Abstract
The term microbiome refers to the existence of multiple microbial genomes present in an environment in an association with a host. With the development of more precise sequencing approaches, identification of genus and families that were uncultivable microbes has been made possible. The current chapter explores the importance of understanding microbial communities and their association with agricultural production systems with particular attention to endophytic microorganisms. Agri-management practices and their relationship to the selection of microbial variation of taxa by plants and soil have been discussed in detail. The article also discusses how farming practices such as cover cropping and mulching mediate microbial community dynamics. Future perspectives on advancing sustainability by microbiome optimization are discussed.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
5.1 Introduction: Evolving Concepts of the Plant Microbiome
5.1.1 General
The soil is a complex environment where there is a vast mix of organic matter, minerals, nutrients, and gases, among others, enclosing a myriad of organisms – micro and macro – that are capable of supporting and retarding plant life and growth. The heterogeneity that exists in these environments is controlled by a series of biological and ecological interactions combined by soil properties, which allow for the proliferation and establishment of certain groups of microbial organism, changing the dynamics of the ecosystem (Gale et al. 2000).
The importance of understanding microbial communities and their association with agricultural production systems lies on the premises of a future with more sustainable approaches to challenges in agriculture. Although many efforts have been directed toward a better understanding on how these microbial communities work, there are still a great number of questions related to the most influential factors dictating the identity or core participants, the diversity and niche specificity, the establishment and maintenance of association with plants, and retrograde signaling networks that could functionalize associations.
5.1.2 Looking Deeper into the Plant Microbiome Using Developing Technologies
The term microbiome refers more to the existence of multiple microbial genomes present in an environment in an association with a host. For the purpose of this chapter, we are focused on the plant bacterial microbiome in an agricultural context. The soil microbial community has received an abundance of attention over past decades, but the broader plant microbiome includes organisms that dwell in the phyllosphere, inside the plant as endophytic organisms, as well as in the rhizosphere and soil. Bacterial organisms are classified as endophytic if they inhabit plant tissue during its life cycle. In contrast, some rhizospheric bacteria colonize plants as opportunistic organisms that interact at some point with the plant but don’t inhabit it in an obligate manner. An interest in endophytes, particularly obligate endophytes and the benefits they are able to confer to plants, and how some of these changes may be transferred genetically has emerged recently.
Recent advances in next-generation sequencing (NGS) technology have advanced our understanding of this community (Lundberg et al. 2012; Bulgarelli et al. 2012; Wagner et al. 2016). In terms of the plant microbiome and its relationship to agricultural production, studies have proved that the presence of certain groups of organisms is capable of processing and absorbing nutrients (Manzoni et al. 2008) rendering them available for plant growth (Schardl et al. 2004; Barrow et al. 2008; Xia et al. 2013), repression of disease, and the capacity to mediate the impact of extreme environmental stress factors (Plett and Martin 2011). What remains complicated is how to foment the presence of those beneficial groups and how they could be used for improvement of many important agronomical crops. Indeed, it will important to establish how soil conditions and agronomical practices affect the selection of these microbial organisms by the plant. Technologies such as NGS accompanied by fluorescence in situ hybridization (CARD-FISH) for specific microbiome components have broadened what we can identify and how we associate them with the host plant. Agri-management practices and their relationship to the selection for variation of taxa by plants and soil are the main reasons for the development of this chapter. We will be looking at how managing practices could be important when trying to understand the strengths or weaknesses of these relationships, since they are able to influence the development and dominance of a bacterial community.
5.2 The Microbiome and Agriculture
The interaction between plants and individual microorganisms has been studied for the last several decades. Isolation and testing of strains present in soil and plants have largely aimed to understand the capacity that these microorganisms have for plant improvement or pathogenicity. Until the last 5 years, most of the isolation and identification was done via culture-dependent techniques. However, with the development of more precise sequencing approaches, identification of genus and families that were unculturable has been made possible, even to the point of looking at functional genes (Tsurumaru et al. 2015). These advances have provided more insight into the selection and structure of bacterial communities by plants under different environments (Lundberg et al. 2012, 2013; Lebeis et al. 2015; Birtel et al. 2015; Ding and Melcher 2016). Identifying the variability as well as functionality of communities that colonize plants could be used to select for bacteria (or groups of bacterial community members) that can positively modify the plant morphology or interaction with its environment. Despite the attractiveness of being able to inject a single or collection of microorganisms into an agricultural production system to enhance crop performance, there are many reasons that this will be challenging in practice. The complexity of the microbial community and competitiveness of a single microbial factor are unlikely to be dominant enough to sustain any influence on a cropping system. Furthermore, the ability to genetically optimize or engineer microbes to enhance agricultural systems will be a regulatory and environmental containment challenge. As related to agricultural production systems, the notion that understanding the plant microbiome and how it functions and then adapting our management practices to maximize the most interesting members of the microbiome is perhaps the most rational area for future work. Furthermore, plant breeding has not taken into account any influence of a microbiome, and it remains possible that the intersection between plant breeding and microbiome functionality will be a fruitful area for research (Gopal and Gupta 2016). Finally, knowledge of the mechanisms by which a microbiome element influences the plant anatomy is still developing and should shed light on hormonal networks and functional gene networks influenced by the microbiome.
