Abstract
Our understanding of how T cells respond to dengue virus has greatly advanced in the last decade but important questions still remain unanswered. Dengue virus infection elicits a broad anti-viral T cell response with NS3, NS4b and NS5 being the main targets for CD8+ T cells, which dominate the response while the structural proteins capsid, envelope and the secreted protein NS1 are the preferential targets for CD4+ T cells. Upon T cell activation during acute dengue infection, dengue-specific T cells acquire expression of the skin-homing marker cutaneous associated antigen (CLA) and they can be found at high frequencies in the skin of infected patients. This suggests that the skin represents an important site for the immuno surveillance of dengue virus. The immunoprotective role of skin-homing dengue-specific T cells, their potential involvement in pathological skin manifestations and their long-term persistence as tissue resident T cells to provide immediate onsite protection are open questions that we are currently investigating. The contribution of pre-existing dengue-specific T cells towards protective immunity and/or immunopathology during secondary dengue infection remains a major knowledge gap. The evidence supporting these opposing outcomes and our current understanding of the characteristics of the human T cell response to dengue virus will be discussed.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
17.1 Role of T Cell Immunity and “Original Antigenic Sin”
CD4+ and CD8+ T cells are an essential component of protective immunity against viral infections and understanding their role and how they develop is critical for the design of optimal vaccines.
Acute viral infections resulting in viral clearance generally elicit effective anti-viral T cell responses that progress through three phases: a period of activation and expansion of virus-specific T cells, a contraction phase where the majority of effector cells undergo death followed by the establishment and maintenance of a pool of virus-specific memory T cells [35]. These memory T cells, together with other components of the immune system, will confer protection upon re-infection with the same virus. In particular, CD8+ T cells are important for viral clearance as they can directly lyse virus-infected cells through the production of IFN-γ and cytotoxic molecules such as perforin and granzymes. These processes involve the recognition through the T cell receptor (TCR) of viral peptides of 9–10 aminoacids in length, that are presented on the cell surface of antigen presenting cells and virus-infected cells in association with molecules of the human leukocyte antigen (HLA) system. CD4+ T cells recognize longer peptides of 12–15 aminoacids in length and have a more diverse function as they are required to develop a broad and efficient antibody response and they are critical for the generation of both B cell and CD8+ T cell memory responses. In addition, CD4+ T cells can also directly kill virus-infected cells through production of IFN-γ and cytotoxic effector functions in a similar manner to CD8+ T cells.
Successful vaccines such as the vaccinia virus vaccine (which led to the eradication of small pox) and the yellow fever vaccine are known to elicit strong and long-lasting antibody and T cell responses [17, 23]. We believe that the induction of both components of the immune system is key to a successful vaccine. However, designing a vaccine for dengue virus has proved to be particularly challenging because of the co-circulation of four distinct serotypes that share approximately 70% of sequence homology. Thus, individuals living in a dengue-endemic region will be exposed to multiple dengue serotypes during their lifetime. Infection with one dengue serotype will confer life-long protective immunity towards the same serotype but evidence shows that that it increases the risk of DHF/DSS upon secondary infection with another serotype [11]. An explanation for these observations was provided by a mechanism called “antibody dependant enhancement” (ADE) whereby anti-DENV antibodies that are generated during a primary dengue infection will bind to – but poorly neutralize – the secondary infecting serotype due to variations in the peptide sequence between the two viruses. Suboptimal neutralization results in increased uptake of virus-antibody complexes through the FcγR present on antigen presenting cells with subsequent increased viral load and antigen-presentation and enhanced immune activation. In 2003 the group of Gavin Screaton proposed a similar mechanism for T cells whereby during secondary dengue infections the T cell response is dominated by weakly cross-reactive memory T cells specific for the primary infecting virus that have suboptimal anti-viral capacity towards the secondary infecting serotype but secrete pro-inflammatory cytokines such as TNF-α, which contribute to plasma leakage [18]. This process known as “original antigenic sin” was first described in a mice to impair CD8+ T cell recognition and clearance of a secondary-infecting Lymphocytic Choriomeningitis Virus (LCMV) bearing mutations in T cell epitopes [13]. However, subsequent mouse models showed that the presence of cross-reactive T cells could in some cases be beneficial. For example, pre-existing memory cells that are cross-reactive with vaccinia virus can protect mice from an otherwise lethal dose of vaccinia virus [4]. Different mouse models of secondary virus infection have shown that pre-existing cross-reactive T cell immunity can alter the hierarchy of the T cell response to a secondary-infecting virus leading to enhanced or diminished protective immunity and altered immunopathology. There is also evidence of a strong skewing of the T cell repertoire in individuals with secondary DENV infections compared to those with primary infection or in vaccinees receiving a tetravalent live-attenuated dengue vaccine as opposed to the monovalent vaccine [32, 33]. However, the implications of the skewed T cell repertoire for T cell functionality remain unclear (Fig. 17.1).
