Abstract
Developing an effective prophylaxis HIV-1 vaccine is likely to require the elicitation of broadly neutralizing antibodies (bnAbs). As the HIV-1 envelope (Env) glycoprotein – the sole target of bnAbs – has evolved multiple mechanisms to evade antibody neutralization, the processes for bnAb generation are highly selective and time-consuming. Benefiting from antibody isolation technologies of single B cell culturing and direct single B cell sorting and cloning, a new generation of monoclonal bnAbs has been isolated since 2009, exhibiting remarkable breadths and potencies, thus breaking through a nearly 20-year-long limit of four monoclonal bnAbs with moderate breadth and potency. The discovery of a long list of monoclonal bnAbs has provided in-depth understanding of the sites of vulnerability on the HIV-1 Env and the complexity of human B cell immunology to generate such responses, thus presenting both guidance and challenges to move the Env immunogen design effort forward.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
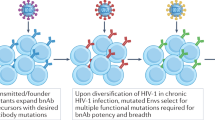
The development of HIV-1 broadly neutralizing antibodies (bnAbs) in a subset of infected individuals has demonstrated the ability of the human immune system to mount effective antibody responses against the virus (Simek et al. 2009; Doria-Rose et al. 2010). As the HIV-1 envelope (Env) glycoprotein is the only viral protein expressed on the surface of the virion, bnAbs target the HIV-1 Env to block viral infection. Because HIV-1 Env has evolved a number of mechanisms to evade antibody neutralization (Wei et al. 2003; Moore et al. 2009), the processes of bnAb generation are proven highly selective and time-consuming. Nonetheless, HIV-1 bnAbs with 50% breadth are developed in half of HIV-1 chronically infected individuals (Hraber et al. 2014), supporting the feasibility of inducing a similar spectrum of antibody responses with optimal Env immunogens. To inform Env immunogen design and improve our understanding of human B cell immunology to mount an effective antibody response, the isolation of monoclonal bnAbs has become critical.
Though high in demand, there were only four monoclonal HIV-1 bnAbs isolated before 2009, largely due to limitations in methods for monoclonal antibody (mAb) isolation and lack of knowledge for HIV-1 bnAbs at that time. Since 2009, improved methods of single B cell culturing and direct single B cell cloning were introduced and applied to HIV-1-infected samples, leading to a major breakthrough in identifying a new generation of bnAbs with remarkable breadths and potencies. The new era of HIV-1 monoclonal bnAb discovery and understanding of these antibodies opened up new avenues for basic research in immunology, structural biology, and vaccinology, as well as in translational and clinical research for HIV-1 infection. As we contributed to the isolation of the first HIV-1 monoclonal bnAb VRC01 by singe B cell sorting and cloning, I review the critical path and scientific thoughts that drove the progress of the project and eventually led to the discovery of this exciting antibody.
2 HIV-1 Neutralization by Polyclonal Sera
Sera or plasma profiling, as a straightforward analysis, usually takes place first by directly sampling the polyclonal antibodies in donor sera or plasma for binding specificities to antigens and for antibody functions against pathogens. Since the identification of HIV as the cause of AIDS in the early 1980s (Barre-Sinoussi et al. 1983; Gallo et al. 1983, 1984), polyclonal sera and plasma samples from HIV-infected individuals have been rigorously tested for binding specificities to various viral antigens and for activity to neutralize the virus. Initial neutralization studies used lab-adapted strains such as IIIB, MN, HXB2, and NL4-3 and found that sera and plasmas from vaccinated or infected donors potently neutralized lab-adapted strains (Mascola et al. 1994; Kovacs et al. 1993; Schwartz et al. 1993). However, subsequent studies using HIV-1 primary isolates showed high levels of resistance to sera and plasma neutralization from vaccinated and infected donors (Moore et al. 1995; Parren et al. 1998). This difference in neutralization sensitivity indicated that the Envs presented on the surface of lab-adapted strains were different from those presented on primary isolates. To categorize the neutralization sensitivity among lab-adapted strains and primary isolates, a “tiered” system (Seaman et al. 2010) was introduced with tier 1A representing the easy-to-neutralize lab-adapted strains, followed by tier 1B, then tier 2 representing the typical primary isolates, and finally tier 3 representing the most difficult-to-neutralize strains.
Tested against various panels of tier 2 strains, many chronically infected polyclonal sera and plasma samples have been ranked based on their neutralization breadths and potencies. Though most samples lacked activities to neutralize genetically diverse tier 2 strains, small factions (10–20%) were identified with such activities (Simek et al. 2009; Doria-Rose et al. 2010). These donors have been referred to as “broad neutralizers” and “elite neutralizers” if ranked in the top 1%. The identification of broad and elite neutralizers was exciting because they demonstrated that the human immune system can mount effective antibody responses against the virus, thus providing a scientific basis for Env immunogen design to elicit similar antibody responses to block infection. The first step to implement this strategy was to understand the antibody specificities that mediated the observed neutralization breadths, i.e., where did the antibodies bind on the HIV Env to block viral entry? As the plasma neutralization test itself did not inform antibody specificities or targets, the analyses followed were sera mapping (Walker et al. 2010; Binley et al. 2008; Gray et al. 2009). The caveat was that, because sera and plasmas are mixes of polyclonal antibodies, it has been extremely difficult to dissect complex antibody specificities and precisely map the epitopes and targets mediating neutralization. Before the isolation of monoclonal bnAbs, there had been an extensive debate on whether polyclonal antibodies or mAbs mediated neutralization breadth. To address this question and to precisely map the epitopes targeted by bnAbs, the solution was to isolate monoclonal bnAbs. For this purpose, samples from the identified broad and elite neutralizers received the highest priority and were selected for the isolation of mAbs that could account for donor’s serological activities.
3 mAb Isolation and the First Generation of Monoclonal bnAbs
Before 2009 only four mAbs, b12, 2G12, 2F5, and 4E10, all isolated in the early 1990s (Table 3.1), demonstrated superior neutralization spectrums over the other mAbs available at that time. For nearly 20 years, these four mAbs had been used for almost all bnAb-related basic, pre-clinical and clinical research, including epitope mapping, Env structural analysis, Env immunogen design, passive immunization in nonhuman primates (Mascola et al. 1999, 2000), and a clinical trial to treat HIV-infected individuals (Trkola et al. 2005). Because of their wide uses for such a prolonged period, the four mAbs became well known for their role in defining HIV-1 monoclonal bnAbs. Given the fact that a mAb only binds to a single epitope on the Env antigen, a single mAb may not be expected to neutralize a broad range of circulating strains. Based on a conservative expectation, the relatively narrow breadths (30–60%) and weak potencies (mean or median IC50s >2 μg/ml) of the four mAbs (Walker et al. 2009) were accepted by the research community as the limit of neutralization for a single mAb. Meanwhile, the breadths and potencies displayed by a number of well characterized broadly neutralizing plasmas from HIV-1 chronically infected donors (Li et al. 2007), including the NIH donor 45 from whom VRC01 was later isolated, greatly exceeded those displayed by the four mAbs. Thus, there was a discrepancy in the observed neutralization breadths between the polyclonal plasmas and the four known monoclonal bnAbs. Because of this discrepancy, HIV neutralization breadth was thought to be mediated by polyclonal antibodies and not by mAbs. This view would disfavor vaccine development strategies to target a conserved site or site of vulnerability on the HIV Env. To address the discrepancy between broadly neutralizing plasmas and few known monoclonal bnAbs, it was necessary to identify additional monoclonal bnAbs, should they exist.
