Abstract
Breast cancer is the most identified reason for death among women worldwide. New developments in the field of biomedical image processing have enabled the early and effective diagnosis of breast cancer. Therefore, this article aims at developing an effective computer-aided diagnosis (CAD) system which can precisely label the mammograms as normal, benign or malignant. In the presented scheme, compound local binary pattern (CLBP) is used to obtain the texture features from the extracted regions of interest (ROI) of mammograms. Then, principal component analysis (PCA) is used to obtain the reduced feature set. Finally, different classifiers like support vector machine (SVM), k-nearest neighbors (KNN), C4.5, artificial neural network (ANN), and Naive Bayes are utilized for classification. The proposed model is validated on two standard datasets, namely, MIAS and DDSM. Further, the proposed model’s performance is assessed in terms of different measures like classification accuracy, sensitivity, and specificity. From the result analysis, it is noticed that the proposed scheme achieves better classification accuracy as compared to the benchmark schemes.
Access provided by CONRICYT-eBooks. Download conference paper PDF
Similar content being viewed by others
Keywords
- Breast cancer
- Computer-aided diagnosis system
- Compound local binary pattern
- Principal component analysis
1 Introduction
Breast cancer is considered to be the major cause of death among women after lung cancer. It is the result of the unrestricted growth of breast cells. According to GLOBOCAN cancer survey [1] about 1.67 million new cases of breast cancer were diagnosed in the year 2012 which constituted about 25% of all the cancers. Moreover, an approximate figure of 266,120 new cases of breast cancer is anticipated in men and women in the year 2018. Early detection and treatment are necessary in order to combat the mortality rate due to breast cancer. Mammography is one of the most genuine methods for screening and detection of breast cancer as compared to other methods such as breast self-examination (BSE), surgery and clinical breast examination(CBE). It uses X-rays for analysis of breasts in order to locate suspicious lesions. It results in the formation of an X-ray image called a mammogram which is studied by a radiologist. Computer-aided diagnosis (CAD) systems assist the radiologists in the understanding of breast images in order to detect the suspicious regions. The CAD system helps in increasing the diagnostic accuracy and thus improves the mammogram interpretation rate.
Talha [2] used discrete wavelet transform (DWT) along with discrete cosine transform (DCT) for extracting features. The obtained features were classified as normal or abnormal using SVM. Beura et al. [3] used two dimensional DWT and gray level co-occurrence matrix (GLCM) for extracting the relevant features from the ROI, followed by the selection of a subset of the extracted features using F-test and t-test and used backpropagation neural network for classification. Pratiwi et al. [4] presented a classification of mammograms using radial basis function neural network (RBFNN) based on GLCM texture based features. A CAD system has been proposed by Mohamed et al. [5] wherein GLCM is used for feature extraction along with three different classifiers, namely, SVM, ANN, and KNN. Dong et al. [6] used dual contourlet transform for feature extraction and an improved KNN classifier. Reyad et al. [7] showed a comparison of statistical, local binary pattern (LBP) and multi-resolution features based on DWT and contourlet transform and SVM as a classifier. Wang et al. [8] presented a mass classification scheme which utilized hidden features of mass to expose the hidden distribution pattern. Phadke et al. [9] proposed a CAD system which utilized a combination of local and global features to find out the abnormalities in the mammograms with the help of SVM. Liu et al. [10] combined a support vector machine based recursive feature elimination technique along with normalized mutual information to eliminate singular disadvantages. Zhang et al. [11] developed an ensemble system for the classification of the region of interest as benign or malignant with the help of SVM by using mass shape features. Gedik [12] introduced a new method for extracting features based on fast finite shearlet transform and used SVM for classification. Elmoufidi et al. [13] used dynamic K-means clustering algorithm for regions of interest (ROI) detection on the mini-MIAS dataset. Hariraj et al. [14] used wiener filter for noise removal, GLCM for feature extraction and SVM and KNN for classification. From the literature, it is realized that the improvement in the modules like feature extraction, feature reduction and classification leads to improvement in the overall performance of a CAD system. There exists an enormous scope to develop an improved CAD system to correctly diagnose the mammograms. Hence, keeping this in mind, authors are motivated to propose a CAD system using the compound local binary pattern for feature extraction, principal component analysis for feature reduction and different classifiers like SVM, KNN, ANN, C4.5, and Naive Bayes. Further, as per the best knowledge of the authors, this is the first attempt to propose a CAD system with this combination (CLBP+PCA+SVM, KNN, ANN, C4.5, and Naive Bayes).
