Abstract
Vitelliform macular dystrophy (VMD) is a group of macular dystrophy characterized by the subretinal accumulation of yellow yolk-like materials which predominantly affect the macula. Best vitelliform macular dystrophy is among the most common autosomal dominant (AD) retinal dystrophy, caused by mutations in the BEST1 gene. Since first identification of BEST1 gene in 1998, molecular biology and pathophysiology of BEST1 gene and vitelliform macular dystrophy were studied. Recent advances in genetic analysis have described over 200 different human BEST1 mutations to date, associated with a broad spectrum of ocular diseases, called bestrophinopathy. However, the genotype-phenotype correlation in VMD is largely unexplored. Genetic test is clinically important in the diagnosis of VMD because the clinical features of VMD are similar to those of exudative age-related macular degeneration (AMD), choroidal neovascularization (CNV), or central serous chorioretinopathy (CSC). Here, in addition to describing the clinical characteristics of VMD, this chapter focuses on the clinical genetics of BEST1 gene in VMD.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Vitelliform macular dystrophy
- Bestrophin-1
- Best vitelliform macular dystrophy
- Adult-onset vitelliform macular dystrophy
- BEST1 gene mutation
- Genome editing
21.1 Introduction
Macular dystrophy is a group of heritable disorders that cause ophthalmoscopically visible macular abnormalities. Vitelliform macular dystrophy (VMD) is a group of macular dystrophy characterized by the subretinal accumulation of yellow yolk-like materials which predominantly affects the macula. Best vitelliform macular dystrophy (BVMD) is named after Friedrich Best who described a family with a history of early-onset macular degeneration in 1905 [1]. BVMD is among the most common autosomal dominant (AD) retinal dystrophy, caused by mutations in the BEST1 gene . Since the first identification of BEST1 gene in 1998 [2], molecular biology and pathophysiology of BEST1 gene in VMD have been studied. Recent advances in genetic analysis have described over 200 different human BEST1 mutations to date, associated with a broad spectrum of ocular diseases , called a bestrophinopathy [3, 4]. Bestrophinopathy includes five clinically distinct categories BVMD, adult-onset vitelliform macular dystrophy (AVMD) , autosomal recessive bestrophinopathy (ARB), autosomal dominant vitreoretinochoroidopathy (ADVIRC), and retinitis pigmentosa . AVMD was first described by Gass in 1974 who initially termed it peculiar foveomacular dystrophy [5]. AVMD is one of the most common forms of macular dystrophy as well [6]. Many investigators suggested that AVMD is a mild form of BVMD within the same spectrum because the clinical features of AVMD were similar to those of early-stage BVMD and the age of onset were highly variable [7,8,9]. Clinically, BVMD is distinguished from AVMD by earlier age of onset, larger lesion size, and an abnormal electrooculogram (EOG). Clinical features of VMD are similar to those of exudative age-related macular degeneration (AMD) , choroidal neovascularization (CNV), or central serous chorioretinopathy (CSC). Thus, genetic test is clinically important in the diagnosis of VMD. Here, in addition to describing the clinical characteristics of VMD, this chapter focuses on the clinical genetics of BEST1 gene in VMD (BVMD and AVMD).
21.2 Epidemiology and Asian Perspective
VMD is an autosomal dominant macular dystrophy with an estimated prevalence of 1 in 10,000 in the USA [10], 2/10,000 in Sweden [11], 1.5/100,000 in Denmark [12], and 1 in 16,500 to 1 in 21,000 in Olmsted County, Minnesota, USA [13]. Males are more affected than females (3:1) [11, 12]. Despite the update of novel mutations of BEST1 in Asian VMD patients, there was no report of the prevalence of VMD in Asian countries. Thus, a study of the prevalence of VMD with a genetic analysis in Asian countries is necessary.
21.3 Molecular Biology
The BEST1 gene consists of 11 exons that encode the bestrophin-1 protein (585 amino acids). Bestrophin-1 is a retinal pigment epithelium (RPE) protein hypothesized to function as a Ca2+-activated Cl- channel (CaCC), or a regulator of ion transport [14]. Bestrophin-1 is predominantly expressed in the basolateral membrane of the RPE [15]. X-ray structure of chicken BEST1-Fab complexes indicates that Bestrophin-1 forms a homo-pentamer and functions as a CaCC [16]. Disease-causing mutations are prevalent within the gating apparatus. In addition, Bestrophin-1 functions as a regulator of intracellular calcium signaling and influences transepithelial electrical properties [17]. Recently, patient’s stem cell-derived RPE is used for the function of bestrophin-1 and reveals that bestrophin-1 assembles into a key calcium-sensing chloride channel in human RPE [18]. Further study using RPE cells from patient-derived induced pluripotent stem cells (iPSc) harboring BEST1 mutations is required to elucidate the exact functional role of bestrophin-1.