Schematic depicting the process of “original antigenic sin” for T cells during dengue infection and its possible implications for protective immunity and/or immunopathology. (a) During a primary infection for example with DENV 3, naïve T cells with high avidity for DENV 3-derived peptides undergo expansion. The resulting dengue-specific T cell pool will comprise of T cells specific for peptides that are unique to DENV 3 (serotype-specific) and those that are shared amongst 2 or more serotypes (conserved). T cells that have high avidity for their cognate antigen undergo optimal TCR triggering which mainly leads to production of IFN-γ and efficient anti-viral function. (b) Upon secondary infection for example with DENV 2, pre-existing T cells generated during the primary DENV 3 infection that are cross-reactive to DENV 2 peptides will preferentially expand compared to their naïve counterpart. This occurs because dengue-specific memory cells are present in higher frequencies and have a lower activation threshold compared to their naïve counterparts. As a result, the responding T cell pool will be dominated by cross-reactive T cells specific for conserved epitopes. Some cross-reactive T cells may have a lower avidity for DENV 2 antigens which results in suboptimal TCR triggering and subsequent production of high levels of TNF-α and poor anti-viral efficacy with a possible contribution to immunopathology. However, variable proportions of cross-reactive memory T cells may have high avidity for DENV 2 antigens, will undergo optimal TCR triggering resulting in high levels of IFN-γ and anti-viral capacity. The outcome of the activation of cross-reactive T cells is therefore difficult to predict and could lead to faster and more efficient protective responses or to different degrees of immunopathology. Both scenarios however should result in skewing of the T cell response towards more conserved epitopes
At a molecular level, because the TCR interacts with only few aminoacids within the 9–10 mer peptide, it can accommodate aminoacid variations providing they do not disrupt the peptide-TCR or the peptide-HLA interactions. However, the result of this variation in terms of avidity of the T cell for the peptide and level of T cell activation is highly variable and difficult to predict. We believe that the complexity of T cell cross-reactivity is at the basis of the contrasting findings reported over the last decade that point to either a protective or a pathologic role of dengue-specific T cells during secondary dengue virus infections [22]. Several studies support the hypotheses proposed in the 2003 study [18], for example dengue-specific T cells are found to be present at higher frequencies in patients with DHF as compared to those with DF [6] and in vitro stimulation of T cells with homologous versus heterologous dengue peptides results in cytokine profiles that are both qualitatively and quantitatively distinct [9, 16, 19]. However, a number of other studies do not confirm these findings [8, 10, 14, 29, 30] and there is now accumulating evidence supporting a protective role of T cell immunity during DENV infection, including data from immune-deficient mouse models [21, 36, 37]. A prospective study in school children shows that asymptomatic secondary dengue infections correlate with increased levels of DENV-specific T cells compared to symptomatic infection22. In healthy adults from dengue endemic regions HLA class I and II molecules associated with decreased susceptibility to severe dengue disease were shown to support memory T cell responses of higher magnitude, suggesting a protective rather than a detrimental role of T cells [31, 34]. Further studies that directly compare the primary versus secondary T cell response to individual DENV serotypes are urgently needed to clarify the role of these cells during infection and to reconcile the contrasting findings reported in the literature. The complex scenario that emerges from the above studies further highlights that it is mandatory for a dengue vaccine to elicit a balanced T cell (as well as antibody) response towards all four DENV serotypes.
