Abstract
Single-layer omics provide limited insight, whereas integrated multi-omics layers allow understanding of their combined influence on the complex biological process. The integrative omics approach has been initially applied to cancer research and later used in understanding host-pathogen interactions and pluripotency regulatory networks in stem cells. Here, different multi-omics layers along with databases and tools specific for multiple data integration, visualization, and integrated network modeling are described. In summary, this chapter focuses on integrative analysis of different multi-omics layers and modeling of interactomes to identify robust biomarkers and biological processes associated with diseases.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
3.1 Introduction
The initial multi-omics data was generated by The Cancer Genome Atlas (TCGA) project on different tumors and cancer cell lines. It provided a comprehensive genomics profiles including genetic mutations, gene expression, microRNA, copy number, and methylation data of 32 types of human tumors. This genomics dataset was possible due to the availability of next-generation sequencing (NGS) technology that provided the complete genome-wide coverage with low cost. After that, the Clinical Proteomic Tumor Analysis Consortium (CPTAC) used the same TCGA tumor samples and generated tandem mass spectrometry (MS/MS)-based proteomics data. All these multi-omics data from TCGA and CPTAC projects were analyzed and stored in LinkedOmics database (Vasaikar et al. 2018). Detailed proteogenomics analysis was performed in TCGA breast cancer samples, where functional consequences of somatic mutations were reported (Mertins et al. 2016). Large-scale protein-protein interactions of human and other model organisms were generated using affinity purification followed by mass spectrometry and yeast two-hybrid-based techniques (Ewing et al. 2007; Rual et al. 2005; Krogan et al. 2006; Uetz et al. 2000). Multi-omics data was not only restricted to cancer, but there were other applications of multi-omics integrative studies such as understanding host-pathogen interaction (Jean Beltran et al. 2017), host signaling regulation by the gut microbiota (Manes et al. 2017), and pluripotency regulatory network in embryonic stem cells (Stumpf et al. 2016).
There are several bioinformatics tools available for integrating, visualizing, and modeling multi-omics data and networks. Bayesian support vector machine and clustering methods have been used to integrate the data of mixed types (Yifeng et al. 2016). Cytoscape is an open-source software that can be used for visualizing the integrated networks (Cline et al. 2007). Network-based approaches used graph theory to integrate multiple homogeneous networks (e.g., protein-protein interaction), where node represents gene or protein and edge represents interaction. There can be two different types of interaction in heterogeneous networks (e.g., protein-protein, protein-DNA and DNA-metabolite interactions), one is the intraspecies interaction (protein-protein) and the other is the interspecies interaction (protein-DNA). The latter interaction is mainly involved in cross talk among multiple layers of the interactome. In summary, multi-omics approaches along with bioinformatics tools allow the integration of data generated from different omic levels and aid in understanding the complex and wired biological networks.
This chapter will first highlight different multi-omics layers and four different types of integrative analysis of multi-omics datasets, including (1) integrative analysis of genomics, epigenomics, and transcriptomics data; (2) integrative analysis of transcriptomics, proteomics, and protein interaction networks; (3) integrative analysis of transcriptomics and metabolomics; and (4) integrative analysis of multi-omics data. Next, the databases and tools used for multi-omics studies will be presented. And finally, the future perspectives and challenges of integrative omics studies will be discussed.
