Abstract
Epigenetics is one of the most rapidly expanding fields in biology over the past decades. Epigenetic mechanisms, including DNA methylation, histone modifications, and RNA-associated editing, lead to the heritable silencing/activation of genes without changes in DNA sequence. The critical role of epigenetic modifications has been demonstrated in normal and disease development in humans. With the advent of next-generation sequencing, the technological breakthrough makes it possible to unveil the genome-wide mapping of epigenetic changes. Here, we give a comprehensive overview of epigenetic mechanisms and focus on the recent progress of epigenetic modifications involved in the pathogenesis or progression of human diseases, in particular, cardiovascular diseases and cancers. In addition, some current epigenetic therapies including the inhibitors of DNA methyltransferases and histone deacetylases that have shown promising therapeutic effects will be also discussed.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
1.1 Cardiovascular Diseases
Cardiovascular diseases (CVDs) are the worldwide leading cause of human morbidity and mortality. CVDs encompass a series of pathological conditions, such as hypertension, coronary artery diseases (CADs), myocardial infarction (MI), and chronic heart failure (CHF). CHF is the final stage of most CVDs; however, recent pharmacotherapy on CHF management is limited to diuretics, renin-angiotensin-aldosterone (RAA) inhibitors, and vasodilators, targeting blood pressure, vasodilation, cardiac contractility, and hypertrophy (Swedberg et al. 2005). Despite efforts in revitalizing therapeutic management over decades, mortality remains unacceptably high with 50% of survival rate 5 years after diagnosis (Roger 2013). Moreover, after the process of thrombosis and coronary atherosclerosis, CAD could evolve into myocardial ischemia by plaque rupture or erosion (Ambrose et al. 1988). CAD is another leading cause of morbidity and mortality affecting the cardiovascular systems in developed countries. Therefore, in addition to the known mechanisms and therapies, there is clearly an unmet need for the exploration of better therapeutic targets for CVDs.
1.2 Epigenetics
The word “epigenesis” was coined by Conrad Waddington in the twentieth century to link the fields of developmental biology and genetics, which is broadly defined as how genotypes give rise to phenotypes during development (Waddington 1952). Several decades later, the term “epigenetics” was proposed as the study of mitotically and meiotically heritable changes in gene function that are not coded in DNA sequence itself (Riggs 1975; Razin and Riggs 1980). Epigenetics explores the mechanisms that make cells adapt quickly to environmental changes and build a bridge between the environment and genes. The mechanisms of heritable epigenetics have been extensively studied in different organisms and encompassed an array of molecular modifications to both DNA and chromatin, including DNA methylation, histone modification, and RNA-associated silencing, which are used to initiate and sustain epigenetic silencing (Egger et al. 2004) (Fig. 3.1). These modulators could modify the binding of transcription activators and repressors with respective gene promoters and/or change the function and conformation of chromatin, further changing the expression and function of some key genes implicated in specific signaling pathways (Gluckman et al. 2009). In general, compared with the short and flexible effect of histone modifications, DNA methylation seems to work as long-term silencers of gene expression, in which the cross talk between them exists in different mechanisms (Goldberg et al. 2007). Furthermore, the function of epigenetics has been extensively evaluated in human cancers, and recent studies have started to explore the research field of epigenetic role implicated in the development and progression of CVDs (Turunen et al. 2009; Loscalzo and Handy 2014).
1.3 DNA Methylation
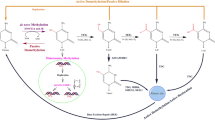
DNA methylation was proposed as the first crucial epigenetic mechanism for the switching and heritability of gene activities (Griffith and Mahler 1969; Holliday and Pugh 1975). As the most widely studied epigenetic modification, DNA methylation is defined as the addition of a methyl group (-CH3) to the carbon 5 position of cytosine ring in a CpG dinucleotide by DNA methyltransferases (DNMTs) (Jones 2001; Robertson 2005). The DNA methylation machinery is made up of two parts, DNMTs, which establish and maintain DNA methylation patterns, and the methyl-CpG-binding proteins (MBDs) family, which are candidates for the readout of DNA methylation (Robertson 2005).
MBD family is composed of five well-characterized proteins, including MeCP2, MBD1, MBD2, MBD3, and MBD4, of which, at least MBD1, MBD2, and MeCP2 could directly repress transcription (Ng et al. 1999; Ballestar and Esteller 2005). Methylation of the promoter CpG island results in the recruitment and binding of MBDs and transcription repressors including histone deacetylases (HDACs) and Sin3A, which further leads to the blockage of transcription initiation (Jones et al. 1998; Ng, Zhang et al. 1999). On the contrary, unmethylated promoters commonly lack MBD proteins with the exception of MBD1 (Lopez-Serra et al. 2006). Upon the treatment of demethylating agent, the promoter CpG islands of cancer cells will be hypomethylated and accompanied by MBD protein release and gene re-expression, suggesting the important association of MBD proteins and methylation status.
DNMT1, DNMT3a, and DNMT3b have the transcription repressor domains which can recruit HDACs and/or other corepressor proteins to DNA in a way similar to MBDs (Rountree et al. 2000; Bachman et al. 2001). Of the three DNMTs, DNMT1 is required for maintaining patterns of methylation, while DNMT3a and DNMT3b are responsible for de novo methylation and mammalian development (Okano et al. 1999; Branco et al. 2008). The maintenance DNMT1 is most abundant in the cells, which is needed to methylate hemimethylated sites that are generated during semiconservative DNA replication (Jones and Liang 2009; Portela and Esteller 2010). Different from DNMT1, DNMT3A and DNMT3B are highly expressed in embryonic stem cells and downregulated in differential cells, which might serve to establish the pattern of methylation during embryonic development (Esteller 2007). Methylation gives some explanations to phenomena in many cellular processes, such as gene imprinting or X chromosome inactivation in female mammals, both of which can be stably maintained (Reik and Walter 2001; Huynh and Lee 2005).
In normal tissues, DNA methylation occurs in 80% of repetitive DNA regions, including tandem repeats (such as satellite DNA) and dispersed repeats (such as long interspersed transposable elements (LINEs), short interspersed transposable elements (SINEs), and endogenous retroviruses) (Yoder et al. 1997; Robertson 2005). CpG islands are small concentrated regions of CpG-rich dinucleotides with the length of 0.5 to several Kb, which show elevated G+C base composition (Bird 1986). The frequency of CpG dinucleotides in human genome is lower than expected; however, approximately 70% of human gene promoters contain CpG islands, whereas some CpG islands are found within the body of genes or even 3′ region (Saxonov et al. 2006). The majority of these promoters are associated with housekeeping genes, as well as half of tissue-specific genes and development regulatory genes (Bird 1986; Larsen et al. 1992).
DNA methylation of these promoter CpG islands correlates with transcriptional repression in mammalian cells (Goll and Bestor 2005) (Fig. 3.2). Normally, the CpG islands associated with the promoters are usually unmethylated, leading to the transcription of the gene. The importance of DNA methylation for inhibition of gene expression and transcriptional silencing is apparent in the exceptions of gene imprinting or X chromosome inactivation in females (Herman and Baylin 2003). The fully methylated CpG islands leading to global gene silencing on X chromosomes and the silenced alleles of some imprinted genes are considered as a form to distinguish maternal and parental determination which are programmed in condition that only one allele of the gene is expressed in normal tissues (Antequera and Bird 1993; Bird 2002; Reik and Lewis 2005). Genomic proper patterns of methylation in mammals are essential for gene regulation and chromatin organization during embryogenesis (Goll and Bestor 2005).
1.4 Histone Modifications
Chromatin is the complex of DNA and proteins that are packaged within the nucleus of eukaryotic cells. Histones are a family of those small proteins that facilitate DNA to condense into chromatin. Importantly, the acetylation and methylation of conserved lysine residues in histone tail domain are known as the posttranslational modifications of histones. Histone acetylation is the best-understood histone modification. Of note, the critical role of acetylation in chromatin structure regulation and gene expression has been recently demonstrated (Cheung et al. 2000; Yang and Seto 2007). Most well-characterized acetylation substrates are nucleus-localized transcription factors and its coregulatory machinery (Blander and Guarente 2004; Yang and Seto 2008). These transcription repressors and activators recruit HDACs and histone acetyltransferase (HAT), respectively. Largely, HDACs lead to transcriptional inhibition by removing acetyl group, and the HATs work to increase the transcription by adding the acetyl group to the conserved lysine residue on N-terminal tail of the histone proteins (Brown et al. 2000).
HDAC
Recent evidence suggests that many HDACs consist of highly conserved domain and are subdivided into four major classes in mammals.
-
1.
Class I HDACs (HDAC1, 2, 3, and 8) comprise of domain surrounded by short NH2 and COOH termini and are widely expressed.
-
2.
Class II HDACs contain one or two catalytic sites and are further subdivided into two subclasses, including subclass IIa (HDAC4, 5, 7, and 9) and subclass IIb (HDAC6 and 10) (Yang and Seto 2008).
-
3.
Class III HDACs are sirtuins (SIRT1-SIRT7).
-
4.
Class IV is known to be homologs of Hda1 and Rpd3 protein of yeast and consists of a solitary membrane HDAC11 (Majumdar et al. 2012; Wang et al. 2014).
HAT
There are two subcategories of HATs including nuclear and cytoplasmic, which are designated as type A HATs and type B HATs, respectively (Thiagarajan et al. 2016).
Type A is transcription related and further divided into five families.
-
1.
GNAT-related acetyltransferase family represented by general control nonderepressible (GCN5), p300/CBP-associated factor (PCAF), and elongator complex protein 3 (ELP3).
-
2.
p300/CBP family represented by p300 and CBP.
-
3.
MYST family consists of MYST1 (HMOF, males absent on the first), MYST2 (HBO1, histone acetyltransferase binding to ORC), MYST3 (MOZ, monocytic leukemia zinc finger), MYST4 (MORF, monocytic leukemia zinc finger protein-related factor), and TIP60 (tat interacting protein 60 kDa).
-
4.
Basal TF family: TFIIIC (transcription factor IIIC), TAF1.
-
5.
Nuclear receptor cofactor (NRCF) family, steroid receptor coactivator (SRC), ACTR/NCOA3 (nuclear receptor coactivator 3).
Type B HATs such as HAT1, HAT2, Hat B3.1, and Rtt109 are relatively characterized.
Briefly, histone acetylation can modify gene transcription at both global acetylation level and promoter-specific acetylation level (Gottesfeld and Forbes 1997; Vaissière et al. 2008). More importantly, there is evidence showing the intimate cross talk between histone modification and DNA methylation in the process of gene silencing. The loss of histone acetylation may initiate gene silencing, which is followed by the induction of DNA methylation (Daujat et al. 2002; Vaissière et al. 2008).
1.5 Noncoding RNAs
Recently, the discovery of the noncoding RNAs (ncRNAs) provided further evidence to posttranslational mechanisms defined as epigenetic changes. It has become apparent that noncoding portion of the genome is of great importance for normal development, differentiation, and multiple diseases (Friedman and Jones 2009; Esteller 2011). Those ncRNAs involved in epigenetic processes could be categorized into the short ncRNAs (<30 nucleotides (nt)) and the long noncoding RNAs (lncRNAs) (>200 nts). The three major classes of short ncRNAs are microRNAs (miRNAs), short interfering RNAs (siRNAs), and piwi-interacting RNAs (piRNAs) (Kowalczyk et al. 2012). Both major groups are found to implicate in a wide range of cellular processes, heterochromatin formation, histone modification, and transcriptional silencing (Grewal and Moazed 2003; Moazed 2009). Here, we mainly focus on the epigenetic role of miRNAs and lncRNAs.
miRNAs belong to a class of single-stranded, small noncoding RNAs of 19–25 nts in length. miRNAs are transcribed from long RNA precursors (pri-miRNAs) by polymerase II and stabilized by 5′ capping and 3′ polyadenylation (Bartel 2009) (Fig. 3.3). The hairpin secondary structure of pri-miRNA is recognized and processed by the microprocessor complex (RNase III enzyme Drosha and DGCR8/Pasha) to release 60–80 nts pre-miRNA (Lee et al. 2003; Denli et al. 2004; Gregory et al. 2004). A pre-miRNA of the hairpin or stem-loop structure is exported into cytoplasm by Exportin 5 and further cleaved by another RNase III enzyme Dicer to generate the mature miRNA (Han et al. 2004; Kim 2005). Mature miRNAs result in translational repression or mRNA degradation of the target protein-coding genes through imperfect base pairing with the 3′ untranslated region (3′ UTR) of targeted mRNAs (Bartel 2004).
On the other hand, with the advent of high-throughput genome studies, a new class of lncRNAs measuring longer than 200 nts in length without evident protein-coding function was discovered (Martin et al. 2012). As a broad definition, lncRNA is encompassed by different classes of RNA transcripts, such as small nucleolar RNA (snoRNA), enhancer RNAs, and intergenic transcripts (Batista and Chang 2013). lncRNAs could regulate gene expression by modification of accessibility of gene promoters, which is further facilitated to tune the gene expression when adapting to the changes of environmental conditions or silence a gene during the developmental program (Batista and Chang 2013; Kornienko et al. 2013). Multiple studies have shown that the expression of miRNAs and lncRNAs is more tissue or cell specific than that of protein-coding genes (Lagos-Quintana et al. 2002; Ravasi et al. 2006; Djebali et al. 2012).