Historically mAbs were isolated by phage display, Epstein-Barr virus (EBV) transformation, electrofusion, and hybridomas. Despite limitations and weaknesses, these technologies dominated the field since their discoveries in the 1970s to 1990s. Before 2009, HIV-1-specific mAbs had been isolated by a few labs that specialize in mAb isolation using these technologies. Disappointingly these technologies suffered from low efficiency when applied to HIV-1-infected samples. Another limiting factor was the mAb screening process that was not based on virus neutralization but instead was based on binding affinities to gp120 monomers, gp120 peptides, or gp41 peptides, thus further reducing the number of bnAbs yielded from these efforts. Despite numbering only four, the discoveries of monoclonal bnAbs still provided tremendous knowledge about HIV-1 neutralization. For example, epitope mapping indicated that b12 targets the CD4-binding site (CD4bs) on gp120, that 2G12 targets a cluster of glycans on gp120, and that 2F5 and 4E10 each binds to a peptide in the membrane-proximal external region (MPER) of gp41. Therefore, we learned that the CD4bs, a cluster of gp120 glycans, and MPER are sites of vulnerability on the HIV-1 Env. Additionally, passive immunizations using a single or combinations among the four mAbs demonstrated protection in nonhuman primate models (Mascola et al. 1999, 2000), thus providing a basis for developing Env immunogens aiming to elicit similar bnAbs. The four monoclonal bnAbs also provided antibody sequences that hinted at common genetic features of HIV-1 bnAbs such as high levels of somatic hypermutation and relatively long heavy chain CDR3 lengths. Nonetheless, after intensive research on the four mAbs for such a long period of time, there was an increasing urgency to identify additional monoclonal bnAbs to verify and address critical questions such as: (1) Were the four monoclonal bnAbs generalizable to other infected donors? If so, could similar mAbs be identified from those donors? (2) Were there mAbs that account for neutralization breadths and potencies observed in donor plasmas? (3) Was HIV-1 neutralization breadth mediated by polyclonal antibodies or mAbs?
4 Sera Mapping and the CD4-Binding Site (CD4bs)
If HIV-1 neutralization breadth were mediated by polyclonal antibodies and not by mAbs, searches for monoclonal bnAbs would likely fail. Though the chance of success was small, there had been sera mapping data strongly supporting the presence of monoclonal bnAbs in some chronically infected donors. Using antibody adsorption and elution from gp120 proteins with and without a point mutation D368R that knocks out CD4 binding, Li et al. mapped the neutralizing specificities of two broadly reactive sera, the NIH donors 1 and 45, to the CD4bs (Li et al. 2007). The fractioned plasmas were tested for neutralization against four HIV-1 strains, with similar results obtained. Therefore, antibodies bound to the D368 site neutralized at least four genetically distant strains, supporting an antibody specificity that targeted the conserved D368 residue and mediated neutralization against different strains.
The rationale for sera mapping studies focusing on the CD4bs was based on the fact that HIV depends on CD4 to initiate viral entry and infection; thus the CD4bs must be functionally conserved. Structurally the CD4bs has been defined at the atomic level in a liganded complex of a gp120 core bound with a 2-domain soluble CD4, along with a mAb 17b antigen-binding fragment (Fab) directed at the co-receptor binding site (Kwong et al. 1998). The crystal structure revealed three domains of gp120 core, an inner domain, an outer domain, and a bridging sheet that connects the two. The gp120 residues in contact with CD4 were discontinuous and spread out in all three domains but were more concentrated in the outer domain. Notably the spans of 365–371, later termed the CD4-binding loop, and 425–430 at β20–21, as part of the bridging sheet, contributed 57% of the total CD4 contact area on gp120 (Kwong et al. 1998). The residue D368 in the middle of the CD4-binding loop was identified as one of the key contact residues for CD4 interaction, and the D368R mutation specifically knocked out CD4 binding. Sera mapping using gp120 proteins with and without the D368R point mutation indicated that novel antibodies to this site were elicited in some HIV-1-infected individuals, and exposure of this conserved site to memory B cells in these individuals might probe the B cells expressing such antibodies. The CD4bs-targeting monoclonal bnAb b12 (Burton et al. 1994) also supported the CD4bs as a site of vulnerability.
5 b12
The mAb b12 was isolated in the year 1994 by phage display using PBMCs from an HIV-1 clade B-infected individual (Burton et al. 1994). The phage display library lost antibody heavy and light chain pairing information and produced Fabs with randomly paired heavy and light chains, thus capturing antibodies not “naturally” produced in donor plasmas. This had raised questions and criticism in the field. Moreover, panning of phage display libraries relied on monomeric gp120 binding. Hence, the phage display effort largely yielded mAbs that were capable of binding to monomeric gp120 but not necessarily neutralizing the virus. Nonetheless, mAb b12 exhibited an overall 35% neutralization breadth against cross-clade HIV strains and a preferential 58% breadth against clade B strains (Walker et al. 2009). Competition ELISA with CD4-Ig indicated that b12 efficiently competed with CD4 to bind to gp120, suggesting that the b12 epitope overlaps with the CD4bs (Moore and Sodroski 1996).
The b12 Fab atomic level crystal structure in complex with HXB2 core was solved in 2007, revealing that only the b12 heavy chain interacted with gp120 and that the vast majority of b12 epitope was on the outer domain (Zhou et al. 2007). The b12 heavy chain CDR1, CDR2, and CDR3 “grasped” all around the CD4-binding loop, making direct contact with each of the 10 consecutive residues from 364 to 373. As expected, the D368R mutation in the CD4-binding loop knocked out b12 binding. Though both primarily targeted the CD4-binding loop, CD4 only formed contacts with one side (six residues), yet b12 grasped on both sides (all ten residues) of the CD4-binding loop (Zhou et al. 2007). Therefore, b12-escape mutations commonly arose on the four residues dispensable for CD4 interaction in the CD4-binding loop (Wu et al. 2009). When the footprints of b12 and CD4 were superimposed on gp120 core, they overlapped primarily on the outer domain, defining an appealing target that was functionally conserved, structurally stable, and antibody accessible. Therefore, new designs of gp120 core proteins to specifically present this region of gp120 to memory B cells in chronically infected donors might probe the B cells expressing antibodies targeting this region. With a powerful single B cell sorting and cloning platform introduced to the field (Tiller et al. 2008; Scheid et al. 2009), new opportunities for functional mAb discovery appeared on the horizon particularly suited to the quest of identifying novel monoclonal bnAbs against HIV.
6 VRC01 and Its Class
The platform of single B cell sorting and cloning was first used to recover influenza-specific mAbs from the plasmablasts of vaccinated individuals (Wrammert et al. 2008) and had not been applied to HIV-infected samples before 2009. With PBMCs from broad neutralizers and a Yu2 gp140 foldon trimer protein readily available, the single B cell sorting platform was first applied and attempted to isolate novel monoclonal bnAbs in 2009. Using the Yu2 gp140 as bait, Scheid et al. processed PBMC samples from four broad neutralizers, including the NIH donor 45, and recovered more than 500 gp140-binding mAbs (Scheid et al. 2009). However, none of the isolated mAbs was broadly neutralizing against HIV-1. It was unclear at that time why the recovered gp140-binding mAbs were not broadly neutralizing, and this result would have supported the view that polyclonal antibodies but not mAbs mediate HIV-1 neutralization breadth. Though inexperienced, our group decided to test the single B cell sorting and cloning platform in our hands using a modified gp120 core as bait.
As mentioned above, the overlapping footprint of b12 and CD4 on gp120 core revealed the portion of CD4bs on the outer domain to be functionally conserved, structurally stable, and antibody accessible; thus new designs of gp120 core proteins to specifically present this region might specifically probe the B cells expressing antibodies targeting this region. With designs from William Schief at the Scripps Institute, Yang and colleagues at the Vaccine Research Center expressed and tested a series of proteins that preserved the part of CD4bs on the outer domain but altered the rest of the protein surface to non-HIV-1 (Wu et al. 2010). These modified gp120 core proteins were termed resurfaced stabilized core (RSC) proteins. Among the expressed RSC proteins, RSC3 performed the best in retaining b12 binding but reducing binding to other non-neutralizing mAbs. Because the inner domain and bridging sheet were altered, RSC3 lost stable binding to CD4, thus abrogating CD4 interference. Using RSC3 along with a negative control ΔRSC3, which deleted a single residue I371 in the CD4-binding loop, we stained a PBMC sample from the NIH donor 45 and sorted individual IgG-expressing B cells that stained RSC3 + ΔRSC3-. From the sorted individual B cells, we recovered three monoclonal bnAbs, VRC01, VRC02, and VRC03, which belonged to the same B cell lineage and same class of bnAbs – the VRC01-class – and recapitulated the RSC3 + ΔRSC3-binding profile (Wu et al. 2010).