3.2 Multi-omics Layers
A single layer of “omics” including genomics, epigenomics, transcriptomics, proteomics, and metabolomics provides specific insight of DNA, RNA, protein, and metabolite level into the biological process of a cell. Genomics, involving the sequencing and analysis of genomes, uses high-throughput DNA sequencing such as next-generation sequencing (NGS), whole-genome sequencing (WGS), whole-exome sequencing, real-time PCR (RT-PCR), and single nucleotide polymorphism (SNP) along with bioinformatics to assemble and analyze the function and structure of the entire genomes (Concepts of genetics 2012; Culver and Labow 2002). Epigenomics, on the other hand, involves the study of reversible modifications on a cell’s DNA or histones that affect gene expression without altering the DNA sequence. The study of epigenetics on a global level has been made possible only recently through the adaptation of genomic high-throughput assays such as chromatin immunoprecipitation followed by microarray (ChIP-chip), chromatin immunoprecipitation followed by sequencing (ChIP-seq), methylated DNA immunoprecipitation (Me-DIP) (Friedman and Rando 2015), and ATAC-seq (Buenrostro et al. 2013). Transcriptomics refers to the study of the information content of an organism present in DNA, which includes mRNA and noncoding RNAs such as tRNA, rRNA, microRNA, and long ncRNA. The various RNA pools differ dramatically in abundance relative to each other and can change across experimental conditions (Yang et al. 2011). The standard protocol for transcriptome analysis involves RNA extraction, reverse transcription, cDNA amplification using quantitative reverse transcription-PCR (qRT-PCR), and hybridization using microarrays followed by library construction and sequencing (RNA-Seq). Proteomics refers to the large-scale analysis of the whole set of proteins which has significantly benefited from the Human Genome Project, accumulation of both DNA and protein sequence databases, improvements in mass spectrometry, and the development of computer algorithms for database searching (Graves and Haystead 2002). Metabolomics aims to measure the low molecular weight compounds called metabolites. The metabolome composition reflects the current status of the organism and is considered to be a chemical reflection of a molecular phenotype (Bujak et al. 2015). Numerous analytical platforms are commonly used in both targeted and untargeted metabolomic studies such as nuclear magnetic resonance (NMR) and mass spectrometry (MS), coupled with different separation techniques (Lindon and Nicholson 2008).
Multi-omics approaches integrate data from different omics levels to understand their combined influence on the biological process. For example, pluripotent stem cells show a high degree of regulation between multiple species of molecules. Studies have shown that the pluripotent state in mouse and human cells is regulated at multiple levels, including transcriptional (Boyer et al. 2005), epigenetic (Lee et al. 2006), signaling (Chen et al. 2008), and metabolic (Moussaieff et al. 2015) layers. Studies by Stumpf et al. shows that in the presence of external stimuli (Ying et al. 2008), the pluripotent state is maintained by a set of TFs, Oct4, Sox2, and Nanog along with secondary factors such as Klf4, Myc, and Lin28 (MacArthur et al. 2012). These core TFs interact with a range of auxiliary TFs via PPIs (Wang et al. 2006) and collectively control transcription of a large number of genes. Transcriptional control is exerted either directly, by binding to gene promoters (Boyer et al. 2005), or indirectly, by mediating the effects of epigenetic remodeling complexes (Orkin and Hochedlinger 2011). To add to this is a network of microRNAs (Wang et al. 2007) which ensures that appropriate protein levels are robustly maintained. Collectively, these reports indicate that pluripotency is regulated by cross talk among multi-omics layers to form interactome (Fig. 3.1) and involves layers of combinatorial regulatory control, including complex feedback relationships between the transcriptional, epigenetic, and signaling strata. Thus, the cross talk between multi-omics layers cannot be determined by single omics reduction approach.
Cross talk of the multi-omics layer to form interactome. Single omics approach like genomics, epigenomics, transcriptomics, proteomics, and metabolomics integrated by combining the interconnections of all layers within species and across species. The molecular species involved at each level is illustrated with nodes of different shapes and colors, and a key is provided below. The edges in dotted red lines show intermolecular species interaction, while the edges solid lines and color matched with nodes show intraspecies interaction, whereas solid arrows indicate external influence. (Partially adapted from Stumpf et al. Proteomics 2016)
3.2.1 Integrative Analysis of Genomics, Epigenomics, and Transcriptomics Data
The TCGA data provides comprehensive genomics profiles including genetic mutations, gene expression, microRNA sequencing, and copy number alterations of over 30 human tumors. Thus, the TCGA data is well studied for integrating multi-omics datasets. The effect of copy number alterations (CNA) on mRNA levels was studied in breast cancer samples, and it was seen that 64% of all genes studied have a positive correlation between CNA and mRNA levels (Mertins et al. 2016). In another study of integrative analysis in liver cancer, it was observed that cancer gene expression could be correlated with DNA copy number (CNVcor) and with DNA methylation (METcor) (Woo et al. 2017). Expression profiles of these CNVcor and METcor genes were able to predict subgroups in hepatocellular cancer. There are few bioinformatics tools available for integrating genomics, epigenomics, and transcriptomics datasets like DINGO, BioWardrobe, and mixOmics (Ha et al. 2015; Kartashov and Barski 2015; Rohart et al. 2017). These tools allow building differential networks and identifying common hub genes found in expression datasets of multiple layers.