2 Epigenetics and Diseases
Gene expression patterns in different cell types should be appropriately maintained responding to environmental and developmental changes, whereas inappropriate expression might result in diseases. The important association of epigenetics with diseases is addressed by the increasing number of human diseases that are known to happen when the epigenetic information is not correctly maintained. It has become clear that aberrant regulation of DNA methylation, miRNAs, and lncRNAs contributes to the initiation or progression of human diseases (Robertson 2005; Calin and Croce 2006; Brunner et al. 2012). Further interests in this field come from the evidence of the interaction and cross talk among DNA methylation, histone modification, and small RNAs, as well as the developing pharmacological ways of reversing epigenetic abnormalities (Brown and Strathdee 2002; Tsai et al. 2010).
2.1 DNA Methylation and Diseases
2.1.1 DNA Methylation and Cancer
A large group of diseases are linked to the abnormalities of DNA methylation, showing the aberrant DNA methylation patterns and the pathological impact of their dysregulation in repeat instability. Among all diseases, the best-known studied disease is cancer. The association of methylation and cancer was firstly reported in 1983, which has been shown that cancer cells are hypomethylated compared with normal counterparts (Feinberg and Vogelstein 1983). Loss of methylation is an early and frequent event in carcinogenesis, which is generally more pronounced with cancer severity and metastatic potential in different tumor types (Widschwendter et al. 2004). For instance, age-dependent hypomethylation of specific DNA sequence was found to precede diploidy loss in a subset of gastrointestinal cancers (Suzuki et al. 2006). On the other hand, hypomethylation in cancer cells may result in the activation of oncogenes, further leading to the genomic rearrangements and instabilities (Herman and Baylin 2003). The synuclein-γ gene seems to be a target of oncogenic effect, demethylation of which serves as a molecular indicator of metastasis in a large group of human cancers, including breast cancer, ovarian cancer, and hepatocellular cancer (Gupta et al. 2003; Zhao et al. 2006).
In contrast to oncogene, biallelic loss of function in tumor suppressor genes (TSGs) is responsible for tumorigenesis as described by the Knudson’s hypothesis. This hypothesis was derived from families with hereditary cancers, in which children who suffer an inactivating germ line mutation of one allele and later a second somatic mutation of the other allele have the chance to predispose to cancer formation (Knudson 1997). Of note, essential TSGs are found to be mapped to frequent regions of homozygous deletion or loss of heterozygosity (LOH) in some human cancers, such as CDKN2A mutation in melanoma, RB1 mutation in retinoblastoma, and TP53 inactivation in Li-Fraumeni syndrome (Malkin 1993; Monzon et al. 1998; Thiagalingam et al. 2002; Wang and Chim 2015). Given that DNA methylation results in gene silencing, promoter methylation of TSGs may work as a second hit in addition to gene mutation or deletion, to fulfill the Knudson’s hypothesis in carcinogenesis (Wang and Chim 2015). Global DNA hypomethylation but gene-specific hypermethylation of promoter-associated CpG islands is a hallmark of human cancers (Esteller 2007). Interestingly, promoter CpG island methylation of TSGs has been noted in a variety of human cancers. In addition to gene silencing and loss of classic tumor suppressor function, it could also affect different cellular signaling pathways with tumor-specific profile and miRNAs with growth inhibitory role (Esteller 2007; Wang and Chim 2015). For instance, a diverse group of TSGs has been reported to be methylated in solid tumors and hematological cancers, suggesting the implication of hypermethylation in the carcinogenesis, including CDKN2A/CDKN2B regulating cell cycle in ependymal tumors, SFRP family (SFRP1/SFRP2/SFRP3/SFRP5) regulating WNT signaling in colorectal cancer and acute myeloid leukemia, and SOCS1/SOCS3/SHP1 regulating the JAK/STAT signaling pathways in hepatocellular carcinoma and multiple myeloma (Yoshikawa et al. 2001; Rousseau et al. 2003; Chim et al. 2004; Suzuki et al. 2004; Niwa et al. 2005; Valencia et al. 2009).
The genome-wide DNA hypomethylation is a consistent finding in human tumors; however, the importance and the detailed mechanism of this change remain an open question. Several animal models are used for studying the effects of changes in DNA hypomethylation implicated in carcinogenesis. Lin et al. investigated the effect of conditional inactivation of de novo methyltransferase DNMT3b in ApcMin/+ mice. The results support the role of DNMT3b in the transition stage between microadenoma formation and macroscopic colonic tumor growth, and suggest the link of DNA hypomethylation and decreased progression of microadenoma to macroscopic tumor growth (Lin et al. 2006). On the other hand, several studies show the important function of maintenance DNA methyltransferase DNMT1 involved in tumor formation. DNA hypomethylation in the transgenic ApcMin/+ mice containing a mutant DNMT1 is shown to promote early formation of hepatocellular adenomas and carcinomas but suppress overal progression of tumorigenesis, indicating the opposite effects of DNA hypomethylation in different cell types (Yamada et al. 2005). Mice carrying a hypomorphic allele of DNMT1 (DNMT1chip/−) developed thymic lymphomas with the generation of intracisternal A particle (IAP) somatic insertion in the middle of the oncogenic Notch1 genomic locus, demonstrating that DNA hypomethylation could induce tumors accompanied by the activation of endogenous retroviral elements and thus the chromosomal instabilities (Howard et al. 2008). Therefore, the different outcomes of animal models may vary from different types of cells, tissues, and tumors that are studied, the background of the animals, as well as the specific DNA sequences and chromosomal variations (Song et al. 2005; Yamada et al. 2005; Illingworth et al. 2008).
2.1.2 DNA Methylation and Atherosclerosis
The role of methylation involved in CVDs remains largely unknown. It has been noted that loss of DNA methylation is present in human atherosclerotic lesions, whereas hypermethylation of several genes is implicated in the pathogenesis of atherosclerosis, including extracellular superoxide dismutase, estrogen receptor-α (ER-α), endothelial nitric oxide synthase (eNOS), and 15-lipoxygenase (Turunen et al. 2009). Hiltunen et al. firstly demonstrated that a global genomic hypomethylation with enhanced transcriptional activity was shown during atherogenesis in advanced human, mouse, and rabbit atherosclerotic lesions, further affecting the cellular proliferation and gene expression (Hiltunen et al. 2002). Interestingly, it has been noted that in mice model during early atherosclerosis, local DNA hypermethylation and hypomethylation occurred in normohomocysteinemic mice, and the atherogenic lipoprotein profiles could influence and promote DNA hypermethylation in cultured human macrophages (Lund et al. 2004; Zaina et al. 2005). This suggested that nutritional factors could modify DNA methylation patterns that are likely to be independent of vitamin or homocysteine levels, and moreover, DNA methylation profiles may serve as early markers of atherosclerosis (Zaina et al. 2005).
Indeed, DNA methylation status could accelerate the process of atherosclerosis through upregulating atherosclerosis-susceptible genes and downregulating atherosclerosis-protective genes (Stenvinkel et al. 2007). The epigenetic changes occur in smooth muscle cells (SMCs), which constitute most of the cellular mass in the arterial wall. For instance, methylation of ER-α promoter was found in SMCs instead of endothelial cells (Post et al. 1999). ER was noted to play the role of active proliferating atheromatous SMCs, and meanwhile, it was the first gene that was well characterized for the association between aging of vascular system and the methylation of CpG islands in normal tissues in vitro (Toyota and Issa 1999; Turunen et al. 2009). Moreover, DNA methylation was also investigated to change monocarboxylate transporters (MCTs) by suppressing MCT3 expression, leading to the regulation of SMC proliferation and the development of atherosclerosis (Zhu et al. 2005). Other genes affected by aging are insulin-like growth factor 2 (IGF-2), N33, MYOD1, and cMYC (Toyota and Issa 1999; Hiltunen and Ylä-Herttuala 2003). Furthermore, inducible nitric oxide synthase (iNOS) promoter was heavily methylated in a variety of normal primary endothelial cells and vascular smooth muscle cells, whereas eNOS is hypomethylated in normal endothelium and controlled by cell-specific methylation or histone modifications (Chan et al. 2005; Fish et al. 2005). However, iNOS is expressed in endothelial cells in atherosclerotic lesions which raises the possibility that modifications in epigenetic mechanisms may lead to the changes (Shirodkar and Marsden 2011). Therefore, recent studies showed that aberrant DNA methylation occurs during atherogenesis and contributes to the lesion development and cell proliferation.
2.1.3 DNA Methylation with Cardiovascular Risk and Diseases
DNA methylation was shown to have the correlation with cardiovascular risk and mortality. The association of repetitive element LINE-1 hypomethylation and cardiovascular events, such as ischemic heart diseases and stroke, in cross-sectional and longitudinal analyses of the population has revealed that DNA hypomethylation might predict the diagnosis of CVDs before the clinical onset (Baccarelli et al. 2010). This finding showed that DNA methylation could be an effective marker that predicts the transgenerational cardiovascular risks in early life of adult diseases (Baccarelli et al. 2010). Moreover, DNA methylation was also measured and analyzed in peripheral blood leukocytes from the group of chronic kidney diseases (CKDs). It showed that inflamed patients with CKDs had global hypermethylation in peripheral blood lymphocyte DNA, suggesting the correlation of DNA methylation with increased cardiovascular mortality (Stenvinkel et al. 2007).
With regard to the DNA methylation involved in the heart failure (HF) or cardiomyopathy, there are two interesting studies supporting the essential role of DNA methylation implicated in the pathogenesis of cardiomyopathies. The global methylation profile within the CpG islands of promoters and within gene bodies was examined between human cardiac tissues from end-stage cardiomyopathic patients who had taken cardiac transplantation and normal controls. This finding showed three angiogenesis-related genes that were differentially methylated, including angiomotin-like 2 (AMOLT2), Rho GTPase-activating protein 24 (ARHGAP24), and platelet endothelial cell adhesion molecule (PECAM1) (Movassagh et al. 2010). Of note, hypermethylation in the 5′ region of PECAM1 in dilated hearts was correlated with its lower expression, and hypomethylation within the gene body of AMOLT2 was associated with reduced gene expression, whereas hypermethylation within the gene body of ARHGAP24 was correlated with increased gene expression (Movassagh et al. 2010). Therefore, the above study showed that aberrant gene expression implicated in the pathogenesis of HF and a part of the etiologies can be ascribed to the differential DNA methylation. In addition, Haas et al. also reported that an altered methylation pattern is related to dilated cardiomyopathies (DCM) compared with normal controls (Haas et al. 2013). In particular, both lymphocyte antigen 75 (LY75) and adenosine receptors A2A (ADORA2A) showed the significant downregulation in the myocardium of DCM which was mediated by promoter hypermethylation; and meanwhile, in vivo studies of LY75 and ADORA2A in zebra fish demonstrated the function role in adaptive or maladaptive pathways implicated in CVDs (Haas et al. 2013).
Nitric oxide (NO) bioavailability is implicated in the pathogenesis and progression of CVDs (Napoli and Ignarro 2009). Interestingly, it has been shown that the synthesis of NO from the precursor L-arginine could be affected by the complex process of arginine methylation pathways upon the inflammation and oxidative stress (Moncada and Higgs 1993; Rochette et al. 2013). All methylated arginine metabolites could inhibit NO synthesis in an indirect way through competitive blockade of arginine transport in CHF (Closs et al. 1997; Usui et al. 1998). Tang et al. investigated that accumulated plasma levels of methylated arginine metabolites (but not methyl-lysine) were correlated with the presence of left ventricular diastolic dysfunction in chronic systolic heart failure patients (Wilson Tang et al. 2008). Meanwhile, the close association between the methylated arginine metabolites and N-terminal pro-brain natriuretic peptide (NT-proBNP) is also confirmed in this study (Dückelmann et al. 2007; Wilson Tang et al. 2008).
2.2 Histone Modification with CVDs
2.2.1 Histone Modification with Hypoxia and Shear Stress
Hypoxia is majorly responsible for the endothelial phenotype, and the global transcription activity is reported to decline under hypoxic conditions. An ancient eukaryotic response to hypoxia is known as hypoxia-inducible factor-1 (HIF-1) transcription paradigm, and it has also been found that hypoxia induces a global drop in H3K9 acetylation via HDAC upregulation (Gluckman et al. 2008; Wadhwa et al. 2009). Across different cell lines, it has been observed that hypoxic conditions mediate an increase in the global H3K9 methylation, owing to a rise in G9 histone methyltransferase expression (Waterland and Michels 2007). Moreover, eNOS is well recognized to be of significant importance in regulating the chromatin structure in a cell-specific manner, and it is noteworthy that eNOS proximal promoter histones undergo H3 and H4 acetylation (Waterland and Jirtle 2003).
Vascular endothelium is known to endure a constant shear stress, owing to the physical force of circulation. Shear stress response element (SSRE) is a transcription factor regulating the expression of a few essential genes which are responsible for adapting to the circulatory pressure changes, such as Krüppel-like factor 2 (KLF2), VEGF receptor 2 (VEGFR2), and eNOS (Bogdarina et al. 2007; Heijmans et al. 2008; Tobi et al. 2009). Notably, it has been evidenced that the chromatin-based mechanisms, in particular global acetylation of H3 and H4 and phosphorylation of serine 10 on histone H3 (H3S10), contribute to the transcriptional regulation of these genes (El-Maarri et al. 2007).