Because the prototype mAb for RSC3 design was b12, we compared the gp120 binding and viral neutralization profiles of VRC01 to b12 (Wu et al. 2010). VRC01 clearly targeted the CD4bs, displaying all known binding characteristics of mAbs directed to the CD4bs, including efficient competition with CD4-Ig and b12 and reduced binding with the D368R mutation. Importantly, VRC01 exhibited superior neutralization breadth and potency than b12 (Fig. 3.1), reaching an overall breadth of 90% with a geometric mean IC50 of 0.33 µg/ml against global circulating strains. Furthermore, VRC01 accounted for a major fraction of the neutralization activities measured in the donor plasma (Wu et al. 2010). Therefore the isolation of VRC01 supported the presence of highly conserved sites of vulnerability on the HIV-1 Env and that neutralization breadths observed in the donor plasma were in a large part mediated by mAbs. Crystal structure analysis of the VRC01 Fab in complex with gp120 core revealed that the VRC01 epitope overlapped with CD4 on the outer domain (Zhou et al. 2010), precisely targeting the site that RSC3 was designed to expose to B cells. Therefore, the RSC3 bait and VRC01 isolation demonstrated an example of successful protein design and engineering. The atomic structure also indicated that the heavy chain CDR2 of VRC01 partially mimicked the CD4 interaction with gp120, forming contacts with only one side of the CD4-binding loop, thus avoiding escape mutations arising from the other side of the CD4-binding loop, a strategy exploited by the virus to escape b12. This mimicry of CD4 by VRC01 partially explained its broader neutralization spectrum than that of b12. Later studies determined that the virus mainly mutates gp120 loop D and V5 to escape VRC01 neutralization (Li et al. 2011).
Neutralization breadth (y-axis) is plotted at the corresponding potency (x-axis) for VRC01 and b12 using the IC50 data from Wu et al. 2010 against a total of 190 HIV-1 Env-pseudotyped viruses representing strains circulating globally
Following the successful isolation of VRC01 by single B cell sorting and cloning with a specific Env bait, more HIV-1 monoclonal bnAbs have been isolated using similar methods. From the B cells sorted with Yu2 gp140, re-PCRs with improved primers yielded VRC01-class of monoclonal bnAbs (Scheid et al. 2011), indicating that the PCR primers used in the first study missed these highly mutated bnAbs because the original primers did not account for possible mutations at the start of the mAb-coding region. To date the VRC01-class of bnAbs has been isolated from more than 10 donors (Table 3.1), rendering it a category of antibody response generalizable across individuals.
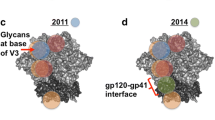
In addition to the VRC01-class, six other classes of bnAbs have been identified that target the CD4bs as exemplified by mAbs HJ16, 8ANC131, CH103, VRC13, VRC16, and 179NC75 (Table 3.1). A collection of crystal structures of the CD4bs-targeting bnAbs in complex with gp120 cores revealed substantially overlapping epitopes and different modes of gp120 recognition (Zhou et al. 2013; Zhou et al. 2015), with the VRC01-class and 8ANC131-class partially mimicking the CD4 interaction with gp120. Though the CD4bs itself is not glycosylated, it is surrounded by glycans, and the bnAbs targeting this site all avoided or accommodated glycans to reach their epitopes. Also, because the CD4bs of gp120 is always readily available to interact with CD4, the Env trimer packaging and conformational changes have minimal impact on bnAbs that target this site. However, this is not the case for non-neutralizing mAbs. Since current vaccines induced non-neutralizing mAbs to the CD4bs, it will be important to consider and adress how to modify and adjust the mode or angle of antibody binding to gp120 in Env immunogen design.
7 Antibody Genetic Composition and Next-Generation Sequencing (NGS)
Because each unique B cell receptor has its own VDJ recombined composition, bnAb nucleotide sequences are essential to establish the corresponding B cell lineages. For mAbs identified by single B cell cloning, their variable region nucleotide sequences are obtained for paired heavy and light chains. As illustrated in Fig. 3.2 using heavy chain as an example, mAbs are usually characterized by their V-gene usage, somatic hypermutation, and CDR3 length. While the antibody’s primary antigen interacting regions CDR1 and CDR2 are coded within the V-gene, its CDR3 is composed of VD junction, D gene, DJ junction, and part of the J gene. Thus, CDR3 is the most diverse region of the antibody. The VRC01-class of bnAbs from multiple donors shared a sequence signature of IGHV1-2 gene usage, high levels of somatic hypermutation, and a short light chain CDR3 length of five amino acids (Zhou et al. 2013; West et al. 2012). This distinct sequence signature has been used to select naïve B cells with the potential to evolve and mature to become the VRC01-class of bnAbs (Jardine et al. 2016).
A schematic of the gene structure for antibody heavy chain showing characteristic elements of V-gene usage, somatic hypermutation, and CDR3. For repertoire studies using next-generation sequencing (NGS), reverse primers annealing to the start of the constant region distinguish between μ chain for IgM, γ chain for IgG, and α chain for IgA
As bnAbs are part of the donor antibody repertoire, the availability of bnAb sequences has inspired and stimulated antibody repertoire analyses by next-generation sequencing (NGS) including IgM, IgG, and IgA (Fig. 3.2) in HIV-1-infected and HIV-1-uninfected donors (Chen et al. 2012; Xiao et al. 2013; Yin et al. 2013; Zhang et al. 2013; He et al. 2014). The NGS data from longitudinal samples of the same donor also allowed for tracking bnAb lineages (Wu et al. 2015; Doria-Rose et al. 2014; Bonsignori et al. 2016; van Gils et al. 2016; MacLeod et al. 2016). However, a caveat to the currently available sequencing platforms is the loss of paired heavy and light chains, and thus antibodies cannot be reconstituted with naturally paired heavy and light chains based on the NGS data alone. Though not yet readily available, new sequencing platforms are being developed to address this issue (DeKosky et al. 2013, 2015). As of today, NGS combining with single B cell sorting and sequencing, which maintains the heavy and light chain pairing information, is still the most practical and comprehensive system to study B cell lineages of interest.
8 Other HIV-1 bnAbs
Pioneered by the Burton group using individual B cell cultures with micro-neutralization screening to isolate monoclonal bnAbs PG9 and PG16 (Walker et al. 2009), a similar system has been applied by other laboratories to isolate many more HIV-1 bnAbs targeting Env regions outside of the CD4bs. Along with single B cell sorting using soluble or cellular gp140 trimer and virus-like particle (VLP) baits (Table 3.1), HIV-1 bnAbs targeting Env regions outside of CD4bs have also been isolated by the single B cell sorting and cloning platform. HIV-1 bnAbs have been extensively reviewed in the literature (Mascola and Haynes 2013; Kwong and Mascola 2012; Kwong et al. 2013; Wu and Kong 2016). Briefly, there are currently seven known categories of bnAbs targeting different regions of the HIV-1 Env. From the Env trimer apex to gp41, known sites of vulnerability include (1) the V1 V2 apex targeted by PG9, CAP256-VRC26, PGT145, N90-VRC38, and others; (2) a gp120 glycan cluster targeted by 2G12; (3) the V3 base glycans targeted by PGT121, PGT128, and others; (4) the CD4bs targeted by VRC01 and others; (5) the gp120 and gp41 interface targeted by 35O22 and others; (6) the fusion peptide targeted by PGT151, VRC34, and ACS202; and (7) the MPER targeted by 10E8, 2F5, and 4E10. These bnAbs play a central role in informing and guiding HIV-1 Env immunogen designs, including defining conserved sites of vulnerability on HIV-1 Env and validating Env immunogens for proper presentation of intact bnAb epitopes. Although still a work in progress, sequential Env variants that evolved during the natural course of infection and stimulated bnAb responses are being identified and cloned (Gao et al. 2014; Bhiman et al. 2015; MacLeod et al. 2016), and the identification of these Envs would rely on the identification of both autologous and heterologous neutralizing antibodies. Additional studies have focused on longitudinal analyses to track bnAb lineages and delineate their maturation pathways (Doria-Rose et al. 2014; Bonsignori et al. 2016; van Gils et al. 2016; MacLeod et al. 2016). As single B cell sorting and cloning has been increasingly included in post-immune analyses (Sundling et al. 2012, 2014; Navis et al. 2014), a relatively complete list of bnAb genetic compositions has been highly informative regarding whether similar bnAb precursors have been engaged by the tested immunizations. The broadest and most potent bnAbs have also been high in demand for research in passive immunization and in HIV-1 treatment and cure.