3.2.2 Integrative Analysis of Transcriptomics, Proteomics, and Protein Interaction Networks
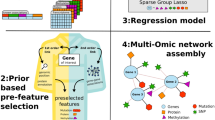
Integrating transcriptomics and proteomics data with protein interaction networks have been used for discovery of biomarkers and novel biological processes. In the field of biomarker discovery, the overlapping genes and proteins observed in multiple layers are common targets or a part of a feedback loop and so possibly better targets for therapeutics (Chakravorty et al. 2017). In a study by Mertins et al. from Broad Institute, results show a correlation between protein expression and gene expression across breast cancer samples taken from TCGA data (Mertins et al. 2016). These results demonstrate the utility of integrated transcriptome and proteome analysis for confirmation of regulatory mechanisms and identification of candidate regulators.
There is a higher coverage of transcriptome data as compared with mass spectrometry-based proteomics approach. Thus, gene expression datasets are merged with protein-protein interaction (PPI) network for the identification of novel biological process and active subnetworks as shown in Fig. 3.2. NetworkAnalyst and jActiveModules allow to merge gene expression and PPI networks. This approach has been studied for a better understanding of cancer and host-pathogen interactions (Jean Beltran et al. 2017; Saha and Ewing 2011).
Integrating transcriptomics and proteomics data to generate an integrative omics network. The gene expression profile from microarray data of disease versus control is combined with protein-protein interaction network to generate an integrative network. The red and blue color gradients represent overexpression and under-expression of differentially regulated genes. Yellow node indicates the proteins, and black lines indicate the edges of the PPI network, and thicknesses of the edges indicate the confidence of interaction. In the integrative network, the red to blue color gradients indicate the gene expression profile of the proteins involved in PPI. Green dotted circle highlights the subnetwork
3.2.3 Integrative Analysis of Transcriptomics and Metabolomics
Metabolomics is an important functional layer in studying multi-omics datasets, since it links genotype to phenotype. Integrative approaches for metabolomics and transcriptomics have been well established in the plant system (Urbanczyk-Wochniak et al. 2003). Datasets from metabolomics and transcriptomics studies are integrated using the correlation-based method, multivariate-based method that uses partial least square (PLS) regression and principal component analysis (PCA), and finally pathway-based method (Cavill et al. 2016). Integrated Molecular Pathway-Level Analysis (IMPaLA) is a web-based freely available tool frequently used for integration of two types of datasets (Kamburov et al. 2011). Other tools like Metscape 2 and Paintomics also perform similar kind of integrative analysis.
3.2.4 Integrative Analysis of Multi-omics Data
With the availability of TCGA and LinkedOmics resources, analyzing multi-omics dataset is possible. Various bioinformatics tools like Lemon-Tree and Omics Integrator allow network-based interpretation of multi-omics datasets (Bonnet et al. 2015). These are open-source, platform-independent and allow integrating multiple types of high-throughput datasets for creating networks.