2.2.2 Histone Modifications with CAD
As mentioned, the main cause of CADs is atherosclerosis-mediated arterial damage. It is also known that acetylation status is linked to the occurrence of atherosclerosis. HAT modulation is derived from CBP via thrombin signaling through mitogen-activated protein kinase (MAPK) pathway (Reddy and Natarajan 2011). SMCs, serving as the most abundant cell type in the arterial wall, are known to be involved in all the stages of lesion formation in atherosclerosis (Doran et al. 2008). Serum response factor (SRF) and SRF cofactor have the ability to activate CArG elements of SMC gene transcription, the binding of which is associated with methylation and acetylation to histone H3 and H4 residues (Manabe and Owens 2001; Cao et al. 2005; McDonald et al. 2006). In addition, HDAC inhibitors have shown to exacerbate the neointimal lesions in low-density lipoprotein (LDL) receptor-deficient (Ldlr−/−) mice, which are known to be atherosclerosis-prone (Choi et al. 2005). Myocardial ischemia and reperfusion injury are also heavily influenced by the HDAC activity, in particular in the case of MI (Granger et al. 2008).
2.2.3 Histone Modifications and HF
HF is regarded as the final stage of all CVDs, and the common causes include ischemic heart disease, hypertrophy, and cardiomyopathy (McMurray and Pfeffer 2005). There are many studies presenting the epigenetic regulation involved in cardiomyocytes. The expression of both class I and class II HDACs has been shown to be associated with cardiac hypertrophy (Kee and Kook 2011). In particular, class I HDACs are commonly considered as injurious to cardiac function (Montgomery et al. 2007; Xie and Hill 2013). It is known that upon the treatment of phenylephrine (PE), the hypertrophic response of cardiomyocytes could be induced with enhanced CBP/p300 transcription activity (Gusterson et al. 2002). Ectopic overexpression of CBP/p300 is shown to stimulate cardiac growth in decompensated HF (Yanazume et al. 2003). On the global level, trimethylation of histone H3 on lysine-4 (K4TM) or lysine-9 (K9TM) markers has been demonstrated to be linked with the development of HF in a rat model by high-throughput pyrosequencing performed with chromatin immunoprecipitation (ChIP) (Kaneda et al. 2009). Furthermore, the evidence showing the increase of histone acetylation only in the complication-free diabetic subjects in vivo and in vitro may signify that histone acetylation functions as a protective mechanism in the heart (Chen et al. 2009a).
2.2.4 Histone Modifications with Hypertension and Pulmonary Arterial Hypertension (PAH)
Recently, some studies have shown that core histone acetylation is responsible for hypertension and PAH (Mu et al. 2011; Yang et al. 2012; Lee et al. 2013). Environment-induced gene expression alteration in PAH is shown to be caused by certain epigenetic mechanisms (Kim et al. 2011). Therefore, the regulators of acetylation could be used as potential drug targets for treating hypertension and PAH. Valproic acid, a HDAC inhibitor, has shown to prevent the development of hypertension through diminishing transcriptional activity of mineralocorticoid receptor (MR) via increasing its acetylation (Lee et al. 2013). Moreover, HDAC inhibitor-treated spontaneously hypertensive rats (SHR) have successfully decreased vascular inflammation along with blood pressure (Usui et al. 2012). Increased activity of class I HDACs could enhance the pro-inflammatory phenotype of pulmonary adventitial fibroblasts from chronically hypoxic hypertensive calves (PH-Fibs), and therefore, the inhibition of HDACs was able to attenuate PH-Fibs functional activity to induce monocyte migration and adhesion (Li et al. 2011).
2.3 RNA Silencing with CVDs
2.3.1 miRNAs and Cardiac Aging
Aging is a predominant risk factor for CVDs, and the physiological cardiovascular aging is mainly linked with left ventricular hypertrophy, enhanced cardiac fibrosis, diastolic dysfunction, and valvular degeneration (Dai et al. 2012). A variety of miRNAs are found to be differently expressed during the process of cardiac aging with cell-type-specific characteristic (de Lucia et al. 2017). Importantly, miR-21 was the first example of miRNA which was well illustrated to play some role implicated in cardiac function. It has been demonstrated that miR-21 was upregulated selectively in fibroblasts of the failing heart compared with non-failing hearts, activating ERK-MAPK signaling by the inhibition of sprouty homologue 1 (SPRY1) (Thum et al. 2008). Pressure overload of the left ventricle was induced in mice by transverse aortic constriction (TAC), which was followed by the treatment of antagomir-21 or a control oligonucleotide. Results showed that silencing of miR-21 could reverse the expression of SPRY1 with the reduction of MAPK activity. Meanwhile, the interstitial fibrosis, cardiac hypertrophy, and dysfunction were repressed 3 weeks after in vivo silencing of miR-21 (Thum et al. 2008). Moreover, the myocardial and plasma levels of miR-21 were significantly higher in aortic stenosis (AS) patients compared with normal donors and correlated with echocardiographic mean transvalvular gradients and myocardial collagen expression (Villar et al. 2013). Furthermore, in the investigation of how myocardial ischemia affects miRNA expression, Roy et al. identified that phosphatase and tensin homologue (PTEN) is a direct target of miR-21 in isolated cardiac fibroblasts and miR-21 could modulate the expression of matrix metalloprotease-2 (MMP-2) through PTEN pathway in the infarct zone (Roy et al. 2009). A decreased expression of PTEN in infarct zone was correlated with enhanced MMP-2 expression (Roy et al. 2009).
miR-34a was another representative miRNA showing an important role in cardiovascular aging (Boon et al. 2013). Boon et al. found that the expression of miR-34a was higher in the heart tissues of C57B16 aging mice compared with young mice. In vivo silencing of miR-34a with antisense antagomir Ant-34a in 18-month-old mice decreased the dead cells, and moreover, the age-induced reduction in cardiac function was preserved in miR-34a−/− mice with less cardiomyocyte cellular death and hypertrophy when compared with wild-type controls (Boon et al. 2013). Meanwhile, miR-34a was also assessed in the mouse model of acute myocardial infarction (AMI), and it has been shown that miR-34a was induced after AMI and inhibition of miR-34a by antagomir Ant-34a or LNA-based antimiRs was able to increase cardiac contractile function after AMI (Boon et al. 2013). In addition, phosphatase 1 nuclear targeting subunit (PNUTS), which could repress telomere erosion and DNA damage response, was demonstrated as a novel target gene of miR-34a, further identifying the pro-apoptotic function of miR-34a in cardiomyocytes implicated in the aging process of CVDs (Kim et al. 2009; Landsverk et al. 2010; Boon et al. 2013).
Furthermore, Jazbutyte et al. determined the miRNA expression profiles in neonatal, 4-week, 6-month, and 19-month healthy C57/B16N mice and found that miR-22 was the most prominent upregulated miRNA accompanied by the downregulation of its direct target gene osteoglycin (OGN) during cardiac aging (Jazbutyte et al. 2013). Interestingly, OGN has been shown to be implicated in the regulation of cardiac inflammation, collagen assembly, and cardiomyocyte hypertrophy (Petretto et al. 2008; Van Aelst et al. 2014). Functionally, overexpression of miR-22 was able to accelerate cell senescence and increase the migratory activity of cardiac fibroblasts (Jazbutyte et al. 2013).
2.3.2 miRNAs and HF
Several studies have reported the potential of circulating miRNAs for the diagnosis of HF. The miRNA signature of failing myocardium is comprised of the dysregulated miRNAs such as miR-1-1, miR-195, miR-199a-1, miR-199b, and miR-221 (Akat et al. 2014). miR-1 family has been investigated as the muscle-specific miRNA and one of the essential regulating miRNAs implicated in cardiogenesis and skeletal muscle development (Lagos-Quintana et al. 2002; Zhao et al. 2007). miR-1 is highly expressed in the heart muscles, and deletion of miR-1-2 could lead to the cardiac defects of left ventricular malfunction progressing to a DCM (Zhao et al. 2007; Rao et al. 2009). Moreover, miR-1 was shown to downregulate in the early stage of cardiac hypertrophy in the hearts of C57BL/6 mice with TAC (Sayed et al. 2007). Several important regulatory genes are demonstrated as the target genes of miR-1, such as insulin-like growth factor-1 (IGF-1), twinfilin-1, a cytoskeleton regulatory protein, and heart-type fatty acid-binding protein-3 (FABP3), which are implicated in the process of cell growth, cellular differentiation or cardiac cytoskeleton, and metabolism (Elia et al. 2009; Li et al. 2010; Varrone et al. 2013).
In acute heart failure (AHF), a subset of 7 circulating miRNAs (miR-18a-5p, miR-26b-5p, miR-27a-3p, miR-30e-5p, miR-106a-5p, miR-199a-3p, and miR-652-3p) were validated to differ mostly in healthy donor controls and AHF patients (Ovchinnikova et al. 2016). In particular, AHF patients were shown to get further downregulation of the above circulating miRNAs within 48 h after admission to hospital (Ovchinnikova et al. 2016). Voellenkle et al. also investigated the miRNA expression profile in peripheral blood derived from a population of CHF patients affected by ischemic and nonischemic DCMs (Voellenkle et al. 2010). Three circulating miRNAs, including miR-107, miR-139, and miR-142-5p, were found to significantly downregulate in all HF patients when compared with healthy control donors (Voellenkle et al. 2010). Interestingly, miR-125b and miR-497 were only shown to decrease in ischemic patients, whereas miR-142-3p and miR-29b were significantly enhanced in nonischemic patients (Voellenkle et al. 2010).
2.3.3 miRNAs as Potential Biomarkers
miRNAs can be easily detected by real-time polymerase chain reaction (qRT-PCR) or microarrays in body fluid, including blood, saliva, or urine, which provides the opportunities to use these small molecules as biomarkers for human diseases (Mitchell et al. 2008; Kroh et al. 2010; Gallo et al. 2012; Tijsen et al. 2012). Importantly, cell-free blood serum and plasma miRNAs are the most stable and suitable sources for clinical application such as the diagnosis of disease given that the process of sample collection is simple and reproducible as well as the expression of miRNAs in the blood is stable even in the long term (Chim et al. 2008; Mitchell et al. 2008). Interestingly, it has been found that genetic exchange of mRNA and extracellular miRNAs between cells could be accomplished through microvesicles (MVs) or exosomes (Hunter et al. 2008). These microvesicles and exosomes containing miRNAs could facilitate the communication between cells or organs in the body. Exosomes are a population of extracellular vesicles (50–100 nm in diameter). They are released from cells in the process of inward budding of endosomal membrane that are fused with plasma membrane. Compared with exosomes, MVs are larger-sized vesicles (50–1000 nm in diameter) that are released from cells via outward budding (Raposo and Stoorvogel 2013; Lawson et al. 2016).
In CVDs, it has been noted that exosomes are manipulated by cardiovascular cells for intercellular communication as paracrine signaling mediators. Normally, the passenger strands of miRNAs, called star miRNAs (miRNA*), are considered to have no important biological function since they undergo intracellular degradation (Bartel 2004). Surprisingly, cardiac fibroblasts could secrete exosomes that are enriched with several miRNA passenger strands, in particular miR-21-3p, which has been identified to take the essential role to induce the cardiomyocyte hypertrophy (Bang et al. 2014). Moreover, the inhibition against miR-21-3p in mice with Ang II-induced cardiac hypertrophy could also reduce the pathological development of cardiac hypertrophy. Accordingly, it showed that cardiac fibroblast-derived miR-21-3p could serve as an important paracrine signaling effector in the process of fibroblast-derived cardiomyocyte hypertrophy, suggesting the potential therapeutic potential in HF patients in the future (Bang et al. 2014). Furthermore, Hergenreider et al. reported that endothelial-derived miR-143/145-containing vesicles could induce an atheroprotective SMC phenotype (Hergenreider et al. 2012). The shear-responsive transcription factor KLF2, which is a crucial mediator of endothelial gene expression patterns induced by atheroprotective flow, could directly bind to the promoter of miR-143/145 cluster and induce its upregulation (Dekker et al. 2006). In addition, extracellular vesicles generated from KLF2-transduced mouse endothelial cells could protect the mice from atherosclerotic lesion formation in the aorta of ApoE−/− mice, further demonstrating the protective role of KLF2 in miR-143/145-dependent manner (Hergenreider et al. 2012). Altogether, the atheroprotective stimuli could initiate the cross talk between endothelial cells and SMCs by the miRNAs-extracellular vesicle mechanisms, implying the promising therapy for atherosclerosis.
3 The Environment, Epigenetics, and CVD Risk
Today, approximately 7.6 billion people around the world from diverse natural environments live in their own lifestyle in different societies. Of note, lifestyle factors, society, and natural environment not only influence on daily living but also on the genome function through the dynamic and plastic epigenetic mechanisms including DNA methylation, histone modification, or miRNA aberrant expression (Alegría-Torres et al. 2011). As the life expectancy increases, the prevalence of chronic diseases will increase. In particular, it is estimated that by 2030, 40.5% of the population in the USA may have some forms of CVDs (Heidenreich et al. 2011). Therefore, the better understanding and early identification of how the natural, social, and personal environments modify the outcome of CVDs are highlighted for our prevention and management.