21.4 Clinical Features
21.4.1 BVMD
BVMD is an early-onset autosomal dominant disorder showing extremely variable penetrance and expressivity. The diagnosis of BVMD shows a bimodal age distribution; the first maximum peak was made during the childhood, but the second peak was made following puberty and extending into the sixth decade of life [19]. Before the era of genetic analysis, the diagnosis of BVMD was based on typical fundus findings, family history, and a decreased Arden ratio (light peak/dark trough) of EOG with a normal electroretinogram (ERG), which may contribute to variability of penetrance, expressivity, and onset age.
BVMD is caused by dysfunction of Bestrophin-1 protein, a CaCC protein located on the basolateral membrane of RPE, which causes abnormal fluid and ion exchange that decreases pumping of fluid from the subretinal space, and results in swelling of RPE and subretinal lipofuscin accumulation [20]. Histopathologically, autofluorescent material was accumulated in the outer retina and the subretinal space in BVMD, which is considered as indigestible components of photoreceptor outer segments that accumulate due to the lack of direct apposition of the outer segments and the RPE [21]. Eventual phagocytosis of these older materials over time would load the RPE cells and may account for excessive accumulation of abnormal lipofuscin in RPE cells across the entire fundus [22]. These findings coincide with the decreased Arden ratio of EOG, less than 1.5, seen in BVMD, which suggest generalized dysfunction of the RPE. Even otherwise asymptomatic carriers of BEST1 mutations will exhibit an altered EOG [23]. Full-field ERG is generally normal, but the multifocal ERG amplitudes of the central and pericentral responses were significantly reduced in the majority of patients [24]. However, the photoreceptor structure evaluated cellular imaging with adaptive optics scanning light ophthalmoscopy was retained within active BVMD lesions, even in apparently advanced disease [25, 26].
Five progressive stages can be defined based on fundus examination [20, 27]. However, these stages are not observed in all patients, nor do they occur consecutively. The first previtelliform stage is characterized by the absence of symptoms and subtle RPE changes such as RPE mottling and a small yellow spot. On optical coherence tomography (OCT), RPE and ellipsoid zone (EZ) disruption was detectable in a small fraction of eyes [28, 29]. A slight thickening of the interdigitation zone was also observed [30]. EOG is abnormal and fluorescein angiogram (FA) shows window defects. Visual acuity remains intact in most patients. The previtelliform lesions are characterized by absence or only slight autofluorescence on fundus autofluorescence (FAF) imaging.
The second vitelliform shows a well-circumscribed, circular, homogeneous, yellow-opaque, 0.5–3 disc diameter sized, yolk-like macular lesions. The remaining part of fundus usually has normal appearance, but multifocal lesions also can be seen. The accumulation of hyperreflective vitelliform material is clearly visible on OCT below the neurosensory retina , located between the EZ and the RPE. The disruption of outer retinal layers and neurosensory retinal detachment with subretinal fluid occur in many cases [28, 29]. The yellowish subretinal material is intensely hyperautofluorescent in FAF imaging. FA shows marked hypofluorescence in the zone covered by lesion by blockage of fluorescence. Metamorphopsia, blurred vision, and a decrease of central vision can occur.
In the third pseudohypopyon stage, the vitelliform material accumulates inferiorly and develops a fluid level. On OCT, the upper part of the lesion is observed as hyporeflective area located between RPE and EZ, with clumping of hyperreflective material on the posterior retinal surface. The lower part of the lesion, where the vitelliform material is still accumulated, shows a highly reflective area located in the subretinal space. FA shows hypofluorescence in the lower part resulted from the blockage by the vitelline material. The superior part shows hyperfluorescent due to transmission defects linked to RPE and chorioretinal atrophy in the early phase. FAF shows loss of autofluorescence, particularly in the upper part.
The fourth vitelliruptive stage is characterized by the partial reabsorption of the vitelliform material. This vitelliform material becomes less homogeneous to develop a “scrambled-egg” appearance. OCT shows an optically empty lesion between EZ and RPE, with clumping of hyperreflective material on the posterior retinal surface like the upper part of the pseudohypopyon. The areas of focal RPE hypertrophy can be observed as hyperreflective mottling on the RPE layer on some parts. FAF shows decreased of autofluorescence centrally but increased autofluorescence at the outer border of the lesion.
In the last atrophic/fibrotic stage, RPE atrophy and loss of central vision occur after rupture and reabsorption of the cystic lesion. FA shows hyperfluorescence without leakage. OCT reveals thinning of all the retinal layers and diffuse disappearance of outer retinal layers within the macular area, with highly hyperreflective thickening at the RPE level [29, 31]. Atrophic lesions are characterized by decreased autofluorescence on FAF.
Choroidal neovascularization (CNV) may develop and can lead to form a disciform scar. Patients usually underwent sudden visual disturbance with central scotoma and/or metamorphopsia, showing a macular hemorrhage on fundus examination. In that case, FA shows hyperfluorescence because of CNV and leakage. Intravitreal injection of anti-vascular endothelial growth factor (VEGF) agent was effective in treating CNV complicated with BVMD and safe even in children [32,33,34].
Patients with BVMD undergo a progressive decrease of vision over time. In a study that evaluated the course of visual decline of 53 patients in BVMD with BEST1 mutation [35], the median age of onset of visual symptoms was 33 years. Twenty-five percent of patients retain visual acuity of 20/40 or better at the age of 66 years. Other study evaluated 47 patient with BVMD; 74% of patient older than 30 years had 20/100 or worse visual acuity at least one eye [36].