17.2 T Cell Responses during Dengue Virus Infection
To better understand the role of T cells during dengue infection we sought to characterize the nature of the T cell response during the course of dengue infection, from acute infection to convalescence. Dengue patients were recruited by the team of Prof Leo Yee Sin at CDC Tan Tock Seng Hospital in Singapore and blood was drawn from the patients at three different time points after diagnosis of dengue infection (days from fever onset: acute day 6–9, post-febrile day 14–21 and convalescent day 60–120). Peripheral blood mononuclear cells (PBMCs) were analyzed at all time points. The first striking observation that we made was that acute dengue infection induces a massive activation and proliferation of CD8+ T cells and a significant but more modest activation and expansion of CD4+ T cells. Activated (HLA-DR+ CD38+) and proliferating (Ki67+ Bcl2-) T cells are found at highest frequencies during acute dengue and represent in some patients up to 70% of all CD8+ T cells (highlighted with a red box in Fig. 17.2a). The proportion of activated and proliferating T cells decreases in the post-febrile and convalescent phases of disease reaching levels comparable to those found in healthy individuals. Representative flow cytometry profiles of the activation and proliferation of CD4+ and CD8+ T cells from a dengue patient are shown in Fig. 17.2a and results from 18 patients are summarized in Fig. 17.2b. Given the large numbers of activated CD8+ T cells observed in the blood of acute dengue patients, it is likely that a proportion of these cells will be bystander-activated and will thus not be specific for DENV. We and others have shown that acute dengue infection is accompanied by the activation and proliferation of CD8+ T cells that are specific for persistent viruses such as human cytomegalovirus or Epstein-Barr virus [25, 26]. These cells may be activated in vivo in a TCR-independent manner through the action of cytokines such as IL-15 that are abundantly produced upon viral infection [26]. However the exact enumeration of dengue-specific T cells is challenging as we rely on functional assays such as IFN-γ production after DENV peptide stimulation which are unable to detect T cells with impaired cytokine-producing capacities and are thus likely to underestimate the number of dengue-specific T cells.
High frequencies of activated and proliferating T cells during acute dengue infection. PBMC samples from dengue patients are analyzed during the acute (A), post-febrile and (PF) convalescent (C) phases of disease. Live cells are surface stained with antibodies against CD3, CD4, CD8, CD38 and HLA-DR followed by intracellular staining for Ki67 and Bcl2. Isotype controls are used to define negativity for CD38, HLA-DR, Ki67 and Bcl2. (a) Activation and proliferation profiles of CD8+ (top panel) and CD4+ T cells (bottom panel) for one representative patient. Activated and proliferating T cells are defined as CD38+ HLA-DR+ and Ki67+ Bcl2- cells, respectively (see red box for CD8+ T cells). (b) Results from 18 patients are summarized for CD8+ (top panel) and CD4+ T cells (bottom panel). Statistics are calculated by Mann-Whitney non-parametric test. Values are considered significant when p < 0.05
Our knowledge of the kinetics and the protein targets of the dengue-specific T cell response during acute dengue infection has greatly advanced over the last decade. We know that dengue infection elicits a DENV-specific T cell response that peaks around days 8–10 from fever onset [6, 24, 29]. The T cell response is broad as it targets all viral proteins to some extent, with a preferential recognition of the non-structural proteins NS3, NS4b and NS5. Similarly to what occurs in other acute viral infections dengue-specific CD8+ T cells are present at higher frequencies compared to CD4+ T cells. While CD8+ T cell responses preferentially target the non-structural proteins NS3 and NS5, CD4+ T cells mainly target the structural proteins capsid and envelope and the secreted proteins NS1, consistent with virions being the main source of antigen for CD4+ T cells [24]. More recent studies have reported that these patterns of immunodominance vary as a function of the infecting serotype with DENV 3 eliciting CD8+ T cells that target both structural and non-structural proteins while DENV 1, 2 and 4 elicit CD8+ T cells targeting mainly NS3, NS4b and NS5 [32, 33].