3.3 Databases and Tools Used for Multi-omics Data
3.3.1 Database
Several databases contain multi-omics data as shown in Table 3.1. The first multi-omics database is The Cancer Genome Atlas (TCGA) that provides an interactive data system for researchers to search, download, upload, and analyze various cancer genomic datasets (Wang et al. 2016). The Library of Integrated Network-Based Cellular Signatures (LINCS) program provides an extensive reference library of cell-based perturbation-response signatures (Koleti et al. 2018). The LinkedOmics database includes information about mass spectrometry-based global proteomics data on TCGA tumor samples along with clinical data (Vasaikar et al. 2018). Multi-Omics Profiling Expression Database (MOPED) contains processed data for gene, protein, and pathway expression of human and model organism (Montague et al. 2015). Very few organ-specific diseases like heart and kidney diseases have multi-omics databases available (Alexandar et al. 2015; Fernandes and Husi 2017). Taken together, most of the integrative resources compiled various types of multi-omics datasets of tumors and cancer cell lines.
3.3.2 Tools
The availability of multi-omics cancer data from the same samples from TCGA allows developing various tools specific for multiple data integration, visualization, and integrated network modeling. The list of software dedicated for multi-omics data along with brief description is shown in Table 3.2. Tools like DINGO, BioWardrobe, and mixOmics are used for integrated analysis of mRNA/miRNA expression, DNA copy number, and methylation (Ha et al. 2015; Kartashov and Barski 2015; Rohart et al. 2017). Tools like jActiveModules (Cytoscape plugins) and NetworkAnalyst are used for integrating gene expression and PPI networks. Similarly, there are tools like Metscape 2 and Paintomics for integration of mRNA expression and metabolites data (Karnovsky et al. 2012; Garcia-Alcalde et al. 2011) and tools like ZikaVR and Immunet for Zika virus and immunological disease research, respectively (Gorenshteyn et al. 2015; Gupta et al. 2016). Omics Integrator software integrates several types of omics data and constructs a heterogeneous network of phosphorylated proteins, metabolites, and mRNA expression (Tuncbag et al. 2016). Lemon-Tree software uses large-scale multi-omics datasets and predicts network modules and pathways (Bonnet et al. 2015). In summary, there are several integrative analysis tools for multi-omics datasets and inferring network modules and pathways for understanding complex biological processes.
3.4 Future Prospective and Challenges
The primary requirement of the integrative multi-omics is that all the omics studies have to be performed in the same sample. So, there are few challenges in integrating multi-omics datasets. First, for integrating protein-protein interactions data, it was observed that most of the data available was from HEK 293 cell line in case of AP-MS studies. There is a considerable gap in generating PPIs of all the proteins from other human cell lines and tissues including healthy and diseased states. Second, for integrating metabolomics and transcriptomics data, it was seen that the metabolites are mainly isolated from blood or urine, while transcriptomics data can be derived from all tissue samples related to the disease. As there is a need for experimental sample source parity, there is also the need for establishing data processing standards and data normalization procedures across different omics layers. So far, most of the multi-omics studies are mainly focused on tumor and cancer cell lines. Besides cancer, there are various diseases like respiratory and cardiac diseases, which need urgent attention for understanding biological mechanisms of these diseases using integrative analysis of multi-omics data.