9 Conclusion
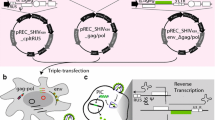
With 41 unique bnAbs isolated to date (Table 3.1) that define 7 general categories of effective targets on the HIV-1 Env, current research has moved on to unveil the mechanisms of human B cell responses that produce bnAbs and to investigate the entire bnAb maturation pathway starting from naïve B cell engagement. The pathways for bnAb generation remain mysterious because current knowledge of basic B cell immunology has been gained primarily from mouse studies, yet mouse may not possess the proper B cell repertoire to generate similar highly functional antibodies against HIV (Hu et al. 2015). Since SHIV-infected nonhuman primates have demonstrated the ability to produce HIV-1 bnAbs (Jia et al. 2016; Walker et al. 2011b; Shingai et al. 2012), emphasis on nonhuman primate models may hold promise to elucidate details of HIV-1 bnAb production in vivo and fill gaps of knowledge in this regard.
References
Barre-Sinoussi F, Chermann JC, Rey F, Nugeyre MT, Chamaret S, Gruest J, Dauguet C, Axler-Blin C, Vezinet-Brun F, Rouzioux C, Rozenbaum W, Montagnier L (1983) Isolation of a T-lymphotropic retrovirus from a patient at risk for acquired immune deficiency syndrome (AIDS). Science 220(4599):868–871
Bhiman JN, Anthony C, Doria-Rose NA, Karimanzira O, Schramm CA, Khoza T, Kitchin D, Botha G, Gorman J, Garrett NJ, Abdool Karim SS, Shapiro L, Williamson C, Kwong PD, Mascola JR, Morris L, Moore PL (2015) Viral variants that initiate and drive maturation of V1V2-directed HIV-1 broadly neutralizing antibodies. Nat Med 21(11):1332–1336. https://doi.org/10.1038/nm.3963
Binley JM, Lybarger EA, Crooks ET, Seaman MS, Gray E, Davis KL, Decker JM, Wycuff D, Harris L, Hawkins N, Wood B, Nathe C, Richman D, Tomaras GD, Bibollet-Ruche F, Robinson JE, Morris L, Shaw GM, Montefiori DC, Mascola JR (2008) Profiling the specificity of neutralizing antibodies in a large panel of plasmas from patients chronically infected with human immunodeficiency virus type 1 subtypes B and C. J Virol 82(23):11651–11668. https://doi.org/10.1128/JVI.01762-08
Bonsignori M, Hwang KK, Chen X, Tsao CY, Morris L, Gray E, Marshall DJ, Crump JA, Kapiga SH, Sam NE, Sinangil F, Pancera M, Yongping Y, Zhang B, Zhu J, Kwong PD, O’Dell S, Mascola JR, Wu L, Nabel GJ, Phogat S, Seaman MS, Whitesides JF, Moody MA, Kelsoe G, Yang X, Sodroski J, Shaw GM, Montefiori DC, Kepler TB, Tomaras GD, Alam SM, Liao HX, Haynes BF (2011) Analysis of a clonal lineage of HIV-1 envelope V2/V3 conformational epitope-specific broadly neutralizing antibodies and their inferred unmutated common ancestors. J Virol 85(19):9998–10009. https://doi.org/10.1128/JVI.05045-11
Bonsignori M, Zhou T, Sheng Z, Chen L, Gao F, Joyce MG, Ozorowski G, Chuang GY, Schramm CA, Wiehe K, Alam SM, Bradley T, Gladden MA, Hwang KK, Iyengar S, Kumar A, Lu X, Luo K, Mangiapani MC, Parks RJ, Song H, Acharya P, Bailer RT, Cao A, Druz A, Georgiev IS, Kwon YD, Louder MK, Zhang B, Zheng A, Hill BJ, Kong R, Soto C, Mullikin JC, Douek DC, Montefiori DC, Moody MA, Shaw GM, Hahn BH, Kelsoe G, Hraber PT, Korber BT, Boyd SD, Fire AZ, Kepler TB, Shapiro L, Ward AB, Mascola JR, Liao HX, Kwong PD, Haynes BF (2016) Maturation pathway from germline to broad HIV-1 neutralizer of a CD4-mimic antibody. Cell 165:449. https://doi.org/10.1016/j.cell.2016.02.022
Buchacher A, Predl R, Strutzenberger K, Steinfellner W, Trkola A, Purtscher M, Gruber G, Tauer C, Steindl F, Jungbauer A et al (1994) Generation of human monoclonal antibodies against HIV-1 proteins; electrofusion and Epstein-Barr virus transformation for peripheral blood lymphocyte immortalization. AIDS Res Hum Retrovir 10(4):359–369. https://doi.org/10.1089/aid.1994.10.359
Burton DR, Pyati J, Koduri R, Sharp SJ, Thornton GB, Parren PW, Sawyer LS, Hendry RM, Dunlop N, Nara PL et al (1994) Efficient neutralization of primary isolates of HIV-1 by a recombinant human monoclonal antibody. Science 266(5187):1024–1027
Cale EM, Gorman J, Radakovich NA, Crooks ET, Osawa K, Tong T, Li J, Nagarajan R, Ozorowski G, Ambrozak DR, Asokan M, Bailer RT, Bennici AK, Chen X, Doria-Rose NA, Druz A, Feng Y, Joyce MG, Louder MK, O’Dell S, Oliver C, Pancera M, Connors M, Hope TJ, Kepler TB, Wyatt RT, Ward AB, Georgiev IS, Kwong PD, Mascola JR, Binley JM (2017) Virus-like particles identify an HIV V1V2 apex-binding neutralizing antibody that lacks a protruding loop. Immunity 46(5):777–791 e710. https://doi.org/10.1016/j.immuni.2017.04.011
Chen W, Prabakaran P, Zhu Z, Feng Y, Streaker ED, Dimitrov DS (2012) Characterization of human IgG repertoires in an acute HIV-1 infection. Exp Mol Pathol 93(3):399–407. https://doi.org/10.1016/j.yexmp.2012.09.022
Corti D, Langedijk JP, Hinz A, Seaman MS, Vanzetta F, Fernandez-Rodriguez BM, Silacci C, Pinna D, Jarrossay D, Balla-Jhagjhoorsingh S, Willems B, Zekveld MJ, Dreja H, O’Sullivan E, Pade C, Orkin C, Jeffs SA, Montefiori DC, Davis D, Weissenhorn W, McKnight A, Heeney JL, Sallusto F, Sattentau QJ, Weiss RA, Lanzavecchia A (2010) Analysis of memory B cell responses and isolation of novel monoclonal antibodies with neutralizing breadth from HIV-1-infected individuals. PLoS One 5(1):e8805. https://doi.org/10.1371/journal.pone.0008805
DeKosky BJ, Ippolito GC, Deschner RP, Lavinder JJ, Wine Y, Rawlings BM, Varadarajan N, Giesecke C, Dorner T, Andrews SF, Wilson PC, Hunicke-Smith SP, Willson CG, Ellington AD, Georgiou G (2013) High-throughput sequencing of the paired human immunoglobulin heavy and light chain repertoire. Nat Biotechnol 31(2):166–169. https://doi.org/10.1038/nbt.2492
DeKosky BJ, Kojima T, Rodin A, Charab W, Ippolito GC, Ellington AD, Georgiou G (2015) In-depth determination and analysis of the human paired heavy- and light-chain antibody repertoire. Nat Med 21(1):86–91. https://doi.org/10.1038/nm.3743
Doria-Rose NA, Klein RM, Daniels MG, O’Dell S, Nason M, Lapedes A, Bhattacharya T, Migueles SA, Wyatt RT, Korber BT, Mascola JR, Connors M (2010) Breadth of human immunodeficiency virus-specific neutralizing activity in sera: clustering analysis and association with clinical variables. J Virol 84(3):1631–1636. https://doi.org/10.1128/JVI.01482-09