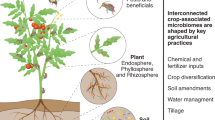
How a bacterial microbiome colonizes and establishes itself in living plant tissue will involve not just the physical entry into the plant but also how to avoid the plant immune system (friend versus foe association) (Downie and Walker 1999; Iniguez et al. 2005). As the field of microbial inoculums matures, it will be important to understand the complexity of this association window and whether it is under passive or active control by the host plant. It is expected that numerous non-obligate bacterial genera enter the plant during germination and seedling establishment. As the main contact point for the plant with the microbe-rich soil, microbes are thought to enter into their host (plant) through the root system due to their vast adhering area with soil particles (Hansen et al. 1997; Tokala et al. 2002; Iniguez et al. 2005; Rosenblueth and Martinez-Romero 2006; Seipke et al. 2012) (Fig. 5.1). The rhizosphere is the area that is described as the zone of the soil that is subjected to the influence of the roots. At the same time, another term that will be highly important to mention while talking about entrance of microbial organism to the plant is the spermosphere. This is related to the seed exterior layers that are in contact with the soil and over which microbes will be interacting before germination.
Schematic presentation of the microbiome. Image of a broadleaf seedling planted (left) and conceptualizing the overlay of management practices. As the seedling grows, bacterial community members from the soil, which are represented as orange, blue, purple, and red dots, occupy various components of the root (red arrow) and phyllosphere (green components of plant aerial tissue)
5.3 Inspection Between Agricultural Management Practices and Microbiome
It seems that through the use of culturing and next-generation sequencing, there have been signs that point toward a more consistent and numerous amounts of organisms being identified as endophytic microbiome elements in organic production systems when compared to conventional farming practices (Xia et al. 2015; Schlaeppi and Bulgarelli 2015; Hartmann et al. 2015). The reasons behind those differences among bacterial communities still remain slightly unclear, but data supporting increased soil microbial diversity in organically managed soils have been well documented (Wang et al. 2016). More work has been put toward the elucidation of the effect that the systems may have on the selection of the taxa present in the soil. These results supported findings by Soltani et al. (2010) and Bacon and White (2016) that many endophytic bacterial genotypes increased plant growth and induced a defense system with low cost.
As mentioned before, the differences found among isolates identified as endophytic microbial species comparing conventional and organic crops are of interest as they may be linked to if they can link crop productivity, since one of the main goals is to be able to replicate these environments for crop enhancements or at least to influence selection by plants toward some of these communities. Hall and Davis (1990) suggest that certain bacilli move through the plant using the vascular system rather than symplastic movement. Base on physiological aspects, the older the plant may be, the harder will be for certain endophytic bacteria to translocate from tissue type to tissue, and therefore it is anticipated that as we develop a more sophisticated understanding of tissue-type endophyte colonization, we may see different levels of abundance or community members. Some research supports that the age of the plant may not be one of the limitations for the colonization of obligated bacteria when tissue type was held consistent (roots) (Lundberg et al. 2012). This could be due to the fact that some of these endophytes may be present at early stages and stay there and that the variation of the presence or absence of other species may be related to those that are not strictly necessary to inhabit the plant. Interestingly, it was found by Lundberg et al. (2012) that genotype was a critical determinant in root microbiome community analysis suggesting that the intersection between breeding and agricultural farming practices may be critical for future work.