17.3 Phenotype and Tissue-Homing of Dengue-Specific T Cells
The identification of a large number of T cell epitopes, the majority of which are CD8+ T cell epitopes, has provided us with tools to better characterize T cells during dengue infection or vaccination. As of September 2017 1613 immunogenic peptides have been deposited in the Immune Epitope Database (http://www.iedb.org/). In particular, the study of CD8+ T cells has been facilitated by the use of peptide-HLA tetramers which are streptavidin-labelled MHC multimers bound to the peptide of interest. These tools have revolutionized the way we can look at T cell immunity as peptide-specific CD8+ T cells can be enumerated and characterized phenotypically directly ex vivo by flow cytometry without relying on a functional read-out [1]. However, the definition of immunodominant epitopes restricted to HLA molecules that are commonly expressed in populations from dengue-endemic regions remains elusive and the selection of T cell epitopes to use for peptide-HLA tetramer analyses of dengue-infected individuals can still be a difficult task.
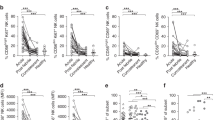
By using peptide-HLA tetramers for the previously described HLA-A*1101-restricted dengue NS3 27 epitope [18] we addressed the functional and phenotypic characteristics of dengue-specific T cells during the course of dengue infection. We show that during secondary acute dengue infection NS3–27-specific T cells are highly activated and proliferating, produce IFN-γ but not TNF-γ and are able to lyse dengue-infected cells, suggesting that at least in vitro their anti-viral function does not appear to be impaired [25]. To address the capacity of dengue-specific T cells to migrate to different organs and to potentially contribute to immunopathology, we studied the homing receptors present on the surface of these cells. During acute dengue infection NS3 27-specific CD8+ T cells express the skin homing molecule cutaneous lymphocyte-associated antigen (CLA), the chemokine receptors CXCR3 and CCR5 which allow migration to inflamed tissues, but not the gut-homing receptor CCR9. Expression of CLA but not of CCR9 was confirmed for the total population of CD4+ and CD8+ T cells specific for NS3 and NS5 (Fig. 17.3a, b). We show that expression of the skin-homing marker CLA by dengue-specific T cells correlates with their presence in the skin tissue where they could be detected at higher frequencies as compared to peripheral blood [25] (Fig. 17.3c). The exact role of dengue-specific T cells in the skin during acute infection, whether they persist long-term in this site as tissue-resident memory T cells upon resolution of the infection and their possible protective role upon secondary infections is currently under investigation. The expression of CLA on dengue-specific T cells during acute dengue is consistent with their initial activation by skin-derived dendritric cells. Dengue virus enters the body through the skin during the blood meal of an infected mosquito, infects local dendritic cells [3, 28] and travels to skin-draining lymph-nodes where it interacts with virus-specific T cells. Studies have shown that during activation T cells acquire the capacity to migrate to certain tissues by expressing different sets of receptors that serve as “address-codes”, the expression of which is determined by the antigen presenting cell and by the local microenvironment [7]. It was shown for example that during T cell priming, tissue-derived antigen presenting cells within cutaneous versus intestinal secondary lymphoid organs imprint the corresponding tissue-specific homing phenotype to the T cell, such that the primed T cells have a predisposition to home back to the skin or gut, respectively [2, 20]. More recently, studies have shown that the route of immunization strongly impacts the migratory capacity and consequently the protective efficacy of T cells upon re-infection, as T cells need to localize to the sites of virus replication in order to exert their function [15, 27]. For example, vaccination with live vaccinia virus was 105 times more effective in protecting against viral re-infection if delivered by skin scarification as compared to subcutaneous, intradermal, and intramuscular vaccination [12]. The authors show that immunization by skin scarification was associated with local keratinocyte infection and generation of a long-lived tissue resident CD8+ T cell memory populations. The presence of tissue resident CD8+ T cells was sufficient to protect against re-infection with vaccinia virus and did not require antibodies or recruitment of circulating blood T cells [15]. These considerations highlight an important aspect of T cell immunity that is often not considered in the context of vaccination. We now know that pathogen infection generates populations of non-recirculating tissue-resident memory T cells that persist long-term within tissues such as the skin, lung, gastro-intestinal tract and reproductive tract. These cells have potent effector functions and provide rapid on-site protection in peripheral tissues upon re-infection with previously-encountered pathogens [5]. A better understanding of the localization of dengue-specific memory T cells in dengue-immune individuals and how these cells can be generated in the context of vaccination would greatly aid the design of dengue vaccines with improved protective efficacy.