21.4.2 AVMD
Gass reported a three-generation family and six sporadic patients characterized by one-third disc diameter sized bilateral subfoveal vitelliform lesions with onset between the ages of 30 and 50 years accompanied by slowly progressive visual loss as “peculiar foveomacular dystrophy.” They also showed occasional paracentral drusen, normal to slightly subnormal response on EOG but normal ERG and color vision [5]. AVMD shows a variable genetic inheritance, although most cases are sporadic [37]. Patients with AVMD may be asymptomatic but become symptomatic in the fourth or fifth decade of life with blurred vision, metamorphopsia, or scotoma and typically have slow progression of vision loss [38]. Patients with AVMD typically present a round, yellowish subretinal deposit in one-third to one disc diameter size within the macular area, similar fundus finding to the vitelliform stage of BVMD.
The initial yellow lesion may present in only one eye and appear as small yellow flecks in the paracentral area. EOG shows a normal or slightly reduced Arden ratio, which is obviously abnormal in BVMD. The macular lesion appears as hyperautofluorescent in FAF. The vitelliform deposit usually appears as initially hypofluorescent but gradually becomes hyperfluorescent on the edges by staining of the dye in FA [39] and hypofluorescent on indocyanine green angiography (ICGA). OCT reveals a dome-shaped hyperreflective lesion located between the retina and RPE [40]. The foveal thinning and EZ disruption are also observed and probably explain the progressive visual loss [41, 42].
AVMD progression is characterized by fragmentation and reabsorption of the vitelliform material [6]. Macular atrophy progressively replaces the vitelliform deposits at the advanced stages of the disease in most cases [42], but most patients retain reading vision throughout life [43, 44]. CNV may be complicated in few cases; 6 out of 51 patients developed CNV after a 6-year follow-up [45]. Anti-VEGF therapies have shown to be effective in the treatment of CNV associated with AVMD [46].
21.5 Genetic Aspects
21.5.1 BVMD
Currently, only genetic test for mutation analysis of the BEST1 gene leads to confirmation of a clinical diagnosis of BVMD. Note that individuals with clinical findings of BVMD occasionally have a normal EOG, turning out to have a pathogenic variant of BEST1 [47]. In case of atypical BVMD [3], genetic test for confirmation should be performed. Over 200 BEST1 mutations with significant clinical heterogeneity require a thorough genetic analysis and clinical examinations to better understanding of genotype -phenotype correlations in BVMD. Most mutations of BEST1 gene in BVMD and AVMD are missense mutations. Table 21.1 shows a list of missense mutations of BEST1 gene in BVMD and AVMD.
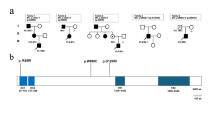
Most genetic studies were performed in Western countries including the USA, England, Sweden, Denmark, Germany, the Netherlands, Italy, and France. BEST1 mutations are extremely heterogenous, but several mutations have been frequently found (Thr6Pro, Arg25Trp, Arg218Cys, Tyr227Asn, Arg243Val, Ile295del, Gle300Asp, Asp301Glu, and Asp302Asn). Interestingly, these frequent mutations are ethnic specific (44.4% of Asp302Asn in Danish [12] and 36.8% of Arg25Trp in Italian [86]).
Currently, only limited reports are available in Asian genetic studies of BEST1 from Chinese [50, 54, 70, 76, 80, 87,88,89], Japanese [48, 83], and Korean [9]. The mutation spectrum of the BEST1 gene in Asian patients of BVMD is differed from those in Western patients [88]. Six novel missense mutations (Thr2Asn, Leu75Phe, Ser144Asn, Arg255Trp, Pro297Thr, and Asp301Gly) and one previously reported mutation (Arg218Cys) were identified [50]. Three novel mutations Tyr4Ile [54], Ala291Val [54], and Phe113Leu [76] in BVMD were reported. Lin [80] reported two novel heterozygous mutations 304delAsp and Trp229Gly in Chinese BVMD patients. Liu [70] reported four previously reported mutations (Ser16Phe, Ser144Asn, Glu292Lys, and Glu300Lys) and two novel disease-causing mutations (Thr307Asp, Arg47His) in Chinese patients with BVMD.
In Japanese study [48], 22 patients including 16 probands from 16 families with BVMD were analyzed. All 16 probands exhibited characteristic BVMD fundus appearances, abnormal EOG, and normal ERG responses with the exception of one diabetic retinopathy proband. Genetic analysis identified 12 BEST1 variants in 13 probands (81%). Of these, ten variants (Tyr2Arg, Arg25Trp, Phe80Leu, Val81Met, Ala195Val, Arg218His, Gly222Glu, Val242Met, Asp304del, and Glu306Asp) have been previously reported in BVMD, while two variants (Ser7Asn and Pro346His) were novel disease-causing mutations.