An interesting concept to examine is how farming practices and the types of crops that are being produced display variance in microbial community metrics. For instance, cover cropping, mulching and soil composition (Kumar et al. 2014), the use of alternative tillage systems (Carbonetto et al. 2014), and overall soil nutrient composition (Stagnari et al. 2014) have an impact in the structure and composition of the soil microbial communities. Carbonetto and co-workers (2014) suggested that soils exposed to high use of fertilizers displayed a shift in the metabolic strategies used by the microbial communities which exasperated community shifts. Metabolism seems to also become more “flexible” for those organisms that were present under tillage practices vs those in non-till areas, but the metabolic flexibility does not mean that they were better adapted; on the contrary, they showed that if conditions were considered unfavorable (e.g., lower nutrient content in soil), some of those microbial organisms are unlike to adapt, which differed from the non-tillage system. Similar results were found in cotton crops that were maintained under conventional tillage and no tillage (Feng et al. 2003). It seems like the use of non-tillage, for example, and not so many applications of fertilizers, among other things, can have a positive effect in microbial communities in the soil. Kennedy and Smith (1995) support that heavy tillage as a farming practice can be negative for microbial diversity and abundance by the alteration of the properties of the soil. Overall, high population and biodiversity of microorganisms in the soil is an indicator of soil health. Healthy soil has a normal amount of aggregation and percent of air, water, and nutrients; thus, the soil does not need many fertilizers or pesticides to increase plant productivity or to control stresses as the plant will be tolerant (Paul 2007). This parlays with good farming practices, not necessarily organic versus conventional practices.
Both practices, organic and conventional, have systems that follow the application of chemicals to treat and maintain their crops during their production process. Some of the chemicals used tend to be more long lasting within the farming system than others and could have small but progressive impacts on an indigenous microbial community present in the soil. Thus, when comparing results in this area, one must consider numerous environment and cultural factors that vary greatly and are different to compare. A question remains whether the use of pesticides affects microbial communities in the soil in a nontarget manner and in turn influences the selection of the plant microbiome. Even though pesticides are made to target insects and other types of organism that have no relationship with the fungi or bacteria present in soil, it is feasible that in a more individual scale, some species in particular may be affected (Foley et al. 2005). To date, further research is needed on a case-by-case basis to interrogate this postulate.
Herbicides or the surfactants used in their application to a target crop may also have an impact in the microbial communities since some of these, for instance, octylamines, can be slightly bacteriotoxic (https://www.echa.europa.eu/sv/web/guest/registration-dossier/-/registered-dossier/1996/7/7/2) but are nontarget and have been unstudied as environmental risk factors in agricultural microbiome systems. Other herbicidal or pesticidal molecules will remain in the soil (predominantly in conventional systems) for years, for example, the preemergent herbicide used on railroad lines indaziflam (Brabham and Debolt 2013) has an extremely long residual time. While off-target influences of commercially available pesticides and herbicides are typically nonlethal and modest, if a product can be mildly class specific bacteriotoxic, it can easily be envisioned how this could shift the balance in an agricultural crop microbiome (Wilkinson and Lucas 1969). To date, we have an unsatisfactory understanding of this process and whether subtle influences could even alter a microbiome in an agricultural setting.
It is important to take in consideration that longtime exposure to a specific managing practice could alter the soil environment by a simple selection mechanism. It seems that although change is part of both systems, organic farming may be a better option to also increase richness, among others, by shifting the structure of the microbiota compared to conventional practices (Hartmann et al. 2015). Still, more parameters and variables need to be tested to fully confirm these hypotheses and address better the full impact that these practices have on the microbial communities’ structure (Hartmann and Widmer 2006).
5.4 Employing Microbial Elements in Agricultural Systems
It is known that obligated microbes have to follow usually a more elaborated process for their colonization. They can be considered pathways, which usually ramify into production of exudates, rates of production of them, quorum sensing, and hormone metabolisms among others. Exudates are considered to be molecules produced and released either by the plant or bacteria to the rhizosphere (Li et al. 2016 PNAS). Some of the molecules present in these exudates are combination of sugars, amino acids, alkaloids, flavonoids, and others (Biedrzycki et al. 2010; Kumar and Bais 2012). Rates of the exudate production can also have an impact on how the plant selects the microbes from the rhizosphere. Now, the fact that some microbes are capable to produce their own chemicals and modulate the communication with the plant through molecule signaling, it is probably one of the future uses of studying the microbiomes of different systems. Indeed, some endophytic microbiome elements have been used to identify target herbicides in plants (Xia et al. 2014). The idea will be to find ways into isolating, producing, or stimulating the production of these chemicals for the manipulation of the selection power of the plant and at least inhabit it for a small time frame (or long, depending on the effect that it has in the host development and health). It may be suitable to bypass the microbial soil feature and grow it in vitro to harvest the target chemical for organic farming purposes, which is already the case for Bacillus thuringiensis.