Dengue-specific T cells have skin-tissue migratory properties during acute dengue infection. Expression of the skin-homing receptor CLA (a) and the gut-homing receptor CCR9 (b) is assessed by flow cytometry on dengue-specific T cells from the peripheral blood of acute dengue patients. PBMCs are briefly stimulated with or without NS3 and NS5 peptides and live cells are stained for expression of CD3, CD4, CD8, CLA and CCR9 followed by intracellular cytokine staining for IFN-γ and TNF-α. Expression of CLA and CCR9 is analysed on cytokine-producing CD4+ and CD8+ T cells (defined as “DENV-specific”) or on those that lack cytokine production (defined “non-DENV specific”). The percentage of CLA+ or CCR9+ cells is shown for CD4+ (white circles) and CD8+ T cells (black circles). Isotype controls are used to define negativity for CLA and CCR9 expression. (c) The presence of dengue-specific T cells in the skin was investigated by raising skin suction blisters on the forearm of dengue patients. PBMCs or cells obtained from skin suction blisters are stimulated with or without NS3 and NS5 peptides and analysed for production of IFN-γ by ELISPOT. Results obtained from cells derived from the skin (white bar) or the peripheral blood (black bar) are expressed as Spot Forming Cells relative to 105 cells (SFC/105). Statistics were calculated using a Kruskal-Wallis test, followed by a non-parametric Mann-Whitney test. (Adapted from Rivino et al. Sci Trans Med [25])
References
Altman JD, Moss PA, Goulder PJ, Barouch DH, McHeyzer-Williams MG, Bell JI, McMichael AJ, Davis MM (1996) Phenotypic analysis of antigen-specific T lymphocytes. Science 274(5284):94–96
Campbell DJ, Butcher EC. (2002) Rapid acquisition of tissue-specific homing phenotypes by CD4(+) T cells activated in cutaneous or mucosal lymphoid tissues. J Exp Med 195(1):135–141. The Rockefeller University Press. https://doi.org/10.1084/jem.20011502
Cerny D, Haniffa M, Shin A, Bigliardi P, Tan BK, Lee B, Poidinger M, Tan EY, Ginhoux F, Fink K (2014) Selective susceptibility of human skin antigen presenting cells to productive dengue virus infection (Kuhn RJ (ed)). PLoS Pathogens 10(12):e1004548. Public Library of Science. https://doi.org/10.1371/journal.ppat.1004548
Chen HD, Fraire AE, Joris I, Brehm MA, Welsh RM, Selin LK (2001) Memory CD8+ T cells in heterologous antiviral immunity and immunopathology in the lung. Nat Immunol 2(11):1067–1076. https://doi.org/10.1038/ni727
Clark RA (2015) Resident memory T cells in human health and disease. Sci Transl Med 7(269):: 269rv1–269rv1. American Association for the Advancement of Science. https://doi.org/10.1126/scitranslmed.3010641
Duangchinda T, Dejnirattisai W, Vasanawathana S, Limpitikul W, Tangthawornchaikul N, Malasit P, Mongkolsapaya J, Screaton G (2010) Immunodominant T-cell responses to dengue virus NS3 are associated with DHF. Proc Natl Acad Sci 107(39):16922–16927. https://doi.org/10.1073/pnas.1010867107
Dudda JC, Simon JC, Martin S (2004) Dendritic cell immunization route determines CD8+ T cell trafficking to inflamed skin: role for tissue microenvironment and dendritic cells in establishment of T cell-homing subsets. J Immunol 172(2):857–863
Dung NTP, Duyen HTL, Thuy NTV, Ngoc TV, Chau NVV, Hien TT, Rowland-Jones SL et al (2010) Timing of CD8+ T cell responses in relation to commencement of capillary leakage in children with dengue. J Immunol 184(12):7281–7287. https://doi.org/10.4049/jimmunol.0903262
Friberg H, Burns L, Woda M, Kalayanarooj S, Endy TP, Stephens HAF, Green S, Rothman AL, Mathew A (2010, April) Memory CD8+ T cells from naturally acquired primary dengue virus infection are highly cross-reactive. Immunol Cell Biol:1–8. Nature Publishing Group. https://doi.org/10.1038/icb.2010.61
