. 2016; Fuchs et al. 2013). Some examples of PP2C involvement in immunity include kinase-associated protein phosphatase (KAPP), PLL4 and PLL5 of Arabidopsis, and XB15 of rice (Holton et al. 2015; Park et al. 2008).
Another approach to regulate immune response is via vesicle-mediated internalization of activated receptors or degradation of the receptor/signaling intermediate (Wang et al. 2016a, b). These proteins are polyubiquitinated by E3 ubiquitin ligases and degraded by 26S proteasomes. Some examples of this pathway include XB3 of rice and PUB12 and PUB13 of Arabidopsis (Lu et al. 2011; Wang et al. 2006).
3.3 Transcriptional Regulation
Activation of immune responses involves rapid transcriptional and translational changes (Li et al. 2016). Transcriptional events are modulated by transcription factors (TFs) which get activated by MAP kinases, Ca2+ signaling or hormonal response (Kang et al. 2015; Li et al. 2016). Some key TF families involved in defense responses include WRKY, MYC, TCP, ZIP, MVQ, AP2/ERF, etc. (Birkenbihl et al. 2017). TFs enhance expression of various defense genes such as PR genes, secondary metabolism, and hormone biosynthesis as well as regulation of related genes.
3.4 Secondary Signaling Molecules
Many non-proteinaceous molecules are key signaling intermediates in plant innate immunity. These molecules include Ca2+, ROS, NO, etc.
3.4.1 Burst of Ca2+
Ca2+ ions play an important role in defense signaling during pathogen attack. Ca2+ burst occurs when MAMPs/DAMPs are perceived and Ca2+ from the extracellular milieu is transported into the cytoplasm (Jeworutzki et al. 2010; Ranf et al. 2011). The permeability of plasma membrane to Ca2+ is mediated by elicitor responsive ion channels. The calcium levels accumulate in distinct signature patterns and generate a particular defense response pathway against the pathogen (Lecourieux et al. 2006). Influx of Ca2+ is followed by opening of other membrane ion transporters such as H+, K+, Cl-, and NO3- channels which lead to alkalization of extracellular space and membrane depolarization (Jeworutzki et al. 2010).
EF-hand motif-containing proteins are known to bind with calcium and serve as sensors of Ca2+ concentration (Schulz et al. 2013). These proteins mainly include Ca2+-dependent protein kinases (CDPK) and calmodulin (CaM). Ca2+ binding causes conformational changes in structure of these proteins leading either to phosphorylation or binding with downstream signaling intermediates (Ishida and Vogel 2006; Wernimont et al. 2010).
3.4.2 ROS Burst
Production of extracellular reactive oxygen species (ROS) also referred to as ROS burst has been found to be associated with pathogen attack (Ranf et al. 2011; Chinchilla et al. 2007; Nühse et al. 2007). MAMP perception is often associated with ROS production by respiratory burst oxidase homolog D (RBOHD), a member of NADPH oxidase family in Arabidopsis (Bigeard et al. 2015). ROS can be present in membranes as impermeable superoxide (O2-) or as permeable hydrogen peroxide (H2O2) and it can be readily translocated from one cell to another. Also it is often associated with elevated Ca2+ levels in the cytosol (Ranf et al. 2011; Bigeard et al. 2015). ROS signaling is accompanied by alteration in plant defense hormone levels such as JA, SA, and ethylene indicating a complex crosstalk between different pathways (Baxter et al. 2014).
3.4.3 NO Signaling
Nitric oxide (NO) along with its derivatives has also been involved in signal transduction pathway upon perception of pathogen attack. The role of NO in activating plant defense was first reported in tobacco mosaic virus infection wherein increase in NO synthase (NOS) resulted in activation of several downstream defense genes (Klessig et al. 2000). Interestingly, NO together with ROS plays a synergistic role in activation of plant defense responses (Domingos et al. 2015). NO can cause a rapid change in cellular glutathione levels in the cell associated with accumulation of SA and activation of NPR1-mediated defense responses (Kovacs et al. 2015).
3.4.4 Lipid Signaling
Lipid-based signaling molecules are also known to play a crucial role in defense signaling upon pathogen attack. These lipid molecules are produced as a result of degradation/destabilization of the cell wall upon pathogen attack. For example, phosphatidic acid (PA) and ceramides have been found to be involved in signal transduction upon pathogen infection (Okazaki and Saito 2014). PA is also involved in release of other signaling intermediates such as DAG, free fatty acids, and lysoPA which in turn induce downstream defense signaling (Wang 2004). Phospholipase A (PLA) which catalyzes the hydrolysis of phospholipids is involved in release of free fatty acids which are utilized during biosynthesis of defense hormone jasmonic acid (Shah 2005).
3.4.5 Hormonal Signaling
Major phytohormones such as jasmonic acid (JA), salicylic acid (SA) and ethylene have been found to play an important role in coordinating cell-to-cell communication during perception of pathogen attack. Each of these phytohormones activates its own downstream targets which lead to diverse immune and signaling events. There are also reports that other phytohormones such as auxin, cytokinin, abscisic acid, gibberellins and brassinosteroids are involved in plant immunity. There is a complex crosstalk among different phytohormones occurring at the cellular level that tailors a specific defense response upon attack by a specific pathogen. Here we summarize the role of some key defense-related phytohormones.
3.4.5.1 Salicylic Acid
SA is a phenolic hormone that is synthesized from chorismate via phenylalanine ammonia pathway (PAL) or isochorismate synthase (ICS) pathway (Chen et al. 2009). It is a key component of PTI as well as ETI and is known to enhance tolerance against various biotrophic, hemi-biotrophic, and viral infections (Dodds and Rathjen 2010; Malamy et al. 1990; Shigenaga and Argueso 2016). It is also necessary for the activation of various PR genes. Arabidopsis ICS1 mutant (ics1), also called SA deficient 2 (sid2), was compromised in SA-mediated immune response (Dewdney et al. 2000; Wildermuth et al. 2001). Interestingly, the non-expressor of PR genes 1 (NPR1) acts as a transcription co-activator and plays a key role in SA-mediated immune responses (Cao 1994). Generally at normal SA levels, NPR1 is localized in the cytoplasm in oligomeric form (Mou et al. 2003). However, at elevated SA level, the NPR1 binds to SA, adopts monomeric form, and gets transported to the nucleus (Kinkema et al. 2000; Mou et al. 2003). In the nucleus, NPR1 binds to TGA transcription factors and activates expression of defense-related genes including PR genes (Kesarwani et al. 2007). Infection studies on ics1 mutant (that fails to increase SA level), NahG (salicylate hydroxylase that degrades SA) expressing transgenic lines (that fail to accumulate SA), and npr1 mutant (that does not respond to SA) indicate that although SA can enhance tolerance towards biotrophic and hemi-biotrophic pathogens, it reduces resistance towards necrotrophic pathogens (Delaney et al. 1994; Glazebrook et al. 1996; Thomma et al. 1998). It is worth noting that phytopathogens utilizes various effectors (such as HopJ, HaRxL44, HopM1 and PsIcs1) to target SA signaling pathway during host colonization (Caillaud et al. 2013; DebRoy et al. 2004; Liu et al. 2014).
3.4.5.2 Jasmonic Acid
JA is a lipid-derived hormone that is involved in many developmental and defense response pathways (Santino et al. 2013; Carvalhais et al., 2017). JA is synthesized by oxygenation of α-linolenic by lipoxygenase (Lox) enzymes and is converted into JA-Ile (JA-isoleucine; the active form of JA) by JA amido synthetase (JAR1) (Staswick 2004; Wasternack and Hause 2013). Coronatine insensitive 1 (COI1), an E3 ubiquitin ligase, is a receptor of JA, and a transcription factor jasmonate ZIM domain 1 (JAZ1) is a negative regulator of JA pathway (Sheard et al. 2010; Yan et al. 2009). At low JA levels, JAZ1 represses JA-responsive genes (Pauwels et al. 2010). After perception of pathogen attack, JA-Ile binds to COI1, which ubiquitinates JAZ1 leading to degradation of JAZ1. Degradation of JAZ1 leads to enhanced expression of JA-responsive genes (Thines et al. 2007).
JA and SA are believed to play antagonistic roles against each other in very complex plant defense response-activating pathways depending on the nature of the pathogen (Thaler et al. 2012; Robert-Seilaniantz et al. 2011). Pathogens have evolved mechanisms to utilize this crosstalk to suppress plant immune responses (Pieterse et al. 2012). A well-studied example is synthesis of the JA mimic molecule coronatine (COR) by Pseudomonas sp. COR activates the JA pathway and suppresses SA pathway leading to increased susceptibility towards biotrophic and hemi-biotrophic pathogens including Pseudomonas (Zheng et al. 2012). Interestingly a hemi-biotrophic pathogen, i.e., Pseudomonas, utilizes effector molecules such as HopZ1 and HopX1 to induce JA pathway during pathogenicity process (Gimenez-Ibanez et al. 2014; Jiang et al. 2013).
3.4.5.3 Ethylene
Ethylene is a gaseous plant hormone known for its role in fruit ripening. However, it is also known to be involved in plant defense responses. ET and JA phytohormones work in a synergistic manner (Robert-Seilaniantz et al. 2011). The activation of JA pathway leads to enhanced expression of ET pathway genes (Penninckx et al. 1998; Zhu et al. 2011). Alike JA, ET also enhances tolerance towards necrotrophic pathogens but increases susceptibility towards biotrophic pathogens (Lawton et al. 1994, 1995). Similar to other phytohormones, ET pathway is also targeted by pathogens to overcome immunity. For example, AvrPto and AvrPtoB effectors of Pseudomonas sp. and XopD effector of Xanthomonas sp. have been found to alter the ET pathway (Cohn and Martin 2005; Kim et al. 2013).