Promoting plant growth by manipulating microbiomes may have a modest capacity to support the positive traits in a cropping species, thus decreasing the use of synthetic chemicals or nutrients (Singh et al. 2010). Using microbes in agriculture as bio-fertilizers and biopesticides has been well established, but lately it has received more attention, and scientists are currently focusing on the plant microbiome itself instead of just using microbes (Deakeret al. 2004). Using microbes is less practical than using synthetic chemicals because variation in soil and environmental conditions will almost certainly be a selection force and will therefore require regional solutions in agriculture. Modern agriculture has not accepted regionality of trait solutions from major crop biotechnology companies, and therefore it is unclear whether microbial systems will be poorly accepted. Organic farmers may be more willing to work with such regional/environment-specific products simply due to scale (Bacon and White 2016).
There are select studies that show that application of bacterial isolates could support plant growth and productivity under specific conditions, possibly modulating plant microbiomes (Xia et al. 2015). However, these rarely translate from greenhouse or in vitro conditions to the field and even more rarely into a wide variety of agricultural eco-zones. The plant growth-promoting fungal inoculum Trichoderma sp. is still the best example of a successful strategy for this (Altmore et al. 1999). It is hoped that the use of beneficial microbes in organic production system could buffer plant productivity by providing nutrients and other growth-promoting compounds to the crop not only for a short time but also for many seasons because this organic system maintains soil fertility and health.
Treatments and inoculation with bacterial organisms showed in Xia et al. (2014) that plant cell walls are susceptible to the colonization and production of certain chemicals (exudates) by the bacteria. This is a good growth indicator for studies of interactions between plant and microbes because of the importance of the plant cell wall, since it plays an essential role in being a barrier against stresses, connecting extracellular and intracellular environments, and regulating plant growth. Their work also showed that the combination of techniques for identification and isolation of organisms was crucial for a proper selection of candidate strains and their capacity of inhabiting the plant during long periods of its life. Even though manipulating the microbiome is important to increase plant productivity, it is currently a challenge to adopt bacterial strains grown in a lab environment and implement their use in the farmers’ fields. These artificially cultured “strains may lack key characteristics for widespread distribution in sustainable and productive agricultural systems” (Parnell et al. 2016). Most of the studies related to bacterial strains as an alternative to synthetic chemicals represent either lab or greenhouse experiments (Adesemoye et al. 2009), making the results obtained from these approaches not an accurate representation of the real environment that plants may be exposed to in a farm setting (Parnell et al. 2016). Although microbial organisms have potential for changing agriculture, there are still a lot of questions that will need to be answered before their acceptance.
5.5 Conclusion
The overall outcome of studies into the functionality of the plant microbiome has been satisfactory to maintain research and agricultural interest. The compelling idea of establishing a more sustainable production system through increasing the abundance or functionality of members of a natural community is highly attractive and potentially cost-effective. Several conclusions and future directions exist. A combined focus on plant breeding in association with detailed microbiome assessment is needed based on the genotype specificity identified in recent studies (Lundberg et al. 2012). Organic farming systems are modestly less likely to drive force and selection on the microbiome community due their inherent focus on soil quality rather than external inputs. Because genotype and environmental conditions both influence the microbiome in plants, long-term studies are needed across numerous species and eco-zones to adequately assess results.