Friberg H, Bashyam H, Toyosaki-Maeda T, Potts JA, Greenough T, Kalayanarooj S, Gibbons RV et al (2011) Cross-reactivity and expansion of dengue-specific T cells during acute primary and secondary infections in humans. Sci Rep 1(August). https://doi.org/10.1038/srep00051
Halstead SB (2014) Dengue antibody-dependent enhancement: knowns and unknowns. Microbiol Spect 2(6). American Society of Microbiology. https://doi.org/10.1128/microbiolspec.AID-0022-2014
Jiang X, Clark RA, Liu L, Wagers AJ, Fuhlbrigge RC, Kupper TS (2012) Skin infection generates non-migratory memory CD8+ TRM cells providing global skin immunity. Nature 483(7388):227–231. https://doi.org/10.1038/nature10851
Klenerman P, Zinkernagel RM (1998) Original antigenic sin impairs cytotoxic T lymphocyte responses to viruses bearing variant epitopes. Nature 394(6692):482–485. https://doi.org/10.1038/28860
Kurane I, Matsutani T, Suzuki R, Takasaki T, Kalayanarooj S, Green S, Rothman AL, Ennis FA (2011) T-Cell Responses to Dengue Virus in Humans. Trop Med Health 39(4 Supplement):S45–S51. https://doi.org/10.2149/tmh.2011-S09
Liu L, Zhong Q, Tian T, Dubin K, Athale SK, Kupper TS (2010) Epidermal injury and infection during poxvirus immunization is crucial for the generation of highly protective T cell – mediated immunity. Nat Med 16(2):224–227. Nature Publishing Group. https://doi.org/10.1038/nm.2078
Mangada MM, Rothman AL (2005) Altered cytokine responses of dengue-specific CD4+ T cells to heterologous serotypes. J Immunol 175(4):2676–2683
Miller JD, van der Most RG, Akondy RS, Glidewell JT, Albott S, Masopust D, Murali-Krishna K et al (2008) Human effector and memory CD8+ T cell responses to smallpox and yellow fever vaccines. Immunity 28(5):710–722. https://doi.org/10.1016/j.immuni.2008.02.020
Mongkolsapaya J, Dejnirattisai W, Xu X-n, Vasanawathana S, Tangthawornchaikul N, Chairunsri A, Sawasdivorn S et al (2003) Original antigenic sin and apoptosis in the pathogenesis of dengue hemorrhagic fever. Nat Med 9(7):921–927. https://doi.org/10.1038/nm887
Mongkolsapaya J, Duangchinda T, Dejnirattisai W, Vasanawathana S, Avirutnan P, Jairungsri A, Khemnu N et al (2006) T cell responses in dengue hemorrhagic fever: are cross-reactive T cells suboptimal? J Immunol 176(6):3821–3829
Mora JR, Bono MR, Manjunath N, Weninger W, Cavanagh LL, Rosemblatt M, von Andrian UH (2003) Selective imprinting of gut-homing T cells by Peyer’s patch dendritic cells. Nat Publ Group 424(6944):88–93. https://doi.org/10.1038/nature01726
Prestwood TR, Morar MM, Zellweger RM, Miller R, May MM, Yauch LE, Lada SM, Shresta S (2012) Gamma interferon (IFN-Γ) receptor restricts systemic dengue virus replication and prevents paralysis in IFN-Α/Β receptor-deficient mice. J Virol 86(23):12561–12570. American Society for Microbiology. https://doi.org/10.1128/JVI.06743-11
Rivino (2016) T cell immunity to dengue virus and implications for vaccine design. Expert Rev Vaccines 15(4):443–453. https://doi.org/10.1586/14760584.2016.1116948
Rivino L, Messi M, Jarossay D, Lanzavecchia A, Sallusto F, Geginat J (2004) Chemokine receptor expression identifies Pre-T helper (Th)1, Pre-Th2, and nonpolarized cells among human CD4+ central memory T cells. J Exp Med 200(6):725–735. https://doi.org/10.1084/jem.20040774
Rivino L, Kumaran EAP, Jovanovic V, Nadua K, Teo EW, Pang SW, Teo GH et al (2013) Differential targeting of viral components by CD4+ versus CD8+ T lymphocytes in dengue virus infection. J Virol 87(5):2693–2706. https://doi.org/10.1128/JVI.02675-12