In Korea, we report a BVMD patient (Fig. 21.1) carrying Asn296Lys mutation which is a causative mutation of multifocal BVMD in German patient [60]. Arg218Leu is a novel disease-causing mutation in BVMD (Fig. 21.2). These findings expand the spectrum of BEST1 genetic variation in Asian and will be valuable for genetic counseling for patients with BVMD [88].
Best vitelliform macular dystrophy (BVMD). A 32-year-old man carrying p.Asn296Lys mutation in the BEST1 gene was incidentally found on routine fundus examination for a pilot license. The visual acuities (VA) were 20/20 in both eyes. (a, f) Bilateral BVMD of vitelliruptive stage shows scattered yellow-white vitelliform deposits. (b, g) Vertical optical coherent tomography (OCT) shows serous retinal detachment and hyperreflective vitelliform materials at RPE in both eyes. (c, h) Fluorescein angiography (FA) shows late pooling of fluorescein dye at the egg lesion. (d, i) Fundus autofluorescence (FAF) image of the vitelliruptive lesion shows increased autofluorescence at inferior part of ruptured vitelliform lesions and at the border of the serous retinal detachment. (e, j) Indocyanine green angiography (ICGA) shows active leakage spot in the left eye
Best vitelliform macular dystrophy (BVMD). A 39-year-old man carrying Arg218Leu mutation in the BEST1 gene had multiple injections of anti-VEGF agents (ten for right eye and five for left eye) in both eyes. At initial visiting in our institute, vitelliform stage of right eye (a) reveals highly reflective subfoveal pillar without surrounding SRF (b). Small round vitelliform lesion with central cicatricial change was found in the left eye (e). OCT reveals marked RPE loss at the fovea (f). Six months later, FAF shows dispersed materials with hyperautofluorescent (c), and OCT reveals the disappearance of subfoveal pillar with a progression to vitelliruptive stage (d) in the right eye. FAF shows central hypoautofluorescent and surrounding hyperfluorescent lesions. OCT reveals that hypoautofluorescent lesion corresponds to the enlarged RPE loss (h)
BVMD shows variable expressivity and incomplete penetrance at the clinical level. Disease-causing effect of BEST1 mutations seems to be cumulative over time [79]. In genotype -phenotype relationship of Dutch study [59], median age of onset of visual symptoms was 33 years (range, 2–78). The cumulative risk of VA below 0.5 (20/40) was 50% at 55 years and 75% at 66 years. The cumulative risk of VA decline less than 0.3 (20/63) was 50% by age 66 years and 75% by age 74 years. Most patients (96%) had missense mutations; the Thr6Pro, Ala10Val, and Tyr227Asn mutations were most common. Visual decline was significantly faster in patients with an Ala10Val mutation than either the Thr6Pro or the Tyr227Asn mutation.
In the recent Chinese study, despite typical macular appearance of BVMD, no clear genotype-phenotype correlation was observed [88]. In Asian BVMD cohort, genetic tests should be performed for the diagnosis with thorough clinical examinations to elucidate a genotype -phenotype correlation.
21.5.2 AVMD
In AVMD, several mutations in BEST1 gene have been identified including p.Ala146Lys [90], p.Thr6Pro, p.Arg47His, P.Ala243Val, p.Asp312Asn [61], and p.Ile38Ser [9]. Table 21.1 includes the list of missense mutations in AVMD. In addition, AVMD is associated with mutations in PRPH2 [91], IMPG1 [92], IMPG2 [93].
Age of onset is a major criterion to distinguish BVMD from AVMD [64]. Thus, systematic screening of BEST1 and PRPH2 has been suggested in BVMD and AVMD. BEST1 screening should be recommended to patients with an age of onset less than 40 years, and PRPH2 screening should be recommended to patients with an age of onset more than 40 years. For an onset between 30 and 40 years, PRPH2 can be screened if no mutation has been detected in BEST1. In this screening approach, we found PRPH2 mutation of p.Pro219_Pro221delinsPro in a 39-year-old female without BEST1 mutation (Fig. 21.3).
21.6 Future Perspectives for Therapy
The development of gene and cell therapies is promising in various retinal diseases . Indeed, the results of clinical trials using iPSC-derived RPE cells in wet age-related macular degeneration [94] or AAV/RPE65 vectors in Leber’s congenital amaurosis [95] were already reported. Therapeutic intervention of inherited retinal dystrophy should be primarily aimed at the restoration of normal gene (i.e., BEST1 gene in BVMD and AVMD) . However, until decade ago, this therapeutic goal was ideal but unachievable due to the lack of a proper biotechnology. Recent advances in genome editing technology using CRISPR system and gene delivery system are promising and harness the CRISPR-based genome editing for the therapeutic applications. Since its first therapeutic applications in retinal disease using wet AMD animal models [96, 97], in vivo genome editing using CRISPR-Cas9 enlarged its therapeutic applications both in genetic diseases harboring mutations [98, 99] and nongenetic degenerative diseases [96, 97, 100].
Conventional concept of gene therapy to deliver normal copy of BEST1 gene into RPE would be effective in the treatment of VMD of haploinsufficiency phenotype , which is caused by BEST1 mutations that exclusively result in a loss of sufficient wild-type protein. In addition, simple destruction of mutant proteins at the DNA level is achievable by genome editing of mutant BEST1 allele using CRISPR-Cas9.