4 Plant Defense Responses
Upon perception of pathogen attack, plants mount a strong immune response to restrict the spread of pathogen/predator. These immune responses involve strengthening of the cell wall, localized cell death, production of antimicrobial compounds, etc. The strength of the immune response depends upon the type of danger. Many pathogens have evolved mechanisms to suppress PTI/ETI directly by secreting effector molecules into the plant cell. This is known as effector-triggered susceptibility (ETS). However, recognition of effectors by host R genes leads to activation of ETI that includes robust defense responses such as programmed cell death to restrict the growth of the pathogen at the site of infection.
4.1 Stomatal Closure
Several phytopathogens use stomata to enter inside the host. Closure of stomata is one of the early defense responses used by the host to prevent pathogens from colonization. Upon perception of pathogen cues (flg22, elf18, elf26, LPS, chitin, oligogalacturonan, etc.), plants close their stomata (Arnaud and Hwang 2015; Murata et al. 2015). This process involves various signaling events including activation of MAP kinase pathway, synthesis of hormones, Ca2+ influx, ROS and NO production, etc. (Desclos-Theveniau et al. 2012; Melotto et al. 2006, 2017). SA and ABA pathways are known to promote stomatal closure while JA-Ile serves as a negative regulator of stomatal closure.
However, successful phytopathogens have evolved various mechanisms to avoid plant stomatal closure. For example, P. syringae secretes various effectors such as HopM1, HopF2, HopZ1, HopZ1a, Hopx1 and AvrB to suppress closure of stomata (Gimenez-Ibanez et al. 2014; Hurley et al. 2014; Jiang et al. 2013; Lozano-Durán et al. 2014; Zhou et al. 2014, 2015). XopR, a Xoo-secreted effector, suppresses flg22-induced stomatal closure in rice (Wang et al. 2016a, b). On the other hand, some of the bacterial pathogens secrete phytotoxins to open stomatal pores to assist colonization. Some of the notable phytotoxins used by bacterium to open stomata are coronatin (COR) (Bender et al. 1999) and syringolin A secreted by P. syringae (Groll et al. 2008), plant natriuretic peptide-like (Gottig et al. 2008) and diffusible signaling factor (DSF) (Gudesblat et al. 2008) molecules secreted by Xanthomonas species.
4.2 Cell Wall Strengthening
Cell wall serves as a key barrier to phytopathogens. Pathogens need to degrade the cell wall to gain access to nutrients that are inside the plant cell. Strengthening of the cell wall is achieved by deposition of callose (β-1,3 glucan) and lignin (phenolic polymers). This is one of the basic mechanisms used by the host plant to suppress the growth of pathogen (Malinovsky et al. 2014). Treatment with various MAMPs, DAMPs or avirulent pathogen strains causes callose deposition in the infected tissues (Luna et al. 2011). Synthesis of callose usually leads to papillae formation that contains antimicrobial compounds such as thionins, H2O2, etc. (McLusky et al. 1999; Thordal-Christensen et al. 1997; Voigt 2016). Besides callose, lignin is also deposited at the secondary cell wall to provide mechanical strength (Malinovsky et al. 2014). Loss of function mutations in various genes involved in lignin synthesis pathway makes the plants more susceptible to pathogens (Miedes et al. 2014).
4.3 Pathogenesis-Related Proteins
Expression of pathogenesis-related (PR) proteins is upregulated in plants after pathogen infection. These proteins are key components of plant immune responses. Many PR proteins are also observed to be upregulated after MAMP and DAMP treatment, wounding, ETI activation, and treatment with immune response-associated hormones (Sels et al. 2008). PR genes encode diverse classes of proteins which can be classified under 17 different families (van Loon et al. 2006). Most of the PR proteins have antimicrobial activities. PR3, PR4, PR8 and PR10 are chitinases which can degrade fungal cell wall, while PR2 proteins are β-1,3-glucanases. PR1 is a most common PR protein accumulated in various plant species upon pathogen attack and is known to have antimicrobial activity (Ménard et al. 2005; Segarra et al. 2013; Song et al. 2015). Some PR genes encode small peptides such as the PR6 family containing proteinase inhibitor peptides (Green and Ryan 1972), PR12s are cysteine-rich defensins (Terras 1995), PR13 encodes thionins (Epple et al. 1995) and PR14 codes for lipid transfer proteins (LPT) (García-Olmedo et al. 1995).
Interestingly, AtPR1, AtPR2, and AtPR5 are SA-responsive genes known to provide resistance against biotrophic and hemi-biotrophic pathogens in Arabidopsis. AtPR3 and AtPR4 are JA-responsive genes and provide tolerance against necrotrophic pathogens and herbivores (van Loon et al. 2006).
4.4 Secondary Metabolites
Plants produce various types of secondary metabolites upon infection by phytopathogens (Piasecka et al. 2015). These metabolites usually have antimicrobial activity and have a toxic effect on phytopathogens. One type of secondary metabolites that are constitutively produced are called phytoanticipins (VanEtten 1994). These are usually produced in an inactive form and are activated by hydrolysis after perception of danger. Plants produce various kinds of phytoanticipins including saponins such as α-tomatine and avenacin, glucosinolates, cyanogenic glucosides and benzoazinone glucoside compounds (Faizal and Geelen 2013; Halkier and Gershenzon 2006; Burkhardt et al. 1964; Papadopoulou et al. 1999; Sandrock and VanEtten 1998). Secondary metabolites that are de novo synthesized in response to biotic stress are called phytoalexins. The major types of phytoalexins include camalexins, phenylalanine-derived phytoalexins, and terpenoids (VanEtten 1994). Mutations in secondary metabolite synthetic genes make plants more susceptible to pathogens (Toyomasu et al. 2014; Xu et al. 2012).
4.5 Hypersensitive Response
Sometimes, plants undergo programmed cell death in the infected area to restrict the spread of a pathogen. This process is called hypersensitive response (HR). PCD is generally involved in developmental processes and stress responses including tolerance towards biotic stress (Bozhkov and Lam 2011; Pennell 1997). Upon pathogen perception by the host R gene, an intricate signaling cascade is triggered that leads to HR. This signaling cascade involves MAP kinase activation, SA production, ROS production, NO accumulation, cytosolic Ca2+ increase, membrane depolarization, etc. (Kadota et al. 2004; Kärkönen and Kuchitsu 2015; Kurusu et al. 2011). HR can be observed as the lesion phenotypes during infection or elicitor treatment or as lesion mimic phenotype if the immune response is constitutively activated (Coll et al. 2011; Lorrain et al. 2003). Although HR is a strong immune response, sometimes it can act like a double-edged sword for plants as necrotrophic pathogens that flourish on dead plant tissues have evolved various pathways to utilize this immune response of plants to colonize host tissues (Mukhtar et al. 2016). These pathogens modulate plant signaling to enhance ROS production and induce HR (Shetty et al. 2008).
4.6 Secretion of Volatile Compounds
Upon pathogen attack, plants often emit gaseous compounds known as VOCs (volatile organic compounds). Emission of volatile derivatives of certain plant hormones such as jasmonic acid and ethylene have been found to be responsible for the systemic activation of plant defense responses (Champigny and Cameron 2009; Fiers et al. 2013; Wasternack and Kombrink 2010; Tamogami et al. 2008). Plants also secrete volatile components that can attract predators or parasitoids such as parasitic wasps to forage upon the feeding insects or induce a systemic defense response in distal uninfected plant parts (Heil 2008; Heil and Silva Bueno 2007; Frost et al. 2007). Volatile compounds that are thus secreted are known as herbivore-induced plant volatiles (HIPV). Lima bean plants secrete certain compounds which are not only involved in attraction of predatory insects (natural enemies of herbivores) but also in production of certain extrafloral nectars (EFN) which serve as a food source for these predatory insects (Choh and Takabayashi 2010). Apple plants have been found to emit certain VOCs upon infection by the bacterial pathogen Erwinia amylovora which can activate the defense responses even in surrounding healthy uninfected plants (Cellini et al. 2018). Interestingly, it has been observed that VOCs produced upon infection by fungal pathogen Colletotrichum lindemuthianum in resistant bean plants can trigger defense responses in neighbouring susceptible plants (Quintana-Rodriguez et al. 2015).
5 Conclusion
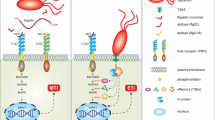
Plants have specialized receptors to sense pathogen attack and mount potent defense responses. Various conserved structural components, damaged cell wall products or effector molecules produced by the pathogens are recognized by these receptors. Generally, these receptor proteins are maintained in a dephosphorylated inactive state and get activated at the time of pathogen attack. Various signaling intermediates like MAPKs, CDPKs, 14-3-3 and heterotrimeric G proteins participate in interception, amplification and transduction of the signal from the receptor to the target defense genes. Also several secondary messengers such as ROS, Ca2+, NO, lipids, and hormones help in the relay of signal (Fig. 20.1). Interestingly, the induction of defense responses is not merely restricted to the infected tissue but it is also elaborated in uninfected as well as distal parts of the plant. Interestingly, phytohormones such as salicylic acid, jasmonic acid and ethylene play a significant role in activation of immune responses.