References
Adesemoye AO, Torbert HA, Kloepper JW (2009) Plant growth-promoting rhizobacteria allow reduced application rates of chemical fertilizers. Microb Ecol 58(4):921–929
Altmore C, Norvell WA, Bjorkman T, Harman GE (1999) Solubilization of phosphates and micronutrients by the plant growth promoting and biocontrol fungus Trichoderma harzianum Rifai. Appl Environ Microbiol 65:2926–2933
Bacon CW, White JF (2016) Functions, mechanisms and regulation of endophytic and epiphytic microbial communities of plants. Symbiosis 68(1):87–98
Barrow JR, Lucero ME, Reyes-Vera I, Havstad KM (2008) Do symbiotic microbes have a role in plant evolution, performance and response to stress? Commun Integr Biol 1:69–73
Biedrzycki ML, Jilany TA, Dudley SA, Bais HP (2010) Root exudates mediate kin recognition in plants. Commun Integr Biol 3:28–35
Birtel J, Walser J-C, Pichon S, Bürgmann H, Matthews B (2015) Estimating bacterial diversity for ecological studies: methods, metrics, and assumptions. PLoS One 10(4):e0125356. doi:10.1371/journal.pone.0125356
Brabham C, Debolt S (2013) Chemical genetics to probe the cell wall. Front Plant Biotechnol 3:309
Bulgarelli D, Rott M, Schlaeppi K, Ver Loren, van Themaat E et al (2012) Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488:91–95
Carbonetto B, Rascovan N, Álvarez R, Mentaberry A, Vázquez MP (2014) Structure, composition and metagenomic profile of soil microbiomes associated to agricultural land use and tillage systems in Argentine Pampas. PLoS One 9:1–11
Deaker R, Roughley RJ, Kennedy IR (2004) Legume seed inoculation technology – a review. Soil Biol Biochem 36(8):1275–1288
Ding T, Melcher U (2016) Influences of plant species, season and location on leaf endophytic bacterial communities of non-cultivated plants. PLoS One 11(3):e0150895. doi:10.1371/journal.pone.0150895
Downie JA, Walker SA (1999) Plant responses to nodulation factors. Curr Opin Plant Biol 2:483–489
Feng Y, Motta AC, Reeves DW, Burmester CH, Van Santen E, Osborne JA (2003) Soil microbial communities under conventional till and no-till continuous cotton systems. Soil Biol Biochem 35:1693–1703
Foley JA, DeFries R, Asner GP, Barford C, Bonan G, Carpenter SR et al (2005) Global consequences of land use. Science 309:570–574
Gale WJ, Cambardella CA, Bailey TB (2000) Surface residue and root-derived carbon in stable and unstable aggregates. Soil Sci Soc Am J 64:196–201
Gopal M, Gupta A (2016) Microbiome selection could spur next-generation plant breeding strategies. Front Microbiol 7:1971–1977
Hall TJ, Davis WEE (1990) Survival of Bacillus subtilis in silver sugar maple seedlings over a two-year period. Plant Dis 74:608–609
Hansen ML, Kregelund L, Nybroe O, Sorensen J (1997) Early colonization of barley roots by Pseudomonas fluorescens studied by immunofluorescence technique and confocal laser scanning microscopy. FEMS Microbiol Ecol 23:353e360
Hartmann M, Widmer F (2006) Community structure analyses are more sensitive to differences in soil bacterial communities than anonymous diversity indices. Appl Environ Microbiol 72:7804–7812
Hartmann M, Frey B, Mayer J, Mader P, Widmer F (2015) Distinct soil microbial diversity under long-term organic and conventional farming. ISME J 9:1177–1194
Iniguez LA, Dong Y, Carter HD, Ahmer BMM, Stone JM, Triplett E (2005) Regulation of enteric endophytic bacterial colonization by plant defenses. Mol Plant-Microbe Interact 18:169–178
Kennedy A, Smith K (1995) Soil microbial diversity and the sustainability of agricultural soils. Plant Soil 170:75–86
Kumar AS, Bais HP (2012) Wired to the roots: impact of root beneficial microbe interactions on the above ground plant physiology and protection. Plant Sign Behav 72:694–706
Kumar A, Maurya BR, Raghuwanshi R (2014) Isolation and characterization of PGPR and their effect on growth, yield and nutrient content in wheat (Triticum aestivum L.) Biocatal Agric Biotechnol 3:121–128
Lebeis SL, Paredes SH, Lundberg DS, Breakfield N, Gehring J, McDonald M, Malfatti S, Glavina del Rio T, Jones CD, Tringe SG, Dangl JL (2015) Plant microbiome. Salicylic acid modulates colonization of the root microbiome by specific bacterial taxa. Science 349:860–864
Li B, Lia Y-Y, Wua H-M, Zhanga F-F, Lia C-J, Lia X-X, Lambersb H, Long L (2016) Root exudates drive interspecific facilitation by enhancing nodulation and N2 fixation. PNAS 113:236496–236501