Currently, many BEST1 mutations cause VMD through dominant negative effect. In addition, over 200 mutations of BEST1 gene, large amounts of BEST1 mutations are missense mutations; thus, a precise base-editing using base-editors enables a literally complete recovery of normal gene [101, 102]. According to the recent advances in genome editing technology using CRISPR system, in vivo genome editing has emerged as a potential treatment strategy for inherited retinal dystrophies [103].
21.7 Summary
VMD is among the most common autosomal dominant macular dystrophy. Multimodal imaging with SD-OCT, FAF, FA, and ICGA is useful to the diagnosis of VMD. Genetic test is clinically important in the diagnosis of VMD because the clinical features of VMD can be similar to those of exudative AMD , CNV, or CSC. Future studies are needed to identify the prevalence with precise genetic mutations of BEST1 in Asian VMD patients. This could provide a clear genotype -phenotype correlation in VMD. In vitro studies using RPE cells from patient-derived iPSC help to understand molecular biology of bestrophin-1 protein. Furthermore, in vivo genome editing using CRISPR-based base-editors might be a potential treatment strategy for the correction of missense mutations in VMD.
References
Best F. Uber eine hereditare maculaafektion; Beitrage zur verergslehre. Zschr Augenheilkd. 1905;13:199–212.
Petrukhin K, et al. Identification of the gene responsible for Best macular dystrophy. Nat Genet. 1998;19:241–7.
Boon CJ, et al. The spectrum of ocular phenotypes caused by mutations in the BEST1 gene. Prog Retin Eye Res. 2009;28:187–205.
Johnson AA, et al. Bestrophin 1 and retinal disease. Prog Retin Eye Res. 2017;58:45–69.
Gass J. A clinicopathologic study of a peculiar foveomacular dystrophy. Trans Am Ophthalmol Soc. 1974;72:139.
Chowers I, Tiosano L, Audo I, Grunin M, Boon CJ. Adult-onset foveomacular vitelliform dystrophy: a fresh perspective. Prog Retin Eye Res. 2015;47:64–85.
Johnson AA, et al. Bestrophin 1 and retinal disease. Prog Retin Eye Res. 2017;58:45–69.
Boon CJ, et al. The spectrum of ocular phenotypes caused by mutations in the BEST1 gene. Prog Retin Eye Res. 2009;28:187–205.
Jun I, et al. Adult-onset Vitelliform macular dystrophy caused by BEST1 p.Ile38Ser mutation is a mild form of Best Vitelliform macular dystrophy. Sci Rep. 2017;7:9146.
Mohler CW, Fine SL. Long-term evaluation of patients with Best’s vitelliform dystrophy. Ophthalmology. 1981;88:688–92.
Nordstrom S. Hereditary macular degeneration--a population survey in the country of Vsterbotten, Sweden. Hereditas. 1974;78:41–62.
Bitner H, Schatz P, Mizrahi-Meissonnier L, Sharon D, Rosenberg T. Frequency, genotype, and clinical spectrum of best vitelliform macular dystrophy: data from a national center in Denmark. Am J Ophthalmol. 2012;154:403–412 e404.
Dalvin LA, Pulido JS, Marmorstein AD. Vitelliform dystrophies: prevalence in Olmsted County, Minnesota, United States. Ophthalmic Genet. 2017;38:143–7.
Marmorstein AD, Cross HE, Peachey NS. Functional roles of bestrophins in ocular epithelia. Prog Retin Eye Res. 2009;28:206–26.
Marmorstein AD, et al. Bestrophin, the product of the Best vitelliform macular dystrophy gene (VMD2), localizes to the basolateral plasma membrane of the retinal pigment epithelium. Proc Natl Acad Sci U S A. 2000;97:12758–63.
Kane Dickson V, Pedi L, Long SB. Structure and insights into the function of a Ca(2+)-activated Cl(−) channel. Nature. 2014;516:213–8.
Marmorstein AD, et al. Bestrophin-1 influences transepithelial electrical properties and Ca2+ signaling in human retinal pigment epithelium. Mol Vis. 2015;21:347–59.
Moshfegh Y, et al. BESTROPHIN1 mutations cause defective chloride conductance in patient stem cell-derived RPE. Hum Mol Genet. 2016;25:2672–80.
Nordstrom S, Barkman Y. Hereditary maculardegeneration (HMD) in 246 cases traced to one gene-source in Central Sweden. Hereditas. 1977;84:163–76.
Mohler CW, Fine SL. Long-term evaluation of patients with Best’s vitelliform dystrophy. Ophthalmology. 1981;88:688–92.
Zhang Q, Small KW, Grossniklaus HE. Clinicopathologic findings in Best vitelliform macular dystrophy. Graefes Arch Clin Exp Ophthalmol. 2011;249:745–51.
O’Gorman S, Flaherty WA, Fishman GA, Berson EL. Histopathologic findings in Best’s vitelliform macular dystrophy. Arch Ophthalmol. 1988;106:1261–8.