Schematic overview of cellular responses induced in host upon perception of pathogen attack. The pathogen possesses certain conserved structural components (MAMPS, DAMPs, HAMPs, NAMPs, etc.) which are recognized by cognate receptors present in host plant. Upon signal perception, a cascade of signal transduction events including induction of phosphorylation events (involving MAP kinases, CDPKs, etc.) as well as secondary signaling molecules (such as calcium, NO, ROS, etc.) are triggered. All these events culminate in activation of potent immune responses which combat most of the potential pathogens to cause disease. Notably, several phytohormones such as SA, JA, ET, etc. also play important roles in elucidation of plant defense responses. Additionally, pathogens secrete effector molecules to inhibit plant immune responses, but plants have evolved resistance genes (R genes) to directly or indirectly recognize them and mount a strong defense response
References
Abramovitch RB, Kim YJ, Chen S, Dickman MB, Martin GB (2003) Pseudomonas type III effector AvrPtoB induces plant disease susceptibility by inhibition of host programmed cell death. EMBO J 22:60–69
Albert I, Böhm H, Albert M, Feiler CE, Imkampe J, Wallmeroth N, Brancato C, Raaymakers TM, Oome S, Zhang H, Krol E, Grefen C, Gust AA, Chai J, Hedrich R, Van Den Ackerveken G, Nürnberger T (2015) An RLP23-SOBIR1-BAK1 complex mediates NLP-triggered immunity. Nat Plants 1:15140
Alfano JR, Collmer A (2004) TYPE III secretion system effector proteins: double agents in bacterial disease and plant defense. Annu Rev Phytopathol 42:385–414
Andolfo G, Ercolano MR (2015) Plant innate immunity multicomponent model. Front Plant Sci 6:987
Aranda-Sicilia MN, Trusov Y, Maruta N, Chakravorty D, Zhang Y, Botella JR (2015) Heterotrimeric G proteins interact with defense-related receptor-like kinases in Arabidopsis. J Plant Physiol 188:44–48
Arnaud D, Hwang I (2015) A sophisticated network of signaling pathways regulates stomatal defenses to bacterial pathogens. Mol Plant 8:566–581
Bacete L, Mélida H, Miedes E, Molina A (2018) Plant cell wall-mediated immunity: cell wall changes trigger disease resistance responses. Plant J 93:614–636
Bai S, Liu J, Chang C, Zhang L, Maekawa T, Wang Q, Xiao W, Liu Y, Chai J, Takken FLW, Schulze-Lefert P, Shen QH (2012) Structure-function analysis of barley NLR immune receptor MLA10 reveals its cell compartment specific activity in cell death and disease resistance. PLoS Pathogens 8:e1002752
Bailey BA, Dean JF, Anderson JD (1990) An Ethylene biosynthesis-inducing endoxylanase elicits electrolyte leakage and necrosis in nicotiana Tabacum cv Xanthi leaves. Plant Physiol 94:1849–1854
Bailey BA, Korcak RF, Anderson JD (1993) Sensitivity to an ethylene biosynthesis-inducing endoxylanase in nicotiana tabacum L. cv. Xanthi is controlled by a single dominant gene. Plant Physiol 101:1081–1088
Bar M, Avni A (2009) EHD2 inhibits ligand-induced endocytosis and signaling of the leucine-rich repeat receptor-like protein LeEix2. Plant J 59:600–611
Bar M, Sharfman M, Schuster S, Avni A (2009) The coiled-coil domain of EHD2 mediates inhibition of LeEix2 endocytosis and signaling. PLoS ONE 4:e7973
Bartels S, Lori M, Mbengue M, van Verk M, Klauser D, Hander T, Böni R, Robatzek S, Boller T (2013) The family of peps and their precursors in arabidopsis: Differential expression and localization but similar induction of pattern-Triggered immune responses. J Exp Bot 64:5309–5321
Baxter A, Mittler R, Suzuki N (2014) ROS as key players in plant stress signalling. J Exp Bot 65:1229–1240
Belfanti E, Silfverberg-Dilworth E, Tartarini S, Patocchi A, Barbieri M, Zhu J, Vinatzer BA, Gianfranceschi L, Gessler C, Sansavini S (2004) The HcrVf2 gene from a wild apple confers scab resistance to a transgenic cultivated variety. Proc Natl Acad Sci USA 101:886–890
Bender CL, Alarcón-Chaidez F, Gross DC (1999) Pseudomonas syringae phytotoxins: mode of action, regulation, and biosynthesis by peptide and polyketide synthetases. Microbiol Mol Biol Rev 63:266–292
Bigeard J, Colcombet J, Hirt H (2015) Signaling mechanisms in pattern-triggered immunity (PTI). Mol Plant 8:521–539
Birkenbihl RP, Liu S, Somssich IE (2017) Transcriptional events defining plant immune responses. Curr Opin Plant Biol 38:1–9
Böhm H, Albert I, Oome S, Raaymakers TM, Van den Ackerveken G, Nürnberger T (2014) A conserved peptide pattern from a widespread microbial virulence factor triggers pattern-induced immunity in arabidopsis. PLoS Pathogens 10:e1004491
Bonardi V, Cherkis K, Nishimura MT, Dangl JL (2012) A new eye on NLR proteins: focused on clarity or diffused by complexity? Curr Opin Immunol 24:41–50
Bonaventure G, Baldwin IT (2010) Transduction of wound and herbivory signals in plastids. Commun Integr Biol 3:313–317
Bonaventure G, VanDoorn A, Baldwin IT (2011) Herbivore-associated elicitors: FAC signaling and metabolism. Trends Plant Sci 16:294–299
Boudsocq M, Sheen J (2013) CDPKs in immune and stress signaling. Trends Plant Sci 18:30–40
Boudsocq M, Willmann MR, McCormack M, Lee H, Shan L, He P, Bush J, Cheng SH, Sheen J (2010) Differential innate immune signalling via Ca(2+) sensor protein kinases. Nature 464:418–422
Boutrot F, Zipfel C (2017) Function, discovery, and exploitation of plant pattern recognition receptors for broad-spectrum disease resistance. Annu Rev Phytopathol 55:257–286
Bozhkov PV, Lam E (2011) Green death: revealing programmed cell death in plants. Cell Death Differ 18:1239–1240
Bozkurt TO, Mcgrann GRD, Maccormack R, Boyd LA, Akkaya MS (2010) Cellular and transcriptional responses of wheat during compatible and incompatible race-specific interactions with Puccinia striiformis f. sp. tritici. Mol Plant Pathol 11:625–640
Brutus A, Sicilia F, Macone A, Cervone F, De Lorenzo G (2010) A domain swap approach reveals a role of the plant wall-associated kinase 1 (WAK1) as a receptor of oligogalacturonides. Proc Natl Acadf Sci 107:9452–9457
Burkhardt HJ, Maizel JV, Mitchell HK (1964) Avenacin, an antimicrobial substance Isolated from avena sativa II structure. Biochemistry 3:426–431
Caillaud MC, Asai S, Rallapalli G, Piquerez S, Fabro G, Jones JDG (2013) A downy mildew effector attenuates salicylic acid-triggered immunity in arabidopsis by interacting with the host mediator complex. PLoS Biol 11:e1001732
Camoni L, Visconti S, Aducci P, Marra M (2018) 14-3-3 proteins in plant hormone signaling: doing several things at once. Front Plant Sci 9:297
Cao H (1994) Characterization of an arabidopsis mutant that is nonresponsive to inducers of systemic acquired resistance. Plant Cell 6:1583–1592
Cao Y, Liang Y, Tanaka K, Nguyen CT, Jedrzejczak RP, Joachimiak A, Stacey G (2014) The kinase LYK5 is a major chitin receptor in Arabidopsis and forms a chitin-induced complex with related kinase CERK1. eLife 3:e03766
Carvalhais LC, Schenk PM, Dennis PG (2017) Jasmonic acid signalling and the plant holobiont. Curr Opin Microbiol 37:42–47
Cellini A, Buriani G, Rocchi L, Rondelli E, Savioli S, Rodriguez Estrada MT, Cristescu SM, Costa G, Spinelli F (2018) Biological relevance of volatile organic compounds emitted during the pathogenic interactions between apple plants and erwinia amylovora. Mol Plant Pathol 19:158–168
Champigny MJ, Cameron RK (2009) Action at a distance. Long-distance signals in induced resistance. Adv Bot Res 51:123–171, Academic Press
Chang IF, Curran A, Woolsey R, Quilici D, Cushman JC, Mittler R, Harmon A, Harper JF (2009) Proteomic profiling of tandem affinity purified 14-3-3 protein complexes in Arabidopsis thaliana. Proteomics 9:2967–2985
Chen Z, Zheng Z, Huang J, Lai Z, Fan B (2009) Biosynthesis of salicylic acid in plants. Plant Signal Behav 4:493–496
Chen X, Zuo S, Schwessinger B, Chern M, Canlas PE, Ruan D, Zhou X, Wang J, Daudi A, Petzold CJ, Heazlewood JL, Ronald PC (2014) An XA21-associated kinase (OsSERK2) regulates immunity mediated by the XA21 and XA3 immune receptors. Mol Plant 7:874–892
Cheng W, Munkvold KR, Gao H, Mathieu J, Schwizer S, Wang S, YBin Y, Wang J, Martin GB, Chai J (2011) Structural analysis of pseudomonas syringae AvrPtoB bound to host BAK1 reveals two similar kinase-interacting domains in a type III effector. Cell Host Microbe 10:616–626
Cheng Z, Li JF, Niu Y, Zhang XC, Woody OZ, Xiong Y, Djonović S, Millet Y, Bush J, McConkey BJ, Sheen J, Ausubel FM (2015) Pathogen-secreted proteases activate a novel plant immune pathway. Nature 521:213–216
Chevalier D, Morris ER, Walker JC (2009) 14-3-3 and FHA domains mediate phosphoprotein interactions. Annu Rev Plant Biol 60:67–91