Doria-Rose NA, Schramm CA, Gorman J, Moore PL, Bhiman JN, DeKosky BJ, Ernandes MJ, Georgiev IS, Kim HJ, Pancera M, Staupe RP, Altae-Tran HR, Bailer RT, Crooks ET, Cupo A, Druz A, Garrett NJ, Hoi KH, Kong R, Louder MK, Longo NS, McKee K, Nonyane M, O’Dell S, Roark RS, Rudicell RS, Schmidt SD, Sheward DJ, Soto C, Wibmer CK, Yang Y, Zhang Z, Mullikin JC, Binley JM, Sanders RW, Wilson IA, Moore JP, Ward AB, Georgiou G, Williamson C, Abdool Karim SS, Morris L, Kwong PD, Shapiro L, Mascola JR (2014) Developmental pathway for potent V1V2-directed HIV-neutralizing antibodies. Nature 509(7498):55–62. https://doi.org/10.1038/nature13036
Falkowska E, Le KM, Ramos A, Doores KJ, Lee JH, Blattner C, Ramirez A, Derking R, van Gils MJ, Liang CH, McBride R, von Bredow B, Shivatare SS, Wu CY, Chan-Hui PY, Liu Y, Feizi T, Zwick MB, Koff WC, Seaman MS, Swiderek K, Moore JP, Evans D, Paulson JC, Wong CH, Ward AB, Wilson IA, Sanders RW, Poignard P, Burton DR (2014) Broadly neutralizing HIV antibodies define a glycan-dependent epitope on the prefusion conformation of gp41 on cleaved envelope trimers. Immunity 40(5):657–668. https://doi.org/10.1016/j.immuni.2014.04.009
Freund NT, Horwitz JA, Nogueira L, Sievers SA, Scharf L, Scheid JF, Gazumyan A, Liu C, Velinzon K, Goldenthal A, Sanders RW, Moore JP, Bjorkman PJ, Seaman MS, Walker BD, Klein F, Nussenzweig MC (2015) A new glycan-dependent CD4-binding site neutralizing antibody exerts pressure on HIV-1 in vivo. PLoS Pathog 11(10):e1005238. https://doi.org/10.1371/journal.ppat.1005238
Gallo RC, Sarin PS, Gelmann EP, Robert-Guroff M, Richardson E, Kalyanaraman VS, Mann D, Sidhu GD, Stahl RE, Zolla-Pazner S, Leibowitch J, Popovic M (1983) Isolation of human T-cell leukemia virus in acquired immune deficiency syndrome (AIDS). Science 220(4599):865–867
Gallo RC, Salahuddin SZ, Popovic M, Shearer GM, Kaplan M, Haynes BF, Palker TJ, Redfield R, Oleske J, Safai B et al (1984) Frequent detection and isolation of cytopathic retroviruses (HTLV-III) from patients with AIDS and at risk for AIDS. Science 224(4648):500–503
Gao F, Bonsignori M, Liao HX, Kumar A, Xia SM, Lu X, Cai F, Hwang KK, Song H, Zhou T, Lynch RM, Alam SM, Moody MA, Ferrari G, Berrong M, Kelsoe G, Shaw GM, Hahn BH, Montefiori DC, Kamanga G, Cohen MS, Hraber P, Kwong PD, Korber BT, Mascola JR, Kepler TB, Haynes BF (2014) Cooperation of B cell lineages in induction of HIV-1-broadly neutralizing antibodies. Cell 158(3):481–491. https://doi.org/10.1016/j.cell.2014.06.022
Georgiev IS, Doria-Rose NA, Zhou T, Kwon YD, Staupe RP, Moquin S, Chuang GY, Louder MK, Schmidt SD, Altae-Tran HR, Bailer RT, McKee K, Nason M, O’Dell S, Ofek G, Pancera M, Srivatsan S, Shapiro L, Connors M, Migueles SA, Morris L, Nishimura Y, Martin MA, Mascola JR, Kwong PD (2013) Delineating antibody recognition in polyclonal sera from patterns of HIV-1 isolate neutralization. Science 340(6133):751–756. https://doi.org/10.1126/science.1233989
Gray ES, Taylor N, Wycuff D, Moore PL, Tomaras GD, Wibmer CK, Puren A, DeCamp A, Gilbert PB, Wood B, Montefiori DC, Binley JM, Shaw GM, Haynes BF, Mascola JR, Morris L (2009) Antibody specificities associated with neutralization breadth in plasma from human immunodeficiency virus type 1 subtype C-infected blood donors. J Virol 83(17):8925–8937. https://doi.org/10.1128/JVI.00758-09
He L, Sok D, Azadnia P, Hsueh J, Landais E, Simek M, Koff WC, Poignard P, Burton DR, Zhu J (2014) Toward a more accurate view of human B-cell repertoire by next-generation sequencing, unbiased repertoire capture and single-molecule barcoding. Sci Rep 4:6778. https://doi.org/10.1038/srep06778
Hraber P, Seaman MS, Bailer RT, Mascola JR, Montefiori DC, Korber BT (2014) Prevalence of broadly neutralizing antibody responses during chronic HIV-1 infection. AIDS 28(2):163–169. https://doi.org/10.1097/QAD.0000000000000106
Hu JK, Crampton JC, Cupo A, Ketas T, van Gils MJ, Sliepen K, de Taeye SW, Sok D, Ozorowski G, Deresa I, Stanfield R, Ward AB, Burton DR, Klasse PJ, Sanders RW, Moore JP, Crotty S (2015) Murine antibody responses to cleaved soluble HIV-1 envelope trimers are highly restricted in specificity. J Virol 89(20):10383–10398. https://doi.org/10.1128/JVI.01653-15
Huang J, Ofek G, Laub L, Louder MK, Doria-Rose NA, Longo NS, Imamichi H, Bailer RT, Chakrabarti B, Sharma SK, Alam SM, Wang T, Yang Y, Zhang B, Migueles SA, Wyatt R, Haynes BF, Kwong PD, Mascola JR, Connors M (2012) Broad and potent neutralization of HIV-1 by a gp41-specific human antibody. Nature 491(7424):406–412. https://doi.org/10.1038/nature11544
Huang J, Kang BH, Pancera M, Lee JH, Tong T, Feng Y, Imamichi H, Georgiev IS, Chuang GY, Druz A, Doria-Rose NA, Laub L, Sliepen K, van Gils MJ, de la Pena AT, Derking R, Klasse PJ, Migueles SA, Bailer RT, Alam M, Pugach P, Haynes BF, Wyatt RT, Sanders RW, Binley JM, Ward AB, Mascola JR, Kwong PD, Connors M (2014) Broad and potent HIV-1 neutralization by a human antibody that binds the gp41-gp120 interface. Nature 515(7525):138–142. https://doi.org/10.1038/nature13601
Huang J, Kang BH, Ishida E, Zhou T, Griesman T, Sheng Z, Wu F, Doria-Rose NA, Zhang B, McKee K, O’Dell S, Chuang GY, Druz A, Georgiev IS, Schramm CA, Zheng A, Joyce MG, Asokan M, Ransier A, Darko S, Migueles SA, Bailer RT, Louder MK, Alam SM, Parks R, Kelsoe G, Von Holle T, Haynes BF, Douek DC, Hirsch V, Seaman MS, Shapiro L, Mascola JR, Kwong PD, Connors M (2016) Identification of a CD4-binding-site antibody to HIV that evolved near-pan neutralization breadth. Immunity 45(5):1108–1121. https://doi.org/10.1016/j.immuni.2016.10.027
Jardine JG, Kulp DW, Havenar-Daughton C, Sarkar A, Briney B, Sok D, Sesterhenn F, Ereno-Orbea J, Kalyuzhniy O, Deresa I, Hu X, Spencer S, Jones M, Georgeson E, Adachi Y, Kubitz M, deCamp AC, Julien JP, Wilson IA, Burton DR, Crotty S, Schief WR (2016) HIV-1 broadly neutralizing antibody precursor B cells revealed by germline-targeting immunogen. Science 351(6280):1458–1463. https://doi.org/10.1126/science.aad9195
Jia M, Lu H, Markowitz M, Cheng-Mayer C, Wu X (2016) Development of broadly neutralizing antibodies and their mapping by monomeric gp120 in human immunodeficiency virus type 1-infected humans and simian-human immunodeficiency virus SHIVSF162P3N-infected macaques. J Virol 90(8):4017–4031. https://doi.org/10.1128/JVI.02898-15