Deutman A. Electro-oculography in families with vitelliform dystrophy of the fovea: detection of the carrier state. Arch Ophthalmol. 1969;81:305–16.
Scholl HPN, Schuster AM, Vonthein R, Zrenner E. Mapping of retinal function in Best macular dystrophy using multifocal electroretinography. Vis Res. 2002;42:1053–61.
Scoles D, et al. Photoreceptor inner segment morphology in Best vitelliform macular dystrophy. Retina (Philadelphia, Pa.). 2017;37:741–8.
Kay DB, et al. Outer retinal structure in best vitelliform macular dystrophy. JAMA ophthalmology. 2013;131:1207–15.
Deutman AF. Ph.D. thesis, Koninklijke Van Gorcum, Assen; 1971.
Battaglia Parodi M, et al. Optical coherence tomography in Best vitelliform macular dystrophy. Eur J Ophthalmol. 2017;27:201–4.
Battaglia Parodi M, Iacono P, Romano F, Bandello F. Spectral domain optical coherence tomography features in different stages of best vitelliform macular dystrophy. Retina (Philadelphia, Pa.). 2017;38(5):1041–6.
Qian CX, et al. Optical coherence tomography examination of the retinal pigment epithelium in Best vitelliform macular dystrophy. Ophthalmology. 2017;124:456–63.
Querques G, et al. High-definition optical coherence tomography features in vitelliform macular dystrophy. Am J Ophthalmol. 2008;146:501–7. e501.
Ruiz-Moreno O, Calvo P, Ferrández B, Torrón C. Long-term outcomes of intravitreal ranibizumab for choroidal neovascularization secondary to Best’s disease: 3-year follow-up. Acta Ophthalmol. 2012;90:e574–5.
Chhablani J, Jalali S. Intravitreal bevacizumab for choroidal neovascularization secondary to Best vitelliform macular dystrophy in a 6-year-old child. Eur J Ophthalmol. 2012;22:677–9.
Leu J, Schrage NF, Degenring RF. Choroidal neovascularisation secondary to Best’s disease in a 13-year-old boy treated by intravitreal bevacizumab. Graefes Arch Clin Exp Ophthalmol. 2007;245:1723–5.
Booij JC, et al. Course of visual decline in relation to the Best1 genotype in vitelliform macular dystrophy. Ophthalmology. 2010;117:1415–22.
Fishman GA, et al. Visual acuity in patients with best vitelliform macular dystrophy. Ophthalmology. 1993;100:1665–70.
Brecher R, Bird A. Adult vitelliform macular dystrophy. Eye. 1990;4:210–5.
Glacet-Bernard A, Soubrane G, Coscas G. Macular vitelliform degeneration in adults. Retrospective study of a series of 85 patients. J Fr Ophtalmol. 1989;13:407–20.
Chowers I, Tiosano L, Audo I, Grunin M, Boon CJ. Adult-onset foveomacular vitelliform dystrophy: a fresh perspective. Prog Retin Eye Res. 2015;47:64–85.
Puche N, et al. High-resolution spectral domain optical coherence tomography features in adult onset foveomacular vitelliform dystrophy. Br J Ophthalmol. 2010;94(9):1190–6.
Benhamou N, et al. Adult-onset foveomacular vitelliform dystrophy: a study by optical coherence tomography. Am J Ophthalmol. 2003;135:362–7.
Querques G, Forte R, Querques L, Massamba N, Souied EH. Natural course of adult-onset foveomacular vitelliform dystrophy: a spectral-domain optical coherence tomography analysis. Am J Ophthalmol. 2011;152:304–13.
Felbor U, Schilling H, Weber BH. Adult vitelliform macular dystrophy is frequently associated with mutations in the peripherin/RDS gene. Hum Mutat. 1997;10:301–9.
Yamaguchi K, et al. Adult-onset foveomacular vitelliform dystrophy with retinal folds. Jpn J Ophthalmol. 2001;45:533–7.
Da Pozzo S, Parodi MB, Toto L, Ravalico G. Occult choroidal neovascularization in adult-onset foveomacular vitelliform dystrophy. Ophthalmologica. 2001;215:412–4.
Mimoun G, et al. Ranibizumab for choroidal neovascularization associated with adult-onset foveomacular vitelliform dystrophy: one-year results. Retina (Philadelphia, Pa.). 2013;33:513–21.
Testa F, et al. A normal electro-oculography in a family affected by best disease with a novel spontaneous mutation of the BEST1 gene. Br J Ophthalmol. 2008;92:1467–70.
Katagiri S, et al. Mutation analysis of BEST1 in Japanese patients with Best’s vitelliform macular dystrophy. Br J Ophthalmol. 2015;99:1577–82.
Kinnick TR, et al. Autosomal recessive vitelliform macular dystrophy in a large cohort of vitelliform macular dystrophy patients. Retina (Philadelphia, Pa). 2011;31:581–95.
Wong RL, et al. Novel and homozygous BEST1 mutations in Chinese patients with Best vitelliform macular dystrophy. Retina (Philadelphia, Pa.). 2010;30:820–7.