References
Alexandar V, Nayar PG, Murugesan R, Mary B, Darshana P et al (2015) CardioGenBase: a literature based multi-omics database for major cardiovascular diseases. PLoS One 10:e0143188
Bonnet E, Calzone L, Michoel T (2015) Integrative multi-omics module network inference with Lemon-Tree. PLoS Comput Biol 11:e1003983
Boyer LA, Lee TI, Cole MF, Johnstone SE, Levine SS et al (2005) Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122:947–956
Buenrostro JD, Giresi PG, Zaba LC, Chang HY, Greenleaf WJ (2013) Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat Methods 10:1213–1218
Bujak R, Struck-Lewicka W, Markuszewski MJ, Kaliszan R (2015) Metabolomics for laboratory diagnostics. J Pharm Biomed Anal 113:108–120
Cavill R, Jennen D, Kleinjans J, Briede JJ (2016) Transcriptomic and metabolomic data integration. Brief Bioinform 17:891–901
Chakravorty D, Jana T, Das Mandal S, Seth A, Bhattacharya A et al (2017) MYCbase: a database of functional sites and biochemical properties of Myc in both normal and cancer cells. BMC Bioinformatics 18:224
Chen X, Xu H, Yuan P, Fang F, Huss M et al (2008) Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133:1106–1117
Cline MS, Smoot M, Cerami E, Kuchinsky A, Landys N et al (2007) Integration of biological networks and gene expression data using Cytoscape. Nat Protoc 2:2366–2382
Concepts of genetics (2012) (10th ed.). San Francisco: Pearson Education. ISBN:978-0-321-72412-0
Culver KW, Labow MA (2002) Genomics. In: Robinson R (ed) Genetics. Macmillan Science Library. Macmillan Reference USA, New York ISBN:978-0-02-865606-9
Ewing RM, Chu P, Elisma F, Li H, Taylor P et al (2007) Large-scale mapping of human protein-protein interactions by mass spectrometry. Mol Syst Biol 3:89
Fernandes M, Husi H (2017) Establishment of a integrative multi-omics expression database CKDdb in the context of chronic kidney disease (CKD). Sci Rep 7:40367
Friedman N, Rando OJ (2015) Epigenomics and the structure of the living genome. Genome Res 25:1482–1490
Garcia-Alcalde F, Garcia-Lopez F, Dopazo J, Conesa A (2011) Paintomics: a web based tool for the joint visualization of transcriptomics and metabolomics data. Bioinformatics 27:137–139
Gorenshteyn D, Zaslavsky E, Fribourg M, Park CY, Wong AK et al (2015) Interactive big data resource to elucidate human immune pathways and diseases. Immunity 43:605–614
Graves PR, Haystead TA (2002) Molecular biologist’s guide to proteomics. Microbiol Mol Biol Rev 66:39–63
Gupta AK, Kaur K, Rajput A, Dhanda SK, Sehgal M et al (2016) ZikaVR: an integrated Zika virus resource for genomics, proteomics, Phylogenetic and Therapeutic Analysis. Sci Rep 6:32713
Ha MJ, Baladandayuthapani V, Do KA (2015) DINGO: differential network analysis in genomics. Bioinformatics 31:3413–3420
Jean Beltran PM, Federspiel JD, Sheng X, Cristea IM (2017) Proteomics and integrative omic approaches for understanding host-pathogen interactions and infectious diseases. Mol Syst Biol 13:922
Kamburov A, Cavill R, Ebbels TM, Herwig R, Keun HC (2011) Integrated pathway-level analysis of transcriptomics and metabolomics data with IMPaLA. Bioinformatics 27:2917–2918
Karnovsky A, Weymouth T, Hull T, Tarcea VG, Scardoni G et al (2012) Metscape 2 bioinformatics tool for the analysis and visualization of metabolomics and gene expression data. Bioinformatics 28:373–380
Kartashov AV, Barski A (2015) BioWardrobe: an integrated platform for analysis of epigenomics and transcriptomics data. Genome Biol 16:158
Koleti A, Terryn R, Stathias V, Chung C, Cooper DJ et al (2018) Data portal for the library of integrated network-based cellular signatures (LINCS) program: integrated access to diverse large-scale cellular perturbation response data. Nucleic Acids Res 46:D558–D566
Krogan NJ, Cagney G, Yu H, Zhong G, Guo X et al (2006) Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440:637–643
Lee TI, Jenner RG, Boyer LA, Guenther MG, Levine SS et al (2006) Control of developmental regulators by Polycomb in human embryonic stem cells. Cell 125:301–313