2 Proposed CAD Framework
The proposed CAD system comprises of mainly three modules, namely, feature extraction using compound local binary pattern (CLBP), feature reduction using principal component analysis (PCA) and classification using SVM, KNN, ANN, C4.5, and Naive Bayes. The complete design of the presented scheme is represented in Fig. 1.
2.1 Preprocessing and ROI Extraction
Noise and unwanted pectoral muscles are removed from the mammograms in the preprocessing stage. The mammograms are provided with information regarding the size of the abnormality. Hence to extract the ROI, a suitable cropping mechanism is used. Figures 2 and 3 represents the ROIs of the MIAS and DDSM databases respectively.
2.2 Feature Extraction Using Compound Local Binary Pattern
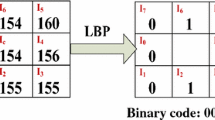
The output of a classifier is determined by the quality of the extracted features. The local binary pattern (LBP) is a simple and efficient texture feature extraction technique. However, it does not take into consideration the difference in magnitude between the center and neighboring pixel values. Therefore, this method produces conflicting results. In order to incorporate the magnitude information along with the sign, a new technique called compound local binary pattern (CLBP) which is an extension of LBP is introduced [15, 16]. CLBP allocates a code of 2P-bit to the middle pixel depending on the P number of neighboring pixels. Each of the P neighbors gets encoded with two bits. The first bit encodes the sign information while the second bit encodes the magnitude of difference with respect to a threshold value. This is illustrated in Eq. (1).
where, \(i_{m}\) is the pixel intensity of the middle pixel, \(i_{n}\) is the pixel intensity of the surrounding pixel and Avg is the average magnitude of the difference between \(i_{n}\) and \(i_{m}\) in the local neighborhood.
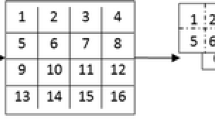
For example, in a 3\(\,\times \,\)3 neighborhood with 8 neighboring pixels, the center pixel is assigned a 16-bit code. This increases the number of features. Thus the two 8 bit patterns which are obtained by dividing the 16-bit pattern helps in reducing the number of features. The first one is generated by joining the bit values in the up, right, down, and left directions of the center pixel, respectively and the other one is formed by combining the bit values in the north-east, south-east, south-west, and north-west directions of the center pixel respectively. Figure 4 illustrates a CLBP example. Therefore, each pixel gets two 8-bit binary codes after the application of the CLBP operator on all pixels followed by dividing the obtained 16-bits into two 8-bits. Thus, two encoded images are obtained for an image from which two histograms are generated. These two histograms are then combined to obtain a histogram which serves as a feature vector for the whole image.
2.3 Feature Reduction Using Principal Component Analysis
PCA converts the features into a set of linearly uncorrelated variables called principal components [17]. It helps in reducing the dimensionality of the original feature set. It maps the data from a higher dimensionality space to a lower dimensionality space thus reducing the number of redundant features. The obtained reduced set contains maximum variability of the original data.
2.4 Classification
SVM is a supervised learning model which is used for classification and regression purposes [5]. It constructs a hyperplane that has the maximum distance from the data. ANN imitates the biological neural networks. It has an input layer, one or more hidden layers, and an output layer [5]. It is a supervised learning model. The generated output is compared with the actual output and an error (difference) is generated. Based on this error, the weights are adjusted unless and until the desired output is obtained. KNN is used for classification and regression [5]. The unknown sample is given a label which is most common among its k neighbors. C4.5 is used for generating decision trees [18]. It is an extension of ID3. It is also called a statistical classifier as the decision tree generated by it can be used for classification. Naive Bayes is based on Bayes’ theorem and is used in medical imaging [19]. It belongs to a family of probabilistic classifiers. Based on training, it classifies features and gives them labels taken from a finite set. In all the above classifiers, training is carried out with 70% data and the rest 20% data is utilized for testing.