3.1 Natural Environmental Exposures
In environmental studies, it is noted that air pollution, circadian rhythm, and sunlight exposure are important factors that proved to be linked with CVDs that are partly regulated by epigenetic modifications. Firstly, exposure to air pollutants has been associated with the increased risk of CVDs (Du et al. 2016). Evidence showed that exposure level to fine metal-rich particulates (particulate matter <2.5 μm in diameter [PM2.5]) was demonstrated to be inversely correlated with the methylation of mitochondrial DNA (mtDNA) in the blood and participants with higher level of mtDNA methylation are more susceptible to heart rate viability due to the effect of PM2.5 exposure (Byun et al. 2016). Moreover, it was also shown that older participants with higher toll-like receptor 2 (TLR2) methylation were more prone to adverse autonomic effect of PM2.5 exposure (Zhong et al. 2015). Dietary modulation by flavonoids might reduce the above effect through the attenuation of TLR2 methylation. In addition, cardiovascular events, such as MI, sudden cardiac death, and pulmonary embolism, are recognized to have circadian rhythmicity with the high incidence in the morning, suggesting that chronic circadian disruption may enhance the susceptibility to CVDs (Oishi 2009). The evidence demonstrated that CLOCK, a transcription factor for circadian function, has the intrinsic HAT activity, whereas SIRT1 was shown to function as a HDAC that counteracts CLOCK activity to manage circadian chromatin remodeling (Bellet and Sassone-Corsi 2010). It has been identified that vitamin D deficiency has contributed to cardiovascular events such as MI, stroke, and HF (Nemerovski et al. 2009). More importantly, adequate vitamin D supplementation or sunlight exposure are beneficial for the prevention of CVDs (Wimalawansa 2016). Interestingly, vitamin D could be interacted with epigenome on multiple levels, including the methylation alteration of several critical genes such as vitamin D receptor (VDR) and the enzyme 25-hydroxylase (CYP2R1), as well as the histone modification of VDR proteins through HATs or HDACs (Fetahu et al. 2014).
3.2 Personal Environment
Smoking, diet, alcohol consumption, and sleeping habits are important factors that may have impacts on CVDs. Prenatal exposure to tobacco smoke increases the risk of diseases later in child’s life which is partly mediated by the hypomethylation of AluYb8 and hypermethylation of AXL and PTPRO in exposed children compared with controls (Breton et al. 2009). It suggested that lifelong effect of in utero exposures could be mediated by aberrant DNA methylation. Jiang et al. also found that persistent nicotine exposure selectively inhibited the expression of TBX5 and GATA4 by promoter hypermethylation, leading to cardiac function impairment with repressed left ventricular ejection fraction (Jiang et al. 2017). On the other hand, diet nutrition is another crucial factor influencing chronic disease susceptibility by the epigenetic mechanism. It is supported by the evidence that maternal diet habit during pregnancy could modify the DNA methylation pattern of specific genes, resulting in some phenotypic changes such as blood pressure, body weight, and coat color (Waterland and Jirtle 2003; Bogdarina et al. 2007). Calorie restriction (CR) is proved to be the most effective nutritional intervention to increase longevity and prevent chronic diseases such as CVDs, hypertension, and diabetes (Omodei and Fontana 2011). DNA methylation drift with age was demonstrated to be delayed by CR in mice and monkeys, showing that epigenetic drift is a good biomarker of life span (Maegawa et al. 2017). In addition, sleeping deprivation also increases the risk of CVDs, which may be mediated by the enhanced global methylation status that leads to systemic cell injury and uncompensated oxidative stress (Everson et al. 2005; Trivedi et al. 2017).
4 Technical Overview of Platform Used in Epigenetic Study
4.1 DNA Methylation with Regional and Genome-Wide Approach
In the genome, usually a proportion of potential target sequences could be methylated so that the distribution of methyl marks can carry and express epigenetic information through transcriptional silencing (Laird 2010). Both the genome-wide scale and regional DNA methylation profiles have critical implications for better understanding of how the specific loci of genome express under some condition and how the epigenetic aberration contributes to the pathogenesis of diseases (Robertson 2005). Recent advances in examination and quantification of DNA modifications facilitate the epigenetic changes to be connected to phenotypic consequences on an unprecedented scale (Plongthongkum et al. 2014).
With regard to the sequence-specific DNA methylation detection and analysis, methylation-dependent pre-treatment of DNA should be performed at first to reveal the presence or absence of methyl group at cytosine residues before DNA amplification or hybridization (Fraga and Esteller 2002; Laird 2003). There are three kinds of methods for methylation-dependent pre-treatments including bisulfite conversion, endonuclease digestion, and affinity enrichment (Laird 2010).
4.1.1 Bisulfite Conversion
The utility of sodium bisulfite on denatured genomic DNA is based on the function of bisulfite to selectively deaminate the unmethylated cytosine residue, providing the basis to distinguish the methylated and unmethylated CpG sites (Shapiro et al. 1970; Hayatsu 2008). The bisulfite treatment of DNA could deaminate unmethylated cytosine to uracil, leaving the methylated cytosine unchanged. Therefore, bisulfite conversion enables the application of new techniques to detect the methylation status at a particular region (Dahl and Guldberg 2003; Laird 2003; Dupont et al. 2004). Bisulfite-treated genomic DNA was at first analyzed and demonstrated by Sanger sequencing of cloned polymerase chain reaction (PCR) amplification at the single base pair level (Frommer et al. 1992; Paul and Clark 1996). Methylation-specific PCR (MSP) can detect the DNA methylation status of specific genes with high sensitivity and specificity (Herman et al. 1996).
4.1.2 Endonuclease Digestion
Restriction enzyme was found in bacteria and archaea by working as a defense mechanism against invading viruses, which was now used as a powerful tool in molecular biology (Roberts et al. 2003). Each sequence-specific restriction enzyme was accompanied by DNMTs that could protect DNA from restriction defense system through the methylation of recognition site (Laird 2010). Predicted modifications in restriction enzyme sites following bisulfite reaction and amplification of the product by the PCR were used to assess the methylation of CpG sites (Sadri and Hornsby 1996). HpaII and SmaI are the most widely used methylation-sensitive restriction enzymes for DNA methylation studies, which are not repressed by CpG methylation (Singer-Sam et al. 1990; Laird 2010).
4.1.3 Affinity Enrichment
The most recent way to enrich genomic methylated DNA is to do affinity purification and enrichment. One approach is to use MBDs with affinity for methylated CpG sites; alternatively, another way is to use monoclonal antibodies that are specific for 5-methylcytosine (5meC, in the context of denatured DNA) to immunoprecipitate methylated DNA (Cross et al. 1994; Selker et al. 2003; Weber et al. 2005). Of note, the affinity of methylated DNA was firstly identified with the rat chromosomal protein MeCP2 (Cross et al. 1994). Meanwhile, the immunoprecipitation approach, which is followed by hybridization to a tiling array or microarray, is regarded as MeDIP or mDIP (Weber et al. 2005; Keshet et al. 2006; Weber et al. 2007). Affinity-based approaches are to measure the density of methylation in a selected region (Zilberman and Henikoff 2007). Thus, the target regions with spare methylation sites might be difficult to differentiate the methylated and unmethylated sites, especially for mammalian genomes (Weber et al. 2007; Zilberman and Henikoff 2007).
4.1.4 Genome-Wide Approaches for Methylation
Over the past years, a large amount of approaches for genome-wide methylation analysis have been reported. As a type of two-dimensional electrophoresis, restriction landmark genomic scanning (RLGS) was the first technique in the genome-wide assessment of both genomic and epigenetic modifications (Fischer and Lerman 1979; Rush and Plass 2002). RLGS uses methylation-sensitive restriction enzymes to create the “landmarks”, and therefore, methylation difference could be detected by the presence of radiolabeled locus in unmethylated profile. In methylated profile, no radiolabeled RLGS fragment is identified because the methylation-sensitive enzyme fails to digest the DNA (Hatada et al. 1991; Rush and Plass 2002). RLGS analysis of different tumor samples has showed the relevant cancer-specific patterns of methylation, contributing to our understanding of the role of methylation implicated in carcinogenesis (Costello et al. 2000; Allegrucci et al. 2007). Nowadays with the advent of next-generation sequencing (NGS) techniques, the use of RLGS is replaced by the sequencing approaches. Sequence-based technique is more flexible and high-throughput to show the methylation patterns on a genome-wide scale within specific sequence since it can read allele-specific DNA methylation after restriction enzyme enrichment (Cokus et al. 2008; Laird 2010; Davey et al. 2011). However, it is restricted to the sequence library biases (Laird 2010).
4.2 Approaches for Histone Modification Analysis
Chromatin state could directly regulate transcription, and the cross talk between chromatin and transcription is dynamic in cellular process (Berger 2007). Importantly, histone modifications are crucial epigenetic mechanisms for eukaryotic gene regulation. ChIP assay serves as the primary powerful technique to investigate the defined genomic locations associated with histone modifications within cells or tissues (Spencer et al. 2003). In standard ChIP protocol, chromatin is released from lysed cells, digested into fragments and then selectively bound with specific antibodies against the protein of interest (Geng et al. 2011). The chromatin-antibody complex is removed, and then ChIP DNA could be further purified and analyzed by a series of approaches, such as conventional PCR, quantitative PCR, hybridization to array (ChIP-chip), or sequencing (Milne et al. 2009). Standard ChIP procedure is subjected to the multiple steps, the size of cell population, as well as the inefficient immunoprecipitation. The introduction of ChIP-chip enables the genome-wide profile of DNA-protein interaction (Ren et al. 2000). During recent years, ChIP coupled with high-throughput sequencing (ChIP-Seq) is increasingly used to map the high-resolution genome-scale views of protein-DNA interactions including transcription factor bindings, histone modification, DNA methylation, and nucleosome positioning (Park 2009; O’Geen et al. 2011). With wider coverage, less noise, single base pair resolution, and deeper sequencing, ChIP-Seq provides several advantages when compared to array-based ChIP-chip (Zhang et al. 2008; Park 2009). In 2007, the first comprehensive genome-wide map produced by ChIP-Seq showed 20 histone methylation markers, histone variant H2A.Z, RNA polymerase II, and CTCF (Barski et al. 2007; Mikkelsen et al. 2007). As the cost of sequencing decreases and institutional support grows, ChIP-Seq is likely to work as the main approach for exploring histone modifications in the future (Park 2009).
4.3 Approaches for miRNA Studies
Over the past years, miRNAs have been extensively studied in cellular processes, development, human diseases, and in particular, human cancers (Bartel 2009). It was reported that approximately 2600 miRNAs regulate more than 60% of human genomes involved in the regulation of cellular growth and death (Friedman et al. 2009; Shu et al. 2017). Therefore, a number of measurement platforms have been developed to examine the relative miRNA expression in different biological samples for better understanding of miRNA roles implicated in human diseases (Shingara et al. 2005; Chen et al. 2008; Willenbrock et al. 2009; Mestdagh et al. 2014). For miRNA studies, three main techniques are included including qRT-PCR, hybridization (DNA microarrays), and small RNA sequencing (Kong et al. 2009). The choice of platform depends on the specific experimental goals. It has been found that TaqMan qPCRarray might show superior sensitivity and specificity when compared with microarray (Chen et al. 2009b). However, microarray and qPCR platforms are limited to the number of probes or assays available, which in turn is subjected to the update of miRBase (Chen et al. 2009b; Mestdagh et al. 2014). In addition, by comparison of microarray (Exiqon’s LNA-based microarrays) and NGS (Illumina’s GA-II sequencing platform), microarray expression analysis might offer both high specificity and sensitivity, while NGS appears the ability to discover new sequence variants (Willenbrock et al. 2009).
5 The Application of Epigenetic Therapy to Human Diseases
Given that the epigenetic etiology may contribute to the pathogenesis of human diseases, this encourages the development of new promising “epigenetic therapy.” To date, several agents have been discovered that could modify the epigenetic information in a manner that is not specific to genes, including inhibitors of DNMTs and HDACs (Egger et al. 2004; Miyamoto and Ushijima 2005). Some of these agents are tested in clinical trials, while gene-specific approaches are under exploration (Lubbert 2000; Issa et al. 2004; Wagner et al. 2010).
5.1 Demethylation by 5-Azacytidine, Decitabine, and Zebularine
Since DNA methylation can be reversible silencing, the expression and function of TSGs, miRNAs, or lncRNAs that have undergone inactivation by DNA methylation will be restored by state-of-the-art epigenetic therapy (Wang and Chim 2015). DNMTs, especially DNMT1, are known to maintain CpG methylation (Robert et al. 2003). The DNMT1 inhibitor, 5-aza-2′-deoxycytidine (5-aza-dC, decitabine), forms covalent attachment with DNMT1, resulting in the degradation of DNMT1 and therefore DNA hypomethylation (Šorm et al. 1964; Christman 2002). Importantly, pharmacological grade 5-aza-dC has been identified to increase response rate and survival in patients with myelodysplastic syndrome (Silverman et al. 2002; Fenaux et al. 2009). Compared with 5-aza-dC used in high dose of 75–100 mg/m2/day, low-dose treatment of 5–20 mg/m2/day for chronic myelogenous leukemia patients in phase I study appeared more effective (Kantarjian et al. 2003; Issa et al. 2004). The major adverse effect of 5-aza-dC was myelosuppression and toxic, and the administration of 5-aza-dC is restricted to the short half-life and cannot be given orally (Issa et al. 2004). To further improve the stability and efficacy of 5-aza-dC, some cytidine analogs are under development (Yang et al. 2010). For instance, zebularine is a chemically stable cytidine analog, which could inhibit DNMTs and be administered orally or intraperitoneally (Cheng et al. 2003; Yoo et al. 2008). Moreover, zebularine showed the preference of depleting DNMT1 and induction of cancer-related antigens in cancer cells rather than normal fibroblasts (Cheng et al. 2004). However, the drawback is that higher concentration of zebularine is needed to get similar effect of demethylation in cells when compared with 5-aza-dC (Cheng et al. 2004).