Lundberg D et al (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488:86–90
Lundberg DS, Yourstone S, Mieczkowski P, Jones CD, Dangl DL (2013) Practical innovations for high-throughput amplicon sequencing. Nat Methods 10:999–1002
Manzoni S, Jackson RB, Trofymow JA, Porporato A (2008) The global stoichiometry of litter nitrogen mineralization. Science 321:684–686
Parnell JJ, Berka R, Young HA, Sturino JM, Kang Y, Barnhart DM, DiLeo MV (2016) From the lab to the farm: an industrial perspective of plant beneficial microorganisms. Frontiers Plant Sci 7:1110. doi:10.3389/fpls.2016.01110
Paul EA (2007) Soil microbiology, ecology, and biochemistry in perspective. Soil microbiology, ecology and biochemistry, 3rd edn. Academic, San Diego, pp 3–24
Plett JM, Martin F (2011) Blurred boundaries: lifestyle lessons from ectomycorrhizal fungal genomes. Trends Genet 27:14–22
Rosenblueth M, Martinez-Romero E (2006) Bacterial endophytes and their interactions with hosts. Mol Plant-Microbe Interact 19:827–837
Schardl CL, Leuchtmann A, Spiering MJ (2004) Symbioses of grasses with seed borne fungal endophytes. Annu Rev Plant Biol 55:315–340
Schlaeppi K, Bulgarelli D (2015) The plant microbiome at work. Mol Plant-Microbe Interact 28:212–217
Seipke RF, Kaltenpoth M, Hutchings MI (2012) Streptomyces as symbionts: an emerging and widespread theme? FEMS Microbiol Rev 36:862–876
Singh BK, Bardgett RD, Smith P, Reay DS (2010) Microorganisms and climate change: terrestrial feedbacks and mitigation options. Nat Rev Micro 8:779–790
Soltani AA, Khavazi K, Asadi-Rahmani H, Omidvari M, Abaszadeh DP, Mirhoseyni H (2010) Plant growth promoting characteristics in some Flavobacterium spp. isolated from soils of Iran. J Agr Sci 4:106–115
Stagnari F, Perpetuini G, Tofalo R, Campanelli G, Leteo F, Della Vella U, Schirone M, Suzzi G, Pisante M (2014) Long-term impact of farm management and crops on soil microorganisms assessed by combined DGGE and PLFA analyses. Front Microbiol 5:644
Tokala R, Strap JL, Jung CM, Crawford DL, Salove MH, Deobald LA, Bailey FJ, Morra MJ (2002) Novel plant-microbe rhizosphere interaction involving Streptomyces lydicus WYEC108 and the pea plant (Pisum sativum). Appl Environ Microbiol 68:2161–2171
Tsurumaru H et al (2015) Metagenomic analysis of the bacterial community associated with the taproot of sugar beet. Microbes Environ 30:63–69
Wagner MR, Lundberg DS, del Rio T, Tringe SG, Dangl JF, Mitchell-Olds T (2016) Host genotype and age shape the leaf and root microbiomes of a wild perennial plant. Nat Comm 7:12151
Wang W, Wang H, Feng Y, Wang L, Xiao X, Xi Y, Luo X, Sun R, Ye X, Huang Y, Zhang Z, Cui Z (2016) Consistent responses of the microbial community structure to organic farming along the middle and lower reaches of the Yangtze River. Scientific Reports 6: Article number 35046
Wilkinson V, Lucas RL (1969) Effects of herbicides on the growth of soil fungi. New Phytol 68:709–719
Xia Y, Greissworth E, Mucci C, Williams MA, DeBolt S (2013) Characterization of culturable bacterial endophytes of switchgrass (Panicum virgatum L.) and their capacity to influence plant growth. GCB Bioenergy 5:674–682
Xia Y, Petti C, Williams MA, DeBolt S (2014) Experimental approaches to study plant cell walls during plant-microbe interactions. Front Plant Sci 5:540
Xia Y, Debolt S, Dreyer D, Scott D, Williams M (2015) Characterization of culturable bacterial endophytes and their capacity to promote plant growth from plants grown using organic or conventional practices. Front Plant Sci 6. doi:10.3389/fpls.2015.00490
Acknowledgment
Work was supported by the National Science Foundation under Cooperative Agreement No. 1355438 and IOS-1256029. US Department of Agriculture Hatch funding and Altria Graduate Research Fellowship to ASB also supported this work.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Sanchez-Barrios, A., Sahib, M.R., DeBolt, S. (2017). “I’ve Got the Magic in Me”: The Microbiome of Conventional vs Organic Production Systems. In: Singh, D., Singh, H., Prabha, R. (eds) Plant-Microbe Interactions in Agro-Ecological Perspectives. Springer, Singapore. https://doi.org/10.1007/978-981-10-5813-4_5
Download citation
DOI: https://doi.org/10.1007/978-981-10-5813-4_5
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-5812-7
Online ISBN: 978-981-10-5813-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)