Chinchilla D, Zipfel C, Robatzek S, Kemmerling B, Nürnberger T, Jones JDG, Felix G, Boller T (2007) A flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature 448:497–500
Choh Y, Takabayashi J (2010) Herbivore-induced plant volatiles prime two indirect defences in lima bean BT. In: Sabelis MW, Bruin J (eds) Trends in acarology. Springer, Dordrecht, pp 255–258
Choi J, Tanaka K, Cao Y, Qi Y, Qiu J, Liang Y, Lee SY, Stacey G (2014) Identification of a plant receptor for extracellular ATP. Science 343:290–294
Choi HW, Manohar M, Manosalva P, Tian M, Moreau M, Klessig DF (2016) Activation of plant innate immunity by extracellular high mobility group box 3 and its inhibition by salicylic acid. PLoS Pathogens 12:e1005518
Chung EH, El-Kasmi F, He Y, Loehr A, Dangl JL (2014) A plant phosphoswitch platform repeatedly targeted by type III effector proteins regulates the output of both tiers of plant immune receptors. Cell Host Microbe 16:484–494
Coaker G, Falick A, Staskawicz B (2005) Activation of a phytopathogenic bacterial effector protein by a eukaryotic cyclophilin. Science 308:548–550
Cohn JR, Martin GB (2005) Pseudomonas syringae pv. tomato type III effectors AvrPto and AvrPtoB promote ethylene-dependent cell death in tomato. Plant J 44:139–154
Coll NS, Epple P, Dangl JL (2011) Programmed cell death in the plant immune system. Cell Death Differ 18:1247–1256
Collier SM, Moffett P (2009) NB-LRRs work a “bait and switch” on pathogens. Trends Plant Sci 14:521–529
Couto D, Zipfel C (2016) Regulation of pattern recognition receptor signalling in plants. Nat Rev Immunol 16:537–552
Cui H, Tsuda K, Parker JE (2015) Effector-triggered immunity: from pathogen perception to robust defense. Annu Rev Plant Biol 66:487–511
Dangl JL, Jones JD (2001) Plant pathogens and integrated defence responses to infection. Nature 411:826–833
DebRoy S, Thilmony R, Kwack Y-B, Nomura K, He SY (2004) A family of conserved bacterial effectors inhibits salicylic acid-mediated basal immunity and promotes disease necrosis in plants. Proc Natl Acad Sci 101:9927–9932
Decreux A, Messiaen J (2005) Wall-associated kinase WAK1 interacts with cell wall pectins in a calcium-induced conformation. Plant Cell Physiology 46:268–278
Decreux A, Thomas A, Spies B, Brasseur R, Van Cutsem P, Messiaen J (2006) In vitro characterization of the homogalacturonan-binding domain of the wall-associated kinase WAK1 using site-directed mutagenesis. Phytochemistry 67:1068–1079
Delaney TP, Uknes S, Vernooij B, Friedrich L, Weymann K, Negrotto D, Gaffney T, Gut-Rella M, Kessmann H, Ward E, Ryals J (1994) A central role of salicylic acid in plant disease resistance. Science 266:1247–1250
Delteil A, Gobbato E, Cayrol B, Estevan J, Michel-Romiti C, Dievart A, Kroj T, Morel JB (2016) Several wall-associated kinases participate positively and negatively in basal defense against rice blast fungus. BMC Plant Biol 16:17
Desaki Y, Kouzai Y, Ninomiya Y, Iwase R, Shimizu Y, Seko K, Molinaro A, Minami E, Shibuya N, Kaku H, Nishizawa Y (2017) OsCERK1 plays a crucial role in the lipopolysaccharide-induced immune response of rice. New Phytol 217:1042–1049
Desclos-Theveniau M, Arnaud D, Huang TY, Lin GJC, Chen WY, Lin YC, Zimmerli L (2012) The Arabidopsis lectin receptor kinase LecRK-V.5 represses stomatal immunity induced by Pseudomonas syringae pv. tomato DC3000. PLoS Pathogens 8:e1002513
Dewdney J, Lynne Reuber T, Wildermuth MC, Devoto A, Cui J, Stutius LM, Drummond EP, Ausubel FM (2000) Three unique mutants of Arabidopsis identify eds loci required for limiting growth of a biotrophic fungal pathogen. Plant J 24:205–218
Dodds PN, Rathjen JP (2010) Plant immunity: towards an integrated view of plant–pathogen interactions. Nat Rev Genet 11:539–548
Domingos P, Prado AM, Wong A, Gehring C, Feijo JA (2015) Nitric oxide: a multitasked signaling gas in plants. Mol Plant 8:506–520
Dong J, Xiao F, Fan F, Gu L, Cang H, Martin GB, Chai J (2009) Crystal structure of the complex between pseudomonas effector AvrPtoB and the tomato Pto kinase reveals both a shared and a unique interface compared with AvrPto-Pto. Plant Cell 21:1846–1859
Duan G, Christian N, Schwachtje J, Walther D, Ebenhöh O (2013) The metabolic interplay between plants and phytopathogens. Metabolites 3:1–23
Durian G, Rahikainen M, Alegre S, Brosché M, Kangasjärvi S (2016) Protein phosphatase 2A in the regulatory network underlying biotic stress resistance in plants. Front Plant Sci 7:812
Engelsdorf T, Gigli-Bisceglia N, Veerabagu M, McKenna JF, Augstein F, van der Does D, Zipfel C, Hamann T (2017) Pattern-triggered immunity and cell wall integrity maintenance jointly modulate plant stress responses. Biorxiv 130013
Epple P, Apel K, Bohlmann H (1995) An Arabidopsis thaliana thionin gene is inducible via a signal transduction pathway different from that for pathogenesis-related proteins. Plant Physiol 109:813–820
Erbs G, Molinaro A, Dow JM, Newman M-A (2010) Lipopolysaccharides and plant innate immunity. Subcell Biochem 53:387–403
Faizal A, Geelen D (2013) Saponins and their role in biological processes in plants. Phytochem Rev 12:877–893
Feng F, Yang F, Rong W, Wu X, Zhang J, Chen S, He C, Zhou JM (2012) A Xanthomonas uridine 5′-monophosphate transferase inhibits plant immune kinases. Nature 485:114–118
Ferrari S (2013) Oligogalacturonides: plant damage-associated molecular patterns and regulators of growth and development. Front Plant Sci 4:49
Fiers M, Lognay G, Fauconnier ML, Jijakli MH (2013) Volatile compound-mediated interactions between barley and pathogenic fungi in the soil. PLoS ONE 8: e66805
Frost CJ, Appel HM, Carlson JE, De Moraes CM, Mescher MC, Schultz JC (2007) Within-plant signalling via volatiles overcomes vascular constraints on systemic signalling and primes responses against herbivores. Ecol Lett 10:490–498
Fuchs Y, Saxena A, Gamble HR, Anderson JD (1989) Ethylene biosynthesis-inducing protein from cellulysin is an endoxylanase. Plant Physiol 89:138–143
Fuchs S, Grill E, Meskiene I, Schweighofer A (2013) Type 2C protein phosphatases in plants. FEBS J 280:681–693
Furukawa T, Inagaki H, Takai R, Hirai H, Che F-S (2014) Two distinct EF-Tu epitopes induce immune responses in rice and arabidopsis. Mol Plant Microbe Interactions 27:113–124
Garcia AV, Al-Yousif M, Hirt H (2012) Role of AGC kinases in plant growth and stress responses. Cell Mol Life Sci 69:3259–3267
García-Olmedo F, Molina A, Segura A, Moreno M (1995) The defensive role of nonspecific lipid-transfer proteins in plants. Trends Microbiol 3:72–74
Gimenez-Ibanez S, Hann DR, Ntoukakis V, Petutschnig E, Lipka V, Rathjen JP (2009) AvrPtoB targets the LysM receptor kinase CERK1 to promote bacterial virulence on plants. Curr Biol 19:423–429
Gimenez-Ibanez S, Boter M, Fernández-Barbero G, Chini A, Rathjen JP, Solano R (2014) The bacterial effector HopX1 targets JAZ transcriptional repressors to activate jasmonate signaling and promote infection in arabidopsis. PLoS Biol 12:e1001792
Giraldo MC, Dagdas YF, Gupta YK, Mentlak TA, Yi M, Martinez-Rocha AL, Saitoh H, Terauchi R, Talbot NJ, Valent B (2013) Two distinct secretion systems facilitate tissue invasion by the rice blast fungus Magnaporthe oryzae. Nat Commun 4:1996
Glazebrook J, Rogers EE, Ausubel FM (1996) Isolation of Arabidopsis mutants with enhanced disease susceptibility by direct screening. Genetics 143:973–982
Göhre V, Spallek T, Häweker H, Mersmann S, Mentzel T, Boller T, de Torres M, Mansfield JW, Robatzek S (2008) Plant pattern-recognition receptor FLS2 is directed for degradation by the bacterial ubiquitin ligase AvrPtoB. Curr Biol 18:1824–1832
Gómez-Gómez L, Boller T (2000) Fls2: an LRR receptor-like kinase involved in the perception of the bacterial elicitor flagellin in Arabidopsis. Mol Cell 5:1003–1011
Gottig N, Garavaglia BS, Daurelio LD, Valentine A, Gehring C, Orellano EG, Ottado J (2008) Xanthomonas axonopodis pv. citri uses a plant natriuretic peptide-like protein to modify host homeostasis. Proc Natl Acad Sci 105:18631–18636
Goverse A, Smant G (2014) The activation and suppression of plant innate immunity by parasitic nematodes. Annu Rev Phytopathol 52:243–265
Green TR, Ryan CA (1972) Wound-induced proteinase inhibitor in plant leaves: a possible defense mechanism against insects. Science 175:776–777
Groll M, Schellenberg B, Bachmann AS, Archer CR, Huber R, Powell TK, Lindow S, Kaiser M, Dudler R (2008) A plant pathogen virulence factor inhibits the eukaryotic proteasome by a novel mechanism. Nature 452:755–758