Lindon JC, Nicholson JK (2008) Analytical technologies for metabonomics and metabolomics, and multi-omic information recovery. Trac-Trend Anal Chem 27:194–204
MacArthur BD, Sevilla A, Lenz M, Muller FJ, Schuldt BM et al (2012) Nanog-dependent feedback loops regulate murine embryonic stem cell heterogeneity. Nat Cell Biol 14:1139–1147
Manes NP, Shulzhenko N, Nuccio AG, Azeem S, Morgun A et al (2017) Multi-omics comparative analysis reveals multiple layers of host signaling pathway regulation by the gut microbiota. mSystems 2:e00107
Mertins P, Mani DR, Ruggles KV, Gillette MA, Clauser KR et al (2016) Proteogenomics connects somatic mutations to signalling in breast cancer. Nature 534:55–62
Montague E, Janko I, Stanberry L, Lee E, Choiniere J et al (2015) Beyond protein expression, MOPED goes multi-omics. Nucleic Acids Res 43:D1145–D1151
Moussaieff A, Rouleau M, Kitsberg D, Cohen M, Levy G et al (2015) Glycolysis-mediated changes in acetyl-CoA and histone acetylation control the early differentiation of embryonic stem cells. Cell Metab 21:392–402
Orkin SH, Hochedlinger K (2011) Chromatin connections to pluripotency and cellular reprogramming. Cell 145:835–850
Rohart F, Gautier B, Singh A, Le Cao KA (2017) mixOmics: an R package for ‘omics feature selection and multiple data integration. PLoS Comput Biol 13:e1005752
Rual JF, Venkatesan K, Hao T, Hirozane-Kishikawa T, Dricot A et al (2005) Towards a proteome-scale map of the human protein-protein interaction network. Nature 437:1173–1178
Saha S, Ewing R (2011) Systemic discovery of condition specific Wnt signalling subnetworks. In: IEEE international conference on bioinformatics and biomedicine workshops
Stumpf PS, Ewing R, MacArthur BD (2016) Single-cell pluripotency regulatory networks. Proteomics 16:2303–2312
Tuncbag N, Gosline SJ, Kedaigle A, Soltis AR, Gitter A et al (2016) Network-based interpretation of diverse high-throughput datasets through the omics integrator software package. PLoS Comput Biol 12:e1004879
Uetz P, Giot L, Cagney G, Mansfield TA, Judson RS et al (2000) A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature 403:623–627
Urbanczyk-Wochniak E, Luedemann A, Kopka J, Selbig J, Roessner-Tunali U et al (2003) Parallel analysis of transcript and metabolic profiles: a new approach in systems biology. EMBO Rep 4:989–993
Vasaikar SV, Straub P, Wang J, Zhang B (2018) LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res 4:D956–D963
Wang J, Rao S, Chu J, Shen X, Levasseur DN et al (2006) A protein interaction network for pluripotency of embryonic stem cells. Nature 444:364–368
Wang Y, Medvid R, Melton C, Jaenisch R, Blelloch R (2007) DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nat Genet 39:380–385
Wang Z, Jensen MA, Zenklusen JC (2016) A practical guide to the Cancer genome atlas (TCGA). Methods Mol Biol 1418:111–141
Woo HG, Choi JH, Yoon S, Jee BA, Cho EJ et al (2017) Integrative analysis of genomic and epigenomic regulation of the transcriptome in liver cancer. Nat Commun 8:839
Xia J, Gill EE, Hancock RE (2015) NetworkAnalyst for statistical, visual and network-based meta-analysis of gene expression data. Nat Protoc 10:823–844
Yang L, Duff MO, Graveley BR, Carmichael GG, Chen LL (2011) Genomewide characterization of non-polyadenylated RNAs. Genome Biol 12:R16
Yifeng L, Fang-Xiang W, Alioune N (2016) A review on machine learning principles for multi-view biological data integration. Brief Bioinform 1–16
Ying QL, Wray J, Nichols J, Batlle-Morera L, Doble B et al (2008) The ground state of embryonic stem cell self-renewal. Nature 453:519–523
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Chakravorty, D., Banerjee, K., Saha, S. (2018). Integrative Omics for Interactomes. In: Singh, S. (eds) Synthetic Biology. Springer, Singapore. https://doi.org/10.1007/978-981-10-8693-9_3
Download citation
DOI: https://doi.org/10.1007/978-981-10-8693-9_3
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-8692-2
Online ISBN: 978-981-10-8693-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)