5.2 Demethylation by Non-nucleoside Analogs
Different from cytidine analogs, non-nucleoside DNMT inhibitors, such as hydralazine, procaine, and procainamide, have limited activity of demethylation and less cytotoxicity as they are not incorporated into DNA (Chuang et al. 2005). For instance, the vasodilator agent hydralazine was shown to induce demethylating activity in human breast adenocarcinoma cells (Segura-Pacheco et al. 2006). Moreover, the inhibitory function of hydralazine with DNMTs was rationalized through the interaction of similar pharmacophore as established by deoxycytidine (Singh et al. 2009). Another approach to inhibit DNMT1 is to use the short-chain oligodeoxynucleotides. MG98 is a 20 base pair antisense oligodeoxynucleotide, which specifically binds to the 3′ UTR of human DNMT1 mRNA (Amato 2007). Preclinical data showed that a patient with renal cell carcinoma treated at 80 mg/m2 of MG98 showed partial response (Stewart et al. 2003). A number of phase I/II clinical trials are investigating the safety and efficacy of MG98 in several solid and hematopoietic cancers (Amato 2007).
5.3 HDAC and HAT Inhibition
Histones can be acetylated and deacetylated, and two opposing classes of enzymes are involved in determining the pattern of histone acetylation, HDACs and HATs. Several leukemogenic transcription factors depress the expression of specific genes through aberrant recruitment of HDAC, and consequently, HDAC-dependent abnormal transcription repression contributes to the oncogenic mechanism of acute promyelocytic leukemia and non-Hodgkin’s lymphoma (Dhordain et al. 1998; Lin et al. 1998). Acetylation results in the relaxation of chromatin structure and transcriptional activation of many genes, which might serve as a powerful mediator of tumor response to chemotherapy or adaptation environment to changes such as hypoxia (Grunstein 1997; Cao et al. 2011).
5.3.1 HDAC Inhibitors
Importantly, a number of HDAC inhibitors are found to induce cellular growth, differentiation, and/or apoptosis in transformed cells in culture or in tumors (Yoshida et al. 1990; Kijima et al. 1993; Kim et al. 1999). In particular, the cell cycle kinase inhibitor, p21WAF1, can be induced with the specific HDAC inhibitor trichostatin A (TSA) in p53-independent manner in NIH3T3 cells, which provides new insights to cancer therapy that could arrest tumor cells in p53-independent way (Xiao et al. 1999). Of note, as autophagy is found to be involved in the pathological cardiac remodeling, TSA treatment has been shown to reduce both load- and agonist-induced hypertrophic growth and depress the autophagy in Beclin 1 transgenic mice, suggesting the promising strategy of the HDAC inhibitor targeting autophagy (Cao et al. 2011).
To date, the HDAC inhibitors, vorinostat and romidepsin, have been used for the treatment of refractory cutaneous T cell lymphoma (CTCL) at clinic (Duvic et al. 2007; Piekarz et al. 2009). Approved by FDA for clinical application, either of them was demonstrated to have the single-agent clinical activity with significant and sustainable responses in CTCL patients, making an important therapeutic option for CTCL treatment (Olsen et al. 2007; Whittaker et al. 2010). At the same time, some newer compounds such as panobinostat, givinostat, and mocetinostat are currently being clinically assessed (Qiu et al. 2013; West and Johnstone 2014).
5.3.2 HAT Inhibitors
Different subtypes of HATs enable to catalyze the histone acetylation on specific lysine residues (Dekker and Haisma 2009). HAT families have been extensively studied including the GNAT family, the p300/CBP family, the MYST family, and the Rtt109 family (Hodawadekar and Marmorstein 2007; Nagy and Tora 2007). HATs are identified to implicate in several crucial cell proliferation pathways, and therefore disruption of theses enzymes leading to chromosomal translocations might be involved in the pathogenesis of leukemia (Sobulo et al. 1997; Perez-Campo et al. 2009; Chen et al. 2010). Therefore, the above findings make HAT an attractive target for cancer therapy. The first HAT inhibitor to be described for selectively targeting the HAT p300/CBP family was bisubstrate analog inhibitor (Lau et al. 2000). Spermidinyl-CoA-based HAT inhibitors are found to block DNA repair and serve as cancer-specific and radiosensitive drug (Bandyopadhyay et al. 2009). A novel inhibitor of p300 HAT activity, C646, is a relatively potent and cell-permeable small molecule, showing the ability to slow the cancer cell growth and impede intracellular histone acetylation (Bowers et al. 2010). Of note, in heart diseases, the balance of histone acetylation and deacetylation plays the role of regulating hypertrophic response of cardiomyocytes to phenylephrine (Gusterson et al. 2003). Inhibition of transcription coactivator p300 or CBP HAT reduced the cardiac hypertrophy, whereas activating them promoted hypertrophy in HAT-dependent manner (Gusterson et al. 2003).
5.4 Combined Therapies with Demethylating Drug and HDAC Inhibitors
Epigenetic drug, no matter demethylating drug or HDAC inhibitor, results in the reexpression and reactivation of tumor-suppressive genes, miRNAs, and lncRNAs, which play important roles involved in the cellular process and function (Wang and Chim 2015). Interestingly, these two inhibitors can be used together to improve its effectiveness. It has been firstly demonstrated that sequential exposure to 5-azacytidine and butyrate (HDAC inhibitor) had a strong synergistic effect to induce the β-adrenergic receptors in HeLa cells (Jahangeer et al. 1982). Moreover, the combination of 5-azacytidine and TSA (HDAC inhibitor) treatment on esophageal squamous cell carcinoma could lead to the functional reactivation of several TSGs (Yamashita et al. 2002). Moreover, DNMT1 contributed to the development of lung cancer in murine model, and importantly, simultaneous 5-azacytidine (DNMT inhibitor) and phenyl butyrate (HDAC inhibitor) treatment led to the re-expression of TSGs, indicating that the combined epigenetic therapy could be effective for preventing murine lung cancer (Belinsky et al. 2003). In addition, clinical trials are developed targeting two epigenetic modifications in cancer patients. It has shown that combined epigenetic therapy with low-dose 5-azacitidine and entinostat (HDAC inhibitor) led to the durable response in patients with recurrent metastatic non-small cell lung cancer (Juergens et al. 2011). In addition, epigenetic therapy can be used for sensitive drug-resistant tumors followed by chemotherapy or immunotherapy, showing a good response (Weber et al. 1994; Plumb et al. 2000).
Despite the advantages of epigenetic therapy, the therapeutic use is still very preliminary, and there are some concerns about the clinical application. One concern is regarding the nonspecific effect to normal cells, in which DNA methylation might enforce the inactivation of genes in normal cells or activation of oncogenes, leading to the potential mutagenicity (Jackson-Grusby et al. 1997; Liang et al. 2002; Sato et al. 2003; Egger et al. 2004). Moreover, both HDAC and methylation inhibitors are cytotoxic, which could induce the cellular apoptosis and/or cell cycle arrest by activating p21 or p53 (Karpf et al. 2001; Marks et al. 2003). Therefore, the clinical application of epigenetic therapy needs to be examined in patients to get a better understanding and response.
6 Future and Prospective
Epigenetic modifications have shown to contribute to several human diseases, including CVDs. Of them, DNA methylation is major epigenetic modification, which is more extensively studied compared with histone modification and miRNA alternations. In particular, DNA methylation of specific genes or miRNAs, as well as histone modifications and aberrant miRNA expression, have been noted in different subtypes of CVDs, suggesting the critical role of epigenetics involved in the pathogenesis of CVD. Interestingly, some environmental exposures could change the epigenetic markers, and therefore, it is promising to find epigenetic therapies to such reversible modifications. Some environmental factors including air pollution, smoking, aging, circadian rhythm, and diet nutrition, are found to be associated with epigenetic alternations in CVD patients when compared with those in normal controls (Dong et al. 2002; Oka et al. 2009; Baccarelli and Ghosh 2012). Recent studies have discovered how epigenetic mechanisms are involved in the pathogenesis of CVDs. As a result, there has been a rapid increase in epigenetic therapies for CVDs, such as the demethylating drug, HDAC inhibitors, or miRNA mimics/inhibitors. However, the difficulties of clinical application for CVDs cannot be underestimated. The dosage and adverse effect of drug, route of administration, and drug interactions, are crucial factors to be considered to avoid the toxicity. It is apparent that we are still at the beginning of understanding the substantial contribution of epigenetics to CVDs, and elucidating the complete epigenetic information map will be an exciting challenge for us and eventually result in a clearer direction to illustrating the diseases as well as the novel therapeutic concepts.
Abbreviations
- 3′ UTR:
-
3′ untranslated region
- 5-aza-dC:
-
5-aza-2′-deoxycytidine
- CADs:
-
Coronary artery diseases
- ChIP:
-
Chromatin immunoprecipitation
- CVDs:
-
Cardiovascular diseases
- DCM:
-
Dilated cardiomyopathies
- DNMTs:
-
DNA methyltransferases
- eNOS:
-
Endothelial nitric oxide synthase
- HATs:
-
Histone acetyltransferases
- HDACs:
-
Histone deacetylases
- HF:
-
Heart failure
- iNOS:
-
Inducible nitric oxide synthase
- lncRNAs:
-
Long noncoding RNAs
- MBDs:
-
Methyl-CpG-binding proteins
- miRNAs:
-
microRNAs
- NGS:
-
Next-generation sequencing
- NO:
-
Nitric oxide
- PAH:
-
Pulmonary arterial hypertension
- SMCs:
-
Smooth muscle cells
- TSGs:
-
Tumor suppressor genes
References
Akat KM, et al. Comparative RNA-sequencing analysis of myocardial and circulating small RNAs in human heart failure and their utility as biomarkers. Proc Natl Acad Sci. 2014;111(30):11151–6.
Alegría-Torres JA, et al. Epigenetics and lifestyle. Epigenomics. 2011;3:267–77.
Allegrucci C, et al. Restriction landmark genome scanning identifies culture-induced DNA methylation instability in the human embryonic stem cell epigenome. Hum Mol Genet. 2007;16(10):1253–68.
Amato RJ. Inhibition of DNA methylation by antisense oligonucleotide MG98 as cancer therapy. Clin Genitourin Cancer. 2007;5(7):422–6.
Ambrose JA, et al. Angiographic progression of coronary artery disease and the development of myocardial infarction. J Am Coll Cardiol. 1988;12(1):56–62.
Antequera F, Bird A. CpG islands. DNA methylation. New York: Springer; 1993. p. 169–85.
Baccarelli A, Ghosh S. Environmental exposures, epigenetics and cardiovascular disease. Curr Opin Clin Nutr Metab Care. 2012;15(4):323.
Baccarelli A, et al. Ischemic heart disease and stroke in relation to blood DNA methylation. Epidimiology. 2010;21(6):819.
Bachman KE, et al. Dnmt3a and Dnmt3b are transcriptional repressors that exhibit unique localization properties to heterochromatin. J Biol Chem. 2001;276(34):32282–7.
Ballestar E, Esteller M. Methyl-CpG-binding proteins in cancer: blaming the DNA methylation messenger. Biochem Cell Biol. 2005;83(3):374–84.
Bandyopadhyay K, et al. Spermidinyl-CoA-based HAT inhibitors block DNA repair and provide cancer-specific chemo-and radiosensitization. Cell Cycle. 2009;8(17):2779–88.
Bang C, et al. Cardiac fibroblast–derived microRNA passenger strand-enriched exosomes mediate cardiomyocyte hypertrophy. J Clin Invest. 2014;124(5):2136.
Barski A, et al. High-resolution profiling of histone methylations in the human genome. Cell. 2007;129(4):823–37.
Bartel D. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281–97.
Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136(2):215–33.
Batista PJ, Chang HY. Long noncoding RNAs: cellular address codes in development and disease. Cell. 2013;152(6):1298–307.
Belinsky SA, et al. Inhibition of DNA methylation and histone deacetylation prevents murine lung cancer. Cancer Res. 2003;63(21):7089–93.
Bellet MM, Sassone-Corsi P. Mammalian circadian clock and metabolism–the epigenetic link. J Cell Sci. 2010;123(22):3837–48.
Berger SL. The complex language of chromatin regulation during transcription. Nature. 2007;447(7143):407–12.
Bird AP. CpG-rich islands and the function of DNA methylation. Nature. 1986;321(6067):209–13.
Bird A. DNA methylation patterns and epigenetic memory. Genes Dev. 2002;16(1):6–21.
Blander G, Guarente L. The Sir2 family of protein deacetylases. Annu Rev Biochem. 2004;73(1):417–35.
Bogdarina I, et al. Epigenetic modification of the renin-angiotensin system in the fetal programming of hypertension. Circ Res. 2007;100(4):520–6.
Boon RA, et al. MicroRNA-34a regulates cardiac ageing and function. Nature. 2013;495(7439):107–10.
Bowers EM, et al. Virtual ligand screening of the p300/CBP histone acetyltransferase: identification of a selective small molecule inhibitor. Chem Biol. 2010;17(5):471–82.
Branco MR, et al. Safeguarding parental identity: Dnmt1 maintains imprints during epigenetic reprogramming in early embryogenesis. Genes Dev. 2008;22(12):1567–71.
Breton CV, et al. Prenatal tobacco smoke exposure affects global and gene-specific DNA methylation. Am J Respir Crit Care Med. 2009;180(5):462–7.
Brown R, Strathdee G. Epigenomics and epigenetic therapy of cancer. Trends Mol Med. 2002;8(4):S43–8.
Brown CE, et al. The many HATs of transcription coactivators. Trends Biochem Sci. 2000;25(1):15–9.