Gudesblat GE, Torres PS, Vojnov AA (2008) Xanthomonas campestris overcomes arabidopsis stomatal innate immunity through a DSF cell-to-cell signal-regulated virulence factor. Plant Physiol 149:1017–1027
Gust AA, Biswas R, Lenz HD, Rauhut T, Ranf S, Kemmerling B, Götz F, Glawischnig E, Lee J, Felix G, Nürnberger T (2007) Bacteria-derived peptidoglycans constitute pathogen-associated molecular patterns triggering innate immunity in Arabidopsis. J Biol Chem 282:32338–32348
Gust AA, Willmann R, Desaki Y, Grabherr HM, Nürnberger T (2012) Plant LysM proteins: modules mediating symbiosis and immunity. Trends Plant Sci 17:495–502
Halkier BA, Gershenzon J (2006) Biology and biochemistry of glucosinolates. Annu Rev Plant Biol 57:303–333
Hayafune M, Berisio R, Marchetti R, Silipo A, Kayama M, Desaki Y, Arima S, Squeglia F, Ruggiero A, Tokuyasu K, Molinaro A, Kaku H, Shibuya N (2014) Chitin-induced activation of immune signaling by the rice receptor CEBiP relies on a unique sandwich-type dimerization. Proc Natl Acad Sci 111:E404–E413
Heil M (2008) Indirect defence via tritrophic interactions. New Phytol 178:41–61
Heil M, Silva Bueno JC (2007) Within-plant signaling by volatiles leads to induction and priming of an indirect plant defense in nature. Proc Natl Acad Sci 104:5467–5472
Hind SR, Strickler SR, Boyle PC, Dunham DM, Bao Z, O’Doherty IM, Baccile JA, Hoki JS, Viox EG, Clarke CR, Vinatzer BA, Schroeder FC, Martin GB (2016) Tomato receptor FLAGELLIN-SENSING 3 binds flgII-28 and activates the plant immune system. Nat Plants 2:16128
Hirsch AM, Bauer WD, Bird DM, Cullimore J, Tyler B, Yoder JI (2003) Molecular signals and receptors: controlling rhizosphere interactions between plants and other organisms. Ecology 84:858–868
Hogenhout SA, Bos JIB (2011) Effector proteins that modulate plant-insect interactions. Curr Opin Plant Biol 14:422–428
Holbein J, Grundler FMW, Siddique S (2016) Plant basal resistance to nematodes: an update. J Exp Bot 67:2049–2061
Holton N, Nekrasov V, Ronald PC, Zipfel C (2015) The phylogenetically-related pattern recognition receptors EFR and XA21 recruit similar immune signaling components in monocots and dicots. PLoS Pathogens 11:e1004602
Hu K, Cao J, Zhang J, Xia F, Ke Y, Zhang H, Xie W, Liu H, Cui Y, Cao Y, Sun X, Xiao J, Li X, Zhang Q, Wang S (2017) Improvement of multiple agronomic traits by a disease resistance gene via cell wall reinforcement. Nat Plants 3:17009
Hurley B, Lee D, Mott A, Wilton M, Liu J, Liu YC, Angers S, Coaker G, Guttman DS, Desveaux D (2014) The Pseudomonas syringae type III effector HopF2 suppresses arabidopsis stomatal immunity. PLoS ONE 9:e114921
Ishida H, Vogel H (2006) Protein-peptide interaction studies demonstrate the versatility of calmodulin target protein binding. Protein Pept Lett 13:455–465
Jalmi SK, Sinha AK (2016) Functional involvement of a mitogen activated protein kinase module, OsMKK3-OsMPK7-OsWRK30 in mediating resistance against Xanthomonas oryzae in rice. Sci Rep 6:37974
Jeworutzki E, Roelfsema MRG, Anschütz U, Krol E, Elzenga JTM, Felix G, Boller T, Hedrich R, Becker D (2010) Early signaling through the Arabidopsis pattern recognition receptors FLS2 and EFR involves Ca2+-associated opening of plasma membrane anion channels. Plant J 62:367–378
Jha G, Rajeshwari R, Sonti RV (2005) Bacterial type two secretion system secreted proteins: double-edged swords for plant pathogens. Mol Plant Microbe Interact 18:891–898
Jiang S, Yao J, Ma KW, Zhou H, Song J, He SY, Ma W (2013) Bacterial effector activates jasmonate signaling by directly targeting JAZ transcriptional repressors. PLoS Pathogens 9:e1003715
Jones KM, Kobayashi H, Davies BW, Taga ME, Walker GC (2007) How rhizobial symbionts invade plants: the Sinorhizobium – Medicago model. Nat Rev Microbiol 5:619–633
Kadota Y, Goh T, Tomatsu H, Tamauchi R, Higashi K, Muto S, Kuchitsu K (2004) Cryptogein-induced initial events in tobacco BY-2 cells: pharmacological characterization of molecular relationship among cytosolic Ca2+ transients, anion efflux and production of reactive oxygen species. Plant Cell Physiol 45:160–170
Kaku H, Nishizawa Y, Ishii-Minami N, Akimoto-Tomiyama C, Dohmae N, Takio K, Minami E, Shibuya N (2006) Plant cells recognize chitin fragments for defense signaling through a plasma membrane receptor. Proc Natl Acad Sci 103:11086–11091
Kang S, Yang F, Li L, Chen H, Chen S, Zhang J (2015) The Arabidopsis transcription factor BRASSINOSTEROID INSENSITIVE1-ETHYL METHANESULFONATE-SUPPRESSOR1 is a direct substrate of MITOGEN-ACTIVATED PROTEIN KINASE6 and regulates immunity. Plant Physiol 167:1076–1086
Kärkönen A, Kuchitsu K (2015) Reactive oxygen species in cell wall metabolism and development in plants. Phytochemistry 112:22–32
Kesarwani M, Yoo J, Dong X (2007) Genetic interactions of TGA transcription factors in the regulation of pathogenesis-related genes and disease resistance in Arabidopsis. Plant Physiol 144:336–346
Kim H-S, Desveaux D, Singer AU, Patel P, Sondek J, Dangl JL (2005) The Pseudomonas syringae effector AvrRpt2 cleaves its C-terminally acylated target, RIN4, from Arabidopsis membranes to block RPM1 activation. Proc Natl Acad Sci 102:6496–6501
Kim JG, Stork W, Mudgett MB (2013) Xanthomonas type III effector XopD desumoylates tomato transcription factor SlERF4 to suppress ethylene responses and promote pathogen growth. Cell Host Microbe 13:143–154
Kinkema M, Fan W, Dong X (2000) Nuclear localization of NPR1 is required for activation of PR gene expression. Plant Cell 12:2339
Klauser D, Desurmont GA, Glauser GDS, Vallat A, Flury P, Boller T, Turlings TCJ, Bartels S (2015) The Arabidopsis Pep-PEPR system is induced by herbivore feeding and contributes to JA-mediated plant defence against herbivory. J Exp Bot 66:5327–5336
Klessig DF, Durner J, Noad R, Navarre DA, Wendehenne D, Kumar D, Zhou JM, Shah J, Zhang S, Kachroo P, Trifa Y, Pontier D, Lam E, Silva H (2000) Nitric oxide and salicylic acid signaling in plant defense. Proc of the Natl Acad Sci 97:8849–8855
Kourelis J, van der Hoorn RAL (2018) Defended to the nines: 25 years of resistance gene cloning identifies nine mechanisms for R protein function. Plant Cell 30:285–299
Kouzai Y, Nakajima K, Hayafune M, Ozawa K, Kaku H, Shibuya N, Minami E, Nishizawa Y (2014) CEBiP is the major chitin oligomer-binding protein in rice and plays a main role in the perception of chitin oligomers. Plant Mol Biol 84:519–528
Kovacs I, Durner J, Lindermayr C (2015) Crosstalk between nitric oxide and glutathione is required for NONEXPRESSOR OF PATHOGENESIS-RELATED GENES 1 (NPR1)-dependent defense signaling in Arabidopsis thaliana. New Phytol 208:860–872
Krasileva KV, Dahlbeck D, Staskawicz BJ (2010) Activation of an Arabidopsis resistance protein is specified by the in planta association of its leucine-rich repeat domain with the cognate oomycete effector. Plant Cell 22:2444–2458
Krol E, Mentzel T, Chinchilla D, Boller T, Felix G, Kemmerling B, Postel S, Arents M, Jeworutzki E, Al-Rasheid KAS, Becker D, Hedrich R (2010) Perception of the Arabidopsis danger signal peptide 1 involves the pattern recognition receptor AtPEPR1 and its close homologue AtPEPR2. J Biol Chem 285:13471–13479
Kunze G (2004) The N Terminus of bacterial elongation factor Tu elicits innate immunity in Arabidopsis plants. Plant Cell 16:3496–3507
Kurusu T, Hamada H, Sugiyama Y, Yagala T, Kadota Y, Furuichi T, Hayashi T, Umemura K, Komatsu S, Miyao A, Hirochika H, Kuchitsu K (2011) Negative feedback regulation of microbe-associated molecular pattern-induced cytosolic Ca2+ transients by protein phosphorylation. J Plant Res 124:415–424
Larkan NJ, Lydiate DJ, Parkin IAP, Nelson MN, Epp DJ, Cowling WA, Rimmer SR, Borhan MH (2013) The Brassica napus blackleg resistance gene LepR3 encodes a receptor-like protein triggered by the Leptosphaeria maculans effector AVRLM1. New Phytol 197:595–605
Lawton KA, Potter SL, Uknes S, Ryals J (1994) Acquired resistance signal transduction in Arabidopsis is Ethylene independent. Plant Cell 6:581–588
Lawton K, Weymann K, Friedrich L, Vernooij B, Uknes S, Ryals J (1995) Systemic acquired resistance in Arabidopsis requires salicylic acid but not ethylene. Mol Plant Microbe Interact 8:863–870
Lecourieux D, Ranjeva R, Pugin A (2006) Calcium in plant defence-signalling pathways: Tansley review. New Phytol 171:249–269
Lee J, Manning AJ, Wolfgeher D, Jelenska J, Cavanaugh KA, Xu H, Fernandez SM, Michelmore RW, Kron SJ, Greenberg JT (2015) Acetylation of an NB-LRR plant immune-effector complex suppresses immunity. Cell Rep 13:1670–1682