Matson ME, Ly SV, Monarrez JL. Novel mutation in BEST1 associated with atypical Best Vitelliform dystrophy. Optom Vis Sci. 2015;92:e180–9.
Boon CJ, et al. Clinical and molecular genetic analysis of best vitelliform macular dystrophy. Retina (Philadelphia, Pa.). 2009;29:835–47.
Querques G, et al. Functional and clinical data of Best vitelliform macular dystrophy patients with mutations in the BEST1 gene. Mol Vis. 2009;15:2960–72.
Tian R, Yang G, Wang J, Chen Y. Screening for BEST1 gene mutations in Chinese patients with bestrophinopathy. Mol Vis. 2014;20:1594–604.
Lacassagne E, et al. Phenotypic variability in a French family with a novel mutation in the BEST1 gene causing multifocal Best vitelliform macular dystrophy. Mol Vis. 2011;17:309–22.
Apushkin MA, Fishman GA, Taylor CM, Stone EM. Novel de novo mutation in a patient with Best macular dystrophy. Arch Ophthalmol. 2006;124:887–9.
Mullins RF, Kuehn MH, Faidley EA, Syed NA, Stone EM. Differential macular and peripheral expression of bestrophin in human eyes and its implication for best disease. Invest Ophthalmol Vis Sci. 2007;48:3372–80.
Lotery AJ, et al. Allelic variation in the VMD2 gene in best disease and age-related macular degeneration. Invest Ophthalmol Vis Sci. 2000;41:1291–6.
Booij JC, et al. Course of visual decline in relation to the Best1 genotype in vitelliform macular dystrophy. Ophthalmology. 2010;117:1415–22.
Boon CJ, et al. Clinical and genetic heterogeneity in multifocal vitelliform dystrophy. Arch Ophthalmol. 2007;125:1100–6.
Kramer F, et al. Mutations in the VMD2 gene are associated with juvenile-onset vitelliform macular dystrophy (Best disease) and adult vitelliform macular dystrophy but not age-related macular degeneration. Eur J Hum Genet. 2000;8:286–92.
Bakall B, et al. The mutation spectrum of the bestrophin protein--functional implications. Hum Genet. 1999;104:383–9.
Maia-Lopes S, et al. Gene symbol: BEST1. Disease: Best macular dystrophy. Hum Genet. 2008;123:111.
Meunier I, et al. Systematic screening of BEST1 and PRPH2 in juvenile and adult vitelliform macular dystrophies: a rationale for molecular analysis. Ophthalmology. 2011;118:1130–6.
Renner AB, et al. Late onset is common in best macular dystrophy associated with VMD2 gene mutations. Ophthalmology. 2005;112:586–92.
Marquardt A, et al. Mutations in a novel gene, VMD2, encoding a protein of unknown properties cause juvenile-onset vitelliform macular dystrophy (Best’s disease). Hum Mol Genet. 1998;7:1517–25.
Kramer F, Mohr N, Kellner U, Rudolph G, Weber BH. Ten novel mutations in VMD2 associated with Best macular dystrophy (BMD). Hum Mutat. 2003;22:418.
Caldwell GM, et al. Bestrophin gene mutations in patients with Best vitelliform macular dystrophy. Genomics. 1999;58:98–101.
Glavac D, et al. Clinical and genetic heterogeneity in Slovenian patients with BEST disease. Acta Ophthalmol. 2016;94:e786–94.
Liu J, Zhang Y, Xuan Y, Liu W, Wang M. Novel BEST1 mutations and special clinical features of Best Vitelliform macular dystrophy. Ophthalmic Res. 2016;56:178–85.
Marchant D, et al. Identification of novel VMD2 gene mutations in patients with best vitelliform macular dystrophy. Hum Mutat. 2001;17:235.
White K, Marquardt A, Weber BH. VMD2 mutations in vitelliform macular dystrophy (Best disease) and other maculopathies. Hum Mutat. 2000;15:301–8.
Sodi A, et al. A novel mutation in the VMD2 gene in an Italian family with Best maculopathy. J Fr Ophtalmol. 2007;30:616–20.
Schatz P, et al. Evaluation of macular structure and function by OCT and electrophysiology in patients with vitelliform macular dystrophy due to mutations in BEST1. Invest Ophthalmol Vis Sci. 2010;51:4754–65.
Eksandh L, Bakall B, Bauer B, Wadelius C, Andreasson S. Best’s vitelliform macular dystrophy caused by a new mutation (Val89Ala) in the VMD2 gene. Ophthalmic Genet. 2001;22:107–15.
Li Y, et al. A novel mutation of the VMD2 gene in a Chinese family with best vitelliform macular dystrophy. Ann Acad Med Singap. 2006;35:408–10.
Marchant D, et al. New VMD2 gene mutations identified in patients affected by Best vitelliform macular dystrophy. J Med Genet. 2007;44:e70.
Palomba G, et al. A novel spontaneous missense mutation in VMD2 gene is a cause of a best macular dystrophy sporadic case. Am J Ophthalmol. 2000;129:260–2.