Rivino L, Kumaran EA, Thein TL, Too CT, Hao Gan VC, Hanson BJ, Wilder-Smith A et al (2015) Virus-specific T lymphocytes home to the skin during natural dengue infection. Sci Transl Med 7(278):278ra35–278ra35. https://doi.org/10.1126/scitranslmed.aaa0526
Sandalova, E, Laccabue D, Boni C, Tan AT, Fink K, Ooi EE, Chua R, Shafaeddin Schreve B, Ferrari C, Bertoletti A (2010) Contribution of herpesvirus specific CD8 T cells to anti-viral T cell response in humans (Selin L (ed)). PLoS Pathogens 6(8): e1001051. https://doi.org/10.1371/journal.ppat.1001051
Sandoval F, Terme M, Nizard M, Badoual C, Bureau MF, Freyburger L, Clement O et al (2013) Mucosal imprinting of vaccine-induced CD8+ T cells is crucial to inhibit the growth of mucosal tumors. Sci Transl Med 5(172):172ra20–172ra20. https://doi.org/10.1126/scitranslmed.3004888
Schmid MA, Harris E (2014) Monocyte recruitment to the dermis and differentiation to dendritic cells increases the targets for dengue virus replication (Richard JK (ed)). PLoS Pathogens 10(12):e1004541. Public Library of Science. https://doi.org/10.1371/journal.ppat.1004541
Simmons CP, Dong T, Chau NV, Dung NTP, Chau TNB, Thao LTT, Dung NT, Hien TT, Rowland-Jones S, Farrar J (2005) Early T-cell responses to dengue virus epitopes in Vietnamese adults with secondary dengue virus infections. J Virol 79(9):5665–5675. https://doi.org/10.1128/JVI.79.9.5665-5675.2005
Townsley E, Woda M, Thomas SJ, Kalayanarooj S, Gibbons RV, Nisalak A, Srikiatkhachorn A et al (2014) Distinct activation phenotype of a highly conserved novel HLA-B57-restricted epitope during dengue virus infection. Immunology 141(1):27–38. https://doi.org/10.1111/imm.12161
Weiskopf, Daniela, Angelo MA, de Azeredo EL, Sidney J, Greenbaum JA, Fernando AN, Broadwater A et al (2013) Comprehensive analysis of dengue virus-specific responses supports an HLA-linked protective role for CD8+ T cells. Proc Nat Acad Sci U S A 110(22):E2046–2053. National Acad Sciences. https://doi.org/10.1073/pnas.1305227110
Weiskopf D, Angelo MA, Sidney J, Peters B, Shresta S, Sette A (2014a) Immunodominance changes as a function of the infecting dengue virus serotype and primary versus secondary infection. J Virol 88(19):11383–11394. https://doi.org/10.1128/JVI.01108-14
Weiskopf D, Angelo MA, Bangs DJ, Sidney J, Paul S, Peters B, de Silva AD et al (2014b) The human CD8 +T cell responses induced by a live attenuated tetravalent dengue vaccine are directed against highly conserved epitopes (Diamond MS (ed)). J Virol 89(1):120–128. https://doi.org/10.1128/JVI.02129-14
Weiskopf D, Bangs DJ, Sidney J, Kolla RV, de Silva AD, de Silva AM, Crotty S, Peters B, Sette A (2015) Dengue virus infection elicits highly polarized CX3CR1 +Cytotoxic CD4 +T cells associated with protective immunity. Proc Natl Acad Sci 112(31):E4256–E4263. https://doi.org/10.1073/pnas.1505956112
Wherry EJ, Ahmed R (2004) Memory CD8 T-cell differentiation during viral infection. J Virol 78(11):5535–5545. https://doi.org/10.1128/JVI.78.11.5535-5545.2004
Yauch LE, Zellweger RM, Kotturi MF, Qutubuddin A, Sidney J, Peters B, Prestwood TR, Sette A, Shresta S (2009) A protective role for dengue virus-specific CD8+ T cells. J Immunol 182(8):4865–4873. https://doi.org/10.4049/jimmunol.0801974
Yauch, Lauren E, Prestwood TR, May MM, Morar MM, Zellweger RM, Peters B, Sette A, Shresta S (2010) CD4+ T cells are not required for the induction of dengue virus-specific CD8+ T cell or antibody responses but contribute to protection after vaccination. J Immunol (Baltimore MD, 1950) 185(9):5405–5416. American Association of Immunologists. https://doi.org/10.4049/jimmunol.1001709.185
Acknowledgments
This work is supported by a Cooperative Basic Research Grant- New Investigator Grant (CBRG-NIG R-913-301-289-213) awarded by the Singapore National Medical Research Council to Laura Rivino.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Discussion of Chapter 17 in Dengue and Zika: Control and Antiviral Treatment Strategies