Li B, Meng X, Shan L, He P (2016) Transcriptional regulation of pattern-triggered immunity in plants. Cell Host Microbe 19:641–650
Liu B, Li J-F, Ao Y, Qu J, Li Z, Su J, Zhang Y, Liu J, Feng D, Qi K, He Y, Wang J, Wang H-B (2012) Lysin motif-containing proteins LYP4 and LYP6 play dual roles in peptidoglycan and chitin perception in rice innate immunity. Plant Cell 24:3406–3419
Liu B, Li JF, Ao Y, Li Z, Liu J, Feng D, Qi K, He Y, Zeng L, Wang J, Wang HB (2013a) OsLYP4 and OsLYP6 play critical roles in defense signal transduction. Plant Signal Behav 8:e22980
Liu J, Ding P, Sun T, Nitta Y, Dong O, Huang X, Yang W, Li X, Botella JR, Zhang Y (2013b) Heterotrimeric G proteins serve as a converging point in plant defense signaling activated by multiple receptor-like kinases. Plant Physiol 161:2146–2158
Liu T, Song T, Zhang X, Yuan H, Su L, Li W, Xu J, Liu S, Chen L, Chen T, Zhang M, Gu L, Zhang B, Dou D (2014) Unconventionally secreted effectors of two filamentous pathogens target plant salicylate biosynthesis. Nat Commun 5:4686
Lorrain S, Vailleau F, Balagué C, Roby D (2003) Lesion mimic mutants: Keys for deciphering cell death and defense pathways in plants? Trends Plant Sci 8:263–271
Lozano-Durán R, Bourdais G, He SY, Robatzek S (2014) The bacterial effector HopM1 suppresses PAMP-triggered oxidative burst and stomatal immunity. New Phytol 202:259–269
Lozano-Durán R, Robatzek S, Lozano-dur R (2015) 14-3-3 proteins in plant-pathogen interactions. Mol Plant Microbe Interact 28:511–518
Lu D, Lin W, Gao X, Wu S, Cheng C, Avila J, Heese A, Devarenne TP, He P, Shan L (2011) Direct ubiquitination of pattern recognition receptor FLS2 attenuates plant innate immunity. Science 332:1439–1442
Luna E, Pastor V, Robert J, Flors V, Mauch-Mani B, Ton J (2011) Callose deposition: a multifaceted plant defense response. Mol Plant Microbe Interact 24:183–193
Ma X, Xu G, He P, Shan L (2016) SERKing coreceptors for receptors. Trends Plant Sci 21:1017–1033
Maffei ME, Mithöfer A, Boland W (2007) Before gene expression: early events in plant-insect interaction. Trends Plant Sci 12:310–316
Malamy J, Carr JP, Klessig DF, Raskin I (1990) Salicylic acid: a likely endogenous signal in the resistance response of tobacco to viral infection. Science 250:1002–1004
Malinovsky FG, Fangel JU, Willats WGT (2014) The role of the cell wall in plant immunity. Front Plant Sci 5:178
Manosalva P, Manohar M, Von Reuss SH, Chen S, Koch A, Kaplan F, Choe A, Micikas RJ, Wang X, Kogel KH, Sternberg PW, Williamson VM, Schroeder FC, Klessig DF (2015) Conserved nematode signalling molecules elicit plant defenses and pathogen resistance. Nat Commun 6:7795
Maruta N, Trusov Y, Brenya E, Parekh U, Botella JR (2015) Membrane-localized extra-large G proteins and Gβγ of the heterotrimeric G proteins form functional complexes engaged in plant immunity in Arabidopsis. Plant Physiol 167:1004–1016
Mathieu J, Schwizer S, Martin GB (2014) Pto kinase binds two domains of AvrPtoB and its proximity to the effector E3 ligase determines if it evades degradation and activates plant immunity. PLoS Pathogens 10:e1004227
McLusky SR, Bennett MH, Beale MH, Lewis MJ, Gaskin P, Mansfield JW (1999) Cell wall alterations and localized accumulation of feruloyl-3′-methoxytyramine in onion epidermis at sites of attempted penetration by Botrytis allii are associated with actin polarisation, peroxidase activity and suppression of flavonoid biosynthesis. Plant J 17:523–534
Meindl T (2000) The bacterial elicitor flagellin activates its receptor in tomato cells according to the address-message concept. Plant Cell 12:1783–1794
Melotto M, Underwood W, Koczan J, Nomura K, He SY (2006) Plant stomata function in innate immunity against bacterial invasion. Cell 126:969–980
Melotto M, Zhang L, Oblessuc PR, He SY (2017) Stomatal defense a decade later. Plant Physiol 174:561–571
Ménard R, De Ruffray P, Fritig B, Yvin JC, Kauffmann S (2005) Defense and resistance-inducing activities in tobacco of the sulfated β-1,3 glucan PS3 and its synergistic activities with the unsulfated molecule. Plant Cell Physiol 46:1964–1972
Mendy B, Wang’ombe MW, Radakovic ZS, Holbein J, Ilyas M, Chopra D, Holton N, Zipfel C, Grundler FMW, Siddique S (2017) Arabidopsis leucine-rich repeat receptor–like kinase NILR1 is required for induction of innate immunity to parasitic nematodes. PLoS Pathogens 13:e1006284
Meng X, Zhang S (2013) MAPK cascades in plant disease resistance signaling. Annu Rev Phytopathol 51:245–266
Miedes E, Vanholme R, Boerjan W, Molina A (2014) The role of the secondary cell wall in plant resistance to pathogens. Front Plant Sci 5:358
Mithofer A, Boland W (2008) Recognition of herbivory-associated molecular patterns. Plant Physiol 146:825–831
Miya A, Albert P, Shinya T, Desaki Y, Ichimura K, Shirasu K, Narusaka Y, Kawakami N, Kaku H, Shibuya N (2007) CERK1, a LysM receptor kinase, is essential for chitin elicitor signaling in Arabidopsis. Proc Natl Acad Sci 104:19613–19618
Mou Z, Fan W, Dong X (2003) Inducers of plant systemic acquired resistance Regulate NPR1 function through redox changes. Cell 113:935–944
Mukhtar MS, McCormack ME, Argueso CT, Pajerowska-Mukhtar KM (2016) Pathogen tactics to manipulate plant cell death. Curr Biol 26:R608–R619
Murata Y, Mori IC, Munemasa S (2015) Diverse stomatal signaling and the signal integration mechanism. Annu Rev Plant Biol 66:369–392
Nitta Y, Ding P, Zhang Y (2015) Heterotrimeric G proteins in plant defense against pathogens and ABA signaling. Environ Exp Bot 114:153–158
Ntoukakis V, Balmuth AL, Mucyn TS, Gutierrez JR, Jones AME, Rathjen JP (2013) The tomato Prf complex is a molecular trap for bacterial effectors based on Pto transphosphorylation. PLoS Pathogens 9:e1003123
Nühse TS, Bottrill AR, Jones AME, Peck SC (2007) Quantitative phosphoproteomic analysis of plasma membrane proteins reveals regulatory mechanisms of plant innate immune responses. Plant J 51:931–940
Oh CS, Martin GB (2011) Tomato 14-3-3 protein TFT7 interacts with a MAP kinase kinase to regulate immunity-associated programmed cell death mediated by diverse disease resistance proteins. J Biol Chem 286:14129–14136
Oh C-S, Pedley KF, Martin GB (2010) Tomato 14-3-3 protein 7 positively regulates immunity-associated programmed cell death by enhancing protein abundance and signaling ability of MAPKKK. Plant Cell 22:260–272
Okazaki Y, Saito K (2014) Roles of lipids as signaling molecules and mitigators during stress response in plants. Plant J 79:584–596
Oldroyd GED (2013) Speak, friend, and enter: Signalling systems that promote beneficial symbiotic associations in plants. Nat Rev Microbiol 11:252–263
Oome S, Van den Ackerveken G (2014) Comparative and functional analysis of the widely occurring family of Nep1-like proteins. Mol Plant Microbe Interact 27:1081–1094
Oome S, Raaymakers TM, Cabral A, Samwel S, Böhm H, Albert I, Nürnberger T, Van den Ackerveken G (2014) Nep1-like proteins from three kingdoms of life act as a microbe-associated molecular pattern in Arabidopsis. Proc Natl Acad Sci 111:16955–16960
Ortiz D, de Guillen K, Cesari S, Chalvon V, Gracy J, Padilla A, Kroj T (2017) Recognition of the magnaporthe oryzae effector AVR-Pia by the decoy domain of the rice NLR immune receptor RGA5. Plant Cell 29:156–168
Papadopoulou K, Melton RE, Leggett M, Daniels MJ, Osbourn AE (1999) Compromised disease resistance in saponin-deficient plants. Proc Natl Acad Sci 96:12923–12928
Park CJ, Peng Y, Chen X, Dardick C, Ruan D, Bart R, Canlas PE, Ronald PC (2008) Rice XB15, a protein phosphatase 2C, negatively regulates cell death and XA21-mediated innate immunity. PLoS Biol 6:e231
Pauwels L, Barbero GF, Geerinck J, Tilleman S, Grunewald W, Pérez AC, Chico JM, Bossche RV, Sewell J, Gil E, García-Casado G, Witters E, Inzé D, Long JA, De Jaeger G, Solano R, Goossens A (2010) NINJA connects the co-repressor TOPLESS to jasmonate signalling. Nature 464:788–791
Pennell RI (1997) Programmed cell death in plants. Plant Cell 9:1157–1168
Penninckx IAMA, Thomma BPHJ, Buchala A, Métraux J-P, Broekaert WF (1998) Concomitant activation of jasmonate and ethylene response pathways is required for induction of a plant defensin gene in Arabidopsis. Plant Cell 10:2103–2113
Petutschnig EK, Jones AME, Serazetdinova L, Lipka U, Lipka V (2010) The Lysin Motif Receptor-like Kinase (LysM-RLK) CERK1 is a major chitin-binding protein in Arabidopsis thaliana and subject to chitin-induced phosphorylation. J Biol Chem 285:28902–28911
Piasecka A, Jedrzejczak-Rey N, Bednarek P (2015) Secondary metabolites in plant innate immunity: conserved function of divergent chemicals. New Phytol 206:948–964