In the proposed scheme, SVM, KNN, ANN, C4.5, and Naive Bayes are used for segregating the images into normal, benign or malignant.
3 Results
MATLAB 2017a environment is used for carrying out the experiments. All images are taken from Mammographic Image Analysis Society (MIAS) [20] and Digital Database for Screening Mammography (DDSM) [21] repositories. MIAS dataset comprises of 319 images out of which 207 are normal, 64 are benign and 48 are malignant ones. A total of 291 images are collected from DDSM dataset out of which 180 are normal, 55 are benign and 56 are malignant images. The ROIs are extracted by cropping the original images and resizing them to 256\(\,\times \,\)256. Then from each of the ROIs, texture features are extracted using CLBP. A feature vector consisting of 512 features is generated. It may be possible that all the 512 features which are extracted do not contribute towards the overall performance of the proposed model. Hence, to reduce the feature set and to curb the curse of dimensionality problem, PCA is applied which reduces the feature vector length to 20 keeping 95% variance of the original data. The reduced feature set is thus fed to different classifiers to classify the mammograms.
Table 1 lists the values of different performance metrics like accuracy (Acc), sensitivity (Sn) and specificity (Sp) obtained with the proposed model for different classifiers for MIAS dataset.
From the table, it is noticed that SVM has the highest accuracy of 100% followed by C4.5 with an accuracy of approximately 95.92%, ANN with 88.1%, and KNN and Naive Bayes both with an accuracy of 83.3856% for normal and abnormal images. In the case of Benign-Malignant, SVM has an accuracy of 100%, followed by C4.5 with an accuracy of approximately 91.07%, ANN with 80.4%, KNN with an accuracy of 76.7857%, and Naive Bayes with an accuracy of 71.4286%. Similarly, the results obtained for DDSM dataset are shown in Table 2.
It is observed that SVM and ANN both have an accuracy of 100% followed by KNN with an accuracy of 99.66%, C4.5 with an accuracy of 98.9691%, and Naive Bayes with an accuracy of 98.6254% for normal and abnormal images. In the case of Benign-Malignant, SVM has an accuracy of 100%, followed by C4.5 with an accuracy of 95.4955%, ANN with an accuracy of 93.7%, KNN with an accuracy of 81.08%, and Naive Bayes with an accuracy of 80.18%. The performance of the proposed scheme is matched with some of the recent approaches with respect to accuracy as depicted in Table 3.
4 Conclusion
Detection and diagnosis of breast cancer at an early stage helps in reducing the fatality rate to a greater extent. Hence, it becomes utmost important to develop an efficient and reliable CAD system which can classify the mammograms accurately. In this article, a model CAD system (CLBP+PCA+SVM, KNN, ANN, C4.5, and Naive Bayes) is proposed. In the presented scheme, compound local binary pattern (CLBP) which is a texture feature extraction technique is used. A total of 512 features are extracted which are then converted to a reduced feature set of size 20, with the help of PCA. The reduced feature set is fed to various classifiers like SVM, KNN, ANN, C4.5 and Naive Bayes to evaluate the performance measures.
It has been observed that SVM obtains the highest accuracy rate among all the classifiers for both Normal-Abnormal and Benign-Malignant classification. Further, it has also been observed that in the majority of the cases, the proposed model achieves better results than that of the competent schemes.
The proposed work can be extended towards the formulation of alternative feature extraction, feature reduction, and classification schemes to obtain an improved classification accuracy.