Brunner AL, et al. Transcriptional profiling of long non-coding RNAs and novel transcribed regions across a diverse panel of archived human cancers. Genome Biol. 2012;13(8):R75.
Byun HM, et al. Effects of air pollution and blood mitochondrial DNA methylation on markers of heart rate variability. J Am Heart Assoc. 2016;5(4):e003218.
Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6:857–66.
Cao D, et al. Modulation of smooth muscle gene expression by association of histone acetyltransferases and deacetylases with myocardin. Mol Cell Biol. 2005;25(1):364–76.
Cao DJ, et al. Histone deacetylase (HDAC) inhibitors attenuate cardiac hypertrophy by suppressing autophagy. Proc Natl Acad Sci. 2011;108(10):4123–8.
Chan GC, et al. Epigenetic basis for the transcriptional hyporesponsiveness of the human inducible nitric oxide synthase gene in vascular endothelial cells. J Immunol. 2005;175(6):3846–61.
Chen J, et al. Highly sensitive and specific microRNA expression profiling using BeadArray technology. Nucleic Acids Res. 2008;36(14):e87.
Chen SS, et al. Elevated plasma prostaglandins and acetylated histone in monocytes in type 1 diabetes patients. Diabet Med. 2009a;26(2):182–6.
Chen Y, et al. Reproducibility of quantitative RT-PCR array in miRNA expression profiling and comparison with microarray analysis. BMC Genomics. 2009b;10(1):407.
Chen J, et al. Leukaemogenesis: more than mutant genes. Nat Rev Cancer. 2010;10(1):23–36.
Cheng JC, et al. Inhibition of DNA methylation and reactivation of silenced genes by zebularine. J Natl Cancer Inst. 2003;95(5):399–409.
Cheng JC, et al. Preferential response of cancer cells to zebularine. Cancer Cell. 2004;6(2):151–8.
Cheung WL, et al. Acetylation and chromosomal functions. Curr Opin Cell Biol. 2000;12(3):326–33.
Chim CS, et al. SOCS1 and SHP1 hypermethylation in multiple myeloma: implications for epigenetic activation of the Jak/STAT pathway. Blood. 2004;103(12):4630–5.
Chim SS, et al. Detection and characterization of placental microRNAs in maternal plasma. Clin Chem. 2008;54(3):482–90.
Choi J-H, et al. Trichostatin a exacerbates atherosclerosis in low density lipoprotein receptor–deficient mice. Arterioscler Thromb Vasc Biol. 2005;25(11):2404–9.
Christman JK. 5-Azacytidine and 5-aza-2′-deoxycytidine as inhibitors of DNA methylation: mechanistic studies and their implications for cancer therapy. Oncogene. 2002;21(35):5483–95.
Chuang JC, et al. Comparison of biological effects of non-nucleoside DNA methylation inhibitors versus 5-aza-2′-deoxycytidine. Mol Cancer Ther. 2005;4(10):1515–20.
Closs EI, et al. Interference of L-arginine analogues with L-arginine transport mediated by the y+ carrier hCAT-2B. Nitric Oxide. 1997;1(1):65–73.
Cokus SJ, et al. Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature. 2008;452(7184):215–9.
Costello JF, et al. Aberrant CpG-island methylation has non-random and tumour-type–specific patterns. Nat Genet. 2000;24(2):132.
Cross SH, et al. Purification of CpG islands using a methylated DNA binding column. Nat Genet. 1994;6(3):236–44.
Dahl C, Guldberg P. DNA methylation analysis techniques. Biogerontology. 2003;4(4):233–50.
Dai D-F, et al. Cardiac aging: from molecular mechanisms to significance in human health and disease. Antioxid Redox Signal. 2012;16(12):1492–526.
Daujat S, et al. Crosstalk between CARM1 methylation and CBP acetylation on histone H3. Curr Biol. 2002;12(24):2090–7.
Davey JW, et al. Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nat Rev Genet. 2011;12(7):499–510.
de Lucia C, et al. microRNA in cardiovascular aging and age-related cardiovascular diseases. Front Med. 2017;4(74):74.
Dekker FJ, Haisma HJ. Histone acetyl transferases as emerging drug targets. Drug Discov Today. 2009;14(19):942–8.
Dekker RJ, et al. KLF2 provokes a gene expression pattern that establishes functional quiescent differentiation of the endothelium. Blood. 2006;107(11):4354–63.
Denli AM, et al. Processing of primary microRNAs by the microprocessor complex. Nature. 2004;432(7014):231–5.
Dhordain P, et al. The LAZ3 (BCL-6) oncoprotein recruits a SMRT/mSIN3A/histone deacetylase containing complex to mediate transcriptional repression. Nucleic Acids Res. 1998;26(20):4645–51.
Djebali S, et al. Landscape of transcription in human cells. Nature. 2012;489(7414):101–8.
Dong C, et al. DNA methylation and atherosclerosis. J Nutr. 2002;132(8):2406S–9S.
Doran AC, et al. Role of smooth muscle cells in the initiation and early progression of atherosclerosis. Arterioscler Thromb Vasc Biol. 2008;28(5):812–9.
Du Y, et al. Air particulate matter and cardiovascular disease: the epidemiological, biomedical and clinical evidence. J Thorac Dis. 2016;8(1):E8.
Dückelmann C, et al. Asymmetric dimethylarginine enhances cardiovascular risk prediction in patients with chronic heart failure. Arterioscler Thromb Vasc Biol. 2007;27(9):2037–42.
Dupont J-M, et al. De novo quantitative bisulfite sequencing using the pyrosequencing technology. Anal Biochem. 2004;333(1):119–27.
Duvic M, et al. Phase 2 trial of oral vorinostat (suberoylanilide hydroxamic acid, SAHA) for refractory cutaneous T-cell lymphoma (CTCL). Blood. 2007;109(1):31–9.
Egger G, et al. Epigenetics in human disease and prospects for epigenetic therapy. Nature. 2004;429(6990):457–63.
Elia L, et al. Reciprocal regulation of microRNA-1 and insulin-like growth factor-1 signal transduction cascade in cardiac and skeletal muscle in physiological and pathological conditions. Circulation. 2009;120(23):2377–85.
El-Maarri O, et al. Gender specific differences in levels of DNA methylation at selected loci from human total blood: a tendency toward higher methylation levels in males. Hum Genet. 2007;122(5):505–14.
Esteller M. Epigenetic gene silencing in cancer: the DNA hypermethylome. Hum Mol Genet. 2007;16(R1):R50–9.
Esteller M. Non-coding RNAs in human disease. Nat Rev Genet. 2011;12(12):861–74.
Everson CA, et al. Antioxidant defense responses to sleep loss and sleep recovery. Am J Phys Regul Integr Comp Phys. 2005;288(2):R374–83.
Feinberg AP, Vogelstein B. Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature. 1983;301(5895):89–92.
Fenaux P, et al. Efficacy of azacitidine compared with that of conventional care regimens in the treatment of higher-risk myelodysplastic syndromes: a randomised, open-label, phase III study. Lancet Oncol. 2009;10(3):223–32.
Fetahu IS, et al. Vitamin D and the epigenome. Front Physiol. 2014;5:164.
Fischer SG, Lerman LS. Length-independent separation of DNA restriction fragments in two-dimensional gel electrophoresis. Cell. 1979;16(1):191–200.
Fish JE, et al. The expression of endothelial nitric-oxide synthase is controlled by a cell-specific histone code. J Biol Chem. 2005;280(26):24824–38.
Fraga MF, Esteller M. DNA methylation: a profile of methods and applications. BioTechniques. 2002;33(3):632–49.
Friedman JM, Jones PA. MicroRNAs: critical mediators of differentiation, development and disease. Swiss Med Wkly. 2009;139(33–34):466–72.
Friedman RC, et al. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009;19(1):92–105.
Frommer M, et al. A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proc Natl Acad Sci. 1992;89(5):1827–31.
Gallo A, et al. The majority of microRNAs detectable in serum and saliva is concentrated in exosomes. PLoS One. 2012;7(3):e30679.
Geng T, et al. Histone modification analysis by chromatin immunoprecipitation from a low number of cells on a microfluidic platform. Lab Chip. 2011;11(17):2842–8.
Gluckman PD, et al. Effect of in utero and early-life conditions on adult health and disease. N Engl J Med. 2008;359(1):61–73.
Gluckman PD, et al. Epigenetic mechanisms that underpin metabolic and cardiovascular diseases. Nat Rev Endocrinol. 2009;5(7):401–8.
Goldberg AD, et al. Epigenetics: a landscape takes shape. Cell. 2007;128(4):635–8.
Goll MG, Bestor TH. Eukaryotic cytosine methyltransferases. Annu Rev Biochem. 2005;74:481–514.
Gottesfeld JM, Forbes DJ. Mitotic repression of the transcriptional machinery. Trends Biochem Sci. 1997;22(6):197–202.
Granger A, et al. Histone deacetylase inhibition reduces myocardial ischemia-reperfusion injury in mice. FASEB J. 2008;22(10):3549–60.
Gregory RI, et al. The microprocessor complex mediates the genesis of microRNAs. Nature. 2004;432(7014):235–40.
Grewal SI, Moazed D. Heterochromatin and epigenetic control of gene expression. Science. 2003;301(5634):798–802.
Griffith J, Mahler H. DNA ticketing theory of memory. Nature. 1969;223:580–2.
Grunstein M. Histone acetylation in chromatin structure and transcription. Nature. 1997;389(6649):349–52.
Gupta A, et al. Hypomethylation of the synuclein γ gene CpG island promotes its aberrant expression in breast carcinoma and ovarian carcinoma. Cancer Res. 2003;63(3):664–73.
Gusterson R, et al. The transcriptional co-activators CBP and p300 are activated via phenylephrine through the p42/p44 MAPK cascade. J Biol Chem. 2002;277(4):2517–24.
Gusterson RJ, et al. The transcriptional co-activators CREB-binding protein (CBP) and p300 play a critical role in cardiac hypertrophy that is dependent on their histone acetyltransferase activity. J Biol Chem. 2003;278(9):6838–47.
Haas J, et al. Alterations in cardiac DNA methylation in human dilated cardiomyopathy. EMBO Mol Med. 2013;5(3):413–29.
Han J, et al. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 2004;18(24):3016–27.
Hatada I, et al. A genomic scanning method for higher organisms using restriction sites as landmarks. Proc Natl Acad Sci. 1991;88(21):9523–7.
Hayatsu H. Discovery of bisulfite-mediated cytosine conversion to uracil, the key reaction for DNA methylation analysis – a personal account. Proc Jpn Acad Ser B. 2008;84(8):321–30.
Heidenreich PA, et al. Forecasting the future of cardiovascular disease in the United States. Policy Statement Am Heart Assoc. 2011;123(8):933–44.
Heijmans BT, et al. Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc Natl Acad Sci U S A. 2008;105(44):17046–9.
Hergenreider E, et al. Atheroprotective communication between endothelial cells and smooth muscle cells through miRNAs. Nat Cell Biol. 2012;14(3):249–56.
Herman JG, Baylin SB. Gene silencing in cancer in association with promoter hypermethylation. N Engl J Med. 2003;349(21):2042–54.
Herman J, et al. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci U S A. 1996;93:9821–6.
Hiltunen MO, Ylä-Herttuala S. DNA methylation, smooth muscle cells, and atherogenesis. Arterioscler Thromb Vasc Biol. 2003;23(10):1750–3.
Hiltunen MO, et al. DNA hypomethylation and methyltransferase expression in atherosclerotic lesions. Vasc Med. 2002;7(1):5–11.
Hodawadekar S, Marmorstein R. Chemistry of acetyl transfer by histone modifying enzymes: structure, mechanism and implications for effector design. Oncogene. 2007;26(37):5528–40.
Holliday R, Pugh JE. DNA modification mechanisms and gene activity during development. Science. 1975;187(4173):226–32.
Howard G, et al. Activation and transposition of endogenous retroviral elements in hypomethylation induced tumors in mice. Oncogene. 2008;27(3):404–8.
Hunter MP, et al. Detection of microRNA expression in human peripheral blood microvesicles. PLoS One. 2008;3(11):e3694.
Huynh KD, Lee JT. X-chromosome inactivation: a hypothesis linking ontogeny and phylogeny. Nat Rev Genet. 2005;6(5):410–8.
Illingworth R, et al. A novel CpG island set identifies tissue-specific methylation at developmental gene loci. PLoS Biol. 2008;6(1):e22.
Issa J-PJ, et al. Phase 1 study of low-dose prolonged exposure schedules of the hypomethylating agent 5-aza-2′-deoxycytidine (decitabine) in hematopoietic malignancies. Blood. 2004;103(5):1635–40.
Jackson-Grusby L, et al. Mutagenicity of 5-aza-2′-deoxycytidine is mediated by the mammalian DNA methyltransferase. Proc Natl Acad Sci. 1997;94(9):4681–5.
Jahangeer S, et al. β-Adrenergic receptor induction in HeLa cells: synergistic effect of 5-azacytidine and butyrate. Biochem Biophys Res Commun. 1982;108(4):1434–40.
Jazbutyte V, et al. MicroRNA-22 increases senescence and activates cardiac fibroblasts in the aging heart. Age. 2013;35(3):747–62.
Jiang X-Y, et al. Inhibition of gata4 and Tbx5 by nicotine-mediated DNA methylation in myocardial differentiation. Stem Cell Rep. 2017;8(2):290–304.
Jones PA. Cancer: death and methylation. Nature. 2001;409(6817):141–4.