Pieterse CMJ, Van der Does D, Zamioudis C, Leon-Reyes A, Van Wees SCM (2012) Hormonal modulation of plant immunity. Annu Rev Cell Dev Biol 28:489–521
Postma J, Liebrand TWH, Bi G, Evrard A, Bye RR, Mbengue M, Kuhn H, Joosten MHAJ, Robatzek S (2016) Avr4 promotes Cf-4 receptor-like protein association with the BAK1/SERK3 receptor-like kinase to initiate receptor endocytosis and plant immunity. New Phytol 210:627–642
Pruitt RN, Schwessinger B, Joe A, Thomas N, Liu F, Albert M, Robinson MR, Chan LJG, Luu DD, Chen H, Bahar O, Daudi A, De Vleesschauwer D, Caddell D, Zhang W, Zhao X, Li X, Heazlewood JL, Ruan D, Majumder D, Chern M, Kalbacher H, Midha S, Patil PB, Sonti RV, Petzold CJ, Liu CC, Brodbelt JS, Felix G, Ronald PC (2015) The rice immune receptor XA21 recognizes a tyrosine-sulfated protein from a Gram-negative bacterium. Sci Adv 1:e1500245
Quintana-Rodriguez E, Morales-Vargas AT, Molina-Torres J, Adame-Alvarez RM, Acosta-Gallegos JA, Heil M (2015) Plant volatiles cause direct, induced and associational resistance in common bean to the fungal pathogen Colletotrichum lindemuthianum. J Ecol 103:250–260
Ranf S, Eschen-Lippold L, Pecher P, Lee J, Scheel D (2011) Interplay between calcium signalling and early signalling elements during defence responses to microbe- or damage-associated molecular patterns. Plant J 68:100–113
Ranf S, Gisch N, Schäffer M, Illig T, Westphal L, Knirel YA, Sánchez-Carballo PM, Zähringer U, Hückelhoven R, Lee J, Scheel D (2015) A lectin S-domain receptor kinase mediates lipopolysaccharide sensing in Arabidopsis thaliana. Nat Immunol 16:426–433
Rasmussen MW, Roux M, Petersen M, Mundy J (2012) MAP kinase cascades in Arabidopsis innate immunity. Front Plant Sci 3:169
Ravensdale M, Bernoux M, Ve T, Kobe B, Thrall PH, Ellis JG, Dodds PN (2012) Intramolecular interaction influences binding of the Flax L5 and L6 resistance proteins to their AvrL567 LIgands. PLoS Pathogens 8:e1003004
Robert-Seilaniantz A, Grant M, Jones JDG (2011) Hormone crosstalk in plant disease and defense: more than just JASMONATE-SALICYLATE antagonism. Ann Rev Phytopathol 49:317–343
Ron M, Kantety R, Martin GB, Avidan N, Eshed Y, Zamir D, Avni A (2000) High-resolution linkage analysis and physical characterization of the EIX-responding locus in tomato. Theor Appl Genet 100:184–189
Ronald PC, Salmeron JM, Carland FM, Staskawicz BJ (1992) The cloned avirulence gene avrPto induces disease resistance in tomato cultivars containing the Pto resistance gene. J Bacteriol 174:1604–1611
Rooney HCE, Van’t Klooster JW, van der Hoorn RAL, Joosten MHAJ, Jones JDG, de Wit PJGM (2005) Cladosporium Avr2 inhibits tomato Rcr3 protease required for Cf-2-dependent disease resistance. Science 308:1783–1786
Rotblat B, Enshell-Seijffers D, Gershoni JM, Schuster S, Avni A (2002) Identification of an essential component of the elicitation active site of the EIX protein elicitor. Plant J 32:1049–1055
Rouxel T, Balesdent M-H (2013) From model to crop plant-pathogen interactions: cloning of the first resistance gene to Leptosphaeria maculans in Brassica napus. New Phytol 197:356–358
Russell AR, Ashfield T, Innes R (2015) Pseudomonas syringae effector AvrPphB suppresses AvrB-induced activation of RPM1, but not AvrRpm1-induced activation. Mol Plant Microbe Interact 28:727–735
Sandrock RW, VanEtten HD (1998) Fungal sensitivity to and enzymatic degradation of the phytoanticipin α-tomatine. Phytopathology 88:137–143
Santino A, Taurino M, De Domenico S, Bonsegna S, Poltronieri P, Pastor V, Flors V (2013) Jasmonate signaling in plant development and defense response to multiple (a)biotic stresses. Plant Cell Rep 32:1085–1098
Saur IML, Kadota Y, Sklenar J, Holton NJ, Smakowska E, Belkhadir Y, Zipfel C, Rathjen JP (2016) NbCSPR underlies age-dependent immune responses to bacterial cold shock protein in Nicotiana benthamiana. Proc Natl Acad Sci 113:3389–3394
Schreiber KJ, Baudin M, Hassan JA, Lewis JD (2016) Die another day: molecular mechanisms of effector-triggered immunity elicited by type III secreted effector proteins. Semin Cell Dev Biol 56:124–133
Schulz P, Herde M, Romeis T (2013) Calcium-dependent protein kinases: hubs in plant stress signaling and development. Plant Physiol 163:523–530
Segarra G, Santpere G, Elena G, Trillas I (2013) Enhanced botrytis cinerea resistance of arabidopsis plants grown in compost may be explained by increased expression of defense-related genes, as revealed by microarray analysis. PLoS ONE 8:e56075
Selote D, Kachroo A (2010) RIN4-like proteins mediate resistance protein-derived soybean defense against Pseudomonas syringae. Plant Signal Behav 5:1543–1546
Sels J, Mathys J, De Coninck BMA, Cammue BPA, De Bolle MFC (2008) Plant pathogenesis-related (PR) proteins: a focus on PR peptides. Plant Physiol Biochem 46:941–950
Seo S, Katou S, Seto H, Gomi K, Ohashi Y (2007) The mitogen-activated protein kinases WIPK and SIPK regulate the levels of jasmonic and salicylic acids in wounded tobacco plants. Plant J 49:899–909
Seybold H, Trempel F, Ranf S, Scheel D, Romeis T, Lee J (2014) Ca2+ signalling in plant immune response: From pattern recognition receptors to Ca2+ decoding mechanisms. New Phytol 204:782–790
Seybold H, Boudsocq M, Romeis T (2017) CDPK activation in PRR signaling. Methods in Molecular Biology Humana Press, New York, pp 173–183
Shah J (2005) Lipids, lipases, and lipid-modifying enzymes in plant disease resistance. Annu Rev Phytopathol 43:229–260
Shan L, He P, Li J, Heese A, Peck SC, Nürnberger T, Martin GB, Sheen J (2008) Bacterial effectors target the common signaling partner BAK1 to disrupt multiple MAMP receptor-signaling complexes and impede plant immunity. Cell Host Microbe 4:17–27
Sheard LB, Tan X, Mao H, Withers J, Ben-Nissan G, Hinds TR, Kobayashi Y, Hsu F-F, Sharon M, Browse J, He SY, Rizo J, Howe GA, Zheng N (2010) Jasmonate perception by inositol-phosphate-potentiated COI1-JAZ co-receptor. Nature 468:400–405
Shetty NP, Jørgensen HJL, Jensen JD, Collinge DB, Shetty HS (2008) Roles of reactive oxygen species in interactions between plants and pathogens. Eur J Plant Pathol 121:267–280
Shigenaga AM, Argueso CT (2016) No hormone to rule them all: Interactions of plant hormones during the responses of plants to pathogens. Semin Cell Dev Biol 56:174–189
Song Y, Chen D, Lu K, Sun Z, Zeng R (2015) Enhanced tomato disease resistance primed by arbuscular mycorrhizal fungus. Front Plant Sci 6:786
Souza CA, Li S, Lin AZ, Boutrot F, Grossmann G, Zipfel C, Somerville SC (2017) Cellulose-derived oligomers act as damage-associated molecular patterns and trigger defense-like responses. Plant Physiol 173:2383–2398
Staswick PE (2004) The oxylipin signal Jasmonic acid is activated by an enzyme that conjugates it to isoleucine in Arabidopsis. Plant Cell 16:2117–2127
Stergiopoulos I, de Wit PJGM (2009) Fungal effector proteins. Annu Rev Phytopathol 47:233–263
Stotz HU, Mitrousia GK, de Wit PJGM, Fitt BDL (2014) Effector-triggered defence against apoplastic fungal pathogens. Trends Plant Sci 19:491–500
Sun Y, Li L, Macho AP, Han Z, Hu Z, Zipfel C, Zhou JM, Chai J (2013) Structural basis for flg22-induced activation of the Arabidopsis FLS2-BAK1 immune complex. Science 342:624–628
Tamogami S, Rakwal R, Agrawal GK (2008) Interplant communication: airborne methyl jasmonate is essentially converted into JA and JA-Ile activating jasmonate signaling pathway and VOCs emission. Biochem Biophys ResCommun 376:723–727
Terras F (1995) Small cysteine-rich antifungal proteins from radish: their role in host defense. Plant Cell 7:573–588
Thaler JS, Humphrey PT, Whiteman NK (2012) Evolution of jasmonate and salicylate signal crosstalk. Trends Plant Sci 17:260–270
Thines B, Katsir L, Melotto M, Niu Y, Mandaokar A, Liu G, Nomura K, He SY, Howe GA, Browse J (2007) JAZ repressor proteins are targets of the SCF(COI1) complex during jasmonate signalling. Nature 448:661–665
Thomma BPHJ, Eggermont K, Penninckx IAMA, Mauch-Mani B, Vogelsang R, Cammue BPA, Broekaert WF (1998) Separate jasmonate-dependent and salicylate-dependent defense-response pathways in Arabidopsis are essential for resistance to distinct microbial pathogens. Proc Natl Acad Sci 95:15107–15111
Thordal-Christensen H, Zhang Z, Wei Y, Collinge DB (1997) Subcellular localization of H2O2 in plants. H2O2 accumulation in papillae and hypersensitive response during the barley-powdery mildew interaction. Plant J 11:1187–1194