Klein F, Gaebler C, Mouquet H, Sather DN, Lehmann C, Scheid JF, Kraft Z, Liu Y, Pietzsch J, Hurley A, Poignard P, Feizi T, Morris L, Walker BD, Fatkenheuer G, Seaman MS, Stamatatos L, Nussenzweig MC (2012) Broad neutralization by a combination of antibodies recognizing the CD4 binding site and a new conformational epitope on the HIV-1 envelope protein. J Exp Med 209(8):1469–1479. https://doi.org/10.1084/jem.20120423
Kong L, Ju B, Chen Y, He L, Ren L, Liu J, Hong K, Su B, Wang Z, Ozorowski G, Ji X, Hua Y, Deller MC, Hao Y, Feng Y, Garces F, Wilson R, Dai K, O’Dell S, McKee K, Mascola JR, Ward AB, Wyatt RT, Li Y, Wilson IA, Zhu J, Shao Y (2016a) Key gp120 Glycans pose roadblocks to the rapid development of VRC01-class antibodies in an HIV-1-infected Chinese donor. Immunity 44:939. https://doi.org/10.1016/j.immuni.2016.03.006
Kong R, Xu K, Zhou T, Acharya P, Lemmin T, Liu K, Ozorowski G, Soto C, Taft JD, Bailer RT, Cale EM, Chen L, Choi CW, Chuang GY, Doria-Rose NA, Druz A, Georgiev IS, Gorman J, Huang J, Joyce MG, Louder MK, Ma X, McKee K, O’Dell S, Pancera M, Yang Y, Blanchard SC, Mothes W, Burton DR, Koff WC, Connors M, Ward AB, Kwong PD, Mascola JR (2016b) Fusion peptide of HIV-1 as a site of vulnerability to neutralizing antibody. Science 352(6287):828–833. https://doi.org/10.1126/science.aae0474
Kovacs JA, Vasudevachari MB, Easter M, Davey RT, Falloon J, Polis MA, Metcalf JA, Salzman N, Baseler M, Smith GE et al (1993) Induction of humoral and cell-mediated anti-human immunodeficiency virus (HIV) responses in HIV sero-negative volunteers by immunization with recombinant gp160. J Clin Invest 92(2):919–928. https://doi.org/10.1172/JCI116667
Kwong PD, Mascola JR (2012) Human antibodies that neutralize HIV-1: identification, structures, and B cell ontogenies. Immunity 37(3):412–425. https://doi.org/10.1016/j.immuni.2012.08.012
Kwong PD, Wyatt R, Robinson J, Sweet RW, Sodroski J, Hendrickson WA (1998) Structure of an HIV gp120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody. Nature 393(6686):648–659. https://doi.org/10.1038/31405
Kwong PD, Mascola JR, Nabel GJ (2013) Broadly neutralizing antibodies and the search for an HIV-1 vaccine: the end of the beginning. Nat Rev Immunol 13(9):693–701. https://doi.org/10.1038/nri3516
Li Y, Migueles SA, Welcher B, Svehla K, Phogat A, Louder MK, Wu X, Shaw GM, Connors M, Wyatt RT, Mascola JR (2007) Broad HIV-1 neutralization mediated by CD4-binding site antibodies. Nat Med 13(9):1032–1034. https://doi.org/10.1038/nm1624
Li Y, O’Dell S, Walker LM, Wu X, Guenaga J, Feng Y, Schmidt SD, McKee K, Louder MK, Ledgerwood JE, Graham BS, Haynes BF, Burton DR, Wyatt RT, Mascola JR (2011) Mechanism of neutralization by the broadly neutralizing HIV-1 monoclonal antibody VRC01. J Virol 85(17):8954–8967. https://doi.org/10.1128/JVI.00754-11
Liao HX, Lynch R, Zhou T, Gao F, Alam SM, Boyd SD, Fire AZ, Roskin KM, Schramm CA, Zhang Z, Zhu J, Shapiro L, Mullikin JC, Gnanakaran S, Hraber P, Wiehe K, Kelsoe G, Yang G, Xia SM, Montefiori DC, Parks R, Lloyd KE, Scearce RM, Soderberg KA, Cohen M, Kamanga G, Louder MK, Tran LM, Chen Y, Cai F, Chen S, Moquin S, Du X, Joyce MG, Srivatsan S, Zhang B, Zheng A, Shaw GM, Hahn BH, Kepler TB, Korber BT, Kwong PD, Mascola JR, Haynes BF (2013) Co-evolution of a broadly neutralizing HIV-1 antibody and founder virus. Nature 496(7446):469–476. https://doi.org/10.1038/nature12053
MacLeod DT, Choi NM, Briney B, Garces F, Ver LS, Landais E, Murrell B, Wrin T, Kilembe W, Liang CH, Ramos A, Bian CB, Wickramasinghe L, Kong L, Eren K, Wu CY, Wong CH, Kosakovsky Pond SL, Wilson IA, Burton DR, Poignard P (2016) Early antibody lineage diversification and independent limb maturation lead to broad HIV-1 neutralization targeting the Env high-mannose patch. Immunity 44(5):1215–1226. https://doi.org/10.1016/j.immuni.2016.04.016
Mascola JR, Haynes BF (2013) HIV-1 neutralizing antibodies: understanding nature’s pathways. Immunol Rev 254(1):225–244. https://doi.org/10.1111/imr.12075
Mascola JR, Louwagie J, McCutchan FE, Fischer CL, Hegerich PA, Wagner KF, Fowler AK, McNeil JG, Burke DS (1994) Two antigenically distinct subtypes of human immunodeficiency virus type 1: viral genotype predicts neutralization serotype. J Infect Dis 169(1):48–54
Mascola JR, Lewis MG, Stiegler G, Harris D, VanCott TC, Hayes D, Louder MK, Brown CR, Sapan CV, Frankel SS, Lu Y, Robb ML, Katinger H, Birx DL (1999) Protection of macaques against pathogenic simian/human immunodeficiency virus 89.6PD by passive transfer of neutralizing antibodies. J Virol 73(5):4009–4018
Mascola JR, Stiegler G, VanCott TC, Katinger H, Carpenter CB, Hanson CE, Beary H, Hayes D, Frankel SS, Birx DL, Lewis MG (2000) Protection of macaques against vaginal transmission of a pathogenic HIV-1/SIV chimeric virus by passive infusion of neutralizing antibodies. Nat Med 6(2):207–210. https://doi.org/10.1038/72318
Moore JP, Sodroski J (1996) Antibody cross-competition analysis of the human immunodeficiency virus type 1 gp120 exterior envelope glycoprotein. J Virol 70(3):1863–1872
Moore JP, Cao Y, Qing L, Sattentau QJ, Pyati J, Koduri R, Robinson J, Barbas CF 3rd, Burton DR, Ho DD (1995) Primary isolates of human immunodeficiency virus type 1 are relatively resistant to neutralization by monoclonal antibodies to gp120, and their neutralization is not predicted by studies with monomeric gp120. J Virol 69(1):101–109
Moore PL, Gray ES, Morris L (2009) Specificity of the autologous neutralizing antibody response. Curr Opin HIV AIDS 4(5):358–363. https://doi.org/10.1097/COH.0b013e32832ea7e8
Muster T, Steindl F, Purtscher M, Trkola A, Klima A, Himmler G, Ruker F, Katinger H (1993) A conserved neutralizing epitope on gp41 of human immunodeficiency virus type 1. J Virol 67(11):6642–6647
Navis M, Tran K, Bale S, Phad GE, Guenaga J, Wilson R, Soldemo M, McKee K, Sundling C, Mascola J, Li Y, Wyatt RT, Karlsson Hedestam GB (2014) HIV-1 receptor binding site-directed antibodies using a VH1-2 gene segment orthologue are activated by Env trimer immunization. PLoS Pathog 10(8):e1004337. https://doi.org/10.1371/journal.ppat.1004337
Parren PW, Wang M, Trkola A, Binley JM, Purtscher M, Katinger H, Moore JP, Burton DR (1998) Antibody neutralization-resistant primary isolates of human immunodeficiency virus type 1. J Virol 72(12):10270–10274