Discussion of Chapter 17 in Dengue and Zika: Control and Antiviral Treatment Strategies
This discussion was held at the 2nd Advanced Study Week on Emerging Viral Diseases at Praia do Tofo, Mozambique.Transcribed by Hilgenfeld R and Vasudevan SG (Eds); approved by Dr. Laura Rivino.
-
Anuja Mathew: So you found higher frequencies of CD8+ T cells in the skin. Do you have an hypothesis of why you would see a preference for CD8+ or CD4+ T cells?
-
Laura Rivino: We have preliminary data showing that the dengue-specific T cells that we find in the skin are mostly CD8+ T cells, so we think these cells could account in part for the increase of total CD8+ T cells that we find during acute dengue in the skin. We do not expect this influx of CD8+ T cells to comprise of only dengue-specific T cells. We know, for example from studies performed by us and others that in the blood of acute dengue patients there is also “bystander-activation” of CD8+ T cells that are specific for non-related, persistent viruses such as HCMV and EBV. So the influx of CD8+ T cells that occurs in the skin of acute dengue patients could comprise of both dengue-specific and “bystander-activated” T cells.
-
This higher frequncies of CD8+ T cells that we observe in the skin of acute dengue patients was not surprising to us since for other skin viral infections, such as Herpex simplex virus, the anti-viral response at the skin level is mainly a CD8+ T cell response.
-
Katja Fink: The cytokines that you find in the blister fluid are more or less the same as what is observed in the plasma or in the skin?
-
Laura Rivino: No. So the increase in pro-inflammtory cytokines is more pronounced in the skin compared to the blood. We have done quite a lot of cytokine profiling from the plasma of dengue patents and the increase in pro-inflammatory cytokines at the time points we look at are obvious only for the more severe dengue cases. We generally do not see significant increases of these cytokines in the blood, especially of TNF-α.
-
Katja Fink: So you compared skin and blood samples from the same patients?
-
Laura Rivino: Yes.
-
Aravinda da Silva: Were these cases primary infections?
-
Laura Rivino: Most of our patients are experiencing secondary infections. We had few primary infections so it’s difficult to really make a statement, but when I looked seperately at patients with primary and secondary infection I didn’t see a very big difference in terms of immunodominance of the different dengue proteins. The immunodominance was similar in primary and secondary cases and also for different dengue serotypes as well.
-
Aravinda da Silva: Right, but in terms of seeing the cells so early in the skin by day four of onset of symptoms, it seems very fast for a primary response. Do you see it that fast and in those numbers even in primary infection?
-
Laura Rivino: We don’t have the data yet of which patients were experiencing primary or secondary infection for the skin samples.
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Rivino, L. (2018). Understanding the Human T Cell Response to Dengue Virus. In: Hilgenfeld, R., Vasudevan, S. (eds) Dengue and Zika: Control and Antiviral Treatment Strategies. Advances in Experimental Medicine and Biology, vol 1062. Springer, Singapore. https://doi.org/10.1007/978-981-10-8727-1_17
Download citation
DOI: https://doi.org/10.1007/978-981-10-8727-1_17
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-8726-4
Online ISBN: 978-981-10-8727-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)