Toyomasu T, Usui M, Sugawara C, Otomo K, Hirose Y, Miyao A, Hirochika H, Okada K, Shimizu T, Koga J, Hasegawa M, Chuba M, Kawana Y, Kuroda M, Minami E, Mitsuhashi W, Yamane H (2014) Reverse-genetic approach to verify physiological roles of rice phytoalexins: characterization of a knockdown mutant of OsCPS4 phytoalexin biosynthetic gene in rice. Physiol Plant 150:55–62
Urano D, Jones AM (2014) Heterotrimeric G protein-coupled signaling in plants. Annu Rev Plant Biol 65:365–384
van der Hoorn RAL, Kamoun S (2008) From guard to decoy: a new model for perception of plant pathogen effectors. Plant Cell 20:2009–2017
van Loon LC, Rep M, Pieterse CMJ (2006) Significance of inducible defense-related proteins in infected plants. Annu Rev Phytopathol 44:135–162
VanEtten HD (1994) Two classes of plant antibiotics: phytoalexins versus “phytoanticipins”. Plant Cell 6:1191–1192
Voigt CA (2016) Cellulose/callose glucan networks: the key to powdery mildew resistance in plants? New Phytol 212:303–305
Wan J, Zhang X-C, Neece D, Ramonell KM, Clough S, S-y K, Stacey MG, Stacey G (2008) A LysM receptor-like kinase plays a critical role in chitin signaling and fungal resistance in Arabidopsis. Plant Cell 20:471–481
Wan J, Tanaka K, Zhang X-C, Son GH, Brechenmacher L, Nguyen THN, Stacey G (2012) LYK4, a lysin motif receptor-like kinase, is important for chitin signaling and plant innate immunity in Arabidopsis. Plant Physiol 160:396–406
Wang X (2004) Lipid signaling. Curr Opin Plant Biol 7:329–336
Wang YS, Pi LY, Chen X, Chakrabarty PK, Jiang J, De Leon AL, Liu GZ, Li L, Benny U, Oard J, Ronald PC, Song WY (2006) Rice XA21 binding protein 3 is a ubiquitin ligase required for full Xa21-mediated disease resistance. Plant Cell 18:3635–3646
Wang Y, Li J, Hou S, Wang X, Li Y, Ren D, Chen S, Tang X, Zhou J-M (2010) A Pseudomonas syringae ADP-Ribosyltransferase inhibits Arabidopsis mitogen-activated protein kinase kinases. Plant Cell 22:2033–2044
Wang S, Sun J, Fan F, Tan Z, Zou Y, Lu D (2016a) A Xanthomonas oryzae pv. oryzae effector, XopR, associates with receptor-like cytoplasmic kinases and suppresses PAMP-triggered stomatal closure. Sci China Life Sci 59:897–905
Wang WM, Liu PQ, Xu YJ, Xiao S (2016b) Protein trafficking during plant innate immunity. J Integr Plant Biol 58:284–298
Wasternack C, Hause B (2013) Jasmonates: biosynthesis, perception, signal transduction and action in plant stress response, growth and development. An update to the 2007 review in annals of botany. Ann Bot 111:1021–1058
Wasternack C, Kombrink E (2010) Jasmonates: structural requirements for lipid-derived signals active in plant stress responses and development. ACS Chem Biol 5:63–77
Weerasinghe RR, Bird DM, Allen NS (2005) Root-knot nematodes and bacterial Nod factors elicit common signal transduction events in Lotus japonicus. Proc Natl Acad Sci 102:3147–3152
Weerasinghe RR, Swanson SJ, Okada SF, Garrett MB, Kim SY, Stacey G, Boucher RC, Gilroy S, Jones AM (2009) Touch induces ATP release in Arabidopsis roots that is modulated by the heterotrimeric G-protein complex. FEBS Lett 583:2521–2526
Wernimont AK, Artz JD, Finerty P, Lin YH, Amani M, Allali-Hassani A, Senisterra G, Vedadi M, Tempel W, MacKenzie F, Chau I, Lourido S, Sibley LD, Hui R (2010) Structures of apicomplexan calcium-dependent protein kinases reveal mechanism of activation by calcium. Nat Struct Mol Biol 17:596–601
Wildermuth MC, Dewdney J, Wu G, Ausubel FM (2001) Isochorismate synthase is required to synthesize salicylic acid for plant defence. Nature 414:562–565
Williams CE, Wang B, Holsten TE, Scambray J, De Assis Goes Da Silva F, Ronald PC (1996) Markers for selection of the rice Xa21 disease resistance gene. Theor Appl Genet 93:1119–1122
Willmann R, Lajunen HM, Erbs G, Newman M-A, Kolb D, Tsuda K, Katagiri F, Fliegmann J, Bono J-J, Cullimore JV, Jehle AK, Gotz F, Kulik A, Molinaro A, Lipka V, Gust AA, Nurnberger T (2011) Arabidopsis lysin-motif proteins LYM1 LYM3 CERK1 mediate bacterial peptidoglycan sensing and immunity to bacterial infection. Proc Natl Acad Sci 108:19824–19829
Wilton M, Subramaniam R, Elmore J, Felsensteiner C, Coaker G, Desveaux D (2010) The type III effector HopF2 Pto targets Arabidopsis RIN4 protein to promote Pseudomonas syringae virulence. Proc Natl Acad Sci 107:2349–2354
Wu J, Hettenhausen C, Meldau S, Baldwin IT (2007) Herbivory rapidly activates MAPK signaling in attacked and unattacked leaf regions but not between leaves of nicotiana attenuata. Plant Cell 19:1096–1122
Wu S-J, Liu Y-S, Wu J-Y (2008) The signaling role of extracellular ATP and its dependence on Ca2+ flux in elicitation of salvia miltiorrhiza hairy root cultures. Plant Cell Physiol 49:617–624
Xiang T, Zong N, Zou Y, Wu Y, Zhang J, Xing W, Li Y, Tang X, Zhu L, Chai J, Zhou JM (2008) Pseudomonas syringae effector AvrPto blocks innate immunity by targeting receptor kinases. Curr Biol 18:74–80
Xu M, Galhano R, Wiemann P, Bueno E, Tiernan M, Wu W, Chung IM, Gershenzon J, Tudzynski B, Sesma A, Peters RJ (2012) Genetic evidence for natural product-mediated plant-plant allelopathy in rice (Oryza sativa). New Phytol 193:570–575
Yan J, Zhang C, Gu M, Bai Z, Zhang W, Qi T, Cheng Z, Peng W, Luo H, Nan F, Wang Z, Xie D (2009) The Arabidopsis CORONATINE INSENSITIVE1 protein is a jasmonate receptor. Plant Cell 21:2220–2236
Zeng L, Velásquez AC, Munkvold KR, Zhang J, Martin GB (2012) A tomato LysM receptor-like kinase promotes immunity and its kinase activity is inhibited by AvrPtoB. Plant J 69:92–103
Zhang L, Kars I, Essenstam B, Liebrand TWH, Wagemakers L, Elberse J, Tagkalaki P, Tjoitang D, van den Ackerveken G, van Kan JAL (2014) Fungal endopolygalacturonases are recognized as microbe-associated molecular patterns by the Arabidopsis receptor-like protein RESPONSIVENESS TO BOTRYTIS POLYGALACTURONASES1. Plant Physiol 164:352–364
Zheng XY, Spivey NW, Zeng W, Liu PP, Fu ZQ, Klessig DF, He SY, Dong X (2012) Coronatine promotes pseudomonas syringae virulence in plants by activating a signaling cascade that inhibits salicylic acid accumulation. Cell Host Microbe 11:587–596
Zhou J, Wu S, Chen X, Liu C, Sheen J, Shan L, He P (2014) The Pseudomonas syringae effector HopF2 suppresses Arabidopsis immunity by targeting BAK1. Plant J 77:235–245
Zhou Z, Wu Y, Yang Y, Du M, Zhang X, Guo Y, Li C, Zhou J-M (2015) An Arabidopsis plasma membrane proton ATPase modulates JA signaling and is exploited by the pseudomonas syringae effector protein AvrB for stomatal invasion. Plant Cell 27:2032–2041
Zhu Z, An F, Feng Y, Li P, Xue LAM, Jiang Z, Kim J-M, To TK, Li W, Zhang X, Yu Q, Dong Z, Chen W-Q, Seki M, Zhou J-M, Guo H (2011) Derepression of ethylene-stabilized transcription factors (EIN3/EIL1) mediates jasmonate and ethylene signaling synergy in Arabidopsis. Proc Natl Acad Sci 108:12539–12544
Zipfel C (2014) Plant pattern-recognition receptors. Trends Immunol 35:345–351
Zipfel C, Kunze G, Chinchilla D, Caniard A, Jones JDG, Boller T, Felix G (2006) Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts agrobacterium-mediated transformation. Cell 125:749–760
Zuo W, Chao Q, Zhang N, Ye J, Tan G, Li B, Xing Y, Zhang B, Liu H, Fengler KA, Zhao J, Zhao X, Chen Y, Lai J, Yan J, Xu M (2015) A maize wall-associated kinase confers quantitative resistance to head smut. Nat Genet 47:151–157
Acknowledgments
The authors acknowledge various researchers who have significantly contributed in this field, but due to lack of space, their work has not been cited in this book chapter. SG and KM are supported by fellowship from the Council of Scientific and Industrial Research (Govt. of India). GJ was supported by core research grant from the National Institute of Plant Genome Research, India, and research funding from DBT, Government of India. RVS was supported by core research grants from the National Institute of Plant Genome Research, and Centre for Cellular and Molecular Biology, India, along with research funding from DBT, ICAR, and CSIR, Government of India. RVS is also supported by a J C Bose fellowship from the Science and Engineering Research Board, Government of India.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Ghosh, S., Malukani, K.K., Chandan, R.K., Sonti, R.V., Jha, G. (2019). How Plants Respond to Pathogen Attack: Interaction and Communication. In: Sopory, S. (eds) Sensory Biology of Plants. Springer, Singapore. https://doi.org/10.1007/978-981-13-8922-1_20
Download citation
DOI: https://doi.org/10.1007/978-981-13-8922-1_20
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-8921-4
Online ISBN: 978-981-13-8922-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)