Scheid JF, Mouquet H, Feldhahn N, Seaman MS, Velinzon K, Pietzsch J, Ott RG, Anthony RM, Zebroski H, Hurley A, Phogat A, Chakrabarti B, Li Y, Connors M, Pereyra F, Walker BD, Wardemann H, Ho D, Wyatt RT, Mascola JR, Ravetch JV, Nussenzweig MC (2009) Broad diversity of neutralizing antibodies isolated from memory B cells in HIV-infected individuals. Nature 458(7238):636–640. https://doi.org/10.1038/nature07930
Scheid JF, Mouquet H, Ueberheide B, Diskin R, Klein F, Oliveira TY, Pietzsch J, Fenyo D, Abadir A, Velinzon K, Hurley A, Myung S, Boulad F, Poignard P, Burton DR, Pereyra F, Ho DD, Walker BD, Seaman MS, Bjorkman PJ, Chait BT, Nussenzweig MC (2011) Sequence and structural convergence of broad and potent HIV antibodies that mimic CD4 binding. Science 333(6049):1633–1637. https://doi.org/10.1126/science.1207227
Schwartz DH, Gorse G, Clements ML, Belshe R, Izu A, Duliege AM, Berman P, Twaddell T, Stablein D, Sposto R et al (1993) Induction of HIV-1-neutralising and syncytium-inhibiting antibodies in uninfected recipients of HIV-1IIIB rgp120 subunit vaccine. Lancet 342(8863):69–73
Seaman MS, Janes H, Hawkins N, Grandpre LE, Devoy C, Giri A, Coffey RT, Harris L, Wood B, Daniels MG, Bhattacharya T, Lapedes A, Polonis VR, McCutchan FE, Gilbert PB, Self SG, Korber BT, Montefiori DC, Mascola JR (2010) Tiered categorization of a diverse panel of HIV-1 Env pseudoviruses for assessment of neutralizing antibodies. J Virol 84(3):1439–1452. https://doi.org/10.1128/JVI.02108-09
Shingai M, Donau OK, Schmidt SD, Gautam R, Plishka RJ, Buckler-White A, Sadjadpour R, Lee WR, LaBranche CC, Montefiori DC, Mascola JR, Nishimura Y, Martin MA (2012) Most rhesus macaques infected with the CCR5-tropic SHIV(AD8) generate cross-reactive antibodies that neutralize multiple HIV-1 strains. Proc Natl Acad Sci U S A 109(48):19769–19774. https://doi.org/10.1073/pnas.1217443109
Simek MD, Rida W, Priddy FH, Pung P, Carrow E, Laufer DS, Lehrman JK, Boaz M, Tarragona-Fiol T, Miiro G, Birungi J, Pozniak A, McPhee DA, Manigart O, Karita E, Inwoley A, Jaoko W, Dehovitz J, Bekker LG, Pitisuttithum P, Paris R, Walker LM, Poignard P, Wrin T, Fast PE, Burton DR, Koff WC (2009) Human immunodeficiency virus type 1 elite neutralizers: individuals with broad and potent neutralizing activity identified by using a high-throughput neutralization assay together with an analytical selection algorithm. J Virol 83(14):7337–7348. https://doi.org/10.1128/JVI.00110-09
Sundling C, Li Y, Huynh N, Poulsen C, Wilson R, O’Dell S, Feng Y, Mascola JR, Wyatt RT, Karlsson Hedestam GB (2012) High-resolution definition of vaccine-elicited B cell responses against the HIV primary receptor binding site. Sci Transl Med 4(142):142ra196. https://doi.org/10.1126/scitranslmed.3003752
Sundling C, Zhang Z, Phad GE, Sheng Z, Wang Y, Mascola JR, Li Y, Wyatt RT, Shapiro L, Karlsson Hedestam GB (2014) Single-cell and deep sequencing of IgG-switched macaque B cells reveal a diverse Ig repertoire following immunization. J Immunol 192(8):3637–3644. https://doi.org/10.4049/jimmunol.1303334
Tiller T, Meffre E, Yurasov S, Tsuiji M, Nussenzweig MC, Wardemann H (2008) Efficient generation of monoclonal antibodies from single human B cells by single cell RT-PCR and expression vector cloning. J Immunol Methods 329(1–2):112–124. https://doi.org/10.1016/j.jim.2007.09.017
Trkola A, Pomales AB, Yuan H, Korber B, Maddon PJ, Allaway GP, Katinger H, Barbas CF, 3rd, Burton DR, Ho DD, et al. (1995) Cross-clade neutralization of primary isolates of human immunodeficiency virus type 1 by human monoclonal antibodies and tetrameric CD4-IgG. J Virol 69 (11):6609–6617
Trkola A, Kuster H, Rusert P, Joos B, Fischer M, Leemann C, Manrique A, Huber M, Rehr M, Oxenius A, Weber R, Stiegler G, Vcelar B, Katinger H, Aceto L, Gunthard HF (2005) Delay of HIV-1 rebound after cessation of antiretroviral therapy through passive transfer of human neutralizing antibodies. Nat Med 11(6):615–622. https://doi.org/10.1038/nm1244
van Gils MJ, van den Kerkhof TL, Ozorowski G, Cottrell CA, Sok D, Pauthner M, Pallesen J, de Val N, Yasmeen A, de Taeye SW, Schorcht A, Gumbs S, Johanna I, Saye-Francisco K, Liang CH, Landais E, Nie X, Pritchard LK, Crispin M, Kelsoe G, Wilson IA, Schuitemaker H, Klasse PJ, Moore JP, Burton DR, Ward AB, Sanders RW (2016) An HIV-1 antibody from an elite neutralizer implicates the fusion peptide as a site of vulnerability. Nat Microbiol 2:16199. https://doi.org/10.1038/nmicrobiol.2016.199
Walker LM, Phogat SK, Chan-Hui PY, Wagner D, Phung P, Goss JL, Wrin T, Simek MD, Fling S, Mitcham JL, Lehrman JK, Priddy FH, Olsen OA, Frey SM, Hammond PW, Kaminsky S, Zamb T, Moyle M, Koff WC, Poignard P, Burton DR (2009) Broad and potent neutralizing antibodies from an African donor reveal a new HIV-1 vaccine target. Science 326(5950):285–289. https://doi.org/10.1126/science.1178746
Walker LM, Simek MD, Priddy F, Gach JS, Wagner D, Zwick MB, Phogat SK, Poignard P, Burton DR (2010) A limited number of antibody specificities mediate broad and potent serum neutralization in selected HIV-1 infected individuals. PLoS Pathog 6(8):e1001028. https://doi.org/10.1371/journal.ppat.1001028
Walker LM, Huber M, Doores KJ, Falkowska E, Pejchal R, Julien JP, Wang SK, Ramos A, Chan-Hui PY, Moyle M, Mitcham JL, Hammond PW, Olsen OA, Phung P, Fling S, Wong CH, Phogat S, Wrin T, Simek MD, Koff WC, Wilson IA, Burton DR, Poignard P (2011a) Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 477(7365):466–470. https://doi.org/10.1038/nature10373
Walker LM, Sok D, Nishimura Y, Donau O, Sadjadpour R, Gautam R, Shingai M, Pejchal R, Ramos A, Simek MD, Geng Y, Wilson IA, Poignard P, Martin MA, Burton DR (2011b) Rapid development of glycan-specific, broad, and potent anti-HIV-1 gp120 neutralizing antibodies in an R5 SIV/HIV chimeric virus infected macaque. Proc Natl Acad Sci U S A 108(50):20125–20129. https://doi.org/10.1073/pnas.1117531108
Wei X, Decker JM, Wang S, Hui H, Kappes JC, Wu X, Salazar-Gonzalez JF, Salazar MG, Kilby JM, Saag MS, Komarova NL, Nowak MA, Hahn BH, Kwong PD, Shaw GM (2003) Antibody neutralization and escape by HIV-1. Nature 422(6929):307–312. https://doi.org/10.1038/nature01470