Wabbels B, Preising MN, Kretschmann U, Demmler A, Lorenz B. Genotype-phenotype correlation and longitudinal course in ten families with Best vitelliform macular dystrophy. Graefes Arch Clin Exp Ophthalmol. 2006;244:1453–66.
Lin Y, et al. Two novel mutations in the bestrophin-1 gene and associated clinical observations in patients with best vitelliform macular dystrophy. Mol Med Rep. 2015;12:2584–8.
Wittstrom E, et al. Morphological and functional changes in multifocal vitelliform retinopathy and biallelic mutations in BEST1. Ophthalmic Genet. 2011;32:83–96.
Sohn EH, et al. Phenotypic variability due to a novel Glu292Lys variation in exon 8 of the BEST1 gene causing Best macular dystrophy. Arch Ophthalmol. 2009;127:913–20.
Yanagi Y, Sekine H, Mori M. Identification of a novel VMD2 mutation in Japanese patients with Best disease. Ophthalmic Genet. 2002;23:129–33.
Arora R, et al. Unilateral BEST1-associated retinopathy. Am J Ophthalmol. 2016;169:24–32.
Marchant D, et al. Use of denaturing HPLC and automated sequencing to screen the VMD2 gene for mutations associated with Best’s vitelliform macular dystrophy. Ophthalmic Genet. 2002;23:167–74.
Sodi A, et al. BEST1 sequence variants in Italian patients with vitelliform macular dystrophy. Mol Vis. 2012;18:2736–48.
Lin Y, et al. Bestrophin 1 gene analysis and associated clinical findings in a Chinese patient with Best vitelliform macular dystrophy. Mol Med Rep. 2017;16:4751–5.
Lin Y, et al. Genetic variations in Bestrophin1 and associated clinical findings in two Chinese patients with juvenile-onset and adult-onset best vitelliform macular dystrophy. Mol Med Rep. 2017;17(1):225–33.
Tian L, et al. Screening of BEST1 gene in a Chinese cohort with Best Vitelliform macular dystrophy or autosomal recessive Bestrophinopathy. Invest Ophthalmol Vis Sci. 2017;58:3366–75.
Allikmets R, et al. Evaluation of the best disease gene in patients with age-related macular degeneration and other maculopathies. Hum Genet. 1999;104:449–53.
Wells J, et al. Mutations in the human retinal degeneration slow (RDS) gene can cause either retinitis pigmentosa or macular dystrophy. Nat Genet. 1993;3:213–8.
Manes G, et al. Mutations in IMPG1 cause vitelliform macular dystrophies. Am J Hum Genet. 2013;93:571–8.
Meunier I, et al. Frequency and clinical pattern of vitelliform macular dystrophy caused by mutations of interphotoreceptor matrix IMPG1 and IMPG2 genes. Ophthalmology. 2014;121:2406–14.
Mandai M, et al. Autologous induced stem-cell-derived retinal cells for macular degeneration. N Engl J Med. 2017;376:1038–46.
Maguire AM, et al. Safety and efficacy of gene transfer for Leber’s congenital amaurosis. N Engl J Med. 2008;358:2240–8.
Kim E, et al. In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni. Nat Commun. 2017;8:14500.
Kim K, et al. Genome surgery using Cas9 ribonucleoproteins for the treatment of age-related macular degeneration. Genome Res. 2017;27:419–26.
Suzuki K, et al. In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration. Nature. 2016;540:144–9.
Ruan GX, et al. CRISPR/Cas9-mediated genome editing as a therapeutic approach for Leber congenital Amaurosis 10. Mol Ther. 2017;25:331–41.
Huang X, et al. Genome editing abrogates angiogenesis in vivo. Nat Commun. 2017;8:112.
Gaudelli NM, et al. Programmable base editing of A*T to G*C in genomic DNA without DNA cleavage. Nature. 2017;51(7681):464–71.
Komor AC, Kim YB, Packer MS, Zuris JA, Liu DR. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature. 2016;533:420–4.
Yanik M, et al. In vivo genome editing as a potential treatment strategy for inherited retinal dystrophies. Prog Retin Eye Res. 2017;56:1–18.
Compliance with Ethical Requirements
Sung Wook Park, Chang ki Yoon, Dae Joong Ma, Un Chul Park, and Hyeong Gon Yu declare that they have no conflict of interest.
All procedures followed were in accordance with the ethical standards of the responsible committee on institutional review board and with the Helsinki Declaration of 1975, as revised in 2000. Informed consent was obtained from all patients for being included in the study.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Park, S.W., Yoon, C.K., Ma, D.J., Park, U.C., Yu, H.G. (2019). Clinical Genetics of Vitelliform Macular Dystrophy: An Asian Perspective. In: Prakash, G., Iwata, T. (eds) Advances in Vision Research, Volume II. Essentials in Ophthalmology. Springer, Singapore. https://doi.org/10.1007/978-981-13-0884-0_21
Download citation
DOI: https://doi.org/10.1007/978-981-13-0884-0_21
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-0883-3
Online ISBN: 978-981-13-0884-0
eBook Packages: MedicineMedicine (R0)