References
The International Agency for Research on Cancer: Globocan 2012: estimated cancer incidence, mortality and prevalence worldwide in 2012 (2012)
Uppal, M.T.N.: Classification of mammograms for breast cancer detection using fusion of discrete cosine transform and discrete wavelet transform features. Biomed. Res. 27(2) (2016)
Beura, S., Majhi, B., Dash, R.: Mammogram classification using two dimensional discrete wavelet transform and gray-level co-occurrence matrix for detection of breast cancer. Neurocomputing 154, 1–14 (2015)
Pratiwi, M., Harefa, J., Nanda, S.: Mammograms classification using gray-level co-occurrence matrix and radial basis function neural network. Procedia Comput. Sci. 59, 83–91 (2015)
Mohamed, H., Mabrouk, M.S., Sharawy, A.: Computer aided detection system for micro calcifications in digital mammograms. Comput. Methods Programs Biomed. 116(3), 226–235 (2014)
Dong, M., Wang, Z., Dong, C., Mu, X., Ma, Y.: Classification of region of interest in mammograms using dual contourlet transform and improved KNN. J. Sens. (2017)
Reyad, Y.A., Berbar, M.A., Hussain, M.: Comparison of statistical, LBP, and multi-resolution analysis features for breast mass classification. J. Med. Syst. 38(9), 100 (2014)
Wang, Y., Li, J., Gao, X.: Latent feature mining of spatial and marginal characteristics for mammographic mass classification. Neurocomputing 144, 107–118 (2014)
Phadke, A.C., Rege, P.P.: Fusion of local and global features for classification of abnormality in mammograms. Sādhanā 41(4), 385–395 (2016)
Liu, X., Tang, J.: Mass classification in mammograms using selected geometry and texture features, and a new SVM-based feature selection method. IEEE Syst. J. 8(3), 910–920 (2014)
Zhang, Y., Tomuro, N., Furst, J., Raicu, D.S.: Building an ensemble system for diagnosing masses in mammograms. Int. J. Comput. Assist. Radiol. Surg. 7(2), 323–329 (2012)
Gedik, N.: A new feature extraction method based on multi-resolution representations of mammograms. Appl. Soft Comput. 44, 128–133 (2016)
Elmoufidi, A., El Fahssi, K., Jai-Andaloussi, S., Sekkaki, A.: Detection of regions of interest in mammograms by using local binary pattern and dynamic k-means algorithm. Int. J. Image Video Process. Theory Appl. 1(1), 2336-0992 (2014)
Hariraj, V., Wan, K., Zunaidi, I., et al.: An efficient data mining approaches for breast cancer detection and segmentation in mammogram (2017)
Doshi, N.P.: Multi-dimensional local binary pattern texture descriptors and their application for medical image analysis. Ph.D. thesis (2014). Niraj P. Doshi
Tyagi, D., Verma, A., Sharma, S.: An improved method for facial expression recognition using hybrid approach of CLBP and Gabor filter. In: 2017 International Conference on Computing, Communication and Automation (ICCCA), pp. 1019–1024. IEEE (2017)
Buciu, I., Gacsadi, A.: Directional features for automatic tumor classification of mammogram images. Biomed. Signal Process. Control. 6(4), 370–378 (2011)
Martens, D., De Backer, M., Haesen, R., Vanthienen, J., Snoeck, M., Baesens, B.: Classification with ant colony optimization. IEEE Trans. Evol. Comput. 11(5), 651–665 (2007)
Yang, M.C., Huang, C.S., Chen, J.H., Chang, R.F.: Whole breast lesion detection using Naive Bayes classifier for portable ultrasound. Ultrasound Med. Biol. 38(11), 1870–1880 (2012)
Suckling, J., Parker, J., Dance, D., Astley, S., Hutt, I., Boggis, C., Ricketts, I., Stamatakis, E., Cerneaz, N., Kok, S.: The mammographic image analysis society digital mammogram database. Exerpta Medica. Int. Congr. Series. 1069, 375–378 (1994)
Heath, M., Bowyer, K., Kopans, D., Moore, R., Kegelmeyer, P.: The digital database for screening mammography. In: Digital mammography, pp. 431–434 (2000)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Bagchi, M.J., Mohanty, F., Rup, S., Dash, B., Majhi, B. (2018). Digital Mammogram Classification Using Compound Local Binary Pattern Features with Principal Component Analysis Based Feature Reduction Approach. In: Singh, M., Gupta, P., Tyagi, V., Flusser, J., Ören, T. (eds) Advances in Computing and Data Sciences. ICACDS 2018. Communications in Computer and Information Science, vol 905. Springer, Singapore. https://doi.org/10.1007/978-981-13-1810-8_27
Download citation
DOI: https://doi.org/10.1007/978-981-13-1810-8_27
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-1809-2
Online ISBN: 978-981-13-1810-8
eBook Packages: Computer ScienceComputer Science (R0)