West AP Jr, Diskin R, Nussenzweig MC, Bjorkman PJ (2012) Structural basis for germ-line gene usage of a potent class of antibodies targeting the CD4-binding site of HIV-1 gp120. Proc Natl Acad Sci U S A 109(30):E2083–E2090. https://doi.org/10.1073/pnas.1208984109
Wibmer CK, Gorman J, Ozorowski G, Bhiman JN, Sheward DJ, Elliott DH, Rouelle J, Smira A, Joyce MG, Ndabambi N, Druz A, Asokan M, Burton DR, Connors M, Abdool Karim SS, Mascola JR, Robinson JE, Ward AB, Williamson C, Kwong PD, Morris L, Moore PL (2017) Structure and recognition of a novel HIV-1 gp120-gp41 Interface antibody that caused MPER exposure through viral escape. PLoS Pathog 13(1):e1006074. https://doi.org/10.1371/journal.ppat.1006074
Wrammert J, Smith K, Miller J, Langley WA, Kokko K, Larsen C, Zheng NY, Mays I, Garman L, Helms C, James J, Air GM, Capra JD, Ahmed R, Wilson PC (2008) Rapid cloning of high-affinity human monoclonal antibodies against influenza virus. Nature 453(7195):667–671. https://doi.org/10.1038/nature06890
Wu X, Kong XP (2016) Antigenic landscape of the HIV-1 envelope and new immunological concepts defined by HIV-1 broadly neutralizing antibodies. Curr Opin Immunol 42:56–64. https://doi.org/10.1016/j.coi.2016.05.013
Wu X, Zhou T, O’Dell S, Wyatt RT, Kwong PD, Mascola JR (2009) Mechanism of human immunodeficiency virus type 1 resistance to monoclonal antibody B12 that effectively targets the site of CD4 attachment. J Virol 83(21):10892–10907. https://doi.org/10.1128/JVI.01142-09
Wu X, Yang ZY, Li Y, Hogerkorp CM, Schief WR, Seaman MS, Zhou T, Schmidt SD, Wu L, Xu L, Longo NS, McKee K, O’Dell S, Louder MK, Wycuff DL, Feng Y, Nason M, Doria-Rose N, Connors M, Kwong PD, Roederer M, Wyatt RT, Nabel GJ, Mascola JR (2010) Rational design of envelope identifies broadly neutralizing human monoclonal antibodies to HIV-1. Science 329(5993):856–861. https://doi.org/10.1126/science.1187659
Wu X, Zhou T, Zhu J, Zhang B, Georgiev I, Wang C, Chen X, Longo NS, Louder M, McKee K, O’Dell S, Perfetto S, Schmidt SD, Shi W, Wu L, Yang Y, Yang ZY, Yang Z, Zhang Z, Bonsignori M, Crump JA, Kapiga SH, Sam NE, Haynes BF, Simek M, Burton DR, Koff WC, Doria-Rose NA, Connors M, Mullikin JC, Nabel GJ, Roederer M, Shapiro L, Kwong PD, Mascola JR (2011) Focused evolution of HIV-1 neutralizing antibodies revealed by structures and deep sequencing. Science 333(6049):1593–1602. https://doi.org/10.1126/science.1207532
Wu X, Zhang Z, Schramm CA, Joyce MG, Kwon YD, Zhou T, Sheng Z, Zhang B, O’Dell S, McKee K, Georgiev IS, Chuang GY, Longo NS, Lynch RM, Saunders KO, Soto C, Srivatsan S, Yang Y, Bailer RT, Louder MK, Mullikin JC, Connors M, Kwong PD, Mascola JR, Shapiro L (2015) Maturation and diversity of the VRC01-antibody lineage over 15 years of chronic HIV-1 infection. Cell 161(3):470–485. https://doi.org/10.1016/j.cell.2015.03.004
Xiao M, Prabakaran P, Chen W, Kessing B, Dimitrov DS (2013) Deep sequencing and Circos analyses of antibody libraries reveal antigen-driven selection of Ig VH genes during HIV-1 infection. Exp Mol Pathol 95(3):357–363. https://doi.org/10.1016/j.yexmp.2013.10.004
Yin L, Hou W, Liu L, Cai Y, Wallet MA, Gardner BP, Chang K, Lowe AC, Rodriguez CA, Sriaroon P, Farmerie WG, Sleasman JW, Goodenow MM (2013) IgM repertoire biodiversity is reduced in HIV-1 infection and systemic lupus erythematosus. Front Immunol 4:373. https://doi.org/10.3389/fimmu.2013.00373
Zhang Y, Yuan T, Li J, Xu J, Shao Y, Chen Z, Zhang MY (2013) The potential of the human immune system to develop broadly neutralizing HIV-1 antibodies: implications for vaccine development. AIDS 27(16):2529–2539. https://doi.org/10.1097/QAD.0000000000000015
Zhou T, Xu L, Dey B, Hessell AJ, Van Ryk D, Xiang SH, Yang X, Zhang MY, Zwick MB, Arthos J, Burton DR, Dimitrov DS, Sodroski J, Wyatt R, Nabel GJ, Kwong PD (2007) Structural definition of a conserved neutralization epitope on HIV-1 gp120. Nature 445(7129):732–737. https://doi.org/10.1038/nature05580
Zhou T, Georgiev I, Wu X, Yang ZY, Dai K, Finzi A, Kwon YD, Scheid JF, Shi W, Xu L, Yang Y, Zhu J, Nussenzweig MC, Sodroski J, Shapiro L, Nabel GJ, Mascola JR, Kwong PD (2010) Structural basis for broad and potent neutralization of HIV-1 by antibody VRC01. Science 329(5993):811–817. https://doi.org/10.1126/science.1192819
Zhou T, Zhu J, Wu X, Moquin S, Zhang B, Acharya P, Georgiev IS, Altae-Tran HR, Chuang GY, Joyce MG, Kwon YD, Longo NS, Louder MK, Luongo T, McKee K, Schramm CA, Skinner J, Yang Y, Yang Z, Zhang Z, Zheng A, Bonsignori M, Haynes BF, Scheid JF, Nussenzweig MC, Simek M, Burton DR, Koff WC, Mullikin JC, Connors M, Shapiro L, Nabel GJ, Mascola JR, Kwong PD (2013) Multidonor analysis reveals structural elements, genetic determinants, and maturation pathway for HIV-1 neutralization by VRC01-class antibodies. Immunity 39(2):245–258. https://doi.org/10.1016/j.immuni.2013.04.012
Zhou T, Lynch RM, Chen L, Acharya P, Wu X, Doria-Rose NA, Joyce MG, Lingwood D, Soto C, Bailer RT, Ernandes MJ, Kong R, Longo NS, Louder MK, McKee K, O’Dell S, Schmidt SD, Tran L, Yang Z, Druz A, Luongo TS, Moquin S, Srivatsan S, Yang Y, Zhang B, Zheng A, Pancera M, Kirys T, Georgiev IS, Gindin T, Peng HP, Yang AS, Mullikin JC, Gray MD, Stamatatos L, Burton DR, Koff WC, Cohen MS, Haynes BF, Casazza JP, Connors M, Corti D, Lanzavecchia A, Sattentau QJ, Weiss RA, West AP, Jr., Bjorkman PJ, Scheid JF, Nussenzweig MC, Shapiro L, Mascola JR, Kwong PD (2015) Structural repertoire of HIV-1-neutralizing antibodies targeting the CD4 supersite in 14 donors. Cell 161 (6):1280–1292. doi:https://doi.org/10.1016/j.cell.2015.05.007
Acknowledgments
Xueling Wu and her laboratory are funded by internal funds from the Aaron Diamond AIDS Research Center and by research grants R01AI114380 and R01AI122593 from the National Institute of Allergy and Infectious Diseases (NIAID), National Institutes of Health (NIH), USA.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Wu, X. (2018). HIV Broadly Neutralizing Antibodies: VRC01 and Beyond. In: Zhang, L., Lewin, S. (eds) HIV Vaccines and Cure . Advances in Experimental Medicine and Biology, vol 1075. Springer, Singapore. https://doi.org/10.1007/978-981-13-0484-2_3
Download citation
DOI: https://doi.org/10.1007/978-981-13-0484-2_3
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-0483-5
Online ISBN: 978-981-13-0484-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)