Jones PA, Liang G. Rethinking how DNA methylation patterns are maintained. Nat Rev Genet. 2009;10(11):805–11.
Jones PL, et al. Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nat Genet. 1998;19(2):187–91.
Juergens RA, et al. Combination epigenetic therapy has efficacy in patients with refractory advanced non–small cell lung cancer. Cancer Discov. 2011;1(7):598–607.
Kaneda R, et al. Genome-wide histone methylation profile for heart failure. Genes Cells. 2009;14(1):69–77.
Kantarjian HM, et al. Results of decitabine (5-aza-2′ deoxycytidine) therapy in 130 patients with chronic myelogenous leukemia. Cancer. 2003;98(3):522–8.
Karpf AR, et al. Activation of the p53 DNA damage response pathway after inhibition of DNA methyltransferase by 5-aza-2′-deoxycytidine. Mol Pharmacol. 2001;59(4):751–7.
Kee HJ, Kook H. Roles and targets of class I and IIa histone deacetylases in cardiac hypertrophy. J Biomed Biotechnol. 2011;2011:928326.
Keshet I, et al. Evidence for an instructive mechanism of de novo methylation in cancer cells. Nat Genet. 2006;38(2):149–53.
Kijima M, et al. Trapoxin, an antitumor cyclic tetrapeptide, is an irreversible inhibitor of mammalian histone deacetylase. J Biol Chem. 1993;268(30):22429–35.
Kim VN. MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev Mol Cell Biol. 2005;6(5):376–85.
Kim YB, et al. Oxamflatin is a novel antitumor compound that inhibits mammalian histone deacetylase. Oncogene. 1999;18(15):2461–70.
Kim H, et al. TRF2 functions as a protein hub and regulates telomere maintenance by recognizing specific peptide motifs. Nat Struct Mol Biol. 2009;16(4):372–9.
Kim GH, et al. Epigenetic mechanisms of pulmonary hypertension. Pulm Circ. 2011;1(3):347–56.
Knudson AG. Karnofsky memorial lecture. Hereditary cancer: theme and variations. J Clin Oncol. 1997;15(10):3280–7.
Kong W, et al. Strategies for profiling microRNA expression. J Cell Physiol. 2009;218(1):22–5.
Kornienko AE, et al. Gene regulation by the act of long non-coding RNA transcription. BMC Biol. 2013;11(1):59.
Kowalczyk MS, et al. Molecular biology: RNA discrimination. Nature. 2012;482(7385):310–1.
Kroh EM, et al. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods. 2010;50(4):298–301.
Lagos-Quintana M, et al. Identification of tissue-specific microRNAs from mouse. Curr Biol. 2002;12(9):735–9.
Laird PW. The power and the promise of DNA methylation markers. Nat Rev Cancer. 2003;3(4):253.
Laird PW. Principles and challenges of genome-wide DNA methylation analysis. Nat Rev Genet. 2010;11(3):191–203.
Landsverk HB, et al. The protein phosphatase 1 regulator PNUTS is a new component of the DNA damage response. EMBO Rep. 2010;11(11):868–75.
Larsen F, et al. CpG islands as gene markers in the human genome. Genomics. 1992;13(4):1095–107.
Lau OD, et al. HATs off: selective synthetic inhibitors of the histone acetyltransferases p300 and PCAF. Mol Cell. 2000;5(3):589–95.
Lawson C, et al. Microvesicles and exosomes: new players in metabolic and cardiovascular disease. J Endocrinol. 2016;228(2):R57–71.
Lee Y, et al. The nuclear RNase III Drosha initiates microRNA processing. Nature. 2003;425(6956):415–9.
Lee HA, et al. Histone deacetylase inhibition attenuates transcriptional activity of mineralocorticoid receptor through its acetylation and prevents development of hypertension. Circ Res. 2013;112(7):1004–12.
Li Q, et al. Attenuation of microRNA-1 derepresses the cytoskeleton regulatory protein twinfilin-1 to provoke cardiac hypertrophy. J Cell Sci. 2010;123(14):2444–52.
Li M, et al. Emergence of fibroblasts with a proinflammatory epigenetically altered phenotype in severe hypoxic pulmonary hypertension. J Immunol. 2011;187(5):2711–22.
Liang G, et al. Analysis of gene induction in human fibroblasts and bladder cancer cells exposed to the methylation inhibitor 5-aza-2′-deoxycytidine. Cancer Res. 2002;62(4):961–6.
Lin RJ, et al. Role of the histone deacetylase complex in acute promyelocytic leukaemia. Nature. 1998;391(6669):811–4.
Lin H, et al. Suppression of intestinal neoplasia by deletion of Dnmt3b. Mol Cell Biol. 2006;26(8):2976–83.
Lopez-Serra L, et al. A profile of methyl-CpG binding domain protein occupancy of hypermethylated promoter CpG islands of tumor suppressor genes in human cancer. Cancer Res. 2006;66(17):8342–6.
Loscalzo J, Handy DE. Epigenetic modifications: basic mechanisms and role in cardiovascular disease (2013 Grover conference series). Pulm Circ. 2014;4(2):169–74.
Lubbert M. DNA methylation inhibitors in the treatment of leukemias, myelodysplastic syndromes and hemoglobinopathies: clinical results and possible mechanisms of action. Curr Top Microbiol Immunol. 2000;249:135.
Lund G, et al. DNA methylation polymorphisms precede any histological sign of atherosclerosis in mice lacking apolipoprotein E. J Biol Chem. 2004;279(28):29147–54.
Maegawa S, et al. Caloric restriction delays age-related methylation drift. Nat Commun. 2017;8(1):539.
Majumdar G, et al. Pan-histone deacetylase inhibitors regulate signaling pathways involved in proliferative and pro-inflammatory mechanisms in H9c2 cells. BMC Genomics. 2012;13:709.
Malkin D. p53 and the Li-Fraumeni syndrome. Cancer Genet Cytogenet. 1993;66(2):83–92.
Manabe I, Owens GK. Recruitment of serum response factor and hyperacetylation of histones at smooth muscle-specific regulatory regions during differentiation of a novel P19-derived in vitro smooth muscle differentiation system. Circ Res. 2001;88(11):1127–34.
Marks PA, et al. Histone deacetylases. Curr Opin Pharmacol. 2003;3(4):344–51.
Martin L, et al. Systematic reconstruction of RNA functional motifs with high-throughput microfluidics. Nat Methods. 2012;9(12):1192–4.
McDonald OG, et al. Control of SRF binding to CArG box chromatin regulates smooth muscle gene expression in vivo. J Clin Invest. 2006;116(1):36–48.
McMurray JJ, Pfeffer MA. Heart failure. Lancet. 2005;365(9474):1877–89.
Mestdagh P, et al. Evaluation of quantitative miRNA expression platforms in the microRNA quality control (miRQC) study. Nat Methods. 2014;11(8):809–15.
Mikkelsen TS, et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature. 2007;448(7153):553–60.
Milne TA, et al. Chromatin immunoprecipitation (ChIP) for analysis of histone modifications and chromatin-associated proteins. Methods Mol Biol (Clifton, N.J.). 2009;538:409–23.
Mitchell PS, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci. 2008;105(30):10513–8.
Miyamoto K, Ushijima T. Diagnostic and therapeutic applications of epigenetics. Jpn J Clin Oncol. 2005;35(6):293–301.
Moazed D. Small RNAs in transcriptional gene silencing and genome defence. Nature. 2009;457(7228):413–20.
Moncada S, Higgs A. The L-arginine-nitric oxide pathway. N Engl J Med. 1993;329(27):2002–12.
Montgomery RL, et al. Histone deacetylases 1 and 2 redundantly regulate cardiac morphogenesis, growth, and contractility. Genes Dev. 2007;21(14):1790–802.
Monzon J, et al. CDKN2A mutations in multiple primary melanomas. N Engl J Med. 1998;338(13):879–87.
Movassagh M, et al. Differential DNA methylation correlates with differential expression of angiogenic factors in human heart failure. PLoS One. 2010;5(1):e8564.
Mu S, et al. Epigenetic modulation of the renal beta-adrenergic-WNK4 pathway in salt-sensitive hypertension. Nat Med. 2011;17(5):573–80.
Nagy Z, Tora L. Distinct GCN5/PCAF-containing complexes function as co-activators and are involved in transcription factor and global histone acetylation. Oncogene. 2007;26(37):5341–57.
Napoli C, Ignarro LJ. Nitric oxide and pathogenic mechanisms involved in the development of vascular diseases. Arch Pharm Res. 2009;32(8):1103–8.
Nemerovski CW, et al. Vitamin D and cardiovascular disease. Pharmacother: J Hum Pharmacol Drug Ther. 2009;29(6):691–708.
Ng H-H, et al. MBD2 is a transcriptional repressor belonging to the MeCP1 histone deacetylase complex. Nat Genet. 1999;23:58–61.
Niwa Y, et al. Methylation silencing of SOCS-3 promotes cell growth and migration by enhancing JAK/STAT and FAK signalings in human hepatocellular carcinoma. Oncogene. 2005;24(42):6406–17.
O’Geen H, et al. Using ChIP-seq technology to generate high-resolution profiles of histone modifications. Epigenetics Protoc. 2011;791:265–86.
Oishi K. Plasminogen activator inhibitor-1 and the circadian clock in metabolic disorders. Clin Exp Hypertens. 2009;31(3):208–19.
Oka D, et al. The presence of aberrant DNA methylation in noncancerous esophageal mucosae in association with smoking history. Cancer. 2009;115(15):3412–26.
Okano M, et al. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell. 1999;99(3):247–57.
Olsen EA, et al. Phase IIb multicenter trial of vorinostat in patients with persistent, progressive, or treatment refractory cutaneous T-cell lymphoma. J Clin Oncol. 2007;25(21):3109–15.
Omodei D, Fontana L. Calorie restriction and prevention of age-associated chronic disease. FEBS Lett. 2011;585(11):1537–42.
Ovchinnikova ES, et al. Signature of circulating microRNAs in patients with acute heart failure. Eur J Heart Fail. 2016;18(4):414–23.
Park PJ. ChIP–seq: advantages and challenges of a maturing technology. Nat Rev Genet. 2009;10(10):669–80.
Paul CL, Clark SJ. Cytosine methylation: quantitation by automated genomic sequencing and GENESCAN analysis. BioTechniques. 1996;21(1):126–33.
Perez-Campo FM, et al. The histone acetyl transferase activity of monocytic leukemia zinc finger is critical for the proliferation of hematopoietic precursors. Blood. 2009;113(20):4866–74.
Petretto E, et al. Integrated genomic approaches implicate osteoglycin (Ogn) in the regulation of left ventricular mass. Nat Genet. 2008;40(5):546–52.
Piekarz RL, et al. Phase II multi-institutional trial of the histone deacetylase inhibitor romidepsin as monotherapy for patients with cutaneous T-cell lymphoma. J Clin Oncol. 2009;27(32):5410–7.
Plongthongkum N, et al. Advances in the profiling of DNA modifications: cytosine methylation and beyond. Nat Rev Genet. 2014;15(10):647–61.
Plumb JA, et al. Reversal of drug resistance in human tumor xenografts by 2′-deoxy-5-azacytidine-induced demethylation of the hMLH1 gene promoter. Cancer Res. 2000;60(21):6039–44.
Portela A, Esteller M. Epigenetic modifications and human disease. Nat Biotechnol. 2010;28(10):1057–68.
Post WS, et al. Methylation of the estrogen receptor gene is associated with aging and atherosclerosis in the cardiovascular system. Cardiovasc Res. 1999;43(4):985–91.
Qiu T, et al. Effects of treatment with histone deacetylase inhibitors in solid tumors: a review based on 30 clinical trials. Future Oncol. 2013;9(2):255–69.
Rao PK, et al. Loss of cardiac microRNA-mediated regulation leads to dilated cardiomyopathy and heart failure. Circ Res. 2009;105(6):585–94.
Raposo G, Stoorvogel W. Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol. 2013;200(4):373–83.
Ravasi T, et al. Experimental validation of the regulated expression of large numbers of non-coding RNAs from the mouse genome. Genome Res. 2006;16(1):11–9.
Razin A, Riggs AD. DNA methylation and gene function. Science. 1980;210(4470):604–10.
Reddy MA, Natarajan R. Epigenetic mechanisms in diabetic vascular complications. Cardiovasc Res. 2011;90(3):421–9.
Reik W, Lewis A. Co-evolution of X-chromosome inactivation and imprinting in mammals. Nat Rev Genet. 2005;6(5):403–10.
Reik W, Walter J. Genomic imprinting: parental influence on the genome. Nat Rev Genet. 2001;2(1):21–32.
Ren B, et al. Genome-wide location and function of DNA binding proteins. Science. 2000;290(5500):2306–9.
Riggs AD. X inactivation, differentiation, and DNA methylation. Cytogenet Genome Res. 1975;14(1):9–25.
Robert M-F, et al. DNMT1 is required to maintain CpG methylation and aberrant gene silencing in human cancer cells. Nat Genet. 2003;33(1):61–5.
Roberts RJ, et al. A nomenclature for restriction enzymes, DNA methyltransferases, homing endonucleases and their genes. Nucleic Acids Res. 2003;31(7):1805–12.
Robertson KD. DNA methylation and human disease. Nat Rev Genet. 2005;6(8):597–610.
Rochette L, et al. Nitric oxide synthase inhibition and oxidative stress in cardiovascular diseases: possible therapeutic targets? Pharmacol Ther. 2013;140(3):239–57.
Roger VL. Epidemiology of heart failure. Circ Res. 2013;113(6):646–59.
Rountree MR, et al. DNMT1 binds HDAC2 and a new co-repressor, DMAP1, to form a complex at replication foci. Nat Genet. 2000;25(3):269–77.
Rousseau E, et al. CDKN2A, CDKN2B and p14 ARF are frequently and differentially methylated in ependymal tumours. Neuropathol Appl Neurobiol. 2003;29(6):574–83.
Roy S, et al. MicroRNA expression in response to murine myocardial infarction: miR-21 regulates fibroblast metalloprotease-2 via phosphatase and tensin homologue. Cardiovasc Res. 2009;82(1):21–9.
Rush LJ, Plass C. Restriction landmark genomic scanning for DNA methylation in cancer: past, present, and future applications. Anal Biochem. 2002;307(2):191–201.
Sadri R, Hornsby PJ. Rapid analysis of DNA methylation using new restriction enzyme sites created by bisulfite modification. Nucleic Acids Res. 1996;24(24):5058–9.
Sato N, et al. Frequent hypomethylation of multiple genes overexpressed in pancreatic ductal adenocarcinoma. Cancer Res. 2003;63(14):4158–66.
Saxonov S, et al. A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters. Proc Natl Acad Sci. 2006;103(5):1412–7.
Sayed D, et al. MicroRNAs play an essential role in the development of cardiac hypertrophy. Circ Res. 2007;100(3):416–24.
Segura-Pacheco B, et al. Global DNA hypermethylation-associated cancer chemotherapy resistance and its reversion with the demethylating agent hydralazine. J Transl Med. 2006;4(1):32.
Selker EU, et al. The methylated component of the Neurospora crassa genome. Nature. 2003;422(6934):893–7.
Shapiro R, et al. Reactions of uracil and cytosine derivatives with sodium bisulfite. J Am Chem Soc. 1970;92(2):422–4.
Shingara J, et al. An optimized isolation and labeling platform for accurate microRNA expression profiling. RNA. 2005;11(9):1461–70.
Shirodkar AV, Marsden PA. Epigenetics in cardiovascular disease. Curr Opin Cardiol. 2011;26(3):209.
Shu J, et al. Dynamic and modularized microRNA regulation and its implication in human cancers. Sci Rep. 2017;7(1):13356.
Silverman LR, et al. Randomized controlled trial of azacitidine in patients with the myelodysplastic syndrome: a study of the cancer and leukemia group B. J Clin Oncol. 2002;20(10):2429–40.
Singer-Sam J, et al. A quantitative HpaII-PCR assay to measure methylation of DNA from a small number of cells. Nucleic Acids Res. 1990;18(3):687.
Singh N, et al. Molecular modeling and molecular dynamics studies of hydralazine with human DNA methyltransferase 1. Chem Med Chem. 2009;4(5):792–9.
Sobulo OM, et al. MLL is fused to CBP, a histone acetyltransferase, in therapy-related acute myeloid leukemia with a t (11; 16)(q23; p13. 3). Proc Natl Acad Sci. 1997;94(16):8732–7.
Song F, et al. Association of tissue-specific differentially methylated regions (TDMs) with differential gene expression. Proc Natl Acad Sci U S A. 2005;102(9):3336–41.
Šorm F, et al. 5-Azacytidine, a new, highly effective cancerostatic. Cell Mol Life Sci. 1964;20(4):202–3.
Spencer VA, et al. Chromatin immunoprecipitation: a tool for studying histone acetylation and transcription factor binding. Methods. 2003;31(1):67–75.
Stenvinkel P, et al. Impact of inflammation on epigenetic DNA methylation–a novel risk factor for cardiovascular disease? J Intern Med. 2007;261(5):488–99.
Stewart D, et al. A phase I pharmacokinetic and pharmacodynamic study of the DNA methyltransferase 1 inhibitor MG98 administered twice weekly. Ann Oncol. 2003;14(5):766–74.
Suzuki H, et al. Epigenetic inactivation of SFRP genes allows constitutive WNT signaling in colorectal cancer. Nat Genet. 2004;36(4):417–22.
Suzuki K, et al. Global DNA demethylation in gastrointestinal cancer is age dependent and precedes genomic damage. Cancer Cell. 2006;9(3):199–207.
Swedberg K, et al. Guidelines for the diagnosis and treatment of chronic heart failure: executive summary (update 2005). Eur Heart J. 2005;26(11):1115–40.
Thiagalingam S, et al. Loss of heterozygosity as a predictor to map tumor suppressor genes in cancer: molecular basis of its occurrence. Curr Opin Oncol. 2002;14(1):65–72.
Thiagarajan D, et al. Mechanisms of transcription factor acetylation and consequences in hearts. Biochim Biophys Acta. 2016;1862(12):2221–31.
Thum T, et al. MicroRNA-21 contributes to myocardial disease by stimulating MAP kinase signalling in fibroblasts. Nature. 2008;456(7224):980–4.
Tijsen AJ, et al. Circulating microRNAs as diagnostic biomarkers for cardiovascular diseases. Am J Phys Heart Circ Phys. 2012;303(9):H1085–95.
Tobi EW, et al. DNA methylation differences after exposure to prenatal famine are common and timing- and sex-specific. Hum Mol Genet. 2009;18(21):4046–53.
Toyota M, Issa J-PJ. CpG island methylator phenotypes in aging and cancer. Semin Cancer Biol. 1999;9(5):349–57.
Trivedi MS, et al. Short-term sleep deprivation leads to decreased systemic redox metabolites and altered epigenetic status. PLoS One. 2017;12(7):e0181978.
Tsai M-C, et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010;329(5992):689–93.
Turunen MP, et al. Epigenetics and atherosclerosis. Biochim Biophys Acta (BBA)-Gen Subj. 2009;1790(9):886–91.
Usui M, et al. Increased endogenous nitric oxide synthase inhibitor in patients with congestive heart failure. Life Sci. 1998;62(26):2425–30.
Usui T, et al. HDAC4 mediates development of hypertension via vascular inflammation in spontaneous hypertensive rats. Am J Physiol Heart Circ Physiol. 2012;302(9):H1894–904.
Vaissière T, et al. Epigenetic interplay between histone modifications and DNA methylation in gene silencing. Mutat Res/Rev Mutat Res. 2008;659(1):40–8.
Valencia A, et al. Wnt signaling pathway is epigenetically regulated by methylation of Wnt antagonists in acute myeloid leukemia. Leukemia. 2009;23(9):1658–66.
Van Aelst LN, et al. Osteoglycin prevents cardiac dilatation and dysfunction after myocardial infarction through infarct collagen strengthening. Circ Res. 2014;116:425–36.
Varrone F, et al. The circulating level of FABP3 is an indirect biomarker of microRNA-1. J Am Coll Cardiol. 2013;61(1):88–95.
Villar AV, et al. Myocardial and circulating levels of microRNA-21 reflect left ventricular fibrosis in aortic stenosis patients. Int J Cardiol. 2013;167(6):2875–81.
Voellenkle C, et al. MicroRNA signatures in peripheral blood mononuclear cells of chronic heart failure patients. Physiol Genomics. 2010;42(3):420–6.
Waddington C. The epigenetics of birds. New York: Cambridge University Press; 1952.
Wadhwa PD, et al. Developmental origins of health and disease: brief history of the approach and current focus on epigenetic mechanisms. Semin Reprod Med. 2009;27(5):358–68.
Wagner JM, et al. Histone deacetylase (HDAC) inhibitors in recent clinical trials for cancer therapy. Clin Epigenetics. 2010;1(3):117.
Wang LQ, Chim CS. DNA methylation of tumor-suppressor miRNA genes in chronic lymphocytic leukemia. Epigenomics. 2015;7(3):461–73.
Wang Y, et al. Dysregulation of histone acetyltransferases and deacetylases in cardiovascular diseases. Oxidative Med Cell Longev. 2014;2014:641979.
Waterland RA, Jirtle RL. Transposable elements: targets for early nutritional effects on epigenetic gene regulation. Mol Cell Biol. 2003;23(15):5293–300.
Waterland RA, Michels KB. Epigenetic epidemiology of the developmental origins hypothesis. Annu Rev Nutr. 2007;27:363–88.
Weber J, et al. Expression of the MAGE-1 tumor antigen is up-regulated by the demethylating agent 5-aza-2′-deoxycytidine. Cancer Res. 1994;54(7):1766–71.
Weber M, et al. Chromosome-wide and promoter-specific analyses identify sites of differential DNA methylation in normal and transformed human cells. Nat Genet. 2005;37(8):853–62.
Weber M, et al. Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nat Genet. 2007;39(4):457–66.
West AC, Johnstone RW. New and emerging HDAC inhibitors for cancer treatment. J Clin Invest. 2014;124(1):30–9.
Whittaker SJ, et al. Final results from a multicenter, international, pivotal study of romidepsin in refractory cutaneous T-cell lymphoma. J Clin Oncol. 2010;28(29):4485–91.
Widschwendter M, et al. DNA hypomethylation and ovarian cancer biology. Cancer Res. 2004;64(13):4472–80.
Willenbrock H, et al. Quantitative miRNA expression analysis: comparing microarrays with next-generation sequencing. RNA. 2009;15(11):2028–34.
Wilson Tang WH, et al. Differential effects of arginine methylation on diastolic dysfunction and disease progression in patients with chronic systolic heart failure. Eur Heart J. 2008;29(20):2506–13.
Wimalawansa SJ. Vitamin D and cardiovascular diseases: causality. J Steroid Biochem Mol Biol. 2016;175:29–43.
Xiao H, et al. Both Sp1 and Sp3 are responsible for p21waf1 promoter activity induced by histone deacetylase inhibitor in NIH3T3 cells. J Cell Biochem. 1999;73(3):291–302.
Xie M, Hill JA. HDAC-dependent ventricular remodeling. Trends Cardiovasc Med. 2013;23(6):229–35.
Yamada Y, et al. Opposing effects of DNA hypomethylation on intestinal and liver carcinogenesis. Proc Natl Acad Sci U S A. 2005;102(38):13580–5.
Yamashita K, et al. Pharmacologic unmasking of epigenetically silenced tumor suppressor genes in esophageal squamous cell carcinoma. Cancer Cell. 2002;2(6):485–95.
Yanazume T, et al. Cardiac p300 is involved in myocyte growth with decompensated heart failure. Mol Cell Biol. 2003;23(10):3593–606.
Yang XJ, Seto E. HATs and HDACs: from structure, function and regulation to novel strategies for therapy and prevention. Oncogene. 2007;26(37):5310–8.
Yang XJ, Seto E. Lysine acetylation: codified crosstalk with other posttranslational modifications. Mol Cell. 2008;31(4):449–61.
Yang X, et al. Targeting DNA methylation for epigenetic therapy. Trends Pharmacol Sci. 2010;31(11):536–46.
Yang Q, et al. Pulmonary artery smooth muscle cell proliferation and migration in fetal lambs acclimatized to high-altitude long-term hypoxia: role of histone acetylation. Am J Physiol Lung Cell Mol Physiol. 2012;303(11):L1001–10.
Yoder JA, et al. Cytosine methylation and the ecology of intragenomic parasites. Trends Genet. 1997;13(8):335–40.
Yoo CB, et al. Long-term epigenetic therapy with oral zebularine has minimal side effects and prevents intestinal tumors in mice. Cancer Prev Res. 2008;1(4):233–40.
Yoshida M, et al. Potent and specific inhibition of mammalian histone deacetylase both in vivo and in vitro by trichostatin A. J Biol Chem. 1990;265(28):17174–9.
Yoshikawa H, et al. SOCS-1, a negative regulator of the JAK/STAT pathway, is silenced by methylation in human hepatocellular carcinoma and shows growth-suppression activity. Nat Genet. 2001;28(1):29–35.
Zaina S, et al. Nutrition and aberrant DNA methylation patterns in atherosclerosis: more than just hyperhomocysteinemia? J Nutr. 2005;135(1):5–8.
Zhang Y, et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 2008;9(9):R137.
Zhao W, et al. Abnormal activation of the synuclein-gamma gene in hepatocellular carcinomas by epigenetic alteration. Int J Oncol. 2006;28(5):1081–8.
Zhao Y, et al. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell. 2007;129(2):303–17.
Zhong J, et al. Cardiac autonomic dysfunction: particulate air pollution effects are modulated by epigenetic immunoregulation of toll-like receptor 2 and dietary flavonoid intake. J Am Heart Assoc. 2015;4(1):e001423.
Zhu S, et al. Inactivation of monocarboxylate transporter MCT3 by DNA methylation in atherosclerosis. Circulation. 2005;112(9):1353–61.
Zilberman D, Henikoff S. Genome-wide analysis of DNA methylation patterns. Development. 2007;134(22):3959–65.
Acknowledgment
This work was supported by Hong Kong Research Grants Council – General Research Fund (HKU 17127215; project title: Molecular Mechanisms Underlying the Progressive Development of Pulmonary Arterial Hypertension in Secretin Knockout Mice; PI: Professor BKC Chow, School of Biological Sciences; 1,100,302 HKD, ongoing).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Wang, L.Q., Singh, K., Zaw, A.M., Chow, B.K.C. (2018). The Emerging Role of Epigenetics. In: Jiang, H., Liu, M. (eds) Heart Genomics. Translational Bioinformatics, vol 16. Springer, Singapore. https://doi.org/10.1007/978-981-13-1429-2_3
Download citation
DOI: https://doi.org/10.1007/978-981-13-1429-2_3
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-1428-5
Online ISBN: 978-981-13-1429-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)